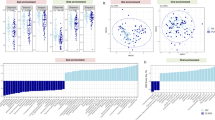
Abstract
Late presentation of HIV-1 infection is linked to gut dysbiosis, impaired immune reconstitution, excess inflammation, immune activation, and increased morbidity and mortality. It is unclear if antiretroviral therapy initiation can reverse HIV-associated gut dysbiosis at all, or if specific antiretroviral regimens are more effective in restoring the gut microbiota than others. This has important implications for the long-term health of individuals with HIV. In this multicenter, open-label, randomized clinical trial (NCT02337322), 88 antiretroviral-naïve individuals with advanced HIV-1 infection (median CD4+ T cells of 34 cells/mm3) were randomized (1:1) to initiate lamivudine/abacavir plus either dolutegravir or ritonavir-boosted darunavir, and were followed for 2 years. Both groups had similar HIV-1 suppression rates and recovery of CD4+ T cells. However, treatment with dolutegravir led to increased gut microbial richness and diversity and enrichment of specific microbial taxa and metabolic pathways. These changes were associated with reduced inflammation and lower immune activation, outcomes that did not occur with darunavir/ritonavir. After two years, participants on dolutegravir-based therapy had gut microbiota profiles more closely resembling those of people without HIV, compared to individuals taking darunavir/ritonavir. In summary, dolutegravir-based therapy restores the gut microbiota more effectively than darunavir/ritonavir in patients who present late with HIV.
Similar content being viewed by others
Introduction
Nearly half of all new diagnoses of human immunodeficiency virus type 1 (HIV-1) infection occur in individuals with advanced disease and severe immunological deterioration, i.e., CD4+ T cell counts <350 cells/mm31,2. Initiating antiretroviral therapy as early as possible is crucial for improving prognosis3,4. However, often this only results in partial immune reconstitution5,6. Late presenters remain at high risk of opportunistic infections, malignancies, and death5,7,8, even years after starting antiretroviral therapy3,4. It is thus essential to implement measures for the early diagnosis and treatment of HIV-1 infections, identify optimal antiretroviral treatment options for late presenters, and uncover modifiable factors that may help mitigate the comorbidity gap associated with late presentation. Persistent gut dysbiosis is commonly found in individuals with low nadir CD4+ T cell counts9, and may partially account for such a gap.
The gut-associated lymphoid tissue (GALT) plays a critical role in human health, as well as in HIV pathogenesis10. It houses the majority of CD4+ T cells in the body, serves as the largest site for HIV-1 replication, and is essential for maintaining intestinal barrier integrity11,12,13. A diverse and rich gut microbiome is crucial for human health14,15,16,17, as commensal microbes help preserve mucosal integrity18, modulate immune homeostasis16, and promote enterocyte health19,20. Disruption of enterocytes and GALT during acute HIV-1 infection leads to intestinal and systemic inflammation, immune dysregulation, and the translocation of bacterial products into the bloodstream, which exacerbates chronic systemic inflammation21. Chronic gut inflammation and immune dysregulation result in decreased microbial richness, an increase in generalist microbes better adapted to oxidative stress, and a loss of highly specialized oxygen-sensitive species9,22,23.
The initiation of antiretroviral treatment generally enhances immune reconstitution. However, its effect on restoring gut dysbiosis remains uncertain. Additionally, it is not clear if certain antiretroviral regimens with superior intestinal penetration, such as HIV integrase strand-transfer inhibitors, are more effective in restoring gut microbiota compared to those with lower intestinal penetration, like protease inhibitors12,24,25. This has significant implications for the long-term well-being of individuals with HIV, as a rich and diverse gut microbiota is one of the key indicators of better overall health26. Here, we examined these questions in a 2-year, two-arm randomized clinical trial comparing first-line antiretroviral therapy with dolutegravir versus ritonavir-boosted darunavir/ritonavir in subjects presenting with advanced-stage HIV. The primary and most secondary outcomes of this trial have been previously published27; here, we report a pre-specified secondary outcome focusing on the longitudinal restoration of gut microbiota composition and function.
Results
Similar immune reconstitution, but greater reductions in inflammation and immune activation with dolutegravir—than with darunavir/ritonavir—including ART
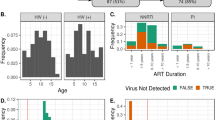
Eighty-eight ART-naïve participants with baseline CD4⁺ T cell counts below 100 cells/mm³ (median, IQR: 34, 12–67) were randomized 1:1 to receive either darunavir/ritonavir or dolutegravir, each in combination with lamivudine and abacavir (Fig. 1). Both study groups were well balanced regarding demographic factors, body mass index, soluble immune activation markers, cytokines, and microbial gene richness at study entry (Table 1). Fecal specimens were collected at baseline (week 0) and at weeks 24, 48, and 96. Following DNA extraction and whole-genome shotgun metagenomic sequencing, samples failing DNA quantity requirements or predefined sequencing quality control thresholds were excluded. Metagenomic analyses were therefore restricted to participants with either a high-quality baseline sample or at least two high-quality follow-up samples available. All metagenomic analyses were conducted in the modified intention-to-treat population (mITT2) (Figs. 1 and 2A).
This diagram shows the patient enrollment, allocation, and final analysis sets. Out of 149 assessed individuals, 104 were randomized into two treatment arms: DTG (n = 52) and DRV/r (n = 52). The panels detail the number of participants excluded before randomization, those who did not start treatment, and the specific reasons why 13 patients were not evaluable for the final analysis (such as loss to follow-up, virologic failure, or death). DTG dolutegravir, DRV/r darunavir/ritonavir, mITT modified intention-to-treat.
A Schematic representation of microbiome study design. B Longitudinal analyses of α-diversity (Shannon, observed richness, Gini dominance, gene richness) using linear mixed-effects models (metric ~ treatment * time_point + covariates + (1 | participant_id)). Top: within-arm changes vs baseline; bottom: between-arm contrasts (dolutegravir − darunavir/ritonavir) at each time point. In all panels, symbols estimated marginal means and error bars denote 95% confidence intervals; multiple testing was controlled with Benjamini–Hochberg FDR. Statistics are based on n = 88 independent biological replicates (participants: n = 46 DTG, n = 42 DRV/r) and N = 257 longitudinal observations. C Repeated-measures correlations (rmcorr) between α-diversity metrics and clinical/immunologic markers (CD4, CD8, activated T cell subsets, CRP, IL-6, TNF-α, sCD14, BMI), shown separately for darunavir/ritonavir and dolutegravir. Colors encode correlation direction and magnitude; outlined cells indicate significant associations (q < 0.05). D Representative rmcorr plots for selected associations (e.g., CD4 vs gene richness, BMI vs gene richness, CRP vs gene richness, BMI vs observed richness). Points are colored by treatment (darunavir/ritonavir, dolutegravir); lines depict participant-level fitted trends. Each panel reports the correlation coefficient (r), q value, and sample size (n). Source data are provided as a Source data file.
Both treatment arms achieved similar immune reconstitution and viral suppression rates over 96 weeks. CD4⁺ T-cell counts increased from baseline (Δ = +4.66-fold, 95% CI = 3.44–5.88, q value < 1 × 10−¹⁵ in darunavir/ritonavir; Δ = 3.33-fold, 95% CI = 2.23–4.43, q value = 2.5 × 10⁻¹¹ in dolutegravir). Alongside these improvements, there were also significant CD8⁺ T-cell increases. Markers of immune activation and inflammation, including TNF-α, IL-6, and sCD14, decreased in both groups (all q value < 0.001). Participants on dolutegravir had a greater reduction in sCD14 at week 96 (Δ = −1.92-fold, 95% CI = −2.12 to −1.73) than those on darunavir/ritonavir (Δ = −1.63-fold, 95% CI = −1.83 to −1.43), with a between-group difference of −0.36-fold (95% CI = −0.62 to −0.10; q value = 0.015). CRP showed a similar pattern (dolutegravir Δ = −1.67-fold, 95% CI = −2.46 to −0.88; darunavir/ritonavir Δ = −0.43-fold, 95% CI = −1.26 to 0.40), but the between-group difference was not statistically significant after false discovery rate (FDR) adjustment (q value = 0.144). Body mass index increased slightly and similarly in both groups at week 96 (Δ = +0.16-fold; 95% CI = 0.89 to 2.261, q value < 0.001) (Supplementary Figs. 1 and 2).
Dolutegravir initiation leads to sustained improvements in microbial gene diversity and overall alpha diversity
Microbial Shannon diversity, observed microbial richness, Gini dominance, and gene richness were assessed next (Fig. 2B). Subjects initiating dolutegravir showed an increase in gene richness by week 48 relative to baseline (Δ = 0.27-fold, 95% CI = 0.06–0.49; q value = 0.004), which was sustained through week 96 (Δ = 0.29-fold, 95% CI = 0.07–0.51; q value = 0.004). Shannon diversity showed a smaller increase, which reached significance at week 96 (Δ = 0.13-fold, 95% CI = 0.01–0.24; q value = 0.025). Gini dominance decreased over time in dolutegravir at week 48 (Δ = −0.007-fold, 95% CI = −0.013 to −0.001; q value = 0.005) and week 96 (Δ = −0.009-fold, 95% CI = −0.015 to −0.003; q value < 0.001). In contrast, no temporal changes reached significance in the darunavir/ritonavir arm (all q value > 0.300). Between-group comparisons (dolutegravir vs darunavir/ritonavir) at weeks 48–96 were directionally positive for gene and observed richness but did not meet the study-wide adjusted threshold (gene richness q value ≈ 0.089; observed richness q value ≈ 0.105).
