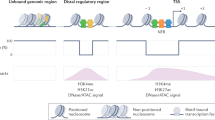
Abstract
Transcription factors (TFs) search for and bind specific DNA target sites to control gene expression. However, eukaryotic TF target search within the chromatinized genome is poorly characterized. Here, we combine in vivo and in vitro single-molecule imaging to dissect the role of differentially-charged DNA-binding domain flanking regions in the target search of Sox TFs. We demonstrate that the flanking region of Sox2 markedly enhances search efficiency compared to the negatively-charged flanking region of Sox17. The involved mechanisms are distinct for naked DNA compared to chromatin. On DNA, the enhanced search of Sox2 is driven by an increased target recognition rate during 1D sliding, despite reduced sliding speed. Conversely, enhanced nonspecific interactions between the Sox2’s DNA-binding domain flanking region and nucleosomes facilitate binding to compact chromatin and reinforce pioneer activity. These findings provide critical insights into biophysical mechanisms governing TF target search within the chromatinized genome.
Similar content being viewed by others
Introduction
Transcription factors (TFs) are central regulators of gene expression in all living organisms. They identify and bind target DNA sites in a sequence-specific manner via their DNA-binding domains (DBDs) (specific interactions), navigating a vast landscape of nonspecific DNA (nonspecific interactions). An efficient target search is crucial to allow a limited number of TFs (103–106 molecules per cell1,2) to locate and discriminate tens of thousands of target sites identified by chromatin immunoprecipitation sequencing (ChIP-seq)3,4,5,6 from millions of nonspecific decoy sites within the complex nuclear environment7,8,9,10,11.
The mechanisms by which TFs maximize their target search efficiency within chromatin remain poorly understood. The facilitated diffusion model, formulated for free DNA, states that TFs enhance search efficiency by engaging in nonspecific DNA interactions, alternating between 1D diffusion along DNA and 3D diffusion within the nucleoplasm12,13,14,15. This dimensionality-reduced diffusion is believed to substantially increase search efficiency compared to free diffusion alone, although the magnitude of this effect is still debated16. Pioneering experiments showed directly that the bacterial TF LacI follows the facilitated diffusion model in live E. coli17 and in vitro18,19,20. However, in eukaryotes chromatin acts as a barrier to TF-DNA interactions, potentially impairing TF target search and site recognition21,22,23,24.
Eukaryotic TFs contain intrinsically disordered regions (IDRs) outside of their DBDs. How these regions influence target search remains unclear. Recent findings indicate that IDRs play an important role in defining the binding specificity of TFs in vivo25. TFs with identical DBDs but distinct IDRs recognize the same motifs in vitro, but often bind distinct genomic subsets in vivo, highlighting the role of DBD-proximal IDRs in modulating target selection. While IDRs can mediate protein-protein interactions26,27,28,29, they can also play a key role in determining chromatin binding and diffusion behavior, even in the absence of interaction partners30,31,32,33,34.
A subset of eukaryotic TFs, known as pioneer TFs (pTFs), can bind nucleosomal DNA and initiate local chromatin opening, facilitating access to previously inaccessible regulatory elements35,36,37,38. Sox TFs, which include several pTFs24,39, bind their targets through their high mobility group box DBD, either alone or as heterodimers with other TFs. We previously observed that individual Sox TFs show markedly different degrees of association with mitotic chromosomes, a property that correlates with nonspecific DNA binding. We also showed that a higher content of positively charged amino acids promotes TF association to mitotic chromosomes40. Sox TFs exhibit a broad charge distribution in regions outside the DBD, going from −9z to +9z, while their net charge within the DBDs is comparable (Fig. 1a). The overall charge in these DBD-flanking IDRs is positively correlated with their ability to bind mitotic chromosomes (Fig. 1b).
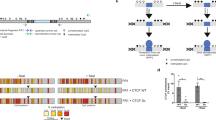
a Net charge in DBD (red) and non-DBD (black) of Sox TFs. b Left, Pearson correlation between the mitotic bound fraction (MBF)40 and the net charge in non-DBD of Sox TFs in a; Right, representative images40 of H2B-mCherry, Sox2-YPet (top) and Sox17-YPet (bottom) localization in metaphase. Scale bar: 10 μm. c Charge distributions for Sox2 (top) and Sox17 (bottom), calculated as sliding-window averages of residue charges over five-amino-acid segments (see the “Methods” section). d Scheme representation of the Sox TF library. e Top, representative images of in vivo SMT. Nuclei (left, red) and sparsely labeled Sox TFs (right, yellow) are visualized in the SiR647 and JF549 emission channels, respectively. Scale bar: 15 μm; Bottom, scheme of in vivo single-molecule imaging using HILO microscopy (left) and a labeling strategy for Sox TFs using the Halo-JF549 dye (right). f Representative images of JF549-labeled Halo-Sox2 molecules, acquired at different time intervals in time-lapse movies. Scale bar: 2 μm. The kymographs below each image correspond to the molecules in the highlighted squares. g Reverse cumulative density function (1-CDF) of Sox TF bound times. Curve shadings: standard deviation (SD) calculated from 100 resampling iterations, each using a randomly selected subset (80%) of the full dataset selected randomly (N = 281, 169, 223, 91, 164, 159 cells). h Residence time distributions obtained from GRID analysis of 100 resampling iterations (randomly selected 80% subset of the full dataset shown in g), assembled into histograms (see the “Methods” section). The GRID amplitude indicates the bound fraction per unit of time for the corresponding residence times.
Among the Sox TFs, Sox2 exhibited the most positively-charged DBD flanking region (DFR), which correlated with the highest fraction of TF molecules associated with mitotic chromosomes40. This has been proposed to reflect its superior ability to nonspecifically bind chromatinized DNA and to efficiently search the genome. Indeed, Sox2 functions as a potent pTF5,41 and is a core pluripotency TF42,43, heterodimerizing with Oct444,45. In contrast, the DFR of another Sox TF, Sox17, was found to carry the most negative charges, and Sox17 only showed weak mitotic binding, potentially indicating a lower ability to nonspecifically scan chromatin, which may limit its pTF function. Sox17 is expressed during differentiation towards extraembryonic endoderm46,47, and can also heterodimerize with Oct448,49. Importantly, Sox2 and Sox17 alone bind to a virtually identical motif50 due to their similar DBD, which allows a direct comparison between the two TFs. Still, upon heterodimerization with Oct4, they bind a slightly distinct composite motif because of differences in their heterodimerization interface located within the DBD.
Here, we investigate the role of differentially-charged DBD-flanking IDR stretches of the Sox2 and Sox17 TFs (DFRSox2 and DFRSox17, respectively) in regulating their ability to search for their binding sites on naked DNA and chromatin, using both in vivo and in vitro single-molecule approaches.
Results
Sox2 and Sox17 mutants to study the DFR impact on target search
We defined the DFRs (Supplementary Fig. 1a) as the segment C-terminal to the DBD, starting at the residue that immediately follows the prominent positive charge peak in each TF’s respective charge distribution (Fig. 1c). The DFR length (116 amino acids) was determined to achieve a large charge difference between DFRSox2 (+1z) and DFRSox17 (−9.5z) while limiting the total number of amino acids included (Supplementary Fig. 1b). We then generated two DFR-swapped constructs: (i) Sox2 carrying the acidic DFRSox17 (Sox2a) and (ii) Sox17 carrying the DFRSox2 (Sox17b) (Fig. 1d). Additionally, constructs containing only the DBDs of Sox2 and Sox17 (Sox2D and Sox17D) were produced to isolate the influence of the DFRs on Sox-DNA binding dynamics.
We engineered mouse embryonic stem cell (mESC) lines containing HaloTag (Halo) fusions to either Sox2, Sox2a, Sox17, Sox17b, Sox2D or Sox17D stably integrated under the control of a doxycycline (dox)-inducible promoter, enabling their expression at low levels for single-molecule imaging51 (Fig. 1e). We then characterized their genome-wide binding profiles by ChIP-seq (Supplementary Fig. 1c). Motif enrichment analysis of ChIP-seq peaks indicated that all Sox TFs bound to nearly identical motifs, including their respective canonical motifs for Sox2 and Sox17 (Supplementary Fig. 1d). Therefore, although Sox17 is not expressed endogenously in mESCs, Sox17 and its mutant Sox17b can still recognize their target sites in this context. Notably, Sox2a bound to motifs that are more similar to those bound by Sox2 than Sox17, confirming that Sox2a still retains the DNA-binding specificity of Sox2. Since Sox2 is crucial for maintaining pluripotency in mESCs, we also determined whether Sox2a can fulfill this role. To this end, we performed a pluripotency rescue assay by using the 2TS22C cell line52, which allows us to fully deplete endogenous Sox2 upon dox addition within 48 h. These cells were also engineered to express Halo-Sox2 or Halo-Sox2a at the same level. We then treated these cell lines with dox for 5 days and quantified the percentage of undifferentiated cell colonies and NANOG expression levels by immunofluorescence (IF) (Supplementary Fig. 1e, f). Halo-Sox2a was able to rescue ESC colony morphology and NANOG expression, even though less potently than Halo-Sox2, demonstrating that Sox2a can partially replace Sox2 in maintaining pluripotency of mESCs.
DFRs impact the residence times of TFs on DNA in vivo
Single-molecule tracking (SMT) allows direct observation of the TF search process within living cells51. We expressed our six different TFs at a low level and labeled them with JF54953. To visualize chromatin, we used SNAP-H2B labeled with silicon rhodamine (SiR)54. We then quantified TF-DNA interaction dynamics using single-molecule highly inclined and laminated optical sheet (smHILO) microscopy. We used four different imaging conditions by fixing the illumination time per frame at 60 ms, and interspersing variable dark times to obtain frame intervals of 120, 240, 480, and 960 ms (Supplementary Fig. 2a). This approach allows us to determine TF binding behavior over a broad temporal range (Fig. 1f).
Most individual molecules were confined within nuclei (Fig. 1e, f). We then reconstructed trajectories for individual molecules using TrackIt55 and plotted the reverse cumulative density function (1-CDF) of bound times. Sox2 and Sox17b exhibited slower 1-CDF decays compared to Sox2a and Sox17 or their DBD alone (Fig. 1g, Supplementary Fig. 2b), indicating slightly longer residence times.
We employed the genuine rate identification (GRID)56 method to resolve the populations of bound molecules in an unbiased manner, categorizing them into distinct bound states. For all TFs, we identified six populations of bound states (Supplementary Fig. 2c) from a distribution of residence times ranging from 0.1 to 1000 s (Supplementary Table 1). While the residence time distributions for Sox2 and Sox17b were narrow and well defined, Sox2a and Sox17 exhibited broader distributions (Fig. 1h), suggesting complex interactions associated with a continuum of binding energies. We note that the uncertainty on the duration and amplitudes of extremely long-lived states in GRID analysis is inherently high, as the determination of these bound times approaches is limited by dye lifetimes. Moreover, the amplitudes of the extremely long-lived states are very low in the spectra. We therefore interpret the slowest components as indicative of rare, very stable binding events rather than precisely quantified lifetimes.
The mean frequency of long-lived binding events (>10 s) in GRID spectra was higher for Sox2 (0.042) compared to either Sox2a (0.018) or Sox17 (0.025) (Supplementary Fig. 2d, Supplementary Table 2). We also calculated fractional occupancies55,56,57,58, i.e., the fraction of total bound time spent in each state (Supplementary Fig. 2e; see the “Methods” section). This revealed that Sox2 (0.66) and Sox17b (0.60) spend more time in long-lived states (>10 s) than Sox2a (0.47) and Sox17 (0.47) (Supplementary Fig. 2f, Supplementary Table 2). Overall, these results suggest that DFRs have an impact on Sox TF-DNA residence times.
DFRs regulate in vivo target search efficiency
Next, we dissected the target search process of the different Sox TFs. We first performed continuous imaging using a 10 ms exposure time to observe both diffusing and bound molecules (Fig. 2a, b). In 2D HILO imaging, fast-diffusing molecules often appear motion-blurred or leave the focal plane, leading to their under-detection59,60. To make sure that this limitation does not bias our results, we applied the state array (SA) method61, which estimates diffusion coefficients and localization error variances on a per-trajectory basis while also correcting for defocalization biases. We obtained diffusion coefficient distributions spanning 10−3 to 102 μm2s−1 (Fig. 2c, d; see the “Methods” section). Strikingly, swapping the DFRs between Sox2 and Sox17 led to a reciprocal swap in their diffusion profiles (Fig. 2d), indicating that the electrostatic properties of the DFRs modulate the diffusion landscape of Sox TFs.
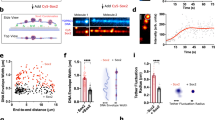
a Representative images from a movie showing H2B-SNAP (red) and Halo-Sox2 (yellow), with color-coded trajectories of Sox2 molecules. Scale bar: 5 μm. b Scheme of the three-state diffusion model with kinetic parameters for diffusion coefficients (D) and corresponding fractions (F) for fast (f), bound (b), and slow (s) diffusion modes. State-array analysis of diffusion coefficients of Sox TF molecules. Each row represents a different cell (N = 86, 40, 38, 80 cells). Posterior state populations are shown as heatmaps (c) and as marginal distributions of diffusion coefficients (d). The dashed black vertical line indicates the diffusion coefficient threshold for bound states at D = 0.01 μm2 s−1. Colored lines: mean. Shaded area: standard error of mean (SEM). Diffusion coefficients from Spot-On single-cell analysis considering the three-state diffusion model for the bound state (e), slow diffusion (f) and fast diffusion (g) (N = 86, 38, 38, 72 cells). Black line: median. Number: median value. Cap: SD. Statistical significance was assessed using Dunn’s test following a nonparametric Kruskal–Wallis ANOVA (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001), and non-significant comparisons are not indicated. h Bound fractions of molecules with D < 0.01 μm2 s−1 corresponding to data shown in (c) and (d). Line: mean. Cap: SD. Box: SEM. Pseudo on-rates of nonspecific (N = 85, 40, 38, 78 cells) (i) and specific (N = 70, 81, 78, 54 cells) (j) interactions per cell, normalized to the mean of Sox2. Line: mean. Cap: SD. Box: SEM. Statistical significance was evaluated using Dunn test following a nonparametric Kruskal–Wallis ANOVA (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001).
Notably, the peak of the diffusion spectrum displayed a pronounced left-skewed tail, suggesting the presence of at least one intermediate state in addition to the bound and freely diffusing populations. Consistent with previous observations for Sox259,62, we modeled the diffusion behavior of all Sox TFs using a three-state diffusion model encompassing fast-diffusing, slow-diffusing, and bound states (Fig. 2b, Supplementary Fig. 3a). Spot-On analysis of jump distance distributions59 revealed that Sox2 and Sox17b generally exhibited lower diffusion coefficients for fast and slow diffusing populations (Fig. 2e–g, Supplementary Table 2).
To quantify the bound population, we defined molecules with diffusion coefficients <0.01 μm2 s−1 as chromatin-bound63,64,65. Under this criterion, Sox2 and Sox17b exhibit higher bound fractions than Sox2a and Sox17 (Fig. 2h), which was also in line with Spot-On analysis (Supplementary Fig. 3b, Supplementary Table 2). Furthermore, Sox2 compared to Sox2a, and Sox17b compared to Sox17 spend less time in fast and slow diffusing modes, respectively, and also transition to a bound state more rapidly (Supplementary Fig. 3c). Taken together, this suggests that individual Sox2 and Sox17 molecules spend less time in a diffusing state before transitioning to a bound state.
