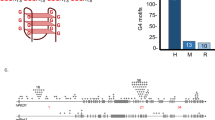
Abstract
The human genome contains approximately 800 G protein-coupled receptors (GPCRs), all characterized by a common 7-transmembrane domain architecture. Here, we show that PKD1, an 11-transmembrane protein with a noncanonical transient receptor potential (TRP) channel architecture, functions as a GPCR with unique biochemical properties. PKD1 acts as a WNT-activated receptor, directly coupling to heterotrimeric Gαi1-3 subunits to inhibit cellular cAMP accumulation. While PKD1 contains both ligand-binding and G protein recruitment sites, PKD2, an associating TRP channel subunit, chaperones PKD1 to the plasma membrane to operate as a GPCR. This represents a striking departure from classical GPCR architecture and expands the functional repertoire of the TRP channel family. Given that mutations in either PKD1 or PKD2 are linked to autosomal dominant polycystic kidney disease, a multisystemic disorder marked by elevated cAMP levels, our results provide molecular insights into disease pathogenesis and highlight potential new therapeutic avenues for this debilitating and costly condition.
Similar content being viewed by others
Introduction
Vertebrate genomes contain ~1000 G protein-coupled receptors (GPCRs), forming the largest class of cell surface receptors mediating a plethora of cellular functions. All known GPCRs have a structural signature consisting of a 7-transmembrane (TM) domain flanked by an extracellular N-terminal segment and an intracellular C-terminal tail. Coupling of these receptors to a diverse group of heterotrimeric G proteins leads to the generation of second messengers, such as signaling lipids, Ca2+, and cAMP, that initiate a complex array of cellular responses1,2,3.
Autosomal dominant polycystic kidney disease (ADPKD), a multisystemic genetic disease affecting roughly 12.5 million people worldwide4,5, has been mechanistically linked to altered cAMP metabolism6,7,8,9. ADPKD is caused by heterozygous germline mutations in PKD1 and PKD2 genes10,11,12. PKD1 is a large (~500 kDa) integral membrane protein with 11 membrane-spanning segments13, an extensive extracellular N-terminal fragment, and an intracellular C-terminal tail14. PKD2 (~130 kDa) spans the membrane 6 times and belongs to the transient receptor potential (TRP) ion channel superfamily12,15. PKD1 and PKD2 assemble to form a heteromeric complex with a noncanonical “TRP” channel architecture16 for which the function has not been clearly defined. A unique and paradoxical feature of the PKD1/PKD2 complex is that the side chains of three positively-charged residues of PKD1 protrude into the ionic pore of PKD2, rendering it unlikely to conduct Ca2+ and other cations. Thus, while these subunits do assume a TRP-like pore architecture, their function as a cation-permeable channel complex is unsettled.
Evidence accumulated over 30 years has shown that cells derived from ADPKD patients have abnormally high cytosolic cAMP levels6,7,8,9 and an aberrant response to cAMP17, which is critical for disease development. Deletion of phosphodiesterase 1 and 3 family members (Pde1a, Pde1c, and Pde3a) aggravate PKD18, and pharmacologic inhibition of PDE4 promotes cystogenesis in mouse Pkd1-null embryonic kidneys19. Conversely, pharmacologic and genetic manipulations to lower cAMP suppress cyst progression and improve renal function20,21,22. These studies culminated in the identification of Tolvaptan, which serves as the only FDA-approved agent for the treatment of ADPKD23,24,25. Tolvaptan is a potent antagonist of the vasopressin 2 receptor26, which is coupled to the heterotrimeric Gαs subunit, leading to cAMP production. Whilst the use of Tolvaptan is limited to a subset of ADPKD patients due to side effects, these studies collectively suggest that elevated cellular cAMP concentration is a driver of cystogenesis in ADPKD. However, as the molecular function(s) of PKD1 and PKD2 remain elusive, a major knowledge gap is the mechanisms by which PKD1 and/or PKD2 keep cytosolic levels of cAMP in check.
Fragmented evidence suggests that PKD1 may function as an atypical GPCR27. Indirect support for the function of PKD1 as a GPCR comes from the presence of a GPCR-autoproteolysis-inducing (GAIN) domain in its ectodomain, a defining feature of cell adhesion GPCRs28,29. However, there is no evidence linking autoproteolysis to cAMP regulation. Independent biochemical studies show that several Gα subunits can directly bind the C-terminal tail of PKD130,31, and a naturally occurring PKD1 mutation (L4132Δ) within a predicted G protein-binding motif leads to severe cystic disease32. Finally, functional studies using various cell culture heterologous systems33,34,35,36,37, a mouse model38, or the Xenopus pronephric model system support the role of PKD1 in GPCR-mediated signaling31. However, direct evidence that PKD1 functions as a GPCR in response to a specific ligand in any system is lacking. Combined with the fact that PKD1 lacks the typical “7-TM” signature architecture of classical GPCRs, these knowledge gaps have left the questions open of whether PKD1 is truly a GPCR and, if so, how this function is linked to cellular cAMP levels.
Results
Activation of PKD1 by WNT ligands induces Gα-Gβγ dissociation
We have shown previously that WNT-9B functions as the activating ligand of PKD1 in a signaling pathway determining kidney tubular diameter during renal development39. Disruption of this pathway results in the failure of developing tubules to achieve and maintain the proper tubular diameter, leading to cystogenesis40,41. Here, we sought to determine whether PKD1 mediates heterotrimeric G protein activation in response to WNT-9B. Activated GPCRs promote the exchange of GDP for GTP on Gα subunits, followed by the dissociation of Gα-GTP and free Gβγ1. To assess PKD1-mediated Gα-Gβγ dissociation, we employed a set of bioluminescence resonance energy transfer (BRET)-based G protein activity sensors (G-CASE) covering all four major G protein families42. These sensors comprise Gα, Gβ, and Gγ on a single plasmid, with Gα subunits tagged with the luminescent donor, NanoLuciferase (Nluc), and Gγ subunits N-terminally labeled with circularly permuted Venus (cpVenus173) to serve as the acceptor fluorophore (Fig. 1a). We conducted ΔBRET time courses for six of the available G protein sensors in human embryonic kidney (HEK) 293 T cells transiently co-transfected with the different biosensors and PKD1 (Fig. 1b–g). Kinetic analysis covered a time span of 10 min, 1 min before and 9 min after stimulation with 1 μg/ml (30 nM) WNT-9B or vehicle control. WNT-9B did not induce the dissociation of Gαs or Gα13 (Fig. 1b, c). However, WNT-9B induced PKD1-mediated dissociation of Gαq and Gαi1-3, with the largest response seen for Gαi3 (Fig. 1d–h). The EC50 for WNT-9B was 405.6 ng/ml (11 nM) (Fig. 1i), which was consistent with the estimated EC50 of WNT-9B binding to extracellular fragments of PKD139 and channel activation43. The magnitude of PKD1-mediated Gαi3 activation by WNT-9B was comparable to the magnitude of responses elicited by the activation of known GPCRs, such as the muscarinic acetylcholine receptor M1 (M1R) and calcium-sensing receptor (CaSR) using G-CASE sensors (Supplementary Fig. 1a, b). Cell surface expression of PKD1 was confirmed using a BRET-based saturation assay; human PKD1 C-terminally-tagged with Nluc (PKD1-Nluc) was titrated with increasing concentrations of a HaloTag-fused cell surface marker bearing the RAS CAAX motif44 (Supplementary Fig. 1c). Specificity of this interaction is indicated by saturable binding (Supplementary Fig. 1d). Live-cell imaging verified expression of PKD1-eGFP at the cell surface, as seen by co-localization with HaloTag-fused CAAX (Supplementary Fig. 1e).
a Schematic representation of the G-CASE BRET assay setup. Created in BioRender. Tsiokas, L. (2026) https://BioRender.com/5p1mc32. b–g ΔBRET time courses of G-CASE BRET sensors upon WNT-9B, WNT-5A, or WNT-3A addition (1 μg/ml, time = 0 min). h Heat map representation of the area under curve (AUC, left panel) and rate constants [k (min−1), right panel] determined from the data shown in (b–g). WNT-induced kinetics were analyzed with a plateau followed by one phase dissociation equation. i Ligand dose-response curve for Gαi3-CASE, plotted as the maximum ΔBRET (%) determined within 6 min following WNT-9B stimulation in the kinetic experiments. The curve was fit using a nonlinear three-parameter model. j ΔBRET time course of Gαi3-CASE in HEK293T cells transiently transfected with pcDNA3 (Control, gray), CRISPR human PKD1 KO #1-3 (PKD1KO, red), mouse wild-type PKD1 (PKD1WT, blue), or PKD1KO + PKD1WT (purple). Cells were stimulated with WNT-9B (1 μg/ml, time = 0 min). Statistical significance was assessed by one-way ANOVA using Tukey’s post-hoc test for multiple comparisons. k Quantification of PKD1-dependent luminescence to assess PKD1 knockout efficiency in HEK293T cells transiently transfected with PKD1-Nluc alone (Control), and with different CRISPR human PKD1 KO constructs (PKD1KO #1, PKD1KO #2, PKD1KO #3, combined PKD1KO #1-3). Measurements were conducted 72 hours post-transfection (j and k). Data are represented as mean ± SEM of n = 3 (b–i), n = 5 (j), or n = 4 (k) experimental replicates.
Next, we determined if G protein selectivity differs in response to various WNT family members, as we have shown that in addition to WNT-9B, WNT-5A and WNT-3A bind the extracellular domain of PKD139. These experiments offered a unique opportunity to compare (a) canonical/β-catenin dependent (WNT-3A), (b) noncanonical/β-catenin independent (WNT-5A), and (c) mixed (both canonical and noncanonical, i.e., WNT-9B) pathway activating WNTs. Similarly to WNT-9B, neither WNT-5A nor WNT-3A stimulated coupling between PKD1 and Gαs or Gα13 (Fig. 1b, c). PKD1-Gα selectivity in response to WNT specificity is demonstrated by the reduction in the rate of Gαq, Gαi1, and Gαi2 dissociation from Gβγ induced by WNT-3A compared to other WNTs. WNT-3A induced slower dissociation of these subunits compared to WNT-5A and WNT-9B (Fig. 1d–f, h). Maximal dissociation of Gαi3 was seen with all three WNT ligands (Fig. 1g, h). The slower Gαq dissociation by WNT-3A compared to the other two WNTs raises the possibility that ligand binding specificity, perhaps by involving different subdomains in the extracellular domain of PKD1 and/or co-receptors, determines selectivity in heterotrimeric G protein coupling. The overall BRET response and relative PKD1-mediated dissociation rates of all six Gα subunits in response to three WNT ligands are shown in the heat map diagrams (Fig. 1h).