We then quantified within-arm consistency of longitudinal trajectories. For each participant, we estimated a subject-level slope of change from baseline (Δ per visit) and compared the dispersion of slopes across arms. Across alpha-diversity metrics, subjects on dolutegravir showed lower within-arm dispersion than those receiving darunavir/ritonavir (e.g., gene richness SDdolutegravir = 0.006 vs SDdarunavir/ritonavir = 0.008; Shannon diversity SDdolutegravir = 0.003 vs SDdarunavir/ritonavir = 0.004). Variance-ratio estimates (dolutegravir/darunavir/ritonavir) were consistently <1—0.495 (95% CI = 0.141–3.040) for gene richness, 0.536 (95% CI = 0.220–2.210) for Shannon diversity, 0.905 (95% CI = 0.252–2.970) for observed richness, and 0.719 (95% CI = 0.303–1.990)—indicating more homogeneous changes within the dolutegravir arm. Robust tests of equality of variances (Brown–Forsythe and Fligner–Killeen) were not significant after FDR correction (q value ≥ 0.317), consistent with limited power, but the directionally consistent pattern supports that mean changes are more readily detected under dolutegravir owing to reduced within-arm heterogeneity (Supplementary Fig. S3A).
Higher microbial richness associates with greater CD4⁺ counts and lower inflammation in dolutegravir recipients
To explore host–microbiome interactions over time, we then computed repeated-measure correlations between alpha diversity metrics and markers of systemic inflammation, immune activation, and bacterial translocation (Fig. 2C, D). Microbial gene richness correlated positively with CD4⁺ T cell counts (r = 0.44, q value = 0.004) and body mass index (r = 0.35, q value = 0.047), and inversely with plasma CRP levels (r = −0.43, q value = 0.047) in the dolutegravir arm. Observed richness also showed a positive association with body mass index (r = 0.37, q value = 0.023). These associations suggest that higher microbial richness is linked to improved immune reconstitution and lower systemic inflammation in dolutegravir recipients. No significant correlations were detected in the darunavir/ritonavir arm.
Dolutegravir induces progressive gut microbiota homogenization over 96 weeks
Between-sample beta diversity was first explored by non-metric multidimensional scaling (NMDS) of Bray–Curtis distances (Fig. 3A). Samples were clustered by treatment arm, with 95% confidence ellipses shown for each group and time point. To quantify the contribution of clinical covariates to the overall community structure, we performed a PERMANOVA analysis (Fig. 3B). Treatment arm influenced community structure (R² = 0.013, p value = 0.001), with a magnitude of effect similar to that of having an AIDS-defining event (R² = 0.013, p value = 0.001) or cytomegalovirus (CMV) seronegativity (R² = 0.006, p value = 0.001). Time points also explained a small but significant portion of the variance (R² = 0.014, p value = 0.003), while age, gender, HIV-1 load, and other covariates were not significant contributors (all p value > 0.100). This indicates that, although each of these factors influences the gut microbiota composition, none of them dominates on its own. Assessment of the taxonomic structure of the top 25 most abundant genera further showed that, as previously described23,28, the microbiome of men who had sex with men (MSM) was enriched in Prevotella relative to Bacteroides, independent of ART regimen and other factors (Supplementary Fig. 3A).
A NMDS ordination based on Bray–Curtis dissimilarities of relative-abundance profiles, showing individual samples colored by treatment and grouped by study time point. Ellipses denote 95% confidence intervals for each treatment–time point combination. B Results of the PERMANOVA model. Statistical significance was determined using two-sided permutation tests (999 permutations) constrained by participant. The table reports degrees of freedom, pseudo-F statistics, effect size (R²), and exact permutation-based p values. C Longitudinal changes in within-group dispersion, expressed as the Euclidean distance of each sample to its treatment–time point centroid in NMDS space. Left: model-based estimates of within-arm changes vs baseline; right: between-treatment contrasts (dolutegravir − darunavir/ritonavir) at each time point, both derived from linear mixed-effects models. Symbols denote estimated marginal means, and error bars denote 95% confidence intervals. Multiple testing was controlled with the Benjamini–Hochberg. Statistics are based on n = 88 independent biological replicates (participants: n = 46 DTG, n = 42 DRV/r) and N = 257 longitudinal observations. D Mean Bray–Curtis dissimilarity to HIV-negative gut microbiomes for MetaHIV and ADVANZ-4 samples, stratified by treatment and time point. In boxplots, the center line denotes the median; box limits denote the 25-75th percentiles; whiskers extend to 1.5x the IQR. Individual points represent independent biological samples. E Estimated differences in dissimilarity to HIV-negative microbiomes relative to the HIV-negative reference group, derived from linear mixed-effects models with FDR correction (points = estimate; bars = 95% confidence intervals). For both (D, E), statistics are based on independent biological replicates from the ADVANZ-4 trial (n = 88 participants; n = 46 DTG, n = 42 DRV/r) and the metaHIV datasets (n = 156 subjects). F Repeated-measures correlations between NMDS centroid distance and host clinical or immunologic markers, shown separately for treatment group. Colors indicate correlation direction and magnitude; black outlines denote significant associations (q value < 0.05). G Representative rmcorr plots illustrating significant associations. Points are colored by treatment; lines depict participant-level fitted trends. Source data are provided as a Source data file.
Moreover, samples from dolutegravir-treated subjects progressively migrated toward a common region in the ordination space over the 96 weeks of follow-up—reflected by increasingly tight confidence ellipses—whereas samples from subjects receiving darunavir/ritonavir remained broadly dispersed. To determine whether this visual convergence represented a real reduction in community heterogeneity, we calculated for each sample the Euclidean distance to the centroid of its treatment- and time-specific group in the NMDS space (Fig. 3C). In the dolutegravir arm, centroid distances decreased markedly from baseline at week 24 (Δ = −0.35-fold, 95% CI = −0.59 to −0.11; q value = 0.001), week 48 (Δ = −0.40-fold, 95% CI = −0.62 to −0.17; q value < 0.001) and week 96 (Δ = −0.47-fold, 95% CI = −0.69 to −0.24; q value < 0.001), indicating progressive microbiota homogenization over time. In contrast, no significant temporal changes were detected in the darunavir/ritonavir arm (all q value > 0.800). Between-group comparisons confirmed that dolutegravir recipients exhibited lower centroid distances than darunavir/ritonavir at weeks 48 (Δ = −0.31-fold, 95% CI = −0.56 to −0.06; q value = 0.030) and 96 (Δ = −0.35-fold, 95% CI = −0.61 to −0.10; q value = 0.027), demonstrating a greater, sustained, and statistically robust reduction in microbial community dispersion under dolutegravir relative to darunavir/ritonavir.
To assess whether the longitudinal convergence observed under dolutegravir represented a shift toward a healthier gut microbiota configuration, we integrated our dataset with the metaHIV cohort9,28, which includes fecal microbiomes from HIV-negative individuals and several clinical phenotypes of people with HIV (Fig. 3D, E). We computed each sample’s mean Bray–Curtis dissimilarity to the HIV-negative reference group. Both treatment arms remained significantly distinct from HIV-negative controls throughout follow-up (all q values < 0.001). However, dolutegravir-treated participants exhibited a consistent reduction in dissimilarity over time (Δ ≈ −0.05-fold between weeks 0–96), resulting in lower distances than those of darunavir/ritonavir at week 96. Mixed-effects modeling confirmed that, while all HIV-positive profiles—including darunavir/ritonavir and metaHIV subgroups such as concordant, discordant, and elite controllers—differed significantly from HIV-negative individuals (Δ = 0.08–0.14; q value < 0.001), dolutegravir samples at week 96 displayed the smallest deviation (Δ = 0.08 ± 0.02; q value = 8.6 × 10⁻⁵). These results indicate that dolutegravir-associated microbiota homogenization after 2 years of treatment partially restores the gut microbial community composition towards that of HIV-uninfected individuals, although full normalization is not achieved and might require longer treatment exposure.
To explore the functional implications of dolutegravir-specific homogenization, we assessed repeated-measures correlations between individual centroid distances and systemic immune or inflammatory markers (Fig. 3F, G and Supplementary Fig. 3B, C). In dolutegravir-treated participants, shorter centroid distances were strongly associated with higher CD4⁺ T cell counts (r = −0.48; q value = 1.8 × 10⁻⁵) and BMI (r = −0.53; q value = 2.5 × 10⁻⁶), and inversely correlated with markers of immune activation and inflammation including CD4⁺CD38⁺HLA-DR⁺ T cells (r = 0.50; q value = 0.018), TNF-α (r = 0.40; q value = 0.005), sCD14 (r = 0.38; q value = 0.008) and CRP (r = 0.35; q value = 0.023). No significant associations were observed in the darunavir/ritonavir arm (all q value > 0.180). This suggests that the extent of gut microbiota convergence under dolutegravir is associated with enhanced immune reconstitution and lower systemic inflammation, which aligns with a partial restoration of host–microbiome homeostasis.
Dolutegravir-enriched microbial taxa correlate with immunologic and inflammatory profiles
Next, we sought to understand which specific microbial taxa were involved in the observed variations in gut microbiome ecology and their associations with systemic inflammation and immune activation. A differential abundance analysis at multiple taxonomic levels using ANCOMBC2, and implementing a treatment * time_point interaction model that adjusted for baseline covariates and included patient identifier as a random effect to account for repeated measurements. Although no taxa reached study-wide significance after FDR correction under this stricter longitudinal framework, several genera displayed consistent effect sizes and narrow confidence intervals, suggesting biologically meaningful shifts between treatment arms.
