Abstract
Efficient water uptake and transport through xylem vessels are essential for plant growth and development. The patterned secondary cell wall (SCW) structure of xylem vessels provides robust mechanical support to withstand the strong negative pressure generated by transpiration and facilitates long-distance water transport. However, the key factors governing SCW patterning in xylem vessels and their potential for enhancing water use efficiency (WUE) remain undetermined. Here, we report the identification of a recessive maize (Zea mays) mutant drought-sensitive 1 (ds1), which is highly susceptible to water deficit. ds1 defects in SCW patterning and xylem vessel differentiation, and exhibits significantly reduced hydraulic conductivity. DS1 is the ortholog of Arabidopsis Exo70A1 and is regulated by the NAC transcription factor NECROTIC UPPER TIPS1 (NUT1) in vascular tissues. Overexpressing Exo70A1 enhanced hydraulic conductivity and consequently boosted biomass and grain yield under both well-watered and drought conditions. Thus, the NUT1–Exo70A1 module represents a promising genetic target for improving WUE in crops.
Similar content being viewed by others
Introduction
Drought causes huge losses to global agriculture and crop production1. Global freshwater use continues to grow at a rate of approximately 1% per year, with agriculture accounting for ~70% of freshwater use2. Global climate change is exacerbating this challenge. Maize (Zea mays) is a major food and forage crop whose yield is highly sensitive to water-deficit stress3. Improving water use efficiency (WUE, grain yield or biomass production per unit volume of consumed water in agricultural systems) in maize is therefore critical for ensuring global food security.
Water deficit imposes significant constraints on crop growth, ultimately leading to yield losses. Plant WUE is influenced by water uptake from roots and xylem-mediated transport, affecting photosynthetic carbon fixation and metabolic processes throughout the plant lifecycle. Water transport from root to shoot is primarily driven by transpiration-induced tension within xylem vessels and by adhesive forces between water molecules and the secondary cell walls (SCWs) of xylem vessels4,5,6. The continuity and SCW structure of xylem vessels are therefore crucial for efficient water transport in plants.
Xylem vessels consist of a series of highly specialized tubular cells interconnected by perforations at their termini7,8. Xylem vessel differentiation is a precisely regulated process involving cell specialization and elongation, localized deposition of SCWs, programmed cell death (PCD), and autolysis9. Unlike SCWs in xylem fiber cells, SCWs in xylem vessel cells are deposited as annular or spiral thickenings in protoxylem and as reticulated or pitted thickenings in metaxylem10,11. This SCW deposition pattern is essential for the generation of adhesive forces between water molecules and cell walls that enable efficient water transport through the vessels12. Xylem vessels thus facilitate the efficient transport of large volumes of water over long distances, ensuring optimal water-transport efficiency13,14,15. Disturbances in SCW deposition weaken the ability of xylem vessels to withstand the negative pressures generated by the gradient of decreasing water potential between the soil and atmosphere, causing vessel collapse and reducing water-transport efficiency16,17. In rice (Oryza sativa), Irregular xylem10 (IRX10) encodes a xylan synthase responsible for the formation of a xylan-rich nanodomain at the pit borders of xylem vessels, which are critical for water transport and leaf transpiration18. A loss-of-function mutation in maize (Zea mays) α-tubulin4 impairs SCW patterning of the protoxylem, resulting in reduced vessel diameter and water-transport efficiency12.
Xylem cell differentiation is controlled by a hierarchical network of transcriptional regulators. A subclade of NAC transcription factors (VASCULAR-RELATED NAC DOMAIN1–7) function as master switches for differentiation and SCW deposition by directly controlling the expression of genes involved in SCW biosynthesis and PCD10,19,20,21. By contrast, the NAC transcription factors XYLEM NAC DOMAIN 1 (XND1), VND-INTERACTING 1 (VNI1), and VNI2 negatively regulate xylem vessel development22,23. Disrupting XND1 and its regulatory partner Dof4.6 increased root hydraulic conductivity and drought resistance in Arabidopsis24,25. However, the factors that govern patterned SCW thickening in xylem vessels and their potential to enhance water transport and WUE remain to be characterized.
Here, we identified a drought-sensitive mutant from a maize ethyl methanesulfonate (EMS) mutant library named drought-sensitive 1 (ds1). DS1 is an ortholog of Arabidopsis Exo70A1, which encodes the Exo70 subunit of the exocyst complex. The ds1 mutant is defective in xylem vessel differentiation and patterned SCW deposition, which reduces hydraulic conductivity throughout the plant. Exo70A1 expression is regulated by the maize VND homolog NUT1 (NECROTIC UPPER TIPS 1) in vascular tissues. Overexpressing Exo70A1 enhanced hydraulic conductivity, leading to increased WUE and yield under both normal and drought conditions, making it a promising target for improving these traits in maize.
Results
ds1 is a drought-sensitive mutant
To identify key genes that function in drought resistance, we screened a maize EMS mutant library (in the B73 inbred-line background) for drought-hypersensitive mutants (Supplementary Fig. 1a, b). Using the survival rate of 3-week-old seedlings under severe drought stress as an index, we identified the drought-sensitive mutant ds1 (Fig. 1a). Under drought stress, the seedling survival rate of ds1 was significantly lower than that of B73, but under normal watering conditions, the growth of ds1 and B73 seedlings was comparable (Fig. 1a, b). After 6 weeks of growth under normal watering conditions, ds1 plants were shorter than B73 plants. The upper leaves of ds1 plants were curled and wilted when transpiration increased at noon (11:00 a.m.), but not in the morning (6:00 a.m.) (Fig. 1c–e). However, the water loss rates of detached leaves were comparable between ds1 and B73, indicating that the drought sensitivity and leaf-curling phenotype of ds1 were not due to rapid water loss from leaves (Supplementary Fig. 1c, d).
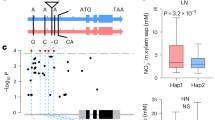
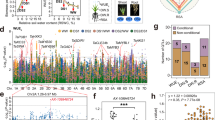
a Seedling survival of B73 and ds1. Photographs were taken before and after drought treatment. Scale bar, 5 cm. b Survival rates of B73 and ds1 after drought treatment. The experiment was performed three times, and 15 plants per genotype were tested in each experiment. Morphology of B73 and ds1 plants at 6:00 a.m. (c) and 11:00 a.m. (d) at the V7 stage (when the seventh leaf ligule is visible). Scale bars, 10 cm. e Representative photographs of the top three leaves of B73 and ds1 plants at 6:00 a.m. and 11:00 a.m. Scale bar, 5 cm. f MutMap analysis mapped ds1 to the coding region of Zm00001d001765 on chromosome 2. g Drought resistance of LH244 (WT) and exo70a1-ko seedlings. Scale bars, 5 cm. Survival rates of exo70a1-ko1 (h) and exo70a1-ko2 (i) compared to WT. j Drought resistance of WT and Exo70A1-OE seedlings. Scale bars, 5 cm. Survival rates of Exo70A1-OE1 (k) and Exo70A1-OE2 (l) compared to WT. m Relative leaf water content (RLWC) from different positions of WT, exo70a1-ko, and Exo70A1-OE plants under well-watered (WW) conditions. The leaves from three individual adult plants with similar growth were measured. n Relative Exo70A1 expression levels in the leaf (V4), leaf (V6), root, stem, tassel, and silk. o In situ RT-PCR assay of Exo70A1 expression. Dark blue signal indicates the presence of Exo70A1 transcripts in cross-sectional images of the root (upper panel) and stem (lower panel). No reverse transcription (No RT) and 18S rRNA (18S) were used as controls. Images are representative of three independent replicates with similar results. Scale bars, 100 μm. In b, h, i, k, l, m, and n, data are presented as mean ± SD of 3 biological replicates; p-values were obtained by two-sided Student’s t-test.
