Abstract
Pulmonary fibrosis represents a progressive interstitial lung disease marked by excessive extracellular matrix deposition and architectural distortion. Vascular endothelial cells critically contribute to fibrogenesis through paracrine secretion of pro-fibrotic mediators, yet their mechanobiological regulation remains elusive. Using integrated single-cell multi-omics profiling of human pulmonary fibrosis specimens and experimental fibrosis models induced by bleomycin or silica, we identify mechanosensitive Piezo1 upregulation in Endothelial cells as a hallmark of fibrotic progression. Endothelial-specific Piezo1 knockout significantly attenuates Bleomycin-induced fibrotic remodeling in male mice, establishing its pathogenic necessity. Mechanistically, PIEZO1 activation promotes pulmonary fibrosis development via CAPN2-mediated STAT3 phosphorylation, which may regulate the secretion of the pro-fibrotic molecule interleukin-33. These findings suggest that the endothelial PIEZO1-CAPN2-STAT3-IL33 axis is a potential therapeutic target for PF intervention.
Similar content being viewed by others
Introduction
Pulmonary fibrosis (PF) represents a spectrum of chronic progressive interstitial lung disorders characterized by excessive extracellular matrix deposition, culminating in substantial morbidity and mortality. Idiopathic pulmonary fibrosis (IPF), the most prevalent and clinically significant subtype, presents particular therapeutic challenges due to its indeterminate pathogenesis and restricted treatment modalities. While two antifibrotic agents (pirfenidone and nintedanib) have received clinical approval, these palliative therapies only modestly decelerate disease progression without achieving fibrosis resolution1. This therapeutic limitation underscores the critical need for elucidating pathogenic mechanisms and developing targeted interventions. Current research implicates dysregulated repair mechanisms in PF pathogenesis, particularly abnormal wound healing responses characterized by progressive fibrotic extracellular matrix accumulation2. This complex process involves coordinated interactions among pulmonary structural cells (epithelial, endothelial) and recruited effector cells (immune populations, activated fibroblasts), mediated through an evolving network of soluble mediators (e.g., cytokines, chemokines) that regulate intercellular communication3.
Vascular endothelial cells (ECs), constituting the luminal interface of blood vessels, are ubiquitously distributed across tissues and critically regulate diverse pathophysiological processes4. Within the pulmonary system, these cells extend their functional repertoire beyond vascular permeability maintenance to actively modulate tissue homeostasis5. Emerging evidence indicates spatial adjacency between pulmonary fibroblasts and vascular cells, with dysregulated intercellular communication driving progressive fibrotic remodeling6. Experimental models demonstrate that bleomycin (BLM)-induced repetitive lung injury triggers pathological ligand presentation from pulmonary vascular endothelial cells to perivascular fibroblasts, initiating maladaptive repair cascades that culminate in fibrosis6. Furthermore, the NRP1-HIF2α-EPCR signaling axis has been implicated in aging-associated endothelial reprogramming, fostering aberrant macrophage-endothelial crosstalk that impairs regenerative capacity and accelerates fibrotic progression7. These findings collectively highlight pulmonary EC-derived molecular pathways as novel therapeutic targets for PF intervention.
ECs employ mechanosensors to transduce mechanical stimuli through a process termed mechanotransduction8. This biological mechanism converts mechanical forces (e.g., shear stress, cyclic stretch) into intracellular biochemical signals, thereby inducing chromatin remodeling and transcriptional reprogramming8. Biomechanical stresses orchestrate critical endothelial functions spanning angiogenesis, apoptosis regulation, inflammatory responses, and vasoactive mediator production, with dysregulation implicated in both physiological adaptation and pathological progression9,10,11. Cellular traction forces transmitted through the extracellular matrix (ECM) establish reciprocal mechanical feedback, wherein ECM stiffening perturbs mechanosensitive signaling cascades, ultimately compromising EC functional plasticity in disease states12. Pathological ECM accumulation—a hallmark of fibrotic remodeling—likely disrupts mechanical homeostasis during alveolar morphogenesis, consequently altering pulmonary vascular mechanobiology. Nevertheless, the precise molecular mechanisms linking aberrant mechanical signaling to pulmonary vascular dysfunction remain incompletely defined.
In this study, through single-nucleus RNA sequencing (snRNA-seq) profiling of IPF patient lung tissues, we identified mechanosensitive transcriptional signatures in ECs correlating with fibrotic progression. Leveraging multi-omics analysis across human IPF specimens and preclinical PF models (BLM/SiO2-induced), we demonstrated pathological upregulation of PIEZO1 mechanotransduction pathways within the pulmonary vasculature. Conditional Piezo1 knockout in ECs (Cdh5-CreERT2) significantly attenuated fibrotic remodeling in BLM-challenged mice. Pharmacological modulation using PIEZO1 antagonist GsMTx4 or agonist Yoda1 reciprocally mitigated or exacerbated fibrotic outcomes, respectively. Mechanistic investigations revealed that PIEZO1 activation induces endothelial IL33 secretion through CAPN2/STAT3-mediated transcriptional activation, establishing this mechanochemical axis as a driver of fibroblast activation. Our findings establish endothelial mechanotransduction via PIEZO1-IL33 signaling as a druggable target for pulmonary fibrosis intervention.
Results
Mechanical Stress in Pulmonary ECs Associated with Deterioration of Lung Function
To understand pulmonary fibrosis (PF) through clinical prognostic indicators, such as the relationship between lung function and cell subpopulations, we collected lung tissues from four idiopathic pulmonary fibrosis (IPF) lung transplant recipients and five non-IPF normal controls at West China Hospital of Sichuan University and Shenzhen People’s Hospital. These samples were processed using 10x Genomics for single-nucleus RNA sequencing (snRNA-seq) (Fig. 1A, Supplementary Data S1-S3), and 35,310 cells from IPF patients and 35,997 cells from normal controls were obtained for subsequent analysis after quality control. Uniform manifold approximation and projection (UMAP) was performed for dimensionality reduction. Fifteen cell subpopulations were characterized based on marker gene expression, including alveolar type I cells (AT1), alveolar type II cells (AT2), ciliated cells, club cells, basal cells, macrophages, fibroblasts, smooth muscle cells (SMC), lymphoid cells, arterial endothelial cell (arterial ECs), general capillary ECs, aerocyte ECs, systemic venous ECs, pulmonary venous ECs, lymphatic ECs (Fig. 1B, Supplementary Fig. 1A-C). We then employed Scissor, a method that identifies cell subpopulations from single-cell data associated with lung function phenotypes13.
A The schematic workflow of this study integrated single-nucleus RNA sequencing with bulk RNA-seq data. B Uniform Manifold Approximation and Projection (UMAP) clustering of cells in snRNA-seq data of IPF patients and normal lung tissues. C The “Scissor” algorithm co-analyzed snRNA and bulk-RNA data, selecting lung function-related cell subpopulations from snRNA-seq data using bulk-RNA lung function data. D Stacked bar chart showed the proportions of Scissor subpopulations in each cell type. E The UMAP plot of endothelial cells (ECs) subpopulations in snRNA-seq data. F UMAP plot displaying ECs of Scissor groups. G Volcano plots depicting pseudobulk differentially expressed genes between FLP ECs of IPF group and FLP ECs of Normal group in snRNA-seq data. P values were calculated using the Wald test. Vertical dashed lines represented log2(fold change) threshold of 1, and horizontal dashed lines represented the P-value threshold of 0.05. UMAP plot (H) and violin plot (I) showed mechanical stress (MS) scores of FLP ECs in IPF (n = 4061 cells) and Normal (n = 1826 cells) groups. J Correlation analysis (Pearson Correlation) between mechanical stress score of ECs and fibrosis score of samples shown with error band representing 95% confidence interval. K The UMAP plot showed scissor-positive and scissor-negative cells associated with low lung function (FLP) and high lung function (FHP). UMAP (L) and violin plot (M) showed MS scores of FLP ECs in IPF (n = 5020 cells) and Normal (n = 1383 cells) groups of five GEO datasets. N Meta-analysis of five GEO datasets showing the MS score of ECs in IPF and Normal groups. Standardized mean difference (SMD) were calculated as Hedges’ g with corresponding 95% confidence intervals. O Correlation analysis (Pearson Correlation) of mechanical stress scores and fibroblast subpopulations in GSE136831 dataset. I, M: Violin plots displayed the mean (yellow diamond) the median (center line), the interquartile range, and the whiskers (1.5 × interquartile range). P value was calculated using the two-tailed unpaired t-test. Source data are provided as a Source Data file.
The clinical trajectory of idiopathic pulmonary fibrosis is variable, with a decline in forceful lung volume (FVC) being the main indicator of progression14. Given that an FVC ≤ 50% is indicative of severe pulmonary dysfunction and significant physiological impairment15,16, we established 50% of FVC as the diagnostic cut-off value. Subjects were stratified into two phenotypes: those with FVC ≤ 50% were defined as the FVC-low phenotype (FLP, designated Scissor+), while those with FVC > 50% comprised the FVC-high phenotype (FHP, designated Scissor–).We then integrated the bulk data (GSE47460) with the snRNA-seq data using Scissor, identifying 18,272 Scissor+ (FVC% ≤ 50%) and 29,897 Scissor- cells associated with FLP and FHP, respectively (Fig. 1C). After calculating the proportion of FLP cells of each cell type, we found that the proportion of FLP ECs within the total population of ECs was the most predominant compared with other types of cells (Fig. 1D). To delineate aberrant signaling pathways in ECs associated with severe lung function impairment during pulmonary fibrosis progression, we performed Scissor-based analysis on ECs from IPF patients and normal controls. Endothelial cells were further partitioned into six subtypes (Fig. 1E). FLP cells from IPF patients and controls were then selected using the Scissor method and subjected to pseudobulk and differential expression analysis (Fig. 1F-G). Gene set scoring revealed significant upregulation of the mechanical stress (MS) signaling pathway in the IPF group (Fig. 1H-I).
