Abstract
Germline mutations in the chromatin remodelling protein ATRX cause a severe developmental disorder associated with α-thalassemia. In addition, ATRX is amongst the twenty genes most frequently mutated in cancer. How ATRX mutations alter gene expression remains unclear. Using the α-globin locus as a model, here we show that ATRX deficiency downregulates α-globin in a subset of cells exhibiting DNA damage. A G-rich repeat at the α-globin locus serves as a potential site of G-quadruplex formation and DNA damage. ATRX binds this repeat co-transcriptionally, and its loss increases R-loop accumulation at this site, leading to local DNA damage and transcriptional disruption in cis. Deletion of this repeat abolishes this effect, while targeted DNA damage reinstates it. These findings reveal a mechanism linking ATRX’s role in genome stability to transcriptional regulation and uncover a molecular basis of human genetic disease mediated via a distal G-rich repeat.
Similar content being viewed by others
Introduction
The ATRX gene was first identified because when mutated it causes a specific, severe form of syndromic intellectual disability associated with alpha-thalassaemia: so-called alpha-thalassemia X-linked intellectual disability (ATR-X) syndrome1. ATRX thus plays a key role in normal development2, and it is also one of the twenty genes most frequently mutated in cancer3,4,5. ATRX is a highly conserved SWI/SNF chromatin remodelling protein, which forms a multiprotein complex with the histone variant H3.3 and its chaperone DAXX to maintain chromatin organisation6,7,8,9. ATRX plays a part in many nuclear processes including DNA replication, transcription, DNA damage and DNA repair10,11,12,13,14,15,16,17. Although previous studies have mapped ATRX binding to heterochromatin, CpG islands, and active regulatory elements18,19,20,21,22, the precise mechanisms by which ATRX regulates gene expression remain unclear. Notably, ATRX selectively affects human alpha-globin (α-globin), but not beta-globin (β-globin) expression during erythropoiesis -- a process requiring coordinated expression of α- and β-globin genes to form haemoglobin heterotetramers (α2β2), providing a tractable model to explore its regulatory function.
In marked contrast to the β-globin cluster, the human α-globin cluster lies within an early replicating, GC-rich, gene dense, sub-telomeric region of chromosome 16. The human α-globin genes are associated with prominent unmethylated CpG islands23,24 and are embedded within a region of the genome containing a high density of repeat sequences including several G-rich variable number tandem repeats (VNTRs) (Fig. 1a). In ATR-X syndrome, mutations in ATRX downregulate gene expression at the human α-globin locus, with the α-like globin gene HBM being most affected, followed by α-globin genes HBA2 and HBA119, suggesting a unique mechanism of gene regulation.
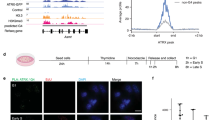
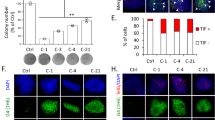
a Schematic representation of human α-globin locus on chromosome 16p13.3. From the telomeric end, the locus contains the ζ-globin gene HBZ, the minor μ-globin gene HBM, the two α-globin genes HBA2 and HBA1, and lastly the minor θ-globin gene HBQ1. Two pseudogenes are present: ψζ HBZP1 between HBZ and HBM, and ψα1 HBAP1 between HBM and HBA2. Four distal enhancers (R1-R4) are situated upstream of the genes. b Schematic overview of the experimental workflow to generate ATRX knockout erythroid cells from healthy donor CD34+ HSPCs, followed by downstream analysis. c Western blot analysis of ATRX expression in day 7 differentiated cells, confirming efficient knockout. Quantification is presented on the right (n = 3 biological replicates, data shown as mean ± SEM). **p = 0.009, two-tailed unpaired Welch’s t-test. d Relative expression of HBA, HBM and α to β-like globin ratios by RT-qPCR analysis in day 10 and day 13 differentiated erythroid cells (n = 3 biological replicates, data shown as mean ± SEM). Differences between AAVS1 and ATRX KO were compared using a two-tailed unpaired Welch’s t-test on log₂-transformed fold-change values. e Genotyping summary of the collected 351 BFU-E colonies derived from ATRX KO samples showing genotype distribution. f Three-dimensional plot showing Biomark gene expression analysis of ATRX, HBA and HBM in single BFU-E colony. Each dot represents one colony, and the edited AAVS1 colonies are shown in blue; ATRX KO colonies with frameshift mutations (ATRX_FS) are shown in red. AAVS1 controls gather around the internal space whereas the majority of the ATRX KO colonies assemble around the origin indicating low expression of ATRX, HBM and HBA. g Quantified gene expression analyses of BFU-E colonies show statistical differences between AAVS1 (n = 15) and ATRX KO groups (n = 57) (Mann-Whitney U test). Individual colonies are selected from two biological repeats. ***p = 1.493 × 10-5 for ATRX expression, **p = 0.003632 for HBM expression, and p = 0.9083 for HBA expression. ns: not significant. Source data are provided as a Source Data file.
In this study, we investigate how ATRX loss leads to α-thalassemia, a key disease phenotype of ATR-X syndrome, by elucidating the mechanisms that link ATRX’s gene regulatory functions to human disease. Here we show that dysregulation of α-globin expression in the absence of ATRX depends on a G-rich VNTR within the human α-globin locus. Loss of ATRX promotes G-quadruplex (G4) and R-loop formation at this site, resulting in DNA damage and distance-dependent disruption of gene expression. Removal of this VNTR abolished the gene downregulation caused by ATRX loss and inducing DNA damage at this site restored the effect. Together, our findings uncover a systematic mechanism of ATRX-mediated gene regulation, highlighting its role in orchestrating genome stability and transcription regulation. In addition, by directly linking molecular dysfunction at tandem repeats to human pathology, our work reveals a pathogenic mechanism underlying human genetic disease, underscoring the clinical significance of genome stability and chromatin dynamics at repetitive loci.
Results
ATRX loss perturbs α-globin expression in a subset of cells
To unravel the underlying mechanism by which ATRX regulates α-globin expression, we established an ATRX-deficient model by inducing a knockout (KO) of ATRX, using CRISPR-Cas9, in CD34+ haematopoietic stem and progenitor cells (HSPCs) from healthy male donors. Cells were electroporated with ribonucleoprotein (RNP, Cas9 protein complexed with guide RNA) targeting either ATRX, or as a control, the safe-harbour locus AAVS1. These CD34+ cells were then differentiated into erythroid cells using a three-phase differentiation system25 for genotype and phenotype analysis (Fig. 1b). Genotype analysis detected an average of 84.7% indels and 72% of knockout (out-of-frame) effect (Supplementary Fig. 1a), and the abolishment of ATRX was confirmed at the protein level (Fig. 1c). Consistent with ATRX’s role in replication10,14,15, ATRX KO cells showed slightly slower cell proliferation (Supplementary Fig. 1b). Phenotypically, ATRX KO cells demonstrated normal erythroid differentiation (Supplementary Fig. 1c) and cellular morphology (Supplementary Fig. 1d). Haemoglobin analysis revealed a similar pattern to the controls, with dominant adult haemoglobin A (HbA) expression and minor fetal haemoglobin (HbF) expression (Supplementary Fig. 1e). As in the peripheral blood of many patients with ATR-X syndrome, ATRX KO cells only showed a very mild (non-significant) reduction in HBM expression (Fig. 1d) and no erythroid cells with haemoglobin H (HbH, β4) inclusions were found2.
The absence of an α-thalassemia phenotype in ATRX KO cells may be due to the ATRX defect affecting only a small subset of cells, which is often obscured in a bulk population analysis. To address this, early erythroid progenitors (EEP) cells were sorted (Supplementary Fig. 1f) and plated for a colony-forming unit (CFU) assay, which allowed us to determine the genotype and the pattern of gene expression in individual erythroid colonies. ATRX KO sample showed similar haematopoietic progenitor frequencies (Supplementary Fig. 1g) and colony-forming capacity (Supplementary Fig. 1h-i) as the AAVS1 control. Single burst-forming unit-erythroid (BFU-E) colonies were then manually picked and analysed for their genotype and gene expression. A total of 96 BFU-E colonies from the AAVS1 sample and 351 colonies from the ATRX KO sample were analysed. Out of the 351 collected colonies, 240 (68.38%) had frameshift (FS) indels (Fig. 1e). Fifty AAVS1 “edited” colonies and 190 colonies from the ATRX sample harbouring ATRX frameshift (FS) indels were randomly selected for Biomark gene expression analysis. This showed reduced ATRX expression in most ATRX_FS colonies, however, only a subset showed decreased HBM and HBA expression (Fig. 1f). As expected from previous work19, HBM was more affected than HBA (Fig. 1g). Together, these results indicate that as in the peripheral blood of ATRX patients20, loss of ATRX affects gene expression at the α-globin locus in only a subset of cells, and HBM is more affected than HBA.
scRNA-seq links the gene perturbation to DNA damage response
To uncover the associated molecular signatures in the affected cells and elucidate ATRX’s regulatory mechanisms, we next performed single-cell RNA sequencing (scRNA-seq) on AAVS1 and ATRX KO cells. Characterisation during erythroid differentiation identified day 13 as optimal for single-cell analysis, in that ATRX KO cells showed pronounced HBM downregulation with a 77% knockout efficiency (Supplementary Fig. 2a-c). Accordingly, single day 13 nucleated erythroid cells were sorted for Smart-seq2 analysis (Supplementary Fig. 2d). After quality filtering and identification of highly variable genes (HVGs), principal component analysis (PCA) and t-distributed stochastic neighbour embedding (t-SNE) were performed. Both analyses indicated a closely overlapping distribution pattern between ATRX KO cells and the controls (Fig. 2a, Supplementary Fig. 2e). Differential gene expression analysis identified only 53 differentially expressed genes (DEGs) that were significantly downregulated, including HBM, and 18 upregulated genes (Fig. 2b, full list provided in Supplementary Data 1).
