Abstract
In vivo astrocyte-to-neuron (AtN) conversion suffers low efficiency due to pre-existing intrinsic barriers. However, it is unclear whether astrocytes have inducible barriers to reprogramming. Here, we identify Olig2, a basic helix-loop-helix (bHLH) transcription factor, as an inducible barrier to Ngn2-mediated AtN conversion. Olig2 is strongly upregulated in cortical astrocytes following the ectopic expression of bHLH neuronal reprogramming factors such as Ngn2, NeuroD1, and Ascl1, but is barely expressed in normal astrocytes. Knocking down Olig2 in Ngn2-transduced astrocytes reduces astrocyte-specific gene expression, enhances neurogenesis-related gene expression, and increases AtN conversion efficiency by approximately threefold. Further multi-omics analysis shows that astrocytic Olig2 directly binds to regulatory regions of pro-neurogenic genes, including Ngn2, inhibiting their expression and impeding the expression of neural progenitor genes. Collectively, our findings reveal Olig2 as an inducible barrier to AtN conversion, providing insights into the regulation of neuronal reprogramming.
Similar content being viewed by others
Introduction
The adult mammalian central nervous system has limited neurogenic capacity after neuronal loss, while glial cells can proliferate and typically produce glial scars throughout life1. Among these glia, astrocytes exhibit neural stem cell-like properties in certain diseased conditions2,3, and are thus considered potential endogenous candidate cells for in vivo AtN conversion4,5. In general, the bHLH proneural transcription factors (TFs, e.g., Ngn2, Ascl1 and NeuroD1) are frequently selected as the key pioneer factors for driving AtN conversion because of their essential contribution to neurogenesis during development6,7,8. Despite significant progress in AtN conversion in animal studies, the barriers that inhibit AtN conversion are still not fully understood. Therefore, the low efficiency of AtN conversion will inevitably hinder its future clinical application.
Terminal differentiated cells maintain their identity through a complex interplay of genetic and epigenetic mechanisms that hinder cellular reprogramming9. In general, to achieve efficient cell reprogramming, we have to overcome these resistances existing within the cells10,11. For instance, the RE1 silencing transcription factor (REST) complex, known for its competition with Ngn2 for binding to the NeuroD4 promoter, acts as a barrier to AtN conversion12; the Notch signaling pathway, a highly conserved signaling system that is critical for regulating neurogenesis and promoting astrocytogenesis, is identified as another obstacle to AtN conversion3,13,14; unfolded protein response caused by gene mutation is recently identified as a novel hurdle for astrocyte reprogramming15; one study used a transcriptionally activated form of Olig2-VP16 to antagonize the function of Olig2 in dividing glial cells, promoting the generation of immature neurons, imply that Olig2 acts as a suppressor of neurogenesis in these dividing glial cells16. In addition, DNA methylation and microRNA (miRNA) inside cells also exert an important influence on AtN conversion at epigenetic level4,8,17. Apparently, the aforementioned obstacles are intrinsic and pre-existed in astrocytes regardless of AtN initiation, acting as the “innate” barriers to AtN conversion. Whether there is any obstacle that is absent in the normal astrocytes but induced after the ectopic expression of proneural TFs to induce neuronal reprogramming remains largely unknown.
In this study, we discovered the increase of Olig2+ cells during the in vivo TFs-mediated AtN conversion in the adult mouse cortex. These Olig2+ cells did not originate from the proliferation of oligodendrocyte precursor cells, but rather from astrocytes that ectopically expressed bHLH proneural TFs (Ngn2, Ascl1, or NeuroD1). Knockdown of Olig2 expression in astrocytes significantly enhanced the efficiency of Ngn2-mediated AtN conversion. ScRNA-seq study demonstrated that Olig2 knockdown in Ngn2-transduced astrocytes significantly altered their transcriptome, including further suppression of astrocyte-related genes and upregulation of neurogenesis-related genes. CUT&Tag-seq analysis revealed that astrocytic Olig2 directly bound to pro-neurogenesis genes and repressed their expression. Collectively, our study identifies a distinct inhibitory mechanism for AtN conversion, where the induced Olig2 upregulation in astrocytes counteracts the effect of proneural TFs, revealing a concept of “inducible” cellular defense against fate switching.
Results
bHLH pioneer transcription factors induce the expression of Olig2 in astrocytes
The brain parenchyma is predominantly composed of neurons and glial cells, encompassing astrocytes, oligodendrocytes, and microglia, which exhibit relatively stable population densities under normal conditions. In vivo AtN conversion utilizes endogenous astrocytes to generate neurons, which may affect the cell composition and microenvironments in the conversion regions. Therefore, to illuminate how AtN conversion affects the local cell populations, we examined the density of different cell types within the conversion regions. AAV9-based gene delivery system was used to target astrocytes via a synthetic GFAP promoter (hGFAP). The specificity of AAV9-hGFAP system was verified by immunostaining at 7 days post injection (dpi) of AAV9-hGFAP::tdTomato (tdT, control, Fig. 1a). Approximately 100% of tdT+ cells were co-labeled with the pan-astrocytic marker S100β (Fig. 1b, c). tdT reporter gene was not expressed in oligodendrocyte lineage cells (Sox10), microglia (Iba1), and neurons (NeuN, Fig. 1b). When AAV9-hGFAP::Ngn2-tdT, AAV9-hGFAP::Ascl1-tdT, or AAV9-hGFAP::NeuroD1-tdT was respectively injected in the mouse cortex (AAV9-hGFAP::TF-tdT, Fig. 1a), they were specifically expressed in the transduced astrocytes, as demonstrated by immunostaining (tdT+, Supplementary Fig. 1a–c). These results indicate that AAV9 combined with the hGFAP promoter is a highly specific gene delivery system for targeting astrocytes.
a Experimental design for testing control and transcription factors (TFs) in adult mouse cortex. Created in BioRender.com. Hou, KY. (https://BioRender.com/b8j6fdr). b Confocal images displaying tdT co-staining with S100β, Sox10, Iba1 and NeuN (Nuclei: DAPI). Scale bar, 20 μm. c Quantified data indicating high astrocyte targeting rate by the AAV9 system; n = 5 mice per group, mean ± SD. d Olig2 staining in control vs TF groups; white dashed line indicates the AAV9 infection area. Scale bar, 100 μm. e Quantified data indicating a significant increase of Olig2 density with TF overexpression; n = 5 mice per group, one-way ANOVA followed by Bonferroni’s post hoc test, mean ± SD. f The schematic treatment of Sox10-CreERT2; Ai14 transgenic mice (Top). Created in BioRender.com. Hou, KY. (https://BioRender.com/0vi388m). Confocal images showing GFP, tdT, and Olig2 co-staining. Scale bars, left 50 μm, right 10 μm. g Quantification of Olig2+ cell density; n = 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD. h The percentage of Olig2 distribution in tdT+ cells (Oligodendrocyte lineage cells) or GFP+ cells (AAV9-infected astrocytes) between the control and the Ngn2 group; n = 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD. i Pie chart showing increased Olig2 in GFP+ astrocytes with Ngn2 overexpression. j Quantified data showing no significant change of tdT+ cell number between the two groups; n = 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD. k Orthogonal images showing Olig2/GFAP co-expression (arrowhead). Olig2 presented only in Ngn2-transduced astrocytes (tdT+). Scale bars, 10 μm. l A significantly increased Olig2 expression in Ngn2-infected astrocytes labeled with different astrocytic markers; n = 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD. m, Sustained Olig2 upregulation from 3 dpi to 60 dpi after AAV9-GFAP::Ngn2 injection. Arrowheads: Ngn2-infected astrocytes with Olig2 (tdT+), arrows: endogenous Olig2+ cells (tdT-). DAPI for nuclei. Scale bar, 20 μm. n Quantification of Olig2 expression in Ngn2-tdT+ cells over the different time points (gray dotted line showing expression level in the control group); n = 5 mice per group, mean ± SD.
Then we examined the density of local cells in the AAV-TF transduced regions at 7 dpi and found no significant difference in the density of neurons (NeuN), astrocytes (S100β), microglia (Iba1), mature oligodendrocytes (CC1), and NG2 cells in the cortex of Ngn2-transduced mice compared with mice that transduced with tdT (Supplementary Fig. 1d). Furthermore, Ngn2 overexpression did not affect the morphology of microglia, that are the primary immune cell inside the brain (Supplementary Fig. 2). However, the density of Olig2+ cells increased 1.5-fold in AAV9-Ngn2 infected regions compared to the control group at 7 dpi (Fig. 1d, e). This phenomenon was also observed in the cortex of mice injected with AAV9-Ascl1 and AAV9-NeuroD1 (Fig. 1d, e; 7 dpi). Olig2+ oligodendrocyte progenitor cells (OPCs) are the major proliferating cells in the adult brain, so we wondered whether the increased density of Olig2+ cells was caused by OPC proliferation. To test this hypothesis, Sox10-CreERT2 mice were crossed with Rosa-tdTomato reporter mice (Ai14) to label oligodendrocytes and OPCs in the mouse brain18. As expected, Sox10 and Olig2 immunosignals were specifically observed in tdT+ cells in the cortex, indicating that these tdT+ cells are derived from the oligodendrocyte lineage (Supplementary Fig. 3a–e). Then, the lineage-tracing transgenic mice were administrated with AAV9-hGFAP::GFP (control) or AAV9-hGFAP::Ngn2-P2A-GFP in cortical regions at 2–5 months of age 10 days after tamoxifen induction, and mice were sacrificed for immunohistochemistry (IHC) at 7 days after AAV9 injection (Fig. 1f). Consistently, the reporter gene GFP was not expressed in the oligodendrocyte lineage cells (tdT+, Fig. 1f left) and the Olig2+ cells were dramatically increased in AAV9-Ngn2 injected cortical regions compared to the control (Fig. 1f right, 1 g). In the control group, Olig2 was essentially detected in all oligodendrocyte lineage cells (Ai14-tdT+), but not in AAV-infected cells (GFP+, Fig. 1h), showing that AAV-GFP didn’t change the Olig2 expression pattern and that Olig2 was restricted to oligodendrocyte lineage cells in the adult mouse cortex. In contrast, when AAV9-Ngn2-GFP was transduced into the astrocytes, Olig2 immunoreactivity was not only detected in oligodendrocyte lineage cells (Ai14-tdT+) but also in the Ngn2-transduced cells (GFP+, Fig. 1f right, 1 h). Quantitatively, about 30% of Olig2+ cells were not co-labeled with tdTomato but with GFP in AAV9-Ngn2 injected areas (Fig. 1i). However, the number of oligodendrocyte lineage cells (Ai14-tdT+) was comparable in both groups (Fig. 1j). In addition, there was no significant difference in the Olig2 expression intensity between the two groups in oligodendrocyte lineage cells (Supplementary Fig. 3f, g). Together, these results suggest that the increase of Olig2+ cells after the overexpression of Ngn2 in cortical astrocytes is not derived from the proliferation of OPCs.
Olig2 is primarily expressed in oligodendrocytes and their progenitors in the normal adult brain. However, Olig2 transient expression in the astrocytes has been reported after acute cortical injury19. Since AAV9 combined with the hGFAP promoter specifically targets astrocytes (Fig. 1b, c and Supplementary Fig. 1a–c), we wanted to know the relationship between the upregulation of Olig2 and the ectopic expression of proneural TFs in the astrocytes. In the control group, there were few Olig2 immunostaining signals in tdT+ cells (Fig. 1k left and Supplementary Fig. 1e left), whereas almost all tdT+ cells that expressed Ngn2, Ascl1, or NeuroD1 were Olig2 positive (Supplementary Fig. 1f–h). To further investigate that Olig2 was upregulated in cortical astrocytes, immunostaining for three typical astrocyte markers (GFAP, S100β, and Aldh1l1) was performed to identify astrocytes. About 5% of astrocytes were observed with Olig2 in the control group, however, this co-labeling rate was dramatically increased to nearly 100% after overexpression of Ngn2 at 7 dpi (Fig. 1k right, 1 l and Supplementary Fig. 1e right). Notably, astrocytic Olig2 was induced as early as 3 dpi following AAV9-Ngn2 administration and remained at a high level until 90 dpi in our examined time window (Fig. 1m, n). Collectively, these results indicate that Olig2 is significantly upregulated in cortical astrocytes following the ectopic expression of bHLH proneural transcriptional factors.
Non-bHLH transcription factors such as Sox2 and Dlx2 are also commonly selected to drive AtN conversion in mouse brain20,21,22. We then asked whether non-bHLH transcription factors could induce the expression of Olig2 in the astrocytes. In contrast to the bHLH proneural TFs, neither Sox2 (AAV9-hGFAP::Sox2-GFP) nor Dlx2 (AAV9-hGFAP::Dlx2-GFP) was able to induce the expression of Olig2 in the transduced cells (Supplementary Fig. 4). Thus, these data suggest that ectopic expression of the bHLH pioneer TFs (Ngn2, Ascl1, and NeuroD1) in cortical astrocytes results in upregulation of Olig2 in the transduced astrocytes, whereas other non-bHLH transcription factors do not exhibit this effect.
