Abstract
Copy number variation (CNV) plays a fundamental role in modulating plant agronomic traits and tumorigenesis in animals. While frequently linked to replication stress, the mechanisms giving rise to CNVs are not fully elucidated. Here we characterize the mutational consequences associated with losing the conserved TONSOKU (TSK/TONSL) pathway (CAF-1-H3.1-TSK), which is required to resolve impaired DNA replication forks. Using Arabidopsis thaliana, we demonstrate that tsk mutants rapidly accumulate large, heritable tandem duplications within their genomes that are consistent with DNA Polymerase θ (Pol θ) activity. These duplications are associated with late replicating heterochromatin enriched in sources of replication stress. We also show that stochastic developmental phenotypes in tsk plants are the result of the DNA Damage Response (DDR), with phenotype suppression occurring when ATR–WEE1 checkpoint signaling is removed. We thus describe a previously uncharacterized source of large tandem duplications that are relevant to understanding genome stability in diverse eukaryotes, and in disease contexts.
Similar content being viewed by others
Introduction
CNVs, where large segments of the genome are either lost or duplicated, are a frequently observed class of structural variants. In many cases, they underlie key agricultural traits selected for during plant domestication and, in humans, they occur somatically in many cancers1,2,3,4,5. In the latter case, CNVs are considered key components of intratumor heterogeneity6,7,8, which can give rise to resistance to specific cancer treatments. The classification of CNVs therefore has important biomedical applications in categorizing tumors, guiding appropriate courses of treatment and predicting patient prognosis4,8,9.
Tandem duplications are a common class of CNV. When such duplications arise frequently across the genome of cancerous tissue, it is referred to as a tandem duplicator phenotype, which is further classified by the size of the duplications5,10. For example, cells deficient in the homologous recombination (HR) DNA repair protein BRCA1 form tandem duplications that span ~11 kb (Group 1), while accumulations of larger duplications (~231 kb, Group 2; ~1.7 Mb, Group 3) are associated with other genetic drivers found in specific cancers5. All three groups of tandem duplications are linked to the occurrence of stress at replication forks, leading to DNA breaks and the opportunity for duplications to arise11.
Central to the resolution of stress occurring at stalled or collapsed replication forks is the widely conserved protein TONSOKU-LIKE (TONSL), which directly interacts with unmethylated histones H3.1/H3.2 and H4 incorporated into newly synthesized DNA by CAF-112,13,14,15,16. In concert with its partner MMS22L, TONSL participates in the removal of RPA at resected ssDNA and loading of the HR recombinase RAD5112,13,17. Loss of TONSL functionality is thus expected to induce genome instability, however, whether this is the case is unknown. Null tonsl mutations in vertebrates are lethal at the organism level18,19, thus making it difficult to assess the mutational profile associated with lack of TONSL activity.
In Arabidopsis thaliana (Arabidopsis), the TONSL ortholog (TONSOKU/TSK) is not required for viability or fertility, although tsk mutants exhibit many developmental phenotypes20,21,22,23,24. Arabidopsis is thus a unique system to study the genomic and developmental consequences that occur when TSK is lost. The TSK gene has long been associated with genome stability in Arabidopsis, as tsk mutants are highly sensitive to genotoxic stress21,23. This suggests that tsk plants are unable to adequately repair DNA damage, which we hypothesized may lead to elevated levels of mutations over developmental time and/or generations.
Results
Large CNVs arise in the absence of TSK
To investigate the genomic stability of tsk mutants, we performed whole-genome sequencing (WGS; Illumina) on three tsk-4 mutant plants (SALK_034207) that had been grown for multiple generations (i.e., Gen. X plants). Across the genomes of these plants, we detected background levels of single-nucleotide variants (SNVs) and small insertions/deletions (indels; <50 bp) (Supplementary Fig. 1A). In contrast, a substantial number of large CNVs were readily apparent in the genomes of tsk-4 mutants (Fig. 1A). While we did not observe CNVs consistent with fixed deletions events (one exception being a 699 bp deletion present in Chromosome 4 of plants #2 and #3), we found 42, 31 and 44 amplifications in the three tsk-4 Gen. X plants (Fig. 1B; Supplementary Data 1). The mean coverage level of these amplifications was 1.85× over Col-0, consistent with duplications of these regions (Supplementary Fig. 1B). These duplications range in size from less than 1 kb to nearly 1.5 Mb, with 44.6–74.4% being greater than 100 kb (Fig. 1B). Together, they add 13.5–18.8 Mb of DNA to the genomes of the three tsk-4 Gen. X plants, an increase of 6.4–7.9% over the diploid genome size of Arabidopsis (TAIR10) (Fig. 1C). While some duplications were shared between the tsk-4 Gen. X plants, a majority were only observed in a single plant. This suggests that the duplications arose independently.