a t-SNE visualisation of scRNA-seq data showing individual cells in AAVS1 and ATRX KO samples using HVGs. b Volcano plot of differentially expressed genes in ATRX KO versus AAVS1 control. Each dot represents one gene. The horizontal dashed line indicates a significance threshold of 0.05, and the vertical dashed lines bound the minimal log2 fold-change (absolute value > 0.5) for the most-differentially-expressed genes. Upregulated genes are shown in red, downregulated genes in blue. The downregulated HBM is highlighted in blue and upregulated RNF168 is highlighted in pink. c Violin plot of RNF168 expression in both samples, with each dot representing a single cell. Percentages below sample labels show the number and proportion of RNF168-expressing cells. d Representative immunofluorescence images and quantification of γH2AX foci in day 13 AAVS1 and ATRX KO samples. Scale bar = 10 μm. 250 cells were randomly selected for quantification and lines show mean ± SEM. A two-tailed Mann-Whitney U test was used for statistical analysis. *p = 0.0111 for number of foci per nucleus, ****p < 0.0001 for median intensity. e Expression of globin genes (HBM, HBA, HBB, HBG) in both samples. Red dashed frames highlight a subpopulation of ATRX KO cells with low HBM expression. f Single-cell expression of HBB and HBM in AAVS1 (top) and ATRX KO (bottom) samples, coloured by HBM expression level, showing a low-HBM subpopulation in ATRX KO sample. g Upregulated pathways in affected ATRX KO HBMlow cells by GO analysis, emphasising histone H2A ubiquitination. x-axis indicates p-value; dot size represents the number of DE genes attributed to each GO term; and dot colour indicates term abundance. h Representative immunofluorescence images and quantification of H2Aub intensity in day 13 AAVS1 and ATRX KO samples. Scale bar = 10 μm. A total of 1000 cells were quantified, lines show mean ± SEM (****p < 0.0001, two-tailed Mann-Whitney U test). i Violin plot of UIMC1 expression in ATRX KO HBMhigh, HBMmid, and the affected HBMlow cells. Source data are provided as a Source Data file.
Gene Ontology (GO) analysis of these DEGs highlighted pathways related to cell cycle, cell division, and erythrocyte function (Supplementary Fig. 2f), consistent with ATRX’s roles in facilitating DNA replication10,14,15 and the observation of slower proliferation of ATRX KO cells in culture (Supplementary Fig. 2a). Of interest, RNF168, a key DNA damage response regulator, was significantly upregulated in ATRX KO cells (Fig. 2b-c), suggesting increased DNA damage in the absence of ATRX. The MRN-ATM-γH2AX-RNF168 pathway is essential for DNA damage response and repair26. ATRX has previously been shown to interact with the MRN complex to overcome replication stress and facilitate DNA repair14. To assess DNA damage in ATRX KO cells, we performed immunofluorescence staining for phosphorylated H2AX (γH2AX) and observed an increased number and intensity of γH2AX foci compared to the controls, indicating elevated DNA damage following ATRX loss (Fig. 2d).
To look into the transcriptomic signature of the affected subpopulation, cells were categorised based on globin gene expression. As a group, the ATRX KO sample showed similar levels of α-globin (HBA), β-globin (HBB), and \(\gamma\)-globin (HBG) expression to AAVS1 control but significantly reduced α-like-globin HBM expression (Fig. 2e). However, scatter plots revealed a subpopulation of ATRX KO cells at quiescent G1 phase with low α(-like)-globin expression (HBM and HBA) but normal HBB expression (Fig. 2f, Supplementary Fig. 3a-b), indicating this subpopulation of cells exhibited α-thalassemia. This enables us to group cells into HBMlow, HBMmid and HBMhigh based on their HBM expression (HBMlow < 2.5, HBMhigh > 7) (Supplementary Fig. 3c), with HBMlow exhibiting more separated clustering in the tSNE plot (Supplementary Fig. 3d). DEG and GO analysis between HBMhigh and HBMlow cells (Supplementary Fig. 3e) identified 56 downregulated pathways exclusively in the HBMlow ATRX KO cells (Supplementary Fig. 3f, Supplementary Data 2). The analysis also indicated 5 upregulated pathways exclusively in the affected HBMlow ATRX KO cells (Supplementary Data 2), including mitochondrial dysfunction, cell responses, and importantly, histone H2A ubiquitination pathways (Fig. 2g). Interestingly, histone H2A is the primary histone substrate of RNF168 and closely associated with transcriptional silencing27. To validate whether H2A ubiquitination is dysregulated in ATRX KO cells, immunofluorescence staining for ubiquitinated H2A (H2Aub) was performed. A subset of ATRX KO cells exhibited increased H2Aub intensity, indicating activation of the DNA damage response pathway (Fig. 2h). Consistently, UIMC1 (encoding RAP80), a ubiquitin-binding protein that recognises ubiquitinated histones at DNA damage sites28, was significantly upregulated in the affected ATRX KO HBMlow subpopulation (Fig. 2i, Supplementary Fig. 3g). Collectively, these findings suggest ATRX loss leads to increased DNA damage and enhanced RNF168-mediated H2A ubiquitination, which may contribute to the repression of genes lying within or adjacent to regions of DNA damage.
ATRX regulates gene expression at the α-globin locus via a G-rich VNTR
A potential site of DNA damage at the human α-globin locus is a highly repetitive VNTR region with 96% GC content, located within the second intron of the pseudogene HBZP1. This is located approximately 1 kb upstream of HBM, 8 kb upstream of HBA2, 12 kb upstream of HBA1 and is referred to as the pseudo-zeta (ψζ) VNTR (Fig. 1a). This VNTR comprises highly G-rich repeats (CGCGGGGCGGGG)n that can form G4 structures in vitro19,29, making it a prime candidate for causing DNA damage at the α-globin locus in the absence of ATRX. Of importance, we have previously shown that the size of ψζ VNTR positively correlates with the degree of α-globin downregulation in patients with ATR-X syndrome19. This led us to hypothesise that the ψζ VNTR acts as a transcriptional modulator of human α-globin expression regulated via ATRX. Of interest, this VNTR is absent in the mouse genome and ATRX deficiency in mouse models of ATR-X syndrome do not have α-thalassaemia30.
Using a well-established human model to study erythropoiesis (HUDEP-2 erythroid progenitor cell line), we tracked ATRX binding at the ψζ VNTR during erythroid differentiation. ATRX ChIP-seq demonstrated that ATRX starts to bind at the ψζ VNTR on day 4 of differentiation coincident with the onset of transcription of the zeta (ζ) pseudogene HBZP1 and α-like-globin gene HBM (Fig. 3a-c). This binding pattern, correlated with transcriptional activation, aligns with prior evidence of ATRX associating with G-rich sequences when they are transcribed31. To study the influence of the ψζ VNTR on α-globin gene expression, we first generated an inducible ATRX degron system in HUDEP-2 cells (Fig. 3d) and subsequently knocked out the ψζ VNTR. Characterisation of the ATRX degron cell line indicated normal erythroid differentiation, even under dTAG-13 treatment (Supplementary Fig. 4a-b). dTAG-13 induced rapid and potent degradation of ATRX (Fig. 3e, Supplementary Fig. 4c). As expected, this was associated with a significantly reduced level of HBM expression and sightly decreased HBA expression (Fig. 3f, Supplementary Fig. 4d), faithfully mimicking the mild α-thalassaemia phenotype seen in ATR-X syndrome. To test whether the degradation of ATRX downregulates α-globin expression via an effect on the ψζ VNTR, we next generated a HUDEP-2 cell line in which the ψζ VNTR was deleted from both alleles via CRISPR-Cas9 (Supplementary Fig. 5a) and repeated the dTAG experiment. The VNTR KO line showed normal erythroid differentiation (Supplementary Fig. 5b-c), and ATRX can be potently degraded by dTAG-13 (Fig. 3e). Interestingly, in striking contrast to the previous experiment, when the ψζ VNTR was removed, HBM was no longer downregulated in the absence of ATRX (Fig. 3f), showing that downregulation of HBM expression in the absence of ATRX is mediated via the G-rich ψζ VNTR.
a mRNA expression analysis of HBM by qRT-PCR during erythroid differentiation, normalised to RPS18, data are shown as mean ± SEM (n = 3 biological replicates). b cDNA PCR amplification of the ζ-globin gene HBZ and its pseudogene HBZP1 where the ψζ VNTR resides, in HUDEP-WT cells. The red arrows point the active and increasing transcription of the pseudogene HBZP1 during erythroid differentiation. c Representative ChIP-seq tracks showing ATRX occupancy at the ψζ VNTR-HBM locus across different stages of erythroid differentiation. The black blocks above peaks within each track show the peak call. d Schematic representation of the generation of the ATRX degron cell line. e ATRX expression in ATRX degron cells (left) and ATRX degron-VNTR KO cells (right) during erythroid differentiation, demonstrating dTAG-13-induced ATRX depletion. f Expression of HBM and HBA to β-like (HBB + HBG) globin ratios on differentiation day 8 cells. ATRX + refers to ATRX degron cell treated with 100 nM of dTAG-13-Neg (a negative control compound for dTAG-13) from day 0 to day 8; ATRX - represents ATRX degron cell treated with 100 nM of dTAG-13 from day 0 to day 8; ATRX + / VNTR KO is ATRX degron-ψζ VNTR KO cell treated with dTAG-13-Neg from day 0 to day 8; ATRX - / VNTR KO is ATRX degron-ψζ VNTR KO cell treated with dTAG-13 from day 0 to day 8. Comparisons between two groups were performed using a two-tailed unpaired Welch’s t-test on log₂-transformed fold-change values. Data are shown as mean ± SEM (n = 5 biological replicates). **p = 0.0047 for HBM to β-like ratio between ATRX+ and ATRX- samples. Source data are provided as a Source Data file.