F1 plants from a cross between ds1 and B73 grew normally and did not have visible differences from B73. When the F2 progeny were planted in the field, approximately 25% of the F2 plants showed curled and wilted leaves resembling those of the original ds1 mutant at noon, indicating that ds1 was caused by a single recessive mutation (Supplementary Fig. 1e). MutMap analysis26 revealed a strong co-segregation signal of single-nucleotide polymorphisms (SNPs) on chromosome 2 with the mutant phenotype. The most significant SNP that co-segregated with the mutant phenotype was a G-to-A transition in the ninth exon of Zm00001d001765, which led to premature termination of Zm00001d001765 protein translation (Fig. 1f). To determine whether Zm00001d001765 was the causal gene, we obtained three more EMS mutants of Zm00001d001765 from a public B73 EMS mutant library (http://maizeems.qlnu.edu.cn)27 and named them ems1, ems2, and ems3 (Supplementary Fig. 2a). The seedling survival rate was significantly lower in these three mutants than the wild type (Supplementary Fig. 2b–e), and after 6 weeks of growth, they phenocopied the original ds1 mutant (Supplementary Fig. 2f–h). Curling and wilting of the upper leaves at noon was more evident in plants grown in the field than in the greenhouse. Moreover, under drought conditions, the upper leaves were almost completely dried out and burned (Supplementary Fig. 2i). The allelic test by crossing the original ds1 mutant with each of the new mutants (ds1 × ems1, ds1 × ems2, and ds1 × ems3) showed that all F1 plants exhibited the mutant phenotypes, supporting that Zm00001d001765 is the causal gene of ds1 (Supplementary Fig. 2j).
DS1 encodes the Exo70 subunit of the exocyst complex and functions in xylem vessel development
Phylogenetic analysis showed that Zm00001d001765 is an ortholog of Arabidopsis Exo70A1, a member of a small gene family encoding the Exo70 subunit of the exocyst complex (Supplementary Fig. 3). This multi-subunit protein complex tethers trafficking vesicles to a specific region of the plasma membrane in eukaryotes28,29. The Arabidopsis exo70a1 mutant is defective in tracheary-element development and Casparian strip deposition in root endodermal cells30,31. Through CRISPR-Cas9-based gene editing, we generated gene knockout mutants (exo70a1-ko1 and exo70a1-ko2), as well as Exo70A1-overexpressing (Exo70A1-OE) plants with a translational in-frame MYC-tag fusion, in the LH244 genetic background, which is amenable to genetic transformation and is hereafter referred to as the wild type (WT) (Supplementary Fig. 4a–c). The exo70a1-ko lines had phenotypes similar to those of ds1, including lower seedling survival rates under drought stress, leaf wilting at noon, and dwarfism at the adult stage (Fig. 1g–i, Supplementary Fig. 4d–e). By contrast, the Exo70A1-OE lines exhibited significantly higher survival rates under drought stress than WT plants, without other visible phenotypic differences (Fig. 1j–l, Supplementary Fig. 4d–e). There were no significant differences in the water loss rates of detached leaves among the genotypes, and their stomatal apertures and stomatal densities were comparable (Supplementary Fig. 4f–j). However, the relative leaf water content (RLWC) of the top five leaves at noon was significantly lower in exo70a1-ko than in WT plants, even though the water supply was sufficient, and there were no significant differences in RLWC between Exo70A1-OE and WT plants (Fig. 1m). These results suggest that Exo70A1 is important for water transport to top leaves but does not control leaf water loss.
Exo70A1 is highly expressed in roots, stems, and silks (Fig. 1n). In situ PCR assays revealed that Exo70A1 is expressed in vascular tissues of the root and stem (Fig. 1o). To identify putative interaction partners of Exo70A1, we performed immunoprecipitation-coupled mass spectrometry analysis (IP-MS) of the transgenic maize plants expressing an Exo70A1-GFP fusion protein using a GFP antibody. Other components of the exocyst complex (EXO84b, SEC15b, SEC10, SEC8, and SEC6) were identified, providing evidence that Exo70A1 functions as a component of the exocyst complex in vivo (Fig. 2a, b, Supplementary Data 1). Enzymes related to SCW biosynthesis were also identified, including peroxidases, xyloglucan endotransglucosylases, and cellulose synthases (Fig. 2a).
a Interacting proteins identified by immunoprecipitation-mass spectrometry (IP-MS) of Exo70A1-OE plants. b Cryo-electron microscopy structure of the exocyst complex28 (PDB ID: 5YFP). Subunits identified are color-coded based on the IP-MS score; undetected subunits are gray. c Cleared sheath tissue stained with toluidine blue. The red arrow indicates lignified annular secondary cell wall (SCW) thickenings in protoxylem. Scale bars, 30 μm. d Distance between two adjacent annular thickenings. n = 90 distances between two adjacent annular thickenings measured from 3 plants. e Semi-thin cross-sections of primary roots. Scale bars, 100 μm. f Root hydraulic conductivity (Lpr) regression slope, based on 3 replicated experiments. g Dye uptake and movement assay using stems of 14-day-old seedlings. Red arrows indicate dye front. h Rate of dye movement. n = 9 plants. i Scanning electron microscopy (SEM) images of stem vascular bundles. Red asterisks highlight undifferentiated vessels. Px, protoxylem; Mx, metaxylem. j Percentage of abnormal vascular bundles per genotype. k Xylem vessels area for indicated genotypes. n = 9 vascular bundles from 3 plants. l Pitted metaxylem vessels stained with Direct Red in the indicated plants (with identical genetic background of WT). Scale bar, 100 μm. m Length of pitted vessels. n = 50 vessels from 3 plants. n Schematic of metaxylem vessel longitudinal section. PCW, Primary cell wall. o Transmission electron microscopy (TEM) of root metaxylem vessel longitudinal sections. Brown and green lines delineate SCW thickenings and PCWs. XPs, xylem parenchyma cells. Scale bars, 2 μm. Thickness (p) and adjacent distance (q) of SCW thickenings in root metaxylem vessels. n = 60 SCW thickenings from 3 plants in p. n = 60 distances between two adjacent SCW thickenings from 3 plants in q. In e, and i, the experiment was repeated at least three times, and a representative image was selected for display. In d, f, h, j, k, m, p, and q, data are presented as mean ± SD; p-values were obtained by two-sided Student’s t-test. In c, e, g, i, l, and o, representative images are shown.
Given that the exocyst complex targets vesicles to the plasma membrane and that some Exo70A1-interacting proteins function in SCW biosynthesis, we examined the patterns of SCW thickening in WT, exo70a1-ko, and Exo70A1-OE plants. The density of annular thickenings in the protoxylem of leaf sheaths was significantly reduced in exo70a1-ko plants compared to the WT, and approximately 15% of exo70a1-ko roots exhibited defects in metaxylem vessel differentiation (Fig. 2c–e). Importantly, root hydraulic conductivity was significantly lower in exo70a1-ko than WT plants, whereas Exo70A1-OE plants exhibited the opposite phenotype (Fig. 2f). Additionally, a dye-movement assay demonstrated that transport rates through the xylem vessels of stems and leaf veins were significantly lower in exo70a1-ko plants but higher in Exo70A1-OE plants than in WT plants (Fig. 2g, h; Supplementary Fig. 5a, b).
In cross sections from exo70a1-ko stems and leaf veins, over 90% of vascular bundles exhibited defects characterized by undeveloped xylem vessels (Fig. 2i, j; Supplementary Fig. 5c). Therefore, the area of xylem vessels was significantly reduced in exo70a1-ko compared to WT plants (Fig. 2k). The Exo70A1-OE lines showed normal metaxylem and protoxylem development, with larger xylem vessels than WT plants (Fig. 2i, k). In addition, metaxylem vessels were significantly shorter in exo70a1-ko but longer in Exo70A1-OE vs. WT plants (Fig. 2l, m). Transmission electron microscopy (TEM) of longitudinal sections through metaxylem vessels of the stem and root revealed that SCW thickenings were markedly thinner and less dense in exo70a1-ko than the WT but significantly denser and thicker in Exo70A1-OE plants (Fig. 2n–q; Supplementary Fig. 5d–f). Therefore, the compromised water conductivity of exo70a1-ko mutants is likely due to defects in xylem vessel development, including reductions in the number and size of vessels and changes in the pattern of SCW thickenings. By contrast, overexpressing Exo70A1 promoted xylem development and increased water-transport capacity in maize.
NECROTIC UPPER TIPS1 regulates Exo70A1 expression in vascular tissues
The upper-leaf wilting observed in exo70a1 plants at noon resembled that of the maize necrotic upper tips1 (nut1) mutant, which harbors a mutation in a NAC-type transcription factor gene32. Notably, two NUT1-binding sequences (TTGCTT/TNNCNTNNNNNNNAAGN) were identified in the promoter of Exo70A1, as evidenced by the NUT1 DNA-Affinity Purification-Sequencing (DAP-seq) analysis32 (Fig. 3a, Supplementary Fig. 6a). In an electrophoretic mobility shift assay (EMSA), prokaryotically expressed MBP-NUT1 fusion protein bound to the biotin-labeled NUT1-binding sequence within the Exo70A1 promoter. This binding was inhibited by adding a 100-fold excess of an unlabeled competitor probe but not by a 200-fold excess of a mutated competitor probe, confirming the specificity of the binding (Fig. 3b). A yeast one-hybrid assay further supported this finding (Fig. 3c). Chromatin immunoprecipitation (ChIP)-qPCR analysis using a GFP antibody in maize protoplasts transfected with an ubi:NUT1-GFP construct revealed significant enrichment of the DNA fragment containing the NUT1-binding sequences at the Exo70A1 promoter region (Fig. 3d). In transient transactivation assays in maize leaf protoplasts, NUT1 activated the expression of the luciferase reporter gene driven by a 1.0-kb promoter fragment of Exo70A1. However, when the NUT1-binding sequence was deleted from the promoter, this activation was abolished (Fig. 3e, f).