Several dolutegravir-enriched species and genus showed 95% CIs excluding zero at specific visits (Fig. 4A and Supplementary Fig. 5A), including Methanobrevibacter smithii (week 48: log₂FC = 2.35; 95% CI = 0.55–4.14), Parasutterella excrementihominis (week 48: log₂FC = 3.63; 95% CI = 0.94–6.32), Olsenella scatoligenes (week 48: log₂FC = 1.68; 95% CI = 0.23–3.13) and Mogibacterium diversum (week 48: log₂FC = 2.49; 95% CI = 0.58–4.40) (Fig. 4A). Conversely, several Bacteroides-associated species were consistently depleted, notably Bacteroides thetaiotaomicron (week 24: log₂FC = −3.51; 95% CI = −5.67 to −1.36; week 48: log₂FC = −1.99; 95% CI = −3.91 to −0.08; week 96: log₂FC = −2.15; 95% CI = −4.00 to −0.30) and Parabacteroides distasonis (week 24: −1.65; 95% CI = −3.03 to −0.28; week 48: −1.57; 95% CI = −2.92 to −0.23).
A Species-level differential abundance between dolutegravir and darunavir/ritonavir arms across study time points (0, 24, 48, and 96 weeks), estimated with ANCOMBC2 (abundance ~ treatment * time_point + covariates + (1 | participant_id)). Points represent estimated log₂ fold changes (dolutegravir − darunavir/ritonavir) and error bars denote 95% confidence intervals. Genera with q value < 0.05 and confidence intervals excluding zero are highlighted; the accompanying heatmap shows per-time-point log₂ fold changes and taxon prevalence. Statistics are based on n = 88 independent biological replicates (participants: n = 46 DTG, n = 42 DRV/r) and N = 257 longitudinal observations. B Repeated-measures correlations (rmcorr) between CLR-transformed species abundances and host clinical/immunologic markers (CD4, CD8, activated T cell subsets, CRP, IL-6, TNF-α, sCD14, BMI), shown separately for darunavir/ritonavir and dolutegravir. Colors indicate correlation direction and magnitude; black outlines denote significant associations (q value < 0.05). Source data are provided as a Source data file.
Although multiple-testing correction in this repeated-measures setting is conservative, the direction of effects is consistent across time, pointing to a reconfiguration characterized by the enrichment of fermentative lineages—including lactate- and succinate-producing genera such as Olsenella, Prevotella, and Parasutterella—alongside the methanogenic archaeon Methanobrevibacter, which can facilitate hydrogen disposal and sustain anaerobic fermentation; in parallel, we observe a relative depletion of Bacteroides-dominated profiles and reductions in Lachnoclostridium/Erysipelatoclostridium. This community shift is compatible with increased cross-feeding toward SCFA production and a more even, less dominance-driven structure, which aligns with the longitudinal increases in richness/evenness and the inverse associations with systemic inflammation.
To explore potential host–microbiome interactions, we next assessed the association between longitudinal changes in microbial species and systemic immunologic and inflammatory markers using repeated-measures correlation (Fig. 4B and Supplementary Fig. 5B). Despite modest overall effect sizes, several taxa exhibited consistent and biologically coherent relationships within treatment arms. Within the dolutegravir arm, Methanobrevibacter smithii abundance was positively correlated with BMI (r = 0.32; q value = 0.017), while Parasutterella excrementihominis showed an inverse association with plasma sCD14 levels (r = –0.42; q value = 0.012), consistent with its previously reported anti-inflammatory profile29,30,31. Conversely, Lactobacillus sakei (r = –0.31; q value = 0.022) and Clostridium aldenense (r = –0.32; q value = 0.017) both exhibited negative correlations with BMI, and Olsenella scatoligenes correlated positively with BMI (r = 0.30; q value = 0.022) but inversely with CRP (r = –0.39; q value = 0.046), suggesting context-dependent metabolic and inflammatory associations. Alistipes indistinctus negatively correlated with activated CD8⁺CD38⁺HLA-DR⁺ T cells (r = –0.59; q value = 0.014), indicating a potential link between this taxon and reduced T cell activation. In contrast, under darunavir/ritonavir, only Parabacteroides distasonis displayed significant correlations, positively associated with BMI (r = 0.34; q value = 0.039). Together, these findings indicate that the immunometabolic landscape accompanying dolutegravir initiation is intertwined with species-specific microbial trajectories, particularly involving hydrogenotrophic archaea and commensals with fermentative or anti-inflammatory capacities.
Dolutegravir remodels gut microbial co-occurrence networks and increases local motif cohesion by week 96
At week 96, co-occurrence networks revealed clear topological differences by ART regimen (Fig. 5A). In participants treated with dolutegravir, the network divided into 32 connected components, whereas under darunavir/ritonavir treatment, it divided into 55 components. The largest connected component (LCC) included 53% of taxa in the dolutegravir group compared to 15% in the darunavir/ritonavir group, indicating a more consolidated core in the former. Global network metrics further highlight this increased cohesion: dolutegravir networks demonstrated a non-zero clustering coefficient (0.127 compared to 0.000), slightly greater density (0.0191 compared to 0.0076), and marginally greater natural connectivity (0.0151 compared to 0.0145). Eigenvector-based hub analysis at week 96 identified Bacteroides, Gordonibacter, Lachnoclostridium, and Parabacteroides as central nodes in the dolutegravir network, whereas darunavir/ritonavir hubs included Catenibacterium, Eggerthella, Erysipelatoclostridium, and Holdemanella, with fewer cross-phyla links. Together, these measures indicate that long-term dolutegravir treatment is associated with a more integrated and cohesive network structure.
A Side-by-side SPRING-derived genus-level networks for patients treated with dolutegravir (left) versus darunavir/ritonavir (right) at week 96. Nodes represent genera (colored by phylum), and node size is proportional to CLR-transformed relative abundance. Only the 200 most prevalent genera (prevalence ≥15%) are shown, with edges weighted by signed association strength. Layouts are held constant between panels for direct visual comparison; isolated singletons present in both networks have been removed. B Heatmap of the differential Graphlet Correlation Matrix (ΔGCM = GCM_dolutegravir – GCM_darunavir/ritonavir) computed on the largest connected component at week 96. Each cell displays the change in Spearman correlation between a pair of non-redundant graphlet orbit counts; red shades indicate orbit pairs more positively correlated in dolutegravir; blue shades indicate stronger correlations in darunavir/ritonavir. Numerical labels give Δρ, and asterisks denote FDR-adjusted significance of the difference (*q value < 0.05; **q value < 0.01; ***q value < 0.001). Bounding boxes group orbits by their topological role (degree, chain, cycle, terminal). See Supplementary Table 4 for a detailed description of the graphlet orbits. Source data are provided as a Source data file.
To better understand how local structure in the network is organized, we performed a graphlet correlation analysis, which examines how often small patterns of connections—called graphlets—appear together (Fig. 5B). This approach allowed us to detect two distinct types of structural motifs in the dolutegravir network. One group of motifs was made up of linear, chain-like patterns, while the other formed more circular, loop-like structures. These motifs tended to co-occur consistently in the dolutegravir network, as shown by strong correlations between their frequencies (Spearman |ρ| > 0.8, p < 0.01). In contrast, the darunavir/ritonavir network also exhibited several strong motif–motif correlations, but these were fewer in number, and the overall correlation structure was more dispersed, indicating a less coherent local organization. Quantitative comparison of graphlet correlation distances confirmed that the week 96 dolutegravir network diverged significantly from both baseline and darunavir/ritonavir architectures, supporting a more pronounced structural reorganization under dolutegravir. This demonstrates that 96 weeks of dolutegravir not only increased global connectivity but also promoted the emergence of more stable and internally coherent higher-order interaction motifs. Network and graphlet analysis for weeks 0, 24, and 48 showed intermediate connectivity and motif organization that progressively converged toward the structures observed at week 96 (Supplementary Fig. 5A, B).
Dolutegravir-specific remodeling of metabolic and biosynthetic pathways is linked to host immune and inflammatory markers
Applying ANCOMBC2 to MetaCyc pathways with the same treatment * time_point model, including a random intercept for participant, no pathway met the pre-specified multiple-testing criteria after adding random effects (q value < 0.05; prevalence >10%). However, several functions showed visit-specific effects whose 95% CIs excluded zero (Fig. 6A). At Week 24, dolutegravir was associated with higher relative abundances of central-carbon and biosynthetic routes, including the incomplete reductive TCA cycle (P42-PWY; log₂FC = 0.54; 95% CI = 0.03–1.06), phospholipid biosynthesis I (PHOSLIPSYN-PWY; log₂FC = 0.52; 0.05–1.00), tetrapyrrole biosynthesis II (PWY-5189; log₂FC = 0.97; 95% CI = 0.36–1.57), TCA cycle II (PWY-5690; log₂FC = 0.50; 95% CI = 0.09–0.91), and ubiquinol-8 biosynthesis (UBISYN-PWY; log₂FC = 1.36; 95% CI = 0.24–2.48). In the reverse comparison (darunavir/ritonavir > dolutegravir), we noted increased activity in the urea cycle (PWY-4984; log₂FC = –0.89; 95% CI = –1.58 to –0.19), chondroitin sulfate degradation I (PWY-6572; log₂FC = –1.48; 95% CI = –2.53 to –0.43), acetyl-CoA fermentation to butanoate II (PWY-5676; log₂FC = –0.48; 95% CI = –0.94 to –0.01), and 4-hydroxybenzoate biosynthesis V (PWY-6435; log₂FC = –1.44; 95% CI = –2.59 to –0.30).
A Differential abundance of MetaCyc pathways between dolutegravir and darunavir/ritonavir treatment arms across study time points (0, 24, 48, and 96 weeks), estimated with ANCOMBC2 (abundance ~ treatment * time_point + covariates + (1 | participant_id), prevalence ≥10%). Points represent estimated log₂ fold changes (dolutegravir − darunavir/ritonavir) and error bars denote 95% confidence intervals. Pathways with q value < 0.05 and confidence intervals excluding zero are highlighted. The adjacent heatmap displays per-time-point log₂ fold changes (dolutegravir − darunavir/ritonavir) and pathway prevalence. Statistics are based on n = 88 independent biological replicates (participants: n = 46 DTG, n = 42 DRV/r) and N = 257 longitudinal observations. B Repeated-measures correlations (rmcorr) between CLR-transformed MetaCyc pathway abundances and host clinical or immunologic markers (CD4, CD8, activated T cell subsets, CRP, IL-6, TNF-α, sCD14, BMI), shown separately for darunavir/ritonavir and dolutegravir. Colors indicate correlation direction and magnitude; black outlines denote significant associations after FDR correction (q value < 0.05).