Next, we investigated whether the set of TFs differs in their association rate to genomic targets. We measured their pseudo on-rate (kpn)40, defined as the time-averaged binding frequency of TFs to the whole genome, as determined by the number of immobilized molecules per imaging frame (see the “Methods” section). We defined 1 s as a minimal trajectory length to separate nonspecific and specific interactions to determine kpn. Sox2 (kns,pn = 1.00 ± 0.53) and Sox17b (kns,pn = 0.89 ± 0.45) displayed a higher nonspecific kpn than Sox2a (kns,pn = 0.66 ± 0.22) and Sox17 (kns,pn = 0.68 ± 0.39) (Fig. 2i), in line with the previous diffusion analysis. This suggests that the DFRSox2 leads to a higher frequency of nonspecific DNA-binding events for Sox2 and Sox17b. Importantly, Sox2 (ks,pn = 1.00 ± 0.62) exhibits a faster specific kpn than the other TFs (ks,pn = 0.40 ± 0.24 for Sox2a, 0.59 ± 0.46 for Sox17, 0.28 ± 0.19 for Sox17b) (Fig. 2j). In summary, our in vivo data demonstrate that TFs containing a DFRSox2 interact more frequently with DNA and display enhanced target search efficiency compared to TFs containing DFRSox17.
DFRs do not impact 3D diffusion-mediated association to naked DNA and residence times in vitro
A limitation of in vivo measurements is that specific and nonspecific binding events can only be inferred from residence times, and the chromatin context in which they occur is unknown. Therefore, we performed in vitro studies to examine how the DFRs impact DNA binding dynamics of Sox TFs on naked and chromatinized DNA, focusing mainly on Sox2, Sox2a, and Sox17.
We employed a fluorescence colocalization approach37 (Fig. 3a) using single-molecule total internal reflection fluorescence (smTIRF) microscopy. Here, Alexa Fluor 647 (AF647)-labeled DNA or chromatin substrates are immobilized in a flow channel, and their positions are detected. Subsequently, Sox TFs carrying a JF549-dye are introduced. When bound to DNA, they appear as fluorescent spots colocalizing with immobilized DNA molecules. Recording movies enables us to directly determine on-rates and residence times for specific DNA or chromatin environments.
a Left, scheme of a single-molecule experiment to detect Sox TFs binding to DNA. bt: biotin, nt: neutravidin; Right, microscopy images showing DNA loci in the AF647 emission channel (top, red), and bound Sox2 in the JF549 emission channel (bottom, yellow). Scale bar: 2 μm. b EMSA of indicated TFs with Sox motif-containing DNA. c EMSA at different NaCl concentrations for indicated TFs with Sox motif-containing DNA. d DNA templates of different lengths are used for single-molecule binding experiments. e Fluorescence time-trace of Sox2 (2 nM), binding to 50 bp motif-containing DNA, detected by JF549 emission (blue) and fitted with a step-detection algorithm (red). f Fluorescence time-traces of Sox TFs binding to DNA of indicated lengths. The raw data (gray) were fitted (blue, red). g Cumulative distributions of search times for Sox TFs, on various DNA substrates. Curve shading: SD (N = 4 for Sox2 with 50 bp DNA+; N = 3 for all the other cases). h Scheme of the 1D target search model: initially, a TF binds nonspecifically to DNA with ka. Once bound, the TF slides along the DNA with a speed of D1D for ατd until it dissociates from DNA. Upon reaching the target site, the TF either recognizes the target with kr, or bypasses it. sL: sliding length. KR: target recognition constant. i Specific on-rate curve as a function of DNA length. Fit: 1D target search model. Symbol: mean. Cap: SD (N = 4 for Sox2 with 50 bp DNA+; N = 3 for all the other cases). j-n Fitting and derived parameters from the fitting in i (see the “Methods” section): target recognition probability during 1D sliding (j) target recognition constant (k) target recognition rate (l) sliding length (m) and 1D diffusion coefficient (n). Line: mean. Cap: fitting error (N = 4 for Sox2 with 50 bp DNA+; N = 3 for all the other cases).
We produced recombinant Halo-containing Sox2, Sox2a, and Sox17 in insect cells (Supplementary Fig. 4a–c; see the “Methods” section) and validated their DNA-binding ability by electrophoretic mobility shift assays (EMSA), which allowed us to estimate nanomolar DNA affinities for the Sox TFs (Fig. 3b, Supplementary Fig. 4d), consistent with published values41. We also measured the salt dependency of Sox-DNA binding via EMSA by varying NaCl concentrations. Sox2a and Sox17 dissociated from DNA at lower NaCl concentrations (EC50,Sox2a = 14.6 ± 2.1 mM NaCl, EC50,Sox17 = 27.6 ± 3.8 mM NaCl) than Sox2 (EC50,Sox2 = 38.4 ± 5.4 mM NaCl) (Fig. 3c, Supplementary Fig. 4e, f). Therefore, Sox2 binds its consensus site at higher ionic strength, suggesting a charge-dependent effect of the DFRs on the binding mechanism.
We then investigated how charged DFRs influence TF target search on naked DNA. We designed DNA substrates of 25, 50, 100, and 250 bp length, based on the 601 Widom sequence (601WS)66 (Fig. 3d). Each DNA substrate contained a single Sox motif (5’-CTTTGTT-3’) at the center (DNA+), except for a 50 bp control DNA without motif (DNA-). The DNA molecules were immobilized in a flow cell, and the binding dynamics of Sox TFs were monitored using smTIRF imaging. We recorded 10-min movies in the JF549 channel at a frame rate of 5 Hz with a 100 ms illumination time to monitor TF-DNA binding events by colocalization detection (Fig. 3a). Fluorescence time-traces were extracted from the JF549 channel, where high intensity indicates bound states (Fig. 3e). This revealed that the frequency of binding events for all Sox TFs increased with longer DNA, particularly for short binding events (Fig. 3f).
We next determined the on-rates (kon) of Sox TFs by fitting a monoexponential decay to the 1-CDF of unbound times (Supplementary Fig. 5a). This revealed that Sox2, Sox2a, and Sox17 display comparable kon values that increased with DNA length (Supplementary Fig. 5b, Supplementary Table 3).
kon exhibited a nonlinear relationship with DNA length in bp (Supplementary Fig. 5c). The collision radius of two molecules in solution is proportional to their radius of gyration, and this becomes particularly important for DNA lengths above the DNA persistence length of 45 nm67,68,69. Indeed, kon scaled directly with the calculated radii of gyration of the DNA templates (Supplementary Fig. 5d; see the “Methods” section). This relationship allowed us to extrapolate to short DNA lengths and determine a nanoscopic association rate (ka), defined as the association rate to a single site on DNA. We found similar values for all TFs, with ka,Sox2 = 2.14 ± 0.11 × 105 M−1 s−1 bp−1, ka,Sox2a = 1.93 ± 0.21 × 105 M−1 s−1 bp−1, and ka,Sox17 = 1.85 ± 0.13 × 105 M−1 s−1 bp−1 (Supplementary Fig. 5e). This indicates that neither the DFRSox2 nor the DFRSox17 impacts the overall probability of interactions between freely diffusing TFs and naked DNA.
Next, we quantified TF residence times by analyzing the distribution of bound times. The 1-CDFs (Supplementary Fig. 5f) were well-described by a biexponential decay, indicating two distinct binding modes of Sox TFs with DNA+ (Supplementary Fig. 5g–j, Supplementary Table 3): a short-lived (τ1,Sox2 = 1.1 ± 0.3 s, τ1,Sox2a = 0.8 ± 0.3 s, τ1,Sox17 = 1.1 ± 0.4 s) and a long-lived interaction mode (τ2,Sox2 = 14.9 ± 4.5 s, τ2,Sox2a = 12.4 ± 5.2 s, τ2,Sox17 = 10.2 ± 4.5 s). Conversely, the binding of Sox TFs to DNA lacking a motif (DNA-) was characterized by a single short-lived population (τ2,Sox2 = 0.5 ± 0.01 s, τ2,Sox2a = 0.6 ± 0.05 s, τ2,Sox17 = 0.6 ± 0.06 s). This indicates that the long-lived populations, characterized by τ2, correspond to specific, on-target binding. The average specific residence times and bound fractions for all DNA lengths and the three TFs (Supplementary Fig. 5k, l) aligned well with a previously reported value for Sox2 measured by in vitro SMT70. Moreover, for each DNA length, we observed similar specific residence times τ2 for the Sox TFs.
In contrast, it was not possible to directly relate τ1 to nonspecific interactions. Previous measurements using fluorescence correlation spectroscopy in mouse embryos revealed that such nonspecific DNA interactions for Sox2 at individual random sites are extremely short-lived, with microscopic nonspecific residence times of 10 ms71, which is well below the time resolution of our experiment (5 Hz). Similarly, the nonspecific residence time of LacI in E. coli has been reported as less than 5 ms using SMT72, whereas other TFs, such as TetR or p53, exhibited longer nonspecific residence times (158 and 900 ms, respectively)60,73. This suggests that the short-lived interactions we measured as τ1 likely represent the extreme tail of the nonspecific residence time distribution, and may involve numerous transient events of Sox TFs, such as hopping12,19, where TFs repeatedly re-associate and dissociate within a local regime before fully dissociating.
DFRSox2 facilitates target search by increasing the target recognition probability
The overall DNA binding rate (kon) is determined by the intervals between all binding events, regardless of interaction type. Because it encompasses both specific and nonspecific interactions, kon does not directly reflect the efficiency in locating a specific target site. Target search efficiency, in contrast, is defined solely by the intervals between specific interactions, which we refer to as the search time.
To quantify target search efficiency, we introduce a new rate constant: the specific on-rate (ks,on). We used a residence-time threshold of τres > 1 s to identify specific binding events and determined search times (as illustrated in Fig. 3e). We fitted the search time distributions (Fig. 3g) with a monoexponential decay to determine ks,on for the Sox TFs interacting with the different DNA substrates. This analysis revealed that ks,on increased with DNA length for Sox TFs (Supplementary Fig. 5m, Supplementary Table 3). We interpret this increased search efficiency on longer DNA as evidence that 1D sliding facilitates target search. This is consistent with direct experimental evidence for TF sliding for both prokaryotic and eukaryotic TFs and DNA binding proteins19,74,75,76,77. Sox2 consistently exhibited about 1.5 times higher ks,on values compared to both Sox2a and Sox17, suggesting that DFRSox2 enhances target search efficiency over DFRSox17. Furthermore, the nearly identical ks,on values of Sox2a and Sox17 strongly indicate that the DFR plays a key role in enhancing 1D diffusion-mediated target search.
We then implemented a theoretical formulation of the facilitated diffusion model describing the target search of a TF on DNA containing a single target site12,17 (Fig. 3h). Here, the target search process involves three steps. First, a TF binds nonspecifically to a random DNA site. Second, it slides along the DNA to the target until it either dissociates or recognizes and specifically binds the target site. The model is thus defined by four parameters: (i) the nanoscopic association rate (ka), previously determined as about 2 × 105 M−1 s−1 bp−1 for the three Sox TFs (Supplementary Fig. 5e); (ii) the 1D diffusion coefficient on DNA (D1D); (iii) the target recognition rate (kr), which determines the probability of the TF recognizing the specific DNA site when sliding over it; and iv) the nonspecific residence time τd, reporting on the probability of the TF dissociating from nonspecific DNA sites, and set to 10 ms for both TFs71. Since τd sets the intrinsic timescale of the search process, we defined both the sliding and target recognition processes as a function of τd, introducing an average sliding length (sL2 = 2 ∙ D1D ∙ τd) and a target recognition constant (KR=kr ∙ τd) to reduce the number of parameters (Fig. 3h; see the “Methods” section).
The 1D target search model, now a function of two free parameters, KR and sL, was subsequently applied to fit the ks,on dependence on DNA length (Fig. 3i, Supplementary Table 4). The specific binding probabilities during 1D sliding (pbind) were 0.049 ± 0.036 for Sox2, 0.011 ± 0.007 for Sox2a, 0.009 ± 0.010 for Sox17 (Fig. 3j), confirming that DFRSox2 facilitates the TF target search more effectively. Accordingly, KR, which was determined to be 2.3-fold higher for Sox2 (Fig. 3k), resulted in a higher target recognition rate for Sox2 (kr,Sox2 = 2.70 ± 0.99 × 104 s−1) over Sox2a and Sox17 (kr,Sox2a = 1.15 ± 0.16 × 104 s−1, kr,Sox17 = 1.03 ± 0.17 × 104 s−1) (Fig. 3l). Conversely, sL was around 1.4-fold greater for Sox2a and Sox17 (Fig. 3m), corresponding to D1D,Sox2 = 2.63 ± 1.09 × 105 bp2 s−1, D1D,Sox2a = 5.03 ± 2.34 × 105 bp2 s−1, and D1D,Sox17 = 5.55 ± 5.14 × 105 bp2 s−1 (Fig. 3n), in agreement with values reported for LacI19.
Together, this indicates that the DFRSox2 facilitates target recognition of Sox2 during 1D sliding, whereas the DFRSox17 leads to more frequent bypassing of the target site. The slower Sox2 sliding may reflect a trade-off between scanning speed and recognition efficiency78,79.
Sox TFs interact specifically and nonspecifically with nucleosomes
As Sox2 can bind to nucleosomes in vivo, we dissected the impact of DFRSox2 on TF-DNA interactions in the context of chromatin by comparing Sox2 and Sox2a. To examine their binding dynamics to single nucleosomes (Fig. 4a), we reconstituted three different mononucleosomes (MNs), using a 250 bp DNA sequence based on the 601WS and recombinant human histone octamers, resulting in MNs flanked by 50 bp linker DNA. One MN contained a Sox motif positioned at superhelical location −6 (SHL-6), a position close to the nucleosome entry site (MN-6), where Sox2 can bind with Oct444. A second MN had the motif positioned at SHL+2 (MN+2), an internal position where Sox2 can bind and locally distort DNA39 (Fig. 4b; see the “Methods” section). We also reconstituted an MN devoid of any binding motif (MN-). Using the smTIRF colocalization approach, we measured binding kinetics and constructed 1-CDFs of bound times from the resulting fluorescence time-traces (Fig. 4c, Supplementary Fig. 6a–c).
a Scheme of single-molecule experiment to detect Sox TF binding to mononucleosomes. b Structure of nucleosomes bound by Sox2 DBD (Sox2D) together with Oct4 DBD (Oct4POUs) at the heterodimer motif located at SHL-6 (left; PDB: 6T90)44 and by Sox2D at its motif positioned at SHL + 2 (right; PDB: 6T7B)39. Number: SHL. c Representative fluorescence time-traces of Sox2 interacting with MN-6 (blue) and MN- (gray). d Scheme of Sox nucleosome binding modes. States 1 and 2: nonspecific binding to linker DNA and to histones. State 3: Specific binding to target (green). e–h Residence times (e, g) (τ1, gray; τ2, dark gray; τ3, blue) and corresponding bound fractions and fractional occupancies (f, h) of Sox2 (e, f) and Sox2a (g, h) on MNs. Symbol: replicate (N = 4 for MN + 2; N = 3 for MN-6 and MN-). Bar: mean. Cap: SD. Statistical significance between specific bound fraction (blue bars in f, red bars in h) was assessed using two-tailed t-tests (*0.01 < p ≤ 0.05).