Inhibiting Olig2 expression promotes AtN conversion
Olig2 is a member of the bHLH family and plays a critical role during the differentiation and maturation of oligodendrocyte lineage cells in the central nervous system23,24,25, but the impact of Olig2 on AtN conversion needs further elucidation. While the molecular mechanisms of Ngn2-mediated AtN conversion are well understood, most studies have been in vitro rather than in vivo, and none have used the AAV-GFAP system employed here8,12,26. Therefore, we selected AAV-GFAP::Ngn2-mediated AtN conversion as a model to elucidate the role of Olig2 in the AtN conversion. For this purpose, we designed shRNAs to downregulate Olig2 (shOlig2) in astrocytes. We used EGFP for visualization (Supplementary Fig. 5a) and observed that the reporter gene (GFP) was primarily expressed in astrocytes and neurons, with minimal expression in NG2+ OPCs, CC1+ oligodendrocytes, or Iba1+ microglia (Supplementary Fig. 5b, c). Therefore, there was almost no impact on the endogenous Olig2 expression in OPCs and oligodendrocytes with AAV9-shOlig2 injection, as indicated by the similar proportion and expression level of Olig2 in OPCs and oligodendrocytes between the control shRNA (shNC) group and the shOlig2 group (Supplementary Fig. 5d–g). To utilize more channels for immunofluorescence staining, we excluded the EGFP sequence from the knockdown vector in subsequent experiments. When we combined Olig2 knockdown with Ngn2 overexpression, we observed a significant decrease in the mRNA level of Olig2 in fluorescence-activated cell sorting (FACS) purified tdT+ cells in the Ngn2+shOlig2 group (Fig. 2a, b). Further immunostaining studies showed that at 7 dpi, Olig2 was sharply decreased in Ngn2+shOlig2 treated mice (0.1% in tdT+ cells) compared to the Ngn2+shNC group (94.2%; Fig. 2c, d), indicating that the upregulation of Olig2 in Ngn2-overexpressing astrocytes was effectively inhibited by shOlig2.
a Scheme of the Olig2 shRNA (3 in 1) and experimental design combining Ngn2 and shRNA overexpression. Created in BioRender.com. Hou, KY. (https://BioRender.com/vqwedzp). b qPCR showing reduced Olig2 mRNA in FACS-sorted Ngn2-overexpressing astrocytes after Olig2 knockdown; n = 4 mice for Ngn2+shNC and n = 5 mice for Ngn2+shOlig2, unpaired two-sided Student’s t-test, mean ± SD. c Representative images showing efficient Olig2 downregulation in astrocytes by shOlig2 (arrows) compared to shNC (arrowheads). Scale bar, 20 μm. d Quantification of Olig2 knockdown efficiency; n = 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD. e Experimental design testing Ngn2 and shRNA induced astrocyte-to-neuron conversion. Created in BioRender.com. Hou, KY. (https://BioRender.com/uif2top). f Confocal images showing tdT, S100β, and NeuN co-staining among four groups at 45 dpi. White arrowheads indicating tdT⁺NeuN⁺ cells. Scale bar, 20 μm. g–i Conversion efficiency among groups. Note that Ngn2+shOlig2 induced tdT+S100β-NeuN- intermediate stage cells from 21 to 45 dpi (i) and induced more tdT+NeuN+ cells at 45 dpi (g). All time points for the Ngn2 group had n = 5 mice. For the Ngn2+shNC and Ngn2+shOlig2 groups, n = 5 mice at 7, 14, and 45 dpi, n = 7 mice at 21 dpi, n = 6 mice for Ngn2+shNC at 30 dpi, and n = 7 mice for Ngn2+shOlig2 at 30 dpi. One-way ANOVA followed by Bonferroni’s post hoc test, mean ± SD. j Representative morphology changes from 7 to 45 dpi in Ngn2+shOlig2 treated mice. Arrow indicating tdT+S100β+NeuN− cells, arrowhead indicating tdT+S100β-NeuN+ cells, and asterisk for tdT+S100β-NeuN- cells. n = 5 mice at 7, 14, and 45 dpi, n = 7 mice at 21 dpi. Scale bar, 20 μm. k Confocal images showing tdT co-staining with astrocytic markers GFAP/Sox9 and neuronal marker NeuN at 30 dpi. Scale bar, 5 μm. l Quantification showing ~40% of tdT⁺ cells were double-negative for astrocytic and neuronal markers at 30 dpi; n = 5 mice for GFAP and NeuN analysis, n = 4 mice for Sox9 and NeuN analysis. Data are presented as mean ± SD.
Next, we wanted to investigate the effect of Olig2 knockdown on Ngn2-mediated neuronal reprogramming, and four groups (tdT, Ngn2, Ngn2+shNC, Ngn2+shOlig2) were included for comparison (Fig. 2e). At 7 dpi, essentially all virus-infected cells were S100β+ astrocytes among the four groups (Supplementary Fig. 6a). However, at 45 dpi, more tdT+ cells showed NeuN signal in the Ngn2+shOlig2 injected mouse brain (Fig. 2f). Quantitative analyses revealed that approximately 30% of tdT+ cells became neurons (indicated by NeuN signal, Fig. 2g). Parallel to this trend, the percentage of tdT+ cells co-localized with S100β decreased from 98.1% at 7 dpi to 33.3% at 45 dpi in the Ngn2+shOlig2 group (Fig. 2h). Conversely, among the other three groups (tdT, Ngn2, and Ngn2+shNC), the number of tdT+ cells acquiring neuronal identity (NeuN+) was significantly lower than that of the Ngn2+shOlig2 group at 45 dpi (0.1% for tdT, 9.9% for Ngn2, 11.0% for Ngn2+shNC), and the remaining tdT+ cells were immunopositive for S100β (Fig. 2g, h). Notably, most of the converted neurons in the Ngn2+shOlig2 group were labeled by the glutamatergic neuron marker CamKII (Supplementary Fig. 6c, d), which is consistent with previous findings that Ngn2 directs the reprogramming of astrocytes into glutamatergic neurons12,27.
To elucidate the dynamic process of AtN conversion, we examined the morphological changes of tdT+ cells in the Ngn2+shNC and Ngn2+shOlig2 groups from 7 to 45 dpi. In the Ngn2+shOlig2 group, tdT+ cells initially displayed a typical astrocytic morphology during the first 14 days, co-localizing with S100β but not with the neuronal marker NeuN (Fig. 2g, h, j). At 21 dpi, round-shaped tdT+ cells emerged, which were double-negative for both S100β and NeuN, though their proportion was still small (Fig. 2i, j). At this stage, the neural stem cell markers Vimentin and Nestin were expressed in the tdT+ cells in Ngn2+shOlig2 group, while DCX expression was absent (Supplementary Fig. 7). By 30 dpi, the percentage of these double-negative cells increased, and a small number of tdT+ cells began to express NeuN (Fig. 2g, i, j). By 45 dpi, the proportion of tdT+ cells expressing NeuN had further increased, while the proportion of double-negative cells began to decline (Fig. 2g–j). In contrast, the Ngn2+shNC group exhibited minimal morphological changes, with most tdT+ cells retaining a typical astrocytic morphology throughout the time course (Supplementary Fig. 6b). Only at 45 dpi did a small fraction of tdT+ cells (approximately 10%) express NeuN (Fig. 2g–i). These findings indicate that Olig2 knockdown in astrocytes significantly enhances Ngn2-mediated AtN reprogramming and suggest the existence of a transient double-negative intermediate state during the conversion of astrocytes to neurons.
It was worth noting that during the conversion process, some tdT+ cells in the Ngn2+shOlig2 group were double negative for astrocytic marker S100β and neuronal marker NeuN (28.6% at 21 dpi, 44.8% at 30 dpi, 37.9% at 45 dpi), which was rare in other 3 groups (tdT, Ngn2 and Ngn2+shNC; Fig. 2i). We next wondered whether these double-negative cells had lost typical astrocyte features and used two other astrocyte markers, GFAP and Sox9, co-stained with NeuN at 30 dpi, respectively. The three populations of tdT+ cells were detected in the brains of mice treated with Ngn2+shOlig2. First, GFAP and Sox9 were positive in tdT+ cells without NeuN expression, and these cells represented astrocytes (Fig. 2k, left; quantified in 2 l). Second, some tdT+ cells showed neither GFAP, Sox9, nor NeuN signals (Fig. 2k, middle; quantified in 2 l), which may represent intermediate-stage cells as reported by a previous study21. These intermediate cells likely represent a transitional stage during AtN reprogramming, where the cells have lost their astrocytic identity, but neuronal gene expression has not yet been fully upregulated, resulting in the observation of these double-negative intermediate cells. Finally, when NeuN signals became detectable, tdT+ cells showed neuronal morphology and were negative for GFAP and Sox9 (Fig. 2k, right; quantified in 2 l), and these cells represented converted neurons. Together, these data indicate that Olig2 knockdown promotes AtN conversion, which may go through an intermediate stage between astrocytes and neurons.
To further confirm the inhibitory effect of Olig2 on Ngn2-mediated AtN conversion, we used a dominant active form of Olig2 (Olig2-VP16) to antagonize the inhibitory function of endogenous Olig2 in astrocytes16,28. Olig2-VP16 was driven by the hGFAP promoter and co-overexpressed with tdT or Ngn2-tdT (Supplementary Fig. 8a). At 7 dpi, we confirmed that control tdT had high GFAP+ astrocytes targeting rate, Olig2 and VP16 were also strongly expressed in tdT+ cells (Supplementary Fig. 8b–e). Multi-time point analysis revealed that Olig2-VP16 alone failed to convert astrocytes into neurons in the absence of Ngn2 from 7 dpi to 45 dpi, with almost all tdT+ cells still expressing S100β but not NeuN. However, when Olig2-VP16 worked with Ngn2, the AtN conversion efficiency reached up to 40% at 45 dpi, as indicated by a decrease in the proportion of tdT+S100β+ cells and an increase in tdT+NeuN+ cells (Supplementary Fig. 8f–h). Taken together, these findings further demonstrate that Olig2 plays a negative regulatory role in Ngn2-induced AtN conversion, and removing the inhibition of Olig2 could promote cell fate conversion.
Functional analysis of converted neurons
To test whether the converted neurons were functional, we performed whole-cell patch-clamp recordings in acute cortical slices at 40–45 dpi, with the infected cells identified by the expression of tdTomato (Fig. 3a). We first compared the Na+-K+ currents between Ngn2+shNC (control) and Ngn2+shOlig2 (conversion) treated mice and found larger K+ currents in the control group cells, without a significant fast Na+ current, exhibiting typical astrocyte electrophysiological properties (Fig. 3b, top). However, in the conversion group cells, we not only recorded outward K+ currents but also inward fast Na+ currents (Fig. 3b, bottom). Of note, the K+ currents were significantly reduced while Na+ currents were dramatically increased in conversion group cells compared to the control group (Fig. 3c). Next, for testing the probability of action potential firing, step currents were injected into the AAV-infected cells under the current patch-clamp model. Consistently, no evoked action potentials were recorded in the control group (Fig. 3d top), while 95.5% of conversion group cells were able to generate action potentials (18.2% with single spike and 77.3% with regular spikes) upon the step current injection (Fig. 3d bottom and quantified in 3e). Thus, Ngn2+shOlig2 can convert cortical astrocytes from non-excitatory glial cells to excitable neurons.
a Phase and fluorescent images of a tdT+ astrocyte-like cell in the Ngn2+shNC group and a tdT+ neuron-like cell in the Ngn2+shOlig2 group. Scale bar, 10 μm. b Representative traces of Na+-K+ currents. Upward traces represented K+ currents, and the sharp inward traces indicated the fast Na+ currents, respectively. c I-V plot of K+ (left) and Na+ currents (right) from tdT+ cells of Ngn2+shNC and Ngn2+shOlig2; n = 18 cells from 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD. d Representative traces of action potential (AP) firing. e Pie chart showing the percentage of tdT+ cells displaying different AP firing patterns in the Ngn2+shOlig2 group; n = 22 cells from 5 mice. Electrical properties of tdT+ cells: membrane capacitance (f), membrane input resistance (g), resting membrane potential (RMP, h). Ngn2+shNC: n = 35 cells from 5 mice; Ngn2+shNC: n = 24 cells from 5 mice; unpaired two-sided Student’s t-test, mean ± SD. i Typical traces of sEPSCs recorded from tdT+ cells. Quantified data showing sEPSC frequency (j) and amplitude (k); n = 22 cells from 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD. l Typical traces of sIPSCs recorded from tdT+ cells. Quantified data showing sIPSC frequency (m) and amplitude (n); n = 22 cells from 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD. o The pie chart showing the proportion of all converted neurons that could fire APs, with and without PSCs in the Ngn2+shOlig2 group; n = 21 cells from 5 mice.
