Abstract
Plasmids remain by far the most common medium for delivering engineered DNA to microorganisms. However, the reliance on natural plasmid replication mechanisms limits their tunability, compatibility, and modularity. Here we refactored the natural pMB1 origin and created plasmids with customizable copy numbers by tuning refactored components. We then created compatible origins that use synthetic RNA regulators to implement independent copy control. We further demonstrated that the synthetic origin of replication (SynORI) can be engineered modularly to respond to various signals, allowing for multiplexed copy-based reporting of environmental signals. Lastly, a library of 6 compatible SynORI plasmids was created and co-maintained in E. coli for a week. This work establishes the feasibility of creating plasmids with SynORI that can serve as a biotechnology for synthetic biology.
Similar content being viewed by others
Introduction
DNA plasmids play a vital role as mobile genetic elements in bacterial evolution, capable of mobilizing key functions such as antibiotic resistance1,2,3, resistance to heat4, and substrate metabolism5,6,7. Plasmids also remain the most common way to deliver recombinant DNA to a broad range of microbes8,9,10. However, most plasmids used today are limited to a few well-characterized types from the 1980s that have undergone minimal engineering, particularly in their origins of replication (ORI), which dictates plasmid copy number and compatibility (i.e., ability of plasmids to stably co-exist in the same cell)11,12. As a result, researchers are largely restricted to a limited menu of plasmids that have fixed copy numbers and limited compatibility, posing challenges for the advancement of plasmid-based applications, such as implementing division of labor across plasmids. In part, this lack of engineered plasmids is due to the non-modular genetic architectures of ORIs, which have evolved to incorporate features such as overlapping genes and operons. This lack of tunability, compatible variants, and modularity is in stark contrast to other genetic parts (e.g., promoters, RBS, and terminators) that have been extensively engineered13,14,15.
Previously, efforts have been made to extend the tunability of plasmid copy number16,17,18. Sheth et al. created tunable plasmid pTrig by inducible expression of phage P1 lytic replication protein RepL, achieving a 635-fold change in copy number. Rouches et al. created tunable ColE1-based plasmids that allow plasmid copy numbers to range from 1.4 to 50 or 270 to 30 with inducible control, and from 1 to 800 with constitutive promoter engineering. Joshi et al. created a tunable pSC101 plasmid by the inducible expression of RepA protein, allowing a plasmid copy number ranging from 3 to 60. Other than direct regulation of the ORI, horizontal gene transfer can also be harnessed to tune plasmid copy number19. Son et al. achieved population-level plasmid copy tuning through plasmid conjugation and Cas9-based plasmid deletion, resulting in a 4400-fold change in plasmid density within the population. These previous efforts to design plasmids with customizable copy number have generally avoided altering the ORI sequence. Instead, tunability was achieved by replacing peripheral promoters with inducible versions or by adding extra inducible regulatory elements. While these strategies have proven effective in creating plasmids with tunable copy number, the lack of direct engineering of the ORI itself has limited further advances in plasmid compatibility and modularity.
To create synthetic plasmids beyond the bounds of evolution, we focused on creating a synthetic ORI (SynORI) that would address the current limitations of modularity, tunability, and compatibility. To do this, we refactored and reengineered the pMB1 origin of replication (Fig. 1A). pMB1 belongs to the ColE1 family that initiates DNA replication through transcription and processing of an RNA primer (RNAII). DNA replication initiation is regulated by an antisense RNA (RNAI) that binds to the RNA primer co-transcriptionally to induce an alternative conformation that terminates DNA replication20. Because RNAI must act on RNAII within a narrow co-transcriptional window—and is rapidly degraded by RNase E—its regulation of replication is inherently non-linear and highly sensitive to RNAI abundance21. As the concentration of RNAI is directly linked to plasmid copy number, this establishes negative feedback control on the replication of the pMB1 plasmid.
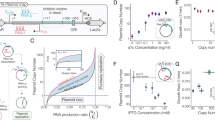
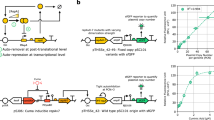
A Schematic of the MB1 plasmid origin of replication (ORI) mechanism. The RNA primer is transcribed by RNA polymerase (RNAP) and, by default, folds into an RNA structure that is processed by RNase H to create a substrate for DNA polymerase I (Pol I) to initiate DNA replication. The antisense RNA is produced from an overlapping gene, which, if allowed to interact with the RNA primer, induces an alternative fold that does not initiate DNA replication. As the concentration of antisense RNA is proportional to the plasmid copy, this serves as a copy-control negative feedback loop. B Schematic of a refactored pMB1 ORI that separates the RNA primer and antisense RNA gene and introduces inactivating mutations in the P1 promoter sequence. C Shows sequences of mutations to inactivate the P1 promoter encoded inside the RNA primer. (Left graph) Whole-cell fluorescent characterization of the mutant promoters using a fluorescent reporter gene. (Right graph) Relative copy number of plasmids containing the original pMB1 ORI or refactored ORI with an RNA primer containing mutant P1 promoters. Copy number was characterized by encoding a constitutive RFP expression cassette onto the plasmid. Fluorescence characterization was performed (measured in units of fluorescence/optical density (OD) at 600 nm) in E. coli cells. Data show mean values ± SD and individual values of n = 4 biological replicates. Source data for this figure is available in the Source Data file.
Results
The pMB1 ORI can be refactored
To refactor the overlapping components of pMB1 ORI, we evaluated whether the antisense RNA promoter inside the RNA primer, called P1, could be removed without disrupting the replication capability of the ORI (Fig. 1B). A collection of P1 promoters mutated in their −10 and −35 regions were designed and their relative strength measured in E. coli cells using RFP expression (Fig. 1C and Supplementary Fig. 1). No promoter activity was observed from any of the modified promoters. We then built a refactored pMB1 ORI by separating the antisense RNA and RNA primer into separate transcription cassettes and using an RNA primer that contained mutations in the P1 promoter. To measure the relative copy number, a constitutively expressed RFP was inserted into each refactored plasmid, and whole-cell fluorescence was measured. A reduction in the relative copy number was seen with greater modification in the RNA primer (Fig. 1C), with the 2-mutation promoter having the closest copy number to the natural pMB1 ORI, which was used for subsequent iterations. Taken together, these results show we can effectively refactor the pMB1 ORI by separating transcription cassettes and engineering the RNA primer.