A triexponential function, associated with three residence times τ1, τ2, and τ3, was required to adequately describe the binding kinetics of both Sox2 and Sox2a on motif-containing MNs (Supplementary Fig. 6d). In contrast, binding kinetics to MN-, lacking a TF binding site, were well-described by a biexponential decay with two residence times τ1 and τ2. The shortest residence time (τ1,Sox2 = 0.8 ± 0.1 s) was observed across all MNs, and given its similarity to τ1 in naked DNA, could be attributed to nonspecific DNA interactions (Fig. 4d). In contrast, the intermediate residence time (τ2,Sox2,MN-6 = 3.4 ± 0.4 s) was not identified in free DNA and is thus specific for MNs. As it appeared in both motif-containing and motif-absent MNs, we attributed it to TF-histone interactions (Fig. 4d). Finally, the longest residence time (τ3,Sox2,MN-6 = 9.3 ± 0.6 s) was unique to motif-containing MNs, confirming its correspondence to specific interactions (Fig. 4d).
We then compared Sox2 binding to the motifs within MNs at SHL + 2 and SHL−6. While the residence times were comparable for the two positions (Fig. 4e), the fraction of specific binding events was higher in MN + 2 compared to MN−6, indicating that Sox2 has a higher probability of binding to the internal position at SHL + 2 (Fig. 4f). This can be explained by the rotational positioning of the motif, which faces outward in MN + 2 but inward in MN-6, consistent with previous reports70. Finally, a comparison between Sox2 and Sox2a (Fig. 4e–h, Supplementary Table 5) showed similar residence times across all tested MNs.
DFRSox2 increases the fraction of specifically bound molecules on chromatin
We next used single-molecule imaging to assess whether the nature of the DFR is critical for the chromatin invasion ability of Sox237 (Fig. 5a). We assembled chromatin fibers (CFs) on DNA templates containing 10 tandem repeats of the 601WS separated by 30 bp linker DNA, either containing a Sox motif at the entry site (SHL−6) of the fifth nucleosome (CF+) or in the absence of any target motif (CF−) (Supplementary Fig. 7a–c; see the “Methods” section).
a Scheme of single-molecule experiment to detect Sox TF binding to chromatin fibers. b Fluorescence time-traces of Sox TFs interacting with CF+ and CF−. c Specific on-rate of Sox TFs to mononucleosomes and chromatin. The background on-rates (gray) measured with control constructs (MN−, CF−) are overlaid with corresponding constructs containing the target site (green). Symbol: replicate (N = 4 for Sox2 with MN + 2 and CF+, and Sox2a with MN + 2; N = 3 for all the other cases). Bar: mean. Cap: SD. d On-rate of Sox TFs for different nucleosomes. Symbol: replicate (N = 4 for Sox2 with MN + 2 and CF+, and Sox2a with MN + 2; N = 3 for all the other cases). Bar: mean. Cap: SD. Statistical significance was assessed using two-tailed t-tests (*p ≤ 0.05; **p ≤ 0.01; ***p ≤ 0.001).
We then imaged Sox2 and Sox2a binding dynamics to these chromatin fibers. Of note, under our imaging buffer conditions, corresponding to an ionic strength of 120 mM monovalent cations, chromatin is partially compacted37,80. With CF+, Sox2 exhibited more frequent long binding events compared to Sox2a (Fig. 5b). Similar to the MN results, both Sox TFs showed three different residence times on CF + , while they displayed only two nonspecific binding populations on CF− (Supplementary Fig. 7d–h). These residence times closely matched the ranges identified in the MN dataset (Supplementary Table 5). On chromatin fibers, nonspecific interactions predominate because of their large size and the consequent challenge for a TF to locate its embedded target site. However, Sox2 exhibited significantly more long binding events (7.0 ± 1.6%) (Supplementary Fig. 7f) compared to Sox2a (1.2 ± 0.3%) (Supplementary Fig. 7h). This suggests that DFRSox2 enhances Sox2’s ability to access and recognize motifs within chromatin.
We next determined ks,on for Sox2 and Sox2a on MNs and CFs, using a residence time threshold (3.8 s for MNs, 3.4 s for CFs) to select specific binding events, and generated 1-CDF curves of the interspaced search times (Supplementary Fig. 7i, j). We found ks,on to be markedly higher for Sox2 than for Sox2a on all substrates (Fig. 5c). This was especially pronounced for MN + 2 compared to MN-6, highlighting Sox2’s superior ability to interact with internal target sites. Of note, we detected a lower frequency of binding events with residence times above the threshold in the absence of the motif, which originate from the tail of the τ2 lifetime distribution and could be assigned as a background on-rate (shown in gray in Fig. 5c).
In general, a target site within chromatin is expected to be less accessible than on a nucleosome because of nucleosomes stacking in a tetranucleosome conformation81,82,83. However, the effective ks,on of Sox2 increased from MN-6 to CF+ by 1.3-fold (Fig. 5c), suggesting that 1D diffusion along the chromatin, or a higher local TF concentration due to the availability of many nonspecific binding sites, might facilitate target search. In contrast, no such increase in effective ks,on was detected for Sox2a (Fig. 5c). Finally, when including nonspecific binding events (Supplementary Fig. 7k,l), Sox2 exhibited approximately three times higher kon with all the nucleosome constructs than Sox2a (Fig. 5d). As the nature of the DFR did not impact the on-rate for naked DNA (Supplementary Fig. 5c–e), this suggests that the nonspecific interactions of DFRSox2 increase the on-rate specifically on nucleosomes.
In summary, our on-rate results demonstrate that DFRSox2 promotes prevalent nonspecific binding from 3D diffusion to nucleosomes, enabling Sox2 to invade chromatin more effectively.
DFRSox2 enhances in vivo pioneer activity by facilitating closed chromatin binding and initiation of chromatin opening
We next asked how the different DFRs control chromatin invasion of Sox TFs in cells. We induced Sox2 and Sox2a expression in engineered mESC lines, and subsequently performed fluorescence-activated cell sorting (FACS) to select cells with near-identical TF expression levels (Fig. 6a, Supplementary Fig. 8a; see the “Methods” section). We validated the genome-wide binding profiles of exogenously-expressed Halo-Sox2 against endogenously expressed Sox2 via ChIP-Seq3 (Supplementary Fig. 8b, c).
a Cell sorting strategy for ChIP-seq and ATAC-seq: mESCs expressing Halo-Sox2 or -Sox2a were treated with SiR647-Halo ligand and FACS-sorted using indicated gating windows (black) corresponding to near-identical TF expression levels (see also Supplementary Table 13). b Genomic regions enriched by Sox2 (blue) and Sox2a (pink). Two replicates for each Sox TF were merged. c Heatmaps of ChIP-seq data (RPKM) across Sox-enriched regions. d Genome tracks of genomic regions displaying ChIP-seq and ATAC-seq (RPKM) signals, representing Sox2/Sox2a-enriched CORs, and ±10% subregions. The values indicate the number of peaks in each region. e Heatmaps of ChIP-seq and ATAC-seq data (RPKM) aligned to the log2-fold change in ChIP scores of Sox2 over Sox2a across CORs (2/2a) and ±10% subregions. 2/C: log2-fold change in ATAC scores of Sox2 over Ctrl; 2a/C: log2-fold change in ATAC scores of Sox2a over Ctrl. Log2-fold changes were calculated based on the peak signal in bed files. f Mean score profile of Ctrl ATAC data for Sox2-enriched and Sox2a-enriched CORs. g Mean score profile of log2-fold change in ATAC scores of Sox TFs over Ctrl across CORs. h Mean score profile of Ctrl ATAC data for mESCs overexpressing Sox TFs across ±10% subregions. i Mean score profile of the log2-fold change in ATAC scores of mESCs overexpressing Sox TFs compared to Ctrl mESCs across ±10% subregions.
To quantitatively compare the genomic occupancy of Sox2 and Sox2a, we applied a spike-in normalization approach using Drosophila chromatin (see the “Methods” section). We merged regions enriched for one or both Sox TFs (Sox-enriched regions) (Fig. 6b). Sox2a enrichment was well-correlated to Sox2 ChIP scores across Sox-enriched regions, although the occupancy level was lower compared to Sox2 (Fig. 6c, Supplementary Fig. 8d). To analyze genome-wide binding patterns, we identified the commonly occupied regions (CORs) containing ChIP-seq peaks for both Sox2 and Sox2a (Fig. 6b). We then plotted the log2-fold change of the COR ChIP scores for Sox2 over Sox2a in descending order. CORs were further sub-clustered into three groups (Fig. 6d) defined by (i) higher Sox2 binding; (ii) higher Sox2a binding; (iii) equal binding of Sox2 and Sox2a within a ±10% margin. The larger number of Sox2-enriched CORs (Fig. 6d) and higher ChIP enrichment of Sox2 across CORs (Supplementary Fig. 8e) confirmed that Sox2 exhibited greater genomic occupancy compared to Sox2a. Besides, motif analysis across CORs and exclusive regions revealed comparable binding specificity of Sox2 and Sox2a (Supplementary Fig. 8f).
To assess the ability of Sox2 and Sox2a to open compact regions, we generated ATAC-seq data for the same cell lines. We aligned ATAC-seq profiles with ChIP log2-fold changes to assess chromatin accessibility across CORs. CGR8 cells expressing only H2B-SNAP (Ctrl) served as a control of initial chromatin accessibility (Fig. 6e). mESCs overexpressing Sox2 exhibited higher chromatin accessibility at CORs than those overexpressing Sox2a (Supplementary Fig. 8g). However, Ctrl ATAC-seq revealed that Sox2-enriched CORs were initially 1.7-fold more closed compared to Sox2a-enriched CORs (Fig. 6f). This confirms that DFRSox2 enhances chromatin invasion of Sox2, as we observed in vitro. We then computed the log2-fold change of ATAC scores for Sox2 and Sox2a over Ctrl. Sox2 increased chromatin accessibility across CORs more than Sox2a (Fig. 6g, Supplementary Fig. 8h).
Finally, we determined changes in chromatin accessibility in regions with comparable Sox2 and Sox2a occupancy (within a range of ±10 %) and initial chromatin accessibility (Fig. 6h). Chromatin accessibility increased more for Sox2 than for Sox2a (Fig. 6i). Altogether, this data indicates that DFRSox2 allows Sox2 to bind to poorly accessible chromatin and to open bound regions more effectively, which endows Sox2 with superior pioneering activity.
Discussion
Eukaryotic TFs search the genome within a highly complex nuclear environment, characterized by chromatinized DNA organized into higher-order structures including chromatin fibers, loops84,85, TADs, and A/B compartments86. Pioneer TFs such as Sox2 evolved mechanisms to penetrate compact chromatin states, to search the underlying DNA, and to open chromatin structure after target motif recognition36,87,88. Earlier in vivo single-molecule imaging revealed that Sox2 spends significant time in free 3D diffusion6,62,87,89, which may reflect a generic strategy used by eukaryotic TFs to overcome obstacles such as nucleosomes that prevent sliding90.
A major challenge in understanding gene regulation is uncovering how biochemical properties of TFs shape target search efficiency, particularly given the role of large disordered regions in search dynamics, target selection, and overall TF function91. Although these domains lack a defined structure, both short sequence elements and broader physicochemical properties (e.g., charge and hydrophobicity), particularly in the DFR, modulate DNA-binding characteristics of TFs. These have significant implications: studies in yeast25,92,93 and mammalian cells30,33,87,94,95 demonstrate that the DBD alone is insufficient for target selection, and that truncating the DFR alters site specificity. Moreover, diffusional properties in the nucleus are modified upon swapping or deleting DFRs, including a loss of diffusional confinement87,96 and reduced scanning of compact chromatin regions33,92. To understand how the DFR impacts target search, we mechanistically dissected this process for Sox2 within the chromatinized genome, combining in vitro and in-cell biophysics and genomics.
Here, we compare the effects of DFRs from Sox2 and Sox17, two TFs that show highly different mitotic chromatin binding40, albeit having homologous DBDs recognizing a very similar DNA motif. We revealed distinct mechanisms by which DFRSox2 and DFRSox17 control TF target search on DNA versus chromatin (Fig. 7a). On naked DNA, DFRSox2 drives rapid target recognition during 1D sliding and reduced sliding speed, whereas DFRSox17 results in an increased diffusion coefficient but more frequent target site bypassing. Indeed, the DFRSox2 may resolve the speed-stability paradox of TF site recognition, which states that the smooth energy landscape required for fast diffusion on DNA limits the ability of TFs to recognize and stably bind their target sites9,15. Here, nonspecific DNA interactions of the DFRSox2 may limit the sliding speed of Sox2 by increasing the roughness of the energy landscape, thereby rendering specific site binding more probable. We propose that the electrostatic properties of the DFRs play a key role in this process. The DFRSox2 is more positively-charged and results in stronger nonspecific DNA interactions compared to other Sox TF DFRs, such as DFRSox17. Such an electrostatic mechanism is also consistent with stronger DNA binding when challenged with high salt concentrations. Overall, we thus show that more positively-charged DFRs result in higher TF search efficiency, in agreement with our earlier observations40. Of note, negatively-charged IDRs were also reported to control nonspecific DNA binding through an autoinhibition mechanism11,93, but both Sox2 and Sox17 lack such highly negatively charged D/E repeat domains94.
a Distinct target search mechanism on chromatinized versus naked DNA. b DFRSox2 enhances the abilities of chromatin invasion and local chromatin opening.
On chromatinized DNA, the DFRSox2 exhibits an even more pronounced effect, as it promotes frequent nonspecific interactions with both histones and nucleosomal DNA, which can transition into specific target binding. This facilitates chromatin invasion by enabling the TF to access DNA sites embedded within compact chromatin (Fig. 7b). Conversely, DFRSox17 is associated with a lower overall on-rate and thus fewer nonspecific binding events, resulting in a strongly reduced chromatin-binding ability. This dual function of DFRSox2 thereby drives efficient target search within the complex chromatin environment, driving potent pioneer activity of Sox2 for local chromatin accessibility control (Fig. 7b). We note that the 601WS positioning sequences, while providing precise nucleosome placement, cannot recapitulate the heterogeneity of native chromatin, where nucleosome positioning, stability, and occupancy vary. However, the strong concordance with in vivo data indicates that the main conclusions extend beyond the reconstituted system.
We propose that Sox2 can diffuse along chromatin by exploiting nucleosome architecture. The close spatial proximity of the two DNA gyres on the nucleosome surface allows Sox2 to bypass extensively wrapped regions, while steric occlusion of the inward-facing gyre restricts sampling to exposed turns. Furthermore, spontaneous chromatin fluctuation97 transiently reveals buried DNA segments. Consequently, TFs with high nonspecific chromatin-binding ability, like Sox2, may exploit these fleeting openings to outcompete histones and invade compact chromatin more efficiently.
In vivo GRID analysis revealed six Sox TF–chromatin binding states spanning from 0.18 to 380 s in line with recent studies investigating TF binding dynamics that also revealed complex residence time distributions34,58,98,99,100. As supported by our in vitro measurements, intermediate residence times (10–20 s) likely correspond to specific DNA binding, while shorter-lived states (sub-second to a few seconds) reflect transient nonspecific interactions with DNA, nucleosomes, or higher-order chromatin. The longest-lived states (>50 s) are consistent with previously reported extended TF residence on regulatory chromatin101,102 and may involve cofactor binding, transcriptional complex formation, or chromatin remodeling. Of note, our experiments focused solely on interactions of individual TFs with DNA or chromatin in the absence of other factors. Possible DFR-mediated changes in protein-protein interactions, in particular potential differences in cofactor recruitment, were not evaluated in our work, and we cannot fully exclude that such effects might contribute to the observed differences between the Sox TF variants in vivo. Moreover, at high local concentrations, Sox2 has the propensity to form condensates95,103, which increases local retention and may play an important role in super-enhancer activation104,105. IDRs often contribute to condensate formation, which may further modulate the search dynamics. While condensates may contribute to cellular dynamics, in our in vitro studies, the experimental conditions did not result in phase separation, and thus the on-rate-enhancing effects observed are independent of condensate formation.