In addition, when comparing the tdT+ cells between the control and conversion group, tdT+ cells with Olig2 knockdown had lower membrane capacitance (Fig. 3f) and higher membrane input resistance (Fig. 3g), implying a reduction in gap junctions when astrocytes were converted into neurons. We also found that tdT+ cells in the control group displayed a more negative resting membrane potential (RMP) than those in the conversion group (Fig. 3h). Moreover, we further examined whether the converted neurons could integrate into the local circuit and form functional synaptic connections. To this end, both spontaneous excitatory postsynaptic currents (sEPSCs) and spontaneous inhibitory postsynaptic currents (sIPSCs) were recorded in tdT+ cells. As expected, none of the tdT+ cells exhibited any spontaneous synaptic events in control group cells, whereas both sEPSCs and sIPSCs were detected in half of the conversion group cells (Fig. 3i–n). Notably, approximately half of the converted neurons (47.6%) acquired the ability to generate action potentials but no detectable postsynaptic currents (PSCs, Fig. 3o), implying that these newly converted cells require a longer period of maturation to establish functional synaptic connections. Taken together, our electrophysiological results demonstrate that cortical astrocytes can be converted into functional neurons by Ngn2+shOlig2 and integrate into local synaptic circuits.
Knockdown of Olig2 enhances AtN conversion efficiency in astrocyte lineage-tracing mice and retro-labeling study
To further validate the enhanced conversion efficiency induced by Olig2 knockdown, we employed astrocytic lineage-tracing mice generated by crossing Aldh1l1-CreERT2 mice with Ai14 mice (Fig. 4a). After tamoxifen treatment, around 98.9 ± 0.7% and 98.1 ± 0.8% tdT+ cells expressed astrocyte marker S100β and Sox9 in cortex, respectively (Figs. 4b, c), indicating highly efficient to pre-label endogenous astrocytes. Next, we found that under normal conditions, around 6.7 ± 2.2% tdT+S100β+ astrocytes expressed Olig2 in the adult mouse cortex, consistent with previous reports in wild-type mice. Seven days after Ngn2 injection, Olig2 was markedly upregulated in tdT+ astrocytes (94.2 ± 1.7% in tdT+Ngn2+ cells, Figs. 4d, e). Thus, these results indicate that overexpression of Ngn2 in lineage-tracing mice also significantly increases Olig2 expression in pre-labeling astrocytes.
a Schematic diagram showing the cross of Aldh1l1-CreERT2 mouse and Ai14 mouse to produce the double transgenic mouse, in which astrocytes can express tdTomato after tamoxifen administration. Created in BioRender.com. Hou, KY. (https://BioRender.com/kvri6mm). b Confocal images showing co-staining of tdT and S100β or Sox9 ten days after tamoxifen induction. Scale bar, 20 μm. c Quantified data showing high level co-expression between tdT and S100β or Sox9, indicating high specificity of tdT signal; n = 5 mice per group, mean ± SD. d The schematic treatment of Aldh1l1-CreERT2; Ai14 transgenic mice (Top). Created in BioRender.com. Hou, KY. (https://BioRender.com/qb5kiom). Immunostaining showing Olig2 expression in tdT+S100β+ astrocytes under normal conditions and in Ngn2+shNC/shOlig2 infected tdT+ astrocytes. Arrows indicated astrocytes without Olig2, and arrowheads indicated astrocytes with Olig2. Scale bar, 10 μm. e Quantification showing the proportion of Olig2 expression in tdT+Ngn2+ cells. Baseline showing the percentage of Olig2 expression in endogenous tdT+S100β+ astrocytes; n = 5 mice per group, One-way ANOVA followed by Bonferroni’s post hoc test, mean ± SD. f Representative images showing co-staining of tdT, S100β and NeuN, indicating the conversion efficiency. Scale bar, 20 μm. White boxes representing the enlarged images. Scale bar, 5 μm. g Quantified data illustrating a reduced proportion of tdT+S100β+ cells and an increased proportion of tdT+NeuN+ cells after Olig2 knockdown; n = 5 mice per group (a total of 1252 cells were included in the analysis for the no-AAV group, 1140 cells for Ngn2+shNC, and 1366 cells for Ngn2+shOlig2), One-way ANOVA followed by Bonferroni’s post hoc test, mean ± SD.
Given that Olig2 knockdown increased conversion efficiency about three-fold in wild-type mice at 45 dpi (Fig. 2g), we extended this analysis to the lineage-traced mice. First, we confirmed the knockdown efficiency at 7 dpi. Quantitative analysis revealed that Ngn2 co-expression with shOlig2, compared to shNC controls, significantly reduced Olig2 protein levels (Fig. 4d-e). In the absence of AAV injection, the lineage-traced mice exhibited over 98% labeling specificity for astrocytes in the cortical area (Fig. 4f top, g left), with tdT+ neurons comprising less than 2% (Fig. 4f top, g right). At 45 dpi, we observed that around 5.0% tdT+ cells became NeuN+ in the Ngn2+shNC group, accompanied by a corresponding decrease in tdT+S100β+ astrocytes compared to the normal conditions. As expect, shOlig2 significantly promoted the generation of tdT+NeuN+ cells, reaching up to 12.2% (Fig. 4f, g), further supporting the inhibitory role of Olig2 in Ngn2- mediated AtN conversion.
To further validate AtN reprogramming in the mouse cortex, we pre-labeled endogenous neurons with GFP by injecting engineered AAV2 variants. Adult mice were first injected with rAAV2-retro-CAG::GFP (Retro-GFP) into the T6 corticospinal tract (CST), followed by intracerebral injections of AAV9-Ngn2-tdT+shNC or AAV9-Ngn2-tdT+shOlig2 viruses (Supplementary Fig. 9a). Forty-five days after the AAV9 injection, we observed 7.6% and 22.3% tdT+ neurons in the brains injected with Ngn2-tdT+shNC and Ngn2-tdT+shOlig2 viruses, respectively (Supplementary Fig. 9b, c). Specifically, in the Ngn2+shNC group, 416 tdT+ AAV-transduced cells were analyzed, of which 34 were neurons. Among these, 5 tdT+ neurons were pre-labeled with GFP, representing approximately 15% of the neuronal population analyzed (Supplementary Fig. 9b, d). In the Ngn2+shOlig2 group, 968 tdT+ AAV-transduced cells were analyzed, with 224 tdT+ neurons identified. Notably, the majority of these tdT+ neurons (96%) were not labeled by GFP (Supplementary Fig. 9b, d). These results suggest that Olig2 knockdown in astrocytes significantly enhances the efficiency of Ngn2-mediated AtN reprogramming, rather than causing nonspecific tdT reporter expression in endogenous neurons.
ScRNA-seq reveals the underlying mechanism of reprogramming after knocking down Olig2 in astrocytes
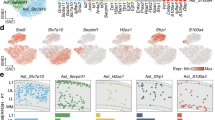
To gain mechanistic insight into how Olig2 impedes AtN conversion, we performed single-cell RNA sequencing (scRNA-seq) of freshly pooled cortical samples infected with Ngn2 either in the presence or absence of shOlig2 vectors at 18, 20 and 22 dpi (n = 2 mice for each time-point, Fig. 5a), when astrocytes infected with Ngn2 and shOlig2 began to lose astrocytic features (Fig. 2h). After data-quality filtering, we obtained 32,095 cells from the Ngn2+shNC group (median gene count: 928) and 32,439 cells from the Ngn2+shOlig2 group (median gene count: 894). Uniform manifold approximation and projection (UMAP) analysis identified 11 cell types based on the respective representative marker genes (Supplementary Fig. 10a, b). The overall cell composition of the Ngn2+shNC group and the Ngn2+shOlig2 group was similar (Supplementary Fig. 10c, d). The tdTomato transcript was mainly distributed in astrocytes, whereas tdTomato reporter gene was barely detected in neurons (Supplementary Fig. 10e, f). This might be due to the limited number of reprogrammed neurons at these time points (Fig. 2g, j), the decreased activity of the GFAP promoter in converted neurons, and neuronal damage caused by tissue dissociation. Additionally, tdT transcripts were detected in a small subset of oligodendrocytes (Supplementary Fig. 10e, f). However, analysis of the distribution and relative expression levels of tdT transcripts revealed that both were much higher in astrocytes than in oligodendrocytes. (Supplementary Fig. 10g). To compare the transcriptomic changes in AAV-infected astrocytes between the two groups, we first separated tdT+ astrocytes from the total cell population for further analysis (Fig. 5b, c). We confirmed the downregulation of Olig2 transcript and protein expression level in AAV-shOlig2 infected cells at 21 dpi (Fig. 5d, e), similar to the immunostaining results at 7 dpi (Fig. 2c, d). We then performed differential expression analysis between Olig2 knockdown tdT+ cells (Ngn2+shOlig2) and the controls (Ngn2+shNC), identifying 1281 upregulated genes including those associated with neurogenesis (e.g., Nes, Sox4, and Pax6) and 597 downregulated genes embracing those related to mature astrocyte function (e.g., Slc1a2, S100β, and Aqp4, Fig. 5f). Gene Ontology (GO) enrichment analysis revealed that the upregulated genes after Olig2 knockdown were related to mRNA processing, translation and neurogenesis, whereas metabolism and gliogenesis-related pathways were significantly downregulated (Fig. 5g, h). Real-time PCR further demonstrated that the mRNA of Nestin, Vimentin and Olig1 were significantly upregulated in Ngn2+shOlig2 infected astrocytes (Fig. 5i). Furthermore, we confirmed that Nestin and Vimentin, known to be expressed in neural progenitor cells, were slightly expressed in Ngn2-transduced astrocytes and then significantly increased after Olig2 knockdown, implying that astrocytic Olig2 represses neurogenesis gene expression (Fig. 5j, k).
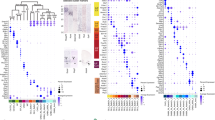
a Experimental design. Created in BioRender.com. Hou, KY. (https://BioRender.com/0zr0vf9). b UMAP showing the astrocytes of the Ngn2+shNC and the Ngn2+shOlig2 groups. c tdTomato projection to (b). There were a total of 557 tdT+ astrocytes. d Comparison of Olig2 gene expression between the Ngn2+shNC and the Ngn2+shOlig2 groups. e tdT and Olig2 double staining. n = 5 mice per group. Scale bar, 20 μm. f Differentially expressed genes (Ngn2+shOlig2 vs Ngn2+shNC) in tdT+ astrocytes. g Gene ontology analysis showing top significantly downregulated biological process (tdT+ astrocytes: Ngn2+shOlig2 vs Ngn2+shNC). h Gene ontology analysis showing top significantly upregulated biological process (tdT+ astrocytes: Ngn2+shOlig2 vs Ngn2+shNC). i RT-qPCR verifying higher mRNA expression level of Nestin, Vimentin, and Olig1 with Olig2 knockdown in tdT+ astrocytes (purified by FCAS); n = 5 mice per group for Nestin and Olig1 quantification, n = 4 mice in Ngn2+shNC group, and n = 5 mice in Ngn2+shOlig2 group for Vimentin quantification. Unpaired two-sided Student’s t-test, mean ± SD. j Immunostaining and fluorescence intensity quantification showing the higher expression level of Nestin in tdT+ cells of the Ngn2+shOlig2 group. Scale bar, 4 μm; n = 5 mice per group, unpaired two-sided Student’s t-test. Data are presented as mean ± SD. k Immunostaining and fluorescence intensity quantification showing the higher expression level of Vimentin in tdT+ cells of the Ngn2+shOlig2 group. Scale bar, 6 μm; n = 5 mice per group, unpaired two-sided Student’s t-test. Data are presented as mean ± SD. l Subpopulation analysis of astrocytes. m Histogram showing the percentage of astrocyte subclusters in each group. n Histogram showing the percentage of each group in each astrocyte subcluster. o Heatmap showing different gene expression patterns among astrocyte subclusters. p Feature plots highlighting differential expression of Sox4, Nestin, Olig1, and Vimentin in control and Olig2 knockdown astrocytes. q Pseudotime analysis of astrocyte subclusters and NPCs. r Relative expression of indicated genes in the pseudotime trajectory.