Synthetic ORI (SynORI) can be created using synthetic transcriptional repressors
We next sought to create a synthetic origin of replication (SynORI) in which the natural pMB1 ORI copy control mechanism was replaced with synthetic transcriptional repressors. Specifically, we focused on the pT181 transcriptional attenuator, a transcriptional regulator discovered in a naturally occurring plasmid from Staphylococcus aureus22, which has been extensively engineered to yield orthogonal libraries23,24. This system is composed of a target RNA that harbors a latent transcriptional terminator that adopts a conformation that prevents downstream transcription only in the presence of its corresponding repressor small RNA (sRNA) (Supplementary Fig. 2). We reasoned that, by inserting the RNA primer downstream of the target and replacing the antisense RNA with the repressor sRNA, we could effectively implement negative feedback to control the plasmid copy number in SynORI (Fig. 2A). By separating copy-control and replication initiation into distinct, interchangeable genetic modules, this method offers enhanced design flexibility. To test this hypothesis, a set of SynORIs using a pair of orthogonal pT181 attenuators that contained all four combinations of target and repressor sRNAs was created, and the relative copy number was characterized. Designs with cognate pairs of target and repressor that can implement negative feedback showed a dramatic decrease in relative copy number in E. coli cells compared to designs using non-cognate pairs (Fig. 2B, Supplementary Fig. 3a). Interestingly, we saw the same trend in Shewanella oneidensis, suggesting that SynORI functions in multiple cell types (Supplementary Fig. 4). These results suggest that SynORIs can be created that implement plasmid copy control using synthetic transcriptional regulators.
A Schematic of synthetic ORI (SynORI) in which the copy-control mechanism is replaced with the synthetic pT181 transcriptional attenuator. In this mechanism, the repressor small RNA (sRNA) inhibits transcription of a target RNA that is placed upstream of the RNA primer to control plasmid copy number. Matrix showing the relative copy number of plasmids containing (B) different combinations of target and repressor sRNA from two orthogonal pT181 variants (WT and LS) and C different strength promoters. D Relative copy number of plasmids with different SynORI genetic architectures. Relative copy number was characterized by encoding a constitutive RFP expression cassette onto the plasmid in E. coli cells (measured in units of Fluorescence/Optical density [OD] at 600 nm). E Schematic of multimer resolving by parCBA. F Percentage of plasmid multimers in E. coli cells with and without parCBA present. Data shows the mass ratio (percentage of total nucleotides in each multimer species) measured by nanopore whole-plasmid sequencing. G Schematic of the RNA primer truncation library. H Relative enrichment of each truncation (5′ to 3′) using a growth-based selection. Relative enrichment was determined by comparing the abundance of each design in a naïve library before the Golden Gate assembly to the abundance of each design after transformation, selection, and plasmid isolation. The enrichment of the plasmid variant without truncation (0 nt) was normalized to 1. Only truncations that preserve plasmid replication should be enriched following the selection. The inset shows the predicted structure of the RNA primer, with each nucleotide’s color corresponding to the relative enrichment at that truncation site. In B–D, H data shows the mean of n = 4 biological replicates and in F data shows the values of n = 3 biological replicates. All error bars indicate SD. Source data for this figure is available in the Source Data file.
To determine whether SynORI offers tunable copy control, we replaced the promoters driving both target and repressor sRNA with four varying strength promoters. Using RFP as a relative measure of copy number, we saw the RFP varied over 26-fold, with the highest copy number seen when target RNA transcription was greater than the repressor sRNA, and the lowest copy number when transcription rates were inverted (Fig. 2C). We note that the combination of high target and low repressor sRNA transcription appeared to lead to plasmid instability (i.e., runaway plasmids), resulting in large errors between replicates (Supplementary Fig. 3b). As an additional measure of copy number, we also performed qPCR on E. coli cells to quantify the abundance of the plasmid-encoded rfp gene relative to a single-copy genome-encoded housekeeping gene, dxs25. This showed plasmid copy varied over 115-fold with estimated plasmid copies from 1.6 to 185 per cell (Supplementary Fig. 5). Comparing plasmid copy quantification by fluorescence and qPCR, we saw good correlation on a log-log scale, consistent with power-law scaling, with fluorescence exhibiting signal saturation at the highest copy numbers (>150). Next, we investigated how the orientation of transcriptional cassettes affected the copy number (Fig. 2D). Tandem-down and convergent orientations showed the lowest plasmid copy, likely due to the repression of RNA primer transcription from DNA supercoiling created by the transcription of the upstream repressor sRNA26,27. Taken together, this shows SynORI copy number can be tuned using promoter libraries and distinct genetic architectures, which are unfeasible with the natural pMB1 ORI.
Plasmid multimerization is a persistent problem seen in the ColE1 plasmid family28,29,30, which in part stems from the deletion of the recombination site, cer, from the laboratory derivatives that resolve multimers31. To investigate multimerization in our SynORI, we sequenced DNA plasmids from E. coli cells using whole-plasmid sequencing. We observed ~10.5% and ~1.5% of plasmids formed dimers and trimers, respectively. Surprisingly, reintroducing cer into SynORI did not reduce multimer formation (Supplementary Fig. 6). However, incorporating a parCBA system, adapted from plasmid RK232 (Fig. 2E), effectively eliminated multimers (Fig. 2F). The addition of a toxin-antitoxin system, parDE, further enhanced plasmid stability4. These findings demonstrate that natural plasmid maintenance mechanisms can effectively mitigate SynORI multimerization.
As SynORI is independent of the natural RNA primer’s copy control mechanism, we reasoned that the RNA primer could be minimized. Deletion of the entire region responsible for interacting with the antisense RNA (termed stem loop I, II, and III) proved non-functional in E. coli. Therefore, we conducted incremental truncations from the 5′ end of the RNA primer using a pooled library approach (Fig. 2G). By sequencing and analyzing the enrichment of various truncations after transformation into E. coli, we found that approximately two-thirds of the interacting sequence (stem loops I and II) could be removed without compromising functionality (Fig. 2H, Supplementary Fig. 7). However, truncations inside stem loop III led to non-functional SynORIs, which we verified by individual plasmid characterization (Supplementary Fig. 8). Interestingly, truncations within stem loop structures often disrupted RNA primer function, suggesting that an incomplete stem loop disrupted RNA primer function. Taken together, these results show that a functional and minimal SynORI design can be achieved using an RNA primer truncated to stem loop III, in combination with a tunable synthetic promoter, a convergent genetic architecture, and a parCBA resolvase system.