A Genome tracks of the log2 ratio of tsk-4 Gen. X normalized coverage (BPM) to that of a Col-0 sample across Chromosome 3. The bar below represents the chromosome with constitutive heterochromatin plotted in black45. B Size of CNVs in the tsk-4 Gen. X plants where each point is an individual CNV. Region ≤100 kb shaded in dark gray, region ≤400 kb shaded in light gray. C Cumulative CNV increase over haploid genome size in the tsk-4 Gen. X plants. D Proportion of transcripts with ≥2-fold (red), or ≤2-fold (blue) abundance (TPM) above a Col-0 sample in the tsk-4 Gen. X. plants. Transcripts are classified by whether their encoding genes are within a CNV (+) or not (−). Source data are provided as a Source Data file.
CNVs in tsk-4 plants are expected to interfere with transcriptional activity25,26, and likely account for the previously observed local clustering of upregulated genes in tsk-4 mutants24. To assess this, we performed paired WGS and RNA-seq on three plants grown without a functional TSK gene for three generations. These plants had 22, 18 and 27 duplications, respectively. RNA-seq analysis showed that transcript abundance was increased across the duplicated regions, with average increases of 5.2, 3.3 and 9.7 fold in the three tsk-4 plants. Overall, 38–46% of transcripts within duplicated regions, and a further 12–15% of transcripts outside of duplications, had abundances >2 fold greater than Col-0 (Fig. 1D).
CNVs in tsk plants and associated mutants correspond to tandem duplications
CNVs detected using short-read WGS can represent different structural arrangements within the genome. Examining alignments at the edges of CNV regions (>100 kb) in tsk-4 mutants revealed novel junctions between the end and the start of duplicated segments based on the presence of chimeric reads partially mapping to both ends of the CNVs. In the three tsk-4 Gen. X plants, 88%, 93% and 79% of the CNVs were associated with these chimeric reads, respectively. This result suggests that the CNVs are tandem duplications, with the chimeric reads representing breakpoint junctions between the original segment and its duplicated copy integrated into the chromosome in the same orientation (Fig. 2A). However, cells experiencing genomic instability have also been commonly reported to contain extrachromosomal DNA (ecDNA), which oftentimes form circular structures27. It is therefore possible that the CNVs in tsk-4 mutants correspond to ecDNA, with the chimeric reads representing the point at which the circles close.
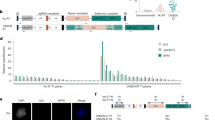
A Schematic of tandem duplication relative to reference sequence (left panel). The tandem duplication creates chimeric reads (green) across the breakpoint junction that traverse the end of the first segment to the start of the second (middle panel). When aligned to the reference genome, tandem duplications manifest as regions of increased coverage (right panel). Created in BioRender. Thomson, G. (2025) https://BioRender.com/w69kcxm. B A representative CNV (157 kb) in tsk-4 plants characterized by optical mapping. The top track depicts the log2 ratio of normalized coverage (BPM) on the TAIR10 genome to that of a Col-0 using WGS (Illumina) sequencing and the distribution of DLE-1 sites on the genome. The lower track shows the repeated pattern of DLE-1 sites anticipated to be present in the genome of the tsk-4 plants if the CNV is a tandem duplication. Representative DNA molecules mapping to this CNV location and spanning 1) a breakpoint junction (red), 2) a breakpoint junction and one breakend (yellow), or 3) a breakpoint junction and both breakends (green). C Genome tracks showing a CNV in a mgo1 CRISPR line. Tracks are the log2 ratio of normalized coverage (BPM) to that of a Col-0 sample. Bar below represents the chromosome/chromosomal region with constitutive heterochromatin plotted in black45. D Genome tracks depicting CNVs present in fas1-4 x fas2-4, H3.1kd, and tsk-4 from previously published data. Tracks are the log2 ratio of normalized coverage (BPM) to that of a Col-0 sample. The bar below represents the chromosome with constitutive heterochromatin plotted in black45. Source data are provided as a Source Data file.
Because short-read WGS is unable to effectively distinguish between tandem duplications and circular ecDNA, we performed optical genome mapping on the pooled progeny of a previously sequenced tsk-4 (third generation) plant. Comparison of the resulting mapped DNA molecules and 11 CNVs observed in the parent plant revealed long DNA molecules covering both copies of the duplication as well as flanking sequences for all CNVs up to 178 kb in size (6 CNVs) (Fig. 2B; Supplementary Data 2). While definitive results for the remaining larger 5 CNVs were limited by technical constraints of the assay, long DNA molecules consistent with tandem duplications were also found for these remaining CNVs (Supplementary Data 2, 3). These results demonstrate that the CNVs detected in the absence of TSK are tandem duplications integrated into the Arabidopsis chromosomes.