FZD6 has been shown to couple to Gαq and Gαi subunits in response to WNT activation45, thus, we sought to determine whether FZD6 and possibly other FZDs endogenously present in HEK293T cells could account for PKD1-induced Gαi3-CASE activation in response to WNT-9B. ZNRF3 is an E3 ubiquitin ligase that constitutively clears FZDs and LRP6 from the cell surface46. Co-expression of ZNRF3 and PKD1 did not affect the magnitude or kinetics of WNT-9B-induced Gαi3 dissociation from Gβγ, indicating that endogenous FZDs could not account for the activation of the Gαi3-CASE sensor by PKD1 (Supplementary Fig. 1g). The functionality of ZNRF3 was confirmed by its effect on overexpressed FZD6-induced activation of Gαi3-CASE (Supplementary Fig. 1h). To provide additional evidence for the preference of PKD1 towards Gαi3 we determined the effect of G protein signaling modulator 1 (GPSM1), which functions as a Gαi3-specific guanine nucleotide dissociation inhibitor47, on WNT-9B-induced dissociation of Gαi3 via PKD1. Co-expression of GPSM1 suppressed PKD1-mediated Gαi3 dissociation in response to WNT-9B (Supplementary Fig. 1i). Altogether, these data suggested that WNT ligands induce the dissociation of Gαi1-3 and Gαq, but not Gαs or Gα13 subunits from Gβγ via PKD1. These effects do not seem to involve endogenous FZD receptors and LRP6 co-receptors in transiently transfected HEK293T cells.
WNTs did not induce significant Gα dissociation for any of the six subtypes in mock (sensor only)-transfected cells in response to WNT ligands (≤5% drop in normalized BRET, Supplementary Fig. 1f) 48 h following transfection or 24 h post-seeding onto 96-well plates. However, 72 h after transfection or 48 h post-seeding, cells showed higher activity (up to 20% drop in normalized BRET) in response to WNT-9B in the absence of transfected PKD1 (Fig. 1j), perhaps by allowing more time to recover endogenous PKD1 from trypsinization after replating. We took advantage of this high signal-to-noise BRET ratio under these conditions to test whether endogenous PKD1 contributed to the WNT-induced activation of Gαi3. We knocked out human PKD1 using a combination of 3 human PKD1-specific sgRNAs and tested for WNT-9B-induced reduction of the Gαi3-CASE sensor activity 72 h following transfections (Fig. 1j). Knockout efficiency was quantitatively determined (95%) on PKD1-Nluc 72 h following transfection (Fig. 1k). Knockout of endogenous PKD1 led to a reduction in normalized BRET compared to sensor-only-transfected cells in response to WNT-9B. To confirm that this effect was PKD1-specific, we added back mouse PKD1, which is resistant to knockout by any of the 3 sgRNAs used to inactivate human PKD1. Mouse PKD1 rescued the effect of human PKD1 knockout (Fig. 1j, red versus purple). These data reveal an essential role of endogenous PKD1 in the WNT-9B-induced activation of Gαi3.
To further confirm the specificity of our results and obtain mechanistic insights of how PKD1 functions as a WNT-activated GPCR, we tested the effects of several PKD1 mutations (Fig. 2a) on Gαi1-3 dissociation from Gβγ in response to WNT-9B (Fig. 2b–d). S99I is a pathogenic mutant that compromises PKD1 cell surface expression39, and thus, it should reduce receptor function. Indeed, expression of PKD1-S99I failed to induce Gα-Gβγ dissociation. Similar results were obtained with W139C, also a pathogenic mutation located within the WNT-9B binding site. A similar effect was seen by the L4132Δ mutation, which contains a pathogenic in-frame deletion of L4132 localized within a predicted G protein-binding domain in the C-terminal tail of PKD132. Interestingly, the T3409V mutation, which renders PKD1 unable to undergo GAIN-mediated autoproteolysis29, led to an approximately 50% reduction in heterotrimeric G protein signaling (Gαi3-CASE, Fig. 2d). This result is consistent with the hypomorphic function of this mutation in mice29. Finally, the longest known pathogenic mutation, S4213X, truncates PKD1 by deleting the majority of its cytosolic C-terminal tail, including the coiled-coil domain essential for interaction with PKD2, and resulted in complete loss of receptor function. Expression levels of mutants in transiently transfected HEK293T cells are shown in Supplementary Fig. 2. Altogether, these data agree with in vivo effects, supporting a biological role of PKD1 as a WNT-activated GPCR.
a Cartoon diagram of PKD1 showing the location (indicated by arrows) mutated in tested constructs (S99I, W139C, T3049V, L4132Δ, and S4213X). Created in BioRender. Tsiokas, L. (2026) https://BioRender.com/5p1mc32. ΔBRET time course (left trace) and corresponding AUC (right bar graph) for wild-type (WT) and mutant PKD1 in response to WNT-9B (1 μg/ml, time = 0 min) using Gαi1- (b), Gαi2- (c), and Gαi3-CASE (d) sensors. Data are represented as mean ± SEM of n = 3 experimental replicates (b–d). Statistical significance was assessed by one-way ANOVA using Dunnett’s post hoc test for multiple comparisons.
Next, we wanted to assess the mechanism of G protein coupling selectivity for PKD1 and compare this with the mechanism of canonical GPCRs. In the conventional mechanism of G protein activation, the distal portion of the Gα C-terminal alpha helix engages the TM domains of the receptor during coupling (Fig. 3a, c)48,49. We designed chimeras for Gαs-CASE and Gαi3-CASE, in which the 16 final C-terminal amino acids were exchanged with that of the other (Fig. 3b, d) and conducted BRET assays as described previously to determine whether the Gα C-terminus is a key component of coupling selectivity for PKD1 in response to WNT-9B (Fig. 3e). As expected, the exchange of the Gαi3 C-terminus for that of Gαs (Gαi3-s) led to a reduction in the maximum response compared to the wild-type Gαi3. Interestingly, the Gαs to Gαi3 C-terminal swap (Gαs-i3) only resulted in a partial gain of function. A range of G protein sensor concentrations were tested to account for possible expression issues, yet the maximum response was still seen using the same concentration as the wild-type sensor (Supplementary Fig. 3a). We also confirmed these results were PKD1-specific, as sensor-only-transfected cells did not produce any response upon WNT stimulation (Supplementary Fig. 3b, c). In contrast to PKD1, Gαi-coupled CaSR and Gαs-coupled beta-2 adrenergic receptor (β2AR) each showed a complete exchange in coupling selectivity correlating with the swap in Gα C-terminal motif (Fig. 3f, g), validating the functionality of the chimeras. Compared to the CaSR and β2AR, our results for PKD1 indicate that the Gα C-terminus contributes only partially to selective coupling and suggests additional/multiple interactions between PKD1 and the G protein may dictate selectivity. Thus, the mechanisms dictating G protein coupling selectivity of PKD1 is likely different from traditional 7-TM containing GPCRs.
a Active state of the CaSR receptor coupled to heterotrimeric Gαi3 (purple), Gβ (light blue), and Gγ (light green). Assembled using PDB code 8SZH, and the extracellular domain of CaSR was removed for simplicity. b Depiction of the Gαi3-s chimera in which the C-terminal 16 amino acid region of Gαi3 (purple) within the α5 helix is swapped with that of Gαs (dark blue). c Active state of the β2AR receptor coupled to heterotrimeric Gαs (dark blue), Gβ (light blue), and Gγ (light green). Assembled using PDB 3SN6. d Depiction of the Gαs-i3 chimera in which the C-terminal 16 amino acid region of Gαs (dark blue) within the α5 helix is swapped with that of Gαi3 (purple). ΔBRET time courses for Gαs- and Gαi3-CASE wild type or chimera sensors with PKD1 (e), CaSR (f), or β2AR (g) following agonist stimulation (time = 0 min). Data are represented as mean ± SEM of n = 3 experimental replicates (e–g).
WNT-9B induces the recruitment of Gα subunits to PKD1 and Gα-GDP to Gα-GTP exchange
A hallmark of heterotrimeric G protein activation downstream of a GPCR is the exchange of GDP for GTP on Gα subunits following the recruitment of Gα subunits to the receptor1. Thus, we sought to determine whether PKD1 and Gαi1-3 and Gαq subunits are recruited to PKD1 in response to WNT-9B activation. In this series of experiments, we used PKD1-Nluc and YFP-tagged Gαq and Gαi1-3 subunits (Fig. 4a). Saturation experiments revealed saturable and thus specific interactions between PKD1 and Gαq or Gαi1-3 subunits (Fig. 4b–e), and in response to WNT-9B, these subunits are recruited to PKD1 in a time-dependent manner (Fig. 4f–i). To determine whether WNT-9B induces a PKD1-dependent Gα-GDP to Gα-GTP exchange, we employed a collection of biosensors (ONE-GO sensors) to measure G protein activation50 (Fig. 4a). WNT-9B induced GDP to GTP exchange on Gαq and Gαi1-3 subunits (Fig. 4j–m), but not Gαs or Gα13 (Supplementary Fig. 4a, b). Altogether, these results are in complete agreement with results obtained using G-CASE biosensors, further supporting the hypothesis that PKD1 is directly coupled to heterotrimeric G protein signaling for a selective subset of Gα subunits.
a Schematic representation of the PKD1-Nluc and Gα-YFP BRET assay (left panel) and ONE-GO BRET assay (right panel). Created in BioRender. Tsiokas, L. (2026) https://BioRender.com/5p1mc32. Gα-YFP was titrated with a fixed amount of PKD1-Nluc to assess WNT-independent basal recruitment of Gαq- (b), Gαi1- (c), Gαi2- (d), and Gαi3-YFP (e) to PKD1. BRET time-courses for PKD1-Nluc and Gαq- (f), Gαi1- (g), Gαi2- (h), and Gαi3-YFP (i) in response to WNT-9B (1 μg/ml, time = 0 min). ΔBRET time courses for Gαq (j), Gαi1 (k), Gαi2 (l), and Gαi3 (m) ONE-GO BRET sensors in the presence or absence of PKD1 upon WNT-9B addition (1 μg/ml, time = 0 min). Data are represented as mean ± SEM of n = 4 (b–e) and n = 3 (f–m) experimental replicates.