To provide detailed insights into the changes in cellular feature of the astrocytes after the knockdown of Olig2 during the Ngn2-mediated AtN conversion, we performed an unbiased astrocyte sub-clustering analysis and further divided the astrocyte population into five subclusters (A1-A5, Fig. 5l) according to the differentially expressed markers (Supplementary Fig. 10h). Gene set variation analysis (GSVA) showed that A1 astrocyte was enriched for genes related to synapse regulations. A2 and A5 astrocytes shared similar functions related to metabolism and energy production. A3 and A4 astrocytes were featured by active gene transcription and translation (Supplementary Fig. 10i). Interestingly, the percentages of A1 and A2 were reduced following Olig2 knockdown, whereas A3 (green) and A4 (red) showed significant increases (Fig. 5m). In addition, tdT+ astrocytes in both groups were detected in the A3 and A4 subpopulations (Fig. 5c, l and supplementary Fig. 12g), indicating that Ngn2 alone is sufficient to change the profile of astrocytes and push them to enter a transcriptionally and translationally active state. But Olig2 knockdown further increased the proportion of A3 and A4 astrocyte populations in the Ngn2-transduced astrocytes (Fig. 5n). Additionally, we performed a supplementary analysis on the Ngn2+shNC group without AAV infection in astrocytes (tdT-). We found that only 9.3% of these astrocytes were distributed in the A3 and A4 subpopulations, while about 90% were observed in the A1 and A2 subpopulations (Supplementary Fig. 10j). Together, these data suggest that the induced astrocytic Olig2 may inhibit the Ngn2-transduced astrocytes to enter a transcriptionally and translationally active state.
We next analyzed detailed features of the A3 and A4 subpopulations. Genes related to mature astrocyte functions, such as Aqp4 for BBB maintenance, Slc1a2 for glutamate transport, Apoe for lipid transport, and Glul for glutamine synthesis, were downregulated in these subpopulations (Fig. 5o). Meanwhile, genes related to cell dedifferentiation, chromatin remodeling, neurogenesis and axonogenesis were significantly upregulated (Fig. 5o). We then selected four representative genes associated with astrocyte dedifferentiation (Sox4, Nestin, Olig1 and Vimentin) for scatter plot visualization. Our analysis revealed that these genes were predominantly concentrated within the A3 and A4 subpopulations and the frequency of these astrocyte populations was significantly elevated in the Olig2 knockdown group (Fig. 5p). Moreover, A3 and A4 astrocytes also upregulated genes involved in cell cycle regulation (e.g., Ccnd2, Fgfr1 and Akt1), chromatin remodeling (e.g., H2aj, Hdgf and Hat1), RNA processing and splicing (e.g., Dhx9, Ddx5 and Akt1), translation (e.g., Rpls and Rpss), and actin cytoskeleton organization (e.g., Ezr, Coro1c and Actg1; Fig. 5o). Importantly, genes associated with neurogenesis and axonogenesis, such as Pax6, Stmn1, Smn1, Robo2, and Carpin1, were also upregulated in A3 and A4 (Fig. 5o). Collectively, these data reveal that A3 and A4 astrocytes undergo a phenotypic transition that exhibits characteristics of reprogramming toward neural progenitor cells.
CytoTRACE is a computational method to predict the potential of cell reprogramming and differentiation potential based on scRNA-seq data, with higher scores suggesting better differentiation potential29. Analysis and quantification showed that knocking down Olig2 in Ngn2-transduced astrocytes led to a significant increase in the CytoTRACE score (Supplementary Fig. 11a). Further analysis of astrocyte subpopulations revealed that A3 and A4 subclusters showed higher CytoTRACE scores compared with A1 and A2 (Supplementary Fig. 11b), suggesting that A3 and A4 astrocytes have stronger reprogramming potential. Next, to investigate the AtN conversion at the single-cell level, we employed cell fate trajectory and pseudotime analysis using Monocle 2, incorporating neural progenitor cells (NPCs) into this analysis. Of note, NPCs were mainly found in Ngn2+shOlig2 treated mouse cortex (Supplementary Fig. 12a–c), which were highly expressed Nnat, Tubb2b, Igfbpl1, and Marcksl1 genes (Supplementary Fig. 12b). These genes are mainly related to axon growth, neuronal differentiation, and regeneration, and are highly expressed in immature neurons. Our analysis revealed a cellular continuum with two distinct branch points that originated from astrocytes with mature astrocyte function (A1 and A2) and differentiated into astrocytes with features of immature astrocytes and neurogenesis (A3 and A4). A branch point emerged from A1 and A2, diverging into two distinct paths: one predominantly leading to A3 (cell fate 1) and the other to A4 (cell fate 2). In control cells (Ngn2+shNC), A1 and A2 were predominantly located in the pre-branch region (Fig. 5q, left panel). However, upon knocking down Olig2, A1, A2, and A3 cells moved beyond the branch point, adopting characteristics of cell fates 1 and 2 (Fig. 4q, right panel). In addition, most A3 cells and some A4 cells were located near NPCs (Fig. 5q), indicating intrinsic similarity, and this distribution pattern was more pronounced after knockdown of Olig2 (Supplementary Fig. 12d–g). Along the cellular trajectory, mature astrocyte markers (e.g., Glul and Aqp4) were downregulated, while neural progenitor marker Vimentin was gradually upregulated (Fig. 5r), indicating a transition towards a neural progenitor fate. Olig1, another bHLH transcription factor reported to have the ability to promote axonogenesis30, also showed a gradual increase and then was maintained at a certain level in cellular trajectory (Fig. 5r). Together, knocking down Olig2 in Ngn2-transduced astrocytes leads to significant transcriptomic changes and upregulates neural stem cell signature genes.
Olig2 inhibits Ngn2-mediated AtN conversion in multiple ways
To further illuminate how Olig2 knockdown in Ngn2-transduced astrocytes facilitates AtN conversion, we investigated the additional alterations in the transcriptome of Ngn2-overexpressing astrocytes resulting from the knockdown of Olig2. We first compared the transcriptome of Ngn2-transduced astrocytes (tdT+) and that of non Ngn2-transduced astrocytes (tdT-) in the Ngn2+shNC group to figure out the changes induced by Ngn2 (Fig. 6a left, DEGs-1). Ngn2 alone is sufficient to change the gene expression profiles of astrocytes (Supplementary Fig. 13a–c). Among these genes, we identified Vim, Sox4, Nes, and Olig1 genes have been upregulated, while Aldoc, Glul and Gjb6 genes have been downregulated in tdT+ astrocytes (Supplementary Fig. 13a). GO analysis revealed that genes associated with biosynthesis and membrane transport were significantly downregulated, while genes related to cellular respiration and translation were markedly upregulated (Supplementary Fig. 13b, c). Notably, genes involved in neurogenesis and axonogenesis, which were enriched in the Ngn2+shOlig2 group (Fig. 5h), were conspicuously absent from the upregulated pathways induced by Ngn2 alone, suggesting that this may underlie the limited efficiency of Ngn2-mediated AtN conversion.
a The top cartoon illustrating the origin of the two DEGs datasets. Created in BioRender.com. Hou, KY. (https://BioRender.com/j2p0bx9). The common and unique DEGs were shown in the Venn diagram at the bottom. b Bar chart showing the top GO-BP enriched pathways unique to Ngn2-overexpressing astrocytes without knockdown of Olig2. c Bar chart showing the top GO-BP enriched pathways specific to Ngn2-overexpressing astrocytes with Olig2 downregulation. d The Venn diagram showed four different regulatory patterns of 367 common DEGs, representing that Olig2 exerted a profound effect on the regulation of gene expression in Ngn2-overexpressing astrocytes. Chord diagram showing the genes and enriched GO pathways for Pattern 1 (e), Pattern 2 (f), Pattern 3 (g), and Pattern 4 (h), respectively.
Then, we compared the two groups of DEGs that were generated from the comparisons between either tdT+ and tdT- astrocytes in Ngn2+shNC group (defined as DEGs-1), or tdT+ astrocytes in Ngn2+shOlig2 and those in Ngn2+shNC (defined as DEGs-2; Fig. 6a). Venn diagram displayed the common and unique DEGs in the 2 datasets (Fig. 6a). There were 367 common DEGs shared between the two datasets, while 542 and 1308 unique DEGs were found in DEGs-1 and DEGs-2, respectively (Fig. 6a). DEGs unique to DEGs-1 represent the profile change induced by the Ngn2 that contained Olig2 in astrocytes, which were associated with cytoplasmic translation and cellular respiration (Fig. 6b). DEGs unique to DEGs-2 represent the profile change induced by knocking down Olig2 upon Ngn2 overexpression in astrocytes, which included specific genes relevant for RNA splicing, processing and protein catabolic process (Fig. 6c). Furthermore, upregulated DEGs related to neurogenesis, such as Pax6, Robo2, and Stmn1, as well as downregulated DEGs associated with mature astrocyte functions, such as Aqp4 and Slc1a2, were also unique to Olig2 knockdown in Ngn2-transduced astrocytes (Fig. 6a, bottom Venn diagram). Thus, knocking down Olig2 in Ngn2-transduced astrocytes further substantially alters the astrocyte transcriptome and promotes neurogenic related genes expression.
Next, we focused on common DEGs shared by DEGs-1 and DEGs-2 that may reflect the patterns of how Olig2 impacts Ngn2-mediated AtN conversion, and identified four distinct patterns (Fig. 6d). Pattern 1 included genes that were upregulated by Ngn2 and further upregulated by Olig2 knockdown (Fig. 6d, left), which were mainly associated with ribosome biogenesis and cytoplasmic translation (Fig. 6e). These results suggest that knocking down Olig2 may alter the expression of genes involved in protein synthesis, which could be linked to the cellular reprogramming process and the need for new functional proteins. Pattern 2 contained genes that were initially downregulated by Ngn2 and further downregulated by Olig2 knockdown (Fig. 6d, middle left). The functions of these genes were mainly associated with membrane and cell junction adjustment, as well as synapse regulation (Fig. 6f), which are typical functions of astrocytes. These results indicate that knockdown of Olig2 further inhibited the expression of astrocyte signature genes, suggesting that knockdown of Olig2 may facilitate Ngn2-mediated astrocyte dedifferentiation. Moreover, our analysis revealed not only a synergistic relationship between Ngn2 and Olig2, but also two distinct patterns of astrocytic Olig2 functionality that exhibit opposing effects. In Pattern 3, genes were upregulated by Ngn2 but downregulated by Ngn2+shOlig2 (Fig. 6d, middle right). We observed several pathways related to metabolism (Fig. 6g), consistent with previous findings that cellular reprogramming is accompanied by many metabolic changes7,31. Notably, among these genes, we also identified Hes5, which has been reported as a repressor for neurogenesis32, and it was significantly upregulated by Ngn2 overexpression but downregulated by Olig2 knockdown (Fig. 6d, middle right). Finally, Pattern 4 exhibited genes that were downregulated by Ngn2 but upregulated upon the knockdown of Olig2 (Fig. 6d, right), which are primarily related to receptor regulation (Fig. 6h). Cellular receptor remodeling represents a pivotal event in the process of reprogramming, which involves the reorganization and functional changes of cell receptors to support the cell fate switch. Collectively, knockdown of Olig2 further alters the gene expression signature of astrocytes and then promotes Ngn2-mediated astrocyte reprogramming in multiple aspects.
Olig2 knockdown alters astrocyte metabolic gene expression
Given the common occurrence of metabolic shifts during AtN conversion7,31, we compared changes in gene expression associated with metabolic preference of tdT+ astrocytes in Ngn2+shNC or Ngn2+shOlig2 treated animals. The results indicated that cells with Olig2 knockdown exhibited an overall low metabolic score (Fig. 7a). Classical metabolic pathways preferred in astrocytes, including canonical glycolysis, glucose metabolism, fatty acid metabolism, β-oxidation, glutamate and pyruvate metabolism were notably downregulated in the Ngn2+shOlig2 group (Fig. 7b–h). Oxidative phosphorylation (OXPHOS), a pathway typically favored in neurons, did not show a significant difference (Fig. 7i), primarily due to the impact of Ngn2 alone on the elevation of OXPHOS levels (Fig. 7j). We then performed a metabolic pathway activity scoring analysis on subclusters of astrocytes and neurons, observing that A3 and A4 exhibited lower metabolic value, a trend also detected in neurons (Supplementary Fig. 14a). ScMetabolism pathway analysis revealed that astrocytes had distinct metabolic transcriptome profiles compared to neurons and neural stem cells. Specifically, metabolic pathways that were highly active in A2 and A5 astrocytes, but relatively quiescent in neurons and NPCs (glycolysis/gluconeogenesis, pentose phosphate pathway, steroid biosynthesis, etc.), were found to be downregulated in A3 and A4 astrocytes. Interestingly, a metabolic pathway (N-glycan biosynthesis) that had high value in excitatory neurons (ENs) but less value in A1, A2 and A5 astrocytes, was upregulated in A3 and A4 (Supplementary Fig. 14b), consistent with Ngn2 primarily inducing ENs (Supplementary Fig. 6c, d).
a Violin plots showing lower metabolic activity with the knockdown of Olig2 in Ngn2-overexpressing astrocytes (tdT+). n = 209 tdT+ astrocytes in Nng2+shNC group, n = 348 tdT+ astrocytes in Nng2+shOlig2 group. Mann-Whitney U test. Data are shown as box plot (boxes, 25–75%; whiskers, 1.5 × Interquartile Range; lines, median). Violin plots showing different levels of diverse metabolic pathways between the two groups, including glycolysis (b), β-oxidation (c), glutamate metabolism (d), glucose metabolism (e), fatty acid metabolism (f), tricarboxylic acid cycle (g), pyruvate metabolism (h), and oxidative phosphorylation (i). The p values were shown at the top of the graphs. n = 209 tdT+ astrocytes in Nng2+shNC group, n = 348 tdT+ astrocytes in Nng2+shOlig2 group. Mann–Whitney U test. Data are shown as box plot (boxes, 25-75%; whiskers, 1.5 × Interquartile Range; lines, median). j Dot plots showing Ngn2 alone upregulating neuron-preferred metabolic pathways. k The Venn diagrams showing the overlap between DEGs from scRNA-seq and the reported mitochondrial genes enriched in astrocytes or neurons. l Heatmap showing the expression levels of the overlapping DEGs indicated in (k) between the two groups. m Immunostaining showing Sfxn5 expression between the Ngn2+shNC and the Ngn2+shOlig2 groups. Scale bar, top: 200 μm and bottom: 5 μm. n Quantification showing significant lower expression level of Sfxn5 after Olig2 knockdown; n = 5 mice per group, unpaired two-sided Student’s t-test, mean ± SD.