Benchmarking the performance of pSynORI
To evaluate the overall performance of the finalized SynORI plasmids (pSynORI) we compared it with pMB1 in E. coli, with regard to several important performance criteria. Because plasmid maintenance can impose a fitness burden on the host, growth rate comparisons were performed to determine whether the engineering of pSynORI alters host physiology. Cells containing pSynORI had generally comparable growth curves to cells containing pMB1, although maximum growth was slightly reduced (Fig. 3A). We next assessed whether differences in copy control mechanisms led to altered heterogeneity of plasmid copy at the single-cell level by measuring single-cell fluorescence. The plasmid copy distribution showed a slightly wider copy distribution for pSynORI (Fig. 3B), likely due to the intrinsic difference in the pMB1 and pSynORI copy control mechanisms. Next, we evaluated the rate of multimer formation since multimerization is a common source of plasmid instability in ColE1-like origins. To do this, whole plasmid sequencing was used, and multimers were quantified. As expected, multimers were not observed for pSynORI; however, significant levels of multimers (dimers, trimers, and tetramers) were seen for pMB1. Plasmid copy number stability was next considered by measuring whole-cell fluorescence over a 7-day continuous culture with antibiotic selection. This assay was designed to assess whether pSynORI could maintain relatively stable copy numbers over many generations under selective conditions. Both pMB1 and pSynORI exhibited relatively stable whole-cell fluorescence levels over 7 days (Fig. 3D). We also compared the single-cell fluorescence distribution. Both pMB1 and pSynORI maintain a stable distribution through the 7-day culture, while the latter shows a wider distribution and slightly more cells (~2%) with no fluorescence (Supplementary Fig. 9). Plasmid integrity was also verified through whole-plasmid sequencing of samples from day 1 and day 7. Finally, we quantified the rate of spontaneous plasmid loss by comparing colony-forming units (CFU) with and without antibiotic selection of plasmids in a 7-day continuous growth experiment (Fig. 3E). This experiment was motivated by the possibility that engineering of pSynORI might compromise plasmid retention, leading to loss in the absence of selection. No plasmid loss was observed in cells with pMB1 and pSynORI after overnight growth with or without antibiotics. Likewise, no loss was seen with the antibiotic across 7 days for either pMB1 or pSynORI. Interestingly, for pSynORI, without antibiotic selection, plasmid loss was seen in 67% of cells after 7 days. Although the underlying cause remains unclear, this finding suggests that the engineering of pSynORI may compromise certain stability features, such as genetic stability or plasmid partitioning. Taken together, these results indicate that pSynORI performs comparably to the original pMB1 plasmid.
A Characterization of growth rate in E. coli cells transformed with pMB1 and pSynORI (measured in units of optical density [OD] at 600 nm). The maximum growth rate is indicated. B Single-cell fluorescence of E. coli cells transformed with pMB1 and pSynORI containing a constitutive RFP expression cassette. The coefficients of variation (CV) of 3 biological replicates of pMB1 are 56.9, 57.8, and 57.9. The CVs of pSynORI are 60.1, 68.2, and 60.0. C Percentage of plasmid multimers in E. coli cells with and without parCBA present. Data show the mass ratio (percentage of total nucleotides in each multimer species) measured by nanopore whole-plasmid sequencing. D Relative copy number of plasmids over time. Relative copy number was characterized by encoding a constitutive RFP expression cassette onto the plasmid in E. coli cells (measured in units of fluorescence/optical density [OD]). E The percentage of cells retaining plasmid after culturing is measured by comparing the percentage of cells surviving with and without antibiotic (+AB) selection over a continuous 7-day culture. Cells retaining plasmid are resistant to AB and should grow in both conditions. Data A–D show n = 3 biological replicates. Data in E show 20 colonies collected from n = 3 biological replicates. All error bars and error bands indicate SD. Source data for this figure is available in the Source Data file.
Converting chemical signals into plasmid copy
An advantage of the pSynORI system is its flexibility in adapting different regulatory systems for plasmid copy control. To demonstrate this, we explored ligand-inducible copy control using inducible promoters and transcriptional riboswitches (Fig. 4A). By using the IPTG- and cumate-inducible promoters to transcribe the pT181-regulated RNA primer, we achieved up to a 13-fold increase in RFP (Fig. 4B, C). We also investigated the impact of IPTG-induced plasmid copy control on cell growth rate. Interestingly, a higher IPTG induction level leads to faster growth. We posit this may be due to higher copy of the antibiotic resistance gene, although further investigation is required (Supplementary Fig. 10). Additionally, we replaced the pT181 target RNA with the ZTP- and 2AP-riboswitches33,34,35, enabling both low-to-high and high-to-low plasmid copy flipping in response to specific chemical signals (Fig. 4D, E). Comparing the response curves of the different inducible systems, we see differences in the copy control dynamic range, which we posit is due to the differences in the dynamic range of each transcriptional regulator. These results highlight pSynORI’s potential as a flexible platform for converting chemical information into plasmid copy number, providing a reporting modality that can be easily analyzed through sequencing.
A SynORI plasmids (pSynORI) can be programmed to convert distinct chemical signals into DNA copy output using transcriptional regulators to control RNA primer transcription. Relationship between relative copy number and chemical input for pSynORI built using (B) Plac (activated by IPTG), C PcymR (activated by cumate), D the ZTP riboswitch (activated by Z), E and the yxjA riboswitch (repressed by 2AP). F Matrix showing the relative copy of a pSynORI that is activated by cumate and repressed by IPTG with inducer titrations. Relative copy number was characterized by encoding a constitutive RFP expression cassette onto the plasmid in E. coli cells (measured in units of fluorescence/optical density [OD] at 600 nm). G Schematic of compatible and inducible pSynORI that converts chemical input signals into a DNA copy, which can be read using sequencing. A pSC101 plasmid with a non-inducible copy number is included as a reference. H The relative copy of pSynORI A and B was calculated using the reference plasmid. Bar plots show the DNA copy ratio of pSynORI A and B. Data in B–E and F shows n = 4 biological replicates, and Data in H show n = 3 biological replicates. All error bands indicate SD. Source data for this figure is available in the Source Data file.