To further validate our findings, we expanded our sample size by collecting scRNA-seq data from 66 IPF patients and 59 normal lung tissues from five public scRNA-seq datasets in the GEO database17,18,19,20,21. Pearson correlation analysis on collected samples demonstrated a positive correlation between mechanical stress score of ECs and fibrosis score of samples (Fig. 1J). We then identified 34,898 ECs (Supplementary Fig. 2A-B) and used the scissor method to identify FLP and FHP cells (Fig. 1K). We also focused on whether mechanical stress was responsible for the progression of IPF and found widespread increases in mechanical stress in the lung cells of IPF patients (Fig. 1L-M and Supplementary Fig. 2C). Additionally, a meta-analysis combining the five GEO datasets validated that ECs from IPF patients exhibited relatively high mechanical stress (Fig. 1N). We performed Pearson correlation analysis on a randomly selected dataset (GSE136831). This analysis revealed a positive correlation between mechanical stress scores and both the myofibroblast percentage in total fibroblasts and fibroblast markers (Fig. 1O and Supplementary Fig. 2D). These results suggest that abnormally elevated mechanical stress in ECs is associated with the development of PF.
Mechanical Stress of ECs Elevated in SiO2 Induced Silicosis
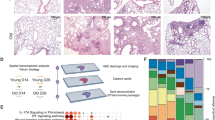
Silicosis is a type of occupational PF caused by the inhalation of large amounts of SiO₂ dust. To determine whether ECs in PF exhibit elevated mechanical stress, we established a mouse model of silicosis by tracheal instillation of SiO₂ over five months. We collected single-cell suspensions from lung tissues to prepare scRNA-seq and scATAC-seq libraries using 10X Genomics kits to measure RNA and chromatin accessibility (Fig. 2A). The H&E and Masson staining results demonstrated significant interstitial lung fibrosis in the PF mouse model induced by SiO₂ (Fig. 2B).
A Schematic diagram of the SiO2 mouse model. Mice were divided into control (saline, n = 3) and experimental (SiO2, n = 3) groups by tracheal injection. After 20 weeks, mice were euthanized, and lung single-cell suspensions were prepared for sequencing. B Validation of the SiO2 mouse model. H&E and Masson staining showed increased inflammation and fibrosis in the lungs of SiO2 treated mice (n = 3 mice per group). C UMAP plots of lung tissue scRNA-seq data from SiO2-induced and control mice. D UMAP plots of lung tissue scATAC-seq data from SiO2-induced and control mice. E Violin plots showed expression of marker genes of scRNA-seq data from SiO2-induced and control mice. F In scATAC-seq data from SiO2-induced and control mice, peaks for marker genes were identified for each subcelltype. G The UMAP plot showed endothelial subpopulations in scRNA-seq data from SiO2-induced and control mice. UMAP plot (H) and violin plot (I) showed the MS scores of endothelial cells in scRNA-seq data of Silica-induced (n = 784 cells) and control (n = 816 cells) mice. J The UMAP plot showed the endothelial subpopulations of scATAC-seq data. UMAP plot (K) and violin plot (L) showed the MS scores of endothelial cells in scRNA-seq data of Silica-induced (n = 387 cells) and control (n = 689 cells) mice. M GO enrichment analysis of motifs showed that signaling pathways associated with mechanical stress were enriched in ECs. Statistical significance of GO terms was calculated using the hypergeometric test. N Table showing TF motifs of endothelial cells enriched in mechanical stress-related pathways by scATAC-seq data. P values were calculated using the hypergeometric test. I, L: Violin plot displayed the mean (yellow diamond) the median (center line), the interquartile range, and the whiskers (1.5 × interquartile range). P value was calculated using the two-tailed unpaired t-test. Source data are provided as a Source Data file.
After quality assessment and screening of SiO₂-treated and saline-treated mice (Supplementary Data S4-S6), scRNA-seq clustered 39,048 cells and identified major cell types in the lung tissues based on marker genes expression. As a result, UMAP plots showed lung cell subpopulations, including endothelial cells, dendritic cells, B cells, AT1 cells, AT2 cells, macrophages, neutrophils, T cells, pericytes, clara cells, natural killer (NK) cells, monocytes, ciliated cells, basal cells, mast cells, smooth muscle cells (SMC), and fibroblasts (Fig. 2C, Supplementary Fig. 3A). We also analyzed the scATAC-seq dataset including 26,575 cells for quality control and batch correction using the R package “Seurat” and used our annotated scRNA-seq dataset to predict cell types via scATAC-seq (Fig. 2D, Supplementary Fig. 3B). Additionally, we identified markers for cellular subtypes in mouse lung tissues post-SiO₂ treatment using scATAC-seq and scRNA-seq (Fig. 2E-F).
To investigate differentially regulated signaling pathways in ECs, we performed GO analysis and MS scoring on ECs of scRNA-seq dataset. We found that ECs from SiO₂-treated mice showed elevated mechanotransduction signals (Fig. 2G-I and Supplementary Fig 3C-D). We also analyzed the different peaks of ECs and performed MS scoring on ECs in the scATAC-seq dataset of SiO₂-treated and saline-treated mice, revealing an increase in mechanical stress scores in the SiO₂-treated group (Fig. 2J-L Supplementary Fig. 3E).
Finally, we analyzed the enrichment of the mechanical stress-related genes in ECs for motifs (Fig. 2M) and observed that the central motifs included Sox9, Stat1, Ets1, Fosl1, Fos, Etv1, Rela, Neurog1, Meis2, Pparg, Jun, Gata4, Foxp2, Nfkb1, and Irf1 (Fig. 2N). These results confirm the critical role of aberrant mechanosensitive signaling pathways in ECs in the pathogenesis of occupational PF.
Mechanosensitive PIEZO1 is Relevant to the Development of PF
BLM-induced pulmonary fibrosis (PF) mice are widely used as an animal model for studying PF due to their ability to mimic human lung fibrosis. To further validate the above findings, we harvested lung tissues from BLM-treated mice for single-cell multi-omics studies (Fig. 3A). H&E and Masson staining revealed marked interstitial fibrosis in the lungs (Fig. 3B), confirming the successful establishment of the PF model. Using the same bioinformatics analysis approaches, scRNA-seq and scATAC-seq clustered 41,522 and 35,791 cells, respectively (Supplementary Data S7-9). Next, we visualized the major lung cell subpopulations using UMAP analysis (Fig. 3C-D) and identified clustering-specific markers for each subpopulation (Supplementary Fig. 4A-F).
A BLM intratracheal model scheme (n = 3 per group). B H&E and Masson validation of inflammation/fibrosis (n = 3 per group). C Lung scRNA-seq UMAP. D scATAC-seq UMAP subclusters. E Venn showing Piezo1 and Piezo2 as co-regulated genes across silica and BLM bulk, scRNA-seq and scATAC-seq data. F scATAC-seq coverage peaks of Piezo1 and Piezo2. G CD31 and CD45-sorted EC bulk-RNA-seq heatmap and bar chart (n = 3 per group). Gene expression levels were Z-score normalized. P values were calculated using the two-tailed unpaired t-test. H PIEZO1 (green) and Ve-cad (red) co-staining and quantification in fibrotic mouse lungs (n = 4). UMAP and bar charts (mean ± SEM) of PIEZO1+ (I) and PIEZO2+ (J) EC counts in normal controls (n = 5) versus IPF (n = 4) patients. P values were calculated using the two-tailed unpaired t-test. K PIEZO1 (green) and CD31 (red) co-staining and quantification in human IPF vs normal lungs (n = 10). L Meta-analysis of five GEO datasets showing elevated PIEZO1 in IPF ECs. Standardized mean difference (SMD) were calculated as Hedges’ g with corresponding 95% confidence intervals. Data were shown as mean ± SEM, H and K: two-tailed unpaired t-tests, Source data are provided as a Source Data file.