G4s, R-loops, and DNA damage at G-rich VNTRs drives gene dysregulation
The ψζ VNTR is predicted to form highly stable G4 structures (G4Hunter32,33 score > 2) (Supplementary Fig. 6a). Treating cells with G4 stabilisers during day 6 to 8 of differentiation revealed that CX-5461 specifically downregulated HBM expression to levels comparable to that seen when ATRX is absent, but had little effect on the β(-like)-globin genes HBB and HBG (Fig. 4a, Supplementary Fig. 6b). A similar downregulation of gene expression was observed at another ATRX target gene, NME4 (Supplementary Fig. 6b), which also contains a G-rich VNTR predicted to form a highly stable G4 structure (G4Hunter score > 2) (Supplementary Fig. 6c). To assess whether this represents a broader phenomenon, we examined the 71 genes dysregulated in CD34 ATRX knockout cells for the presence of GC-rich repetitive elements. Notably, 45 of 71 genes (63.4%) contained GC-rich tandem or low-complexity repeats (Supplementary Data 1), which is significantly higher than the genome-wide background rate of such repeats (23.3%, analysis based on the UCSC RepeatMasker track, p < 1 × 10-15; one-tailed binomial test). This suggests a general mechanism by which the absence of ATRX perturbs the expression of genes associated with such sequences. Consistent with this, when the ψζ VNTR was deleted, CX-5461 no longer had effect on HBM expression (Fig. 4a) but retained its effect on NME4 expression (Supplementary Fig. 6b), implicating the formation of G4 structures at the ψζ VNTR in HBM regulation.
a Expression of HBM and HBA to β-like (HBB + HBG) globin in day 8 ATRX degron and ATRX degron-VNTR KO cells following 48 h treatment with G4 stabiliser CX-5461. NT: untreated; dTAG-13: 100 nM from day 0 to day 8; CX-5461: 0.1 μM from day 6 to day 8. Statistical analysis within each cell line used one-way ANOVA followed by Dunnett test on log₂-transformed values (n = 3 biological replicates, data shown as mean ± SEM). For HBM to β-like ratio, *p = 0.0267 (NT vs dTAG-13); **p = 0.0025 (NT vs CX-5461). b S9.6 dot-blot assay measuring global R-loops in dTAG-13-neg-treated ATRX+ and dTAG-13-treated ATRX- cells. RNase H served as a negative control. Quantification is shown as mean ± SEM from three independent experiments. *p = 0.0454 for 200 ng, two-way ANOVA Šídák’s test. c DRIP-qPCR analysis showing increased R-loop levels around the VNTR region in ATRX-negative cells, but not at a distal region on chromosome 16 (n = 3 biological replicates, data shown as mean ± SEM; *p = 0.0422, two-way ANOVA Šídák’s test). d Expression of HBM to β-like globin following RNase H overexpression in ATRX-negative cells. (n = 3 biological replicates, data shown as mean ± SEM; **p = 0.0084, one-way ANOVA Dunnett test). e Schematic of the ψζ VNTR locus and the sgRNA target sites used in the CRISPR-Cas9-induced DNA damage assay in VNTR KO cells. f Expression of HBM and HBA to β-like globin in day 8 ATRX degron-VNTR KO cells after 48 h CRISPR assay (n = 3 biological replicates, data shown as mean ± SEM). AAVS1 vs sgRNA used one-way ANOVA Dunnett test, and ARTX+ vs ATRX- used a two-tailed unpaired Welch’s t-test. For HBM to β-like ratio, **p = 0.0067, 0.0013, 0.0013 for AAVS1 vs sgRNA-1, sgRNA-2, sgRNA-3 in VNTR KO-ATRX+ cells; **p = 0.0058, 0.0053 for AAVS1 vs sgRNA-2, sgRNA-3 in VNTR KO-ATRX- cells. *p = 0.0332 for ATRX+ vs ATRX- cells. For HBA to β-like ratio, **p = 0.0057, 0.0022, 0.0026 for AAVS1 vs sgRNA-1, sgRNA-2, sgRNA-3 respectively in VNTR KO-ATRX+ cells. Source data are provided as a Source Data file.
It has previously been shown that ATRX binds to known G4 structures when they are transcribed and that accumulate R-loops which stabilise such structures31,34. In line with this, ATRX-deficient cells displayed a genome-wide increase in R-loops (Fig. 4b). DNA:RNA IP (DRIP)-qPCR further revealed significantly elevated R-loop formation at the ψζ VNTR locus in the absence of ATRX (Fig. 4c). Interestingly, overexpression of RNase H in ATRX-deficient cells partially restored HBM downregulation (Fig. 4d), supporting a functional role for R-loops in regulating HBM transcription. Accumulated G4 structures and R-loops are known to cause replicative stress and DNA damage, leading to perturbed chromatin and local transcriptional silencing35,36,37. Hence, we proposed that ATRX depletion induces the formation of unusual secondary structures at the ψζ VNTR, which triggers DNA damage and downregulates the nearby α(-like)-globin genes (HBM and HBA).
To test whether DNA damage alone could drive HBM dysregulation, we introduced local DNA double-strand breaks (DSBs) upstream of HBM via CRISPR-Cas9 targeting in the region where the ψζ VNTR had been deleted (VNTR KO cells) (Fig. 4e). Interestingly, this artificially induced DNA damage led to marked downregulation of HBM and, to a lesser extent, HBA expression (Fig. 4f). This suggests that downregulation of HBM and HBA gene expression and the consequent alpha-thalassaemia occurs in cis as a result of DNA damage at the ψζ VNTR. It has been previously reported that transcription is highly repressed at sites adjacent to DSBs, and that this repression decreases upon movement away from the lesion38,39, as shown at the α-globin locus (HBM > HBA2 > HBA1)19. The mechanism appears to be complex40, but the α-globin locus model presented here offers the opportunity to solve this in detail.
Discussion
ATRX has long been recognised as an important genomic guardian, with its loss associated with increased genomic instability and transcriptional dysregulation. Previous studies have established that ATRX deficiency leads to DNA replication defects10,14,41 and promotes the accumulation of non-canonical DNA structures, such as G-quadruplexes (G4s)29,35,42,43 and R-loops31,34. However, these investigations have largely examined individual aspects of ATRX function in isolation, the molecular mechanisms connecting ATRX’s chromatin remodelling activity to gene regulation therefore remain incompletely resolved. In this study, we leverage the α-globin gene cluster, a well-characterised target of ATRX, to establish an integrated model that unifies these mechanistic elements. Through this, we demonstrate how ATRX orchestrates non-canonical genomic structures, genome stability, and transcriptional output within a physiological context, thereby elucidating how ATRX deficiency leads to transcriptional dysregulation.
We show that loss of ATRX downregulates α-globin expression in only a subset of cells exhibiting elevated levels of DNA damage. Using an inducible ATRX degron system, we found that ATRX normally binds to a G-rich VNTR when the region is transcribed, and prevents or resolves the formation of R-loops. In the absence of ATRX, persistent R-loops accumulate at the transcribed VNTR, triggering DNA damage that consequently disrupts gene expression in cis (Fig. 5). Although G4 structures and R-loops may also form at other G-rich regions within the α-globin locus including the ζ-VNTR44, the close proximity and genetic linkage of α-thalassaemia to the ψζ VNTR19 make ψζ VNTR the leading candidate for the effect proposed here. Of note, VNTRs are inherently variable in size, and such variation is likely to influence their propensity to form G4s and R-loops, which in turn correlates with gene expression, as reported in ATR-X syndrome19. Assessing the impact of VNTR length on R-loop formation and chromatin dynamics in appropriate model systems would provide valuable mechanistic insight.
ATRX binds the G-rich ψζ VNTR region during transcription to facilitate proper gene expression. In the absence of ATRX, this G-rich region forms G4 structures and R-loops, which are prone to inducing DNA damage and thereby disrupting nearby gene expression in cis. The closest gene HBM is most strongly affected, followed by the more distal HBA2/1 genes. Created in BioRender. Shen, Y. (2026) https://BioRender.com/bq12e2p.
While this mechanism is exemplified at α-globin, ATRX binds G-rich regions genome-wide (see examples in Supplementary Data 1), many of which are actively transcribed and prone to forming G4s and R-loops19,35,42,45. Our findings therefore suggest that ATRX-dependent suppression of G4s and R-loops associated with actively transcribed regions is likely to be a general mechanism by which gene expression is maintained without interruption from the secondary consequences of unresolved G4 DNA structures. Disruption of this pathway of maintaining genome integrity may help explain aspects of the developmental defects observed in ATR-X syndrome and raise the possibility that similar processes could contribute to other phenotypes associated with ATRX mutations in ageing and cancer, where genomic instability and epigenetic deregulation are hallmarks of disease progression.