We also explored the integration of multiple signals for plasmid copy regulation by using the cumate- and IPTG-inducible promoters to control RNA primer and repressor sRNA transcription independently. As expected, in most cases, relative plasmid copy was upregulated by cumate and downregulated by IPTG. Interestingly, an exception to this was when 400 μM cumate was used, which showed slight upregulation by IPTG (Fig. 4F, Supplementary Fig. 11). We posit this could be due to runaway plasmid replication at higher cumate concentrations, disrupting the expected trend. This result demonstrates that SynORIs can integrate multiple signals to regulate copy number; however, the relationship between chemical signals and copy number can be more complex and unpredictable. As such, to achieve the predictable conversion of multiple signals into a DNA copy, further work is required.
Next, we investigated the potential of pSynORI for multiplexable reporting of chemical signals through copy control. To test this concept, we constructed two compatible pSynORI (A and B) responding to cumate and IPTG, respectively (Fig. 4G). These pSynORI were co-transformed into E. coli cells alongside a reference plasmid, pSC101, that should maintain a steady copy number. Cells were exposed to different chemical combinations, and plasmid copy numbers were determined through whole plasmid sequencing (Fig. 4H). Relative to the reference plasmid, pSynORI A and B showed increases in copy number upon addition of their corresponding chemical signal, although the exact relative copy level achieved varied between conditions. The relative ratio of these plasmids was also adjusted as expected, which was further validated using fluorescent reporters (Supplementary Fig. 12). Additionally, the dynamic responses of single-plasmid and double-plasmid inducible SynORI systems were characterized using a microfluidic device (Supplementary Fig. 13, Supplementary Movie 1 and Supplementary Movie 2). These results show pSynORI can be used for multiplexing the reporting of chemical signals using easy-to-read plasmid copy numbers.
Six compatible pSynORI can co-exist in the same cell
An advantage of pSynORI is the ability to leverage orthogonal regulatory systems for plasmid copy control, enabling the creation of compatible plasmids that coexist within the same cell. To demonstrate this, we constructed seven pSynORI plasmids using previously described orthogonal pT181 systems (YS, F6, F4, F4m1, F3m1, F15m5, and LS)24. We initially validated the ability of different target and repressor sRNA pairs to regulate copy control in pSynORI. For each pT181 variant, a pSynORI with a matching target and repressor sRNA was compared with designs containing a non-matching target or repressor sRNA from another variant, LS. All tested pT181 pairs showed clear repression of pSynORI copy only when the target and repressor sRNA were matched (Fig. 5A, Supplementary Fig. 14a). Interestingly, the final copy number of each pSynORI varied (3.4-fold), which we reasoned was due to the difference in repression efficiency of these pT181 variants24 or potential sequence-specific interactions with the downstream RNA primer. We also compared growth rates of cells containing each pSynORI and also saw a relatively faster growth in high-copy variant F15m5 (Supplementary Fig. 15). These results demonstrate that a library of pSynORI plasmids can be created using orthogonal transcriptional regulator libraries.
A Matrices showing the relative copy number of pSynORI built with 7 orthogonal pT181 variants. For each variant, combinations with a non-matching target and repressor sRNA (LS) are also included as a control. B A library of 6 pSynORI with different pT181 regulation systems (LS, YS, F6, F4, F4m1, and F3m1) and antibiotic resistances (CmR, SpcR, KanR, AmpR, TetR, and AprR) was created and co-transformed into E. coli cells. The proportions of plasmids in a cell population are measured using nanopore whole-plasmid sequencing. C Plasmid proportion during 8-day continuous culture with antibiotic selection. Cells are re-streaked on day 6, and individual colonies grown from a single cell were picked for day 8 cultures. The samples from days 1, 2, 4, 6, and 8 were processed for sequencing to confirm the presence of plasmids. The ring plot shows the ratio of plasmids on day 1, 6, and 8, with other days shown in Supplementary Fig. 14. Data in A show n = 4 biological replicates and data in C show n = 3 biological replicates. Source data for this figure is available in the Source Data file.
We then picked six SynORI variants that exhibit strong plasmid copy repression to create a library of compatible pSynORI. Each pSynORI was cloned to contain a distinct antibiotic resistance gene. Surprisingly, colonies were obtained from a single transformation of all six pSynORI into E. coli cells. To verify the presence of all six plasmids, we conducted a 6-day continuous culture with antibiotic selection. After 1, 2, 4, and 6 days, plasmids were extracted, and sequenced, and the presence of the six plasmids confirmed (Fig. 5C, Supplementary Fig. 14b). Comparing relative distribution of the six plasmids, we saw it was uneven and did not correlate with individual relative copy numbers (Supplementary Fig. 16), likely due to varying pT181 repression efficiencies and potential crosstalk. Importantly, we observed no plasmid loss throughout the culture. Interestingly, a mutation in the LS-SynORI plasmid on day 4 in one biological repeat led to a change in plasmid copy from low to high (Supplementary Fig. 14b), suggesting that the antibiotic concentrations might require optimization based on plasmid composition and that maintaining six plasmids simultaneously can be challenging for the cell. As a control, we also constructed six pMB1 plasmids with distinct antibiotic-resistance genes. Attempts to transform the six pMB1 were unsuccessful (Supplementary Fig. 17a), although colonies were obtained by transforming four pMB1 (Supplementary Fig. 17b). A mixture of small and large colonies was observed, and attempts to grow cultures resulted in no growth for some colonies. For cultures that grew, analysis of isolated DNA plasmids showed a high frequency of dimer formation (Supplementary Fig. 17c). Taken together, these results suggest our modifications to pSynORI are critical for achieving compatibility.
We next aimed to provide evidence that cultures were composed of cells containing all six plasmids, rather than a consortium of cells that had escaped individual plasmids. To test this, an aliquot of cells from the day 6 culture was plated and grown overnight. Single colonies grown from individual cells were then used to inoculate a culture grown until day 8, and the presence of all six plasmids was confirmed. In conclusion, we have successfully demonstrated the creation of a compatible library of six pSynORI that can co-exist in the same cell, which extends the current state of the art in plasmid design.