In order to identify by which specific gene of mechanosensitive pathway of EC might attribute to PF development, we then used Venn diagram analysis to integrate bulk RNA-seq of ECs from BLM-induced PF of mice with a single multi-omics dataset from above-analyzed human and mice samples, and we identify mechanosensitive Piezo1 and Piezo2 might serve as the candidate genes to involve in the pathogenesis of PF (Fig. 3E). PIEZOs are calcium-permeable nonselective cation channels that sense mechanical stimuli in multicellular organisms and mediate various biological processes. Subsequently, we analyzed the peaks of all mechanical stress-related genes in ECs and found that the increased peak activities of ECs mainly included Piezo1 and Piezo2 (Fig. 3F and Supplementary Fig. 4G). Additionally, motif enrichment analysis identifies motifs associated with mechanical stress in ECs, including Neurog1, Stat1, Nfkb1, Gata4, Rela, Sox9, Foxp2, Etv1, Fos, Fosl1, Meis2, Irf1, Jun, Ets1, and Pparg (Supplementary Fig. 4H). Interestingly, heatmap analysis of bulk-RNA-seq showed the expression of piezo1 increased in ECs from BLM-treated mice, while Piezo2 exhibited an opposite expression pattern (Fig. 3G). Consistent with these findings, immunofluorescence analysis of murine models demonstrated that Piezo1 expression was significantly upregulated in pulmonary ECs of BLM-treated mice (Fig. 3H). To further verify our finding in human samples, we firstly investigated the subpopulation distribution of PIEZOs in ECs, we quantified the proportions of PIEZO1+ and PIEZO2+ ECs, respectively. The analysis revealed that PIEZO1 is predominantly expressed in vascular ECs, whereas PIEZO2 shows preferential expression in lymphatic ECs (Supplementary Fig. 4I-J). Quantitative analysis of PIEZO1+ and PIEZO2+ EC proportions compared with control revealed a significant increase of PIEZO1+ cells in the IPF group, whereas PIEZO2+ cells showed no significant change (Fig. 3I-J). Subsequent immunofluorescence validation using IPF patient and normal control confirmed that PIEZO1 expression was significantly upregulated in IPF vascular ECs compared to control (Fig. 3K). A meta-analysis of the GEO Human IPF Public Database also showed that PIEZO1 was significantly upregulated in ECs (Fig. 3L). Therefore, we selected PIEZO1 in ECs as the promising intervention target of PF for the functional validation in following study.
Gene Deletion and Pharmacological Intervention of Piezo1 Affect PF Development
To determine the functional role of PIEZO1 in pulmonary vascular cell-mediated fibrosis, we systematically knocked out Piezo1 in vascular cells of adult mice using a tamoxifen-inducible EC-specific CreERT2 system. We developed mice with loxP site-flanking Piezo1 crossed with VE-Cad-CreERT2 mice. Tamoxifen was used to specifically activate CreERT2 activity in ECs, inducing EC-specific deletion of Piezo1 (Piezo1 ΔEC) in VE-cad-CreERT2 Piezo1 loxP/loxP mice. Piezo1-haploinsufficient adult mice (Piezo1 WT) served as controls to monitor Cre toxicity.
We observed significant knockdown of PIEZO1 in lung vascular endothelial cells (Fig. 4A-B), confirming the successful generation of EC-specific Piezo1-deficient mice. Subsequently, Masson, picrosirius red (PSR) and H&E staining revealed significant reduction of collagen deposition and inflammation in Piezo1ΔEC mice compared to controls (Fig. 4C, Supplementary Fig. 5A-B). Alpha-smooth muscle actin (αSMA) immunofluorescence staining further verified the inhibition of lung fibrosis by Piezo1 deletion in ECs (Fig. 4D). We induced pulmonary fibrosis in these Piezo1ΔEC mice using BLM and observed a significant reduction in hydroxyproline in lung tissue (Fig. 4E).
A Tracheal BLM model schematic (Piezo1 WT and Piezo1ΔEC mice). B PIEZO1 protein in isolated EC, immune cells, fibroblast and epithelial cells (n = 3). C Representative Masson and PSR images and quantification (n = 6). D αSMA immunofluorescence and quantification (n = 4). E Lung hydroxyproline content in Piezo1 WT and Piezo1ΔEC mice (n = 5). F Scheme of PIEZO1 agonist Yoda1 or antagonist GsMTx4 treatment; mice sacrificed after 3 weeks. G Lung hydroxyproline after i.p. Yoda1 or GsMTx4 (n = 6). H Representative Masson and PSR images and quantification (n = 6). I αSMA immunofluorescence and quantification (n = 5). Error bars represent mean ± SEM, B: two-tailed unpaired t-tests, G-I: one-way ANOVA followed Tukey’s multiple comparisons test was, C-F: two-way ANOVA followed Tukey’s multiple comparisons test. All n denote biologically independent samples. Source data are provided as a Source Data file.
To further investigate the role of PIEZO1 in modulating pulmonary EC function and lung injury, we administered the PIEZO1 agonist Yoda1 via both intratracheal and intraperitoneal routes. We found that, compared to the saline control group, both intratracheal and intraperitoneal administration of Yoda1 led to an increased inflammatory cells and fibrosis in the lungs of mice revealed by the hydroxyproline assay, Masson, picrosirius red (PSR) and H&E (Supplementary Fig. 5C-F).
Additionally, we applied antagonist and agonist to explore if PIEZO1 is a potential therapeutic target for PF (Fig. 4F). Histopathological staining and hydroxyproline assay revealed that the antagonist GsMTx4 reduced the production of hydroxyproline in fibrotic lung tissue as well as deposition of ECM and collagen, indicating GsMTx4 can effectively alleviate PF, whereas the agonist Yoda1 exacerbated the fibrotic response in the BLM-induced PF mouse model (Fig. 4G-I, Supplementary Fig. 5G-H). Importantly, Yoda1 failed to worsen fibrosis in endothelial-specific Piezo1-knockout mice subjected to BLM, confirming that the pro-fibrotic effect of Yoda1 is mediated through endothelial Piezo1 (Supplementary Fig. 6A-D).
In conclusion, our findings demonstrate that targeting PIEZO1 in ECs inhibits the development of pulmonary fibrosis, highlighting PIEZO1 as a promising target for the development of therapeutic interventions for PF.
PIEZO1 Activation Coupling IL-33 to Regulate the Development of PF
Recent studies demonstrate that ECs play a crucial role in the pathogenesis of pulmonary fibrosis through both juxtacrine and paracrine mechanisms, mediated by the secretion of various bioactive molecules, particularly cytokines and chemokines4,6,12,22. Notably, emerging evidence suggests that aberrant mechanical stress can significantly influence the release of these mediators from ECs during pathological processes. Building upon these findings, we propose a novel hypothesis that aberrant mechanical stress contributes to the progression of PF by modulating the secretion of pro-fibrotic factors from ECs.
Firstly, we conducted CellChat analysis on our snRNA-seq dataset, which revealed that PIEZO1+ ECs had significantly higher interaction numbers and strengths with other lung cells compared to PIEZO1- ECs (Fig. 5A). To further explore potential downstream pro-fibrotic factors sensitive to mechanical stress and regulated by PIEZO1 signaling, we analyzed publicly available scRNA-seq data and identified 47 differentially regulated pro-fibrotic genes through Venn diagram analysis (Fig. 5B). Heat map analysis showed that IGFBP5, IL33, MGP, SPRY1, and ACKR1 were specifically expressed in ECs (Fig. 5C), and these genes exhibited a significant correlation with mechanical stress scores (Fig. 5D). However, PIEZO1 expression were only positively correlated with the expression of IL33 and ACKR1 (Fig. 5E). Given that ACKR1 is not classified as a cytokine, whereas IL-33 represents a canonical cytokine, we therefore selected IL33 as the definitive therapeutic target in this study. Feature and violin plot indicated that IL33 was specifically and highly expressed in ECs (Fig. 5F), and IL33 was significantly upregulated in the IPF group compared to normal control (Fig. 5G). The violin plot of our snRNA-seq dataset revealed that IL33 expression was significantly higher in PIEZO1+ cells compared to PIEZO1- cells, with PIEZO1+ ECs in the IPF group exhibiting a higher IL33 expression level than those in the control group (Fig. 5H). We further analyzed bulk RNA-seq data from sorted ECs, observing upregulation of Il33 expression in ECs from BLM-treated mice compared to control, the sequencing results were also validated (Fig. 5I-J), which were further verified by immunofluorescence staining of IL-33 in the lung ECs of fibrosis patients (Fig. 5K).We then examined the relationship between IL33 and PIEZO1. To our expectation, immunofluorescence staining showed that administration of Yoda1 enhanced IL-33 expression, while GsMTx4 had the opposite effect (Fig. 5L). Immunofluorescence analysis further confirmed that specific deletion of Piezo1 in ECs markedly reduced IL-33 expression, reinforcing the hypothesis that PIEZO1 activation regulates IL33 secretion in ECs (Fig. 5M).
A CellChat analysis of human lung snRNA-seq data. B Venn of 47 shared DEGs from four datasets (GSE159354, GSE132771, GSE128033, GSE122960). C Heatmap of 47-gene expression across cell types. D Correlation analysis (Pearson Correlation) of expression of IGFBP5, IL33, MGP, SPRY1, ACKR1 and mechanical stress scores in ECs. E Correlation analysis (Pearson Correlation) of expression of PIEZO1 and IGFBP5, IL33, MGP, SPRY1, ACKR1 in ECs. F Human snRNA-seq UMAP and violin of IL33 distribution and subpopulations. G Bar chart showed expression of IL33 (mean ± SEM) in IPF (n = 7966 cells) vs control (n = 4174 cells). P value was calculated using the two-tailed unpaired t-test. H Violin of IL33 in PIEZO1- vs PIEZO1+ ECs. I Bulk-RNA-seq of saline and BLM (7 days and 14 days) induced mice (n = 3 per group). Gene expression levels were Z-score normalized. P value was calculated using the two-tailed unpaired t-test. J qPCR of Il33 in BLM mouse lung ECs. K IL-33 (green) and CD31 (red) co-staining in human IPF vs normal, quantification is shown at right (n = 10). L IL-33 (green) and CD31 (red) co-staining in Yoda1- or GsMTx4-treated fibrotic mouse lungs, quantification is shown at right (n = 3). M IL-33 (green) and CD31 (red)co-staining in Piezo1 WT and Piezo1ΔEC BLM lungs, quantification is shown at right (n = 3). Error bars represent mean ± SEM, H, K, J, M: two-tailed unpaired t-test; L: one-way ANOVA followed Tukey’s multiple comparwasons test, All n values indicate biologically independent samples. Source data are provided as a Source Data file.