Consistent with ATRX’s role in DNA replication10,14,15,41, we observed that ATRX-deficient erythroid cells exhibit upregulated DNA damage and activated DNA damage response pathways (Fig. 2d, g-i). ATRX has been shown to interact with the MCM helicase complex, a key component of the replication machinery35. Direct visualisation study revealed that G4s form immediately behind the MCM helicase at replication forks, impeding DNA synthesis and inducing replicative stress46. These findings suggest that ATRX may collaborate with MCM to resolve transient G4 structures during replication. In support of this, Teng et al. have shown that the helicase domain of ATRX is required to protect cells from G4-induced replicative stress35. Loss of ATRX compromises this protective mechanism, leading to G4 accumulation, fork stalling, replication stress, and DNA damage.
ATRX also suppresses aberrant nucleic acid structures during transcription. G-rich DNA regions are particularly prone to forming G4s and R-loops during transcription, which may interfere with transcription and replication. ATRX is enriched at such transcribed G-rich loci, where its loss results in increased G4 and R-loop accumulation31,42. Overexpression of RNase H in ATRX-deficient cells partially rescued HBM downregulation (Fig. 4d), suggesting that ATRX modulates R-loops and G4 structures through at least partially independent mechanisms. Given the high GC content (96%) of the ψζ VNTR, G4 formation likely occurs intrinsically and both G4 and R-loop formation are enhanced during transcription47 in ATRX-deficient cells, thereby compromising genome stability. Recent reports also indicate a role of R-loops in facilitating DNA damage repair around DSBs, independent of sequence composition48,49. Although R-loop formation at the α-globin locus is primarily driven by transcriptional activity and the GC richness, it remains an open question whether ongoing DNA damage further promotes R-loop accumulation. Investigating this potential feedback mechanism represents an important direction for future research.
Persistent G4 and R-loop structures may lead to replication-transcription collisions, ultimately resulting in DNA damage and activating DNA damage response pathways. Among the two primary DNA damage response kinases (ATM and ATR) ATM appears to play a more dominant role in responding to ATRX loss-induced damage. In the Atrx-null mouse embryonic brain, phosphorylated ATM is elevated while ATR levels remain unchanged50. ATM signalling pathway is consistently activated in ATRX-deficient cells15,22,51. In addition, ATRX promotes the recovery from replication stress by interacting with the MRN complex10,14, a sensor of DSB that activates ATM52. Activation of this pathway leads to phosphorylation of ATM and downstream targets such as RNF16826, a ubiquitin ligase that marks damaged chromatin by ubiquitinating histone H2A, which could be recognised by UIMC1 (RAP80)28 thereby recruiting DNA repair proteins such as BRCA1 and 53BP1. Consistent with this pathway, we observed upregulation of RNF168, increased H2A ubiquitination, and elevated UIMC1 expression in ATRX-deficient cells (Fig. 2c, g-i).
Importantly, this pathway does not merely facilitate DNA repair but also has direct impact on gene expression. Increased RNF168 recruitment and histone ubiquitination have been observed at DNA damage and G4 sites, and this DNA damage response-related ubiquitin signalling pathway is frequently altered in cancers with aberrant gene expression53,54. Ubiquitination at sites of damage has been linked to transcriptional silencing of nearby genes39. Thus, activation of this pathway may contribute to gene repression at loci like α-globin in the context of ATRX deficiency. In addition, the histone variant macroH2A1 is one important factor in ATM-dependent DNA damage response. Both variants, macroH2A1.1 and macroH2A1.2, are recruited to DNA damage sites and promote local chromatin condensation55,56. Importantly, macroH2A1.2 has been shown to be an epigenetic link between ATRX loss and DNA damage response at sites of recurrent replication stress57,58, and macroH2A1 deposition at α-globin has been shown to be involved in ATRX-mediated α-globin regulation59. The H2A ubiquitination and macroH2A1 deposition60 might be complementary pathways in α-globin regulation, particularly both are associated with DNA damage responses and repressive chromatin states61. Further study is needed to fully elucidate whether these two pathways act synergistically to regulate gene expression at damaged genomic loci.
As shown here, ATRX deficiency exerts these effects on only a subset of cells, suggesting that the consequences of ATRX loss accumulate in a stochastic and progressive manner. This is supported by the BFU-E colony assays in which only a fraction of colonies displayed gene downregulation (Fig. 1f), indicating that impairments in this subpopulation arise cumulatively during otherwise normal erythroid differentiation. Specifically, only cells that accumulate unresolved G4s/R-loops and replication stress are to be affected. The likelihood of a replication or transcription fork encountering these structures during a dynamic process introduces a stochastic element to the effect. Notably, ATRX is not the sole factor capable of destabilising G4s and R-loops; other helicases, such as FANCJ DNA helicase, RecQ helicases, and DDX5, can also resolve these structures to release the cells from stress46,62,63. Therefore, the affected subpopulation likely represents those in which G4s and R-loops persist because ATRX is absent and other complementary helicases fail to resolve these structures in time. With successive rounds of cell division and differentiation, these unresolved structures are predicted to accumulate, causing progressive DNA and chromatin impairment, ultimately suppressing gene expression. The stochastic nature of this phenomenon presents a technical challenge for detecting α-globin locus-specific DNA damage and chromatin changes. A more robust model that exhibits a stronger and more consistent phenotype would be necessary to explore these aspects further, which we identify as an important avenue for future research. Likewise, assessing VNTR length heterogeneity at the single-cell level would help reveal whether the affected subset of cells carries longer repeats during differentiation. Although determining VNTR length in individual cells or single BFU-E colonies remains technically challenging, it represents another direction for future investigation.
Cell-cycle analysis revealed that the affected cells were predominantly in the G1 phase (Supplementary Fig. 3b). It is known that unrepaired DNA damage from G2/M phases is resolved during G1 through 53BP1 nuclear bodies, which often localise at chromosomal fragile sites60. ATRX has been reported to colocalise with 53BP1 nuclear bodies in G1 to facilitate the repair of these fragile regions8. In our single-cell transcriptomic data, TP53BP1 expression was modestly increased in ATRX KO cells, though not significantly, supporting a potential link between ATRX loss and G1-phase repair activity. Collectively, these data suggest that ATRX-deficient cells accumulate DNA damage at fragile sites that require G1-phase repair, resulting in the observed enrichment of cells in this phase.
In summary, our findings highlight the interconnected role of ATRX at G-rich loci in safeguarding genome stability and maintaining transcriptional fidelity, providing a coherent mechanism linking ATRX deficiency to human disease. The working model presented here not only bridges a key mechanistic gap in understanding ATRX function but also implicates clinical relevance. This mechanism accounts for several previously unexplained observations in ATR-X syndrome: the mild α-thalassemia in ATR-X syndrome (affecting only a subset of cells), the phenotypic heterogeneity among patients (distance- and VNTR size-dependent gene dysregulation) observed in patients19, and the absence of α-thalassemia in ATRX-deficient mouse models which do not harbour the ψζ VNTR30. Together, these findings uncover a pathogenic mechanism of perturbed gene expression in ATR-X syndrome mediated by a distal VNTR. More broadly, they demonstrate how subtle underlying perturbations can lead to profound disease consequences.
Methods
CD34+ cell culture and differentiation
The CD34+ cells were separated from healthy adult peripheral blood leucocyte cones purchased from the National Health Service Blood and Transplant (NHSBT), UK. Ethical approval was granted by the Wales Research Ethics Committee (REC5) (13/WA/0371). The mononuclear cells were isolated using Histopaque-1077 (Sigma, Cat No H8889), and CD34+ cells were purified using Human CD34 Microbead kit (Miltenyi Biotec, Cat No 130-046-702) following the manufacturer’s instruction and the purity was confirmed by by FACS. The CD34+ cells were cultured in StemSpan SFEM II media (StemCell Technologies, Cat No 9655) supplemented with 100 ng/mL SCF (PeproTech, Cat No 300-07), 100 ng/mL TPO (PeproTech, Cat No 300-18), 100 ng/mL Flt3L (PeproTech, Cat No 300-19) and 1x Penicillin/Streptomycin (P/S, Gibco, Cat No 15070063) for 24 hours at 37 °C with 5% CO2 incubation prior to electroporation. After electroporation, the cells were recovered in the same media without P/S for 24 hours before starting erythroid differentiation.
CD34+ cells were differentiated along the erythroid lineage using a previously described three-phase liquid culture system25, requiring a base media Iscove’s Modified Dulbecco’s Medium (IMDM) (Gibco, Cat No 31980022) containing 3% Solvent/Detergent pooled Human AB plasma (NHSBT), 10 μg/mL human insulin (Merck, Cat No I9278), 3 U/mL heparin (Merck, Cat No H3149), 3 U/mL erythropoietin (EPO, Oxford University Hospital (OUH) pharmacy), 200 μg/mL iron saturated human holo-transferrin (Bio-Techne, Cat No 2914-HT), 2% (v/v) fetal bovine serum (FBS) and 1x P/S. In phase I (day 0 to day 7), 1 ng/mL IL3 (PeproTech, Cat No 200-03) and 10 ng/mL SCF were added. In phase II (day 7 to day 11), the IL3 was removed and only 10 ng/mL SCF was supplemented. In phase III (post day 11), only 300 μg/mL holo-transferrin was added. The media was changed every two days to maintain cell concentration around 1-2 × 105 cells/mL in phase I and phase II, 0.5-1 × 106 cells/mL in phase III.