PKD2 enables PKD1 to function as a GPCR
PKD2 is endogenously expressed at detectable levels in HEK293T host cells (Supplementary Fig. 5a), which could influence the functional expression of PKD1 as a GPCR. Thus, we inactivated the endogenous PKD2 gene via CRISPR/Cas9-mediated genome editing in these cells using a doxycycline-inducible Cas9 vector (Supplementary Fig. 5a, lane 4). The efficiency of the PKD2-specific CRISPR construct was confirmed on transfected human PKD2 (Supplementary Fig. 5b, lane 4). Inactivation of human PKD2 led to the loss of PKD1 receptor function, indicating that endogenous PKD2 was essential for the receptor function of PKD1 in HEK293T cells (Fig. 5a and Supplementary Figs. 5c and 4, gray versus red). The functional dependence of PKD1 on endogenous PKD2 was also tested in a previously characterized clone of mouse inner medullary collecting duct (mIMCD-3) cells (Fig. 5b, gray versus red), in which endogenous Pkd2 has been permanently inactivated51. Therefore, the requirement of endogenous PKD2 for the functional expression of PKD1 as a GPCR is demonstrated in two different cell lines.
a ΔBRET time course for Gαi3-CASE (left trace) and summary data (AUC, right bar graph) for HEK293T cells co-transfected with PKD1 and pcDNA3 (control, gray), CRISPR human PKD2 KO alone (KO, red) or KO in combination with mouse wild-type PKD2 (KO + WT, orange), PKD2-E442G (KO + E442G, blue), PKD2-D509V (KO + D509V, green), PKD2-W199A (KO + W199A, purple), or PKD2-F602P (KO + F602P, yellow). Cells were stimulated with WNT-9B (1 μg/ml, time = 0 min). b ΔBRET time course (left panel) and summary data (AUC, right panel) for Gαi3-CASE in mIMCD-3 cells. Gαi3-CASE and PKD1 were transiently transfected in wild-type mIMCD-3 cells and assessed either directly in response to WNT-9B (Control, gray) or following pre-treatment with lanthanum (Control+La3+). Pkd2-null mIMCD-3 cells were transiently transfected the Gαi3-CASE, PKD1 and pcDNA3 (KO, red), or with human wild-type PKD2 (KO + WT, orange), PKD2-W201A (KO + W201A, purple), or PKD2-F604P (KO + F604P, yellow). Cells were stimulated with WNT-9B (2 μg/ml, time = 0 min). c Subcellular co-localization of C-terminally tagged mouse PKD1 with enhanced GFP (PKD1-eGFP, green) and HaloTag-CAAX construct (HT-CAAX, red) in the presence of untagged wild-type PKD2 (WT), PKD2-D509V (D509V), PKD2-E442G (E442G), PKD2-W201A (W201A), or PKD2-F604P (F604P) in HEK293T cells. Scale bar: 10 μm. d Co-localization of PKD1-eGFP (green) and C-terminally HaloTag-labeled PKD2 (PKD2-HT, red). Arrows indicate cell surface-expression of PKD1-eGFP and PKD2-HaloTag. Scale bar: 10 μm. e Quantification of BRET (left panel) or PKD1-based luminescence (right panel) for PKD1-Nluc and HaloTag-CAAX in HEK293T cells co-transfected with pcDNA3 (Control, gray), CRISPR human PKD2 KO (KO, red), or KO in combination with mouse PKD2-wildtype (KO + WT, orange) or PKD2-E442G (KO + E442G, blue). f Net BRET between PKD1-Nluc and wild type PKD2-HaloTag (WT, orange) or PKD2-E442G-HaloTag (E442G, blue). g Cartoon illustration to model the chaperone activity of PKD2 in trafficking PKD1 to the cell surface. Created in BioRender. Tsiokas, L. (2026) https://BioRender.com/5p1mc32. Data are represented as mean ± SEM of n = 3 (a, b, e, f) experimental replicates. Statistical significance was assessed by one-way ANOVA using Tukey’s post hoc test for multiple comparisons. Live-cell imaging data (c, d) are shown as representative images from 3 independent experiments.
To gain insight into the mechanism by which PKD2 supported the receptor function of PKD1, we used the following PKD2 variants: D511V (or D509V in mouse PKD2), E442G (mouse PKD2), W201A (or W199A in mouse PKD2), and F604P (or F602P in the mouse). The D509V mutation compromises PKD2 channel activity and impairs maturation and targeting of PKD1 to the plasma membrane52. W199A reduces channel activity of PKD253, whereas F602P increases PKD2 channel activity54,55. Finally, E442G does not affect channel function but impairs PKD2 trafficking to the primary cilium56. Homozygous Pkd2lrm4 (E442G) mice show a strong PKD phenotype, indistinguishable from that produced by complete Pkd1 or Pkd2 deletion57. Adding back PKD2-E442G or PKD2-D509V in PKD2-null cells failed to restore PKD1 activity (Fig. 5a, blue and green, respectively). Co-expression of PKD1-eGFP with wild-type untagged PKD2 or PKD2 C-terminally fused to HaloTag led to robust expression of both proteins at the plasma membrane (Fig. 5c, d). As expected, PKD2-D509V did not drive expression of PKD1-eGFP to the plasma membrane (Fig. 5c, D509V panels). Interestingly, PKD2-E442G also failed to target PKD1-eGFP to the plasma membrane (Fig. 5c, E442G panels). PKD1-Nluc co-localization with HaloTag-CAAX determined by BRET quantifiably supports live-cell imaging data (Fig. 5e). Notably, overexpression of wild-type PKD2 increased overall levels of PKD1 (Fig. 5e and Supplementary Fig. 2), possibly via protein-protein-mediated stabilization, as has been shown using C-terminal tails of PKD1 and PKD258. PKD2-E442G showed reduced stabilization of PKD1 compared to wild type (Fig. 5e), implying that E442G may affect PKD1/PKD2 assembly. The E442G mutation is located in the TOP domain of PKD2, which forms a tetrameric structure with 4-fold symmetry in homomeric PKD2 complexes59 and a similar structure but with a 15° deviation from 4-fold symmetry in the heteromeric PKD1/PKD2 complex16. Since E442G did not impede channel function56, we reasoned that it should not affect self-assembly but rather assembly with PKD1. As expected, BRET experiments showed highly efficient energy transfer between wild-type PKD1-Nluc and PKD2-HaloTag (Fig. 5f). However, BRET was severely reduced by the E442G mutation (PKD2-E442G-HaloTag), suggesting that E442G interferes with PKD1/PKD2 complex assembly and thus fails to target PKD1 to the plasma membrane. Expression of human PKD2-W201A or human PKD2-F604P in HEK293T cells did not affect PKD1 expression at the plasma membrane (Fig. 5c, W201A and F604P panels). In agreement with this finding, expression of mouse PKD2-W199A or mouse PKD2-F602P in Pkd2-null HEK293T cells and human PKD2-W201A or human PKD2-F604P in Pkd2-null mIMCD-3 cells did not impair Gαi3 dissociation from Gβγ in response to WNT-9B (Fig. 5a, b). Furthermore, treatment of wild-type mIMCD-3 cells with La3+ which blocks cation permeation of PKD1/PKD239, PKD2, or even structurally related PKD2-L160 did not have a significant effect on the GPCR function of PKD1 (Fig. 5b). Collectively, these data suggest that PKD2 enables PKD1 to function as a GPCR by chaperoning it to the plasma membrane, independently of its channel function (Fig. 5g). They further implicate the loss of PKD1 GPCR function as an alternative mechanism underlying the cystogenesis observed in Pkd2lrm4 (E442G) mice57.
PKD1-mediated Gαi1-3 activation inhibits cAMP accumulation
Gαi1-3 signaling leads to inhibition of cAMP accumulation (Fig. 6a). Therefore, we tested whether overexpression of PKD1 can inhibit basal or forskolin-induced cAMP accumulation in response to WNTs. We employed GloSensor technology, a cAMP assay to monitor changes in cytosolic cAMP levels in PKD1-transfected cells using two complementary strategies: (i) cells were treated with a concentration of forskolin close to its EC50 (10-6M) (Fig. 6b, c) and soluble WNTs were added when cAMP had reached its half-maximal concentration (Fig. 6c). In strategy (ii), endpoint analysis was used whereby PKD1-transfected cells were pretreated overnight with pertussis toxin (PTX, 100 ng/ml) to block Gαi1-3 signaling, then treated with forskolin in the presence or absence of soluble WNTs for 10 min and cAMP levels determined (Fig. 6d). Kinetic analysis showed that WNTs suppressed the dynamic accumulation of cAMP via PKD1 (Fig. 6c). WNT-3A had the lowest effect (Fig. 6c), which was consistent with the fact that it is most efficiently coupled to Gαi3 (Fig. 1g, h), which shows lower endogenous expression than Gαi2 in HEK293T cells (Supplementary Fig. 6a). Gαi1 is practically undetectable in HEK293T cells (Supplementary Fig. 6a). Similar results were obtained using the endpoint analysis, whereby PTX suppressed WNT-induced inhibition of cAMP accumulation (Fig. 6d). We next asked whether WNT ligands can lower basal (resting) levels of cytosolic cAMP in the presence of PKD1. WNT-9B induced a time-dependent reduction in cytosolic cAMP concentration in cells transfected with PKD1 (Fig. 6e, left trace). A similar but smaller effect was seen with WNT-5A or WNT-3A (Fig. 6e, center and right traces, respectively). The PKD1-mediated reduction in basal cAMP levels in response to WNT-9B was approximately half of the response elicited by the well-characterized muscarinic acetylcholine receptor M2 (M2R) following activation with carbachol in HEK293T cells (Supplementary Fig. 6b). Expression of pathogenic PKD1 mutants showed markedly suppressed inhibition of steady-state cAMP levels compared to wild-type PKD1 (Fig. 6f). To determine whether PKD2 had an essential role in WNT-induced inhibition of cAMP concentration via PKD1, we used our endpoint analysis for cAMP in cells lacking endogenous PKD2 and cells in which wild-type PKD2 was added back. These experiments support the role of PKD2 in inhibiting forskolin-induced cAMP accumulation in response to WNT-9B (Fig. 6g). We confirmed these results in wild-type and Pkd2-null mIMCD-3 cells using the end-point analysis. WNT-9B produced a significant reduction in forskolin-induced cAMP levels in wild-type, but not in Pkd2-null cells (Fig. 6h). Interestingly, deletion of Pkd2 did not affect resting cAMP levels. In summary, these data suggest that Gαi1-3 signaling mediated by WNT-induced activation of PKD1 is sufficient to reduce steady-state and forskolin-induced cAMP accumulation, and this regulation is compromised in Pkd2-null cells (Fig. 6i).