Discussion
Our pSynORI system demonstrates that plasmids can be extensively refactored and modified to achieve desired plasmid features of tunability, compatibility, and modularity in replication control. In contrast to previous works creating tunable copy-control plasmids17,18, pSynORI features refactored regulatory elements that effectively modularize plasmid copy regulation and replication initiation. This modular design of pSynORI creates flexibility by making it easier to add desired regulatory elements without unintentionally impairing replication. This modularity allowed the incorporation of inducible promoters and riboswitches, enabling pSynORI plasmids to respond to diverse input signals and for multiplexed, copy-based reporting of environmental signals. Additionally, this modularity allows pSynORI to address an important limitation of prior efforts, which is the creation of multiple engineered plasmids that are compatible. Together, these features establish pSynORI as a versatile and extensible platform that advances plasmid origin engineering.
We anticipate that pSynORI tunability, modularity, and compatibility could open up applications in the future. Herein, we demonstrate that chemical signals can be converted into DNA copy changes using pSynORI, enabling multiplexable biosensing. We show that two inputs can be simultaneously detected, with results directly read out using portable nanopore sequencing. Unlike fluorescent reporters limited by spectral overlap, pSynORI could potentially offer higher multiplexing. Another application that our technology can support is the rapid prototyping of genetic programs. Specifically, our set of six compatible plasmids, which are also amenable to tunable copy control, can allow researchers to mix and match multiple plasmids in a single E. coli cell to test gene or pathway variants in vivo. This type of flexibility has been utilized in cell-free systems for engineering gene circuits and metabolic pathways36. Looking further ahead, we believe pSynORI, with its tunability, modularity, and compatibility, can be used for other applications. For example, coupling copy control to different inputs could be valuable for biocontainment measures to restrict the spread of plasmids37 or to dynamically control plasmid copy to decouple growth and production phases38. In this way, we expect pSynORI to serve as a next-generation plasmid system with broad utility across biotechnology and synthetic biology.
The SynORI allows compatible co-existence of multiple plasmids in the same cell using synthetic transcriptional regulators. However, their orthogonality is not absolute, which we reason are due to several factors. First, there is an inherent competition for resources between plasmids that coexist in the same cell, leading to interactions that could alter copy number, especially for high-copy plasmids. Second, the potential crosstalk between synthetic pT181 regulators could cause repression of one pSynORI by another.
While we focused here on refactoring and reengineering the pMB1 plasmid, we expect the approach to be broadly applicable to other plasmids39. This would be useful to apply to plasmids suitable for non-model microbes, which often lack plasmids that are functional, compatible, or cover diverse copy ranges. Applying this approach to broad-host range plasmids discovered within plasmidomes40, in combination with broad-host or conjugative genetic parts8,41,42,43,44, could allow the creation of pSynORI adapted for programming diverse microbes or microbiomes. In summary, the development of pSynORI could play a vital role in establishing tunable, compatible, and modular plasmid-based synthetic biological systems.
Methods
Plasmid assembly strategies
All plasmids used in this study are listed in Supplementary Data 1. The key plasmids are visualized in Supplementary Fig. 18. The sequence of the pSynORI plasmid, characterized in Fig. 3 and the sequence of pT181 transcriptional repressors are provided in Supplementary Data 1. The sequences of promoters and riboswitches are provided in Supplementary Tables 1 and 2. The examples of RNA primer truncations used in Fig. 2G, H are provided in Supplementary Table 3. Plasmids were constructed with a combination of inverse PCR, Gibson assembly, or Golden Gate assembly. The promoters and pT181 were replaced using a flexible design of Golden Gate assembly, which is explained in Supplementary Table 4 with key primer sequences. All assembled plasmids were verified using Sanger DNA sequencing (Genewiz) or Nanopore sequencing (Plasmidsaurus).
Plasmid transformation and culture conditions
For experiments using E. coli, plasmids were transformed into chemically competent E. coli (NEB Turbo strain). E. coli cells were recovered at 37 °C for 1 hour, plated on LB-agar plates containing combinations of 34 μg/mL chloramphenicol (Sigma-Aldrich), 50 μg/mL spectinomycin (Sigma-Aldrich), and 100 μg/mL kanamycin (Sigma-Aldrich) depending on the plasmids used, and incubated overnight at 37 °C. For the transformation of six plasmids, the LB-agar plates contained 34 μg/mL chloramphenicol, 50 μg/mL spectinomycin, 50 μg/mL kanamycin, 100 μg/mL carbenicillin (Sigma-Aldrich), 50 μg/mL apramycin (Sigma-Aldrich), and 5 μg/mL tetracycline (Sigma-Aldrich). 1.25 μg/mL tetracycline was used for liquid LB culture, as tetracycline showed higher selection efficiency in liquid LB than on LB-agar plates.
For induction experiments using E. coli, colonies were picked from overnight LB plates to inoculate 200 μL of LB containing the corresponding antibiotic in a 2 mL 96-well block (Costar), and grown overnight at 1000 rpm in a VorTemp 56 bench-top shaker (Labnet) at 37 °C. Then 4 μL of each overnight culture was added to 192 μL of LB containing corresponding antibiotics in a newly prepared 96-well block. Depending on the experiment, 4 μL of inducer was added to the newly prepared culture to reach the final concentrations of: 1, 3, 10, 30, 100, 300, 1000 μM of IPTG (Sigma-Aldrich) for Fig. 4B; 5, 10, 25, 50, 100, 200, 400 μM of cumate (Sigma-Aldrich) for Fig. 4C; 1, 3, 10, 30, 100, 300, 1000 μM of 5-aminoimidazole-4-carboxamide ribonucleotide (Z, Sigma-Aldrich) for Fig. 4C; 1, 3, 10, 30, 100, 300, 1000 μM of 2-aminopurine (2-AP, Sigma-Aldrich) for Fig. 4D. The cells were then incubated at 1000 rpm in VorTemp at 37 °C for 8 h before whole-cell fluorescence measurement.
For experiments using S. oneidensis, plasmids were transformed into electrocompetent cells. The cells were prepared by washing overnight-cultured cells 3 times with 10 % glycerol. Plasmids were electroporated into the cells using Gene Pulser Xcell electroporation system (Bio-Rad) with 1.2 kV, 25 μF, 200 Ω setting in 1 mm electroporation cuvettes (Fisher). S. oneidensis cells were recovered at 30 °C for 2 h, then plated on LB-agar plates containing 50 μg/mL kanamycin and incubated overnight at 30 °C.