CRISPR-Cas9 knocking out experiment in CD34+ cells
Given that ATRX is located on the chromosome X and ATR-X syndrome only affects males, donor gender was confirmed by checking the SRY gene on chromosome Y. Only CD34+ cells derived from the male donors were used for CRISPR experiment to achieve efficient ATRX knockout given the ATRX gene is located on chromosome X. 1-3 × 105 of CD34+ cells were transfected with RNPs (3.2 μg sgRNA (synthesised by Synthego) + 6 μg SpCas9 nuclease (IDT)) using Amaza P3 Primary cell 4D-Nucleofector X Kit (Lonza, Cat No V4XP-3032) and 4D-Nucleofector X Unit device (Lonza). Guide RNA sequences are listed in Supplementary Table 1. Cells were recovered in pre-warmed antibiotic-free SFEM II complete media right after the electroporation for 24 hours before the initiation of differentiation or cell sorting. ATRX knockout efficiency was assessed by Sanger sequencing of the targeted region using primers listed in Supplementary Table 1, followed by Synthego Inference of CRISPR Edits (ICE) analysis.
Colony forming unit assay
CFU assay was performed following the technical manual for Human CFU Assays.
Using MethoCult (STEMCELL Technologies, Cat No H4435). Single-cell suspensions were prepared and diluted in Iscove’s MDM supplemented with 2% FBS medium (STEMCELL Technologies, Cat No 07700) and then mixed into enriched MethoCult (STEMCELL Technologies, Cat No H4435) to allow progenitor cells to expand and differentiate into colonies under incubation at 37 °C with 5% CO2. After 14-16 days of culture, colonies were examined and scored based on their morphology. Then the single individual BFU-E colonies were manually picked under a light inverted EVOS XL Imaging System (ThermoFisher Scientific, Cat No AME3300) and transferred into tubes containing cold PBS. Each collected colony sample was subsequently divided into two aliquots - one for genotyping after whole genome amplification and the other for gene expression analysis using the Fluidigm Biomark system.
Analysis of single BFU-E colonies
Genomic DNA from single BFU-E colonies was amplified directly from cell lysate using llustra Single Cell GenomiPhi DNA Amplification Kit (GE Healthcare, Cat No 25-6600-30) following the manufacturer’s instructions. Then PCR amplification of CRISPR target regions and subsequent Sanger sequencing were conducted using the purified DNA to determine their genotype.
For gene expression analysis, samples were mixed with 2x Reaction Mix containing Superase-In and cDNA was pre-amplified by adding 0.2x Taqman assay mix and RT/Taq mix (from Superscript III One Step Kit, ThermoFisher, Cat No 12574026). The amplification was carried out on a Bio-Rad T100 Thermal Cycler under the following conditions: 50 °C for 15 minutes, 95 °C for 2 minutes, followed by 20 cycles of 95 °C for 15 seconds and 60 °C for 4 minutes. The resulting cDNA was diluted with 4x vol TE buffer and stored at −20 °C for up to 3 months. Then the diluted cDNA and Taqman Assays were loaded onto an IFC chip for gene expression analysis as per protocols provided on the Fluidigm website. Data analysis was finished using the built-in software, adhering to the “Real-Time PCR Analysis” guidelines from Fluidigm. Colonies with undetected reference genes were excluded from further analysis. For gene expression analysis of ATRX, HBA and HBM in individual BFU-E colonies, expression values were normalised to the mean of reference genes (RPL13A, RPS18, ACTB and GAPDH), and a uniform cutoff of 2 was applied to relative expression values during plotting to exclude outlier values.
Cellular morphological analysis - Wright’s stain of cytospin
1 × 105 cells were resuspended in 200 μL of PBS, spun (400 rpm for 5 mins) onto a glass slide in a Cytospin 4 Centrifuge (ThermoFisher). The slides were air-dried, stained with modified Wright’s Stain using Hemateck slide strainer and mounted in DPX (Sigma, Cat No 06522). Cytospins were imaged using an Olympus BX 60 microscope.
Flow cytometry
Cells were washed and resuspended in 100 μL of FACS buffer (1x PBS supplemented with 2% FBS and 2 mM EDTA) prior to staining with a panel of fluorophore-conjugated antibodies at 4 °C for 30 min. Following antibody staining, the cells were washed and resuspended in FACS buffer containing Hoechst-33258 for live/dead cell analysis. FACS analysis was done on an Attune NxT analyser (ThermoFisher). Gating was set using fluorescence-minus-one (FMO) and unstained controls. Analysis was performed using FlowJo software. The details of the FACS panels and flow cytometry antibodies are provided in Supplementary Table 2.
Iso-electric focusing for haemoglobin analysis
Approximately 5 × 105 cultured erythroblasts were lysed in haemoglobin elution solution, and the haemoglobin was analysed using an iso-electric focusing kit (RESOLVE Haemoglobin kit, PerkinElmer, USA) on a water-cooled horizontal electrophoresis rig (GE Healthcare) according to the manufacturer’s instructions. Following electrophoresis, the gels were fixed in 10% trichloroacetic acid and stained using the JB-2 staining system (Perkin Elmer, USA) as per the manufacturer’s guidelines.
HUDEP-2 cell culture and differentiation
Wildtype HUDEP-2 cells were obtained from the Cell Engineering Division, RIKEN BioResource Center, Tsukuba, Japan64. During the expansion phase, cells were cultured in StemSpan SFEM medium (STEMCELL Technologies, Cat No 09650) supplemented with 50 ng/mL SCF, 3 U/mL EPO, 0.4 μg/mL dexamethasone (OUH pharmacy), 2mM L-Glutamine (Gibco, Cat No 25030081), 1x P/S, and 2 μg/mL doxycycline (Sigma-Aldrich, Cat No D9891). Cultures were maintained at 37 °C 5% CO2 incubator, with a cell density of 0.5-1 ×106 cells/mL. The media was replaced every two days to ensure an effective doxycycline concentration.
Differentiation of HUDEP-2 cells used a two-phase liquid culture system, requiring a base media consisted of IMDM supplemented with 3% Human AB plasma, 10 μg/mL human insulin, 3 U/mL heparin, 3 U/mL EPO, 200 μg/mL holo-transferrin, 2% (v/v) FBS and 1x P/S. From day 0 to day 6, 1 ng/mL IL3, 50 ng/mL SCF, and 2 μg/mL doxycycline were added to the base media for cell differentiation. After 6 days, media was changed to the second phase, where only 300 μg/mL holo-transferrin was supplemented to the base media. Media was changed every two days. Cell density was maintained at 1 × 106 cells/mL prior to day 6 and at 1.5 × 106 cells/mL thereafter.
Generation of ATRX degron cell line
A guide RNA targeting the ATRX C-terminal domain (CTD) (gRNA sequence in Supplementary Table 1) were designed and cloned into the pX458-Ruby plasmid (Addgene 110164). A DNA donor template for homologous-directed recombination (HDR) was constructed to contain EGFP-FKBP12 F36V, flanked by 800 bp homologous arms on each side, directly linking to the absolute CTD of ATRX. The donor DNA was synthesised and inserted into pMK-RQ plasmid. To prevent cleavage of the donor DNA, all sgRNAs target sites within the donor plasmid were point-mutated without altering the encoded amino acids. For nucleofection, 2.5 μg of the pX458-Ruby-gRNA plasmid and 2.5 μg of the donor plasmid were co-transfected into 2 × 106 HUDEP-2 WT cells using the 2B Amaxa Human CD34 Cell Nucleofector Kit (Lonza, Cat No VPA-1003) on the AMAXA Nucleofector 2B system following the manufacturer’s instructions. The transfected cells were cultured in HUDEP-2 SFEM expansion media supplemented with 0.75 μM RAD51-stimulatory compound-1 (RS-1) to enhance HDR efficiency. Two days post-transfection, Ruby+ cells were bulk sorted to recovering for 1 week and subsequently single-cell sorted into Terasaki plates (Nunc, Cat No 163118) for clonal screening. Successful grown clones were genotyped using the primers listed in Supplementary Table 1.
Generation of ATRX degron-ψζ VNTR KO cell line
The ψζ VNTR region was knocked out in the ATRX degron cell line using a dual-gRNA CRISPR-Cas9-mediated knockout strategy. Due to the high homology between HBZ and its pseudogene HBZP1 where ψζ VNTR resides, the 5’ sgRNA was designed to target HBZP1 promoter where the sequences have no homology with HBZ, while the 3’ sgRNA targeted a downstream region of HBZP1. This design facilitated the deletion of the entire HBZP1 (~2.5 kb). The dual gRNAs were cloned into the pX458-eGFP (Addgene 48138) and pX458-Ruby (Addgene 110164) plasmids respectively, and 2.5 μg of each was co-transfected into 2 × 106 ATRX degron cells using the same nucleofection protocol as described above. Cells were cultured in SFEM expansion media after transfection, and GFP+ Ruby+ double-positive cells were sorted prior to single clonal screening via a PCR-based strategy. The gRNA and primer sequences used are provided in Supplementary Table 1.