In summary, our results show that rapid, transient nonspecific chromatin-binding events and enhanced target recognition on bare DNA, both relying on a finely tuned charge balance, allow Sox2 to sample the chromatinized genome efficiently. We propose that this mechanism, driven by DFR electrostatic properties, may represent a general principle extending to a broad range of eukaryotic TFs, in particular pioneer TFs that need to access compact chromatin regions.
Methods
mESC culture
mESCs were routinely cultured on 0.1% gelatin-coated (Sigma, G9391) 100 mm Petri dishes at 37 °C with 5% CO2 in GMEM (Sigma, G5154-500ML), supplemented with 10% ESC-qualified fetal bovine serum (Gibco, 16141-079), 1% nonessential amino acids (Gibco, 11140-050), 2 mM L-glutamine (Gibco, 25030-024), 2 mM sodium pyruvate (Sigma, S8636-100ML), 100 μM 2-mercaptoethanol (Sigma, 63689-25), 1% penicillin–streptomycin (BioConcept, 4-01 F00-H), in-house produced leukemia inhibitory factor (LIF), CHIR99021 (Merck, 361559-5MG) at 3 μM and PD184352 (Sigma, PZ0181-25MG) at 0.8 μM. Cells were passaged every 2–3 days using trypsinization (Sigma, T4049-100ML). Once cells reached 60–70% confluency, one-sixth to one-tenth of the culture was used for the next passage.
For imaging, one day prior to measurement, 170 μm glass-bottom 35 mm Petri dishes were pre-coated with a 1:10 dilution of Biolaminin (BioLamina, LN511-0202) in DPBS containing magnesium and calcium ions (Gibco, 14040117). Cells were then plated on the dishes in FluoroBrite DMEM (ThermoFisher, A18967-01) supplemented with 10% ESC-qualified fetal bovine serum, 1% nonessential amino acids, 2 mM L-glutamine, 2 mM sodium pyruvate, 100 µM 2-mercaptoethanol (Sigma-Aldrich, 63689-25ML-F), 1% penicillin/streptomycin, LIF, CHIR99021 at 3 µM, and PD184352 at 0.8 µM.
Lentiviral vector production
Lentiviral vectors were produced by transfection of HEK 293T cells with the envelope (psPAX2, Addgene, 12260), packaging (pMD2.G, Addgene 12259), and the lentiviral construct of interest using calcium phosphate transfection106,107. The HEK293T cells were cultured in GlutaMAX-containing DMEM (Gibco, 31966021) with 10% fetal bovine serum (Gibco, 10270106) and 1% penicillin–streptomycin (BioConcept, 4-01F00-H). At 2 days post-transfection, when the cells reached 80% confluency, the virus-containing medium was harvested and cleared of cell debris by centrifugation. Viral vectors were concentrated 120-fold by ultracentrifugation at 20,000×g for 120 min at 4 °C. 50,000 cells in 1 mL of medium in a 12-well plate were transduced with 50 μL of concentrated lentiviral vector particles to generate stable cell lines.
Generation of dox-inducible Halo-Sox TF-expressing mESCs
Sox TF-expressing embryonic stem (SES) cell lines in Supplementary Table 6 are mESC derivatives of CGR8 cells engineered to express Halo-Sox TFs upon doxycycline induction. Three transgenes were inserted into the genome using a lentiviral vector approach. The lentiviral plasmids (pLV-TRE3G-Halo-Sox) were generated using the pLV-TRE3G-Halo-Sox2 plasmid108 and the cDNA sequence of Sox1740, which were sourced from the laboratory. Using the In-fusion cloning technique (Takara Bio, 638947), we generated lentiviral plasmids containing transgenes for every Halo-Sox TF. The obtained products were transformed into HB101 competent cells. Clones were sequenced to verify the presence of the sequence of interest.
CGR8 cells (Sigma, 07032901-1VL) were transduced to constitutively express rtTA3G, Halo-Sox TFs, or H2B-SNAP. The genes encoding the recombinant proteins, Halo-Sox TFs (sequences in Supplementary Table 7), regulated by seven Tet operators, and H2B-SNAP driven by an hPGK promoter, were integrated into the mouse genome of SES cells as indicated in Supplementary Table 6. To select SES cells containing the transgene of interest, cells were treated with 2 µg/mL of puromycin (Gibco, A11138-03) for the TRE3G-Halo-Sox2 transgene and/or 1 µg/mL of blasticidin for PGK-rtTA3G for a week. PGK-H2B-SNAP positive SES cells were sorted by FACS based on SNAP-SiR647 intensity after 30 min incubation of SNAP-labeling SiR647 dye (NEB, S9102S) in the cells.
Generation of 2TS22C cells expressing Halo-Sox2 or Halo-Sox2a
The 2TS22C cell line52 enables knockdown of endogenous Sox2 upon dox treatment. Similar to the dox-inducible Halo-Sox expressing mESCs (Supplementary Table 6), 2TS22C cells expressing Halo-Sox2 (SES13) or Halo-Sox2a (SES14) were generated using lentiviral vector transduction. To generate pLV-EF1α-Halo-Sox plasmids, mCherry in the pLV-EF1α-mCherry plasmid fragment was exchanged with Halo-Sox2 or Halo-Sox2a using restriction cloning. Finally, SES13 and SES14 were sorted by FACS based on Halo-SiR647 intensity after a 30 min incubation with 50 nM of Halo-SiR647 dye at 37 °C.
Pluripotency maintenance assay
To ensure similar expression levels of Halo-Sox2 and Halo-Sox2a, SES13 and SES14 were incubated with Halo-labeling SiR647 dye for 30 min at 37 °C followed by FACS using the intensity gate, resulting in the same mean fluorescence intensity between SES13 and SES14 samples. Immediately after FACS, 1000 cells were seeded on gelatin-coated, 170 μm glass-bottom 35 mm Petri dishes in GMEM (Sigma, G5154-500ML), supplemented with 10% ESC-qualified fetal bovine serum (Gibco, 16141-079), 1% nonessential amino acids (Gibco, 11140-050), 2 mM L-glutamine (Gibco, 25030-024), 2 mM sodium pyruvate (Sigma, S8636-100ML), 100 µM 2-mercaptoethanol (Sigma, 63689-25), 1% penicillin–streptomycin (BioConcept, 4-01 F00-H), LIF, CHIR99021 (Merck, 361559-5MG) at 3 µM and PD184352 (Sigma, PZ0181-25MG) at 0.8 µM. Following this, 1 µg/mL of dox was added to induce knockdown of endogenous Sox2. The medium supplemented with dox was changed on the third and fifth day after seeding.
On the sixth day after the first dox addition, cells were fixed with 4% formaldehyde (ThermoFisher, 28908) for 10 min at room temperature, gently washed with DPBS (Gibco, 14040091), and stored at 4 °C in 2 mL of DPBS. Since differentiation of mESCs leads to colony flattening and loss of regular edges109,110,111, colony morphology serves as a metric of the pluripotency status. The shape of cell colonies was assessed using phase-contrast microscopy (Zeiss LSM700 UP2, BIOP, EPFL) with a ×10 air objective (Objective EC Plan-Neofluar ×10/0.3 Ph1 M27) on an inverted Zeiss Axio Observer Z1 microscope.
For NANOG IF imaging, on the fifth day after the first dox addition, 170 μm glass-bottom 35 mm Petri dishes were pre-coated with a 1:10 dilution of Biolaminin (BioLamina, LN511-0202) in DPBS containing magnesium and calcium ions (Gibco, 14040117). 600 cells were seeded on the dishes with fresh medium supplemented with dox. After cell fixation on day 6 using 4% formaldehyde (ThermoFisher, 28908), cells were permeabilized with 0.5% Triton (AppliChem, A1388,0500) at room temperature, followed by multiple washes using DPBS (Gibco, 14040091). Blocking was performed with 1% BSA solution (Sigma-Aldrich, A7906; in DPBS) for 30 min. Subsequently, a rabbit anti-NANOG antibody (CST, 8822) was diluted 1:500 in 1% BSA and incubated with the cells overnight at 4 °C. After washing the antibody off with DPBS, cells were incubated for 1 h at room temperature with AF647-conjugated secondary Chicken anti-Rabbit IgG antibody (Invitrogen, A-21443) at a 1:500 dilution in 1% BSA solution. The cells were washed gently twice with 0.1% Tween-20 (Fisher Scientific, 10113103; in DPBS) and once with DPBS. Finally, Fluoromount G with DAPI (SouthernBiotech, 0100-20) was applied to the samples, and cells were incubated for at least 10 min before imaging.
IF imaging was performed on an Inverted Nikon Eclipse Ti2-E motorized microscope using confocal microscopy (Nikon-CSU-W1, BIOP, EPFL) with a ×60 oil objective (Objective CFI Plan Apo Lambda ×60/1.40). 638 and 405 nm diode lasers were used to excite AF647 dye and DAPI, respectively. For quantification, the StarDist plugin112 in Fiji113 was employed to detect nuclei based on DAPI signals. AF647-NANOG IF intensity was then measured within the nuclei of each cell.
In vivo SMT sample preparation
One day prior to imaging, approximately 105 cells were plated on a 170 μm glass-bottom 35 mm Petri dish in FluoroBrite DMEM (ThermoFisher, A18967-01) supplemented with 10% ESC-qualified fetal bovine serum (Gibco, 16141-079), 1% nonessential amino acids (Gibco, 11140-050), 2 mM L-glutamine (Gibco, 25030-024), 2 mM sodium pyruvate (Sigma, S8636-100ML), 100 μM 2-mercaptoethanol (Sigma, 63689-25), 1% penicillin–streptomycin (BioConcept, 4-01 F00-H), LIF, CHIR99021 (Merck, 361559-5MG) at 3 µM and PD184352 (Sigma, PZ0181-25MG) at 0.8 µM. The dishes were already pre-coated with a 1:10 dilution of Biolaminin (BioLamina, LN511-0202) in DPBS containing magnesium and calcium ions (Gibco, 14040117).
The specific concentrations of dyes and dox to visualize Halo-Sox molecules were optimized for each Halo-Sox TF. Cells were incubated overnight with dox to induce Halo-Sox TF expression. Before the measurement, we performed Halo-labeling with the JF549 dye (Promega, GA1110) to measure the residence times and diffusing dynamics. Dox and Halo-JF549 concentrations are indicated in Supplementary Table 8. For all cell lines and conditions, 1–2 nM of SNAP-labeling SiR647 (NEB, S9102S) was applied. Cells were incubated with the dyes for 30 min at 37 °C to visualize Halo-Sox TFs and SNAP-H2B, respectively, before imaging. Excess dye was gently washed out using the imaging medium.
In vivo smHILO microscopy setup
An inverted Nikon Eclipse Ti2-E fluorescence microscope equipped with a ×100 oil immersion NA 1.49 objective (SR HP Apo TIRF) was employed. Fluorophore excitation was achieved using the N-STORM Direct Point illuminator. Two-color imaging was performed with a 30 mW 561 nm diode laser for Halo-JF549 excitation, and a 50 mW 638 nm diode laser for SNAP-SiR647 excitation, with a 405/488/568/647 DM wheel and DM 568 LP emission filter. Dye emission was detected using two Photometrics Prime 95B sCMOS cameras with a physical pixel size of 11 × 11 µm2.
Our multi-time interval approach interleaves 60 ms illumination with dark periods of 60, 180, 420, and 900 ms. This design creates overlapping observation windows that capture both fast off-rates and long-lived binding events, ensuring comprehensive sampling of all intermediate lifetimes. This multi-time interval measurement also enabled correction for photobleaching, since the time-lapse imaging conditions only affect the photobleaching rate without altering residence times. The number of cells per time interval and Sox TF is indicated in Supplementary Table 9. SNAP-H2B was imaged with a 60-ms exposure time once every 9–10 frames of Halo-JF549.
To measure diffusion dynamics, we recorded continuous time-lapse movies for 30 s with a 100 Hz frame rate (cell numbers are indicated in Supplementary Table 9). Prior to capturing Sox TF kinetics, a snapshot of SNAP-H2B was taken with a 100 ms exposure. We used the same microscope settings as for diffusion analysis, except for the laser power for JF549 activation, which was increased to 110 mW.
Expression, labeling, and purification of Sox TFs
Each Halo-Sox TF construct contained a twin-Strep tag at the C-terminus and an 8×His tag at the N-terminus, and was cloned into the donor plasmid pACEBac1 (bearing a gentamicin resistance marker and a YFP reporter gene). Baculovirus production from Sf9 cells and subsequent Halo-Sox TF expression in Hi5 cells were performed by the Protein Production and Structure Core Facility at EPFL. For purification, the cell pellet harvested from 300 mL of cell culture was suspended in a lysis buffer (100 mM KCl, 150 mM NaCl, 50 mM Tris, 10 mM HEPES at pH 7.6, 2 mM DTT, 1 mM EDTA, protease inhibitor cocktail, 0.2 mM PMSF, 0.1% IGEPAL, 20–30 U/mL DNase I, 5 mM MgCl2, and 5 mM CaCl2). The suspension was lysed by sonication at 30% amplitude (750 W/20 kHz) with 3/27 s on/off pulses for 5 cycles on ice. The lysate was clarified by three rounds of centrifugation at 16,000–21,000×g for 15–30 min each.
Twin-Strep affinity purification was performed on ice using 1 mL of Strep-Tactin Superflow high-capacity resin (IBA Lifesciences, 2-1208-010-20ML) according to the standard gravity-flow protocol. The resin was first washed with 2 mL of buffer W (100 mM Tris, pH 7.6, 150 mM NaCl, 2 mM DTT, 1 mM EDTA) and preincubated in cold lysis buffer. After loading the lysate, the column was washed with 7 mL of W buffer and then eluted with 12 mL of E buffer (100 mM Tris, pH 7.6, 150 mM NaCl, 2 mM DTT, 1 mM EDTA, 10 mM desthiobiotin). Elution progress was monitored by SDS–PAGE gel electrophoresis (Supplementary Fig. 4a–c). The eluted sample was incubated overnight on ice with a 1.1 molar equivalent of Halo ligand-JF549 (Promega, GA1110) in buffer S (100 mM KCl, 150 mM NaCl, 10 mM HEPES at pH 7.6, 2 mM DTT). After size-exclusion chromatography on a Superdex 200 Increase 10/300 GL column (Cytiva, 28990944) (Supplementary Fig. 4a–c), the Halo-Sox TFs were assessed for labeling efficiency (92–99%) and for concentration (1–2 µM) by UV–Vis spectroscopy (Supplementary Fig. 4a–c; Supplementary Table 10).