a Schematic of the GloSensor cAMP assay. Created in BioRender. Tsiokas, L. (2026) https://BioRender.com/5p1mc32. b Dose-response curve for forskolin-cAMP accumulation detected by the pGloSensor-22F cAMP biosensor in HEK293T cells. c Analysis of the PKD1-mediated effect on 1 μM forskolin (FSK)-induced time-dependent cAMP accumulation in HEK293T cells. Time course of cAMP levels before and following WNT stimulation (1 μg/ml, left panel) and summary data depicting the changes in cAMP concentration (ΔcAMP) 10 min after treatment. d Endpoint analysis of PKD1-mediated inhibition of cAMP in HEK293T cells treated with different WNT ligands (1 μg/ml) followed by 1 μM FSK in the presence or absence of 0.1 μg/ml pertussis toxin (PTX). e Kinetic analysis of PKD1-mediated basal cAMP changes in response to 1 μg/ml of WNT-9B (left panel), WNT-5A (center panel), or WNT-3A (right panel) at time = 0. f Time course (left trace) and summary data (AUC, right bar graph), of basal cAMP changes mediated by wild type and mutant PKD1 in response to WNT-9B (1 μg/ml, time = 0). g Endpoint analysis of cAMP levels in HEK293T cells transiently transfected with pGloSensor-22F, PKD1, and either pcDNA3 (Control, gray), CRISPR human PKD2 KO (KO, red), or PKD2 KO+mouse wild type PKD2 (KO + WT, orange). h Endpoint analysis of cAMP levels in wild type (Control, gray) or stable PKD2 KO (KO, red) mIMCD-3 cells transiently transfected with pGloSensor-22F and PKD1. Cells were stimulated with 2 μg/ml WNT-9B. i Diagrammatic representation of a model for the regulation of cytosolic cAMP levels by PKD1/PKD2. In the model, regulation of cAMP levels is regulated via a direct effect of WNT-activated PKD1/PKD2 on Gαi1-3 subunits. Created in BioRender. Tsiokas, L. (2026) https://BioRender.com/5p1mc32. Data are represented as mean ± SEM of n = 3 (b–h) experimental replicates. Statistical significance was assessed by one-way ANOVA using Tukey’s post hoc test for multiple comparisons.
GRK6 promotes the internalization of PKD1 and suppresses PKD1-mediated signaling
Typically, GPCR signaling is regulated through the process of desensitization, whereby G protein-coupled receptor kinases (GRKs) are recruited to the activated receptor to phosphorylate specific sites, “priming” the receptor for recruitment of beta-arrestins (ARRBs), which signal receptor internalization and degradation or recycling. Alternatively, GRK recruitment can initiate ARRB-dependent signaling that is independent of Gα/Gβγ signaling. In either case, some intrinsic affinity of the receptor for GRKs is needed to initiate desensitization and/or ARRB-based signaling. Based on this, we tested whether PKD1 is localized proximally to GRK2, GRK3, GRK5, or GRK6. BRET experiments using PKD1-Nluc and the different GRK subtypes fused to HaloTag revealed saturable and specific recruitment of GRK2, GRK5, and GRK6, but not GRK3 to PKD1-Nluc (Fig. 7a–d). Interestingly, the maximum BRET between PKD1 and GRK6 was increased upon overexpression of PKD2, but to a lesser degree by overexpression of PKD2-ΔC which lacks the C-terminal tail of PKD2 (Fig. 7d) and fails to target PKD1-eGFP to the plasma membrane (Fig. 7f). The opposite effect was seen with GRK2 (Fig. 7c). Since GRK2 is expressed predominantly in the cytoplasm, while GRK6 is anchored to the plasma membrane via palmitoylation modifications in its C-terminus61, these data suggest that PKD2 targets the PKD1/PKD2 complex to the plasma membrane where the complex is then in proximity to GRK6. This interpretation is consistent with the reduction in BRET between PKD1-Nluc and GRK2-HaloTag in the presence of PKD2 (Fig. 7c). The PKD2-ΔC construct fails to facilitate co-localization of PKD1 with GRK6 (Fig. 7e, lower panels) because it does not target the complex to the cell surface. To determine whether GRK6 functionally affects PKD1 coupling to Gαi3, we tested whether it promotes internalization and negatively affects WNT-induced PKD1 signaling. Overexpression of GRK6 did not affect overall receptor levels (Fig. 7g, right bar graph) but did result in the reduction of PKD1 from the cell surface (Fig. 7g, left bar graph) and WNT-induced PKD1-mediated Gαi3 dissociation from Gβγ (Fig. 7h, DMSO). Inhibition of GRK6 activity using two different GRK6 inhibitors, GRK6-IN-1 or GRK6-IN-2, for 1 h prior to WNT-9B activation partially reverted PKD1/PKD2-mediated Gαi3 coupling in the presence of GRK6, indicating that the kinase activity of GRK6 was essential for its effect on PKD1/PKD2 function (Fig. 7h). Given the short-term effect of GRK6 inhibition (1 h due to the toxicity of the inhibitors) on PKD1 internalization, we consider the partial reversal on PKD1/PKD2 function significant.
GRK3- (a), GRK5- (b), GRK2- (c), and GRK6-HaloTag (d) were titrated with a fixed amount of PKD1-Nluc to assess basal recruitment to PKD1. Recruitment of GRK2- and GRK6-HaloTag was assessed in HEK293T cells co-transfected with pcDNA3 (- PKD2, cyan for GRK2-HaloTag, blue for GRK6-HaloTag), wild-type PKD2 (WT, orange), or PKD2-ΔC (ΔC, pink). e Co-localization of C-terminally tagged mouse PKD1 with enhanced GFP (PKD1-eGFP, green) and GRK6-HaloTag (red) in the presence of untagged wild type PKD2 (WT, upper panels) or PKD2-ΔC (ΔC, lower panels) in HEK293T cells. f Co-localization of PKD1-eGFP (green) and the HaloTag-CAAX construct (red) in the presence of untagged wild-type PKD2 (WT, upper panels) or PKD2-ΔC (ΔC, lower panels) in HEK293T cells. Arrows indicate cell surface-expression, scale bar: 10 μm (e, f). g Quantification of BRET (left panel) and PKD1-based luminescence (right panel) for PKD1-Nluc and HaloTag-CAAX in HEK293T cells co-transfected with untagged GRK6 at varied concentrations. Statistical significance was assessed by one-way ANOVA using Dunnett’s post hoc test for multiple comparisons. h ΔBRET time courses for Gαi3-CASE in HEK293T cells transiently co-transfected with PKD1 and pcDNA3 (- GRK6, gray square) or with untagged GRK6 (+GRK6, blue circle). Cells were pretreated with DMSO (left panel), GRK6-IN-1 (25 μM, center panel), and GRK6-IN-2 (25 μM, right panel) for 1 h before WNT-stimulation (1 μg/ml, time = 0 min). Data are represented as mean ± SEM of n = 3 (a–d, g, h) experimental replicates. Live-cell imaging data (e, f) are shown as representative images from 3 independent experiments.
ARRB2 functions downstream of GRK6 in mediating internalization of the PKD1/PKD2 complex
PKD1 has been shown to interact with ARRB2 via its Polycystin-1, lipoxygenase, and α-toxin (PLAT) domain located in the intracellular loop between the first and second TM domains62. BRET saturation assays confirmed specific binding of HaloTag-fused ARRB2, but not ARRB1, to C-terminally-tagged PKD1-Nluc (Fig. 8a, b). Kinetic analysis revealed time-dependent recruitment of ARRB2-HaloTag to PKD1-Nluc in response to WNT-9B, which was unaffected by overexpression of PKD2 (Fig. 8c). However, time-dependent recruitment of ARRB2-HaloTag to Nluc-tagged PKD2 (PKD2-Nluc) was found to be PKD1-dependent (Fig. 8c), suggesting that recruitment of ARRB2 to PKD2 is indirect via PKD1 and that ARRB2 is recruited to the PKD1/PKD2 complex. WNT-9B-induced recruitment of ARRB2-HaloTag to PKD1-Nluc was reduced when cells were first incubated with GRK6 inhibitors, supporting the role of endogenous of GRK6 in the recruitment of ARRB2 to the PKD1/PKD2 complex (Fig. 8d).