Mitochondria have been reported to play a crucial role in metabolic shifts, and specific mitochondrial proteins are essential for implementing the AtN conversion7. Studies indicate that astrocytes and neurons exhibit distinct mitochondrial proteomes, and cells lacking the upregulation of neuron-enriched mitochondrial proteins often fail to achieve successful reprogramming7,15. In our data, we identified 31 mitochondrial protein-related genes enriched in astrocytes and 21 enriched in neurons that underwent transcriptional changes following Olig2 knockdown (Fig. 7k). Among the astrocyte-enriched genes, 74.2% (23 out of 31, e.g., Sfxn5, Prdx6, and Cox20) were downregulated in the Ngn2+shOlig2 group. In contrast, 85.7% (18 out of 21, e.g., Prdx2, Bax, and Slc25a22) of neuron-enriched genes were upregulated with knockdown of Olig2 (Fig. 7l). Sfxn5 is a mitochondrial transporter responsible for citrate transport and is enriched in astrocyte-derived mitochondria7. Immunostaining results revealed that the expression of Sfxn5 in astrocytes was significantly decreased in the Ngn2+shOlig2 group (Fig. 7m, n). These findings suggest that the knockdown of astrocytic Olig2 promotes the shift from astrocytic mitochondria transcriptome profile to that of neurons, which may facilitate Ngn2-mediated AtN reprogramming.
CUT&Tag genome binding profiling of astrocytic Olig2
Olig2 belongs to the bHLH family, which forms heterodimers or homodimers to bind to the DNA and serves as a transcription regulator24. To uncover the targets of Olig2, which impede the Ngn2-mediated AtN conversion, we conducted Olig2 CUT&Tag sequencing to analyze the genomic binding loci of Olig2 in the Ngn2-transduced cells. We collected tdT+ astrocytes infected with AAV9-Ngn2 through FACS and then subjected them to Olig2-CUT&Tag sequencing (Fig. 8a). A total of approximately 60,000 sorted cells from 12 mice were pooled for a single experiment, and there two replicates were conducted in this study (Fig. 8b). We identified 8372 Olig2-binding sites within 3 kb upstream or downstream of the annotated transcription start site (TSS, Fig. 8c). Among these Olig2-occupied loci, most of them were in the regulatory regions including intergenic regions (44.3%), introns (35.0%) and promoters (16.4%, Fig. 8d). The closest annotated gene to each Olig2-binding locus was considered as a presumed target, and GO analysis revealed that Olig2-target genes were primarily associated with cell communication, signaling regulation, and development (Fig. 8e). We then performed motif analysis to investigate whether the DNA motifs of Olig2 binding sites in cortical astrocytes were similar to those in OPCs. We did not detect canonical E-box motif (CANNTG) in AAV-Ngn2 transduced astrocytes in two replicate experiments. Instead, Olig2-occupied loci in these astrocytes displayed GC-rich, TC-rich, and AG-rich repeat motifs (Supplementary Fig. 15). To determine whether the motifs enriched in astrocytes are unique, we conducted additional experiments using the same CUT&Tag methodology to investigate Olig2 binding sites in oligodendrocyte lineage cells, a glial cell type that naturally expresses Olig2 in the normal adult mouse brain. We collected tdT+ cells from adult Sox10-CreERT2; Ai14 transgenic mice via FACS, and identified peaks along with their binding distribution after sequencing (Supplementary Fig. 16a–c). We detected a GC-enriched motif similar to that in astrocytes (such as Sp and Klf genes), as well as the previously reported canonical E-box CAGCTG motif (Supplementary Fig. 16d, e). Together, these results suggested that Olig2 may have some unique regulatory mechanisms in astrocytes undergoing Ngn2-mediated reprogramming.
a Experimental workflow of CUT&Tag analysis. Created in BioRender.com. Hou, KY. (https://BioRender.com/yspzfyr). b TdT+ cells were collected by FACS, and about 60,000 sorted cells from 12 mice were pooled together as one sample for an individual experiment. There are two replicates in our study. c Heatmap of Olig2-binding signals in Ngn2-overexpressing astrocytes. d Pie chart illustrating the genomic location of CUT&Tag peaks. e GO analysis of biological processes corresponding to Olig2-targeted genes in Ngn2-overexpressing astrocytes. f Integrative analysis of Olig2 targeting genes from CUT&Tag and DEGs from scRNA-seq. Notably, 90 downregulated DEGs and 201 upregulated DEGs, a total of 291 overlapping genes, were found to be occupied by astrocytic Olig2. g Heatmap showing the different expression levels of the 291 overlapping genes in tdT+ astrocytes between the Ngn2+shNC and the Ngn2+shOlig2 groups. h Representative Olig2 binding profiles on Ngn2 gene locus in Ngn2-overexpressing astrocytes. i RT-qPCR showing higher relative Ngn2 mRNA expression level with Olig2 knockdown; n = 4 mice for Ngn2 group, n = 6 mice for Ngn2+shNC and Ngn2+shOlig2 groups, one-way ANOVA followed by Bonferroni’s post hoc test, mean ± SD. j Confocal images showing Ngn2 expression levels between different groups. Scale bar, 20 μm. k Quantified data showing enhanced Ngn2 intensity after Olig2 knockdown in Ngn2-overexpressing astrocytes; n = 5 mice per group, one-way ANOVA followed by Bonferroni’s post hoc test, mean ± SD. l Work model of Ngn2 and Olig2 interaction: Ngn2 activates Olig2, which in turn inhibits Ngn2 expression and AtN conversion. Created in BioRender.com. Hou, KY. (https://BioRender.com/1usr45d).
To further explore the unique features of Olig2-targeting regions in Ngn2-transduced astrocytes, we compared the Olig2 occupancy profiles in these astrocytes with those in mouse embryonic stem cells (mESCs)33, OPCs23 and Sox10-tdT+ cells, respectively (Supplementary Fig. 17a). Our analysis revealed that there was only a small portion of overlapping genes between astrocytes and ESCs (1180 shared vs 7192 unique), which were primarily associated with functions related to axonogenesis, synapse organization, and forebrain development (Supplementary Fig. 17b). In contrast, the genes unique to Ngn2-transduced astrocytes were more enriched in the regulation of apoptotic signaling pathway, immune cell migration and tissue development, although the enrichment was less significant (Supplementary Fig. 17c). Additionally, the ratio of the shared genes between astrocytes and OPCs was also small (1376 shared vs 6996 unique), with the top GO biological process terms including axonogenesis, synapse organization and regulation of membrane potential (Supplementary Fig. 17d). Genes unique to astrocytes compared to OPCs were notably enriched in response to cell-substrate adhesion, stem cell population maintenance and maintenance of cell number (Supplementary Fig. 17e). Finally, we found that more genes shared between adult astrocytes and Sox10-tdT+ oligodendrocyte lineage cells (2526 shared vs 5846 unique). The overlapping genes were primarily enriched in pathways related to synaptic organization and signal transduction (Supplementary Fig. 17f), whereas the genes unique to astrocytes showed highly close association with axonogenesis and regulation of membrane potentials (Supplementary Fig. 17g). Together, these results reveal that Olig2 may play an important gene regulatory role during astrocyte reprogramming.
To further investigate whether the genes targeted by Olig2 undergo transcriptional changes, we conducted the integrative analysis of the scRNA-seq transcriptome data and CUT&Tag-seq gene binding loci data of Olig2. We discovered that 90 Olig2-targeted genes exhibited downregulated expression after Olig2 knockdown, while 201 Olig2-targeted genes upregulated after reducing the expression of Olig2 (Fig. 8f, g). For the upregulated targets, there was a group of genes related to neuronal development, including Nes, Vim, Gria3, and Pou3f2. Notably, Hes5, a transcriptional repressor of genes that require bHLH proteins for transcription and plays a negative regulator of neurogenesis32, was downregulated following Olig2 knockdown (Fig. 8g). Stub1 (also known as CHIP) acts as an E3 ubiquitin ligase that regulates the ubiquitination and degradation of specific proteins (such as GPX4), promoting ferroptosis34. In Ngn2-transduced astrocytes, Olig2 knockdown reduced Stub1 expression (Fig. 8g), supporting the idea that AtN reprogramming is regulated by programmed cell death31. GO analysis indicated that the downregulated genes were enriched in processes related to steroid metabolism, synaptic maintenance and organization, and actin binging (Supplementary Fig. 18a). Meanwhile, the upregulated genes were mainly related to actin filament organization, intrinsic component of organelle membrane and transcription coregulatory activity (Supplementary Fig. 18b). Overall, the integrated analysis of CUT&Tag and scRNA-seq suggests that Olig2 plays a key regulatory role in the expression of genes involved in Ngn2-mediated astrocyte reprogramming.
Ngn2 acts as a pioneer transcription factor that reprograms astrocytes into functional neurons. Interestingly, we found that Olig2 in astrocytes can directly target the Ngn2 gene (Fig. 8h), potentially revealing a regulatory link. The prominent genomic binding peak of Olig2 within the Ngn2 exon region motivated us to explore the possibility of the direct regulation of Olig2 to the expression of Ngn2 (Fig. 8h), given that Olig2 typically functions as a transcription repressor25,35. qRT-PCR data revealed that Ngn2 mRNAs were significantly increased after knocking down Olig2 in AAV9 transduced cortical regions (Fig. 8i). Next, the expression of Ngn2 was examined through immunostaining in mice injected with AAV-Ngn2, Ngn2+shNC, and Ngn2+shOlig2 at 7 dpi. As expected, the Ngn2 protein was specifically expressed in cortical astrocytes (tdT+) in all groups (Fig. 8j). Noteworthy, the fluorescence intensity of Ngn2 almost doubled after Olig2 knockdown (Fig. 8k), suggesting that removing Olig2 increases Ngn2 expression in astrocytes. Together, these results suggest a negative feedback of the ectopic expression of Ngn2 in the astrocytes, which induces the Olig2 expression to inhibit Ngn2 expression and neuronal reprogramming (Fig. 8i).
Discussion
In this study, we demonstrate that Olig2 works as an inducible barrier during Ngn2-mediated AtN conversion. We first found that Olig2 was upregulated in cortical astrocytes when initiating AtN conversion by the forced expression of conversion pioneer factors Ngn2, Ascl1, and NeuroD1. Knockdown of astrocytic Olig2 promotes the efficiency of Ngn2-mediated AtN conversion and generated more functional neurons in the mouse brain. scRNA-seq studies show that Olig2 acts as a repressor of AtN conversion, and knockdown of Olig2 in Ngn2-overexpressing astrocytes not only further upregulates neuronal genes but also further downregulates astrocyte identity genes. CUT&Tag analysis shows that Olig2 in astrocytes binds to many gene loci involved in regulating cell morphogenesis, development, and multicellular biological processes. Therefore, this study identifies Olig2 as an inducible barrier to AtN conversion and reveals a regulatory mechanism underlying the Ngn2-mediated astrocyte reprogramming.