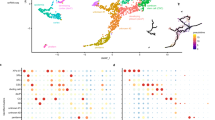
a Diagram of the Exo70A1 promoter, containing two predicted NUT1 binding sequences (NUT1-BS). b Electrophoretic mobility shift assay (EMSA) for binding of NUT1 to the NUT1-BS (NUT1 binding sequence) in the Exo70A1 promoter. Competition was tested by adding excess mutated unlabeled probes as indicated. Red letters represent mutated sequence. Red asterisks indicate the specific shifted bands. c Yeast one-hybrid assay for binding of NUT1 to the Exo70A1 promoter. Six tandem repeats of the NAC recognition motif were ligated in front of the LacZ reporter gene. d Chromatin immunoprecipitation (ChIP)-qPCR showed that NUT1 binds Exo70A1 promoter at P1 region that indicated in a. e Diagram of the effector and reporter constructs used for transient expression assays in maize protoplasts. f NUT1 activates the expression of the firefly-luciferase reporter gene driven by 1.0-kb promoter sequence of Exo70A1. Immunoblot shows levels of NUT1-GFP fusion protein in different samples. Relative NUT1 expression levels in various tissues (g) and root stele and cortex (h). Relative Exo70A1 expression levels in root stele (i), cortex (j), and stem (k) of two nut1-ko lines. In d, f, g–k, data are presented as mean ± SD, based on 3 independent replicates. In d, g–k, p-values were obtained by two-sided Student’s t-test. Different letters indicate significant differences (p < 0.05) determined by one-way ANOVA with Tukey’s multiple-comparison test in f.
Like Exo70A1, NUT1 was also highly expressed in roots, stems, and silks (Fig. 3g, Supplementary Fig. 6b–e). In roots, NUT1 expression was significantly higher in the stele than the cortex (Fig. 3h). We examined the expression of Exo70A1 in two independent nut1 knockout lines (nut1-ko1 and nut1-ko2) generated by CRISPR-Cas9-targeted mutagenesis (Supplementary Fig. 7a). Strikingly, Exo70A1 expression was dramatically reduced in the stele of nut1-ko roots but remained unchanged in the cortex (Fig. 3i, j). In stem tissues, Exo70A1 expression was also markedly reduced in nut1-ko plants (Fig. 3k). These results suggest that NUT1 modulates Exo70A1 expression in vascular tissues.
Exo70A1 acts downstream of NUT1 to regulate xylem vessel development and hydraulic conductivity
Similar to exo70A1-ko plants, seedling survival after drought stress was significantly lower in nut1-ko plants than the WT (Fig. 4a–c), and nut1-ko xylem vessels were significantly reduced in terms of both length and cross-sectional area (Fig. 4d, e; Supplementary Fig. 7b, c). Root hydraulic conductivity was markedly impaired in nut1-ko compared to WT plants (Fig. 4f). The thickness and density of SCW thickenings in root metaxylem vessels were also reduced in nut1-ko plants (Fig. 4g–i). A dye-movement assay indicated that water transport in stems and leaf veins was significantly compromised in nut1-ko compared to the WT (Fig. 4j–m). Moreover, nut1-ko plants were significantly shorter than WT plants under both normal and drought conditions (Fig. 4n; Supplementary Fig. 7d, e). The tips of upper leaves and tassels in nut1-ko plants exhibited signs of wilting or scorching, similar to exo70a1-ko plants (Fig. 4n; Supplementary Fig. 7f–h). However, when the Exo70A1-overexpression construct was introduced into nut1-ko plants, the dwarfism and leaf-tip scorching phenotypes were largely rescued, and the defects in the patterning of the SCW (thickness and density) were also restored, suggesting that Exo70A1 functions downstream of NUT1 (Fig. 4n–p, Supplementary Fig. 7i–k).
a Drought sensitivity of nut1-ko seedlings. Scale bars, 5 cm. Survival rate of nut1-ko1 (b) and nut1-ko2 (c) seedlings after drought treatment. The experiment was performed three times, and 15 plants per genotype were tested in each experiment. d Images of pitted metaxylem vessels stained with Direct Red. Scale bars, 100 μm. e Length of pitted vessels. n = 123 vessels from 5 plants. f Root hydraulic conductivity of WT, nut1-ko1, and nut1-ko2. n = 3 biological replicates. g TEM images of longitudinal sections through root metaxylem vessels. Brown lines delineate SCW thickenings in metaxylem vessels. Green lines delineate PCWs. Scale bars, 2 μm. Thickness (h) and adjacent distance between of SCW thickenings (i) in root metaxylem vessels. n = 60 SCW thickenings from 3 plants in h. n = 60 distances between two adjacent SCW thickenings from 3 plants in i. Dye uptake and movement assays using stems (j) and leaf veins (l) of 14-day-old seedlings. Red arrows indicate distances traveled by dye from the base of the stem or leaf blade within the same period of time. Scale bars, 1 cm. Rate of dye movement in stems (k) and leaf veins (m). n = 20 plants in k, and n = 9 plants in m. n Phenotypes of the indicated plants. Red arrows indicated the withered leaf tip of nut1-ko plant. o Dye uptake and movement assay using stems of 14-day-old seedlings. Red arrows indicate distance traveled by the dye from the base of the stem within the same period of time. p Rate of dye movement. n = 9 plants. In b, c, e, f, h, i, k, m, and p, data are presented as mean ± SD. In b, c, e, f, h, i, k, and m, p-values were obtained by two-sided Student’s t-test. Different letters indicate significant differences (p < 0.05) determined by one-way ANOVA with Tukey’s multiple-comparison test in p. In a, d, g, j, l, n, and o, representative images are shown.
In addition, the stem diameter and puncture force (a measure of the mechanical strength of stems) of both exo70a1-ko and nut1-ko plants were significantly reduced compared to the WT (Supplementary Fig. 8a–f). Consistent with these findings, lignin and cellulose contents were lower in exo70a1-ko and nut1-ko vs. WT stems (Supplementary Fig. 8g–j), as was grain yield (Supplementary Fig. 8k–m). In general, the phenotypes of nut1-ko plants resembled those of exo70a1-ko plants, supporting the notion that NUT1 functions in the same pathway as Exo70A1 in regulating xylem vessel development, thereby affecting water transport and overall plant growth, even under sufficient watering conditions.
Overexpressing Exo70A1 increases water use efficiency in the field
Because Exo70A1-OE plants exhibited significantly higher hydraulic conductivity in vascular tissues than WT plants, we next asked whether they would show increased growth and grain yield under field conditions. We performed field trials under well-watered (WW) and drought conditions over two years (2023 and 2024) in Zhangye (northwest China), where rainfall is scarce during the maize-growing season. Water was applied through controlled drip irrigation. The total amount of water applied to the drought-stressed plots was approximately 62.5% (in 2023) and 72.1% (in 2024) that applied to the well-watered plots (Supplementary Data 2).