ARRB1- (a) and ARRB2-HaloTag (b) ARRB-HaloTag were titrated with a fixed amount of PKD1-Nluc to assess basal recruitment to PKD1. c ΔBRET time course (left panel) and summary data (AUC, right panel) for ARRB2-HaloTag recruitment to PKD1 and/or PKD2. HEK293T cells were transiently transfected with ARRB2-HaloTag, PKD1-Nluc, and pcDNA3 (PKD1-Nluc, gray circle) or PKD2 (PKD1-Nluc + PKD2, pink circle). Vice versa, cells were transfected with ARRB2-HaloTag, PKD2-Nluc, and pcDNA3 (PKD2-Nluc, gray square) or PKD1 (PKD2-Nluc + PKD1, pink square), and responses recorded before and after WNT-9B addition (1 μg/ml, time = 0 min). Statistical significance was assessed by one-way ANOVA using Tukey’s post hoc test for multiple comparisons. d ΔBRET time course (left trace) and summary data (AUC, right bar graph) for ARRB2-HaloTag recruitment to PKD1-Nluc in response to WNT-9B (1 μg/ml, time = 0 min) following pre-incubation with DMSO or GRK6-IN-1 + GRK6-IN-2 (15 μM) for 1 h. Statistical significance was assessed by unpaired t test with Welch’s correction (two-tailed). e Quantification of basal BRET for HaloTag (HT)-CAAX, HT-Rab5, or HT-FYVE and PKD1-Nluc (left panel) or PKD2-Nluc (right panel). f Kinetic analysis of ΔBRET for HT-CAAX, HT-Rab5, or HT-FYVE and PKD1-Nluc (left trace) or PKD2-Nluc (right trace) in response to WNT-9B (1 μg/ml, time = 0 min). g ΔBRET time course for PKD1-Nluc and HT-CAAX in response to WNT-9B following pre-incubation with barbadin (100 μM). h Diagrammatic presentation of a model for the function of GRK6 and ARRB2 in PKD1-mediated signaling. In the model, GRK6 and ARRB2 function within a pathway that signals PKD1 internalization and/or ARRB-mediated signaling following WNT activation of PKD1. Created in BioRender. Tsiokas, L. (2026) https://BioRender.com/5p1mc32. Data are represented as mean ± SEM of n = 3 (a, b, e–g) or n = 4 (c, d) experimental replicates.
To provide further evidence that the PKD1/PKD2 complex, rather than PKD1 alone, is necessary for GPCR function at the plasma membrane, we determined the kinetics for WNT-9B-induced internalization of PKD1-Nluc and PKD2-Nluc from the plasma membrane and for accumulation of these proteins in early endosomes. We reasoned that if PKD2 has an initial chaperone function to deliver PKD1 to the cell surface and then to dissociate from PKD1 after successful delivery, PKD2 would not show the same time-dependent internalization as PKD1 in response to a ligand. Figure 8e–g shows that this is not the case. Steady-state BRET measurements and super-resolution imaging confirmed proximity and co-localization of PKD1-Nluc or PKD2-Nluc to HaloTag-Rab5 and HaloTag-FYVE1 (Fig. 8e and Supplementary Fig. 7), which marks the early endosome. WNT-9B stimulation in BRET time-course assays revealed similar kinetic profiles for PKD1-Nluc and PKD2-Nluc for both internalization and accumulation in the early endosome (Fig. 8f). The internalization of PKD1 in response to WNT-9B was inhibited by barbadin, an ARRB inhibitor (Fig. 8g). These results indicate that PKD2 remains associated with PKD1 during ligand activation and both proteins move together to the early endosome following activation (Fig. 8h).
Discussion
Collectively, our data show that PKD1 mediates outside-in signaling, leading to the reduction of cAMP levels via direct coupling to Gαi1-3 subunits in response to WNT ligands. PKD1-mediated activation of G protein signaling is independent of FZD receptors and occurs immediately after the addition of soluble WNTs, representing the fastest response of PKD1 to an activating event that we observed. PKD1 is the first example of an integral membrane protein without the 7-TM protein domain of typical GPCRs that can be efficiently coupled to heterotrimeric G proteins.
Direct coupling of PKD1 to G protein signaling is supported by three lines of complementary evidence: (a) WNT-induced PKD1-mediated Gα-Gβγ dissociation detected by G-CASE sensors, (b) Gα-GDP to Gα-GTP conversion detected by ONE-GO sensors, and (c) direct WNT-induced Gα subunit recruitment to PKD1. While it has been speculated that PKD1 may function as a GPCR28, this hypothesis has never been tested directly in real time using full-length PKD1 activated in a ligand-dependent manner. Our studies clearly show that this new role for PKD1 is the case. Based on our previous studies, whereby soluble ligands bind with high affinity to a short segment of the extracellular domain of PKD139, we propose that WNT ligands stabilize PKD1 in an active conformation that allows it to interact with Gαi1-3 and Gαq subunits, leading to their activation.
Our data support the hypothesis that PKD2 assembles with PKD1 and facilitates its targeting to the plasma membrane, where they can function together as a complex. The channel activity of PKD2 is not required for the function of the complex as a GPCR. These conclusions are supported by the following lines of evidence: (a) the D509/511 V PKD2 variant fails to target PKD1 to the plasma membrane and does not support its function as a GPCR, (b) E442G, which compromises complex assembly has similar effects on targeting PKD1 to the cell surface and GPCR function, (c) W199A/201 or F602/604 P, which function diametrically opposite in regard to PKD2 channel activity, do not affect PKD1/PKD2’s GPCR function, (d) pre-treatment the cells with La3+, a non-specific inhibitor of the cation permeation of PKD1/PKD2, PKD2, or a structurally-related PKD2-L1 channel did not affect GPCR function of PKD1, and (e) PKD1 or PKD2 follow the same time-dependent removal from the cell surface and accumulation in the early endosome in response to a WNT ligand. As only activated receptors follow β-arrestin-mediated internalization, these data suggest that PKD1 and PKD2 remain bound together during ligand activation. Given that PKD2 does not directly interact with WNTs39, the simpler conclusion is that the PKD1/PKD2 complex, rather than PKD1 alone, functions as a GPCR at the cell surface. Whether PKD2 has functions in addition to its chaperoning function to modulate GPCR signaling at the plasma membrane requires further work.
Previous work has shown that the C-terminal tail of PKD1 binds a wide spectrum of G proteins with the exception of Gα1330,31,63. Knockdown of Gαi1-3 and all Gαq homologs in Xenopus embryos leads to a PKD phenotype31. Furthermore, exogenous expression of full-length PKD1 or a 1,811 residue N-terminally truncated version activates Gαi/o proteins in freshly isolated sympathetic neuron cells37. Overexpression of PKD2, however, suppresses PKD1 coupling to Gαi/o proteins37. Similarly, constitutive activation of the PKD2 channel in dorsal root ganglia cells is suppressed by PKD1 overexpression36. It was suggested that a critical balance of PKD1 and PKD2 is essential to maintain the complex in an inactive or “silent” state, so that it can only be activated by the appropriate stimulus36. Our data support and extend this model in several respects: (1) both models converge to the point that PKD1 can activate Gαi proteins, (2) our model extends the previous model by demonstrating that PKD1 directly activates Gαi in response to an activating ligand, in real-time, (3) our model incorporates an essential role of endogenous PKD2 in the function of PKD1 as a GPCR, which was not examined in the 2002 study37. (4) Both models support the concept that a critical balance of PKD1 and PKD2 is needed to achieve maximal GPCR activity. In the 2002 study, overexpression of PKD2 suppressed the PKD1-mediated activation of Gαi/o, and in our study, excessive expression of PKD2 did not fully rescue PKD1/PKD2 function in Pkd2-null HEK393T or mIMCD-3 cells, probably by exceeding the threshold for the formation of functional complexes. The concept of a critical threshold for a functional PKD1/PKD2 complex formation is supported by in vivo experiments whereby transgenic mice overexpressing PKD1 or PKD2 show cystic disease64,65.
We show that PKD1 is directly and selectively coupled to Gαi1-3 and Gαq subunits, but not to Gαs or Gα12/13, in response to WNT ligands. The coupling of Gαi1-3 and ensuing inhibition of cAMP accumulation is extremely relevant to ADPKD, as abnormal cytosolic cAMP accumulation and cAMP-dependent signaling are hallmarks of cystic cells derived from ADPKD patients or mice lacking Pkd1 or Pkd29. The prevailing dogma was that accumulated cAMP in these cells is due to reduced Ca2+ signaling mediated by the PKD1/PKD2 channel complex20,66,67, thus a more downstream event. Our data support the alternative, but not mutually exclusive possibility that abnormally high levels of cytosolic cAMP levels are due to impaired Gαi1-3 signaling at a much earlier stage, directly involving PKD1/PKD2. As the receptor function of the PKD1/PKD2 complex does not require Ca2+ permeation, our data are compatible with cryo-EM data of the PKD1/PKD2 complex, whereby three positively charged amino acid residues of PKD1 occlude the PKD1/PKD2 ionic pore16. We hypothesize that while individual PKD1 and PKD2 proteins bear “TRP channel-like” domains, when they form a complex, not only is the channel function blocked, but a GPCR function is enabled. Whether channel activation is induced downstream of G protein dissociation requires further studies.
PKD1/PKD2 does not only mediate ligand-dependent Gα-mediated signaling, but also GRK/ARRB-mediated recruitment and internalization. We show that PKD1/PKD2 is in proximity to GRK6, whose activity contributes to PKD1/PKD2 internalization and ARRB2 recruitment. While we show ligand- and time-dependent recruitment of ARRB2 to PKD1 or PKD2, we could not demonstrate time-dependent recruitment of GRK6 to PKD1 or PKD2 in response to WNT-9B by BRET. We speculate that GRK6 and PKD1 are already in such proximity before receptor activation that “additional” GRK6 recruitment cannot be resolved using BRET. Although GRK6 belongs to a subset of GRKs, including GRK4 and GRK5 that can constitutively phosphorylate GPCRs in the non-activated state68,69, we do not believe that GRK6 constitutively primes PKD1 for ARRB2 recruitment in the non-activated state, because ARRB2 does show ligand-dependent recruitment, indicating that an activating event for ARRB2 recruitment is needed.
We show that PKD1 is a unique type of GPCR containing 11- rather than 7-TM segments typically seen in classical GPCRs. Notably, PKD1 has a distinct sequence and structure, not simply an additional extension of the “typical 7-TM protein” domain seen in canonical or non-traditional GPCRs, such as LYCHOS (or GPR155), a hybrid of a transporter and a GPCR containing a total of 17-TM helices70. LYCHOS was shown to function as a GPCR71,72. Thus, our results expand the general signaling paradigm for the GPCR group of cell surface receptors. The unique structural features and functional properties of the Polycystin complex described here lead us to propose that the Polycystins (PKD1 and PKD2) and possibly Polycystin-like molecules (i.e., PKD1-L (1-3), PKD2-L (1-3)) form a new class of GPCRs. Consistently, PKD1-L2 binds a subset of Gα subunits73.