RNA extraction and qPCR
0.5-2 × 106 cells were collected in 350 μL RLT buffer (from QIAGEN RNeasy Mini Kit), snapped frozen on dry ice and stored at −80 °C until use. Total RNA was purified using RNeasy mini kit (QIAGEN, Cat No 74104), and 1 μg of total RNA was used to generate complementary DNA (cDNA) using SuperScript III First Strand Synthesis SuperMix (ThermoFisher, Cat No 11752-250). All quantitative reverse transcription PCR (qRT-PCR) were carried out using inventoried TaqMan assays (Applied Biosystems; TaqMan IDs: HBA2/HBA1-Hs00361191_g1, HBB-Hs00747223_g1, HBG-Hs00361131_g1, HBM-Hs01392876_g1, NME4-Hs00359037_m1, RPL13A-Hs03043885_g1, RPS18-Hs01375212_g1, ACTB-Hs01060665_g1, GAPDH-Hs02758991_g1) and Taqman Universal PCR Master Mix II (ThermoFisher, Cat No 4440040) in technical triplicate on StepOne Plus Real-Time PCR System (Applied Biosystems). Data were first analysed using the delta-delta CT (ΔΔCT) method and then normalised to the control sample. For PCR amplification of pseudo-zeta gene HBZP1, cDNA from HUDEP-2 WT cells was used as template and primers targeting zeta gene HBZ and pseudo-zeta gene HBZP1 (listed in Supplementary Table 1) were used to distinguish the specific amplification of HBZP1. The PCR reaction was carried out using PWO polymerase (Roche, Cat No 11644947001) and Platinum Taq (Invitrogen, Cat No 10966018), with the addition of Applied Biotechnologies Buffer IV (1x), 20 mM MgCl2, and a high G/C enhancer (1x, New England Biolabs, Cat No B9028) to overcome the high GC content and repetitive nature of the template. The thermal cycling conditions were as follows: 94 °C for 2 min, 35 cycles of 94 °C for 1 min, 60 °C for 30 s, 72 °C for 1 min 30 s and 72 °C for 10 min.
Immunofluorescence staining
Cells were seeded on Cell-Tak (Corning, Cat No 354240)-coated CellView slides (Greiner Bio-One, Cat. No. 543978), and then fixed with 4% paraformaldehyde for 10 minutes, permeabilised with 0.1% Triton X-100 for 5 minutes, and blocked with 1% bovine serum albumin + 2% goat serum for 1 hour at room temperature. Cells were next incubated overnight at 4 °C with an anti-γH2AX antibody (Merck, Cat. No. 05-636, use at 1:400) or anti-ubiquityl-histone H2A antibody (Cell Signaling Technology, Cat. No. 8240, use at 1:800). The following day, cells were incubated with Alexa Fluor 647-conjugated secondary antibodies for 1 hour at room temperature, followed by staining with DAPI. Fluorescence images were acquired using a Leica DMi8 Thunder microscope and analysed in Fiji software.
Protein extraction and western blot
At least 1 × 106 cells were lysed in 50-200 μL of RIPA buffer (Sigma, Cat No R0278) supplemented with digitonin (ThermoFisher, Cat No BN2006), Benzonase (Merck, Cat No 71205-3), and protease inhibitor cocktail (Roche, Cat No 11836153001), based on the cell number. Cells lysis was conducted for 40-60 minutes at 4 °C on an end-to-end rotator. The lysate was centrifuged at 16,000 g for 10 min at 4 °C. Supernatant was collected and mixed with an equal volume of 2x Laemmli Sample Buffer (Bio-Rad, Cat No 1610737) containing 1/4 volume of 1 M DTT (ThermoFisher, Cat No R0861).
Protein samples were denatured at 95 °C for 5 minutes, chilled on ice, and loaded onto a NuPAGE Bis-Tris gel (ThermoFisher Scientific) using a Mini-PROTEAN system (Bio-Rad). Proteins were then transferred to a PVDF blotting membrane (Merck, Cat No 03010040001) overnight (14-18 hours) with constant 25 mA current, followed by blocking with 5% skimmed milk (Merck, Cat No 70166) dissolved in TBS-T (Tris buffered saline + 0.1% Tween-20). The membrane was incubated with primary antibodies for 1 hour at room temperature in 2.5% skimmed milk, washed three times with TBS-T, and then incubated with secondary antibodies for 1 hour at room temperature. After three additional washes, the membrane was exposed on X-ray film. Antibodies used in this study include: anti-ATRX (homemade mouse antibody, clone 39 f; and Abcam, Cat No ab97508, use at 1:1000), anti-alpha tubulin (Abcam, mouse monoclonal, ab7291, use at 1:20,000), and HRP-conjugated secondary antibodies Goat anti-mouse IgG (ThermoFisher, Cat No 31430).
Smart-seq2 single cell RNA sequencing
CD34 differentiated day 13 erythroid cells were stained with the viability dye Hoechst 33342 (ThermoFisher, Cat No 62249) before cell sorting. Single live nucleated cells were sorted into each well of a cold 384-well PCR plate (FrameStar 4titude, Cat No 4ti-0384/C) containing 2 µL of lysis buffer with barcoded Oligo(dT) (0.2% Triton X-100 (Sigma), 2 U/μl of RNase inhibitor (TaKara, Cat No 2313 A), 2.5 mM dNTP, 0.025 AU/mL protease, and ERCC RNA spike-in)65 using a Sony SH800S Cell Sorter. Next, retrotranscription and PCR steps were performed65. Briefly, proteinase in the lysis buffer was heat-inactivated at 72 °C for 15 min, during which retrotranscription mix was prepared by mixing SMARTScribe retrotranscriptase (Clontech, Cat No 639537), RNase inhibitor, and template-switching oligo TSO-LNA (sequence: AAGCAGTGGTATCAACGCAGAGTACATrGrG+G, RNase-free HPLC purified). The RT mix was dispensed into each well after heat inactivation and first-strand cDNA was synthesised. Afterwards, the cDNA was amplified by SeqAMP (Clontech, Cat No 638509) using 21 PCR cycles. Amplified products from each plate were pooled, bead-purified, tagmented, and indexed using Tn5 enzyme (Illumina Nextera XT Kit, Cat No FC-131-1096) to construct libraries. The libraries were purified with AMPure beads, assessed on TapeStation System (Agilent), quantified using the NEBNext Library Quant Kit (NEB, Cat No E7630L), and sequenced on a NextSeq platform with 75 bp single-end reads using the NextSeq 500/550 High-output 75 Cycle Kit (Illumina, Cat No 20024906).
scRNA-seq data analysis
Demultiplexed FASTQ files were trimmed for polyA tails using TrimGalore; files from different lanes were merged using samtools and aligned to hg19 genome using STAR. Gene counts for each cell were obtained with FeatureCounts and normalised by dividing total library size of the cell and multiplying by the mean library size across all processed cells. Genes with normalised counts <1 were considered non-detected and their expression values were set to 0. The normalised counts were further applied with log2 normalisation. QC filtering was performed to retain cells with >3000 reads; <10% mitochondrial reads, and >70% mapped reads per cell. Highly variable genes were identified by fitting a LOESS model of log2(mean expression) versus coefficient of variation for each gene. Genes with a coefficient of variation above the fitted model and log2(mean expression) ≥ 0 were selected as HVGs. Dimensionality reduction was performed by PCA (20 dimensions) and t-SNE (‘Rtsne’ package, with “perplexity = 10, max_iter = 500”) using HVGs, and plotted using “ggplot2” package in R. Differentially expressed genes were identified using a non-parametric Wilcoxon test and expressing cell frequency was computed using Fisher′s exact tests on log2-normalised counts for genes expressed in ≥5 cells. Significant genes were defined by an adjusted p value (false discovery rate (FDR) < 0.05) and an absolute log2 fold change ( |log2FC| > 0.5). P values were combined using Fisher’s method, and adjusted p-values were calculated using the Benjamini & Hochberg FDR procedure. Violin plots from selected genes were generated using “ggplot2” package in R. GO analysis was conducted using ClusterProfiler R package. Differentially expressed gene set was compared with the gene list from org.Hs.eg.db from the Bioconductor package for GO analysis with settings ont = “BP”, pAdjustMethod = “fdr”. For cell cycle analysis, a Seurat object was created using the original count table. Cell cycle scores were computed and used to assign cell cycle stages. Fisher’s test was used to examine cell cycle difference between groups.
Chromatin Immunoprecipitation sequencing (ChIP-seq) and data analysis
ATRX ChIP was performed on 5 × 107 cells using the ChIP Assay Kit (Millipore, Cat No 17-295) following an optimised protocol20,45. Briefly, cells were double-fixed with firstly 2 mM ethylene glycol bis succinimidyl succinate (EGS, Sigma, Cat No E3257) for 45 min and then 1% formaldehyde solution for 20 min at room temperature. The cross-linking was quenched with 125 mM glycine (Sigma-Aldrich, Cat No G7126). After cells lysis, the chromatin was fragmented using Covaris ME220 ultrasonicator (Peak Incident Power (PIP): 75, Duty factor (DF): 14, Cycles/burst (cpb): 1000, Duration: 900 s, Temp: 6 °C), followed by pre-clearing with Protein A/G Dynabeads (Invitrogen, Cat No 10001D, 10003D) before immunoprecipitation with 15 μg ATRX antibody (Abcam, Cat No ab97508). After sequential washes at 4 °C, samples were eluted, de-crosslinked and proteinase K treated. DNA was precipitated using phenol/chloroform extraction and ethanol precipitation with GlycoBlue (15 mg/mL) (Thermo Scientific, Cat No AM9516) prior to dissolving in 20 μL of ultrapure water. DNA was quantified using the Qubit dsDNA HS assay (Invitrogen, Cat No Q32854) and indexed using NEBNext Ultra II DNA library prep kit (NEB, Cat No E7645L). Libraries were quantified, and sequenced on an Illumina NextSeq machine with paired-end reads.
ChIP-seq data quality was assessed by FASTQC. Data was analysed using a customised in-house pipeline (https://github.com/Hughes-Genome-Group/NGseqBasic/releases). Paired-end reads were aligned to the hg19 genome using bowtie 2. Output BAM files were sorted and indexed using Samtools, normalised to RPKM using Deeptools (--normalizeUsingRPKM), and converted to BigWig files for visualisation in the USCS genome browser. BigWig files were used for peak calling on LanceOtron (https://lanceotron.molbiol.ox.ac.uk/).