The Exo70A1-OE plants exhibited greater plant height, stem diameter, and stem mechanical strength than WT plants, as well as higher cellulose and lignin contents and longer metaxylem vessels (Fig. 5a–f; Supplementary Fig. 8i–j). Therefore, the biomass production (biomass WUE), a measure of the biomass produced (excluding grains) per unit volume of consumed water (including irrigation water and natural rainfall), was higher in Exo70A1-OE plants than WT plants due to greater biomass under the same water consumption conditions (Fig. 5g; Supplementary Fig. 8n). Moreover, in field trials conducted over two consecutive years (2023 and 2024), the grain yield of Exo70A1-OE plants was consistently superior to that of WT plants under both well-watered and drought conditions, resulting in a significantly enhanced WUE with respect to harvested grains (grain WUE) (Fig. 5h–j; Supplementary Fig. 8o–p).
a Morphology of WT, exo70a1-ko, and Exo70A1-OE plants grown under well-watered (WW) and drought conditions in the field. Scale bars, 30 cm. b Stem cross sections of WT and Exo70A1-OE adult plants. Scale bar, 1.0 cm. c Statistical analysis of stem diameter. n = 20 plants. d Statistical analysis of stem puncture strength. n = 40 plants. Representative photographs (e) and statistical analysis (f) of stem metaxylem vessels under drought conditions. Scale bars, 100 μm. n = 45 vessels from 3 plants. g Biomass WUE of plants grown under WW and drought conditions in the field in 2023. h Representative photographs of maize ears from plants grown under WW and drought conditions. Scale bars, 5 cm. Grain WUE of different genotypes grown in the field under WW and drought conditions in 2023 (i) and 2024 (j). Photographs of maize ears (k) from different F1 plants (l) grown in the field under WW and drought conditions. Scale bars, 5 cm. m Grain WUE of different F1 plants grown under WW and drought conditions in 2024. In g, i, and j, data were obtained from 3 replicate plots, and 8–12 plants were evaluated per plot. In m, data were obtained from 3 replicate plots, and 14–18 plants were evaluated per plot. In c, d, f, g, i, j, and m, data are presented as mean ± SD; p-values were obtained by two-sided Student’s t-test. In a, b, e, h, k, and l, representative photographs are shown.
Finally, since commercial maize cultivars are F1 plants known for their remarkable heterosis, we tested whether the superior performance of Exo70A1-OE plants would persist in a heterologous F1 background. We generated F1 plants by crossing the homozygous Exo70A1-OE or WT plants (as male parents) with T13 (as the female parent), and the enhanced Exo70A1 expression level was confirmed in the F1 plants (Supplementary Fig. 9a). T13 is a tester line that produces excellent heterosis with LH244. The grain WUE of T13 × Exo70A1-OE F1 plants was consistently higher than that of control F1 plants (T13 × WT) under both well-watered and drought conditions (Fig. 5k–m; Supplementary Fig. 9b). T13 × Exo70A1-OE F1 plants also had greater stem diameters and plant heights than T13 × WT F1 plants (Supplementary Fig. 9c–f). These results indicate that overexpressing Exo70A1 increases plant growth and grain yield under the tested irrigation regimes; concordant increases in hydraulic capacity suggest that improved water delivery contributes to these benefits, while the effects on canopy development and resource allocation may also be involved.
Discussion
In this study, we demonstrated that the loss of Exo70A1 function confers drought sensitivity in maize, which is characterized by a significant decrease in seedling survival under drought and a rapid wilting of the upper leaves under high transpirational demand. The drought-sensitive phenotype of the exo70a1-ko mutant was not due to faster leaf water loss but rather to impaired hydraulic conductivity. The exo70a1-ko plants exhibited defects in metaxylem development, including vessel-cell differentiation and patterned SCW thickening, which impaired water-transport efficiency throughout the plant.
The xylem vessels of vascular plants exhibit elaborate patterns of SCW deposition, including annular or spiral patterns in the protoxylem and reticulated or pitted patterns in the metaxylem, which are essential for mechanical support and the development of adhesive forces for water transport. A continuous water column is maintained within xylem vessels, which are capable of withstanding the substantial negative pressures (below −1 MPa) generated by transpiration16. The thickening of xylem vessel-cell walls is crucial for providing the mechanical strength necessary to resist these pressures, thereby ensuring the uninterrupted flow of xylem sap. During periods of high transpirational demand, such as at noon, the generation of strong negative pressures requires robust cell wall support to prevent vessel collapse or blockage. Furthermore, intricate SCW patterning enables the development of capillary forces for hydraulic conductivity, as demonstrated by the adhesive interactions between water molecules and the cell wall4,5. In exo70a1-ko mutants, undeveloped xylem vessels and reduced thickening of the SCW reduced hydraulic conductivity and compromised mechanical strength throughout the plant. Perhaps the reduced hydraulic conductivity led to reductions in plant height and total biomass in the mutants (Fig. 5).
The NAC transcription factor VND proteins play pivotal roles in the regulatory network of vessel-cell differentiation and development. However, little is known about the processes by which elaborated SCW thickenings are formed in xylem vessel cells. In this study, we determined that NUT1, a homolog of Arabidopsis VND6, directly regulates Exo70A1 expression to influence the SCW thickening patterns of xylem vessels (Fig. 6). Arabidopsis VND6 affects xylem vessel differentiation, whereas VND7 influences protoxylem cell development19,33. NUT1 was previously shown to function predominantly in protoxylem development in maize32. Here, analysis of a newly generated maize CRISPR-targeted nut1 loss-of-function mutant revealed substantial deficiencies in patterned SCW thickening of the metaxylem vessels, as well as defects in protoxylem development. Protoxylem plays a critical role in rapidly developing immature tissues, while metaxylem gradually differentiates in the maturation tissues and subsequently supersedes the role of protoxylem as the primary conductive tissue12. Similar to nut1-ko, the observed wilting and scorching phenotypes of the young leaves and tassels in exo70a1-ko adult plants under field conditions may be attributed to both the defects of annular patterning in the protoxylem and impaired metaxylem development (Fig. 1).
In developing xylem vessel cells, the transcription factor NUT1 activates the expression of Exo70A1, which encodes the Exo70 subunit of the exocyst complex. The exocyst complex tethers secretory vesicles, which carry enzymes and substrates required for secondary cell wall (SCW) deposition, to the target membrane in xylem vessel cells. Loss-of-function mutants exhibit defects in patterned SCW thickening and impaired xylem vessel development, leading to a significant reduction in overall plant water conductivity. PCW primary cell wall, SCW secondary cell wall.
Exo70A1 encodes an Exo70 subunit of the exocyst complex, which directs trafficking vesicles to specific sites on the plasma membrane. The C-terminal portion of Exo70A1 interacts specifically with the phospholipid bilayer and is essential for efficient vesicle tethering at the plasma membrane29. Exocytosis is thought to occur during SCW patterning, integrating cellulose synthase complexes into the plasma membrane and enabling the deposition of substrates for cell-wall biosynthesis and lignification at specific sites7,8. In Arabidopsis, loss of Exo70A1 function led to the mislocalization of CASPARIAN STRIP MEMBRANE DOMAIN PROTEINs and CELLULOSE SYNTHASE 7, thereby impairing the formation of ring-like Casparian strips in root endodermal cells and xylem development in the root stele30,34. The Exo70 subunit mediates the tethering of post-Golgi secretory vesicles to plasma membrane for exocytosis35,36. Abnormal membrane-bound compartments accumulate in the developing xylem vessels and endodermal cells of the Arabidopsis exo70a1-1 mutant14,31,37. We suppose that enhancing Exo70A1 function may increase the ability of vesicles to bind to the plasma membrane, thereby reinforcing exocyst complex activity. In the present study, IP-MS revealed that Exo70A1 interacts with enzymes responsible for SCW biosynthesis, such as peroxidases, cellulose synthases, and xyloglucan endotransglucosylase/hydrolases, as well as other components of the exocyst complex, suggesting a role for exocyst-mediated trafficking at the sites of SCW deposition in xylem vessels. Recently, it is reported that in Arabidopsis, the exocyst complex (consisting of Exo70A1 and SEC6) is pre-assembled before fusing with secretory vesicles that carry the cellulose synthase complex, which ensures efficient membrane trafficking38. Extending this analysis to maize will provide direct evidence for Exo70A1-mediated secondary cell-wall patterning. Although enhance SCW thickening was observed in Exo70A1-OE plants, it did not arrest vessel cell axial elongation, suggesting that Exo70A1 may play a major role in SCW deposition of elongated vessels. The defects in xylem vessel development observed in exo70a1 mutants may be attributed to disrupted vessel cell differentiation, a highly regulated process involving cell specialization, elongation, localized secondary cell wall (SCW) deposition, programmed cell death (PCD), and autolysis9. Impaired SCW deposition may lead to asynchronous differentiation of adjacent vessel cells, thereby compromising the continuity of xylem vessels. Arabidopsis Exo70A1 was reported to regulate the polar transport of auxin39. Given auxin’s critical role in determining vessel size and number39,40, the defective metaxylem vessel differentiation in exo70a1 mutants, as well as the enlarged metaxylem vessels in Exo70A1-OE plants, may result from Exo70A1-mediated changes in auxin polar transport.