For continuous culture experiments, single colonies were picked from the overnight-incubated LB-agar plates and inoculated in 50 mL glass culture tubes with 5 mL liquid LB containing corresponding antibiotics (LB-AB) and incubated at 37 °C, shaking at 225 rpm. For every 24 h, 5 μL of cell culture was inoculated into a new culture tube with 5 mL LB-AB (1:1000 dilution) until the end of the experiment.
Whole-cell fluorescence measurement
Bulk fluorescence measurements were performed with 50 μL of culture diluted in 50 μL of phosphate-buffered saline (PBS, Fisher) in a 96-well plate using a microplate reader (Tecan Spark). The GFP was measured with 485/520 nm as excitation/emission wavelengths. The RFP was measured with 560/630 nm as excitation/emission wavelengths. Optical density at 600 nm (OD600) was also measured. Each 96-well block included two sets of controls: a media blank and E. coli transformed with empty control plasmids, or S. oneidensis without plasmids, referred to here as blank cells. Blank cells were used to determine autofluorescence levels. OD and fluorescence values for each colony were first corrected by subtracting the mean value of the media blank from the respective values of the experimental conditions. The ratio of the corrected fluorescence to the corrected OD (fluorescence/OD) was then calculated for each well. Autofluorescence was removed through the subtraction of fluorescence/OD of the blank cells.
For single-cell fluorescence analysis, single colonies were picked from the overnight-incubated LB-agar plates and inoculated in 50 mL glass culture tubes with 5 mL liquid LB containing 34 μg/mL chloramphenicol. After an overnight culture at 37 °C, an aliquot was taken and diluted with PBS (1:100). The fluorescence distribution of the single cells in the sample was measured by flow cytometry (Sony SH800S cell sorter). Each measurement contains 100,000 event reads. The data were then processed and analyzed in the FlowJo software (v10.8.1). An example gating strategy to isolate single cells is shown in Supplementary Fig. 19.
Plasmid copy number quantification with quantitative PCR
To quantify plasmid copy numbers, E. coli cultures were sampled at the time of whole-cell fluorescence measurements. After the 8-h growth, 5 μL of cell culture was mixed with 45 μL water and heated at 98 °C for 5 min. Cell lysates were further diluted 10-fold in water and stored at −20 °C until measured. In total, 2 μL of thawed lysate was used as template DNA in 10 μL Maxima SYBR Green/ROX (Thermo Scientific) qPCR reactions, and measurements were run in a QuantStudio 5 System (Applied Biosystems). For each sample, 2 reactions were run using primers targeting the rfp gene on the plasmids and the housekeeping dxs gene on the E. coli genome25. Plasmid copy number was calculated from the ΔCt between dxs and rfp, using the assumption that dxs is present at a single copy per cell45.
Selection for functional RNA primer truncations
A DNA oligonucleotide library (Twist Bioscience) was ordered and then amplified through polymerase chain reaction (PCR) using forward and reverse primer shown in Supplementary Table 3, with the following conditions: 98 °C (30 s), 98 °C (15 s) 63 °C (45 s) 72 °C (15 s) for 13 cycles, 72 °C (5 min), 4 °C (forever). This amplified product was used to create the plasmid library containing different RNA primer truncations through Golden Gate assembly. The plasmid library was transformed into E. coli cells and then incubated at 37 °C overnight on LB-agar plates with antibiotic selection. Cells were removed from the plate, and DNA plasmids were isolated using a DNA mini plasmid preparation (Qiagen). Using 10 ng of DNA plasmid as a template, the RNA primer (RNAII) region was amplified using primers also containing adapters for NGS sequencing using Amplicon-EZ (Genewiz). The following PCR conditions were used: 98 °C (30 s), 98 °C (15 s) 63 °C (45 s) 72 °C (15 s) for 20 cycles, 72 °C (5 min), 4 °C (forever). As a naïve control, the amplified PCR product used to construct the plasmid library before transformation was also prepared and sequenced in parallel. The relative enrichments were calculated by dividing the reads from each plasmid variant by the reads from the corresponding naïve control, which was then further normalized by the enrichment of the plasmid without truncation (0 nt).
Growth rate characterization
Single colonies were picked from the overnight-incubated LB-agar plates and inoculated in 50 mL glass culture tubes with 5 mL liquid LB containing 34 μg/mL chloramphenicol (LB-AB). After overnight growth at 37 °C, shaking at 225 rpm, an aliquot was taken and diluted 1:10000 with fresh liquid LB-AB and 200 μL of diluted sample was added to each well of a 96-well plate. The plate was then placed in the microplate reader (Tecan Spark) with the humidity cassette for continuous culture at 37 °C, shaking at 108 rpm. The OD600 of each sample was measured every 10 min for 20 h.
Plasmid loss characterization
Single colonies were picked from the overnight-incubated LB-agar plates and inoculated in 50 mL glass culture tubes with 5 mL liquid LB either containing (LB–AB) or lacking 34 μg/mL chloramphenicol (LB) and cultured continuously for 7 days with a 1:1000 dilution every day. On days 1, 3, 5 and 7, an aliquot of the culture was then streaked on LB-agar plates without antibiotics present and grown overnight. For each biological replicate, 20 colonies were picked and transferred to a new LB-agar plate containing 34 μg/mL chloramphenicol and grown overnight. The survival rate of transferred colonies was then determined to characterize plasmid loss.
Dynamic characterization through a microfluidic device
The experiment was conducted in a microfluidic device with dial-a-wave (DAW) junction and hallway traps46. Microfluidic devices were constructed using standard techniques47. E. coli cells were transformed with IPTG-inducible pSynORI, or with both IPTG-inducible and cumate-inducible pSynORIs. Cells were grown in LB media overnight with the corresponding antibiotic. After overnight growth at 37 °C, shaking at 225 rpm, an aliquot was taken and diluted 1:1000 into 25 mL fresh LB media with antibiotics. After 6 h of growth when OD600 reached ~0.4, 15 mL of culture was centrifuged at 2000g for 8 min at 25 °C. The cell pellet was then resuspended in 10 mL prewarmed fresh LB media with antibiotics and 0.5% Tween 20 (Fisher) and loaded into the microfluidic device at 37 °C. 0.5% Tween 20 was used in all media to prevent cells from adhering to the channel without affecting the cell growth and fluorescence expression.