CRISPR-Cas9-induced DNA damage assay
2 × 106 of ATRX degron-VNTR KO differentiated day 6 cells were transfected with RNPs using the 2B Amaxa Human CD34 Cell Nucleofector Kit on the AMAXA Nucleofector 2B system. Following electroporation, cells were cultured in HUDEP-2 day 6 differentiation media for 2 days before being harvested for qPCR analysis. Three different sgRNAs were tested individually, and their sequences are provided in Supplementary Table 1.
DRIP-qPCR
The DRIP protocol was performed as previously described66. 1 × 108 cells were harvested followed by nuclear lysis. Nuclear extracts were incubated with 30 mg proteinase K at 37 °C overnight, followed by DNA isolation. To minimise nonspecific RNA signals detected by the S9.6 antibody, nuclei were pre-treated with RNase I at 2 U per 100 µg for 15 minutes at 37 °C and sonicated to 200-300 bp. Half the fragments were treated with RNase H (Roche) at 10 U per 100 µg overnight at 37 °C. Treated and untreated samples underwent S9.6 antibody immunoprecipitation and enrichment using sheep anti-mouse IgG Dynabeads (ThermoFisher Scientific, Cat No11031). Hybrids were extracted with phenol/chloroform, precipitated with glycogen, and resuspended in nuclease-free water. qPCR analysis was performed on the samples to detect enrichment around ψζ VNTR and a distal region on chromosome 16. The qPCR run was done using SYBR Green master mix with primers listed in Supplementary Table 1.
Dot blot analysis of R-loops
Briefly, nuclear DNA were extracted from ATRX degron cells (treated with dTAG-13-neg or dTAG-13) by potassium acetate and ethanol precipitated. Precipitated DNA was gently spooled and washed with 70% ethanol without centrifugation. DNA was allowed to air dry before resuspended in water followed by brief sonication to reduce DNA viscosity. DNA concentration was measured and adjusted to 400 ng/μl, followed by serial dilutions. For RNase H controls, an aliquot of DNA was incubated with 5 U RNase H (M0297S New England Biolabs) in 0.5 M NaCl (EN0531 Thermo Scientific) at 37 °C for 20 min respectively, then serially diluted. Two microliters of each dilution were spotted on a Hybond-N+ nylon transfer membrane (RPN203B Amersham), air-dried and UV-crosslinked. The membrane was then blocked with 5% milk overnight, followed by a 2 h incubation with anti-R-loop antibody S9.6 at 4 °C before washing, and then incubated with HRP-conjugated secondary antibody. Signals were detected using enhanced chemiluminescence (ECL) kit (P90720 Millipore).
RNase H overexpression assay
Three million of differentiation day 6 ATRX degron cells (with dTAG-13-neg or dTAG-13 treatment) were electroporated with 3 μg of either a plasmid expressing GFP-tagged RNase H1 or a GFP alone control plasmid using Lonza Human CD34+ Cell Nucleofector Kit (Cat No VPA-1003) on the AMAXA Nucleofector 2B system. GFP+ cells were sorted after 12 h for RNA extraction using RNeasy micro kit (QIAGEN, Cat No 74004), followed by qRT-PCR analysis.
Statistical analysis
Statistical analyses between two groups were conducted using a two-tailed unpaired Welch’s t-test for normally distributed data, or the Mann-Whitney U-test for non-normally distributed data. Comparisons among three or more groups were performed using one-way ANOVA followed by Dunnett test. For datasets involving multiple conditions across groups, two-way ANOVA followed by Šídák’s test was applied. For all the fold-change data, relative fold-change values were log₂-transformed before statistical analysis. Statistical significance was defined as p < 0.05, with significance levels represented as *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001. The number of independent experiments was indicated in each figure. Detailed descriptions of the statistical tests used for each dataset, along with exact p-values, are provided in the corresponding figure legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNA-seq and ChIP-seq data generated in this study have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (NCBI GEO) under accession codes GSE291201 and GSE291197 respectively. Source data are provided with this paper.
Code availability
The in-house pipeline used for processing and aligning ChIP-seq data is available at GitHub (https://github.com/Hughes-Genome-Group/NGseqBasic/releases).
References
Gibbons, R. J., Picketts, D. J., Villard, L. & Higgs, D. R. Mutations in a putative global transcriptional regulator cause X-linked mental retardation with α-thalassemia (ATR-X syndrome). Cell 80, 837–845 (1995).
Gibbons, R. Alpha thalassaemia-mental retardation, X linked. Orphanet J Rare Dis 1, 15 (2006).
Jiao, Y. et al. DAXX/ATRX, MEN1, and mTOR pathway genes are frequently altered in pancreatic neuroendocrine tumors. Science (1979) 331, 1199–1203 (2011).
Schwartzentruber, J. et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 2012 482:7384 482, 226–231 (2012).
Chen, X. et al. Recurrent somatic structural variations contribute to tumorigenesis in pediatric osteosarcoma. Cell Rep 7, 104–112 (2014).
Xue, Y. et al. The ATRX syndrome protein forms a chromatin-remodeling complex with Daxx and localizes in promyelocytic leukemia nuclear bodies. Proc Natl Acad Sci USA 100, 10635–10640 (2003).
Lewis, P. W., Elsaesser, S. J., Noh, K. M., Stadler, S. C. & Allis, C. D. Daxx is an H3.3-specific histone chaperone and cooperates with ATRX in replication-independent chromatin assembly at telomeres. Proc Natl Acad Sci USA 107, 14075–14080 (2010).
Pladevall-Morera, D. et al. Proteomic characterization of chromosomal common fragile site (CFS)-associated proteins uncovers ATRX as a regulator of CFS stability. Nucleic Acids Res 47, 8004–8018 (2019).
Wong, L. H. et al. ATRX interacts with H3.3 in maintaining telomere structural integrity in pluripotent embryonic stem cells. Genome Res 20, 351–360 (2010).
Clynes, D. et al. ATRX dysfunction induces replication defects in primary mouse cells. PLoS One 9, (2014).
Conte, D. et al. Loss of Atrx Sensitizes Cells to DNA Damaging Agents through p53-Mediated Death Pathways. PLoS One 7, e52167 (2012).
Raghunandan, M. et al. Functional cross talk between the Fanconi anemia and ATRX/DAXX histone chaperone pathways promotes replication fork recovery. Hum Mol Genet 29, 1083–1095 (2020).
Juhász, S., Elbakry, A., Mathes, A. & Löbrich, M. ATRX Promotes DNA Repair Synthesis and Sister Chromatid Exchange during Homologous Recombination. Mol Cell 71, 11–24.e7 (2018).
Leung, J. W.-C. et al. Alpha thalassemia/mental retardation syndrome X-linked gene product ATRX is required for proper replication restart and cellular resistance to replication stress. J Biol Chem 288, 6342–6350 (2013).
Huh, M. S. et al. Stalled replication forks within heterochromatin require ATRX for protection. Cell Death & Disease 2016 7:5 7, e2220–e2220 (2016).
Elbakry, A., Juhász, S., Chan, K. C. & Löbrich, M. ATRX and RECQ5 define distinct homologous recombination subpathways. Proc Natl Acad Sci USA 118, (2021).
Garbarino, J., Eckroate, J., Sundaram, R. K., Jensen, R. B. & Bindra, R. S. Loss of ATRX confers DNA repair defects and PARP inhibitor sensitivity. Transl Oncol 14, (2021).
McDowell, T. L. et al. Localization of a putative transcriptional regulator (ATRX) at pericentromeric heterochromatin and the short arms of acrocentric chromosomes. Proc Natl Acad Sci USA 96, 13983–13988 (1999).
Law, M. J. et al. ATR-X syndrome protein targets tandem repeats and influences allele-specific expression in a size-dependent manner. Cell 143, 367–378 (2010).
Truch, J. et al. The chromatin remodeller ATRX facilitates diverse nuclear processes, in a stochastic manner, in both heterochromatin and euchromatin. Nature Communications 2022 13:1 13, 1–16 (2022).
Danussi, C. et al. Atrx inactivation drives disease-defining phenotypes in glioma cells of origin through global epigenomic remodeling. Nature Communications 2018 9:1 9, 1–15 (2018).
Qin, T. et al. ATRX loss in glioma results in dysregulation of cell-cycle phase transition and ATM inhibitor radio-sensitization. Cell Rep 38, (2022).
Ekwattanakit, S. et al. Analysis of DNA Methylation at the Human Alpha Globin Cluster during Hematopoiesis. Blood 112, 1861 (2008).
Bird, A. P., Taggart, M. H., Nicholls, R. D. & Higgs, D. R. Non-methylated CpG-rich islands at the human alpha-globin locus: implications for evolution of the alpha-globin pseudogene. EMBO J 6, 999 (1987).
Scott, C. et al. Recapitulation of erythropoiesis in congenital dyserythropoietic anemia type I (CDA-I) identifies defects in differentiation and nucleolar abnormalities. Haematologica 106, 2960–2970 (2020).
Kelliher, J., Ghosal, G. & Leung, J. W. C. New answers to the old RIDDLE: RNF168 and the DNA damage response pathway. FEBS J 289, 2467–2480 (2022).
Barbour, H., Daou, S., Hendzel, M. & Affar, E. B. Polycomb group-mediated histone H2A monoubiquitination in epigenome regulation and nuclear processes. Nature Communications 11, 5947 (2020).