Our study has the following limitations. Most of the experiments were done using overexpression systems, where ratios of PKD1, PKD2, and various sensors were optimized for maximal coupling. However, these ratios may not represent physiological levels of endogenous components. This limitation was mitigated by loss-of-function experiments, where endogenous PKD1 or PKD2 was inactivated. In addition, the BRET assay does not detect direct protein-protein interactions, but proximity between two proteins down to 10 nm distance. Despite this caveat, it allows the detection of protein-to-protein recruitment in real time in response to a ligand, which is critical in our study. Finally, our study does not provide a structural basis for the PKD1/Gα protein coupling. To date, no high-resolution structure of full-length PKD1 has been feasible. The cryo-EM structure of a truncated PKD1/PKD2 complex has been reported16, but omits key regions of PKD1, including much of its extracellular, ligand-binding, and cytoplasmic domains that mediate G protein signaling. Without full structural coverage, critical aspects of PKD1’s interaction with signaling proteins remain speculative. In summary, while the inability to structurally resolve full-length PKD1 remains a substantial limitation, this study successfully circumvents these technical barriers, shifting the field from correlative speculation to functional demonstration of the role of PKD1 as a ligand-responsive GPCR.
Overall, our data define the molecular function of the Polycystin complex and directly link it to cAMP metabolism, putting forth the notion that impaired PKD1-Gαi1-3 coupling is the root cause of cyst formation in ADPKD. This idea gives a broader perspective of the mechanisms underlying cystogenesis in ADPKD and can serve as the springboard for future studies to address specific downstream functions of the PKD1 GPCR in ADPKD. Our results have implications in understanding biological processes that go far beyond ADPKD, as they shed light onto a wide spectrum of biological processes that involve WNT- and PKD1/PKD2-mediated signaling cascades. Therefore, understanding how this unique ligand/receptor system works will impact research efforts on many other biological processes in various tissues and organs.
Methods
Plasmids
Human wild-type PKD1-HA and human PKD1 T3049V were provided by Feng Qian. Human PKD1 L4132Δ-HA and human PKD1 S4213X-HA were provided by Stephen Parnell32. Mouse PKD1-eGFP was provided by Chris Ward (University of Kansas Medical Center). Mouse Flag-PKD1 was provided by J. Yang (Columbia University). Human PKD1 gRNAs (1-3) and Cas9 were purchased from Addgene (cat #76841, #76842, #76843, and #52962, respectively). To generate human PKD1-Nluc, Nluc was cloned out of the pNLF1-C [CMV/Hygro] vector (Promega) and into PKD1-HA before the HA tag. Mouse PKD2 was made by PCR amplification of mouse cDNA derived from a mouse embryonic kidney cDNA library and cloned into pcDNA3. Human PKD1 W139C-HA39, human PKD1 S99I-HA39, mouse PKD2 E442G, mouse PKD2 D509V, mouse PKD2 W199A, mouse PKD2 F602P, human PKD2 W201A, and human PKD2 F604P were generated using the QuikChange mutagenesis kit (Agilent Technologies). Human PKD1 ΔC was generated by cytoplasmic C-terminal region deletion using pCS2+MT-6xMyc-hPKD2 (Addgene #21370). Mouse PKD2-HaloTag was generated by cloning the mouse PKD2 into the pHTC HaloTag CMV-neo vector (Promega), and mouse PKD2 E442G-HaloTag was generated by mutagenesis. Mouse PKD1-Nluc was generated by cloning the mouse PKD2 in the pNLF1-C [CMV/Hygro] vector (Promega). eTLCV-hPKD2 was generated by cloning the guide RNA for human PKD2 into a modified version (eTLCV2) of the TLCV2 vector backbone (Addgene #87360). Gs-CASE (Addgene #168124), G13-CASE (Addgene #168127), Gq-CASE (Addgene #168125), Gi1-CASE (Addgene #168120), Gi2-CASE (Addgene #168121), and Gi3-CASE (Addgene #168122) were gifts from Gunnar Schulte. The Gi3-s and Gs-i3 chimeras were made in the Gi3-CASE and Gs-CASE as follows: For the Gi3-s chimera, the final 16 C-terminal amino acids of human Gi3 were replaced with those of human Gs (CRDIIQRMHLRQYELL). For the Gs-i3 chimera, the final 16 amino acids of human Gs were replaced with those of human Gi3 (VTDVIIKNNLKECGLY). Galpha-s ONE-GO (Addgene #189731), Galpha-13 ONE-GO (Addgene #189740), Galpha-q ONE-GO (Addgene #189738), Galpha-i1 ONE-GO (Addgene #189735), Galpha-i2 ONE-GO (Addgene #189736), and Galpha-i3 ONE-GO (Addgene #189737) were gifts from Mikel Garcia-Marcos. Galpha-q-YFP, Galpha-i1-YFP, Galpha-i2-YFP, and Galpha-i3-YFP were generated by cloning out the IRES and Nluc components from the ONE-GO biosensors. Mouse GRK2-HaloTag, mouse GRK3-HaloTag, GRK5-HaloTag, human GRK6-HaloTag, ARRB1-HaloTag, ARRB2-HaloTag, HaloTag-CAAX. HaloTag-Rab5 and HaloTag-FYVE were obtained from Mohiuddin Ahmad. Human GRK6 and β2AR were obtained from Augen Pioszak. M1R-HA and M2R were generated using RT-PCR from a human fetal kidney cDNA library. CaSR was obtained from Wenhan Chang (UCSF). Mouse FZD6 in SPORT6 was obtained from Horizon Discovery Collection of cDNA & ORF clones and verified by Sanger sequencing. ZNRF3 was obtained from Dr. Cong (Novartis). Human GPSM1 was purchased from Sino Biological (HG19737-UT). The pGloSensor-22F cAMP Plasmid was purchased from Promega.
Recombinant proteins and drugs
Recombinant human WNT-3A (5036-WN), human/mouse WNT-5A (645-WN), and mouse WNT-9B (3669-WN) were purchased from R&D Systems/Biotechne. WNTs were dissolved to 100 µg/ml in filter-sterilized in PBS containing 0.1% bovine serum albumin (BSA) and stored at −20 °C in low-protein-binding microcentrifuge tubes (Eppendorf). For stimulation experiments, WNTs were diluted in Opti-MEM I Reduced Serum Medium, no phenol red (Gibco), containing 0.1% BSA. The following drugs were stored at −20 °C as stock solutions and diluted in PBS or assay medium on the day of experiments: R568, Spermine, Carbachol, Isoproterenol, Doxycycline, Forskolin (Thermofisher, J63292.MA), Pertussis Toxin (Thermofisher, PHZ1174), GRK6-IN-1 (MedChemExpress, HY-142812), GRK6-IN-2 (MedChemExpress, HY-142817), and Barbadin (MedChemExpress, HY-119706). Lanthanum was prepared from powder the day of the experiment (Sigma-Aldrich, 262072).
Cell lines
HEK293T cells (ATCC, CRL-3216) were maintained in DMEM (Corning) supplemented with 10% FBS (GeminiBio). Wild-type and Pkd2-null mouse inner medullary collecting duct (mIMCD-3) cells were obtained from Dr. Steve Kleene51 and maintained in DMEM:F12 (Gibco) supplemented with 10% FBS. All cell lines were cultured in a humidified incubator at 37 °C and 5% CO2. Absence of mycoplasma contamination was routinely confirmed by PCR.
Transient transfections
On the day of transfection, cells were trypsinized and diluted in cell culture medium to a final density of 2 × 105 cells/ml. Subsequently, 2 ml of cell suspension (400,000 cells) was plated in each well in a sterile six-well plate and incubated for 4-6 h at 37 °C, 5% CO2. HEK293T cells were transiently transfected using the calcium phosphate precipitation method. mIMCD-3 cells were transfected using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s recommendations. For all experiments, cells were transiently transfected with up to 4 μg total DNA per well in a 6-well plate. Transfected cells were then incubated for 24 h at 37 °C, 5% CO2. Plasmid amounts and combinations used for transient transfections are detailed in Supplementary Table 2.
Bioluminescence resonance energy transfer assays
Twenty-four hours post-transfection, cells were trypsinized and resuspended in cell culture medium. Following centrifugation, cells were resuspended in assay medium (Opti-MEM I Reduced Serum Medium, no phenol red, supplemented with 4% FBS) at a final density of 2 × 105 cells/ml, and 100 μl dispensed per well of a sterile white 96-well microplate (Greiner Bio-One) and incubated for 24 h at 37 °C, 5 % CO2. For BRET assays using Nluc- and HaloTag-fused proteins, half of the cells were treated with 100 nM HaloTag NanoBRET 618 Ligand (Promega) and the other half with 0.1% DMSO (no-ligand control), and 100 μl of both cell suspensions dispensed into separate wells. 24 h post seeding, 25 μl of furimazine (Promega) was added to each well (final dilution 1:1000) and incubated for 5 min at room temperature. All measurements were performed using a Synergy Neo2 hybrid multi-mode microplate reader (BioTek) equipped with filters to simultaneously measure the donor and acceptor emissions (450/50 and 610LP for BRET experiments with Nluc- and HaloTag-fused proteins, or 460/540 for experiments with G-CASE sensors, ONE-GO sensors, or Gα-YFP), and using an integration time of 200 ms. For stimulation experiments, baseline measurements were acquired for 5 min before manual addition of 25 μl of compound or vehicle control (assay medium) to the final concentration indicated in individual figure panels and/or legends. Signal was then measured for an additional 10–20 min. Assays done using BRET-based G protein biosensors were conducted at 27 °C. Assays to measure Nluc- and HaloTag-fused protein interactions were conducted at room temperature. BRET data were collected using BioTek Gen5 software for detection (Agilent).