Although IL33 is a well-documented profibrogenic cytokine known to drive fibrosis in multiple organs, the specific role of EC-derived IL33 in PF development remains largely unclear. To determine whether IL33 in ECs is essential for PF development, we specifically deleted Il33 in ECs by crossing VE-cad-CreERT2 mice with Il33loxP/loxP mice and applied BLM to create a pulmonary fibrosis mouse model (Fig. 6A). mRNA of Il33 and protein of IL-33 were efficiently deleted in ECs via Vecad-Cre/LoxP (Fig. 6B). Subsequent analyses with Masson, PSR, H&E and αSMA staining, hydroxyproline assay demonstrated that Il33 ΔEC mice had remarkably lower fibrotic responses than those of Il33 WT mice in bleomycin induced PF mouse model (Fig. 6C-E, Supplementary Fig. 6E).
A Schematic of BLM-induced fibrosis in Il33 WT vs Il33 ΔEC mice. B Il33 mRNA (n = 4) and IL-33 protein in lung ECs (n = 3). C Representative Masson/PSR images and quantification (n = 5). D αSMA immunofluorescence and quantification (n = 4). E Lung hydroxyproline content (n = 5). F Schematic: EC-specific Il33 overexpression via AAV-Tie1 in Piezo1 WT and Piezo1ΔEC mice; 4-week pretreatment, BLM at week 4, sacrifice at week 7. G IL-33 expression in ECs, immune and epithelial cells after AAV-OE-Il33 (n = 3). H Masson/PSR images and quantification (n = 5). I αSMA immunofluorescence and quantification (n = 4). J Hydroxyproline content (n = 5). Error bars: mean ± SEM; B,G: two-tailed unpaired t-test; C,D-E,I: two-way ANOVA followed Tukey’s multiple comparisons test; H,J: three-way ANOVA followed Tukey’s multiple comparisons test. All n values indicate biologically independent samples. Source data are provided as a Source Data file.
Moreover, we constructed a mouse model with EC-specific Il33 overexpression (OE-Il33) using an adeno-associated virus (AAV) delivery system to determine if Piezo1-regulated PF development is dependent on Il33. We generated a lung EC-selective adeno-associated virus (AAV) system expressing Il33 under the EC-selective Tie1 promoter (AAV-Tie1-Il33), which was specifically overexpressed in the lung of Piezo1 ΔEC mice. Mice were initially pretreated with AAV through the trachea for four weeks, followed by bleomycin treatment. They were then euthanized in the third week after bleomycin administration (Fig. 6F). mRNA of Il33 and protein of IL-33 were confirmed the endothelial-specific overexpression of Il33 in lung tissue. (Fig. 6G). As expected, specific deletion of Piezo1 in ECs alleviated pulmonary fibrosis, but this effect was significantly reversed by the specific overexpression of Il33 in ECs achieved by AAV-Il33 (Fig. 6H-J, Fig Supplementary Fig. 6F). Collectively, these in vivo functional rescue experiments indicate that the endothelial PIEZO1 activation promoted PF development in a IL33 dependent manner.
CAPN2-STAT3 Axis Regulates IL-33 Expression Upon the PIEZO1 Activation
We investigated the regulatory mechanism of PIEZO1 on IL33 expression in vitro using primary human umbilical vein endothelial cells (HUVECs). Since both cellular tension and changes in matrix stiffness can activate the PIEZO1 channel, we applied tension to the monolayer of HUVECs using a cell stretching device and cultured HUVECs on substrates of varying stiffness to simulate the spontaneous activation of PIEZO1. We found that when subjected to 20% strain, the secretion and mRNA transcription levels of IL33 in HUVECs initially increased at 6 hours and then decreased at 24 hours, and meanwhile we observed that in monolayers cultured on a 25 kPa substrate, HUVECs exhibited higher secretion and transcriptional expression levels of IL-33 (Fig. 7A-B). Notably, silencing PIEZO1 expression with shRNA in HUVECs inhibited the secretion and transcription of IL-33 after cells were subjected to 20% strain for 6 hours or cultured on a 25 kPa substrate (Fig. 7C-E).
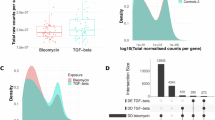
HUVECs under 20% stretch for 0,6,24 h or 2,12,25 kPa substrates; IL-33 secretion (n = 3) A and mRNA (n = 3) B. C PIEZO1 knock-down with shPIEZO1 (n = 3). HUVECs were transduced with shPIEZO1 for 48 h, cultured on 25 kPa substrates for 24 h or subjected to 6 h mechanical stretch before measuring IL-33 secretion (n = 3) D and mRNA (n = 3) E. HUVECs were cultured on 2, 12, 25 kPa substrates for 24 h or stretched (20%) for 0, 6, 24 h, followed by measurement of calpain activity F and CAPN2 protein G (n = 3). HUVECs (shPIEZO1) were cultured on 25 kPa substrates for 24 h or subjected to 6 h mechanical stretch, followed by measurement of calpain activity (n = 3) H and CAPN2 protein levels (n = 3) I. J CAPN2 knock-down with shCAPN2 (n = 3). K HUVECs (shCAPN2) were cultured on 25 kPa substrates for 24 h or subjected to 6 h mechanical stretch, IL-33 secretion(left) and mRNA(right) (n = 3). L Model for paracrine IL-33 regulation. M Cistrome: STAT3 predicted to drive IL-33 regulation. N intersection of top 100 motifs in BLM-treated mouse ECs with top 10 TFs. O The protein levels of P-STAT3 and STAT3 in HUVECs treated with 20% stretch for 0 and 6 hours (n = 3). P The protein levels of P-STAT3 and STAT3 in HUVECs (shPIEZO1 or shCAPN2) treated with 20% stretch for 6 h (n = 3). Q STAT3 knock-down with shSTAT3 (n = 3). R HUVECs (shSTAT3) were cultured on 25 kPa substrates for 24 h or subjected to 6 h mechanical stretch; IL-33 secretion(left) and mRNA(right) (n = 3). Error bars: mean ± SEM; C, G, J, O–Q: two-tailed unpaired t-test; A, B, F: one-way ANOVA followed Šídák’s multiple comparisons test, D–E, H–I, K, R: two-way ANOVA Tukey’s multiple comparisons test. All n denote biologically independent samples. Source data are provided as a Source Data file.
CAPN2 functions as a key downstream effector in the PIEZO1-mediated mechanotransduction pathway, participating in multiple mechanoresponsive biological processes. Interestingly, both the activity and protein level of Calpain2 significantly increased after 20% strain stretching and 25 kPa substrate culturing with a similar trend to IL-33 (Fig. 7F, G), which were inhibited by PIEZO1 knockdown (Fig. 7H, I). Thus, we speculated PIEZO1 activation induced IL33 is dependent on CAPN2. Consistent with our hypothesis, CAPN2 knockdown (shCAPN2) markedly decreased IL-33 expression and secretion upon mechanical stimulation (Fig. 7J, K).
Transcription factors (TFs) are necessary to bind to the promoter region and directly initiate the expression of target genes. To explore which TFs act as downstream molecules of the PIEZO1-CAPN2 axis to regulate IL-33 expression (Fig. 7L), we first utilized the Cistrome (http://dbtoolkit.cistrome.org/) public transcription factor database and predicted the top 10 transcription factors that might regulate IL33 expression (Fig. 7M). Additionally, we performed motif enrichment analysis of differential peaks in ECs using scATAC-seq datasets of the fibrotic mouse model and identified the top 100 motifs highly expressed in the endothelium (Fig. 7N). Venn diagram analysis suggested that STAT3 is a key TF regulating IL33 expression upon mechanical force (Fig. 7N). Importantly, 6 hours of mechanical stretch clearly promoted the activation of STAT3 as revealed by p-STAT3 immunoblotting (Fig. 7O), and this stretch-induced activation was blocked by PIEZO1 and CAPN2 silencing (Fig. 7P). These results suggest that STAT3 might acts as a downstream TF of PIEZO1 that is involved in the regulation of IL33 expression. As demonstrated in Fig. 7Q, R, STAT3 knockdown (shSTAT3) in HUVECs cultured under mechanostimulatory conditions (cyclic 20% tensile strain and 25 kPa substrate stiffness) significantly attenuated both IL33 transcript levels and protein secretion.
In conclusion, endothelial PIEZO1 activation induces the downstream profibrotic molecule IL-33 to regulate the development of PF, might potentially through the CAPN2-STAT3 axis.
Discussion
Pulmonary fibrosis (PF) pathogenesis arises from dynamic interplay between cellular compartments and dysregulated signaling networks. Employing single-cell RNA sequencing (scRNA-seq) of clinical specimens complemented by public dataset integration, we delineated mechanobiological drivers of PF progression. Leveraging the computational algorithm Scissor, we established bi-directional associations between endothelial mechanosensitivity and declining pulmonary function (forced vital capacity [FVC]). Through cross-species validation in BLM/SiO2-induced murine models with scRNA-seq confirmation, we established endothelial PIEZO1 as a central mechanotransducer in fibrotic remodeling. Mechanistic investigations revealed that PIEZO1-mediated calcium flux activates CAPN2-dependent STAT3 phosphorylation, may triggering IL-33 secretion that sustains fibroblast activation through paracrine signaling. These results position the endothelial PIEZO1-CAPN2-STAT3-IL33 axis as a mechanochemical signaling hub with therapeutic potential for PF management.