For cells with IPTG-inducible pSynORI, after cells were loaded, fresh LB media (20 mL in 60 mL syringe) with antibiotic and 0.5% Tween 20 were added to the channel with a flow velocity of 100 μm/s through the channel. After cells filled the trap and stabilized, the channel’s input media was switched to fresh LB media (20 mL in 60 mL syringe) with 100 μM IPTG, 10 μL Oregon Green 488 dye (Fisher), corresponding antibiotic, and 0.5% Tween 20 with the same flow velocity. After 17 h, the input media was switched back to LB media without IPTG till the end of the experiment.
For cells with both IPTG-inducible and cumate-inducible pSynORIs, after cells were loaded, fresh LB media (20 mL in 60 mL syringe) with 100 μM IPTG, antibiotics and 0.5% Tween 20 were added to the channel with a flow velocity of 50 μm/s through the channel. After cells fulfilled the trap and stabilized, the channel’s input media was switched to fresh LB media (20 mL in 60 mL syringe) with 200 μM cumate, corresponding antibiotics, and 0.5% Tween 20 with the same flow velocity. After 17.8 h, the input media was switched back to LB media with 100 μM IPTG till the end of the experiment. Both types of media contained no dye, and the media switch time was recorded manually during the switch.
Phase contrast and fluorescence images were acquired under an inverted fluorescence microscope (Nikon) every 5 minutes at 100× magnification. For cells containing a single pSynORI, the exposure times were set to 100 ms for the RFP fluorescence channel and 400 ms for the GFP fluorescence channel used for the dye. For cells containing a double pSynORI, the exposure times were set to 100 ms for both RFP and GFP fluorescence channels. Experimentally measured time course data were acquired and analyzed with ImageJ and MATLAB.
Nanopore sequencing and data analysis
For sample preparation, single colonies were picked from the overnight-incubated LB-agar plates and inoculated in 50 mL glass culture tubes with 5 mL liquid LB containing corresponding antibiotics at 37 °C, shaking at 225 rpm (LB–AB). For data shown in Fig. 4H, 100 μM IPTG and 200 μM cumate were also added to the liquid culture. Following overnight growth, DNA plasmids were isolated using a DNA mini plasmid preparation (Qiagen). The plasmid sample was then sent for nanopore sequencing (Plasmidsaurus).
Data were processed and displayed as follows. For quantification of multimer species in Figs. 2F, 3C, the mass ratio of multimers from the sequencing result was presented. Mass ratio indicates the distribution of nucleotides, as well as ORI copies, across the different multimer species. For multiplex plasmid reporting in Fig. 4H, the number of nanopore sequencing reads for each plasmid was directly acquired from Plasmidsaurus. For 6-plasmid characterization in Fig. 5C, the raw nanopore sequencing reads from Plasmidsaurus were processed with an alignment workflow (wf-alignment) on the EPI2ME (v5.2.3) platform using all 6 plasmids as the reference. The alignment result is then used to calculate the percentage of plasmid composition.
Statistics and reproducibility
No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper. NGS data are available at the NCBI accession PRJNA1399510 [http://www.ncbi.nlm.nih.gov/bioproject/1399510]. Source data are provided with this paper.
References
Gyles, C. L., Palchaudhuri, S. & Maas, W. K. Naturally occurring plasmid carrying genes for enterotoxin production and drug resistance. Science 198, 198–199 (1977).
MacLean, R. C. & San Millan, A. The evolution of antibiotic resistance. Science 365, 1082–1083 (2019).
Castañeda-Barba, S., Top, E. M. & Stalder, T. Plasmids, a molecular cornerstone of antimicrobial resistance in the One Health era. Nat. Rev. Microbiol 22, 18–32 (2024).
Kamruzzaman, M. & Iredell, J. A ParDE-family toxin antitoxin system in major resistance plasmids of Enterobacteriaceae confers antibiotic and heat tolerance. Sci. Rep. 9, 9872 (2019).
Williams, P. A. & Murray, K. Metabolism of benzoate and the methylbenzoates by Pseudomonas putida (arvilla) mt-2: evidence for the existence of a TOL plasmid. J. Bacteriol. 120, 416–423 (1974).
Ramos, J. L., Wasserfallen, A., Rose, K. & Timmis, K. N. Redesigning metabolic routes: manipulation of TOL plasmid pathway for catabolism of alkylbenzoates. Science 235, 593–596 (1987).
Moraïs, S. et al. Plasmid-encoded toxin defence mediates mutualistic microbial interactions. Nat. Microbiol 9, 108–119 (2023).
Virolle, C., Goldlust, K., Djermoun, S., Bigot, S. & Lesterlin, C. Plasmid transfer by conjugation in gram-negative bacteria: from the cellular to the community level. Genes 11, 1239 (2020).
Kriz, A. et al. A plasmid-based multigene expression system for mammalian cells. Nat. Commun. 1, 120 (2010).
Broothaerts, W. et al. Gene transfer to plants by diverse species of bacteria. Nature 433, 629–633 (2005).
Selzer, G., Som, T., Itoh, T. & Tomizawa, J. The origin of replication of plasmid p15A and comparative studies on the nucleotide sequences around the origin of related plasmids. Cell 32, 119–129 (1983).
Vieira, J. & Messing, J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene 19, 259–268 (1982).
Meyer, A. J., Segall-Shapiro, T. H., Glassey, E., Zhang, J. & Voigt, C. A. Escherichia coli “Marionette” strains with 12 highly optimized small-molecule sensors. Nat. Chem. Biol. 15, 196–204 (2019).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Chen, Y.-J. et al. Characterization of 582 natural and synthetic terminators and quantification of their design constraints. Nat. Methods 10, 659–664 (2013).
Sheth, R. U., Yim, S. S., Wu, F. L. & Wang, H. H. Multiplex recording of cellular events over time on CRISPR biological tape. Science 358, 1457–1461 (2017).
Joshi, S. H.-N., Yong, C. & Gyorgy, A. Inducible plasmid copy number control for synthetic biology in commonly used E. coli strains. Nat. Commun. 13, 6691 (2022).