By Week 48, dolutegravir showed higher L-glutamine biosynthesis III (PWY-6549; log₂FC = 0.36; 95% CI = 0.05–0.66) and methanogenesis from H₂/CO₂ (METHANOGENESIS-PWY; log₂FC = 1.30; 95% CI = 0.22–2.38), whereas darunavir/ritonavir showed higher polyisoprenoid biosynthesis (POLYISOPRENSYN-PWY; log₂FC = –0.62; –1.06 to –0.18), urea cycle (PWY-4984; log₂FC = –0.83; –1.44 to –0.22) y fatty-acid α-oxidation III (PWY66-388; log₂FC = –1.82; 95% CI = –3.17 to –0.46). At Week 96, additional differences included lower creatinine degradation II (PWY-4722; log₂FC = –0.95; 95% CI = –1.87 to –0.03), sulfoglycolysis (PWY-7446; log₂FC = –1.47; 95% CI = –2.92 to –0.03) y L-glutamate→propanoate VIII (PWY-5088; log₂FC = –0.73; 95% CI = –1.36 to –0.11) under dolutegravir (Fig. 6A).
At Week 96, dolutegravir-associated differences persisted but shifted towards pathways involved in nitrogen, sulfur, and amino acid metabolism. Specifically, dolutegravir showed lower relative abundances of creatinine degradation II (PWY-4722; log₂FC = –0.95, 95% CI = –1.87 to –0.03), sulfoglycolysis (PWY-7446; log₂FC = –1.47, 95% CI = –2.92 to –0.03) and L-glutamate → propanoate VIII (PWY-5088; log₂FC = –0.73, 95% CI = –1.36 to –0.11), suggesting a late-phase relative depletion of nitrogen and sulfur turnover routes under dolutegravir.
Applying repeated-measures correlations, we identified treatment-specific couplings between microbial functions and host status (Fig. 6C). Under dolutegravir, CD4⁺ T cell counts tracked positively with central-carbon and biosynthetic pathways—TCA cycle II (ρ = 0.49, q value = 2.1 × 10⁻⁵), methanogenesis from H₂/CO₂ (ρ = 0.37, q value = 5.4 × 10⁻³), and L-glutamine biosynthesis III (ρ = 0.30, q value = 4.8 × 10⁻²). BMI also correlated with polyamine biosynthesis II (ρ = 0.43, q value = 1.6 × 10⁻³), L-glutamate→propanoate degradation (ρ = 0.37, q value = 5.9 × 10⁻³) and methanogenesis (ρ = 0.39, q value = 3.9 × 10⁻³). In contrast, under darunavir/ritonavir, BMI associated positively with the urea cycle (ρ = 0.38, q value = 2.2 × 10⁻²) and negatively with ketogenesis (ρ = −0.37, q value = 2.2×10⁻²) and creatinine degradation II (ρ = −0.34, q value = 4.2 × 10⁻²).
Collectively, these functional data indicate that dolutegravir induces an anabolic remodeling of gut microbial function, enriching central-carbon and biosynthetic pathways that align with higher CD4⁺ T cell counts, while darunavir/ritonavir shows narrower, largely catabolic shifts. In line with these findings, treatment-specific repeated-measures correlations indicate that dolutegravir-enriched functions are associated with improved immune status. In contrast, under darunavir/ritonavir therapy, body mass index (BMI) is correlated with urea cycle activity and inversely linked to ketogenesis, which may reflect a more catabolic and potentially pro-inflammatory state.
Discussion
This randomized trial found that starting antiretroviral therapy with dolutegravir in people with very advanced HIV leads to greater improvement in gut microbiome diversity and function than using darunavir/ritonavir. In our study, dolutegravir initiation was associated with enhanced immune recovery, reduced immune activation, and lower inflammation, while darunavir/ritonavir initiation resulted in minimal microbiome changes over 2 years. This led to gut microbiota profiles more closely resembling those of people without HIV in subjects receiving dolutegravir, compared to those taking darunavir/ritonavir.
Our findings align with Villanueva-Millán et al., who showed that integrase inhibitor regimens normalize gut α-diversity to levels seen in HIV-negative controls, whereas protease inhibitor regimens do not32. In our study, both arms achieved similar virological suppression and peripheral CD4⁺ T cell recovery33, suggesting that the observed microbiome shifts were due to drug-specific qualitative or off-target effects rather than antiviral efficacy. One possible explanation is the superior gastrointestinal penetration of dolutegravir relative to darunavir/ritonavir, which might foster mucosal immune reconstitution, epithelial repair, and create a more favorable niche for commensal gut microbes to thrive. In contrast, darunavir/ritonavir has been previously associated with gut dysbiosis through off-target epithelial damage and increased microbial translocation34,35,36, which may hinder gut microbiota recovery.
Of note, the dolutegravir arm showed concurrent reductions in systemic inflammation and monocyte activation markers, consistent with reduced microbial translocation. Similar findings were observed in a prior trial where an integrase inhibitor regimen led to greater decreases in hsCRP and sCD14 by week 48 compared to non-integrase regimens, with further improvements seen upon switching from a protease to an integrase inhibitor32,36. Elevated inflammation markers (e.g., CRP, IL-6) and monocyte activation and translocation markers (sCD14) in treated patients with HIV have consistently been linked to higher risks of cardiovascular disease, non-AIDS comorbidities, and all-cause mortality37,38. Reductions in such markers with dolutegravir therapy are therefore not only statistically significant but likely carry prognostic value, indicating that integrase inhibitor-based regimens may offer long-term health advantages through the attenuation of ongoing inflammation and progressive microbiome restoration.
In our study, increased gut microbiome α-diversity with dolutegravir was associated with reduced T-cell activation and lower inflammatory cytokines. This is in line with previous research linking higher gut microbial diversity with higher peripheral CD4+ T cell counts and lower endotoxemia in untreated patients with HIV39. Altogether, this suggests that improvements in microbiota composition with dolutegravir may reduce immune activation, fostering a feedback loop that supports ongoing microbiome recovery—unlike the sustained dysbiosis seen with darunavir/ritonavir.
Moreover, we observed a reconfiguration of the gut microbiome toward progressive convergence in microbiota composition (beta-diversity) exclusively among dolutegravir-treated participants, with significant and sustained reductions in centroid distances over 96 weeks. This suggests a shift from the stochastic community structure often seen in advanced immunodeficiency—where dysbiotic microbiomes are uniquely and unpredictably varied—toward a more stable and defined configuration that is likely more closely associated with health than random equilibration40. The persistent microbiome dispersion in the darunavir/ritonavir arm suggests a failure to constrain such heterogeneity. By integrating our data with the metaHIV cohort9,28, we further demonstrated that dolutegravir-treated microbiomes became increasingly similar to those of HIV-uninfected controls, displaying the smallest deviation from the healthy reference group by the end of the study. Whereas full microbiome normalization was not achieved, the inverse correlations between centroid distance and markers of systemic inflammation and immune activation further reinforce the potential clinical relevance of our ecological findings9,15,28,41.
At the taxonomic level, dolutegravir treatment drove a specific reconfiguration of the gut microbial ecosystem characterized by the enrichment of cooperative, fermentative networks rather than just individual taxa. We observed a consistent expansion of hydrogenotrophic archaea (Methanobrevibacter smithii) alongside specialized fermenters such as Olsenella scatoligenes, Parasutterella excrementihominis, and Prevotella species42,43, suggesting restoration of metabolic syntrophy. Methanobrevibacter acts as a hydrogen sink, consuming the H₂ produced by fermenters, thereby thermodynamically favoring the production of short-chain fatty acids (SCFAs) over less favorable metabolic products44,45,46. This environment appears to support the concurrent expansion of lactate producers, including Olsenella and Lactobacillus sakei. The expansion of L. sakei is particularly remarkable, as it is a lactic acid bacterium with documented mucosal anti-inflammatory properties known to reinforce epithelial homeostasis47. Furthermore, the lactate produced by these taxa serves as a critical cross-feeding substrate for butyrate production by commensals such as Clostridium cluster XIVa. The restoration of this pathway may be clinically relevant, as butyrate is essential for maintaining epithelial barrier integrity and promoting the differentiation of colonic regulatory T-cells42. Therefore, functional shifts associated with dolutegravir seem to be more than just changes in microbiome composition, but rather a functional restoration of the H₂–lactate–butyrate pathway, which connects improvements in gut health with better mucosal immunity and reduced systemic inflammation. In contrast, the microbiome of subjects treated with darunavir/ritonavir remained dominated by robust generalists like Bacteroides and potential pathobionts such as Lachnoclostridium. This microbial profile is frequently linked to chronic inflammation and metabolic dysregulation48,49. Notably, recent in vitro data indicate that darunavir lacks direct antibacterial activity against key gut commensals, suggesting that the persistent dysbiosis observed under darunavir/ritonavir is unlikely to be driven by direct antimicrobial pressure50.
The functional translation of the dolutegravir-associated shifts points to a strengthening of the gut barrier. The expanding taxa—particularly Methanobrevibacter and Parasutterella—correlated strongly with reductions in sCD14, a marker of monocyte activation and microbial translocation, and systemic inflammation (CRP, TNF-α). While Parasutterella has been debated as either a commensal or pathobiont, its inverse association with inflammation in our cohort aligns with recent evidence suggesting its role in succinate and bile acid metabolism, supporting intestinal homeostasis30.