Electrophoretic mobility shift assays (EMSA)
The protein concentration was titrated from 0.8 to 50 nM while maintaining a constant concentration (2 nM) of AF647-labeled DNA substrate. Sdna6 (Sox motif: 5’-CATTGTG-3’; Supplementary Fig. 4d) and Sdna4 (Sox motif: 5’-CTTTGTT-3’; Fig. 3b) in Supplementary Table 11 were used as motif-containing DNA substrates. Especially, Sdna6 sequence corresponds to a region (−483 to −454)89 of the human miR302 gene. Additionally, five-fold excess (10 nM) of poly(deoxyinosinic-deoxycytidylic) acid (poly dI-dC, ThermoFisher, 20148E) was added to reduce nonspecific interactions between Sox TFs and DNA114. The EMSA buffer consisted of 20 mM HEPES, 20 mM Tris, 1 mM DTT, 50 mM KCl, 0.5 mg/mL BSA, 0.02% Tween-20, 10% glycerol, 3.2% glucose, and 2 mM Trolox, at pH 7.5. After 30 min of incubation at room temperature, the reactions were separated by native PAGE using a 5% Tris base, boric acid, EDTA (TBE) gel, run in 0.5 x TBE buffer, at 120 V for 50 min on ice. Dissociation constants were estimated by quantifying the proportion of bound versus unbound DNA in each lane as the Sox TF concentration increased, based on AF647 fluorescence emission.
To determine the ionic strength sensitivity of Sox TFs to DNA binding via EMSA, NaCl was titrated from 12.5 to 400 mM into the mixture of 1 nM fluorescently-labeled DNA substrate (Sdna6 in Supplementary Table 11; Sox motif: 5’-CATTGTG-3’), 10 nM Sox TFs in 20 mM HEPES, 20 mM Tris, 1 mM DTT, 12.5 mM KCl, 0.5 mg/mL BSA, 0.02% Tween-20, 10% glycerol, at pH 7.5. After 60 min of incubation at room temperature, the reactions were separated by native PAGE (5% TBE gel, 0.5 × TBE buffer, 50 min at 120 V on ice). EC50 values were estimated by quantifying the proportion of bound versus unbound DNA in each lane as the NaCl concentration increased, based on AF647 fluorescent emission.
Oligonucleotide labeling
Synthetic oligonucleotides containing amino-modified C6 at their 5’-end were subjected to two rounds of ethanol precipitation prior to labeling. To initiate the labeling reaction, 10–20 nmol of oligonucleotide were dissolved in 50 µL of 0.1 mM sodium tetraborate buffer (pH 8.5). A two molar-equivalent amount of the NHS ester-functionalized fluorophore (AF647 (Invitrogen, A20006) or XFD647 (AATbio, 1833), which are chemically identical) was then added to the solution. The mixture was incubated in the dark at room temperature and agitated at 300 rpm overnight. After ethanol precipitation to eliminate unreacted dye, RP-HPLC purification was performed via RP-HPLC on an InertSustain C18 column (GL Sciences, 5020-07445) under a 20-min gradient from 0% to 70% Solvent B (100 mM triethylammonium acetate, pH 7, mixed with acetonitrile), with a flow rate of 1 mL/min and detection at 260 and 640 nm. The labeled oligonucleotides were resuspended in Milli-Q water at a final concentration of 0.1–1 mM and stored at −20 °C.
Generation of naked and mononucleosome DNA
For DNA substrates shorter than 200 bp (Sdna3, Sdna4, Sdna5, Sdna6, Sdna8, in Supplementary Table 11), two complementary oligonucleotides that are either biotinylated or AF647-conjugated (4–40 µM each) were annealed in 50–200 µL of 1× Phusion HF buffer. The mixture was heated to 95 °C and then cooled to 20 °C at a rate of −1 °C/min, followed by a 1-h incubation at 55 °C to ensure complete strand hybridization. For double-stranded DNA longer than 200 bp (Sdna9, Sdna10, Sdna11, Sdna16, in Supplementary Table 11), PCR amplification was performed. Each 50 µL reaction in 1× Phusion HF buffer contained 100–600 nM of a fluorescently labeled primer, a 1.1–1.2 molar equivalent of a second primer, 200 µM dNTPs (NEB, N0447S), and 1–2 U of Phusion DNA polymerase (NEB, M0530S). The two primers were either conjugated with AF647 or biotinylated. Template DNA (1–5 ng) was typically used, except for Sdna4. For Sdna4, both primers were used at 3 µM to generate primer dimers during amplification. The thermocycling program included an initial denaturation at 95 °C for 1 min, followed by 25–30 cycles of 95 °C for 30 s, 60–68 °C for 15 s, and 72 °C for 30 s, with a final 5-min extension at 72 °C. 8–48 parallel reactions were conducted to increase DNA yield. Both the annealed and PCR-amplified products were then purified using QIAquick PCR purification kit (QIAGEN, 28104) and verified by native polyacrylamide gel electrophoresis.
Preparation of 1xMMTV buffer DNA
MMTV buffer DNA (Sdna19 in Supplementary Table 11) for chromatin reconstitution was derived from a plasmid carrying eight tandem repeats of the MMTV sequence (8×MMTV DNA). After growing large-scale plasmid cultures (100 mL), DNA was purified using a midiprep kit (QIAGEN, 12943), digested overnight at 37 °C with EcoRV-HF (NEB, R3195L) and Quick CIP (NEB, M0525L), followed by incubating the digest at 80 °C for 20 min to inactivate the enzymes. The resulting 1× MMTV fragments (151 bp) were purified from the plasmid backbone using PEG6000 precipitation. Residual PEG was removed through PCR purification kits (QIAGEN, 28104), ensuring a clean preparation of the MMTV DNA fragments.
Generation of plasmids containing 10-mer 601WS nucleosome arrays
To obtain a 10×601WS array featuring 30-bp linkers and incorporating a Sox motif at the SHL-6 position of the fifth nucleosome (Supplementary Fig. 7a; Sdna17 in Supplementary Table 11), we adopted a plasmid-based strategy for precision and reproducibility. The final plasmid, Spla38 was assembled through a two-step molecular cloning workflow using RecP1115 and Spla37. Their relevant sequence blocks used in this work are marked below:
RecP1: pWM531-EcoRI-DraIII-1×601WS-BamHI-4×601WS-BglII-BsaI Spla37: pMARQ-BglII-Sox601WS-DraIII
First, the 5×601WS plasmid (Spla42), which contains a Sox motif at the fifth nucleosome position, was generated. The BamHI-4×601WS-BglII segment from RecP1 was inserted into Spla37 (pMARQ-BglII-Sox601WS) by BamHI/BglII restriction cloning. Before digestion, Spla37 was PCR-linearized using a reverse primer (Soli179 in Supplementary Table 12) containing 5’-BamHI-EcoRI-3’ restriction sites. The resulting construct, Spla42 is summarized below:
Spla42: pMARQ-EcoRI-BamHI-4×601WS-BglII-Sox601WS-DraIII
Second, the 10× 601WS plasmid (Spla38) containing a Sox motif at the fifth nucleosome was cloned. The EcoRI-4×601WS-Sox601WS-DraIII segment from Spla42 was transferred back into RecP1 (pWM531-EcoRI-DraIII-5×601WS) by EcoRI/DraIII restriction cloning, yielding the final plasmid, Spla38 shown below:
Spla38: pWM531-4×601WS-Sox601WS-5×601WS-BsaI
To generate a Sox motif-free 10×601WS control plasmid (Sdna18 in Supplementary Table 11), Spla37 was replaced with Spla36 (pMARQ-BglII-601WS-DraIII) and the same cloning workflow was applied. The intermediate and final plasmids are:
Spla43: pMARQ-EcoRI-5×601WS-DraIII Spla35:
pWM531-10×601WS-BsaI
Restriction enzymes used in this workflow included BamHI-HF, BglII, EcoRI-HF, and DraIII-HF (NEB). Both Spla38 and Spla35 were validated by Sanger sequencing and by diagnostic digests (EcoRI-HF, EcoRV-HF, BsaI-v2-HF, and HindIII-HF; NEB). Validated plasmids were expanded in 200 mL cultures of DH5α E. coli and purified using the QIAquick midiprep kits (QIAGEN, 12943) followed by QIAquick PCR purification kits (QIAGEN, 28104).
Purification of chromatin DNA
To purify chromatin DNA for the assembly of CF+ or CF− (Sdna17 and Sdna18, respectively; sequences in Supplementary Table 11), a standard digestion reaction was set up with 1000–2500 pmol of plasmid DNA (Spla38 or Spla35) in a final volume of 1.4 mL of 1× γCutSmart buffer. The reaction was supplemented with 1200 units of EcoRI-HF (NEB, R3101L), 2000 units of EcoRV-HF (NEB, R3195L), and 500 units of Quick CIP (NEB, M0525L), and incubated at 37 °C for 20 h. The enzymes were then inactivated by heating to 80 °C for 30 min.
To generate a BsaI overhang 3’-end, 1000 units of BsaI-HFv2 (NEB, R3733S) were added in a final volume of 1.46 mL of 1× γCutSmart buffer, followed by incubation at 37 °C for 20 h.
The digested DNA was then purified through multiple rounds of 4–7.5% PEG6000 precipitation to remove plasmid backbone fragments. After adjusting the NaCl concentration to 500 mM, samples were incubated on ice for 30 min and centrifuged at 21,000×g for 30 min at 4 °C to pellet short DNA fragments. The supernatant was retained at each step, while the PEG6000 concentration was increased in 0.5% increments. Gel electrophoresis of both the supernatant and the pellet at each step monitored the purification progress. Once the target DNA fragments reached high purity, remaining PEG was removed using QIAquick PCR purification kits (QIAGEN, 28104).
To generate the biotinylated adapter oligo (Sanc1 in Supplementary Table 11) as described previously116, a biotin-labeled oligonucleotide was annealed in 1× T4 DNA ligase buffer to its complementary strand (Soli176 and Soli175, respectively, in Supplementary Table 12), which carries a 5′-phosphorylated BsaI overhang and an AF647 fluorophore introduced via an NHS-ester labeling protocol. Next, 800–1600 pmol of chromatin DNA possessing a BsaI overhang (previously purified by PEG precipitation) were mixed with a 2.5-fold molar excess of Sanc1. The mixture was ligated overnight at 16 °C in the dark using 8000 units of T4 DNA ligase (NEB, M0202S). Excess Sanc1 was removed by PEG precipitation, and the resulting ligated DNA (Sdna17 or Sdna18 in Supplementary Table 11) was further purified with the QIAquick PCR purification kit (QIAGEN, 28104).
Octamer refolding
To refold histone octamers, equimolar ratios of recombinant, purified human H3.2 containing the C110A substitution and wild-type human H4 were combined in an unfolding buffer (6 M Guanidine–HCl, 20 mM Tris–HCl, 5 mM DTT, pH 7.5) alongside 1.1 equivalents of wild-type human H2A and human H2B. The final protein concentration was adjusted to 0.5–1 mg/mL. Refolding proceeded via dialysis in a Slide-A-Lyzer Dialysis Cassette (ThermoFisher, 66332) against a refolding buffer (2 M NaCl, 10 mM Tris, 1 mM EDTA, 1 mM DTT, pH 7.5). Any aggregates were removed by centrifugation at 2500×g for 20 min at 4 °C. The resulting octamers were purified by size-exclusion chromatography on a Superdex 200 Increase 10/300 GL column (Cytiva, 28990944) and analyzed by SDS–PAGE and RP-HPLC. Fractions containing pure octamers were pooled and concentrated to ~40 µM. Glycerol was then added to a final concentration of 50%, and these octamer stocks were stored at −20 °C until use.
Nucleosome assembly
Mononucleosomes (MN−6, MN+2, and MN−) and chromatin fibers (CF+ and CF−) containing up to 10× 601WS nucleosome positioning sequences (NPSs) with 30 bp linkers were assembled through high-salt to low-salt dialysis (from 2 M to 10 mM NaCl) (Supplementary Fig. 7a). In a high-salt buffer (10 mM Tris, 1 mM EDTA, 2 M NaCl), histone octamers were mixed with DNA (Sdna10, Sdna11, Sdna16, Sdna17, or Sdna18 in Supplementary Table 11) at NPS-to-octamer ratios ranging from 1:1 to 1:2.5. To avoid excessive octamer binding to the fibers, 0.25 equivalents of MMTV DNA, which has lower affinity for mononucleosome formation compared to 1× 601 DNA, were added exclusively to the chromatin assembly reaction. The mixture was dialyzed overnight at 4 °C, while the salt concentration was gradually changed using a peristaltic pump from high-salt to low-salt buffer (10 mM Tris, 1 mM EDTA, 10 mM NaCl). After dialysis, any aggregates were removed by centrifugation (21,000×g, 10 min, 4 °C). The dialyzed sample concentrations were determined by UV using a NanoDrop spectrophotometer (ThermoFisher), and nucleosome assembly efficiency was evaluated by native PAGE performed on ice.
The integrity of the chromatin fibers was assessed by ScaI digestion of the fibers, resulting in individual mononucleosomes (Supplementary Fig. 7b). 80 ng of chromatin fibers were incubated with ScaI-HF (NEB, R3122S) at 37 °C for 3 h in the dark. The digested products were then analyzed by native PAGE to confirm nucleosome quality and ensure full nucleosome occupancy as inferred from the absence of free DNA (Supplementary Fig. 7b). Once octamer saturation was verified, magnesium precipitation using 6 mM MgCl2 was used to further purify the chromatin fibers (Supplementary Fig. 7c).
The imaging buffer IB3 contained 100 mM KCl and 20 mM HEPES (with Na+ as the counterion), yielding 120 mM monovalent cations. Under these conditions, chromatin adopts a partially compacted state, as revealed by single-molecule FRET37,80.
Slide and cover glass cleaning and flow-channel assembly
Coverslips (24 × 40 mm2, 1.5 mm thick) and glass slides (76 × 26 mm2), containing holes to access flow channels after assembly, underwent multiple cleaning steps: First, they were sonicated for 20 min in a 10% (w/v) Alconox detergent solution, followed by successive sonication in ethanol and acetone. Between each solvent step, Milli-Q water sonication ensured the removal of the previous solvent. The slides were then cleaned for 2–4 h in a Piranha solution (3 M sulfuric acid and 60% hydrogen peroxide at a 3:1 ratio) and thoroughly rinsed in Milli-Q water. Next, they were sonicated in acetone for 10 min, then silanized for 10 min in 2% (3-aminopropyl)triethoxysilane prepared in acetone. Afterward, they were quenched in Milli-Q water and dried under nitrogen gas.
To assemble flow channels, we cut channel spacers from a 0.12 mm thickness double-sided adhesive sheet (GraceBio, 620001) using previously reported designs37,115,117. Each set of prepared coverslips and slides was assembled into a “sandwich” configuration, with the printed adhesive sheet sandwiched between the silanized coverslip and glass slide. The assembled devices were subsequently vacuum-sealed and stored at −20 °C until use.
Microfluidic channel preparation
To prepare measurement chambers, assembled coverslip-microscopy slide devices were warmed to room temperature. 10-µL pipette tips were glued into the prepared holes using epoxy glue. The individual flow-channels were then passivated by PEGylation: A PEG solution was prepared by dissolving 1 mg of biotinylated PEG-bis-succinimidyl valerate (SVA) (5 kDa) and 20 mg of methoxy PEG-SVA (5 kDa) in 200 µL of 100 mM sodium tetraborate buffer (pH 8.5). This solution was injected into the pre-formed channels and incubated for 2–3 h at room temperature before use in single-molecule experiments.
smTIRF microscopy setup
For single-molecule colocalization imaging, we used a Nikon Ti-E inverted microscope equipped with a Nikon TIRF illuminator and a ×100 oil immersion objective (NA 1.49). For illumination, two lasers were used: a Coherent OBIS 640LX (640 nm, 40 mW) to image AF647-labeled DNA and an OBIS 532LS (532 nm, 50 mW) to image JF549-labeled Halo-Sox TFs. These lasers were coupled into the microscope via an optical fiber, and an acousto-optic tunable filter was used for wavelength selection. Laser power densities at the objective were 44 W/cm2 for 532 nm and 11 W/cm2 for 640 nm. Nikon’s Perfect Focus system was used to compensate for axial fluctuations in the field of view (FOV). Emissions were detected over an 800 × 800 px2 FOV by a Photometrics Prime 95B sCMOS camera, with a physical pixel size of 11 × 11 μm2. Some movies acquired on bare DNA substrates were recorded using an ANDOR iXon EMCCD camera with a 256 × 256 px² FOV and 16 × 16 μm2 pixel size.