The lung maintains lifelong mechanical responsiveness, continuously experiencing biomechanical forces during respiratory cycles that shape its structural and functional adaptation23,24. Pathological fibrotic remodeling manifests as alveolar architectural distortion and compromised pulmonary compliance, driven by myofibroblast dysregulation and aberrant ECM homeostasis25.Emerging evidence implicates vascular biomechanical alterations in pulmonary fibrosis pathogenesis26,27, VECs emerge as central regulators of fibrotic progression through integrated metabolic reprogramming, transcriptional control, and epigenetic modulation of paracrine signaling7,28,29. To date, single-cell studies have revealed a high degree of heterogeneity in ECs across different tissues, along with dynamic and plastic functions of immune cells. Most single-cell studies focus primarily on descriptive mapping, with limitations in linking data to clinical phenotypes. Therefore, we used the “Scissor” algorithm to correlate FVC with single-cell data. Additionally, “Scissor” classified ECs into two groups, revealing that elevated mechanical stress in ECs were strongly associated with worsening lung function. Our bioinformatics approach of correlating single-cell data with clinical phenotypes has strong clinical relevance and can be widely applied to other diseases.
Bleomycin and SiO₂ may induce abnormal mechanical forces in murine lungs. Initially, these agents directly damage alveolar type II epithelial cells (AT2 cells), leading to surfactant dysfunction and aberrant mechanical force generation30. Subsequent epithelial cell death and alveolar collapse further disrupt local mechanical homeostasis, exacerbating parenchymal stress31. Concurrently, inflammatory cascades—marked by immune cell infiltration and alveolar edema—amplify epithelial and endothelial cell injury and dysfunction, thereby intensifying mechanical dysregulation32. During the chronic phase, excessive ECM deposition increases tissue stiffness, initiating a self-reinforcing cycle of mechanical stress33. This cycle activates mechanosensitive pathways in resident cells, including ECs and fibroblasts, which further accelerates ECM remodeling and fibrotic progression. Ultimately, chronic fibrosis results in declined lung function, disrupted respiratory rhythm, and perpetuation of abnormal mechanical forces.
Clinical evidence establishes hemodynamic shear stress perturbations as pathogenic drivers in vascular disorders, notably atherosclerosis. VECs exhibit flow-adaptive spindle-shaped morphology aligned with laminar flow vectors, with pathological progression involving chronic exposure to aberrant shear profiles34. In PF, we posit that ECs undergo pathological shear stress reprogramming. Mechanistically, ECs decode hemodynamic forces through apical mechanosensory complexes comprising PIEZO1, protein kinase cascades, P2X4 receptors, and NOTCH1 signaling nodes35,36. Seminal work by Li et al. demonstrated embryonic lethality in mice with global or endothelial-specific Piezo1 deletion, underscoring its essentiality in vascular morphogenesis12. Although PIEZO1 is recognized as a master mechanoregulator in cardiovascular pathologies (e.g., atherosclerosis, hypertension)37, its pulmonary vascular functions remain unexplored. Through endothelial-specific Piezo1 ablation in conditional knockout (VE-Cad-CreERT2) mice, we established its functional necessity in PF pathogenesis. Single-cell transcriptomic profiling revealed compartment-specific expression patterns: PIEZO1 localized to pulmonary vascular endothelium, whereas PIEZO2 dominated lymphatic endothelial, suggesting distinct mechanobiological roles in respiratory homeostasis.
IL-33, a tissue-derived alarmin of the IL-1 superfamily, is constitutively expressed in nuclear compartments of pulmonary ECs, immune populations, and stromal cells, serving as a key regulator of type 2 immunity and barrier homeostasis38. While its profibrotic role through macrophage-mediated ST2 signaling is established, the endothelial-specific contribution to fibrogenesis remains enigmatic39. scATAC-seq is a useful tool for theoretically investigating potential transcription factors involved in specific cellular processes. Using this tool, we identified STAT3 as a potential upstream transcription factor promoting IL33 expression upon PIEZO1 activation. Thus, STAT3 and IL33 may form a positive feedback loop in the presence of abnormal PIEZO1 signaling. Finally, since ECs are present in almost all human tissues and organs, we suggest that a similar mechanism of PIEZO1-IL33 crosstalk may be involved in other tissue remodeling diseases, including chronic obstructive pulmonary disease (COPD), asthma, and tumors.
Although Yoda1 is widely employed as a PIEZO1 agonist to simulate mechanotransduction pathways, Yoda1 as a surrogate for mechanical stress still presents critical limitations. Firstly, Yoda1 exhibits off-target effects, inducing tonic sustained elevation of intracellular Ca²⁺ that may activate non-physiological pathways, such as aberrant PIEZO-mediated signaling observed in cardiac systems40,41. Secondly, the spatio-temporal Ca²⁺ signaling patterns triggered by Yoda1 fundamentally differ from those induced by physiological mechanical cues (e.g., shear stress, stretch, or stiffness)42,43,44. Unlike the localized (e.g., focal adhesion-associated) and transient activation of PIEZO1 under mechanical stimulation45,46, Yoda1 elicits prolonged, global Ca²⁺ signaling that deviates from in vivo mechanotransduction dynamics. Therefore, to address these limitations, we employed physiologically relevant mechanical stimuli-cyclic stretch and substrate stiffness modulation-to mimic mechanical forces in our experimental models. Our in vitro data demonstrate that mechanical force-induced Ca²⁺ transients (via stretch/stiffness) activate Calpain, consistent with its role in mechanosensitive pathways. Notably, under 24-hour sustained mechanical stimulation, Calpain levels decreased rather than increased, aligning with the hypothesis that prolonged mechanical cues may trigger homeostatic downregulation of Calpain activity to counteract excessive proteolytic stress. This observation underscores the necessity of distinguishing pharmacological agents (e.g., Yoda1) from biomechanical stimuli when interrogating mechanotransduction pathways.
There are several limitations to this study due to technical hurdles. First, although we used bioinformatics tools, we lacked an experimental method to directly measure true mechanical stress levels in ECs. Therefore, we could not demonstrate whether ECs are affected by abnormal shear stress in fibrotic lungs. Second, while our data position STAT3 as an upstream regulator of IL33, exactly how this regulation occurs—whether directly or indirectly—remains to be elucidated.Third, while we used EC conditional knockout mice to show that PIEZO1 activation-mediated IL33 induction is necessary for PF development, we lacked studies on cellular crosstalk between ECs and other cells. IL-33, an EC-derived necrosis factor, may activate perivascular fibroblasts and macrophages during PF development, warranting co-culture experiments to explore this. Although more research is needed, our work elucidates the molecular and cellular nodes of abnormal mechanical stress in ECs, potentially aiding in the discovery of new candidate therapies for treating fibrotic diseases.
Methods
Human Patient Samples
In this study, we enrolled four patients (male: 3; female: 1; age range: 45–64 years, mean ± standard deviation: 57.7 ± 4.3 years) with idiopathic pulmonary fibrosis (IPF) who underwent lung transplantation surgery and five normal individuals (male: 3; female: 2; age range: 38–69 years, mean ± standard deviation: 51.0 ± 6.1 years) whose lung tissue specimens were obtained from surgical resections for other indications. Designed to explore the central role of endothelial cells in pulmonary fibrosis, this study had a limited sample size and was not prospectively stratified or analyzed by sex. Information on participants’ sex was obtained from their clinical medical records. This study was approved by the Medical Ethics Committees of West China Hospital, Sichuan University and Shenzhen People’s Hospital (Approval No.: LL-KY-2021651) in accordance with the guidelines of the Declaration of Helsinki. All participants (or their legal guardians) provided written informed consent for participation in this study, the publication of indirect identifiers mentioned in Supplementary Data S1 and future related research. No financial compensation was provided to participants. For future studies, the collected samples were preserved by freezing or embedding in optimal cutting temperature (OCT) compounds. Detailed demographic and clinical characteristics of the patients are shown in Supplementary Data S1. No individual-level data are presented in a manner that could compromise anonymity, and all data sharing complied with the ethical approvals granted by the Institutional Review Boards of West China Hospital of Sichuan University and Shenzhen People’s Hospital.
Mice
C57BL/6 J mice were obtained from the Model Animal Research Center of Nanjing University. Piezo1loxP/loxP (STOCK Piezo1tm2.1Apat/J; 029213) and Il33loxP/loxP (B6(129S4)-Il33tm1.1Bryc/J; 030619) mice were obtained from GemPharmatech, Nanjing, China. Mice expressing ECspecific Cdh5-(PAC)-CreERT2 were provided by R. H. Adam. Cdh5-(PAC)-CreERT2 mouse was crossed with floxed Piezo1 mouse to generate CreERT2Piezo1loxP/loxP mice and CreERT2Il33loxP/loxP. CreERT2Piezo1loxP/loxP and CreERT2Il33loxP/loxP mice were intraperitoneally treated with tamoxifen (100 mg/kg) 2 months after birth for 6 days (interrupted for 3 days after the third dose), leading to endothelial-specific deletion of Piezo1(Piezo1ΔEC) and Il33(Il33ΔEC). Tamoxifen-treated Cre-negative littermates (Piezo1WT and Il33WT) were used as control for all experiments. All experiments used 8–10-week-old male specific pathogen-free (SPF) mice. The animals were housed in a temperature-controlled (22 ± 2 °C), humidity-regulated (50% ± 10% relative humidity) environment under a standard 12-hour light/dark cycle at the Experimental Center of West China Second University Hospital, Sichuan University. The facility maintains strict SPF conditions with continuous air filtration and regular monitoring of environmental parameters. Mice were group-housed in individually ventilated cages with sterile bedding and had free access to autoclaved standard rodent chow and water ad libitum. Bedding was changed weekly, and health status was monitored regularly. All animal experimental protocols were approved by the Laboratory Animal Ethics Committee of West China Second University Hospital, Sichuan University (approval number: 20220303070), and conducted in strict accordance with the institutional guidelines for the care and use of laboratory animals.