The observed microbial shifts towards syntrophy were corroborated by our functional profiling, including a sustained enrichment of Methanogenesis from the H₂/CO₂ pathway and early increases in TCA Cycle II and later L-Glutamine Biosynthesis III. The latter suggests enhanced central-carbon and biosynthetic capacity, the necessary building blocks for community expansion, SCFA production, and structural repair of the gut barrier. Importantly, such functional changes correlated with peripheral CD4+ T-cell count recovery, suggesting that the functional reconfiguration of the microbiome under dolutegravir may play a role in immune reconstitution. In contrast, protease inhibitor therapy resulted in more limited and predominantly catabolic functional changes, which did not facilitate the establishment of anabolic interactions. These persistent relative increases in nitrogen and sulfur turnover catabolic pathways, such as Creatinine degradation II and sulfoglycolysis under darunavir/ritonavir, are consistent with the use of host-derived nutrients and increased microbial turnover during stress or inflammation51,52,53. The positive relationship between BMI and Urea cycle activity may indicate a distinct and potentially less optimal metabolic profile in subjects receiving darunavir/ritonavir.
Our findings may have relevant implications for the management of advanced HIV infection. First, they suggest that the first-line ART choice may impact long-term prognosis through gut health restoration. A dolutegravir-based regimen achieved superior microbial remodeling, i.e., greater diversity, ecological homogenization, and restoration of key anabolic and fermentative networks than darunavir/ritonavir. Moreover, beyond the well-established benefits on viral control and peripheral immune reconstitution, integrase inhibitor therapy may provide additional benefits in terms of mitigating chronic inflammation and reducing the long-term risk of inflammation-driven comorbidities—such as cardiovascular disease and neurocognitive decline—in late presenters37. Second, our study reinforces the gut microbiome as a modifiable component in people living with HIV. Of note, the functional recovery achieved in our trial with optimized antiretroviral therapy is comparable to that of fecal microbiota transplants in previous studies54. In the future, specific microbiome metrics beyond basic CD4+ T cell counts, such as the rate of community homogenization or the activity of specific microbial pathways, might help clinicians identify patients for whom adjuvant therapies such as dietary interventions, specific pre/probiotics, or FMT may be of help. However, this will require proper validation in larger studies in more diverse populations.
Our study presents several limitations. Its modest sample size limits the detection of subtle effects, stratification by baseline characteristics (e.g., CD4+ cell counts), and full adjustment for potential confounders, even though randomization likely minimized such risk. Whereas the absence of a large healthy HIV-negative control group prevents interpretation relative to a normal reference range, this was mitigated by contextualizing our findings against the MetaHIV cross-sectional study9,28. The assessment of microbial translocation was limited to sCD14, and we only evaluated luminal (stool) rather than mucosa-associated microbial communities. Also, the generalizability of our findings is limited due to the cohort being primarily composed of Caucasian males in Spain, which reflects the epidemiological reality of new HIV diagnoses in the Global North. The relatively low proportion of female participants must still be noted, given established sex-related differences in metabolism and gut microbiota. This highlights the importance of assembling more balanced study groups, as advised by the SAGER guidelines55. It also calls for wider inclusion of diverse global populations—such as those from Africa and Southeast Asia—to enhance external validity. Finally, while randomization likely balanced dietary patterns, standardized dietary data collection was not performed. Clinically, our 96-week follow-up is not powered for “hard” clinical endpoints, including death and non-AIDS-related clinical events, which require studies spanning several years of follow-up and involving at least hundreds of patients.
In summary, this randomized clinical trial demonstrates that initiating antiretroviral therapy with dolutegravir leads to a more comprehensive and robust restoration of gut homeostasis in people with advanced HIV, compared to darunavir/ritonavir. Dolutegravir initiation is associated with progressive microbial homogenization, a sustained increase in alpha diversity, and a functional shift towards anabolic, fermentative metabolism, which correlates with reductions in systemic inflammation and immune activation and increased peripheral immune reconstitution. In conclusion, while these findings are exploratory and hypothesis-generating, and require validation in larger and longer-term studies to determine their impact on morbidity and mortality, in this cohort of individuals with advanced HIV infection, dolutegravir-based regimens were associated over 2 years with greater changes in gut microbiome composition and predicted metabolic functions than darunavir/ritonavir-based regimens. The clinical significance of these findings remains to be established.
Methods
Study design
This substudy was nested within the ADVANZ-4 trial (EudraCT: 2014-002281-70, ClinicalTrials.gov: NCT02337322), a prospective, open-label, two-arm randomized clinical trial enrolling ART-naïve adults presenting with CD4+ T cell counts below 100 cells/mm³ at HIV diagnosis27. Participants were randomized 1:1 to receive either dolutegravir (50 mg QD) or ritonavir-boosted darunavir (800/100 mg QD), each combined with a fixed-dose formulation of lamivudine (300 mg QD) and abacavir (150 mg QD). All participants received cotrimoxazole prophylaxis until achieving CD4+ counts >200 cells/mm³ for 3 consecutive months. Follow-up visits were conducted at baseline and weeks 24, 48, and 96. Fecal samples for microbiome analysis were collected at these time points. We employed a modified intention-to-treat 2 (mITT2) approach, restricting the microbiome analysis to participants providing fecal samples at baseline or at least two follow-up time points.
This study adheres to the STORMS reporting guidelines for human microbiome research, and the completed checklist is provided as Supplementary Table 1.
Endpoints and visit schedule
According to the Statistical Analysis Plan (SAP) of the ADVANZ-4 trial, the primary endpoint was the change in absolute CD4⁺ T-cell count from baseline to week 48. Secondary endpoints included: (i) plasma HIV-1 RNA levels and the proportion of participants achieving viral suppression (<50 copies/mL) at all scheduled visits; (ii) changes in lymphocyte subsets, clinical laboratory parameters, and immune activation, inflammation, apoptosis, and bacterial translocation markers (TNF-α and sCD14), measured at baseline, week 24, and week 48; and (iii) the occurrence of clinical outcomes including new AIDS-defining events, immune reconstitution inflammatory syndrome, adverse events, treatment discontinuations, and mortality.
Per protocol and SAP specifications, clinical assessments, blood sampling, and virological measurements were performed at baseline and weeks 4, 12, 24, 36, and 48, with extended follow-up to week 96 in this substudy. Immunological and inflammation biomarkers were assessed at baseline, week 24, and week 48, consistent with the trial protocol.
Participant exclusions
A total of 45 screened individuals were excluded prior to randomization. Most exclusions were due to medical decision (n = 31), primarily involving severe AIDS-defining conditions that compromised clinical stability (n = 11; including opportunistic infections requiring ICU admission [n = 5, of whom two died during admission], primary CNS lymphoma [n = 3], non-Hodgkin lymphoma [n = 2], and CNS toxoplasmosis [n = 1]). Additional medical reasons included severe potential drug–drug interactions (n = 8; mainly tuberculosis or psychiatric comorbidities), absence of social support (homelessness; n = 4), anticipated lack of adherence (n = 3), liver disease (n = 3), and psychiatric disorders (n = 2). Seven individuals did not meet the inclusion criteria (positive HLA-B*5701 test, n = 4; confirmatory CD4+ T cell count >100 cells/mm³, n = 3), and none had baseline antiretroviral resistance. Finally, seven individuals declined to participate.
Sample collection and processing
Stool samples were collected either at clinical sites or at home using the OMNIgene. GUT collection kit (DNA Genotek, Ottawa, Canada). The kit stabilizes microbial DNA at ambient temperatures, minimizing degradation during transport. Upon receipt at the central laboratory (IrsiCaixa, Badalona, Spain), samples were immediately frozen at −80 °C. DNA extraction was performed using the PowerSoil DNA Isolation Kit (MO BIO Laboratories, Carlsbad, CA, USA) following the Earth Microbiome Project standard protocols56, ensuring reproducibility and minimizing technical biases. Mechanical lysis was achieved via bead-beating.
Sequencing libraries were prepared using the Nextera XT DNA Library Prep Kit (Illumina Inc., San Diego, CA, USA) targeting an average insert size of 300 bp. Paired-end sequencing (2 × 150 bp) was performed on the Illumina HiSeq 2500 platform. A sequencing depth target of 20 million reads per sample was established to ensure sufficient coverage for robust metagenomic profiling.
Flow cytometry
Peripheral blood mononuclear cells (PBMCs) were isolated by density-gradient centrifugation using Ficoll-Hypaque (Biomedics-Biomérieux, Madrid, Spain). Immunophenotyping included quantification of CD4⁺ and CD8⁺ T-cell subsets and characterization of naïve, effector, and memory populations based on CCR7 and CD45RA expression. Markers of immune activation (CD38, HLA-DR), senescence (CD57, CD28), and apoptosis (annexin V, CD95) were assessed in both CD4⁺ and CD8⁺ compartments.
Three-color flow cytometry was performed following previously validated protocols from the parent ADVANZ-4 trial. Data were acquired on a standard flow cytometer and analyzed using FlowJo software. For apoptosis assays, PBMCs were cultured for 18 h at 1 × 10⁶ cells/mL in 24-well plates in the presence or absence of 1% phytohemagglutinin, stained with annexin V-PE (BD PharMingen, San Jose, CA) in buffer containing 5 mM CaCl₂, followed by CD95 detection.
Measurement of inflammatory and immune activation markers by ELISA
Plasma concentrations of inflammatory and immune activation markers were quantified using commercially available enzyme-linked immunosorbent assays (ELISA), performed in accordance with the manufacturers’ protocols. All samples were analyzed under standardized experimental conditions, and measurements were conducted by trained personnel blinded to clinical group allocation.
Baseline plasma samples were available for all participants for all analytes included in the study, and no samples were missing at baseline. Samples were assayed in duplicate, and mean values were used for downstream analyses. Intra- and inter-assay coefficients of variation were within the ranges specified by the manufacturers.