In vitro single-molecule colocalization measurements
Prior to imaging, all buffers were degassed. An oxygen-scavenging system was prepared by dissolving 1 mg catalase in 100 µL of 50 mM phosphate buffer (pH 7.0), and 10 mg glucose oxidase was dissolved in a mixture of 40 µL catalase solution and 60 µL of T50 buffer (Gloxy). This mixture was centrifuged at 21,000×g for 3 min, and the supernatant was taken for use. All DNA and protein samples used for in vitro SMT were spun down at 21,000×g for 10 min at 4 °C to clear aggregates and were transferred to DNA low-binding (Eppendorf, 0030108051) or protein low-binding tubes (Eppendorf, 0030108116), respectively.
Prior to imaging, each PEGylated flow-channel was rinsed with ultra-pure water (ROMIL, H950L) and equilibrated with T50 buffer (10 mM Tris, 50 mM NaCl, pH 8). A peristaltic pump was used for the injection into the microfluidic channel. 30 μL of 0.2 mg/mL neutravidin in the T50 buffer was injected to facilitate DNA-substrate tethering and then washed with the T50 buffer. BSA (5 mg/mL) was incubated for 10 min, followed by washing with T50. Finally, the channels were equilibrated with imaging buffer (IB3), which included 20 mM HEPES, 20 mM Tris, 1 mM DTT, 3 mM EDTA, 100 mM KCl, 2 mg/mL BSA, 0.02% Tween-20, 10% glycerol, 3.2% glucose, Gloxy, and 2 mM Trolox at pH 7.5.
For immobilization, 10–50 pM of AF647-labeled, biotin-conjugated DNA constructs, mononucleosomes, or chromatin fibers in IB3 buffer were injected into the channel, with the degree of immobilization measured by single-molecule imaging. Excess DNA constructs, mononucleosomes, or chromatin fibers were washed away with 200 μL of IB3 buffer. Finally, 2 nM of JF549-labeled Halo-Sox2 or Halo-Sox2a was flowed at 0.5 μL/min.
After injection, Sox TF interaction dynamics with immobilized DNA, nucleosome, and chromatin substrates were recorded using a 10-min acquisition at a 200 ms frame interval. AF647-labeled DNA was excited and imaged once every 200 frames for drift correction and DNA localization, whereas JF549-labeled Halo-Sox TFs were recorded continuously (100 ms illumination + 100 ms dark time per frame).
ChIP-seq
Under the mESC culture conditions described above, SES4.1, SES5.1, SES7.1, and SES9.1 cells (Supplementary Table 6) were incubated for 20 h in the presence of 300, 70, 1000, and 1000 ng/mL dox, respectively, to induce expression of Halo-conjugated Sox2, Sox2a, Sox17, and Sox17b. The Halo-Sox TFs were then labeled with 50 nM Halo-SiR dye. Two biological replicates were processed for Sox2 and Sox2a, while Sox17 and Sox17b were profiled using one biological replicate each.
Fixation was performed in a PBS-based buffer, first with 2 mM of disuccinimidyl glutarate (ThermoFisher, 20593) for 50 min at room temperature, followed by 1% formaldehyde (ThermoFisher, 28908) for 10 min. Fixation was quenched by incubating the samples in 200 mM Tris–HCl (pH 8.0) for 10 min. Fixed cells were pelleted at 600×g for 4 min at 4 °C and resuspended in 1% ESC-qualified fetal bovine serum (Gibco, 16141-079). Cells were sorted by FACS at 4 °C, using the intensity gate adjusted to yield equivalent mean Halo-SiR intensities between SES4.1 and SES5.1 FACS samples (Fig. 6a, Supplementary Table 13), and similarly between SES7.1 and SES9.1 FACS samples.
Sorted cells were resuspended in LB1 buffer (50 mM HEPES–KOH pH 7.4, 140 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, 10% Glycerol, 0.5% NP-40, 0.25% Triton X-100, supplemented with Protease inhibitor cocktail (Sigma, P8340-1ML) at 1:100 dilution), incubated on ice for 10 min, and centrifuged at 1700×g for 5 min at 4 °C. This step was repeated once more. Pellets were then resuspended in LB2 buffer (10 mM Tris–HCl pH 8.0, 200 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, supplemented with Protease inhibitor cocktail (Sigma, P8340-1ML) at 1:100 dilution), incubated for 10 min on ice, and centrifuged again. Pellets were washed twice with SDS shearing buffer (10 mM Tris–HCl pH 8.0, 1 mM EDTA, 0.15% SDS, supplemented with Protease inhibitor cocktail (Sigma, P8340-1ML) at 1:100 dilution)), taking care not to disturb them, and finally resuspended in SDS shearing buffer on ice. Chromatin was sheared using a Covaris E220 (200 cycles, 5% duty cycle, 140 W, 20 min). Lysates were clarified at 10,000×g for 5 min at 4 °C, and supernatants were collected.
ChIP and DNA purification were performed using the ChIP-IT High Sensitivity kit (Active Motif, 53040) according to the manufacturer’s instructions. Input chromatin amounts were 20 μg for Sox2 and Sox2a, and 30 μg for Sox17 and Sox17b. Drosophila spike-in chromatin (Active Motif, 53083) was added at 1 ng per 1 μg of target chromatin. Spike-in antibody (anti-H2Av; Active Motif, 61686) was added at 0.5 μg per 10 ng spike-in chromatin and processed together with 40 μL Halo-Trap agarose beads (ChromoTek, ota-20) per reaction. ChIP reactions were incubated overnight at 4 °C on a rotating platform (30 rpm). Sequencing libraries were prepared using the NEBNext Ultra II DNA Library Prep Kit (NEB, E7645), size-selected using AMPure XP beads (Beckman Coulter, A63880), and sequenced on an Illumina NextSeq 500 using 75-nucleotide read-length paired-end sequencing.
ATAC-seq
SES4.1 and SES5.1 cells were treated with dox under the same conditions used for ChIP-seq and labeled with 50 nM Halo-SiR647 dye. Cells were FACS-sorted at 4 °C using the same intensity gates as in the ChIP-seq workflow (Supplementary Table 13).
ATAC-seq was performed following established methods118. Approximately 50,000 cells were lysed in 50 μL of ATAC–lysis buffer (10 mM Tris–HCl, 10 mM NaCl, 3 mM MgCl2, 0.1% NP-40, pH 7.4), and nuclei were pelleted at 800×g for 5 min. The transposition reaction was carried out by resuspending nuclei in 50 μL of TAPS-DMF buffer (10 mM TAPS–NaOH, 5 mM MgCl2, 10% DMF) supplemented with 0.5 μM in-house-produced Tn5 transposase119 and incubating them at 37 °C for 30 min. DNA was eluted in 10 μL of nuclease-free water, using the DNA Clean & Concentrator kit (Zymo Research, D4004) with a 5:1 binding buffer-to-sample ratio.
For library preparation, transposed DNA was PCR-amplified using NEBNext High-Fidelity 2× PCR Master Mix (NEB, M0541L), 0.5 μM Ad1.1 universal primer, 0.5 μM Ad2.X indexing primer, and 0.6× SYBR Green I (ThermoFisher, S7585) in a 65 μL reaction. Thermocycling proceeded as follows: 72 °C for 5 min; 98 °C for 30 s; then 5 cycles of 98 °C for 10 s, 63 °C for 30 s, and 72 °C for 60 s.
A 10 μL aliquot was analyzed by qPCR to determine the optimal total number of amplification cycles, using the fluorescence signal at one-third saturation to avoid over-amplification. The remaining DNA was amplified for the required number of cycles, purified (Zymo, D4004), size-selected with AMPure XP beads (Beckman Coulter, A63880), and sequenced on an Illumina NextSeq 500 using 75-nucleotide read-length paired-end sequencing.
Electrical charge distribution generation
Along a protein sequence, the average charge of the amino acids within a sliding five-amino-acid window was calculated. The residues Aspartic acid (D) and Glutamic acid (E) were assigned a charge of −1, Histidine (H) a charge of +0.5, and Lysine (K) and Arginine (R) a charge of +1. These calculations were performed using EMBOSS Pepinfo120.
In vivo SMT imaging data processing
Nuclei were detected using the StarDist plugin in Fiji113. Based on these nuclear ROIs, single-cell movies in the JF549 channel were cropped. Using TrackIt55, individual Sox TF molecules within the nuclear ROIs were detected using the detection thresholds listed in Supplementary Table 14. Detected spots were linked across consecutive frames using the nearest-neighbor algorithm to construct the single-molecule trajectories. The tracking-loss probability (due to molecules moving laterally out of the imaging plane or out of focus) was set to 0.005. Considering the input parameters (Supplementary Table 14) and the loss probability, the tracking radii were automatically computed for different dark time conditions (Supplementary Table 15). For the continuous movies with 100 Hz of frame rate, the tracking radius for all Sox TFs was set to 8.1312 pixels with respect to 894 nm of the maximum jump distance.
GRID analysis
GRID is restricted to superimposed exponential reactions with positive amplitudes and converts the inverse Laplace transformation into a minimization problem, exploring a discrete set of potential outcomes56. In the off-rate domain, 200 possible dissociation rates (k) were logarithmically spaced from log(k) = –3 to log(k) = 0.85. Bulk GRID analysis yielded a single photobleaching rate for the pooled trajectories of all the Sox TFs. This photobleaching rate (0.028 s−1) was then applied uniformly to all Sox TF constructs, regardless of the specific TF. The regularization weight parameter was set to 0.1, the default value in the GRID implementation.
For each Sox TF, 100 resampling iterations were performed. In each iteration, 80% of single cells were randomly selected using a custom MATLAB code, and a GRID spectrum was computed. GRID-determined dissociation rates with amplitudes below 0.1% were excluded. The remaining rates from all 100 iterations were combined into a relative frequency histogram using 100 equally spaced bins (Fig. 1h). To compute the fractional occupancy spectrum (\({F}_{i}\)) (Supplementary Fig. 2e), each GRID-determined state’s amplitude (\({A}_{i}\)) was weighted by its residence time (\({{{\rm{\tau }}}}_{i}\)), and the resulting products were normalized by their total sum:
State array analysis
The State array method61, for which the code is openly accessible, was used with the following settings: likelihood_type = RBME, pixel_size_μm = 0.11, frame_interval = 0.01, focal_depth = 0. The diffusion coefficient grid was defined on a logarithmic scale from 10−3 to 102 µm2 s−1 (150 bins) while all the other parameters were kept as default in constant.py. The state occupations marginalized on the diffusion coefficient for individual cells were computed in Fig. 2c and per TF, the single-cell state occupations were averaged in Fig. 2d.
Spot-On analysis
The MATLAB version of Spot-On59 was used in this analysis. To generate the Spot-on data from single-cell analysis, the entire trajectories were used. In the 3-state model, the range of each diffusion coefficient was set to be 1 × 10−5–5 × 10−2 µm2 s−1 for the bound state, 5 × 10−2–5 × 100 µm2 s−1 for the slow diffusing state, and 1 × 100–5 × 101 µm2 s−1 for the fast diffusing state. For the jump length distribution parameters, the number of time points and max jump distance were set as 8 and 2 µm. To correct localization and defocalization errors, for z-correction, an axial resolution of 0.7 µm was applied, and the localization error fitted from data was allowed. The CDFs of jump lengths were fitted globally across all the time points in 3 iterations.
Pseudo on-rate analysis
The pseudo on-rate (kpn) was calculated according to Eq. ( 2)
where \({n}_{{{\rm{f}}}}\) is the number of considered frames, \({{\rm{TI}}}\) is a time interval between movie frames, \({C}_{{{\rm{Sox}}},i}\) is the concentration of target TFs per frame (\(i\)), and \({N}_{i}^{{\tau }_{{{\rm{c}}}}}\) is the number of selected molecules lasting the given bound time (\({\tau }_{{{\rm{c}}}}\)), which appear first at the frame \(i\).
For nonspecific kpn, molecules detected in 100 Hz continuous movies were used and \({\tau }_{{{\rm{c}}}}\) < 1 s was applied. During trajectory reconstruction in TrackIt, the maximum allowed jump distance was constrained by a diffusion coefficient of 0.08 µm2 s−1, yielding a tracking radius of 0.5243 µm. The total number of bound or diffusing trajectories was used as a proxy for the relative nuclear concentration of each Sox TF.
For specific kpn, molecules detected in 120 ms time interval movies were considered and \({\tau }_{{{\rm{c}}}}\) was set to > 1 s. Here, the nuclear concentration of Sox TFs was estimated from the mean Halo-JF549 nuclear intensity after background subtraction. Photobleaching effects were accounted for by measuring the fluorescent intensity per frame over time.
Following the initial phase during which most molecules are detected, a series of consecutive empty frames—where no molecules are detected—could artificially lower the average molecule count, leading to an underestimation of kpn. Such empty frames often occur due to photobleaching or during long recordings with prolonged periods without signal. To correct for this bias, we defined an idle time (5 s for nonspecific kpn and 10 s for specific kpn). When an idle period is detected, \({n}_{{{\rm{f}}}}\) is set to the first frame of that period, and the summation for the average is terminated. This approach excludes empty frames from the calculation, ensuring a more accurate measurement of the kpn. Custom MATLAB scripts were used to perform this calculation.
For statistical analysis in OriginPro 2022, the single-cell distributions of kpn across Sox TFs were first assessed for normality using the Shapiro–Wilk test, with normality rejected at a significance level of 0.05. Consequently, a Kruskal–Wallis ANOVA was performed, followed by Dunn’s test for multiple comparisons.
In vitro single-molecule colocalization data processing
ND2 raw images were converted to TIFF format, followed by background subtraction using a rolling ball filter with a radius of 50 pixels. This process was done in Fiji113. Further data extraction was performed using custom MATLAB codes. The analysis consisted of six main steps. First, the loci of anchored DNA/MN/CF on the FOV were identified. Second, colocalization of Sox TFs with immobilized DNA/MN/CF was determined using 2D-Gaussian fitting of detected single-molecule emitters to determine the shape of the point spread function (PSF) as well as the exact position of each detection. Third, we extracted fluorescence time-traces of JF549-labeled Sox TFs from each DNA/MN/CF location. Fourth, each time-trace was segmented based on the information from the JF549 emission, i.e., sections of the trace were excluded from the analysis: Detections where the PSF exceeded 300 nm at its full width at half maximum were excluded. Moreover, JF549 detections that did not colocalize with DNA/MN/CF positions (as determined by a 300 nm threshold) were excluded from the traces. Lastly, detections that showed multiple stepwise changes in fluorescence intensity or exceeded a threshold value were excluded from the trace, as they arose from multiple TFs or aggregates. If more than 80% of traces from a movie were excluded during trace selection, the entire movie was discarded. Fifth, traces were fitted using a stepfit algorithm121. Finally, bound states were identified using an intensity threshold derived from the fitted trace, and the corresponding bound, unbound, and search times were determined.