Cell culture
HEK293T (ATCC, #CRL-3216,) cells were purchased from the American Tissue Collection Center (ATCC, Manassas, VA) with validated cell identity and eliminated of mycoplasma contamination. HEK293T cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (C11995500BT, Gibco) with high glucose supplemented with 10% fetal bovine serum (FBS) (10099141, Gibco) and penicillin-streptomycin (Gibco). Human umbilical vein endothelial cells (HUVECs) were isolated from neonatal umbilical cord as described and cultured in endothelial Cell Medium (1001, ScienCell) at 37 °C in a humidified atmosphere of 5% CO2.
Public data
Single-cell RNA sequencing (scRNA) data were obtained from the Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/). We retrieved and utilized scRNA sequencing data from IPF patients and healthy individuals (by GSE13683117, GSE13277118, GSE12803319, GSE15935420 and GSE12296021). In addition, we collected extensive RNA sequencing data from IPF patients and normal lung tissue (courtesy of GSE47460)47.
Method specifics
Endothelial cell isolation
Immunomagnetic bead cell sorting is based on the ability of cell surface antigens to bind to specific antibodies attached to magnetic beads. Briefly, the Sheep anti Rat magnetic beads were pre-coated with CD31 (BD, #550274, 1:20 dilution) and CD45 (BD, #550539, 1:20 dilution) respectively. Harvested mouse lungs were washed with pre-cooled DPBS, cut into pieces, digested with digestion DMEM containing 1 mg/mL type I collagenase (Solarbio, #C8140), 1 mg/mL type IV collagenase (Biosharp, C#8160), 1 mg/mL Dispase II (Solarbio, #D6430), and 0.2 mg/mL DNAase (Solarbio, #BS165) in 37 °C for 40 min. After erythrocytes were lysed, cells were resuspended with MACS Buffer (Miltenyi Biotec, #130-091-376) and incubated on ice for 15 minutes with anti-mouse CD31 MicroBeads (for endothelial cell separation, Miltenyi Biotec, #130-097-418), anti-mouse CD45 MicroBeads (for immune cell separation, Miltenyi Biotec, #130-052-301), anti-mouse CD326 MicroBeads (for epithelial cell separation, Miltenyi Biotec, #130-105-958), and anti-mouse CD140a MicroBeads (for fibroblast separation, Miltenyi Biotec, #130-101-502). The MACS MultiStand separation rack (Miltenyi Biotec, #130-042-302), MidiMACS Separator (Miltenyi Biotec, #130-042-302), and LS Separation columns (Miltenyi Biotec, #130-042-401) were assembled and used to separate the cells, the isolated endothelial cells, epithelial cells, immune cells, and fibroblasts are used for Western blot and qPCR experiments.
AAV treatment
Mice were anesthetized by inhalation at 6–8 weeks of age, and after general anesthesia, the limbs of the mice were fixed with sterile tape, the skin of neck was scraped off and disinfected with alcohol, and the trachea was carefully cut open to expose the trachea. 2 × 10*11 Vector Genomes (v.g) of AAVENT-Tie1-NC (Negative control), AAVENT-TIE1-OE-Il33 (GeneChem, Shanghai), were injected into the trachea, respectively. The neck skin was carefully sutured and the efficiency of target gene expression was verified after 4 weeks.
Lung fibrosis model
An intratracheal BLM (Selleck) injection model was used to induce lung fibrosis. BLM sulfate powder was suspended and dissolved in sterile PBS and a dose of 1 unit/kg (50 μL/20 g) was intratracheally injected into mice at 8 weeks of age. During the injection process, mice were anesthetized with Isoflurane. Mice were euthanized 21 days after injection, and the lungs were harvested for the analysis of morphology and fibrosis.
Measurement of hydroxyproline
The hydroxyproline content in lung tissues was measured using Hydroxyproline (HYP) Assay Kit (Solarbio, #BC0255). Briefly, tissues of equal weight were cut into as many pieces as possible. Then, tissues are baked overnight in 12 N hydrochloric acid at 110 °C and neutralized with sodium hydroxide. Next, the samples were added to 1.4% chloramine T in 0.5 M sodium acetate/10% isopropanol and incubated for 20 min at room temperature. Ehrlich’s solution was added into samples for 30 min incubationat 65 °C. Finally, the absorbance values of all samples were measured at 560 nm on a SpectraMax Plus 384 full-spectrum microplate reader (Molecular Devices), and the Hydroxyproline content was calculated based on the fresh weight of tissues and the standard curve.
Histological analysis
The right lung was fixed in 4% paraformaldehyde for 24 h and then sent to Servicebio (Wuhan, China) for dehydration, paraffin embedding, and sectioning at 4 µm. H&E, Sirius Red and Masson staining were performed by Servicebio according to standard protocols. Whole-slide digital images were captured under identical settings using an Olympus VS120-FL-6 slide scanner. Quantitative analysis of H&E staining, Sirius red staining, and Masson staining was performed by evaluating five random fields in each section. The severity of pulmonary inflammation was graded based on the following criteria: 0, normal; 1, inflammatory cells occupying 0–10% of the entire tissue; 2, inflammatory cells occupying 10–40% of the entire tissue; 3, inflammatory cells occupying 40–70% of the entire tissue; 4, inflammatory cells occupying 70–100% of the entire tissue. For Sirius red staining or Masson staining, ImageJ software was used to define the red areas (in Sirius red staining) or blue areas (in Masson staining) in the tissue as positive areas. These areas were then outlined and their ratios to the entire field of view were calculated.
Immunofluorescence staining
Lung tissues from humans or mice were dehydrated after fixation with 4% paraformaldehyde (PFA) and subsequently embedded, cut into 4 μm sections. And then deparaffinized, rehydrated, sections were washed with PBS for a few minutes, then placed in sodium citrate buffer (pH 6.0) or EDTA buffer (pH 9.0) for antigen retrieval (The sections were microwaved on high for 10 min until they reached a slight boil, then the power was reduced to low for an additional 15 min). After cooling to room temperature, the sections were washed again with PBS and blocked with 10% donkey serum in PBS for one hour at room temperature. Then incubated with primary antibodies at 4 °C overnight. The sections were washed and then incubated with fluorochrome-conjugated secondary antibodies. Finally, the sections were counterstained with DAPI (Roche, #10236276001) and observed under an Olympus VS120-FL-6 slide scanner. The primary antibodies used were CD31 (Abcam, #ab9498; Servicebio, #GB13063), VE-cadherin (R&D Systems, #AF938), IL-33 (R&D Systems, #AF3626), PIEZO1(proteintech, #15939-1-AP) and αSMA (Abcam, #ab7817) antibodies.
Cytokine measurement by ELISA
The amounts of IL-33 in HUVECs supernatant were quantified using the Human IL-33 DuoSet ELISA kit (R&D Systems, #D3300B) according to the manufacturer’s instructions. Absorbance was read on a SpectraMax Plus 384 full-spectrum microplate reader (Molecular Devices) and the data were analyzed with OriginPro 9.1.
Protein extraction and western blotting
Isolated cells or tissues were kept on ice and lysed in RIPA buffer (Beyotime, #P0013B) supplemented with protease and phosphatase inhibitor cocktails (MCE). Protein concentration was determined with the BCA assay (beyotime, # P0010). 30–60 μg of protein was loaded per well on 4–20% SDS-PAGE gels (ACE, #ET15420LGel), transferred to ethanol-activated PVDF membranes (Thermo Fisher Scientific, #88518), blocked with 5% skim milk in TBST, and incubated overnight at 4 °C with primary antibodies diluted in TBST: IL-33 (R&D Systems, #AF3626, 1:1000), PIEZO1 (Proteintech, #15939-1-AP, 1:1 000), CALPAIN2 (Labome, #A03492, 1:1000), P-STAT3 (CST, #9134 s, 1:1000), STAT3 (Abcam, #ab68153, 1:2 000), P-STAT1 (CST, #9171, 1:1000), STAT1 (Proteintech, #10144-2-AP, 1:1000), GAPDH (Servicebio, #GB12002, 1:5000), β-actin (Proteintech, #66009-1, 1:5000), β-tubulin (CST, #2146, 1:5000). Membranes were then incubated with HRP-conjugated goat anti-rabbit or anti-goat IgG for 1.5 h at room temperature and developed with ECL Prime. Chemiluminescence was captured using a ChemiDoc MP Imaging System (Bio-Rad). Uncropped Western blot images are provided in the Source Data file.
Real-time quantitative PCR (qPCR)
Purified total RNA from sorted mouse lung cells (include endothelial cell) or HUVECs was extracted using TRIzol (ThermoFisher, #15596026) and reverse transcribed with PrimeScript RT reagent Kit (RR037A, Takara Bio). Gene expression was analyzed by qPCR with SYBR green chemistry (RR420A, Takara Bio). Relative expression level of mRNA was determined by normalizing each gene’s expression level to that of the internal reference gene GAPDH. A CFX96 Connect Real-Time PCR Detection System (Bio-Rad) was used for amplification and data acquisition. Relative expression was calculated by comparing the cycle threshold values using the 2-ΔΔCycle threshold method.