It is supposed that enhancing water-transport efficiency may improve plant WUE. Here, we obtained direct evidence that enhancing the function of Exo70A1 in maize increased plant biomass and grain yield under both sufficient watering and drought conditions, indicating that Exo70A1-OE plants have improved WUE. The reinforced SCW patterning in Exo70A1-OE plants might enhance the adhesive forces between water molecules and the SCW, facilitating water transport in xylem vessels. The increased length of xylem vessels in Exo70A1-OE plants might lead to fewer perforation plates per unit length, thereby enhancing water-transport efficiency. In addition, larger xylem area observed in Exo70A1-OE plants may also be beneficial. It is likely that more efficient water transport enhances the root’s ability to take up water from the soil, thereby competing with water evaporation from the soil surface into the atmosphere. Stem diameter and total cellulose and lignin contents were also enhanced in Exo70A1-OE plants, providing strong mechanical support for the stem and potentially increasing lodging resistance. Currently, no remarkable differences in root morphology, Casparian strip barrier function, or pollen fertility have been observed between Exo70A1-OE and WT plants.
Overall, our findings reveal that the VND-like transcription factor NUT1 regulates Exo70A1 expression in developing xylem vessel cells, providing insight into the mechanisms of xylem vessel development and SCW thickening, which are crucial for enhancing crop WUE. Increasing the expression of Exo70A1 may hold significant potential for improving water transport and WUE in maize and other crops.
Methods
Drought-resistance assay for maize seedlings
Seeds were germinated in soil for 4 days, and then transplanted into cultivation boxes (33 × 19 × 12 cm, length × width × depth). A substrate mixture consisting of vermiculite, nutritious soil, and Pindstrup mix (Denmark) was prepared at a ratio of 1:1:1. After 10 days of growth under well-watered (WW) conditions, the water supply was terminated. Photograph was taken for plant growth under WW conditions within the following 10 days. Watering was resumed when significant wilting differences were observed (typically around 25 days after water deprivation), allowing plants to recover. Five days after re-watering, the number of surviving plants was recorded, and the plants were photographed again.
Phenotypic analysis of leaf wilting in the greenhouse
Seeds of wild type and transgenic lines were germinated, and the seedlings were cultivated in white barrels (35 cm diameter × 35 cm height). After approximately 40 days, the plants reached the V7 stage (when the seventh leaf ligule is visible). At 6:00 a.m. and 11:00 a.m., photographs were taken of the entire plants and their top three leaves. To monitor the water loss rate of detached leaves, fully expanded leaves were excised, and allowed them to air-dry on a desk in the greenhouse. The leaves were weighed at specific time points.
Measurement of the relative leaf water content from the plants grown in the field
Expanded leaf blades from different positions on the plant were sampled at 7:00 a.m., when the vapor pressure deficit (VPD) was approximately 1.15 kPa. The fresh weight (FW) of the leaves was immediately measured. Subsequently, they were placed into mesh bags which were immersed in water in the dark at room temperature for 24 h. Afterward, the samples were removed from the bags, and excess water on the surfaces was gently blotted with absorbent paper. The turgid weight (TW) was then recorded after re-weighing. Following this step, the samples were transferred to an oven at 80 °C for 3 days until they reached a constant dry weight (DW). Finally, the relative leaf water content (RLWC) was calculated using the formula: RLWC (%) = [(FW − DW) / (TW − DW)] × 100.
Transgenic maize generation
For CRISPR-Cas9-mediated targeted gene knockout of Exo70A1 and NUT1, the guide RNA was cloned into pBUE411 vector41. For overexpression constructs, the full-length CDS of Exo70A1 from B73 was cloned into the pBCXUN vector (NCBI GenBank: FJ905215.1), which contains the maize ubiquitin 1 promoter sequence (1991-bp) to drive a constitutive transgene expression. Exo70A1-overexpressing (Exo70A1-OE) plants were generated with a translational fusion to a MYC-tag. All resulting vectors were confirmed by sequencing and transformed into immature embryos of the maize inbred line LH244 through Agrobacterium-mediated transformation42. Transgenic plants were created by the Crop Functional Genomics and Molecular Breeding Research Center, China Agricultural University. To identify the positive mutants, the PCR products encompassing the gRNA-targeted site for each of the knockout transgenic plant was sequenced by Sanger DNA sequencing. Exo70A1 overexpression-positive transgenic lines (Exo70A1-OE) were identified through the transgene specific PCR analysis. Reverse transcription-quantitative (RT-qPCR) was performed to determine the Exo70A1 expression levels in the Exo70A1-OE lines. All primers used in this study are listed in Supplementary Data 3.
RNA extraction and RT-qPCR
Total RNA was extracted from leaves, roots, stems, tassels and silks using TRIzol reagent (Biotopped, Cat#T6631G) and subsequently treated with RNase-free DNase I (Takara, Cat#2270 A) to eliminate genomic DNA contamination. 2 μg of purified total RNA was reverse-transcribed into cDNA using recombinant M-MLV reverse transcriptase (Promega, Cat# M1701). RT-qPCR analysis was conducted on an ABI StepOnePlus real-time PCR system with SYBR premix (Mei5bio, Cat#MF013). The maize Ubi2 gene was served as the internal reference for data normalization.
In-situ PCR
In-situ PCR was performed as previously described43, with minor modifications. The maturation zone of the primary root of a 5-day-old wild type (LH244) plant and the stem of a 40-day-old wild type plant were analyzed. Samples were embedded in low-melting-point agarose, and 50-μm-thick sections of roots and stems were obtained using a microtome (Leica, Wetzlar, Germany). These sections were transferred into 1 mL of cold sterile water containing 100 U of RNase inhibitor (Coolaber, Cat#RE1021) and kept on ice. Sections were treated with RNase-free DNase I (Takara, Cat#2270 A) for 45 min. cDNA synthesis was carried out using recombinant M-MLV reverse transcriptase (Promega, Cat#M1701). Gene specific primers were designed for Exo70A1. The PCR cycling parameters were as follows: initial denaturation at 95 °C for 30 s, followed by 28 cycles of 95 °C for 10 s, 58 °C for 30 s and 72 °C for 20 s, with a final extension at 72 °C for 5 min. Subsequent steps were performed as described previously44. Sections were blocked with 0.1% bovine serum albumin (BSA) for 30 min, incubated with 0.1% digoxin antibody (Roche, Cat#11093274910) for 1 h, rinsed twice with 0.1 M Tris-HCl buffer for 15 min each time, and stained with the staining solution (Roche, Cat#11442074001) for 45–60 min. The reaction was terminated with distilled H2O, and the sections were observed under an optical microscope (OLYMPUS BX51).
Root semi-thin cross-section observation
Roots from 4-day-old seedlings were immersed in a pre-cooled fixation buffer consisting of 1% (w/v) paraformaldehyde (Sigma, Cat#441244) and 2.5% (v/v) glutaraldehyde (Sigma, Cat#8.20603) dissolved in 0.1 M phosphate buffer (PB; pH 7.2). To eliminate air bubbles and ensure complete fixative penetration, samples were fixed under vacuum at room temperature for 2–4 h. After fixation, residual fixative was removed by two 15-min washes with 0.1 M PB (pH 7.2). Dehydration was performed using a graded ethanol series (30%, 50%, 75%, 80%, 90%, 95%, and 100% [v/v] ethanol) to minimize tissue shrinkage. Samples were incubated in each ethanol concentration for 30 min, with the 100% ethanol step repeated twice. Resin infiltration was initiated by incubating samples in a 1:1 (v/v) mixture of 100% ethanol and LR White resin (Sigma-Aldrich, Cat#L9774) for 1 h, followed by two sequential incubations in fresh LR White resin (each ≥8 h; resin was refreshed during the second incubation to optimize infiltration efficiency). Infiltrated samples were transferred to embedding molds filled with fresh LR White resin and polymerized at 60 °C for 14 h using an embedding polymerization system (Zhongxingbairui Model Z-BJ0010) to form solid blocks. Semi-thin cross-sections (1–1.5 μm thick) were cut with a ultramicrotome (Leica EM UC7) equipped with a Diatome diamond knife (Biel Ultra 35°). Sections were carefully transferred onto glass slides pre-mounted with small water droplets to ensure flat spreading, then dried on a slide warmer (Kedi Model KD-H) at 70 °C. Dried sections were stained with 1% (w/v) aqueous crystal violet solution (Sigma-Aldrich, Cat#C6158) at room temperature for 5 min. Excess stain was removed by gentle rinsing with deionized water, and slides were re-dried on the slide warmer. Stained sections were observed and imaged using an optical microscope (OLYMPUS BX51).