In this study, we used the GFAP promoter to drive the overexpression of a neural regeneration transcription factor in cortical astrocytes, which is commonly used for specific gene expression in astrocytes36,37. However, its application in AtN reprogramming may present challenges, including potential cis-regulatory effects leading to non-specific expression38. Notably, while some neural transcription factors, such as Ascl1, NeuroD1, and Pax6, have been shown to cause non-specific expression in endogenous neurons, the leakage of expression from Ngn2 is relatively low38,39. Our results corroborate this, as we observed negligible fluorescence in endogenous neurons following 7 days of Ngn2 overexpression, suggesting that Ngn2 is unlikely to cause significant leakage in endogenous neurons. Moreover, we provide some evidence for the reprogramming of astrocytes to neurons, addressing concerns regarding intermediate or immature neuronal phenotypes. Our immunofluorescence results showed increased double-negative AAV transduced cells (tdT+/S100β−/NeuN−) and upregulation of neural stem cell markers, such as Nestin and Vimentin, in the Ngn2+shOlig2 group. These observations were further supported by single-cell RNA sequencing, which demonstrated a marked downregulation of astrocyte-specific genes (e.g., Slc1a2, S100β, Aqp4) and upregulation of neurogenic genes (e.g., Nes, Sox4, Pax6). Moreover, transcriptomic analysis of mitochondrial energy metabolism further suggested a transition from astrocyte-like to neuron-like metabolic signatures. From a functional perspective, electrophysiological analysis revealed that 18.2% of reprogrammed neurons fired only a single action potential, and nearly half lacked detectable synaptic activity, resembling immature neurons rather than endogenous mature neurons. We also employed astrocyte lineage tracing mice to pre-label astrocytes, a widely recognized “gold standard” technique for assessing AtN reprogramming4,40. These results further demonstrated that Olig2 knockdown in astrocytes significantly enhances the efficiency of Ngn2-mediated AtN reprogramming and provide a foundation for future studies exploring this process.
In addition, as astrocyte reprogramming progresses, the GFAP promoter may gradually lose activity in reprogrammed astrodytes41,42, potentially hindering long-term tracking of converted neurons. GFAP-driven fluorescent protein expression follows the central dogma, involving transcription and translation. While GFAP promoter activity may decrease during astrocyte-to-neuron reprogramming, it does not vanish immediately. Consequently, the fluorescent reporter gene’s transcripts and proteins persist for some time before degradation. Studies have shown that the lifespan of some proteins in cells can range from days to months43,44,45. Furthermore, immunofluorescence staining amplifies antigen signals, allowing for tracking of reprogrammed cells within a certain time window despite the gradual loss of GFAP promoter activity. In the future, technologies such as barcode labeling could further enable long-term, stable tracking of target cells throughout the reprogramming process.
AtN conversion offers a promising approach in neuroregenerative medicine. Different from the transplantation of external cells, in vivo neuronal reprogramming is an endogenous repair technology using local glial cells in the CNS for neuronal regeneration46. Proneural transcription factors such as Ngn2, Ascl1, and NeuroD1 were frequently selected as pioneer factors to drive AtN conversion4,5. In this study, we first found that Olig2 was substantially increased upon overexpressing Ngn2, Ascl1, and NeuroD1 in cortical astrocytes. Olig2 is a member of the bHLH family transcription factors, expressing primarily in oligodendrocyte lineage cells but absent in astrocytes in the adult mouse cortex under normal conditions24,25,47. Transient upregulation of Olig2 in astrocytes has been reported in the context of injury, which is required for reactive astrocyte proliferation19. Here, we found that bHLH proneural TFs-induced AtN conversion resulted in upregulation of Olig2 in cortical astrocytes, which has never been reported in previous studies of astrocyte reprogramming. Unlike transient upregulation caused by injury conditions19, our study observed long-term expression of Olig2 in Ngn2-overexpressing astrocytes, suggesting that Olig2 can produce a lasting impact on AtN conversion.
Direct lineage reprogramming entails a profound transformation in cellular identity, wherein the transcriptomic landscape undergoes significant alterations, marked by differential expression of a substantial array of cell-type-specific genes. Despite the mechanistic heterogeneity of the proneural TFs Ngn2, Ascl1, and NeuroD1 in promoting AtN conversion, a unifying feature of these factors is their ability to transcriptionally upregulate the expression of neuron-specific genes and simultaneously repress astrocyte-associated gene programs8,12,48,49. Olig2 is a bHLH transcription factor involved in regulating cell fate decisions23,24,50, suggesting a potential role in astrocyte reprogramming to neurons. However, the differential expression of Olig2 has not been mentioned in the previous in vitro AtN conversion studies, probably because astrocytes cultured in vitro cannot fully recapitulate their counterparts in vivo51. Most in vitro AtN conversion experiments used the primary astrocytes isolated from the neonatal rodent cortices. During brain development, Olig2 is transiently expressed in developing cortical astrocytes in the neonatal stage, and then gradually downregulates in the adult astrocytes52. In addition, fetal bovine serum (FBS) is widely used as a growth supplement to maintain the astrocyte culture, and growth factors in FBS have been reported to induce Olig2 expression in the cultured astrocytes53. Therefore, cultured astrocytes may contain high background levels of Olig2 expression, which in turn results in the inability to detect further upregulation of Olig2. Collectively, adult cortical astrocytes lacking Olig2 expression are a suitable model to investigate the inducible inhibitory role of Olig2 in the reprogramming of astrocytes.
The bHLH family transcription factors can recognize a DNA motif called E-box and then regulate gene transcription, control essential biological processes such as cell growth, proliferation, and metabolism, neurogenesis, and myogenesis54. As a bHLH transcription factor, Olig2 can recognize and bind to E-box sequences in the promoter or enhancer regions of specific genes, thereby directly regulating the expression of these genes25,55. This regulatory effect can be either activating or inhibitory, depending on the interaction between Olig2 and other transcription factors, co-activators, or co-repressors, as well as the cellular environment56,57. Previous studies have reported that Olig2 functions as a neurogenesis inhibitor in the adult mammalian brain and that antagonizing Olig2 function promotes the reprogramming of reactive glia into neurons16,58. In this study, CUT&Tag analysis reveals that neurogenic genes, such as Ngn2 and Pou3f2, can be targeted by Olig2 in astrocytes. Knockdown of Olig2 in astrocytes significantly increased Ngn2 expression, possibly due to activation of endogenous Ngn2 gene transcription, which is critical for promoting astrocyte reprogramming8,59. Alternatively, since Olig2 has been previously shown to regulate the expression of GFAP52, it may also influence Ngn2 expression in the AAV-GFAP::Ngn2 vector by modulating the activity of the GFAP promoter. Consistent with this idea, we observe an enhancement of Ngn2-mediated AtN conversion when knocking down or antagonizing Olig2 in cortical astrocytes. Furthermore, Olig2 often forms homodimeric or heterodimeric complexes with other bHLH transcription factors, such as Ngn2, thereby synergistically regulating the expression of specific genes25,60. Therefore, the genomic binding profile of Olig2 in Ngn2-overexpressing astrocytes has many different binding sites compared with previous studies in oligodendrocyte lineage cells, possibly due to the presence of Ngn2 affecting DNA binding sites for Olig2 in astrocytes. Notably, the formation of Olig2-Ngn2 heterodimers is regulated by protein phosphorylation, thereby directing cell fate choice and controlling cell proliferation in developing CNS50,61. Recently, a research group has reported a phosphorylated mutant form of Ngn2 (PmutNgn2) that is resistant to phosphorylation and exhibits higher reprogramming efficiency and enhanced epigenetic remodeling activity8,15. This observation may be attributed to a profound alteration in the binding affinity of PmutNgn2 for Olig2, thereby increasing AtN conversion efficiency. Thus, it remains elusive and will be interesting to explore whether PmutNgn2 affects its binding to Olig2 and thus contributes to the AtN conversion efficiency.
In vivo AtN conversion has shown promise in animal models for CNS repair; however, its therapeutic potential is constrained by suboptimal reprogramming efficiencies, highlighting the need for further optimization to facilitate clinical translation4,5. In general, the efficiency of AtN conversion is modulated by two primary determinants: promotional factors that enable cellular reprogramming (energy), and inhibitory factors that resist conversion (barrier). For example, modification of proneural transcription factors8,15,62 or enhancement of their expression has been shown to significantly improve AtN conversion efficiency63. Additionally, removal of the internal obstacles to astrocyte reprogramming can also substantially improve reprogramming efficiency, such as downregulation of REST12, Notch signaling pathway inhibition3,13, epigenetic remodeling (histone deacetylase inhibitor, etc.)51, and suppression of unfolded protein response by using small molecules15. Notably, these internal barriers to reprogramming pre-exist in astrocytes, similar to the nonspecific “innate” immune system, which serves as a first line of defense against cell fate alteration. In the present study, we have identified that astrocytic Olig2 also serves as a barrier to AtN conversion, but in a completely different manner. In adult mouse brains, Olig2 is typically absent in cortical astrocytes, but is strongly upregulated during reprogramming induced by bHLH proneural transcription factors (Ngn2, Ascl1, and NeuroD1). However, non-bHLH reprogramming factors (Sox2 and Dlx2) did not show the ability to upregulate Olig2 in astrocytes. This observation suggests that the induction of Olig2 may be specifically linked to bHLH transcription factor mediated reprogramming. However, it is important to note that Olig2 induction could also reflect a reactive state, as is observed in astrocytes during injury19. In this case, astrocytic Olig2 may play a role similar to a reactive response, thereby inhibiting neurogenesis16. It should be emphasized that our study does not mean that there are no inducible barriers to AtN conversion mediated by non-bHLH pioneer factors, but rather that specific induced barriers may exist, which requires further study.
Although we show here the strong upregulation of Olig2 during Ngn2-mediated AtN conversion and provide evidence that Olig2 works as a reprogramming barrier, we acknowledge several limitations in this study. First, the precise molecular mechanism underlying how Olig2 is induced by bHLH proneural TFs ectopic expression in cortical astrocytes remains to be further elucidated. It has been shown that DNA methylation is involved in suppressing Olig2 expression in acute myeloid leukemia64. Interestingly, previous studies have demonstrated that Ngn2 has the ability to demethylate DNA8,65. Therefore, DNA methylation analysis using Methyl-HiC may help to identify the molecular mechanism of Olig2 upregulation during Ngn2-mediated AtN conversion. Second, Olig2 is a bHLH transcription factor that can regulate gene expression by forming homodimers or heterodimers with other bHLH family members, including Ngn2. In addition, the phosphorylation status can significantly affect the preference of Olig2 dimer formation, thereby affecting cell fate determination50,60. However, whether these influencing factors are involved in the inhibitory effect of Olig2 on Ngn2-mediated AtN conversion has not been addressed in this study. Third, although we compared the transcriptomic differences between astrocytes overexpressing Ngn2 and those not infected with AAV, providing insight into the transcriptomic changes induced by Ngn2, the impact of tdTomato overexpression (AAV-GFAP::tdTomato) on the astrocyte transcriptome remains unknown and needs to be included in future studies. Fourth, our scRNA-seq only tracked the state of neural stem cells, but did not capture the state of neurons. In order to fully reveal the molecular mechanism of the entire AtN conversion, more scRNA-seq studies will be included. Last, although Olig2 knockdown significantly enhances Ngn2-mediated AtN conversion, the efficiency remains low, with only about 12% in lineage-tracing mouse cortex. These results highlight the need for additional strategies to further improve AtN efficiency. Future approaches may include the combined application of neurogenic transcription factors39, phosphorylation modification8, inhibition of astrocytic Notch signaling3, and the removal of epigenetic barriers that impede reprogramming17.
In summary, we characterize an inducible barrier in the process of lineage reprogramming. Extensive alterations in gene networks and signaling pathways have been observed during cellular reprogramming, and Olig2 may be just the tip of the iceberg. Thus, our study not only discovered a factor that hinders astrocyte reprogramming in vivo but, more importantly, expanded the theoretical scope of cell reprogramming barriers.
Methods
Animals
Experiments were conducted on adult (age 2–3 months) wild-type C57BL/6 J mice (Guangdong Yaokang Biotechnology, China) and transgenic mice (The Jackson laboratory). The transgenic mouse strains used in this study include: Sox10-CreERT2 [CBA;B6-Tg(Sox10-icre/ERT2)388Wdr/J, stock number: 027651], Aldh1l1-CreERT2 [B6N.FVB-Tg (Aldh1l1-cre/ERT2)1Khakh/J, stock number: 031008], and Ai14 [B6.Cg Gt(ROSA)26Sortm14(CAG-tdTomato)Hze/J, stock number:007914]. Sox10-CreERT2 mice or Aldh1l1-CreERT2 mice were crossed with the Ai14 mice for cell type-specific lineage tracing. Both male and female mice were used in this study. All mice were housed under a controlled temperature and a 12-h light/dark cycle with free access to water and food in the animal facility. For all surgical experiments, mice were anesthetized using isoflurane. Anesthesia was initiated at 5% and then reduced to 2% to maintain a stable deep anesthetic state. Animal procedures and protocols were approved by the Laboratory Animal Ethics Committee of Jinan University, China (approval no. IACUC-20220613-05 and 20230703-12).