Glosensor cAMP assays
Approximately 400,000 HEK293T cells were seeded on each well of 6-well plates and transfected with plasmids 18 h later. Cell medium was replaced 6 h after transfections, and cells incubated 24 h at 37 °C with 5% CO2. For experiments in which the PKD2 CRISPR KO was transfected, medium was supplemented with doxycycline (2 μg/ml). Subsequently, cells were rinsed with PBS, trypsinized, and resuspended in 2 ml of cell culture medium. Following centrifugation, cells were resuspended in fresh cell culture medium at a final density of 2 × 105 cells/ml, and 100 μl dispensed per well of a white 96-well microplate and incubated for 20 h at 37 °C, 5 % CO2. For end-point analyses to determine PTX-sensitivity, the replacement medium was supplemented with 0.1 μg/ml PTX. The following day, medium was replaced with 100 μl of equilibration medium (88% CO2-independent medium, 10% FBS, 2% GloSensor cAMP Reagent stock solution in HEPES buffer) per well. Cells were then incubated for 2 h at room temperature. For dose-response experiments to assess forskolin-induced cAMP accumulation detection using the pGloSensor-22F cAMP biosensor, a pre-read measurement was taken, followed by the addition of varying concentrations of forskolin. Luminescence was measured after 10 min, and this value was divided by the pre-read measurement to determine fold response. For end-point analysis, a pre-read measurement was taken before compound addition. WNT ligands (1 μg/ml) or vehicle were then manually added to wells using a multichannel pipette and incubated for 5 min. Subsequently, forskolin (1 μM) was then manually added to all wells, and luminescence was measured 25 min later. For kinetic analysis, the plate was pre-equilibrated to the steady-state operating temperature of the microplate reader before compound addition by performing a pre-read kinetic measurement for 10 min to monitor the basal level of luminescence. For experiments to determine the inhibitory effect of WNTs on FSK-induced cAMP, following the pre-read kinetic measurement, 1 μM forskolin was added manually to each well using a multichannel pipette. Measurements were continued for 10 min, followed by manual addition of WNT (1 μg/ml) or vehicle control, and the plate was read for an additional 10 min. For experiments to measure the agonist effect on basal cAMP levels, WNT ligands (1 μg/ml) or vehicle were manually added following the pre-read kinetic measurement, and recordings continued for 10 min.
Live-cell imaging
HEK293T cells were transfected in 8-well chambered cover glasses (Cellvis) with expression plasmids. After a 24-h incubation period post-transfection, cells were labeled with 200 nM JFX 554 HaloTag Ligand (Promega) for 30 min and cultured in phenol red-free Opti-MEM. Live-cell imaging was done using an oil immersion 1.48 NA 100× objective lens in a Nikon CSU-W1 SoRa Spinning Disk confocal microscope. During image acquisition, SoRa 2.8x magnification was used with laser power set at 85%, 200 ms for the 488 channel, and 10%, 100 ms for the 561 channel, all with a step size of 0.1 μm. NIS Elements Software was used to process the images with the LUT set at 100–200 for the 488 channel.
Immunoprecipitations
In experiments where immunoprecipitations were done in lysates, cells were lysed in 1% Triton X-100, 150 mM NaCl, 10 mM Tris-HCl at pH 7.5, 1 mM EDTA, 1 mM EGTA, 0.5% NP-40, and 10% sucrose with protease inhibitor cocktail (Roche Applied Science) at 4 °C for 30 min and lysates were collected by centrifugation (18,000 × g, 20 min). Lysates (500 μl) were incubated at 4 °C for overnight with the indicated antibodies in figures coupled to Sepharose beads. Beads were pelleted by centrifugation, and pellets were washed in lysis buffer three times for 20 min with rotation at 4 °C. Immunoprecipitated proteins immunoblotted with indicated antibodies in the figures.
qRT-PCR
RNA was isolated from HEK293T cells using TRIzol reagent (Thermofisher) and further purified using Qiagen RNA isolation kit (Qiagen). cDNA was prepared from 500 ng RNA using the Maxima First Strand cDNA Synthesis Kit (Thermofisher). qRT-PCR assays were performed using EvaGreen 2× qPCR MasterMix (Bullseye) and primers corresponding to the appropriate cDNA (Supplementary Table 1). Analysis was performed using Bio-Rad CFX96.
Statistics and reproducibility
BRET ratios were defined as the acceptor emission divided by the donor emission. Normalized BRET was determined by subtracting the mean value of the baseline prior to stimulation from all data points. Net BRET was calculated by subtracting the donor-only (transfection without HaloTag or YFP) BRET ratio. To determine ligand-induced effects, ΔBRET was calculated as the difference between vehicle and WNT-treated wells, where each well was normalized to the mean value of the baseline. For analysis of G-CASE experiments, ΔBRET (%) was calculated for each well as a percent over the baseline ([(BRET ratio (treated)—mean BRET ratio (baseline))/mean BRET ratio (baseline)] × 100). Subsequently, the average ΔBRET (%) of vehicle control was subtracted. Max ΔBRET (%) was defined as the maximum change from the baseline following stimulation in the kinetic experiments. Area under curve (AUC) was determined for 6 min post-treatment. Saturation curves were fit using a one-site-specific binding model. BRETmax denotes the maximum net BRET signal. BRET50 represents the acceptor/donor ratio required to reach the half-maximal BRET signal. Dose-response curves were fit using a nonlinear three-parameter model. For all statistical tests, p < 0.05 denoted by one asterisk, was considered significant, and normality was confirmed. All data were analyzed with GraphPad Prism, and details of statistical analyses are provided in the source data file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data are provided with this paper.
References
Weis, W. I. & Kobilka, B. K. The molecular basis of G protein-coupled receptor activation. Annu. Rev. Biochem. 87, 897–919 (2018).
Liu, S., Anderson, P. J., Rajagopal, S., Lefkowitz, R. J. & Rockman, H. A. G protein-coupled receptors: a century of research and discovery. Circ. Res. 135, 174–197 (2024).
Nürnberg, B., Beer-Hammer, S., Reisinger, E. & Leiss, V. Non-canonical G protein signaling. Pharm. Ther. 255, 108589 (2024).
Harris, P. C. & Torres, V. E. Polycystic Kidney Disease, Autosomal Dominant, in GeneReviews(®). (eds M.P. Adam et al.) (University of Washington, Seattle Copyright © 1993-2024, University of Washington, Seattle. GeneReviews is a registered trademark of the University of Washington, Seattle. All rights reserved. (Seattle, 1993).
Grantham, J. J. Clinical practice. Autosomal dominant polycystic kidney disease. N. Engl. J. Med. 359, 1477–1485 (2008).
Hjelle, J. T. et al. Autosomal recessive polycystic kidney disease: characterization of human peritoneal and cystic kidney cells in vitro. Am. J. Kidney Dis. 15, 123–136 (1990).
Belibi, F. A. et al. Cyclic AMP promotes growth and secretion in human polycystic kidney epithelial cells. Kidney Int. 66, 964–973 (2004).
Yamaguchi, T., Nagao, S., Kasahara, M., Takahashi, H. & Grantham, J. J. Renal accumulation and excretion of cyclic adenosine monophosphate in a murine model of slowly progressive polycystic kidney disease. Am. J. Kidney Dis. 30, 703–709 (1997).
Zhou, X. & Torres, V. E. Emerging therapies for autosomal dominant polycystic kidney disease with a focus on cAMP signaling. Front. Mol. Biosci. 9, 981963 (2022).
Consortium, E.P.K.D The polycystic kidney disease 1 gene encodes a 14 kb transcript and lies within a duplicated region on chromosome 16. The European Polycystic Kidney Disease Consortium [published errata appear in Cell 1994 Aug 26;78(4):following 724 and 1995 Jun 30;81(7):following 1170]. Cell 77, 881–894 (1994).
Consortium, I.P.K.D Polycystic kidney disease: the complete structure of the PKD1 gene and its protein. The International Polycystic Kidney Disease Consortium. Cell 81, 289–298 (1995).
Mochizuki, T. et al. PKD2, a gene for polycystic kidney disease that encodes an integral membrane protein. Science 272, 1339–1342 (1996).
Nims, N., Vassmer, D. & Maser, R. L. Transmembrane domain analysis of polycystin-1, the product of the polycystic kidney disease-1 (PKD1) gene: evidence for 11 membrane-spanning domains. Biochemistry 42, 13035–13048 (2003).
Hughes, J. et al. The polycystic kidney disease 1 (PKD1) gene encodes a novel protein with multiple cell recognition domains. Nat. Genet. 10, 151–160 (1995).
Tsiokas, L. et al. Specific association of the gene product of PKD2 with the TRPC1 channel. Proc. Natl. Acad. Sci. USA. 96, 3934–3939 (1999).
Su, Q. et al. Structure of the human PKD1-PKD2 complex. Science 361, eaat9819 (2018).
Yamaguchi, T. et al. Cyclic AMP activates B-Raf and ERK in cyst epithelial cells from autosomal-dominant polycystic kidneys. Kidney Int. 63, 1983–1994 (2003).
Ye, H. et al. Modulation of polycystic kidney disease severity by phosphodiesterase 1 and 3 subfamilies. J. Am. Soc. Nephrol. 27, 1312–1320 (2016).
Pinto, C. S. et al. Phosphodiesterase isoform regulation of cell proliferation and fluid secretion in autosomal dominant polycystic kidney disease. J. Am. Soc. Nephrol. 27, 1124–1134 (2016).
Wang, Q. et al. Adenylyl cyclase 5 deficiency reduces renal cyclic AMP and cyst growth in an orthologous mouse model of polycystic kidney disease. Kidney Int. 93, 403–415 (2018).
Choi, Y. H. et al. Polycystin-2 and phosphodiesterase 4C are components of a ciliary A-kinase anchoring protein complex that is disrupted in cystic kidney diseases. Proc. Natl. Acad. Sci. USA. 108, 10679–10684 (2011).
Rees, S. et al. Adenylyl cyclase 6 deficiency ameliorates polycystic kidney disease. J. Am. Soc. Nephrol. 25, 232–237 (2014).
Zhou, J. X. & Torres, V. E. Autosomal dominant polycystic kidney disease therapies on the horizon. Adv. Kidney Dis. Health 30, 245–260 (2023).
Torres, V. E. et al. Tolvaptan in patients with autosomal dominant polycystic kidney disease. N. Engl. J. Med. 367, 2407–2418 (2012).
Gattone, V. H. 2nd, Wang, X., Harris, P. C. & Torres, V. E. Inhibition of renal cystic disease development and progression by a vasopressin V2 receptor antagonist. Nat. Med. 9, 1323–1326 (2003).
Yamamura, Y. et al. OPC-41061, a highly potent human vasopressin V2-receptor antagonist: pharmacological profile and aquaretic effect by single and multiple oral dosing in rats. J. Pharm. Exp. Ther. 287, 860–867 (1998).