Root hydraulic conductivity
Root hydraulic conductivity was measured as described previously45, with few modifications. 21-day-old hydroponically grown seedlings uniformly growing were selected. The aboveground portion of the plants was carefully excised using a sharp blade, leaving approximately a 2-cm-long stem base intact. The root system, along with the stem base, was then placed into a pressure chamber (Soil Moisture, MODEL 3005F01 Plant Water Status Console), ensuring that the chamber was sealed tightly around the stem base without causing damage. An air pump was activated to maintain constant air pressure at predetermined levels. Exudate was collected under pressures of 0.5, 0.4, 0.3, 0.2, and 0.1 MPa for 2 min (t) each, and its mass (M) was subsequently measured using on an analytical balance. Following the measurements, root samples were scanned using a root scanner (Epson 1680), and the root surface area (S) was calculated using root analysis software (WinRHIZO). Steady-state root water flow (Jv) was computed as Jv = M / (t × S). Root hydraulic conductivity (LPr) was determined by calculating the slope of the regression line between Jv and the applied water pressure gradients, expressed as LPr = Jv / ΔP.
Microscopy analysis of the xylem vessels
For the microscopy analysis of protoxylem vessels, the sheath tissue was cleared using a buffer solution (comprising 20 mL 85% lactic acid, 20 mL dibutyl phthalate, 10 mL benzyl benzoate, 10 mL xylene, 20 g phenol, 20 g trichloroacetaldehyde monohydrate) and subsequently stained with 0.1% toluidine blue. The samples were rinsed three times in autoclaved deionized water for 5 min each. The confocal microscope (Carl Zeiss 880 fast Ariyscan) was set to the basic fuchsin channel with excitation wavelength of 405 nm and an emission range of 422–504 nm, and the images were captured. For observation of vascular bundles in leaf veins, transverse sections of leaf veins were stained in 0.1% toluidine blue solution for 30 s, washed with deionized water for 10 s, and examined under an optical microscope (OLYMPUS BX51). For the microscopy analysis of metaxylem vessels, the 4th internodes of the stem aboveground were immersed in a buffer containing (5% H2O2 and 50% glacial acetic acid) at 80 °C for 12 h to isolate the mature metaxylem vessel cells. Individual vessels were gently peeled off from the vascular bundles to ensure their integrity. The smooth and intact ends of the vessels proved that no breakage occurred during the peeling process. The isolated vessels were stained overnight with 0.01% Direct Red. Samples were observed under a confocal microscope (Carl Zeiss 880 fast Airyscan), with an excitation wavelength of 561 nm and an emission range of 591–649 nm, and images were captured.
For the scanning electron microscopy (SEM) observation of vascular bundles in stem tissues, the transverse sections of the 4th internode from the base of the plant were examined using a scanning electron microscope (Hitachi TM 4000).
For the transmission electron microscopy (TEM) observation of the secondary cell wall (SCW) in metaxylem vessels, root and stem samples were fixed for 1 h at room temperature in a fixative solution containing 2.5% (v/v) glutaraldehyde and 2% (v/v) formaldehyde in 0.05 M phosphate buffer (PB; pH 6.8). The samples were then post-fixed for 1 h in 2% (w/v) osmium tetroxide in PB. Subsequently, the samples were washed twice with distilled water, dehydrated through a graded ethanol series (30%, 50%, 75%, 80%, 90%, 95%, and 100% ethanol for 30 min each), infiltrated with Spurr’s embedding medium (SPI Supplies; ERL 4221, Cat#2386-87-0; DER 736, Cat#41638-13-5; DMAE, Cat#108-01-0; NSA, Cat#28928-97-4), and then polymerized for 48 h at 60 °C. Ultrathin sections (100 nm) were prepared from the embedded samples and observed using a transmission electron microscope (Hitachi HT7800).
Dye uptake and movement assay
Seeds from different lines were germinated in soil and grown under dark conditions. After 14 days, shoots and leaves were excised from the plants. The bases of the shoots or leaf blades were then immersed in a 0.1% toluidine blue solution for 30 min under light conditions with a VPD of 1.26 kPa. Subsequently, the samples were rinsed thoroughly with distilled water to remove surface residual dye. The distance from the tissue base to the farthest point reached by the dye was measured and recorded.
Electrophoretic mobility shift assay (EMSA)
Recombinant MBP-NUT1 fusion proteins were purified from Escherichia coli (DE3) using Amylose Resin (MBP-tag, Cat#E8021S). Oligonucleotide probes were synthesized and 5′ end labeled with biotin (WT: biotin-5‘-GTATCTTGGCCCCCTAAGCAGGG-3’; Mut: biotin-5‘-GTATATAGGCCCCCTTAACAGGG-3’). EMSAs were performed according to the manufacturer’s protocol for the Light Shift Chemiluminescent EMSA Optimization and Control Kit (Thermo Fisher, Cat#89880) and the resulting bands were visualized using a Chemiluminescent imaging system (Thermo Fisher, iBright FL1500).
Chromatin Immunoprecipitation quantitative PCR (ChIP-qPCR)
The ubi:NUT1-GFP construct and empty ubi:GFP vector were individually transfected into maize leaf protoplasts. ChIP experiments were performed as previously described46 with minor modifications. Briefly, protoplasts were crosslinked in buffer containing 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 0.5 mM EDTA (pH 8.0), 0.5% NP-40, 1% formaldehyde, and freshly supplemented 1 mM PMSF, 1 mM DTT, and 1× protease inhibitor cocktail (Roche) for 25 min at room temperature. Crosslinking was quenched by adding glycine to a final concentration of 125 mM and incubating for 5 min. Isolated nuclei were washed 3–4 times with extraction buffer (20 mM Tris-HCl [pH 8.0], 2.5 mM MgCl2, 0.2% Triton X-100, 25% glycerol, 1 mM PMSF, 1 mM DTT, 1× protease inhibitor cocktail) and resuspended in 300 μL high-salt buffer (500 mM NaCl, 0.1% SDS, 1% Triton X-100, 2 mM EDTA [pH 8.0], 20 mM Tris-HCl [pH 8.0]) for 30 min on ice.
Chromatin was extracted and sheared into ~200–500 bp fragments using a Qsonica Q800R3 sonicator (80% power, 3 cycles of 20 s on / 40 s off, 18 min per cycle). The sonicated chromatin was diluted to 600 μL with 20 mM Tris-HCl (pH 8.0), and 50 μL of the mixture was reserved as input control. For immunoprecipitation, 40 μL anti-GFP agarose beads (NuoyiBio, Cat#AGA-50-1K) were added to the remaining chromatin and incubated overnight at 4 °C with gentle rotation to enrich the associated DNA fragments. The beads-chromatin complex was sequentially washed with low-salt wash buffer (150 mM NaCl, 2 mM EDTA [pH 8.0], 0.1% SDS, 1% Triton X-100, 20 mM Tris-HCl [pH 8.0]) and high-salt wash buffer (500 mM NaCl, 2 mM EDTA [pH 8.0], 0.1% SDS, 1% Triton X-100, 20 mM Tris-HCl [pH 8.0]) for 10 min each, followed by two washes with TE buffer (10 mM Tris-HCl [pH 8.0], 1 mM EDTA [pH 8.0]) for 5 min per wash.
Immunoprecipitated chromatin was eluted twice with 250 μL ChIP elution buffer (1% SDS, 0.1 M NaHCO3) on a 65 °C heating block with shaking (1000 rpm, 15 s on / 15 s off cycle) for 15 min per elution. Supernatants from the two elutions were combined, and crosslinks were reversed by incubating the eluted chromatin and input control at 65 °C for ≥8 h with 20 μL 5 M NaCl. Samples were then treated with 10 μL 0.5 M EDTA, 20 μL Tris-HCl (pH 6.5), and 20 μg Proteinase K, followed by incubation at 45 °C for 90 min to digest proteins. DNA was purified via phenol-chloroform extraction and ethanol precipitation with 40 μg glycogen (Roche) as a carrier. Purified DNA was dissolved in 100 μL ddH2O and subjected to qPCR analysis. Protoplasts transfected with the empty ubi:GFP vector served as a negative control. Relative enrichment of target DNA fragments was calculated as a percentage of the input (%). Primers used in this assay are listed in Supplementary Data 3.