Statistical analysis for residence times, on-rates and specific on-rate
The bound, unbound, and search times from each movie (Fig. 3e) were used to generate the corresponding 1-CDF curves (Fig. 3g, Supplementary Fig. 5a, f). First, a cumulative histogram was constructed with a bin size of 0.2 s—matching the movie frame interval. Then, the histogram was normalized to range from 0 to 1.
To estimate the residence times of Sox TFs, we applied multi-exponential decay fits to the 1-CDF curves of bound times. For naked DNA substrates, a biexponential model (n = 2 in Eq. (3) was used, while for nucleosomes, a triexponential model (n = 3 in Eq. (3) was employed.
In these models, the inverse of each decay rate, \({k}_{i}\), corresponds to a residence time, \({{{\rm{\tau }}}}_{i}\).
The on-rate (kon) and specific on-rate (ks,on) were determined by fitting the 1-CDF curves of unbound and search times, respectively, with a monoexponential model (n = 1 in Eq. (3); here, kon and ks,on were calculated by dividing the decay rate k1 by the Sox TF concentration (2 nM). All fitting procedures were carried out using the Levenberg–Marquardt algorithm in OriginPro 2022.
To generate the fractional occupancies (\({F}_{i}\)) (Fig. 4f,h, Supplementary Figs. 5j, 7f,h), each bound fraction (\({A}_{i}\)) was weighted by its residence time (\({{{\rm{\tau }}}}_{i}\)), and the resulting products were normalized by their total sum:
DNA coiling model
DNA, as a polymer, will coil when its length exceeds its persistence length (\({l}_{{{\rm{p}}}}\)), which is ~45 nm67,68,69—roughly equivalent to 132 bp. For naked DNA of length \(2L\), the gyration radius (\({R}_{{{\rm{g}}}}\)) is expressed as follows:
The on-rate (kon) may depend on the overall size of the DNA encountered by a TF. Because DNA coils, kon is proportional to the DNA’s gyration radius (Rg) multiplied by the nanoscopic association rate (ka), which is defined as the rate at which a TF binds to a unit-length DNA segment (2\({L}_{0}\)).
Nonlinear fitting of the kon curve (Supplementary Fig. 5c) using Eq. (6) was performed with the Levenberg–Marquardt algorithm, employing the standard deviation of each data point as the weighting factor. The standard errors of the fitting parameters ka and \({L}_{0}\) were calculated with OriginPro 2022. In this fitting, the persistence length (\({l}_{{{\rm{p}}}}\)) was fixed at 132 bp. \({L}_{0}\) was assumed to be the same among Sox TFs, which was obtained as 5.8 bp.
1D target search model
A 1D search model has been previously introduced by Berg et al.12 and Hammar et al.17. It describes the behavior of a TF on bare DNA containing a single binding site at its center. We can use this model to describe the Sox TF target search for a motif located at the center of a stretch of naked DNA (Fig. 3h). The model involves four parameters: the nanoscopic association rate (ka), nonspecific residence time (\({\tau }_{{{\rm{d}}}}\)), the 1D diffusion coefficient (D1D), and the target recognition rate (kr). A TF binds to any sites on DNA at ka, slides with D1D and then recognizes the target with kr upon encounter. During this process, the TF resides on DNA nonspecifically for \({\tau }_{{{\rm{d}}}}\).
In this model, the specific on-rate (ks,on) was defined as the association rate to the target site on bare DNA, via 1D diffusion as well as 3D diffusion. Consequently, ks,on depends on L, the length of flanking DNA as below:
Although kr cannot be measured directly at present, it can be indirectly estimated (Fig. 3l) by determining the specific on-rate (ks,on) in a defined system under various conditions (Fig. 3i).
To fit the measured ks,on curve (Fig. 3i) with the 1D target search model, we considered the sliding length (\({s}_{{{\rm{L}}}}\)), which was previously introduced as below:
Furthermore, we defined a target recognition constant (KR) as below, since the target recognition process would occur before it dissociates from the site:
Using Eqs. (8) and (9), ks,on in Eq. (7) becomes as a function of L with three parameters of ka, KR, and sL, as below:
Accordingly, the recognition probability \({p}_{{{\rm{bind}}}}\) is expressed as below:
The nonlinear fitting on the ks,on curve (Fig. 3i) with Eq. (10) was conducted based on the Levenberg–Marquardt algorithm using the SD of each data point as the weighting factor. The standard errors of fitting parameters or derived parameters of kr and D1D (from Eqs. (8) and (9) considering \({\tau }_{{{\rm{d}}}}\) = 10 ms71) were also calculated in OriginPro 2022. In the fitting, ka was fixed to measured values of ka,Sox2 = 2.14 M−1 s−1 bp−1, ka,Sox2a = 1.93 × 105 M−1 s−1 bp−1, and ka,Sox17 = 1.85 × 105 M−1 s−1 bp−1 as listed in Supplementary Table 4.
ChIP-seq analysis
ChIP-sequencing libraries were mapped (BAM format) to mm10 version of Mus musculus genome or BDGP6 version of Drosophila melanogaster genome, using STAR122 with parameters --alignMatesGapMax 2000 --alignIntronMax 1 --alignEndsType EndtoEnd --outFilterMultimapNmax 1. Duplicated reads and reads not mapping to chromosome 1–19 and X or Y, were removed using SAMTools123. For the Spike-in normalization, normalization factors were calculated to equalize the total read counts across all Drosophila BAM files. These normalization factors were then applied to downsample the mouse-genome alignment files using SAMTools123. Besides, between replicates, the ratio of their sequencing depths was used to normalize alignment files mapped on the mouse genome using SAMTools123. For score comparisons across samples, the replicate BAM files were merged using SAMTools123. Scores were determined from these merged BAM files using the bamCoverage function from DeepTools v3.3.0124 with setting -- normalizeUsing RPKM.
Peak calling was performed on the normalized BAM files using MACS2 v 3.0.0a7 with settings -f BAMPE -g mm, and regions listed in the ENCODE mm10 blacklist were excluded. ChIP peaks between replicates were consolidated using a specific combining protocol (https://ro-che.info/articles/2018-07-11-chip-seq-consensus), employing lower = 1 and rangesOnly = T in the slice() function from the GenomicRanges library.
These merged peak files were used to generate Sox-enriched regions following the same procedure. Starting from the replicate-merge BED files, CORs were defined via the same protocol, but with lower = 2 and rangesOnly = T in slice(). To quantify enrichment across CORs, two approaches were used: (i) multiBigwigSummary in DeepTools124 or (ii) bigWigToBedGraph from the UCSC genome toolkit. Log2-fold changes in Sox2 scores relative to Sox2a scores were computed for each COR, and regions were classified as Sox2-enriched or Sox2a-enriched based on the sign of the log2-fold values. For exclusive region analysis, each replicate-merged BED file (Sox2 or Sox2a) was processed with the subtract function in BEDTools125 to remove CORs. The resulting intervals were then merged again using the same ChIP peak consolidation protocol. Genome tracks were viewed using Integrated Genome Browser126.
Next, these replicate-merged BED files were input into HOMER2’s findMotifsGenome.pl127 to discover motifs. The search was restricted to 200 bp regions around the peaks, on bigWig files that were generated from merged BAM files spanning the replicates.
ATAC-seq analysis
ATAC-Seq libraries were mapped and filtered as described for ChIP-Seq. Accessible peaks were then called, and scores were computed as described above for ChIP-Seq.
Statistical analysis
Most results are presented as means ± SD unless stated otherwise. Two-group comparisons were conducted using two-tailed, heteroscedastic t-tests. For multiple comparisons, distributions were first evaluated for normality using the Shapiro–Wilk test; When normality was rejected, a nonparametric, one-way Kruskal–Wallis ANOVA was performed, followed by Dunn’s test for pairwise comparisons. When normality was confirmed, one-way parametric ANOVA was used. A p-value ≤ 0.05 was considered statistically significant. All statistical analyses were performed using OriginPro 2022. Statistical significance is indicated in the figures using standard asterisk ranges, while all exact p-values are fully reported in the Source Data.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All single-molecule data are available from Zenodo: https://doi.org/10.5281/zenodo.14948338 (Fig. 1, Supplementary Fig. 2); https://doi.org/10.5281/zenodo.14948340 (Fig. 2, Supplementary Fig. 3); https://doi.org/10.5281/zenodo.14948308, https://doi.org/10.5281/zenodo.16954952, https://doi.org/10.5281/zenodo.16954984 (Fig. 3, Supplementary Fig. 5); https://doi.org/10.5281/zenodo.14948298, https://doi.org/10.5281/zenodo.16955046 (Figs. 4, 5, Supplementary Fig. 6-7). Biochemical gel assay data are available in Zenodo: https://doi.org/10.5281/zenodo.16955178. All sequencing data were deposited at NCBI Gene Expression Omnibus (GEO), accession numbers GSE290873 and GSE290874: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE290873, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE290874. Source Data are provided with this paper. Additional imaging data, detailed protein purification protocols, and plasmid maps of expression vectors are available upon request. Further information and requests for resources and reagents should be directed to and will be fulfilled by Beat Fierz (beat.fierz@epfl.ch) or David Suter (david.suter@epfl.ch). Source data are provided with this paper.
Code availability
Analysis scripts are available in Zenodo: https://doi.org/10.5281/zenodo.16955257 (related to In vivo assay); https://doi.org/10.5281/zenodo.16954687 (related to In vitro assay).
References
Biggin, M. D. Animal transcription networks as highly connected, quantitative continua. Dev. Cell 21, 611–626 (2011).
Strebinger, D. et al. Endogenous fluctuations of OCT4 and SOX2 bias pluripotent cell fate decisions. Mol. Syst. Biol. 15, e9002 (2019).
Placzek, S., Vanzan, L., Deluz, C. & Suter, D. M. Orchestration of pluripotent stem cell genome reactivation during mitotic exit. Cell Rep. 44, 115486 (2025).
Lodato, M. A. et al. SOX2 co-occupies distal enhancer elements with distinct POU factors in ESCs and NPCs to specify cell state. PLoS Genet. 9, e1003288 (2013).
Maresca, M. et al. Pioneer activity distinguishes activating from non-activating SOX2 binding sites. EMBO J. 42, e113150 (2023).
Teves, S. S. et al. A dynamic mode of mitotic bookmarking by transcription factors. eLife 5, e22280 (2016).
Hettich, J. & Gebhardt, J. C. M. Transcription factor target site search and gene regulation in a background of unspecific binding sites. J. Theor. Biol. 454, 91–101 (2018).
Lin, S.-Y. & Riggs, A. D. Lac repressor binding to DNA not containing the Lac operator and to synthetic poly dAT. Nature 228, 1184–1186 (1970).
Mirny, L. et al. How a protein searches for its site on DNA: the mechanism of facilitated diffusion. J. Phys. A: Math. Theor. 42, 434013 (2009).
Mueller, F., Stasevich, T. J., Mazza, D. & McNally, J. G. Quantifying transcription factor kinetics: at work or at play? Crit. Rev. Biochem. Mol. Biol. https://www.tandfonline.com/doi/abs/10.3109/10409238.2013.833891 (2013).
Wang, X. et al. Negatively charged, intrinsically disordered regions can accelerate target search by DNA-binding proteins. Nucleic Acids Res. 51, 4701–4712 (2023).
Berg, O. G., Winter, R. B. & Von Hippel, P. H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 20, 6929–6948 (1981).
Richter, P. H. & Eigen, M. Diffusion controlled reaction rates in spheroidal geometry: application to repressor-operator association and membrane bound enzymes. Biophys. Chem. 2, 255–263 (1974).
Riggs, A. D., Bourgeois, S. & Cohn, M. The lac represser–operator interaction. J. Mol. Biol. 53, 401–417 (1970).
Slutsky, M. & Mirny, L. A. Kinetics of protein–DNA interaction: facilitated target location in sequence-dependent potential. Biophys. J. 87, 4021–4035 (2004).
Halford, S. E. An end to 40 years of mistakes in DNA–protein association kinetics?. Biochem. Soc. Trans. 37, 343–348 (2009).
Hammar, P. et al. The lac repressor displays facilitated diffusion in living cells. Science 336, 1595–1598 (2012).
Hsieh, W. T., Whitson, P. A., Matthews, K. S. & Wells, R. D. Influence of sequence and distance between two operators on interaction with the lac repressor. J. Biol. Chem. 262, 14583–14591 (1987).
Marklund, E. et al. DNA surface exploration and operator bypassing during target search. Nature 583, 858–861 (2020).
Ruusala, T. & Crothers, D. M. Sliding and intermolecular transfer of the lac repressor: kinetic perturbation of a reaction intermediate by a distant DNA sequence. Proc. Natl. Acad. Sci. USA 89, 4903–4907 (1992).
Joseph, S. R. et al. Competition between histone and transcription factor binding regulates the onset of transcription in zebrafish embryos. eLife 6, e23326 (2017).
Kadonaga, J. T. The transformation of the DNA template in RNA polymerase II transcription: a historical perspective. Nat. Struct. Mol. Biol. 26, 766–770 (2019).
Morgunova, E. & Taipale, J. Structural insights into the interaction between transcription factors and the nucleosome. Curr. Opin. Struct. Biol. 71, 171–179 (2021).
Zhu, F. et al. The interaction landscape between transcription factors and the nucleosome. Nature 562, 76–81 (2018).
Brodsky, S. et al. Intrinsically disordered regions direct transcription factor in vivo binding specificity. Mol. Cell 79, 459–471.e4 (2020).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855.e16 (2018).
Dyson, H. J. & Wright, P. E. Intrinsically unstructured proteins and their functions. Nat. Rev. Mol. Cell Biol. 6, 197–208 (2005).
Már, M., Nitsenko, K. & Heidarsson, P. O. Multifunctional intrinsically disordered regions in transcription factors. Chem.—Eur. J. 29, e202203369 (2023).
Soto, L. F. et al. Compendium of human transcription factor effector domains. Mol. Cell 82, 514–526 (2022).
Chen, Y. et al. Mechanisms governing target search and binding dynamics of hypoxia-inducible factors. Elife 11, e75064 (2022).
Iwafuchi, M. et al. Gene network transitions in embryos depend upon interactions between a pioneer transcription factor and core histones. Nat. Genet. 52, 418–427 (2020).
Johnson, T. A., Paakinaho, V., Kim, S., Hager, G. L. & Presman, D. M. Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms. Nat. Commun. 12, 1987 (2021).
Wang, Z. et al. Mesoscale chromatin confinement facilitates target search of pioneer transcription factors in live cells. Nat. Struct. Mol. Biol. 1–12 https://doi.org/10.1038/s41594-024-01385-5 (2024).
Hansen, A. S., Amitai, A., Cattoglio, C., Tjian, R. & Darzacq, X. Guided nuclear exploration increases CTCF target search efficiency. Nat. Chem. Biol. 16, 257–266 (2020).
Cirillo, L. A. et al. Opening of compacted chromatin by early developmental transcription factors HNF3 (FoxA) and GATA-4. Mol. Cell 9, 279–289 (2002).
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151, 994–1004 (2012).
Mivelaz, M. et al. Chromatin fiber invasion and nucleosome displacement by the Rap1 transcription factor. Mol. Cell 77, 488–500.e9 (2020).
Zaret, K. S. & Carroll, J. S. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 25, 2227–2241 (2011).
Dodonova, S. O., Zhu, F., Dienemann, C., Taipale, J. & Cramer, P. Nucleosome-bound SOX2 and SOX11 structures elucidate pioneer factor function. Nature 580, 669–672 (2020).