Virus preparation and injection
Adeno-associated virus (AAV) was used to deliver transcription factors into the mouse cortex. tdTomato was inserted into the AAV vector to generate AAV-tdTomato, and Neurogenin2/Ascl1/NeuroD1 was cloned into the AAV-tdTomato to generate AAV-Ngn2/Ascl1/NeuroD1-tdTomato. Sox2 or Dlx2 was cloned into the AAV-GFP to generate AAV-Sox2/Dlx2-GFP. The transactivation domain of herpes simplex VP16 was fused to bHLH domain of Olig2 to generate AAV-Olig2VP16. Three designed Olig2 shRNA were cloned into one AAV backbone (Olig2 shRNA1: gcacttgtggaatggaatattggatccaatattccattccacaagtgc; Olig2 shRNA2: ggattatggggagtcgaatttggatccaaattcgactccccataatcc; Olig2 shRNA3: gcgagcacctcaaatctaattggatccaattagatttgaggtgctcgc). All transcription factors were under the control of the human GFAP promoter, and shRNA was driven by U6 promoter. Human GFAP promoter (hGFAP) is a synthetic promoter that is composed of GfaABCD1 derived of human GFAP promoter gfa2 enhancer element, derived from the human glial fibrillary acidic protein gene. All recombinant AAVs were produced by PackGene® Biotech, purified through iodixanol gradient ultracentrifuge and subsequent concentration. Purified AAV viruses were tittered using a quantitative PCR-based method. AAV (1 μL) was injected stereotactically into both sides of the mouse cortex with titers of 0.5–2 × 1011 GC mL−1 (transcription factors) and 2 × 1010 GC mL−1 (shRNA). Equal ratios of viruses were mixed for multifactor injections. Injection coordinates are as follows: anterior/posterior, 1.3 mm; medial/lateral, ±1.5 mm; and dorsal/ventral, −0.7 mm. Brain slices with tdT expression within approximately 400 μm surrounding the injection site were included for analysis. For AAV9-Ngn2 (lacking a reporter gene) analysis, we performed serial sectioning and immunofluorescence staining for Ngn2 on every fifth section to confirm Ngn2 expression, and the intervening brain sections were included for conversion validation.
Retrograde labeling and intracerebral injection
The use of AAV to pre-label endogenous neurons was based on a previous study38. Briefly, AAV2-retro-CAG-GFP virus (2 µL per mouse) was injected into the T6 level of the dorsal spinal cord to target the corticospinal tract. Ten days later, we stereotaxically injected AAV9-Ngn2 with shNC or shOlig2 into the cortex at these coordinates: −0.11 mm AP, ±1.5 mm ML, and 0.7 mm DV. Analysis was conducted 45 days after the cortical injection.
Tissue dissociation
Mice were sacrificed, perfused with aCSF, and the virus-infected brain tissue was collected. The brain was dissociated into single-cell suspension using the Papain Dissociation System (Worthington, LK003150) according to the manufacturer’s protocol. Myelin was removed using a debris removal solution (Miltenyi Biotec, 130-109-398) according to the manufacturer’s instructions. For fluorescent-assisted cell sorting (FACS, BD FACS Aria SORP), cells were resuspended in 1× PBS supplemented with 1% BSA and 2 mM EDTA, and kept at 4 °C until sorting. For scRNA-seq, cells were resuspended in 1× PBS.
CUT&Tag and data analysis
For astrocytic Olig2 profiling, tdTomato+ cells were sorted from the cortical tissues of animals (two samples, 12 mice per sample) 7 days after AAV9-GFAP::Ngn2-tdTomato injection using FACS (BD FACSAria SORP). For Olig2 profiling in oligodendrocyte lineage cells, tdTomato+ cells were sorted from the brains of 3-month-old Sox10-CreERT2; Ai14 transgenic mice (two samples, 2 mice per sample) using FACS. Cells were first gated for singlets based on Forward Scatter–Area (FSC-A) versus Forward Scatter–Height (FSC-H), then live cells were selected by excluding DAPI-positive dead cells, and finally, the brightest tdT+ cells were gated (10.0% of total tdT+ cells). The sorted cells were performed with CUT&Tag-IT™ Assay Kit (Active Motif, catalog No. 53160) to get the final CUT&Tag libraries, followed by Illumina NovaSeq. Rabbit anti-Olig2 (Millipore, catalog No. AB9610) was used as the primary antibody. Before read mapping, clean reads were obtained from the raw reads by removing the adaptor sequences. The sequencing data were mapped to the mm10 genome assembly, and peak calling was performed using MACS2 software with a cutoff value < 0.05. The deep tool was used for showing read distributions. A unified peak atlas was generated through the integration of aligned reads (BAM files) derived from two biological samples. Peaks were annotated by the bedtools, and the HOMER’s findMotifsGenome.pl tool was used for Motif analysis.
ScRNA-seq and data analysis
Eighteen, twenty, and twenty-two days after virus injection (n = 2 for each time point), the single-cell suspensions were directly loaded onto a microfluidic chip (GEXSCOPETM Single Cell RNA-seq Kit, Singleron Biotechnologies) and processed for scRNA-seq library preparation according to the manufacturer’s protocol. Raw reads were processed to generate gene expression profiles using CeleScope v1.5.2. Scanpy v1.8.1 was used for quality control, dimensionality reduction, and clustering under Python 3.7. Low quality and duplet cells were removed before further analysis. Cells with less than 200 or with top 2% gene counts were removed. Cells with top 2% UMIs were excluded. Cells with mitochondrial content > 20% were filtered. Cells with genes detected in less than 5 cells were removed. Total 64,534 cells were retained for the downstream analyses, with an average of 972 genes and 1836 UMIs per cell. The raw count matrix was normalized by total counts per cell and logarithmically transformed into normalized data matrix. The top 2000 variable genes were chosen for principal component analysis (PCA), and top 20 principle components were used for clustering and dimensional reduction. Cell clusters were visualized by using Uniform Manifold Approximation and Projection (UMAP). All single-cell analysis results in this study do not incorporate data from other publications.
CytoTRACE analysis
CytoTRACE (Cellular Trajectory Reconstruction Analysis using gene Counts and Expression) is a computational framework designed to infer cellular differentiation potential and reconstruct developmental trajectories from scRNA-seq data29. CytoTRACE first calculates the number of detectably expressed genes per cell. It then identifies genes whose expression patterns correlate with gene counts to derive the Gene Count Signature (GCS). Finally, it employs a local similarity algorithm between cells and applies an iterative two-step smoothing procedure to refine the GCS. By leveraging gene expression heterogeneity, CytoTRACE predicts the relative “stemness” or differentiation status of individual cells, enabling the identification of progenitor-like cells and lineage hierarchies within heterogeneous populations.
Quantitative real-time PCR
Seven days or twenty days after virus injection, brain tissue was dissociated into single-cell suspension, and tdTomato positive cells were collected using FACS. RNA samples were harvested using FlysisAmp Cells-to-CT 1-Step SYBR Green Kit (Vazyme, China) according to the manufacturer’s instructions. QPCR was performed using CFX Duet Real-Time PCR System (Bio-Rad). Relative expression was calculated as 2(−ΔΔCT). The primers for targeted genes were as follows: Olig2 (F: TTACAGACCGAGCCAACACC, R: TGGCCCCAGGGATGATCTAA), Nestin (F: CCTTCTCTAGTGCTCCACGTC, R: CGTCGATTGAGCTCCCACA), Vimentin (F: GCGAGAGAAATTGCAGGAGG, R: CCGTTCAAGGTCAAGACGTG), Olig1 (F: CTCGCCCAGGTGTTTTGTTG, R: TATAAGCCTGCGCTACGACG) and Gapdh (F: AGGTCGGTGTGAACGGATTTG, R: TGTAGACCATGTAGTTGAGGTCA).
Electrophysiology
Brain slices were prepared 40-45 days after virus injection and cut to 300 μm thick coronal sections with a Leica vibratome (VT-1200S) in ice-cold cutting solution (containing 75 mM sucrose, 85 mM NaCl, 2.5 mM KCl, 0.5 mM CaCl2, 4 mM MgCl2, 24 mM NaHCO3, 1.25 mM NaH2PO4, and 25 mM glucose). Slices were transferred to holding solution (containing 93 mM NMDG, 2.5 mM KCl, 1.2 mM NaH2PO4, 30 mM NaHCO3, 20 mM HEPES, 25 mM glucose, 5 mM sodium ascorbate, 2 mM Thiourea, 3 mM sodium pyruvate, 10 mM MgSO4⋅7H2O, 0.5 mM CaCl2, pH 7.3 adjusted with HCl, 300–310 mOsm L−1), with continuous 95% O2 and 5% CO2 bubbling, first at 34 °C for 30 min, and then at room temperature. Whole-cell recordings were performed using Multiclamp 700B patch-clamp amplifier (Molecular Devices, Palo Alto, CA), and the slices were maintained in the artificial cerebral spinal fluid (aCSF) containing 126 mM NaCl, 2.5 mM KCl, 1.25 mM NaH2PO4, 26 mM NaHCO3, 2 mM MgCl2, 2 mM CaCl2, and 10 mM glucose. The pH of bath solution was adjusted to 7.3 with HCl, and osmolarity at 300–310 mOsm L−1. Patch pipettes were pulled from borosilicate glass (~5–8 MΩ) and filled with a pipette solution consisting of 126 mM K-Gluconate, 4 mM KCl, 10 mM HEPES, 4 mM Mg2ATP, 0.3 mM Na2GTP, 10 mM Phospho-Creatnine (pH 7.3 adjusted with KOH, 290 mOsm L−1). Data were collected using pClamp 10 and Clampex 10.4 software (Molecular Devices, Palo Alto, CA), sampled at 10 kHz and filtered at 3 kHz, analyzed with Clampfit 10.4.
Immunofluorescence
Mice were deeply anesthetized with 2.5% Avertin and perfused transcardially with ice-cold artificial cerebrospinal fluid (aCSF), followed by post-fixation in 4% PFA overnight at 4 °C in darkness. Then, the samples were cut into 30-μm sections by a vibratome (Leica, VTS1200). Brain slices were washed three times in 1×PBS for 10 min each. Blocking was performed for 2 h in 0.3% triton PBS + 5% donkey serum. Primary antibody was diluted in 0.05% triton PBS + 5% donkey serum and incubated at 4 °C for two nights. After washing three times with PBS, the samples were incubated with secondary antibodies conjugated to Alexa Flour 488, 555, or 647 (1:500, Jackson ImmunoResearch) for 2 h at room temperature, followed by extensive washing in PBS. The secondary antibody was diluted in 0.05% triton PBS + 5% donkey serum. Sections were counterstained with DAPI and mounted in fluorescent mounting media (DAKO). Images were taken with a Zeiss LSM880 confocal microscope.
Primary antibodies used were listed as follows: rabbit anti-GFAP (1:1000, Cat# AB9610, Millipore), guinea pig anti-Olig2 (1:1000, Cat# OB-PGP040-01, Oasis Biofarm), rat anti-RFP (1:1000, Cat# 5F8, Chromotek), rat anti-GFAP (1:1000, Cat# 13-0300, Invitrogen), rabbit anti-S100β (1:1000, Cat# ab52642, Abcam), rabbit anti- Ascl1 (1:1000, Cat# ab211327, Abcam), rabbit anti-ND1 (1:1000, Cat# AB205300, Millipore), mouse anti-Aldh1l1 (1:500, Cat# AB10673448, NeuroMab), guinea pig anti-NeuN (1:1000, Cat# ABN90, Millipore), rabbit anti-Ngn2 (1:1000, Cat# PA5-78556, Invitrogen), guinea pig anti-Sox10 (1:1000, Cat# OB-PGP001-01, Oasis Biofarm), rabbit anti-Iba1 (1:1000, Cat# 019-19741, Wako), rabbit anti-Sox9 (1:1000, Cat# AB5535, Millipore), rabbit anti-Sox2 (1:1000, Cat# AB5603, Millipore), rabbit anti-Dlx2 (1:1000, Cat# ab135620, Abcam), rabbit anti-NG2 (1:500, Cat# AB5320, Millipore), guinea pig anti-CC1 (1:1000, Cat# OB-PRT039-01, Oasis Biofarm), mouse anti-Nestin (1:500, Cat# Rat-401, DHSB), rabbit anti-Vimentin (1:1000, Cat# ab92527-1002, Abcam), chicken anti-GFP (1:2000, Cat# ab13970, Abcam), mouse anti-VP16 (1:500, Cat# sc-7545, Santa Cruz), rabbit anti-CamkII (1:500, Cat# ab5683, Abcam), rabbit anti-GABA (1:500, Cat# A2052, Sigma), rabbit anti Sfxn5 (1:1000, Cat# ab172971, Abcam), guinea pig anti-DCX (1:1000, Cat# AB2253, Sigma).
Statistics
Statistical analysis was performed with GraphPad Prism 8 and Origin2021. Statistical significance was tested with the unpaired two-sided Student’s t-test and One-Way ANOVA test followed with the Bonferroni’s post hoc test. Significance was determined at a p value ≤ 0.05, and data were presented as mean ± SD.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNA-seq data generated in this study have been deposited in the Gene Expression Omnibus database under accession code GSE289603. The CUT&Tag data generated in this study have been deposited in the Gene Expression Omnibus database under accession code GSE289604 and GSE308070 for astrocytes. The CUT&Tag data generated in this study have been deposited in the Gene Expression Omnibus database under accession code GSE300369 for oligodendrocyte lineage cells. All of the plasmids used in this study are available from the lead contact upon request. All data generated in this study are provided in the Source Data file and. Source data are provided with this paper.