For each analyte, concentration values below the assay-specific lower limit of detection (LLOD) were reported by the assay and retained in the dataset rather than treated as missing values. These values were included in both descriptive and inferential analyses. Apparent gaps in graphical representations therefore reflect measurements below the LLOD and not missing data. As a reference, the LLOD for IL-6 was <2 pg/mL; LLODs for the remaining analytes are provided in the corresponding kit documentation.
All raw concentration values for IL-6 and the remaining inflammatory and immune activation markers measured by ELISA are provided in Supplementary Table 2.
Microbiome data analysis
Raw reads were processed with Trimmomatic (v0.39)57, removing Illumina adapters, trimming reads with a sliding window of 30 nucleotides, and discarding reads with average Phred scores below 20. Human reads were removed by mapping against the hg19 reference genome using Bowtie2 (v2.4.2)58 with end-to-end alignment mode and a maximum mismatch penalty of 6. Only unmapped reads were retained.
Taxonomic profiling was performed using MetaPhlAn3 (v3.0)59 with default parameters, based on clade-specific marker genes from the CHOCOPhlAn database (version 201901). Reads were additionally mapped to the Integrated Gene Catalog (IGC)60 using Bowtie2 for gene-level quantification. Rarefaction curves were generated by subsampling reads to increasing coverage thresholds, and gene richness was defined at the minimum point where curves plateaued. Samples falling below the 2nd percentile of the maximum coverage distribution were excluded to mitigate depth-related biases.
Microbiome data processing
Post-taxonomic assignments, abundance matrices were imported into R as Phyloseq objects (phyloseq v1.42.0)61 for preprocessing. Samples were filtered for completeness of metadata and microbiome data. Taxonomic abundance, taxonomy tables, and sample metadata were integrated into a Tree Summarized Experiment (TreeSE) object (tidyTreeSummarizedExperiment v1.10.0), facilitating consistent downstream analysis.
Alpha diversity analysis
Alpha diversity was evaluated using the mia R package (v1.8.0). Metrics computed included Shannon diversity, Gini dominance, and observed richness (number of detected taxa). To account for uneven sequencing depth, a rarefaction strategy was implemented at the 2nd percentile of library sizes, repeated over 10 random iterations to ensure robustness.
Group and longitudinal differences were analyzed with linear mixed-effects models with the following formula:
This model included fixed effects for treatment, time point, and their interaction, and a random intercept for participant. Multiple testing was controlled using the Benjamini–Hochberg FDR, and model-based effect sizes with 95% confidence intervals are reported. Distributions were visualized with boxplots overlaid with jittered datapoints and estimated marginal means for interpretability.
To characterize within-arm consistency of longitudinal trajectories, we additionally estimated a subject-level slope of change from baseline (Δ per visit) for each diversity metric and compared the dispersion of slopes between treatment arms. Equality of variances was assessed using Brown–Forsythe and Fligner–Killeen tests, and variance ratios (dolutegravir/darunavir/ritonavir) with bootstrap confidence intervals (5000 resamples) were computed to quantify differences in intra-group variability. As a confirmatory analysis within the mixed-model framework, we fitted location–scale linear mixed models allowing arm-specific residual variances (varIdent structure in the nlme package) and compared these to homoscedastic models using likelihood ratio tests.
Gene richness was separately assessed by counting the number of genes mapped above a dynamic threshold determined per sample, ensuring comparability across the cohort. This dynamic threshold was derived based on the rarefaction analysis against the IGC, selecting for each sample the coverage depth at which the rarefaction curve plateaued, thus avoiding biases due to differences in sequencing depth.
A detailed overview of all statistical analyses, including the family of tests and the number of comparisons used for FDR control in each analysis, is provided in Supplementary Table 3.
Correlation analyses
Repeated-measures correlation effects (rmcorr), which account for within-participant dependency across longitudinal samples, were calculated to explore associations between microbiome alpha diversity indices (Shannon diversity, Gini dominance, observed richness, and gene richness) and a comprehensive panel of clinical parameters. The clinical variables included circulating immune cell counts (CD4+, CD8+, CD4 nadir), immune activation markers (expression of CD38 and HLA-DR on CD4+ and CD8+ T cells), systemic inflammatory markers (C-reactive protein [CRP], interleukin-6 [IL-6], tumor necrosis factor-alpha [TNF-α], and soluble CD14 [sCD14]), and body mass index (BMI).
To normalize the distribution of cytokine levels and mitigate the influence of extreme values, cytokine data were log2(x + 1) transformed prior to analysis. Repeated-measures correlations were computed both across the full cohort and within each treatment arm to capture treatment-specific host–microbiome associations.
Two-sided p values from correlation tests were adjusted using the Benjamini–Hochberg method to control for the FDR. Only associations with q < 0.05 were considered significant. Results are presented as correlation coefficients (r) and visualized in heatmaps, with color encoding the strength and direction of associations. Selected relationships were further illustrated with scatterplots showing participant-level trends and fitted regression lines.
Longitudinal evolution of clinical marker levels
Longitudinal changes in clinical markers were analyzed using linear mixed-effects models with the specification:
Here, time_point was modeled as a categorical factor to allow non-linear trajectories, and a random intercept for participant_id accounted for within-individual correlation. The treatment-by-time interaction enabled inference on both between-group differences at each time point and within-group longitudinal changes. Model coefficients (β) and 95% confidence intervals were estimated, and planned contrasts (pairwise time comparisons within treatment and between-treatment comparisons at each time point) were derived from the fitted models. Multiple testing was controlled using the Benjamini–Hochberg FDR. Visualization included boxplots of raw marker distributions by time and treatment with jittered points, forest plots summarizing model-based contrasts with FDR-adjusted p values, and longitudinal effect plots showing time on the x-axis and estimated effects (with 95% CIs) on the y-axis to highlight temporal trends.
Beta diversity analysis
Beta diversity was computed with Bray–Curtis dissimilarities from relative-abundance profiles. Community structure was visualized by NMDS (monoMDS in vegan). To test associations while accounting for repeated measurements, we fit PERMANOVA models with adonis2 using:
Constraining permutations within participants (strata = record_id). Only complete cases for model covariates were included.
To summarize within-group dispersion, we calculated each sample’s Euclidean distance in NMDS space to its leave-one-out centroid for the corresponding treatment × time_point group (“centroid distance”). Centroid distances were analyzed with linear mixed-effects models (lmerTest) to estimate (i) longitudinal changes within treatment arms versus baseline (week 0) and (ii) between-treatment differences at each time point, using:
Planned contrasts were obtained with emmeans; we report estimates, 95% CIs, and Benjamini–Hochberg FDR-adjusted p values. Results are shown as forest plots (between-treatment contrasts) and longitudinal trajectory plots (within-treatment changes vs baseline).
Associations between community variation and clinical/immunologic markers (CD4, CD8, activated T cell subsets, CRP, IL-6, TNF-α, sCD14, BMI) were assessed using repeated-measures correlation (rmcorr), reported as correlation coefficients (r) with FDR-adjusted p values, both globally and stratified by treatment. For compositional summaries, genus-level stacked barplots (top 25 genera; remainder “Other”) were ordered by NMDS1 and annotated with treatment, risk group, time point, and centroid distance. Figures were generated with ggplot2 and arranged with patchwork; PERMANOVA tables were rendered with gt. All analyses were performed in R.
Differential abundance analysis
We first benchmarked microbiome differential-abundance software using the dar R package to guide method selection62. Differentially abundant taxa were then identified with ANCOMBC2 (v2.1.2) (REF) on TreeSummarizedExperiment objects at Genus and Species resolution. The model specification included fixed effects for treatment, time, and covariates, and a subject-level random intercept to account for repeated measures:
Pairwise treatment contrasts at each time point were obtained by setting group = “treatment” and pairwise = TRUE. Analyses applied a prevalence filter of ≥10% across samples and used Benjamini–Hochberg FDR control. We considered taxa significant if the FDR-adjusted q value < 0.05 and the 95% confidence interval for the log fold-change excluded 0. Results are reported as effect sizes (logFC) with 95% confidence intervals and visualized as time-stratified forest plots and heatmaps of logFC (with a side annotation for prevalence).
To relate differential taxa to host immunologic and clinical markers (CD4+ and CD8+ T cell counts, activated T cell subsets, CRP, IL-6, TNF-α, sCD14, BMI), we computed repeated-measures correlations (rmcorr) between CLR-transformed taxon abundances and each marker. Cytokines were log2-transformed with a +1 offset prior to analysis. Repeated-measures correlation coefficients (r) and FDR-adjusted p values were summarized overall and by treatment arm and displayed as dotmaps and per-marker rmcorr plots.
Significance criteria were: |log2 fold-change| ≥ 0.5, FDR-adjusted p value < 0.05, and prevalence >20%. Differential features were visualized using volcano plots and annotated barplots. Significant taxa were additionally correlated with clinical parameters using Spearman’s method.
Functional analysis
Functional profiling focused on MetaCyc pathways derived from HUMAnN3 outputs59. We retained non–taxa-stratified pathways and excluded “UNMAPPED/UNINTEGRATED” entries. Pathway abundance tables were aligned to the study metadata and assembled into TreeSummarizedExperiment objects. Differential pathway abundance was tested with ANCOMBC2 using the model:
With a subject-level random intercept (record_id) to account for repeated measures, group = “treatment”, Benjamini–Hochberg FDR control, and a prevalence filter of ≥10%. We considered effects significant when the FDR-adjusted q < 0.05 and the 95% CI for the log fold-change excluded 0. Results are reported as logFC with 95% CIs and visualized as time-stratified forest plots and heatmaps of logFC (including a prevalence side annotation).