Transient expression assays in maize protoplasts
For the dual-luciferase transient-expression assays, the 1-kb Exo70A1 promoter fragment and the fragment with a deletion of 50-bp NAC-binding sequence were separately constructed into the pGreenII-0800-LUC vectors at the Kpn I site upstream of the firefly luciferase (LUC) gene, generating the ZmExo70A1pro:LUC and ZmExo70A1mpro:LUC plasmids, respectively. The coding sequences of NUT1 was inserted downstream of the ZmUbi1 promoter in the pGreen II-Ubi-GFP vector. The activity of Renilla Luciferase (REN), driven by the CaMV 35S promoter in the same pGreen II-0800-LUC vector, served as an internal control for the transfection efficiency. The empty vector pGreen II-Ubi-GFP was used as a negative control. These vectors were co-transformed into maize protoplasts and incubated at 22 °C for 12 h. Subsequently, the protoplasts were centrifuged at 100 g for 5 min, and the supernatant was discarded. Luciferase signals were measured using the Dual-Luciferase Reporter Assay System (Vazyme, Cat#DL101-01) on a GLOMAX 20/20 Luminometer (Promega, Cat#E5311). Relative LUC/REN activity was calculated by normalizing LUC to REN activity.
Yeast one-hybrid assays
The full-length coding sequence of NUT1 was amplified and cloned into the pB42AD vector to generate the AD-NUT1 plasmid. A 50-bp NAC-binding sequence (including its flanking sequences) within the Exo70A1 promoter, was tandemly repeated six times and then cloned into the LacZ reporter vector (pLacZ). The two resulting plasmids were co-transformed into the yeast strain EGY48. Empty vectors were served as negative controls. Transformants were cultured at 30 °C for 8–12 h on appropriate dropout plates supplemented with X-gal (5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside) and 5% raffinose. The sequences of all primers used in this study were listed in Supplementary Data 3.
Phylogenetic tree construction
The protein sequences of the Exo70 protein family of Arabidopsis and maize were retrieved from the website https://www.arabidopsis.org/ and https://www.maizegdb.org/, respectively. Multiple sequence alignment was conducted using Clustal W. Subsequently, a phylogenetic tree was constructed based on the alignment results using the neighbor-joining method in MEGA X software.
Western blot
To check the Exo70A1-MYC protein level in Exo70A1-OE plants, total protein was extracted from the roots using 2× SDS protein loading buffer (100 mM Tris-HCl, pH 6.8, 0.2% bromophenol blue (v/v), 4% SDS (w/v), 20% glycerol (v/v), followed by boiling at 95 °C for 10 min. The samples were ultracentrifuged at 13,400 g for 10 min, and 10 μL of the supernatants were subjected to immunoblotting analysis. Detection was performed using an anti-MYC antibody (Sigma, Cat#M4439) diluted at 1:2,000. An anti-actin antibody (ABclonal, Cat#AC009) was served as a loading control at a 1:2,000 dilution. An anti-GFP antibody (ABclonal, Cat#AE012) diluted at 1:2,000 was used to detect the transiently expressed NUT1-GFP protein in maize leaf protoplasts in the dual-luciferase transient-expression assay.
Immunoprecipitation coupled mass spectrometry (IP-MS) analysis
The total proteins from transgenic plants overexpressing Exo70A1 coding sequence in a translational fusion of a GFP-tag were extracted from root stele tissues using a lysis buffer composed of 20 mM Tris-HCl pH 7.5, 150 mM NaCl, 0.5 mM EDTA pH 8.0, 0.5% NP-40, and freshly added 1 mM PMSF, 1 mM DTT, 1× protease inhibitor cocktail. After incubation for 40 min at 4 °C, the lysates were centrifuged twice at 13,400 g for 10 min at 4 °C. The cleared protein extracts were then subjected to immunoprecipitation using an GFP Nanoab Agarose beads (NuoyiBio, Cat#AGA-50-1K). The mixture was incubated for 3 h at 4 °C. Subsequently, the precipitated samples were washed 3 times with a wash buffer containing 20 mM Tris-HCl pH 7.5, 150 mM NaCl, 0.5 mM EDTA, pH 8.0, 0.5% NP-40. Finally, the precipitated samples were washed 3 times with 1× PBS (EASYBIO, Cat#BE6258). The resulting protein samples were analyzed by western blotting using an anti-GFP (ABclonal, Cat#AE012) and further processed for LC-MS/MS analysis.
Protein digestion was performed using FASP method with modifications47. Briefly, 100 μg of protein was dissolved in 50 mM ammonium bicarbonate (ABC; NH4HCO3) buffer. Proteins were reduced with dithiothreitol (DTT) at 56 °C for 45 min and alkylated with iodoacetamide (IAM) at room temperature for 30 min in the dark. The reaction mixture was transferred to a 10 kDa ultrafiltration tube (Vivacon 500, Sartorius) and centrifuged at 14,000 g for 20 min. The retentate was washed three times with 50 mM ABC buffer, and 2 μg trypsin (dissolved in 50 μL 50 mM ABC) was added for overnight digestion at 37 °C. Digested peptides were collected by centrifuging the ultrafiltration tube at 14,000 g for 20 min with a new collection tube, followed by washing the tube with additional ABC buffer to maximize peptide recovery. The combined peptide solution was diluted with 0.1% (v/v) formic acid (FA) prior to nanoLC-MS/MS analysis.
NanoLC separation was performed using a Waters nanoAcquity UPLC system (Milford, MA, USA). Peptides were first trapped on a Thermo Acclaim PepMap 100 precolumn (75 μm × 2 mm, C18, 3 μm) and then separated on a homemade analytical column (100 μm I.D. fused silica capillary; Polymicro) packed with 20 cm of C18 stationary phase (Phenomenex, Aqua 3 μm C18 125 A). A linear gradient elution was used: mobile phase B (0.1% FA in acetonitrile) was increased from 1% to 35% over 65 min, with mobile phase A as 0.1% FA in water. Nanoelectrospray ionization (nanoESI)-MS/MS analysis was conducted on a Thermo Orbitrap Fusion Lumos high-resolution mass spectrometer (Thermo Scientific) with the following parameters: full MS scan resolution of 120,000, MS/MS scan resolution of 15,000, automatic gain control (AGC) target of 4 × 10⁵ for full MS, maximum injection time of 50 ms, and a cycle time of 3 s between master scans.
Raw mass spectrometry data were processed with Mascot Distiller 2.7 for peak picking. The generated peak lists were searched against the Maize database using the Mascot 2.7 search engine. Search parameters were as follows: fixed modification (Carbamidomethyl [C]); variable modifications (Oxidation [M]); enzyme (Trypsin); maximum missed cleavages (2); MS mass tolerance (10 ppm); MS/MS mass tolerance (0.02 Da).
Water use efficiency evaluation under field conditions
Evaluation of plant biomass and grain yield under well-watered (WW) and drought conditions in the field was performed as described previously48. Different genotypes were planted in Zhangye, China (38°50′49″ N, 100°22′48″ E; 1551 m above mean sea level) in 2023 and 2024. The mean day length is 14 h, and the average temperature of approximately 21.8 °C during the growing season (May to September). Plants were cultivated in two-row / plot (3 m in length) at a standard planting density of 63,000 plants per hectare. Three replicate plots of each genotype were arranged for both WW and drought treatments. Irrigation in the WW plots was managed according to the local cultivation system. Differential watering in the drought plots commenced at 35 days after sowing, until silking was completed (Supplementary Data 2). After harvest, all ears and plants were dried, and the grain yield and aboveground biomass (excluding ears) of each plant were measured and recorded. The calculation formulas for Biomass Water Use Efficiency (Biomass WUE) and Grain Water Use Efficiency (Grain WUE) are as follows: Biomass WUE = Biomass / water volume (watering + rainfall), Grain WUE = Grain yield / water volume (watering + rainfall).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data in this study were deposited to ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org) via the iProX partner repository49,50, with the dataset identifier PXD073272. All supporting data are included the supplementary information files. A detailed reporting summary and. Source data are provided with this paper.
References
Vadez, V. et al. Crop traits and production under drought. Nat. Rev. Earth Environ. 5, 211–225 (2024).
UNESCO. The United Nations World Water Development Report 2024: Water for Prosperity and Peace (UNESCO, Paris, 2024).
Lobell, D. B. et al. Greater sensitivity to drought accompanies maize yield increase in the U.S. Midwest. Science 344, 516–519 (2014).
Taiz, L. et al. Plant Physiology and Development (Oxford Univ. Press, 2023).
Scharwies, J. D. & Dinneny, J. R. Water transport, perception, and response in plants. J. Plant Res. 132, 311–324 (2019).
Venturas, M. D. et al. Plant xylem hydraulics: what we understand, current research, and future challenges. J. Integr. Plant Biol. 59, 356–389 (2017).