References
Frisen, J. Neurogenesis and gliogenesis in nervous system plasticity and repair. Annu. Rev. Cell Dev. Biol. 32, 127–141 (2016).
Sirko, S. et al. Reactive glia in the injured brain acquire stem cell properties in response to sonic hedgehog. Cell Stem Cell 12, 426–439 (2013).
Zamboni, M., Llorens-Bobadilla, E., Magnusson, J. P. & Frisen, J. A widespread neurogenic potential of neocortical astrocytes is induced by injury. Cell Stem Cell 27, 605–617 (2020).
Bocchi, R., Masserdotti, G. & Gotz, M. Direct neuronal reprogramming: fast forward from new concepts toward therapeutic approaches. Neuron 110, 366–393 (2022).
Qian, H. & Fu, X. D. Brain repair by cell replacement via in situ neuronal reprogramming. Annu. Rev. Genet. 55, 45–69 (2021).
Guo, Z. et al. In vivo direct reprogramming of reactive glial cells into functional neurons after brain injury and in an Alzheimer’s disease model. Cell Stem Cell 14, 188–202 (2014).
Russo, G. L. et al. CRISPR-mediated induction of neuron-enriched mitochondrial proteins boosts direct glia-to-neuron conversion. Cell Stem Cell 28, 524–534 e527 (2021).
Pereira, A. et al. Direct neuronal reprogramming of mouse astrocytes is associated with multiscale epigenome remodeling and requires Yy1. Nat. Neurosci. 27, 1260–1273 (2024).
Holmberg, J. & Perlmann, T. Maintaining differentiated cellular identity. Nat. Rev. Genet. 13, 429–439 (2012).
Wang, H., Yang, Y., Liu, J. & Qian, L. Direct cell reprogramming: approaches, mechanisms and progress. Nat. Rev. Mol. Cell Biol. 22, 410–424 (2021).
Srivastava, D. & DeWitt, N. In vivo cellular reprogramming: the next generation. Cell 166, 1386–1396 (2016).
Masserdotti, G. et al. Transcriptional mechanisms of proneural factors and REST in regulating neuronal reprogramming of astrocytes. Cell Stem Cell 17, 74–88 (2015).
Magnusson, J. P. et al. A latent neurogenic program in astrocytes regulated by Notch signaling in the mouse. Science 346, 237–241 (2014).
Tan, Z. et al. NOTCH1 signaling regulates the latent neurogenic program in adult reactive astrocytes after spinal cord injury. Theranostics 12, 4548–4563 (2022).
Sonsalla, G. et al. Direct neuronal reprogramming of NDUFS4 patient cells identifies the unfolded protein response as a novel general reprogramming hurdle. Neuron 112, 1117–1132.e9 (2024).
Buffo, A. et al. Expression pattern of the transcription factor Olig2 in response to brain injuries: implications for neuronal repair. Proc. Natl. Acad. Sci. USA 102, 18183–18188 (2005).
Kremer, L. P. M. et al. DNA methylation controls stemness of astrocytes in health and ischaemia. Nature 634, 415–423 (2024).
McKenzie, I. A. et al. Motor skill learning requires active central myelination. Science 346, 318–322 (2014).
Chen, Y. et al. The basic helix-loop-helix transcription factor olig2 is critical for reactive astrocyte proliferation after cortical injury. J. Neurosci. 28, 10983–10989 (2008).
Niu, W. et al. In vivo reprogramming of astrocytes to neuroblasts in the adult brain. Nat. Cell Biol. 15, 1164–1175 (2013).
Wu, Z. et al. Gene therapy conversion of striatal astrocytes into GABAergic neurons in mouse models of Huntington’s disease. Nat. Commun. 11, 1105 (2020).
Zhang, Y. et al. A single factor elicits multilineage reprogramming of astrocytes in the adult mouse striatum. Proc. Natl. Acad. Sci. USA 119, e2107339119 (2022).
Yu, Y. et al. Olig2 targets chromatin remodelers to enhancers to initiate oligodendrocyte differentiation. Cell 152, 248–261 (2013).
Zhang, K. et al. The oligodendrocyte transcription factor 2 OLIG2 regulates transcriptional repression during myelinogenesis in rodents. Nat. Commun. 13, 1423 (2022).
Meijer, D. H. et al. Separated at birth? The functional and molecular divergence of OLIG1 and OLIG2. Nat. Rev. Neurosci. 13, 819–831 (2012).
Smith, D. K., Yang, J., Liu, M. L. & Zhang, C. L. Small molecules modulate chromatin accessibility to promote NEUROG2-mediated fibroblast-to-neuron reprogramming. Stem Cell Rep. 7, 955–969 (2016).
Heinrich, C. et al. Directing astroglia from the cerebral cortex into subtype specific functional neurons. PLoS Biol. 8, e1000373 (2010).
Hack, M. A. et al. Neuronal fate determinants of adult olfactory bulb neurogenesis. Nat. Neurosci. 8, 865–872 (2005).
Gulati, G. S. et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 367, 405 (2020).
Fu, X. Q., Zhan, W. R., Tian, W. Y., Zeng, P. M. & Luo, Z. G. Comparative transcriptomic profiling reveals a role for Olig1 in promoting axon regeneration. Cell Rep. 43, 114514 (2024).
Gascon, S. et al. Identification and successful negotiation of a metabolic checkpoint in direct neuronal reprogramming. Cell Stem Cell 18, 396–409 (2016).
Bansod, S., Kageyama, R. & Ohtsuka, T. Hes5 regulates the transition timing of neurogenesis and gliogenesis in mammalian neocortical development. Development 144, 3156–3167 (2017).
Nishi, Y. et al. A direct fate exclusion mechanism by Sonic hedgehog-regulated transcriptional repressors. Development 142, 3286–3293 (2015).
Wang, Y., Yan, D., Liu, J., Tang, D. & Chen, X. Protein modification and degradation in ferroptosis. Redox Biol. 75, 103259 (2024).
Novitch, B. G., Chen, A. I. & Jessell, T. M. Coordinate regulation of motor neuron subtype identity and pan-neuronal properties by the bHLH repressor Olig2. Neuron 31, 773–789 (2001).
Griffin, J. M. et al. Astrocyte-selective AAV gene therapy through the endogenous GFAP promoter results in robust transduction in the rat spinal cord following injury. Gene Ther. 26, 198–210 (2019).
Ortinski, P. I. et al. Selective induction of astrocytic gliosis generates deficits in neuronal inhibition. Nat. Neurosci. 13, 584–U593 (2010).
Wang, L. L. et al. Revisiting astrocyte to neuron conversion with lineage tracing in vivo. Cell 184, 5465–5481 e5416 (2021).
Mattugini, N. et al. Inducing different neuronal subtypes from astrocytes in the injured mouse cerebral cortex. Neuron 103, 1086–1095 e1085 (2019).
Calzolari, F. & Berninger, B. cAAVe phaenomena: beware of appearances!. Cell 184, 5303–5305 (2021).
Wu, Z. et al. Brain-wide neuroregenerative gene therapy improves cognition in a mouse model of Alzheimer’s disease. Adv. Sci. 12, e2410080 (2025).
Su, Z., Niu, W., Liu, M. L., Zou, Y. & Zhang, C. L. In vivo conversion of astrocytes to neurons in the injured adult spinal cord. Nat. Commun. 5, 3338 (2014).
Bomba-Warczak, E., Edassery, S. L., Hark, T. J. & Savas, J. N. Long-lived mitochondrial cristae proteins in mouse heart and brain. J. Cell Biol. 220, e202005193 (2021).
Mohar, B. et al. DELTA: a method for brain-wide measurement of synaptic protein turnover reveals localized plasticity during learning. Nat. Neurosci. 28, 1089–1098 (2025).
Savas, J. N., Toyama, B. H., Xu, T., Yates, J. R. & Hetzer, M. W. Extremely long-lived nuclear pore proteins in the rat brain. Science 335, 942–942 (2012).
Li, H. & Chen, G. In vivo reprogramming for CNS repair: regenerating neurons from endogenous glial cells. Neuron 91, 728–738 (2016).
Wang, H. et al. Region-specific distribution of Olig2-expressing astrocytes in adult mouse brain and spinal cord. Mol. Brain 14, 36 (2021).
Rao, Z. et al. Molecular mechanisms underlying ascl1-mediated astrocyte-to-neuron conversion. Stem Cell Rep. 16, 534–547 (2021).
Ma, N. X., Puls, B. & Chen, G. Transcriptomic analyses of NeuroD1-mediated astrocyte-to-neuron conversion. Dev. Neurobiol. 82, 375–391 (2022).
Li, H., de Faria, J. P., Andrew, P., Nitarska, J. & Richardson, W. D. Phosphorylation regulates OLIG2 cofactor choice and the motor neuron-oligodendrocyte fate switch. Neuron 69, 918–929 (2011).
Lattke, M. et al. Extensive transcriptional and chromatin changes underlie astrocyte maturation in vivo and in culture. Nat. Commun. 12, 4335 (2021).
Cai, J. et al. A crucial role for Olig2 in white matter astrocyte development. Development 134, 1887–1899 (2007).
Dromard, C. et al. NG2 and Olig2 expression provides evidence for phenotypic deregulation of cultured central nervous system and peripheral nervous system neural precursor cells. Stem Cells 25, 340–353 (2007).
Lambert, S. A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Bartosovic, M., Kabbe, M. & Castelo-Branco, G. Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues. Nat. Biotechnol. 39, 825–835 (2021).
Michael, A. K. et al. Cooperation between bHLH transcription factors and histones for DNA access. Nature 619, 385 (2023).
Bertrand, N., Castro, D. S. & Guillemot, F. Proneural genes and the specification of neural cell types. Nat. Rev. Neurosci. 3, 517–530 (2002).
Kronenberg, G. et al. Modulation of fate determinants Olig2 and Pax6 in resident glia evokes spiking neuroblasts in a model of mild brain ischemia. Stroke 41, 2944–2949 (2010).
Zhou, M. et al. Reprogramming astrocytes to motor neurons by activation of endogenous Ngn2 and Isl1. Stem Cell Rep. 16, 1777–1791 (2021).
Gaber, Z. B. & Novitch, B. G. All the embryo’s a stage, and Olig2 in its time plays many parts. Neuron 69, 833–835 (2011).
Sun, Y. et al. Phosphorylation state of Olig2 regulates proliferation of neural progenitors. Neuron 69, 906–917 (2011).
Ghazale, H. et al. Ascl1 phospho-site mutations enhance neuronal conversion of adult cortical astrocytes in vivo. Front. Neurosci. 16, 917071 (2022).
Matsuda-Ito, K., Matsuda, T. & Nakashima, K. Expression level of the reprogramming factor NeuroD1 is critical for neuronal conversion efficiency from different cell types. Sci. Rep. 12, 17980 (2022).
Yalcin, A. et al. The oligodendrocyte lineage transcription factor 2 (OLIG2) is epigenetically regulated in acute myeloid leukemia. Exp. Hematol. 55, 76–85 (2017).
Noack, F. et al. Multimodal profiling of the transcriptional regulatory landscape of the developing mouse cortex identifies Neurog2 as a key epigenome remodeler. Nat. Neurosci. 25, 154–167 (2022).
Acknowledgements
The work was funded by the Jinan University Fund for Distinguished Young Scholars awarded to Z.W., the Natural Science Foundation of Guangdong Province of China (Grant No. 2022A1515012294), Guangzhou Key Projects of Brain Science and Brain-Like Intelligence Technology (Project No. 202206060002), and the Program for Guangdong Introducing Innovative and Entrepreneurial Teams (Grant No. 2021ZT09Y552). We also would like to express our sincere gratitude to Dr. Zhang Liguo from Sichuan University for his invaluable advice and guidance on the Olig2 CUT&Tag experiment.
Author information
Authors and Affiliations
Contributions
Z.W. and G.C. conceived and supervised the entire project. C-Y.L. and K-Y.H. performed all experiments with assistance from W-Y.L., J-H.W., M.Y., X-M.C., Y.S., D-T.Y., and K.L. C-Y.L. performed the bioinformatics analysis. W.L. provided constructive comments during manuscript preparation. C-Y.L., Z.W., and G.C. analyzed the data, made the figures, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.C. is a co-founder of NeuExcell Therapeutics Inc. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Carol Schuurmans, who co-reviewed with Hussein Ghazale; and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lai, C., Hou, K., Li, W. et al. Olig2 acts as an inducible barrier to in vivo astrocyte-to-neuron conversion. Nat Commun 17, 2033 (2026). https://doi.org/10.1038/s41467-026-68869-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68869-4