To relate functional variation to host markers (CD4+ and CD8+ T cell counts, activated T cell subsets, CRP, IL-6, TNF-α, sCD14, BMI), we computed repeated-measures correlations (rmcorr) between CLR-transformed pathway abundances and each marker; cytokines were log2-transformed with a +1 offset. Correlations were summarized globally and by treatment arm, and multiple testing was controlled with FDR.
External cohort integration and comparative dissimilarity analysis (MetaHIV)
To contextualize ADVANZ-4 microbiome profiles against external HIV phenotypes, we integrated samples from the MetaHIV study and reprocessed both datasets with a harmonized pipeline. Short-read profiles were generated with nf-core/taxprofiler using MetaPhlAn4 to obtain species-level relative abundances. Sample metadata were standardized (common IDs, time points, and HIV phenotype labels) and merged. A TreeSummarizedExperiment object was constructed at the species rank, keeping non-missing samples with complete metadata.
Beta diversity (Bray–Curtis) was computed from species-level relative abundances. For each sample iii, we quantified its dissimilarity to HIV-negative microbiomes as the mean Bray–Curtis distance to all HIV-negative samples:
where N indexes HIV-negative samples. Group differences in diversity were tested with a linear mixed-effects model including a subject-level random intercept:
followed by estimated marginal means and treatment-vs-control contrasts with HIV-negative as reference (FDR control via Benjamini–Hochberg). Results are presented as effect estimates with 95% CIs in forest plots.
For ordination, we performed NMDS (Bray–Curtis; monoMDS in vegan) on the integrated species matrix to visualize separation across HIV phenotypes and ADVANZ-4 time points. As a complementary summary, we computed each sample’s Euclidean distance in NMDS space to the HIV-negative centroid:
and assessed differences versus HIV-negative with the same mixed-model framework and FDR-adjusted inference.
Network analysis
To explore genus-level co-occurrence patterns under dolutegravir (dolutegravir) versus darunavir/ritonavir (darunavir/ritonavir) therapy, we began with the pre-processed SingleCellExperiment object containing CLR- and log-transformed assays. Samples were subset by time point (0, 24, 48, and 96 weeks) and, within each subset, counts were agglomerated at the genus rank. Relative-abundance profiles were generated, and taxa present in fewer than 15% of samples were removed. The remaining genera were then transformed by centered log-ratio (pseudocount = 1) to stabilize variance. For each time point, the filtered genus-level data were split by treatment group, yielding separate dolutegravir and darunavir/ritonavir matrices for network inference.
Networks were constructed with the SPRING method in NetCoMi’s netConstruct function (NetCoMi v1.1.0)63,64, restricting to the 200 most prevalent genera and exploring 20 values of the regularization parameter (nlambda = 20, rep.num = 10, thresh = 0.05, Rmethod = “approx”) on signed dissimilarities. The resulting microNet objects were characterized using netAnalyze, applying Louvain community detection, identifying hub taxa via eigenvector centrality, reporting unnormalized degree centrality on the LCC, and computing graphlet orbit counts together with the Graphlet Correlation Matrix (GCM) for that component. To highlight treatment-specific shifts in network micro-architecture, we calculated the difference between the dolutegravir and darunavir/ritonavir GCMs (ΔGCM = GCM_dolutegravir – GCM_darunavir/ritonavir) and visualized this differential matrix as a single heatmap, with statistical significance of Δρ values assessed by FDR-adjusted Spearman tests.
Software and reproducibility
All analyses were conducted in R (v4.5.0) using packages from the Bioconductor (v3.21) and CRAN repositories. Scripted workflows were version-controlled with Git, and data processing pipelines were documented using Quarto notebooks, ensuring transparency. Parallel computing was leveraged via the BiocParallel or future R packages. R Session Info and packages are provided in each Quarto notebook to facilitate reproducibility.
Ethics approval and consent to participate
This study was approved by the institutional ethical review board of the participating institutions (Reference HCB/2014/0905) and by the Spanish Regulatory Authorities (EudraCT 2014-002281-70). All participants provided written informed consent in accordance with the principles expressed in the Declaration of Helsinki and local personal data protection law (LOPD 15/1999).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw metagenomic sequencing data generated in this study have been deposited in the European Nucleotide Archive (ENA) database under accession code PRJEB76817. The clinical, immunological, and microbiome data generated in this study are provided in the Supplementary Information and Source data files. Source data are provided with this paper.
Code availability
The code used to perform the analyses and generate results in this study is publicly available and has been deposited in a GitHub repository at https://github.com/MicrobialGenomics-IrsicaixaOrg/fcatala_advanz4, under MIT license65. Users are permitted to reuse, modify, and distribute the code in accordance with the terms of the license. Any modifications to the code should appropriately credit the original authors as outlined by the license terms.
References
Antinori, A. et al. Late presentation of HIV infection: a consensus definition. HIV Med. 12, 61–64 (2011).
May, M. T. Better to know: the importance of early HIV diagnosis. Lancet Public Health 2, https://doi.org/10.1016/S2468-2667(16)30038-X (2017).
INSIGHT START Study Group et al. Initiation of antiretroviral therapy in early asymptomatic HIV infection. N. Engl. J. Med. 373, 795–807 (2015).
Lundgren, J. D. et al. Long-term benefits from early antiretroviral therapy initiation in HIV infection. NEJM Evid. 2, EVIDoa2200302 (2023).
Paiardini, M. & Müller-Trutwin, M. HIV-associated chronic immune activation. Immunol. Rev. 254, 78–101 (2013).
Stirrup, O. T. et al. (2020) Predictors of CD4 cell recovery following initiation of antiretroviral therapy among HIV-1 positive patients with well-estimated dates of seroconversion. HIV Med. 19, 184–194 (2018).
Estimating the burden of HIV late presentation and its attributable morbidity and mortality across Europe 2010-2016. BMC Infect. Dis. 20, https://doi.org/10.1186/s12879-020-05261-7.
Yang, X. et al. Incomplete immune reconstitution in HIV/AIDS patients on antiretroviral therapy: challenges of immunological non-responders. J. Leukoc. Biol. 107, https://doi.org/10.1002/JLB.4MR1019-189R (2020).
Guillén, Y. et al. Low nadir CD4+ T-cell counts predict gut dysbiosis in HIV-1 infection. Mucosal Immunol. 12, 232–246 (2019).
Veazey, R. S. et al. Gastrointestinal tract as a major site of CD4+ T cell depletion and viral replication in SIV infection. Science 280, 427–431 (1998).
Kuo, H. H. & Lichterfeld, M. Recent progress in understanding HIV reservoirs. Curr. Opin. HIV AIDS 3, https://doi.org/10.1097/COH.0000000000000441 (2018).
Thompson, C. G., Gay, C. L. & Kashuba, A. D. M. HIV persistence in gut-associated lymphoid tissues: pharmacological challenges and opportunities. AIDS Res. Hum. Retrovir. 33, https://doi.org/10.1089/aid.2016.0253 (2017).
Vujkovic-Cvijin, I. et al. Dysbiosis of the gut microbiota is associated with HIV disease progression and tryptophan catabolism. Sci. Transl. Med. 5, 193ra91 (2013).
Blaschitz, C. & Raffatellu, M. Th17 cytokines and the gut mucosal barrier. J. Clin. Immunol. 30, https://doi.org/10.1007/s10875-010-9368-7 (2010).
Dillon, S. M., Frank, D. N. & Wilson, C. C. The gut microbiome and HIV-1 pathogenesis: a two-way street. AIDS 30, https://doi.org/10.1097/QAD.0000000000001289 (2016).
Garidou, L. et al. The gut microbiota regulates intestinal CD4 T cells expressing RORγt and controls metabolic disease. Cell Metab. 22, 100–112 (2015).
Català-Moll, F. & Paredes, R. The rectal microbiome: understanding its role in HIV transmission. Curr. Opin. HIV AIDS 20, 159–164 (2025).
Welch, J. L. M., Hasegawa, Y., McNulty, N. P., Gordon, J. I. & Borisy, G. G. Spatial organization of a model 15-member human gut microbiota established in gnotobiotic mice. Proc. Natl. Acad. Sci. USA 114, E9105–E9114 (2017).
Jenabian, M. A. et al. Immunosuppressive tryptophan catabolism and gut mucosal dysfunction following early HIV infection. J. Infect. Dis. 212, 355–366 (2015).
Rawi, M. H., Zaman, S. A., Pa’ee, K. F., Leong, S. S. & Sarbini, S. R. Prebiotics metabolism by gut-isolated probiotics. J. Food Sci. Technol. 57, https://doi.org/10.1007/s13197-020-04244-5 (2020).
D’Angelo, C., Reale, M. & Costantini, E. Microbiota and probiotics in health and HIV infection. Nutrients 9, https://doi.org/10.3390/nu9060615 (2017).
Crakes, K. R. & Jiang, G. Gut microbiome alterations during HIV/SIV infection: implications for HIV cure. Front. Microbiol. 10, https://doi.org/10.3389/fmicb.2019.01104 (2019).
Vujkovic-Cvijin, I. et al. HIV-associated gut dysbiosis is independent of sexual practice and correlates with noncommunicable diseases. Nat. Commun. 11, 2448 (2020).
Greener, B. N. et al. Dolutegravir pharmacokinetics in the genital tract and colorectum of HIV-negative men after single and multiple dosing. J. Acquir. Immune Defic. Syndr. 64, 39–44 (2013).
Thompson, C. G., Cohen, M. S. & Kashuba, A. D. M. Antiretroviral pharmacology in mucosal tissues. J. Acquir. Immune Defic. Syndr. 63, S240–S247 (2013).
Le Chatelier, E. et al. Richness of human gut microbiome correlates with metabolic markers. Nature 500, 541–546 (2013).