Fukuda, H. Tracheary element differentiation. Plant Cell 9, 1147–1156 (1997).
Turner, S. et al. Tracheary element differentiation. Annu. Rev. Plant Biol. 58, 407–433 (2007).
Furuta, K. M. et al. Molecular control of cell specification and cell differentiation during procambial development. Annu. Rev. Plant Biol. 65, 607–638 (2014).
Yamaguchi, M. et al. Vascular-related nac-domain7 directly regulates the expression of a broad range of genes for xylem vessel formation. Plant J. 66, 579–590 (2011).
Karam, G. N. Biomechanical model of the xylem vessels in vascular plants. Ann. Bot. 95, 1179–1186 (2005).
Huang, S. et al. Tubulin participates in establishing protoxylem vessel reinforcement patterns and hydraulic conductivity in maize. Plant Physiol. 196, 931–947 (2024).
McFarlane, H. E. et al. The cell biology of cellulose synthesis. Annu. Rev. Plant Biol. 65, 69–94 (2014).
Meents, M. et al. The cell biology of secondary cell wall biosynthesis. Ann. Bot. 121, 1107–1125 (2018).
Vanholme, R. et al. Lignin biosynthesis and structure. Plant Physiol. 153, 895–905 (2010).
Groover, A. & Jones, A. M. Tracheary element differentiation uses a novel mechanism coordinating programmed cell death and secondary cell wall synthesis. Plant Physiol. 119, 375–384 (1999).
Ménard, D. & Pesquet, E. Cellular interactions during tracheary elements formation and function. Curr. Opin. Plant Biol. 23, 109–115 (2015).
Wang, H. et al. Xylan-based nanocompartments orchestrate plant vessel wall patterning. Nat. Plants 8, 295–306 (2022).
Kubo, M. et al. Transcription switches for protoxylem and metaxylem vessel formation. Genes Dev. 19, 1855–1860 (2005).
Zhou, J. et al. Arabidopsis NAC domain proteins, VND1 to VND5, are transcriptional regulators of secondary wall biosynthesis in vessels. PLoS One 9, e105726 (2014).
Zhong, R. & Ye, Z. Secondary cell walls: biosynthesis, patterned deposition and transcriptional regulation. Plant Cell Physiol. 56, 195–214 (2015).
Zhao, C. et al. XND1, a member of the NAC domain family in Arabidopsis thaliana, negatively regulates lignocellulose synthesis and programmed cell death in xylem. Plant J. 53, 425–436 (2008).
Yamaguchi, M. et al. VND-INTERACTING2, a NAC domain transcription factor, negatively regulates xylem vessel formation in Arabidopsis. Plant Cell 22, 1249–1263 (2010).
Tang, N. et al. Natural variation at XND1 impacts root hydraulics and trade-off for stress responses in Arabidopsis. Nat. Commun. 9, 3884 (2018).
Ding, B. et al. The transcription factors DOF4.6 and XND1 jointly regulate root hydraulics and drought responses in Arabidopsis. Plant Cell 37, koaf083 (2025).
Abe, A. et al. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat. Biotechnol. 30, 174–178 (2012).
Lu, X. et al. Gene-indexed mutations in maize. Mol. Plant 11, 496–504 (2018).
Mei, K. et al. Cryo-EM structure of the exocyst complex. Nat. Struct. Mol. Biol. 25, 139–146 (2018).
Synek, L. et al. Plasma membrane phospholipid signature recruits the plant exocyst complex via the EXO70A1 subunit. Proc. Natl. Acad. Sci. USA 118, e2105287118 (2021).
Kalmbach, L. et al. Transient cell-specific EXO70A1 activity in the CASP domain and Casparian strip localization. Nat. Plants 3, 17058 (2017).
Li, S. et al. EXO70A1-mediated vesicle trafficking is critical for tracheary element development in Arabidopsis. Plant Cell 25, 1774–1786 (2013).
Dong, Z. et al. Necrotic upper tips1 mimics heat and drought stress and encodes a protoxylem-specific transcription factor in maize. Proc. Natl. Acad. Sci. USA 117, 20908–20919 (2020).
Turco, G. M. et al. Molecular mechanisms driving switch behavior in xylem cell differentiation. Cell Rep. 28, 342–351 (2019).
Vukašinović, N. et al. Microtubule-dependent targeting of the exocyst complex is necessary for xylem development in Arabidopsis. N. Phytol. 213, 1052–1067 (2016).
He, B. et al. Exo70 interacts with phospholipids and mediates the targeting of the exocyst to the plasma membrane. EMBO J. 26, 4053–4065 (2007).
Zhu, J. et al. OsSEC3A, a key exocyst subunit, is required for the emergence of crown root in rice. N. Phytol. 249, 917–929 (2026).
De Bellis, D. et al. Extracellular vesiculo-tubular structures associated with suberin deposition in plant cell walls. Nat. Commun. 13, 1489 (2022).
Jiang, S. et al. Preassembly and independent trafficking of the exocyst complex in Arabidopsis. Proc. Natl. Acad. Sci. USA 122, e2519318122 (2025).
Drdová, E. J. et al. The exocyst complex contributes to PIN auxin efflux carrier recycling and polar auxin transport in Arabidopsis. Plant J. 73, 709–719 (2013).
Miodek, A. et al. What do we know about growth of vessel elements of secondary xylem in woody plants?. Biol. Rev. Camb. Philos. Soc. 96, 2911–2924 (2021).
Xing, H. et al. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 14, 327 (2014).
Ishida, Y. et al. Agrobacterium-mediated transformation of maize. Nat. Protoc. 2, 1614–1621 (2007).
Wang, Y. et al. A dirigent family protein confers variation of Casparian strip thickness and salt tolerance in maize. Nat. Commun. 13, 2222 (2022).
Munns, R. et al. Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat. Biotechnol. 30, 360–364 (2012).
Mees, G. C. & Weatherley, P. E. The mechanism of water absorption by roots. I. Preliminary studies on the effect of hydrostatic pressure gradients. Proc. R. Soc. Lond. B 147, 367–380 (1957).
Gendrel, A. V. et al. Profiling histone modification patterns in plants using genomic tiling microarrays. Nat. Methods 2, 213–218 (2005).
Wiśniewski, J. et al. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359–362 (2009).
Gao, H. et al. Natural variations of ZmSRO1d modulate the trade-off between drought resistance and yield by affecting ZmRBOHC-mediated stomatal ROS production in maize. Mol. Plant 15, 1558–1574 (2022).
Ma, J. et al. iProX: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2021).
Acknowledgements
We thank the great technical support in generating and propagating the transgenic maize provided by the staff in the Center of Crop Functional Genomics and Molecular Breeding, China Agricultural University (CAU). We appreciate the assistance from the electron microscopy Laboratory and the biological mass spectrometry laboratory at CAU for their expertise in electron microscopy imaging and mass spectrometry analysis, respectively. We thank Prof. Xiaoduo Lu and Prof. Chunyi Zhang from Qilu Normal University for providing the EMS mutants used in this study. We thank Dr. Chuanyu Wang and Dr. Ying Zhang at Beijing Academy of Agriculture and Forestry Sciences for their help with xylem vascular imaging. We also thank Dr. Yulong Ren from Institute of Crop Science, Chinese Academy of Agricultural Sciences, for the assistance with transmission electronic microscopy. This research was supported by the National Key Research and Development Program of China (2022YFF1001602), the National Natural Science Foundation of China (32430010, 32301755, 32272024, 32171940), Chinese Postdoctoral Science Foundation (2023M733810, GZB20230854) and Chinese Universities Scientific Fund (2025TC135, 2025TC148).
Author information
Authors and Affiliations
Contributions
F.Q. designed and supervised the study and revised the manuscript. Z.Y. provided important advice for the work. T.Z. and Yanyan W. analyzed the data, performed the experiments and drafted the manuscript. T.Z. and C.W. constructed the EMS mutant library. Yijie W. performed the Mutmap analysis. Yu L. and J.H. provided assistance on ChIP-qPCR experiment and DAP-seq analysis. B.Z., S.L., Yongyan L. and L.W. provided suggestions and assistance for the field trials. J.C. generated the transgenic maize. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Huanzhong Wang and the other anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhu, T., Wang, Y., Wang, Y. et al. NUT1-Exo70A1 Regulates Xylem Vessel Development and Influences Water Use Efficiency in Maize. Nat Commun 17, 2816 (2026). https://doi.org/10.1038/s41467-026-69436-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69436-7