Sobhian, B. et al. RAP80 Targets BRCA1 to Specific Ubiquitin Structures at DNA Damage Sites. Science (1979) 316, 1198–1202 (2007).
Li, Y. et al. Effect of ATRX and G-Quadruplex Formation by the VNTR Sequence on α-Globin Gene Expression. Chembiochem 17, 928–935 (2016).
Tillotson, R. et al. A new mouse model of ATR-X syndrome carrying a common patient mutation exhibits neurological and morphological defects. Hum Mol Genet 32, 2485–2501 (2023).
Nguyen, D. T. et al. The chromatin remodelling factor ATRX suppresses R-loops in transcribed telomeric repeats. EMBO Rep 18, 914–928 (2017).
Bedrat, A., Lacroix, L. & Mergny, J. L. Re-evaluation of G-quadruplex propensity with G4Hunter. Nucleic Acids Res 44, 1746–1759 (2016).
Brázda, V. et al. G4Hunter web application: a web server for G-quadruplex prediction. Bioinformatics 35, 3493–3495 (2019).
Yan, Q. et al. Proximity labeling identifies a repertoire of site-specific R-loop modulators. Nature Communications 2022 13:1 13, 1–17 (2022).
Teng, Y. C. et al. ATRX promotes heterochromatin formation to protect cells from G-quadruplex DNA-mediated stress. Nature Communications 2021 12:1 12, 1–14 (2021).
Crossley, M. P., Bocek, M. & Cimprich, K. A. R-Loops as Cellular Regulators and Genomic Threats. Mol Cell 73, 398–411 (2019).
Jalan, M. et al. RAD52 resolves transcription-replication conflicts to mitigate R-loop induced genome instability. Nat Commun 15, (2024).
Iannelli, F. et al. A damaged genome’s transcriptional landscape through multilayered expression profiling around in situ-mapped DNA double-strand breaks. Nature Communications 2017 8:1 8, 1–12 (2017).
Shanbhag, N. M., Rafalska-Metcalf, I. U., Balane-Bolivar, C., Janicki, S. M. & Greenberg, R. A. ATM-Dependent chromatin changes silence transcription in cis to dna double-strand breaks. Cell 141, 970–981 (2010).
Min, S., Ji, J. H., Heo, Y. & Cho, H. Transcriptional regulation and chromatin dynamics at DNA double-strand breaks. Experimental & Molecular Medicine 2022 54:10 54, 1705–1712 (2022).
Akter, J. et al. Loss of p53 suppresses replication stress-induced DNA damage in ATRX-deficient neuroblastoma. Oncogenesis 10, (2021).
Wang, Y. et al. G-quadruplex DNA drives genomic instability and represents a targetable molecular abnormality in ATRX-deficient malignant glioma. Nat Commun 10, 1–14 (2019).
Levy, M. A., Kernohan, K. D., Jiang, Y. & Bérubé, N. G. ATRX promotes gene expression by facilitating transcriptional elongation through guanine-rich coding regions. Hum Mol Genet 24, 1824–1835 (2015).
Proudfoot, N. J., Gil Sir William, A. & Maniatis, T. The Structure of the Human Zeta-Globin Gene and a Closely Linked, Nearly Identical Pseudogene. Cell vol. 31 (1982).
Truch, J., Telenius, J., Higgs, D. R. & Gibbons, R. J. How to Tackle Challenging ChIP-Seq, with Long-Range Cross-Linking, Using ATRX as an Example. Methods Mol Biol 1832, 105–130 (2018).
Lee, W. T. C. et al. Single-molecule imaging reveals replication fork coupled formation of G-quadruplex structures hinders local replication stress signaling. Nature Communications 2021 12:1 12, 1–14 (2021).
Duquette, M. L., Handa, P., Vincent, J. A., Taylor, A. F. & Maizels, N. Intracellular transcription of G-rich DNAs induces formation of G-loops, novel structures containing G4 DNA. Genes Dev 18, 1618–1629 (2004).
Girasol, M. J. et al. RAD51-mediated R-loop formation acts to repair transcription-associated DNA breaks driving antigenic variation in Trypanosoma brucei. Proceedings of the National Academy of Sciences 120, e2309306120 (2023).
Ngo, G. H. P., Grimstead, J. W. & Baird, D. M. UPF1 promotes the formation of R loops to stimulate DNA double-strand break repair. Nature Communications 12, 3849 (2021).
Watson, L. A. et al. Atrx deficiency induces telomere dysfunction, endocrine defects, and reduced life span. J Clin Invest 123, 2049–2063 (2013).
Huh, M. S. et al. Compromised genomic integrity impedes muscle growth after Atrx inactivation. J Clin Invest 122, 4412 (2012).
Maréchal, A. & Zou, L. DNA Damage Sensing by the ATM and ATR Kinases. Cold Spring Harb Perspect Biol 5, (2013).
Masud, T. et al. Ubiquitin-mediated DNA damage response is synthetic lethal with G-quadruplex stabilizer CX-5461. Scientific Reports 2021 11:1 11, 1–20 (2021).
Rong, G., Pan, Z., Ding, M. & Wang, L. RNF168 suppresses the cancer stem cell-like traits of nonsmall cell lung cancer cells by mediating RhoC ubiquitination. Environ Toxicol 37, 603–611 (2022).
Xu, C., Xu, Y., Gursoy-Yuzugullu, O. & Price, B. D. The histone variant macroH2A1.1 is recruited to DSBs through a mechanism involving PARP1. FEBS Lett 586, 3920–3925 (2012).
Khurana, S. et al. A macrohistone variant links dynamic chromatin compaction to BRCA1-dependent genome maintenance. Cell Rep 8, 1049–1062 (2014).
Kim, J. et al. Replication Stress Shapes a Protective Chromatin Environment across Fragile Genomic Regions. Mol Cell 69, 36–47.e7 (2018).
Kim, J. et al. The macroH2A1.2 histone variant links ATRX loss to alternative telomere lengthening. Nat Struct Mol Biol 26, 213–219 (2019).
Ratnakumar, K. et al. ATRX-mediated chromatin association of histone variant macroH2A1 regulates α-globin expression. Genes Dev 26, 433–438 (2012).
Lukas, C. et al. 53BP1 nuclear bodies form around DNA lesions generated by mitotic transmission of chromosomes under replication stress. Nat Cell Biol 13, 243–253 (2011).
Oberdoerffer, P. & Miller, K. M. Histone H2A variants: Diversifying chromatin to ensure genome integrity. Semin Cell Dev Biol 135, 59–72 (2023).
Mendoza, O., Bourdoncle, A., Boulé, J. B., Brosh, R. M. & Mergny, J. L. G-quadruplexes and helicases. Nucleic Acids Res 44, 1989 (2016).
Yu, Z. et al. DDX5 resolves R-loops at DNA double-strand breaks to promote DNA repair and avoid chromosomal deletions. NAR Cancer 2, (2020).
Kurita, R. et al. Establishment of Immortalized Human Erythroid Progenitor Cell Lines Able to Produce Enucleated Red Blood Cells. PLoS One 8, (2013).
Rodriguez-Meira, A. et al. Unravelling Intratumoral Heterogeneity through High-Sensitivity Single-Cell Mutational Analysis and Parallel RNA Sequencing. Mol Cell 73, 1292–1305.e8 (2019).
Tan-Wong, S. M., Dhir, S. & Proudfoot, N. J. R-Loops Promote Antisense Transcription across the Mammalian Genome. Mol Cell 76, 600–616.e6 (2019).
Acknowledgements
Y.S., R.J.G. and D.R.H. were supported by the Chinese Academy of Medical Sciences Oxford Institute and CAMS Innovation Fund for Medical Science (CIFMS), China (grant number: 2018-I2M-2-002). R.J.G. received funding from Medical Research Council (UK) (grant number MC_UU_12025 / unit programme MC_UU_12009/3). We are very grateful to the WIMM Genome Engineering Facility (Dr. Philip Hublitz) for helping with the construction of the CRISPR plasmids used in this study. We would like to thank WIMM FACS facility and Single Cell facility for their support and guidance. We extend our gratitude to Ms. Gem Flint, Dr. Marco Di Antonio, Prof. Jim Hughes, Mr. Simone Riva, Dr. Ravza Gür, Dr. William Nathan, Dr. Andre Nussenzweig, and Dr. Antonio Rodríguez-Romera for their help and discussion.
Author information
Authors and Affiliations
Contributions
Y.S., R.J.G. and D.R.H. conceptualised the study and conceived the experimental design. Y.S. performed most of the experiments and formally analysed data. K.G. carried out cell culture, CRISPR-induced DNA damage assay, and pseudo-zeta cDNA PCR. S.M.T.-W. performed DRIP-qPCR and R-loops dot blot assay with supervision from N.J.P. Y.S. and S.W. performed the scRNA-seq analysis. C.A.F. designed pseudo-zeta cDNA PCR primers and performed the IEF assay. L.T. contributed to the 3D plotting of Biomark gene expression data. Y.S. and D.R.H. wrote the original manuscript; all authors read and approved the manuscript. R.J.G. and D.R.H. provided supervision and secured funding.
Corresponding authors
Ethics declarations
Competing interests
Y.S. provides consultancy for Alethiomics Ltd, which had no involvement in the study’s design, data collection, analysis, or manuscript preparation. All remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Suming Huang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shen, Y., Gupta, K., Tan-Wong, S.M. et al. ATRX loss couples genome instability at a G-rich repeat to dysregulation of human alpha-globin expression. Nat Commun 17, 2749 (2026). https://doi.org/10.1038/s41467-026-69169-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69169-7