Cell mechanical experiment
Matrigel (356234, Corning BD) was diluted with serum-free medium or PBS. 2-3 ml of mixed matrix gel was added to the chambers (Naturethink) and incubated for 1 h at 37 °C. The unbound Matrigel was aspirated from the chambers and rinsed with serum-free medium or PBS. 1 × 105 Cells were seeded in the Matrigel-coated chambers. The chambers were gently shaken to distribute the cells evenly and then placed in the cell culture incubator for 24 h. When the fusion of cells in the chambers reached 80, chambers were attached to the manual retractor (Naturethink) and pull the chambers to the corresponding scale of the manual retractor. After pulling the chambers by 20%, cells were subjected to the pulling time for 0,6 and 24 h, the cells were then digested from the silica gel chambers with trypsin for further experiments.
Transfection with shRNA plasmid
shRNA oligos were annealed and ligated into AgeI/EcoRI (NEB, #R3552; #R3101)-digested pLKO.1-puro, transformed into competent cells (Tsingke, # DLC101), and sequence-verified. For lentivirus production, 2.5 μg of the pLKO.1-shRNA (or scramble) vector, psPAX2 and pMD2G (5:3:2) were co-transfected into 293 T cells using Lip6000 (Beyotime, #C0526). Viral supernatants were harvested at 48 h, used to infect HUVECs, followed by 1 μg/mL puromycin selection for 2 days. Knockdown efficiency was confirmed by qPCR or Western blot. Primer sequences: shPIEZO1 F 5′-CTCACCAAGAAGTACAATCAT-3′ R 5′-ATGATTGTACTTCTTGGTGAG-3′; shCAPN2 F 5′-TCACCTTGAATGAAGAAATTT-3′ R 5′-AAATTTCTTCATTCAAGGTGA-3′; shSTAT3 F 5′-CTCAGAGGATCCCGGAAATTT-3′ R 5′-AAATTTCCGGGATCCTCTGAG-3′.
Calpain activity
A fluorometric calpain activity assay kit (Abcam, #ab65308) was used to quantify calpain activity in the lysates of HUVECs by fluorescent substrate N-succinyl-LLVY-AMC. All samples were analyzed in triplicate with a multilabel reader (excitation, 400 nm; emission, 505 nm, Thermo, America) and expressed as relative fluorescent units (RFU).
scRNA/snRNA-seq: building libraries with the 10x Genomics platform
Single-cell RNA sequencing (scRNA-seq) and single-nucleus RNA sequencing (snRNA-seq) were performed using the 10X Chromium 3′ v3 kit (10X Genomics, Pleasanton, CA) according to the manufacturer’s instructions. Data processing included quality assessment using FASTQ and quality control using Cell Ranger and was aligned to the Homo sapiens GRCh38.p13 reference genome. The number of unique molecular identifiers (UMIs) and other data are shown in Supplementary Data S2.
scRNA/snRNA-seq data analysis workflow
The scRNA sequencing data were analyzed using the Seurat software package (v4.0)48. The detailed analysis workflow is available on the website (https://satijalab.org/seurat/). Any single cell with less than 200 genes or more than 5000 genes, and cells with more than 25% mitochondrial genes were regarded as low quality cells and excluded. Possible doublets were detected using DoubletFinder. Each dataset was log-normalized using the NormalizeData function. Genes with large differences in expression between cells were identified using the FindVariableFeatures function, and the data was scaled using ScaleData. Integrate each sample and correct batch effects using the Harmony49. Culusters were annotated using reported markers. the Subset function was used to extract endothelial cells etc. for further analysis. Intercellular communication and cellular function orchestration were explored using CellChat25. The Meta and metafor R packages were used for formal meta-analysis and to create forest plots.
Enrichment analysis and gene set scoring
Differential genes were enriched using Gene Ontology (GO) enrichment analysis. Mechanical stress (MS) scoring was performed using AddModuleScore function and the genes for scoring were the pseudobulk DEGs of snRNA-seq data enriched in “response to mechanical stimulus” (GO:0009612). Fibrosis scoring was performed using AddModuleScore function with the genes in GO pathway “collagen fibril organization” (GO:0030199). The gene lists are available as Supplementary Data S10.
“Scissors” identifying cell subpopulations associated with lung function
The “scissors method” integrates single-cell data with phenotypically relevant bulk sequencing data by first quantifying the similarity between each bulk sample and each cell sample. The “scissors method” can then optimize a regression model on a matrix associated with the sample phenotype to identify relevant subpopulations. A detailed description is given in the literature13.
scATAC-seq analysis using the 10x Chromium platform
We use the 10X Genomics Chromium single-cell ATAC-seq solution to prepare single cells for sequencing and the 10X Genomics single-cell ATAC-seq solution technology to prepare scATAC-seq libraries for sequencing. Sample and library preparation as well as instrument and sequencing parameters for generating scATAC-seq data on the 10x Chromium platform are detailed below and also available at https://support.10xgenomics.com/single-cell-atc.
Data processing of scATAC-seq
Data processing included quality assessment using FASTQ and quality control was performed using Cell Ranger and sequencing reads was aligned to the Homo sapiens GRCh38.p13 reference genome.Cells were filtered using the following criteria: (1) 3000<nCount_peaks<50000; (2) pct_reads_in_peaks>15; (3) blacklist_ratio<0.05; (4) Nucleosome signal<4; (5) transcription start site (TSS) enrichment score>1.Peaks calling was performed using MACS2.
Combined analysis of scRNA-seq and scATAC-Seq
To perform an integrated analysis of the scATAC-seq dataset and the scRNA-seq dataset, we utilized the anchoring strategies provided by the R packages Seurat and Signac. Initially, we identified a set of cell pairs or “anchors” between the datasets through the FindTransferAnchors function using typical correlation analysis (CCA) and mutual nearest neighbors (MNNs) methods, which are hypothesized to represent similar biological states. These anchors were the basis for integrating the two datasets. Subsequently, we used the TransferData function to project the clustering labels from the scRNA-seq data onto the scATAC-seq data, which allowed us to evaluate single-cell measurements in different data modalities.
Motif analysis
Motif analysis was performed using Signac package mainly. Transcription factor (TF) binding motifs were downloaded from JASPAR database. Then TF binding motifs were mapped to scATAC-seq peaks using AddMotifs function. RunChromVAR function was used for calculating motif deviation score. We performed differential accessibility analysis with FindMarkers function and obtained corresponding motifs using FindMotifs function. MotifPlot function was used to visualize motifs of peaks with different accessibility between groups.
Bulk RNA sequencing and gene expression analysis
In our study, we performed bulk RNA sequencing of endothelial cells isolated from human and mouse lung tissue. Cells were lysed with RLT Plus buffer immediately after sorting. Total RNA extraction was performed using Qiagen’s RNeasy Plus Mini Kit. We then prepared full-length RNA sequencing libraries and performed 2 × 75 bp paired-end sequencing using a 150-cycle Nextseq 500 High Output V2 kit. The RNA-seq data compilation included at least three independent biological replicates to ensure robust and reproducible results.
Statistical Analysis
All statistical analyses were performed using Prism 10 and R software by investigators who were not blinded to group allocation during sample processing (genotype/treatment identity was required), but all data analyses were subsequently conducted by independent researchers using identical automated scripts to minimize subjective bias; unpaired two-tailed Student’s t-tests were used for two-group comparisons and one- or two-way ANOVA followed by Tukey’s or Šídák’s multiple-comparisons test for multiple groups, with exact tests and p-values reported in the legends, p < 0.05 considered significant, sample sizes (n) stated, and data presented as mean ± s.e.m.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The snRNA-seq data of human lung tissue used in this study are available in the GEO database under accession number GSE314709. The scRNA-seq and scATAC-seq data of mice used in this study are available in the GEO database under accession number GSE314943. The data supporting the findings from this study are available within the manuscript and its supplementary information. Source data are provided with this paper. In this study, public datasets used for analysis including GSE47460, GSE136831, GSE132771, GSE128033, GSE159354 and GSE122960. Source data are provided with this paper.
Code availability
All the bioinformatic analyses were performed according to the official recommended workflow. Codes used in this study could be found at https://github.com/houruijiejerry/RuijieHou-Code, https://doi.org/10.5281/zenodo.17960326.
References
Lederer, D. J. & Martinez, F. J. Idiopathic pulmonary fibrosis. N. Engl. J. Med. 378, 1811–1823 (2018).
Martinez, F. J. et al. Idiopathic pulmonary fibrosis. Nat. Rev. Dis. Prim. 3, 17074 (2017).
Zepp, J. A. & Morrisey, E. E. Cellular crosstalk in the development and regeneration of the respiratory system. Nat. Rev. Mol. Cell Biol. 20, 551–566 (2019).
Bian, F. et al. Lung endothelial cells regulate pulmonary fibrosis through FOXF1/R-Ras signaling. Nat. Commun. 14, 2560 (2023).
Yanagihara, T., Guignabert, C., Kolb, M. R. J. Endothelial cells in pulmonary fibrosis: more than a bystander. Eur. Respir. J. 61, (2023).
Cao, Z. et al. Targeting of the pulmonary capillary vascular niche promotes lung alveolar repair and ameliorates fibrosis. Nat. Med. 22, 154–162 (2016).