Sussman, C. R., Wang, X., Chebib, F. T. & Torres, V. E. Modulation of polycystic kidney disease by G-protein coupled receptors and cyclic AMP signaling. Cell Signal 72, 109649 (2020).
Maser, R. L., Calvet, J. P. & Parnell, S. C. The GPCR properties of polycystin-1- A new paradigm. Front. Mol. Biosci. 9, 1035507 (2022).
Yu, S. et al. Essential role of cleavage of Polycystin-1 at G protein-coupled receptor proteolytic site for kidney tubular structure. Proc. Natl. Acad. Sci. USA. 104, 18688–18693 (2007).
Parnell, S. C. et al. The polycystic kidney disease-1 protein, polycystin-1, binds and activates heterotrimeric G-proteins in vitro [In Process Citation]. Biochem. Biophys. Res. Commun. 251, 625–631 (1998).
Zhang, B., Tran, U. & Wessely, O. Polycystin 1 loss of function is directly linked to an imbalance in G-protein signaling in the kidney. Development 145, dev158931 (2018).
Parnell, S. C. et al. A mutation affecting polycystin-1 mediated heterotrimeric G-protein signaling causes PKD. Hum. Mol. Genet. 27, 3313–3324 (2018).
Parnell, S. C. et al. Polycystin-1 activation of c-Jun N-terminal kinase and AP-1 is mediated by heterotrimeric G proteins. J. Biol. Chem. 277, 19566–19572 (2002).
Parnell, S. C., Magenheimer, B. S., Maser, R. L. & Calvet, J. P. Identification of the major site of in vitro PKA phosphorylation in the polycystin-1 C-terminal cytosolic domain. Biochem. Biophys. Res. Commun. 259, 539–543 (1999).
Kwak, M. et al. Gα(i)-mediated TRPC4 activation by polycystin-1 contributes to endothelial function via STAT1 activation. Sci. Rep. 8, 3480 (2018).
Delmas, P. et al. Gating of the polycystin ion channel signaling complex in neurons and kidney cells. FASEB J. 18, 740–742 (2004).
Delmas, P. et al. Constitutive activation of G-proteins by polycystin-1 is antagonized by polycystin-2. J. Biol. Chem. 277, 11276–11283 (2002).
Kwon, M. et al. G-protein signaling modulator 1 deficiency accelerates cystic disease in an orthologous mouse model of autosomal dominant polycystic kidney disease. Proc. Natl. Acad. Sci. USA. 109, 21462–21467 (2012).
Kim, S. et al. The polycystin complex mediates Wnt/Ca(2+) signalling. Nat. Cell Biol. 18, 752–764 (2016).
Karner, C. M. et al. Wnt9b signaling regulates planar cell polarity and kidney tubule morphogenesis. Nat. Genet. 41, 793–799 (2009).
Castelli, M. et al. Polycystin-1 binds Par3/aPKC and controls convergent extension during renal tubular morphogenesis. Nat. Commun. 4, 2658 (2013).
Schihada, H., Shekhani, R. & Schulte, G. Quantitative assessment of constitutive G protein-coupled receptor activity with BRET-based G protein biosensors. Sci. Signal 14, eabf1653 (2021).
Feng, S. et al. The sorting nexin 3 retromer pathway regulates the cell surface localization and activity of a Wnt-activated polycystin channel complex. J. Am. Soc. Nephrol. 28, 2973–2984 (2017).
George, K. et al. Robust GRK2/3/6-dependent desensitization of oxytocin receptor in neurons. iScience 27, 110047 (2024).
Kilander, M. B. et al. Disheveled regulates precoupling of heterotrimeric G proteins to Frizzled 6. Faseb J. 28, 2293–2305 (2014).
Hao, H. X. et al. ZNRF3 promotes Wnt receptor turnover in an R-spondin-sensitive manner. Nature 485, 195–200 (2012).
Bernard, M. L., Peterson, Y. K., Chung, P., Jourdan, J. & Lanier, S. M. Selective interaction of AGS3 with G-proteins and the influence of AGS3 on the activation state of G-proteins. J. Biol. Chem. 276, 1585–1593 (2001).
Okashah, N. et al. Variable G protein determinants of GPCR coupling selectivity. Proc. Natl. Acad. Sci. USA. 116, 12054–12059 (2019).
Nehme, R. et al. Mini-G proteins: novel tools for studying GPCRs in their active conformation. PLoS ONE 12, e0175642 (2017).
Janicot, R. et al. Direct interrogation of context-dependent GPCR activity with a universal biosensor platform. Cell 187, 1527–1546.e1525 (2024).
Kleene, S. J. & Kleene, N. K. The native TRPP2-dependent channel of murine renal primary cilia. Am. J. Physiol. Renal. Physiol. 312, F96–F108 (2016).
Gainullin, V. G., Hopp, K., Ward, C. J., Hommerding, C. J. & Harris, P. C. Polycystin-1 maturation requires polycystin-2 in a dose-dependent manner. J. Clin. Investig. 125, 607–620 (2015).
Zheng, W. et al. Direct binding between Pre-S1 and TRP-like domains in TRPP channels mediates gating and functional regulation by PIP2. Cell Rep. 22, 1560–1573 (2018).
Ha, K. et al. The heteromeric PC-1/PC-2 polycystin complex is activated by the PC-1 N-terminus. eLife 9, e60684 (2020).
Wang, Z. et al. The ion channel function of polycystin-1 in the polycystin-1/polycystin-2 complex. EMBO Rep. 20, e48336 (2019).
Yoshiba, S. et al. Cilia at the node of mouse embryos sense fluid flow for left-right determination via Pkd2. Science 338, 226–231 (2012).
Walker, R. V. et al. Ciliary exclusion of polycystin-2 promotes kidney cystogenesis in an autosomal dominant polycystic kidney disease model. Nat. Commun. 10, 4072 (2019).
Tsiokas, L., Kim, E., Arnould, T., Sukhatme, V. P. & Walz, G. Homo- and heterodimeric interactions between the gene products of PKD1 and PKD2. Proc. Natl. Acad. Sci. USA. 94, 6965–6970 (1997).
Shen, P. S. et al. The structure of the polycystic kidney disease channel PKD2 in lipid nanodiscs. Cell 167, 763–773.e711 (2016).
DeCaen, P. G., Liu, X., Abiria, S. & Clapham, D. E. Atypical calcium regulation of the PKD2-L1 polycystin ion channel. eLife 5, e13413 (2016).
Stoffel, R. H., Randall, R. R., Premont, R. T., Lefkowitz, R. J. & Inglese, J. Palmitoylation of G protein-coupled receptor kinase, GRK6. Lipid modification diversity in the GRK family. J. Biol. Chem. 269, 27791–27794 (1994).
Xu, Y. et al. The polycystin-1, lipoxygenase, and α-toxin domain regulates polycystin-1 trafficking. J. Am. Soc. Nephrol. 27, 1159–1173 (2016).
Yu, W. et al. Polycystin-1 protein level determines activity of the Galpha12/JNK apoptosis pathway. J. Biol. Chem. 285, 10243–10251 (2010).
Park, E. Y. et al. Cyst formation in kidney via B-Raf signaling in the PKD2 transgenic mice. J. Biol. Chem. 284, 7214–7222 (2009).
Thivierge, C. et al. Overexpression of PKD1 causes polycystic kidney disease. Mol. Cell Biol. 26, 1538–1548 (2006).
Harris, P. C. & Torres, V. E. Genetic mechanisms and signaling pathways in autosomal dominant polycystic kidney disease. J. Clin. Investig. 124, 2315–2324 (2014).
Bergmann, C. et al. Polycystic kidney disease. Nat. Rev. Dis. Prim. 4, 50 (2018).
Baameur, F. et al. Role for the regulator of G-protein signaling homology domain of G protein-coupled receptor kinases 5 and 6 in beta 2-adrenergic receptor and rhodopsin phosphorylation. Mol. Pharm. 77, 405–415 (2010).
Li, L. et al. G protein-coupled receptor kinases of the GRK4 protein subfamily phosphorylate inactive G protein-coupled receptors (GPCRs). J. Biol. Chem. 290, 10775–10790 (2015).
Bayly-Jones, C. et al. LYCHOS is a human hybrid of a plant-like PIN transporter and a GPCR. Nature 634, 1238–1244 (2024).
Xiong, Q. et al. Molecular architecture of human LYCHOS involved in lysosomal cholesterol signaling. Nat. Struct. Mol. Biol. 32, 905–913 (2025).
Zhao, J. et al. Cryo-EM reveals cholesterol binding in the lysosomal GPCR-like protein LYCHOS. Nat. Struct. Mol. Biol. 32, 896–904 (2025).
Yuasa, T., Takakura, A., Denker, B. M., Venugopal, B. & Zhou, J. Polycystin-1L2 is a novel G-protein-binding protein. Genomics 84, 126–138 (2004).
Acknowledgements
We would like to thank Drs. P. DeAngelis and X. Zhang for comments on the manuscript; Dr. Feng Qian for human HA-tagged PKD1 and PKD1-T3049V, Dr. Chris Ward for PKD1-eGFP, and Dr. Wenhan Chang for CaSR. This work was supported by grant number R01DK59599 (LT) and the John S. Gammill Endowed Chair in Polycystic Kidney Disease (LT); F31DK30605 (EPH); R35GM142786 (WLB), R01MH125998 (MA), U54DK126126 (SCP), and R01GM104251 (AAP).
Author information
Authors and Affiliations
Contributions
E.P.H.: performed BRET experiments, analyzed data, and wrote the paper with L.T. A.N.H.: performed imaging experiments. M.M.P.: constructed and characterized the PKD2-CRISPR-KO construct. V.N.: performed immunoblotting experiments on wild-type and PKD1 variants. S.E.G.: prepared PyMOL structures. WLB: assisted with molecular cloning. A.A.P., M.A., H.T.M.H., and S.C.P.: provided reagents. L.T.: supervised the project and wrote the paper with E.P.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Robin Maser, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hardy, E.P., Haider, A.N., Patel, M.M. et al. A heteromeric TRP channel that functions as a WNT-activated G protein-coupled receptor. Nat Commun 17, 3233 (2026). https://doi.org/10.1038/s41467-026-69932-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-69932-w