Miro, J. M. et al. Immune reconstitution in very advanced HIV patients treated with dolutegravir vs. darunavir-based triple antiretroviral therapy: the Advanz-4 randomized clinical trial. Clin. Microbiol. Infect. https://doi.org/10.1016/j.cmi.2025.09.026 (2025).
Noguera-Julian, M. et al. Gut Microbiota Linked to Sexual Preference and HIV Infection. EBioMedicine 5, 135–146 (2016).
Kuehnast, T. et al. Exploring the human archaeome: its relevance for health and disease, and its complex interplay with the human immune system. FEBS J. 292, 1316–1329 (2025).
Galley, J. D. et al. Multifactorial effects of probiotic Parasutterella excrementihominis on gestational inflammation, offspring behaviour and prenatal-stress induced disruptions in tryptophan metabolism. Benef. Microbes 16, 293–310 (2024).
Li, X. et al. The direct and indirect inhibition of proinflammatory adipose tissue macrophages by acarbose in diet-induced obesity. Cell Rep. Med. 6, 101883 (2025).
Villanueva-Millán, M. J., Pérez-Matute, P., Recio-Fernández, E., Rosales, J. M. L. & Oteo, J. A. Differential effects of antiretrovirals on microbial translocation and gut microbiota composition of HIV-infected patients. J. Int. AIDS Soc. 20, 21526 (2017).
Fabbiani, M. et al. Efficacy and durability of dolutegravir- or darunavir-based regimens in ART-Naïve AIDS- or late-presenting HIV-infected patients. Viruses 15, 1123 (2023).
Imahashi, M. et al. Impact of long-term antiretroviral therapy on gut and oral microbiotas in HIV-1-infected patients. Sci. Rep. 11, 960 (2021).
Pinto-Cardoso, S., Klatt, N. R. & Reyes-Terán, G. Impact of antiretroviral drugs on the microbiome: unknown answers to important questions. Curr. Opin. HIV AIDS 13, https://doi.org/10.1097/COH.0000000000000428 (2018).
Gáspár, Z., Nagavci, B., Szabó, B. G. & Lakatos, B. Gut microbiome alteration in HIV/AIDS and the role of antiretroviral therapy—a scoping review. Microorganisms 12, 2221 (2024).
Hileman, C. O. & Funderburg, N. T. Inflammation, Immune Activation, and Antiretroviral Therapy in HIV. Curr. HIV/AIDS Rep. 14, https://doi.org/10.1007/s11904-017-0356-x (2017).
Sandler, N. G. et al. Plasma levels of soluble CD14 independently predict mortality in HIV infection. J. Infect. Dis. 203, 780–790 (2011).
Nowak, P. et al. Gut microbiota diversity predicts immune status in HIV-1 infection. AIDS 29, 2409–2418 (2015).
Zaneveld, J. R., McMinds, R. & Vega Thurber, R. Stress and stability: applying the Anna Karenina principle to animal microbiomes. Nat. Microbiol. 2, 17121 (2017).
Vázquez-Castellanos, J. F. et al. Altered metabolism of gut microbiota contributes to chronic immune activation in HIV-infected individuals. Mucosal Immunol. 8, 760–762 (2015).
Lopetuso, L. R., Scaldaferri, F., Petito, V. & Gasbarrini, A. Commensal Clostridia: leading players in the maintenance of gut homeostasis. Gut Pathog. 5, 23 (2013).
Ju, T., Kong, J. Y., Stothard, P. & Willing, B. P. Defining the role of Parasutterella, a previously uncharacterized member of the core gut microbiota. ISME J. 13, 1520–1534 (2019).
Samuel, B. S. et al. Genomic and metabolic adaptations of Methanobrevibacter smithii to the human gut. Proc. Natl. Acad. Sci. USA 104, 10643–10648 (2007).
Camara, A. et al. Clinical evidence of the role of Methanobrevibacter smithii in severe acute malnutrition. Sci. Rep. 11, 5426 (2021).
van Hoek, M. J. A. & Merks, R. M. H. Emergence of microbial diversity due to cross-feeding interactions in a spatial model of gut microbial metabolism. BMC Syst. Biol. 11, 56 (2017).
Lee, G. H., Choi, S. H., Lee, Y. H. & Park, J. K. Antioxidant and Anti-Inflammatory Activities of Latilactobacillus curvatus and L. sakei Isolated from Green Tripe. Nutrients 17, 2464 (2025).
Boulangé, C. L., Neves, A. L., Chilloux, J., Nicholson, J. K. & Dumas, M.-E. Impact of the gut microbiota on inflammation, obesity, and metabolic disease. Genome Med. 8, 42 (2016).
Dai, W. et al. Uncovering a causal connection between the Lachnoclostridium genus in fecal microbiota and non-alcoholic fatty liver disease: a two-sample Mendelian randomization analysis. Front. Microbiol. 14, 1276790 (2023).
Rubio-Garcia, E. et al. In vitro antibacterial activity of antiretroviral drugs on key commensal bacteria from the human microbiota. Front. Cell. Infect. Microbiol. 13, 1306430. (2024).
Herrera, M. & Byerley, L. O. Dietary nitrogen and its role in the gut microbiome and inflammatory bowel disease: a narrative review. Nutrients 17, 2373 (2025).
Rowland, I. et al. Gut microbiota functions: metabolism of nutrients and other food components. Eur. J. Nutr. 57, 1–24 (2018).
D’Agostino, G. D., Chaudhari, S. N. & Devlin, A. S. Host-microbiome orchestration of the sulfated metabolome. Nat. Chem. Biol. 20, 410–421 (2024).
Serrano-Villar, S. et al. Fecal microbiota transplantation in HIV: a pilot placebo-controlled study. Nat. Commun. 12, 1139 (2021).
Heidari, S., Babor, T. F., De Castro, P., Tort, S. & Curno, M. Sex and Gender Equity in Research: rationale for the SAGER guidelines and recommended use. Res. Integr. Peer Rev. 1, 2 (2016).
Jones, D. S. et al. Community genomic analysis of an extremely acidophilic sulfur-oxidizing biofilm. ISME J. 6, 158–170 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with biobakery 3. eLife 10, e65088 (2021).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841 (2014).
McMurdie, P. J. & Holmes, S. Phyloseq: a bioconductor package for handling and analysis of high-throughput phylogenetic sequence data. In Proc. Pacific Symposium on Biocomputing https://doi.org/10.1142/9789814366496_0023 (2012).
Farré-Badia, J., Noguera-Julian, M., Paredes, R. & Català-Moll, F. dar: a consensus-based framework for differential abundance testing in microbiome data. https://doi.org/10.1101/2025.02.07.637000 (2025).
Peschel, S., Müller, C. L., Von Mutius, E., Boulesteix, A. L. & Depner, M. NetCoMi: network construction and comparison for microbiome data in R. Brief. Bioinform. 22, https://doi.org/10.1093/bib/bbaa290 (2021).
Yoon, G., Gaynanova, I. & Müller, C. L. Microbial networks in SPRING - Semi-parametric rank-based correlation and partial correlation estimation for quantitative microbiome data. Front. Genet. 10, 516 (2019).
Català-Moll, F. et al. Code for dolutegravir restores gut microbiota in late-stage HIV-1 unlike darunavir: an open-label, randomized clinical trial. https://doi.org/10.5281/zenodo.18388866 (2026).
Acknowledgements
The authors want to dedicate this publication to the beloved memory of their esteemed friends and colleagues Carlos Blázquez Bondia and Lluís Force Sanmartí. This work would have been a pillar of Dr. Blaquez’s PhD thesis and was very advanced when he passed away.
Funding
This microbiome study was funded by the European Commission Horizon 2020 Research and Innovation Programme under Grant Agreement N° 847943 (MISTRAL). ViiV Healthcare funded the parent ADVANZ-4 trial and provided additional funds to this microbiome study. J.M.M. received a personal 80:20 research grant from Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), Barcelona, Spain, during 2017–25. None of the funding sources was involved in the study design, data collection, data analysis, result interpretation, or writing of the report.
Author information
Authors and Affiliations
Consortia
Contributions
The study was designed and conceived by J.M.M. and R.P. F.T., C.M., E.B., V.F., P.D., D.P., L.F., A.C., M.P., M.C., J.M.M., and R.P. contributed to data collection. The analysis was conducted by F.C.M., C.B.B., J.F.B., M.N.J., and R.P., and the results interpreted by F.C.M., C.B.B., M.N.J., J.M.M., and R.P. The manuscript was first drafted by F.C.M. and R.P. All authors made substantial contributions to revising the successive drafts and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
V.F., P.D., and A.C. have received fees for educational activities and/or consultancies and/or financial support for attending conferences from Gilead Science, Janssen-Cilag, Merck Sharp & Dohme, and ViiV Healthcare. J.M.M. has received consulting honoraria and/or research grants from Angelini, Basilea, Genentech, Gilead Sciences, Janssen, Lysovant, Medtronic, MSD, Novartis, Pfizer, and ViiV Healthcare, outside the submitted work. R.P. has participated in advisory boards for Pfizer, Gilead, MSD, GSK, Atea, Lilly, Roche, AstraZeneca, ViiV Healthcare, and Theratechnologies, has participated in lectures and seminars funded by Gilead, Pfizer, GSK, and AstraZeneca, and has received research funds awarded to his institution from Gilead, Pfizer, and MSD. The remaining authors declare no competing interests.
Consent for publication
Where individual patient data are included, all patients provided informed consent.
Peer review
Peer review information
Nature Communications thanks Jessica Prodger, Sergio Serrano-Villar, and Souheil A Younes for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Català-Moll, F., Blázquez-Bondia, C., Farré-Badia, J. et al. Dolutegravir restores gut microbiota in late-stage HIV-1 unlike darunavir: an open-label, randomized clinical trial. Nat Commun 17, 2022 (2026). https://doi.org/10.1038/s41467-026-69846-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69846-7