Raccaud, M. et al. Mitotic chromosome binding predicts transcription factor properties in interphase. Nat. Commun. 10, 487 (2019).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
Avilion, A. A. et al. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 17, 126–140 (2003).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Michael, A. K. et al. Mechanisms of OCT4-SOX2 motif readout on nucleosomes. Science 368, 1460–1465 (2020).
Ambrosetti, D. C., Basilico, C. & Dailey, L. Synergistic activation of the fibroblast growth factor 4 enhancer by Sox2 and Oct-3 depends on protein–protein interactions facilitated by a specific spatial arrangement of factor binding sites. Mol. Cell. Biol. 17, 6321–6329 (1997).
Shimoda, M. et al. Sox17 plays a substantial role in late-stage differentiation of the extraembryonic endoderm in vitro. J. Cell Sci. 120, 3859–3869 (2007).
Niakan, K. K. et al. Sox17 promotes differentiation in mouse embryonic stem cells by directly regulating extraembryonic gene expression and indirectly antagonizing self-renewal. Genes Dev. 24, 312–326 (2010).
Aksoy, I. et al. Oct4 switches partnering from Sox2 to Sox17 to reinterpret the enhancer code and specify endoderm. EMBO J. 32, 938–953 (2013).
Jauch, R. et al. Conversion of Sox17 into a pluripotency reprogramming factor by reengineering its association with Oct4 on DNA. Stem Cells 29, 940–951 (2011).
Hamilton, D. J., Hein, A. E., Holmes, Z. E., Wuttke, D. S. & Batey, R. T. The DNA-binding High Mobility Group Box domain of Sox family proteins directly interacts with RNA in vitro. Biochemistry https://doi.org/10.1021/acs.biochem.2c00218 (2022).
Gebhardt, J. C. M. et al. Single-molecule imaging of transcription factor binding to DNA in live mammalian cells. Nat. Methods 10, 421–426 (2013).
Masui, S. et al. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat. Cell Biol. 9, 625–635 (2007).
Grimm, J. B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat. Methods 12, 244–250 (2015).
Lukinavičius, G. et al. A near-infrared fluorophore for live-cell super-resolution microscopy of cellular proteins. Nat. Chem. 5, 132–139 (2013).
Kuhn, T., Hettich, J., Davtyan, R. & Gebhardt, J. C. M. Single molecule tracking and analysis framework including theory-predicted parameter settings. Sci. Rep. 11, 9465 (2021).
Reisser, M. et al. Inferring quantity and qualities of superimposed reaction rates from single molecule survival time distributions. Sci. Rep. 10, 1758 (2020).
Mahadevan, J. et al. Dynamics of endogenous PARP1 and PARP2 during DNA damage revealed by live-cell single-molecule imaging. iScience 26, 105779 (2023).
Engl, W. et al. Single-molecule imaging of SWI/SNF chromatin remodelers reveals bromodomain-mediated and cancer-mutants-specific landscape of multi-modal DNA-binding dynamics. Nat. Commun. 15, 7646 (2024).
Hansen, A. S. et al. Robust model-based analysis of single-particle tracking experiments with Spot-On. Elife 7, e33125 (2018).
Mazza, D., Abernathy, A., Golob, N., Morisaki, T. & McNally, J. G. A benchmark for chromatin binding measurements in live cells. Nucleic Acids Res. 40, e119 (2012).
Heckert, A., Dahal, L., Tjian, R. & Darzacq, X. Recovering mixtures of fast-diffusing states from short single-particle trajectories. eLife 11, e70169 (2022).
Liu, Z. et al. 3D imaging of Sox2 enhancer clusters in embryonic stem cells. eLife 3, e04236 (2014).
Mazzocca, M., Narducci, D. N., Grosse-Holz, S., Matthias, J. & Hansen, A. S. Chromatin dynamics are highly subdiffusive across seven orders of magnitude. Preprint at https://doi.org/10.1101/2025.05.10.653248 (2025).
Wachsmuth, M., Knoch, T. A. & Rippe, K. Dynamic properties of independent chromatin domains measured by correlation spectroscopy in living cells. Epigenet. Chromatin 9, 57 (2016).
Shinkai, S., Nozaki, T., Maeshima, K. & Togashi, Y. Dynamic nucleosome movement provides structural information of topological chromatin domains in living human cells. PLoS Comput. Biol. 12, e1005136 (2016).
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning1. J. Mol. Biol. 276, 19–42 (1998).
Bednar, J. et al. Determination of DNA persistence length by cryo-electron microscopy. separation of the static and dynamic contributions to the apparent persistence length of DNA. J. Mol. Biol. 254, 579–594 (1995).
Hagerman, P. J. Flexibility of DNA. Annu. Rev. Biophys. 17, 265–286 (1988).
Gross, P. et al. Quantifying how DNA stretches, melts and changes twist under tension. Nat. Phys. 7, 731–736 (2011).
Li, S., Zheng, E. B., Zhao, L. & Liu, S. Nonreciprocal and conditional cooperativity directs the pioneer activity of pluripotency transcription factors. Cell Rep. 28, 2689–2703.e4 (2019).
White, M. D. et al. Long-lived binding of Sox2 to DNA predicts cell fate in the four-cell mouse embryo. Cell 165, 75–87 (2016).
Elf, J., Li, G.-W. & Xie, X. S. Probing transcription factor dynamics at the single-molecule level in a living cell. Science 316, 1191–1194 (2007).
Normanno, D. et al. Probing the target search of DNA-binding proteins in mammalian cells using TetR as model searcher. Nat. Commun. 6, 7357 (2015).
Gorman, J. & Greene, E. C. Visualizing one-dimensional diffusion of proteins along DNA. Nat. Struct. Mol. Biol. 15, 768–774 (2008).
Blainey, P. C., van Oijen, A. M., Banerjee, A., Verdine, G. L. & Xie, X. S. A base-excision DNA-repair protein finds intrahelical lesion bases by fast sliding in contact with DNA. Proc. Natl. Acad. Sci. USA 103, 5752–5757 (2006).
Blainey, P. C. et al. Nonspecifically bound proteins spin while diffusing along DNA. Nat. Struct. Mol. Biol. 16, 1224–1229 (2009).
Tafvizi, A., Huang, F., Fersht, A. R., Mirny, L. A. & van Oijen, A. M. A single-molecule characterization of p53 search on DNA. Proc. Natl. Acad. Sci. USA 108, 563–568 (2011).
Rogoulenko, E. & Levy, Y. Skipping events impose repeated binding attempts: profound kinetic implications of protein–DNA conformational changes. Nucleic Acids Res. 52, 6763–6776 (2024).
Leven, I. & Levy, Y. Quantifying the two-state facilitated diffusion model of protein–DNA interactions. Nucleic Acids Res. 47, 5530–5538 (2019).
Korolev, N. et al. Electrostatic origin of salt-induced nucleosome array compaction. Biophys. J. 99, 1896–1905 (2010).
Li, W. et al. FACT remodels the tetranucleosomal unit of chromatin fibers for gene transcription. Mol. Cell 64, 120–133 (2016).
Schalch, T., Duda, S., Sargent, D. F. & Richmond, T. J. X-ray structure of a tetranucleosome and its implications for the chromatin fibre. Nature 436, 138–141 (2005).
Song, F. et al. Cryo-EM study of the chromatin fiber reveals a double helix twisted by tetranucleosomal units. Science 344, 376–380 (2014).
Luger, K., Mäder, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997).
Tremethick, D. J. Higher-order structures of chromatin: the elusive 30 nm fiber. Cell 128, 651–654 (2007).
Rowley, M. J. & Corces, V. G. Organizational principles of 3D genome architecture. Nat. Rev. Genet. 19, 789–800 (2018).
Lerner, J., Katznelson, A., Zhang, J. & Zaret, K. S. Different chromatin-scanning modes lead to targeting of compacted chromatin by pioneer factors FOXA1 and SOX2. Cell Rep. 42, 112748 (2023).
Friman, E. T. et al. Dynamic regulation of chromatin accessibility by pluripotency transcription factors across the cell cycle. Elife 8, e50087 (2019).
Chen, J. et al. Single-molecule dynamics of enhanceosome assembly in embryonic stem cells. Cell 156, 1274–1285 (2014).
Feng, X. A. et al. GAGA zinc finger transcription factor searches chromatin by 1D-3D facilitated diffusion. Nat. Struct. Mol. Biol. 32, 2359–2370 (2025).
Jonas, F., Navon, Y. & Barkai, N. Intrinsically disordered regions as facilitators of the transcription factor target search. Nat. Rev. Genet. 1–12 (2025) https://doi.org/10.1038/s41576-025-00816-3.
Mazzocca, M. et al. Chromatin organization drives the search mechanism of nuclear factors. Nat. Commun. 14, 6433 (2023).
Krois, A. S., Dyson, H. J. & Wright, P. E. Long-range regulation of p53 DNA binding by its intrinsically disordered N-terminal transactivation domain. Proc. Natl. Acad. Sci. USA 115, E11302–E11310 (2018).
Wiebe, M. S., Nowling, T. K. & Rizzino, A. Identification of novel domains within Sox-2 and Sox-11 involved in autoinhibition of DNA binding and partnership specificity *. J. Biol. Chem. 278, 17901–17911 (2003).
Krainer, G. et al. Reentrant liquid condensate phase of proteins is stabilized by hydrophobic and non-ionic interactions. Nat. Commun. 12, 1085 (2021).
Garcia, D. A. et al. An intrinsically disordered region-mediated confinement state contributes to the dynamics and function of transcription factors. Mol. Cell 81, 1484–1498.e6 (2021).
Fierz, B. & Poirier, M. G. Biophysics of Chromatin Dynamics. Annu. Rev. Biophys. 48, 321–345 (2019).
Kuchler, O. et al. Single-molecule tracking (SMT) and localization of SRF and MRTF transcription factors during neuronal stimulation and differentiation. Open Biol. 12, 210383 (2022).
Huynh, D. et al. Effective in vivo binding energy landscape illustrates kinetic stability of RBPJ-DNA binding. Nat. Commun. 16, 1259 (2025).
Garcia, D. A. et al. Power-Law Behaviour of Transcription Factor Dynamics at the Single-Molecule Level Implies a Continuum Affinity Model. https://doi.org/10.1101/637355 (2019).
Pomp, W., Meeussen, J. V. W. & Lenstra, T. L. Transcription factor exchange enables prolonged transcriptional bursts. Mol. Cell 84, 1036–1048.e9 (2024).
Popp, A. P., Hettich, J. & Gebhardt, J. C. M. Altering transcription factor binding reveals comprehensive transcriptional kinetics of a basic gene. Nucleic Acids Res. 49, 6249–6266 (2021).
Nguyen, T. et al. Chromatin sequesters pioneer transcription factor Sox2 from exerting force on DNA. Nat. Commun. 13, 3988 (2022).
Du, M. et al. Direct observation of a condensate effect on super-enhancer controlled gene bursting. Cell 187, 331–344.e17 (2024).
Yoo, W. et al. Molecular basis for SOX2-dependent regulation of super-enhancer activity. Nucleic Acids Res. 51, 11999–12019 (2023).
Dull, T. et al. A third-generation lentivirus vector with a conditional packaging system. J. Virol. 72, 8463–8471 (1998).
Suter, D. M. et al. Rapid generation of stable transgenic embryonic stem cell lines using modular lentivectors. Stem Cells 24, 615–623 (2006).
Deluz, C. et al. A role for mitotic bookmarking of SOX2 in pluripotency and differentiation. Genes Dev. 30, 2538–2550 (2016).
Amit, M. & Itskovitz-Eldor, J. (eds). Atlas of Human Pluripotent Stem Cells, (Humana Press, 2012).
Orozco-Fuentes, S. et al. Quantification of the morphological characteristics of hESC colonies. Sci. Rep. 9, 17569 (2019).
Hunt, C. J. The banking and cryopreservation of human embryonic stem cells. Transfus. Med. Hemother. 34, 293–304 (2007).
Schmidt, U., Weigert, M., Broaddus, C. & Myers, G. Cell detection with star-convex polygons. In Medical Image Computing and Computer Assisted Intervention—MICCAI 2018 (eds Frangi, A. F., Schnabel, J. A., Davatzikos, C., Alberola-López, C. & Fichtinger, G.) 265–273 (Springer International Publishing, 2018).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Larouche, K., Bergeron, M.-J., Leclerc, S. & Guérin, S. L. Optimization of competitor poly(dI-dC)•poly(dI-dC) levels is advised in DNA–protein interaction studies involving enriched nuclear proteins. BioTechniques 20, 439–444 (1996).
Kilic, S. et al. Single-molecule FRET reveals multiscale chromatin dynamics modulated by HP1α. Nat. Commun. 9, 235 (2018).
Teslenko, A. & Fierz, B. Single-molecule analysis reveals the mechanism of chromatin ubiquitylation by variant PRC1 complexes. Sci. Adv. 11, eadt7013 (2025).
Makasheva, K. et al. Multiplexed single-molecule experiments reveal nucleosome invasion dynamics of the Cas9 genome editor. J. Am. Chem. Soc. 143, 16313–16319 (2021).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Madeira, F. et al. The EMBL-EBI job dispatcher sequence analysis tools framework in 2024. Nucleic Acids Res. 52, W521–W525 (2024).
Aggarwal, T., Materassi, D., Davison, R., Hays, T. & Salapaka, M. Detection of steps in single molecule data. Cell. Mol. Bioeng. 5, 14–31 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Freese, N. H., Norris, D. C. & Loraine, A. E. Integrated genome browser: visual analytics platform for genomics. Bioinformatics 32, 2089–2095 (2016).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Acknowledgements
We thank Cedric Deluz for the pLV-PGK-H2B-SNAP plasmid, Dr. Luc Reymond for the Halo-SiR647 dye, and Dr. Armelle Tollenaere for guidance with sequencing sample preparation. We appreciate Dr. Bastien Mangeat, Elisa Cora (Gene Expression Core Facility), Dr. Miguel Garcia, Dr. Valérie Glutz, and Dr. Francesco Palumbo (Flow Cytometry Core Facility) for their support with FACS and next-generation sequencing. Thanks to Dr. Florence Pojer, Dr. Kelvin Lau, and Dr. Amédé Noredine Larabi (Protein Production and Structure Core Facility) for assistance with baculovirus generation and Sox2/Sox2a expression. We also acknowledge LCBM colleagues for histone octamer expression and the Bioimaging and Optics Core Facility for maintaining the Zeiss LSM700 UP2 and Nikon-CSU-W1 microscopes. This work was supported by the Swiss National Science Foundation (310030_200604 to B.F.), EPFL annual donation to D.S., as well as the interdisciplinary Ph.D. program from the School of Life Sciences, EPFL.
Author information
Authors and Affiliations
Contributions
Conceptualization: B.F., D.M.S., and S.S.; Methodology: S.S.; Data curation: S.S.; Formal Analysis: S.S.; Investigation: S.S.; Visualization: S.S.; Software: B.F. and S.S.; Resources: B.F., D.M.S., and S.S.; Supervision: B.F. and D.M.S.; Writing: B.F., D.M.S., and S.S.; Revision: B.F., D.M.S., and S.S.; Funding acquisition: B.F. and D.M.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ziqing Winston Zhao, who co-reviewed with Hendrik Sielaff, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sakong, S., Fierz, B. & Suter, D.M. Electrostatic properties of disordered regions control transcription factor search and pioneer activity. Nat Commun 17, 2512 (2026). https://doi.org/10.1038/s41467-026-69284-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69284-5