Chen, Y. et al. Aging reprograms the hematopoietic-vascular niche to impede regeneration and promote fibrosis. Cell Metab. 33, 395–410.e394 (2021).
Romani, P., Valcarcel-Jimenez, L., Frezza, C. & Dupont, S. Crosstalk between mechanotransduction and metabolism. Nat. Rev. Mol. Cell Biol. 22, 22–38 (2021).
Cui, X. et al. Mechanical forces regulate asymmetric vascular cell alignment. Biophys. J. 119, 1771–1780 (2020).
Baeyens, N., Bandyopadhyay, C., Coon, B. G., Yun, S. & Schwartz, M. A. Endothelial fluid shear stress sensing in vascular health and disease. J. Clin. Invest. 126, 821–828 (2016).
Mehta, V. et al. Mechanical forces regulate endothelial-to-mesenchymal transition and atherosclerosis via an Alk5-Shc mechanotransduction pathway. Sci. Adv. 7, (2021).
Collins, C. et al. Haemodynamic and extracellular matrix cues regulate the mechanical phenotype and stiffness of aortic endothelial cells. Nat. Commun. 5, 3984 (2014).
Sun, D. et al. Identifying phenotype-associated subpopulations by integrating bulk and single-cell sequencing data. Nat. Biotechnol. 40, 527–538 (2022).
Fainberg, H. P. et al. Forced vital capacity trajectories in patients with idiopathic pulmonary fibrosis: a secondary analysis of a multicentre, prospective, observational cohort. Lancet. Digit. Health 4, e862–e872 (2022).
Raghu, G. et al. Idiopathic pulmonary fibrosis (an Update) and progressive pulmonary fibrosis in adults: an official ATS/ERS/JRS/ALAT clinical practice guideline. Am. J. Resp. Crit. Care Med. 205, e18–e47 (2022).
Pastre, J. et al. Idiopathic pulmonary fibrosis patients with severe physiologic impairment: characteristics and outcomes. Respir. Res. 22, 5 (2021).
Adams, T. S. et al. Single-cell RNA-seq reveals ectopic and aberrant lung-resident cell populations in idiopathic pulmonary fibrosis. Sci. Adv. 6, eaba1983 (2020).
Tsukui, T. et al. Collagen-producing lung cell atlas identifies multiple subsets with distinct localization and relevance to fibrosis. Nat. Commun. 11, 1920 (2020).
Morse, C. et al. Proliferating SPP1/MERTK-expressing macrophages in idiopathic pulmonary fibrosis. Eur. Respir. J. 54, (2019).
Gao, X. et al. Osteopontin links myeloid activation and disease progression in systemic sclerosis. Cell Rep. Med. 1, 100140 (2020).
Reyfman, P. A. et al. Single-cell transcriptomic analysis of human lung provides insights into the pathobiology of pulmonary fibrosis. Am. J. Respiratory Crit. Care Med. 199, 1517–1536 (2019).
Ding, B. S. et al. Inductive angiocrine signals from sinusoidal endothelium are required for liver regeneration. Nature 468, 310–315 (2010).
Hogan, B. L. M. Integrating mechanical force into lung development. Dev. Cell 44, 273–275 (2018).
Tang, Z. et al. Mechanical forces program the orientation of cell division during airway tube morphogenesis. Dev. Cell 44, 313–325.e315 (2018).
Mora, A. L., Rojas, M., Pardo, A. & Selman, M. Emerging therapies for idiopathic pulmonary fibrosis, a progressive age-related disease. Nat. Rev. Drug Discov. 16, 755–772 (2017).
Wu, H. et al. Progressive pulmonary fibrosis is caused by elevated mechanical tension on alveolar stem cells. Cell 180, 107–121.e117 (2020).
Shiraishi, K. et al. Biophysical forces mediated by respiration maintain lung alveolar epithelial cell fate. Cell 186, 1478–1492.e1415 (2023).
Zeng, H. et al. Suppression of PFKFB3-driven glycolysis restrains endothelial-to-mesenchymal transition and fibrotic response. Signal Transduct. Target Ther. 7, 303 (2022).
Caporarello, N. et al. Dysfunctional ERG signaling drives pulmonary vascular aging and persistent fibrosis. Nat. Commun. 13, 4170 (2022).
Olajuyin, A. M., Zhang, X. & Ji, H. L. Alveolar type 2 progenitor cells for lung injury repair. Cell Death Discov. 5, 63 (2019).
Snijder, J., Peraza, J., Padilla, M., Capaccione, K. & Salvatore, M. M. Pulmonary fibrosis: a disease of alveolar collapse and collagen deposition. Expert Rev. Respir. Med. 13, 615–619 (2019).
Matthay, M. A. et al. Acute respiratory distress syndrome. Nat. Rev. Dis. Prim. 5, 18 (2019).
Nho, R. S., Ballinger, M. N., Rojas, M. M., Ghadiali, S. N. & Horowitz, J. C. Biomechanical force and cellular stiffness in lung fibrosis. Am. J. Pathol. 192, 750–761 (2022).
Dessalles, C. A., Leclech, C., Castagnino, A. & Barakat, A. I. Integration of substrate- and flow-derived stresses in endothelial cell mechanobiology. Commun. Biol. 4, 764 (2021).
Tamargo, I. A., Baek, K. I., Kim, Y., Park, C. & Jo, H. Flow-induced reprogramming of endothelial cells in atherosclerosis. Nat. Rev. Cardiol. 1–16 (2023).
Mehta, V. et al. The guidance receptor plexin D1 is a mechanosensor in endothelial cells. Nature 578, 290–295 (2020).
Douguet, D., Patel, A., Xu, A., Vanhoutte, P. M. & Honoré, E. Piezo ion channels in cardiovascular mechanobiology. Trends Pharm. Sci. 40, 956–970 (2019).
Cayrol, C. & Girard, J. P. Interleukin-33 (IL-33): a nuclear cytokine from the IL-1 family. Immunol. Rev. 281, 154–168 (2018).
Ding, X. et al. The IL-33-ST2 pathway contributes to ventilator-induced lung injury in septic mice in a tidal volume-dependent manner. Shock 52, e1–e11 (2019).
Zhang, Y. et al. Piezo1-mediated mechanotransduction promotes cardiac hypertrophy by impairing calcium homeostasis to activate calpain/calcineurin signaling. Hypertension 78, 647–660 (2021).
Yu, Z. Y. et al. Piezo1 is the cardiac mechanosensor that initiates the cardiomyocyte hypertrophic response to pressure overload in adult mice. Nat. Cardiovasc. Res. 1, 577–591 (2022).
Chen, X. et al. A feedforward mechanism mediated by mechanosensitive ion channel PIEZO1 and tissue mechanics promotes glioma aggression. Neuron 100, 799–815.e797 (2018).
Yao, M. et al. Force- and cell state-dependent recruitment of Piezo1 drives focal adhesion dynamics and calcium entry. Sci. Adv. 8, eabo1461 (2022).
Xue, Y. et al. The mechanotransducer Piezo1 coordinates metabolism and inflammation to promote skin growth. Nat. Commun. 16, 6876 (2025).
Konig, S., Jayarajan, V., Wray, S., Kamm, R. & Moeendarbary, E. Mechanobiology of the blood-brain barrier during development, disease and ageing. Nat. Commun. 16, 7233 (2025).
Drake, L. Y. et al. Mechanical stretch promotes sustained proliferation and inflammation in developing human airway smooth muscle. Am. J. Physiol. Lung Cell Mol. Physiol. 329, L296–l306 (2025).
Tan, J. et al. Expression of RXFP1 is decreased in idiopathic pulmonary fibrosis. implications for relaxin-based therapies. Am. J. Respiratory Crit. Care Med. 194, 1392–1402 (2016).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Acknowledgments
This study was supported by the National Key R&D Program of China (No. 2022YFF0710803), the Shenzhen Medical Research Fund (Nos. B2302041 and C2302001), the National Key R&D Program of China (Nos. 2022YFF0710800 and 2022YFF0710802), the National Natural Science Foundation of China (Nos. 81900065, 82470033, 82302049, and 82222035), the Fundamental and Applied Basic Research Fund of Guangdong Province (No. 2021A1515111036), the Natural Science Foundation of Sichuan Province (No. 2022NSFSC1394), International Science and Technology Cooperation Project of Chengdu (No. 2023-GH02-00092-HZ), Chengdu Municipal Bureau of Science and Technology, 2023 Open Fund of State Key Laboratory of Respiratory Diseases (No. SKLRD-OP-202304), and the Scientific Application and Foundation Project of Science and Technology Department of Sichuan Province (No. 2023NSFSC1702).
Author information
Authors and Affiliations
Contributions
L.-L.Z., S.C., B.D., L.R, H.Z. conceived the study and wrote the manuscript. L.-L.Z., X.G., R.H., L.J., S.X. performed experiments, analyzed and interpreted data. X.Z., Q.-F.M., Q.L., X.S., B.G., R.L., L.Y., X.C., H.X., P.W., X.T., L.L., B.L., R.C., C.S., W.X. performed the experiments. Y.-Y.F., Z.C., L.-F.Z., L.-W.W., L.-S.Z. participated in discussions. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, L., Gui, X., Hou, R. et al. Single-cell multiomics uncovers an endothelial mechanosensitive PIEZO1-IL-33 axis driving pulmonary fibrosis. Nat Commun 17, 2655 (2026). https://doi.org/10.1038/s41467-026-70193-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70193-w