Rouches, M. V., Xu, Y., Cortes, L. B. G. & Lambert, G. A plasmid system with tunable copy number. Nat. Commun. 13, 3908 (2022).
Son, H.-I. et al. Population-level amplification of gene regulation by programmable gene transfer. Nat. Chem. Biol. 21, 939–948 (2025).
Polisky, B. ColE1 replication control circuitry: Sense from antisense. Cell 55, 929–932 (1988).
Tomizawa, J. Control of colE1 plasmid replication. J. Mol. Biol. 212, 683–694 (1990).
Novick, R. P., Iordanescu, S., Projan, S. J., Kornblum, J. & Edelman, I. pT181 plasmid replication is regulated by a countertranscript-driven transcriptional attenuator. Cell 59, 395–404 (1989).
Lucks, J. B., Qi, L., Mutalik, V. K., Wang, D. & Arkin, A. P. Versatile RNA-sensing transcriptional regulators for engineering genetic networks. Proc. Natl. Acad. Sci. USA 108, 8617–8622 (2011).
Takahashi, M. K. & Lucks, J. B. A modular strategy for engineering orthogonal chimeric RNA transcription regulators. Nucleic Acids Res. 41, 7577–7588 (2013).
Shao, B. et al. Single-cell measurement of plasmid copy number and promoter activity. Nat. Commun. 12, 1475 (2021).
Yeung, E. et al. Biophysical constraints arising from compositional context in synthetic gene networks. Cell Syst. 5, 11–24.e12 (2017).
Johnstone, C. P. & Galloway, K. E. Supercoiling-mediated feedback rapidly couples and tunes transcription. Cell Rep. 41, 111492 (2022).
Bedbrook, J. Recombination between bacterial plasmids leading to the formation of plasmid multimers. Cell 9, 707–716 (1976).
Patient, M. E. & Summers, D. K. ColE1 multimer formation triggers inhibition of Escherichia coli cell division. Mol. Microbiol. 9, 1089–1095 (1993).
Field, C. M. & Summers, D. K. Multicopy plasmid stability: revisiting the dimer catastrophe. J. Theor. Biol. 291, 119–127 (2011).
Colloms, S. D., Sykora, P., Szatmari, G. & Sherratt, D. J. Recombination at ColE1 cer requires the Escherichia coli xerC gene product, a member of the lambda integrase family of site-specific recombinases. J. Bacteriol. 172, 6973–6980 (1990).
Easter, C. L., Schwab, H. & Helinski, D. R. Role of the parCBA Operon of the Broad-Host-Range Plasmid RK2 in Stable Plasmid Maintenance. J. Bacteriol. 180, 6023–6030 (1998).
Strobel, E. J., Cheng, L., Berman, K. E., Carlson, P. D. & Lucks, J. B. A ligand-gated strand displacement mechanism for ZTP riboswitch transcription control. Nat. Chem. Biol. 15, 1067–1076 (2019).
Bushhouse, D. Z. & Lucks, J. B. Tuning strand displacement kinetics enables programmable ZTP riboswitch dynamic range in vivo. Nucleic Acids Res. 51, 2891–2903 (2023).
Cheng, L. et al. Cotranscriptional RNA strand exchange underlies the gene regulation mechanism in a purine-sensing transcriptional riboswitch. Nucleic Acids Res. 50, 12001–12018 (2022).
Dudley, Q. M., Karim, A. S. & Jewett, M. C. Cell-free metabolic engineering: Biomanufacturing beyond the cell. Biotechnol. J. 10, 69–82 (2015).
Wright, O., Delmans, M., Stan, G.-B. & Ellis, T. GeneGuard: a modular plasmid system designed for biosafety. ACS Synth. Biol. 4, 307–316 (2015).
Kang, C. W. et al. Synthetic auxotrophs for stable and tunable maintenance of plasmid copy number. Metab. Eng. 48, 121–128 (2018).
Wagner, E. G. H. & Simons, R. W. Antisense RNA control in bacteria, phages, and plasmids. Annu. Rev. Microbiol. 48, 713–742 (1994).
Jain, A. & Srivastava, P. Broad host range plasmids. FEMS Microbiol. Lett. 348, 87–96 (2013).
Kolatka, K., Kubik, S., Rajewska, M. & Konieczny, I. Replication and partitioning of the broad-host-range plasmid RK2. Plasmid 64, 119–134 (2010).
Scherzinger, E. et al. Replication of the broad host range plasmid RSF1010: requirement for three plasmid-encoded proteins. Proc. Natl. Acad. Sci. USA 81, 654–658 (1984).
Meyer, R. Replication and conjugative mobilization of broad host-range IncQ plasmids. Plasmid 62, 57–70 (2009).
Bethke, J. H. et al. Environmental and genetic determinants of plasmid mobility in pathogenic Escherichia coli. Sci. Adv. 6, eaax3173 (2020).
Lee, C., Kim, J., Shin, S. G. & Hwang, S. Absolute and relative QPCR quantification of plasmid copy number in Escherichia coli. J. Biotechnol. 123, 273–280 (2006).
Alnahhas, R. N. Spatiotemporal Dynamics of Synthetic Microbial Consortia. Dissertation (Rice University, 2019).
Ferry, M. S., Razinkov, I. A. & Hasty, J. Microfluidics for synthetic biology: from design to execution. In Methods in Enzymology, vol. 497, 295–372 (Academic Press, 2011).
Acknowledgments
We would also like to thank Dr. Peter Davenport for helpful discussions and insights regarding this work. This work was supported by National Science Foundation grant 2124306 (J.C.), National Science Foundation grant 2237512 (J.C.), Welch Foundation grant A24-0270-001 (J.C.), National Science Foundation grant 2124308 (M.R.L.), and NSF/NIGMS mathematical biology program grant R01GM144959 (M.R.B.).
Author information
Authors and Affiliations
Contributions
B.L., J.C., and M.R.L. conceived the project. Experiments were performed by B.L., Z.R.D.S. and X.P. B.L. and J.C. wrote the paper, and Z.R.D.S., X.P., M.R.B., and M.R.L. provided feedback.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, B., Seet, Z.R.D., Peng, X. et al. Engineering plasmids with synthetic origins of replication. Nat Commun 17, 2255 (2026). https://doi.org/10.1038/s41467-026-68907-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-68907-1