TSK is also known as MGO3, as tsk-4 plants phenotypically resemble mgo1 and mgo2 single mutants, which have disrupted meristem organization20,28. TSK/MGO3 has been proposed to act in the same pathway as MGO120, which encodes a type 1B topoisomerase (TOP1α)29. WGS of second-generation mgo1 mutants (Supplementary Fig. 1C) revealed tandem duplications (35.5 kb and 74.7 kb in size) (Fig. 2C, Supplementary Fig. 1D), consistent with a shared mechanism involving both TSK and TOP1α. A small number of tandem duplications have also been previously reported in plants deficient in the CAF-1 chaperone complex (≥fourth-generation mutants)26, which loads the specific histone variant that TSK binds, H3.1, onto newly replicated chromatin15,30,31. By re-analyzing published data, we also observed duplications in the genomes of H3.1 knockdown (H3.1kd) lines and other tsk-4 plants (Fig. 2D)32,33. These results point to a previously unrecognized molecular pathway (i.e., CAF-1, H3.1, TSK and TOP1α) required to maintain genome stability at replication forks by preventing tandem duplications.
TSK guards against somatic and germline duplications generated via the activity of Pol θ
The disparity in the number of tandem duplications observed in tsk-4 Gen. X plants (Fig. 1A), and those of the published datasets (Fig. 2D), suggests that tsk-4 plants accumulate CNVs over time. We thus tested if the duplications are heritable. From a population of plants segregating for the tsk-4 mutation, we identified three plants where tsk-4 was fixed for the first time (first-generation mutants). We sequenced those plants, along with seven descendants each, across two generations. Consistent with generational accumulation, the number of observed duplications increased from one generation to the next (Fig. 3A). Intriguingly, the rate at which new duplications arise appeared to increase across generations (Fig. 3B).
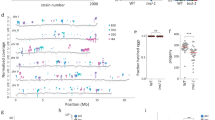
A Family trees of sequenced tsk-4 plants. Each circle represents an individual plant with duplication counts. Numbers in brackets indicate the number of inherited duplications. Generation 1 represents the first time that lineage has been homozygous for tsk-4. B The accumulation of duplications with each generation from three families. Each point is an individual plant (n = 3, 12 and 9 plants for each generation, respectively). Colors correspond to the families shown in panel (A). Bars represent means with 95% confidence intervals. Lower case letters represent pairwise comparisons of a one-way ANOVA (F(2, 21) = 17.01, p = 4.05 × 10−05, ω²=0.57) with Tukey’s Honestly Significant Difference post hoc testing. C Size of duplications arising in each generation of plants from the three families where each point is an individual duplication. Region ≤100 kb shaded in dark gray, region ≤400 kb shaded in light gray. Duplications in yellow are passed on to the next generation and the distribution of their sizes was not detectably different from those not inherited (Wasserstein permutation test, W1 = 0.304, p = 0.45, 10,000 permutations, 95% CI: 0.199–0.733). D Diagram of tissues sampled from a single Arabidopsis plant. Created in BioRender. Thomson, G. (2025) https://BioRender.com/kz793yn. E UpSet plots showing the number of duplications observed in all tissue samples, or only a subset of tissues in two independent second-generation tsk-4 plants. F Genome track snapshots of the log2 ratio of normalized coverage (BPM) for different tissues from the same plant to that of a Col-0 sample. Green arcs represent observed chimeric reads indicative of internal breakpoint junctions. Tissue icons created in BioRender. Thomson, G. (2025) https://BioRender.com/xzcijqz. G Size of individual duplications in tsk-4 ku70-2 and tsk-4 ku80-7 plants, and single mutants grown alongside. Each point is an individual duplication. Region ≤100 kb shaded in dark gray, region ≤400 kb shaded in light gray. PERMANOVA on pairwise 1-Wasserstein distances between plant-level log2 tandem duplication size distributions were performed for tsk-4 and tsk-4 ku70-2 (pseudo-F = 0.656, R2 = 0.099, adj. p = 0.609) and tsk-4 and tsk-4 ku80-7 (pseudo-F = 1.73, R2 = 0.257, adj. p = 0.381). H Representative siliques of Col-0 and tsk-4+/− teb-5 plants. Arrows indicate aborted seeds. The scale bar is 1 cm. I Histogram of mutation size at the junctions of a set of 204 duplications. Mutations less than zero indicate deletions with microhomologies, and mutations greater than zero have insertions. Insertions with adjacent templates are plotted in yellow (limited to the sequence of the read itself), while those without are in blue. Source data are provided as a Source Data file.
Of the duplications that could be tracked across generations, 69% (33/48) were first observed to have normalized read depths ~50% larger than WT, consistent with heterozygous events in which one allele was duplicated and inherited via the germline. Additionally, 25% (13/51) of inheritance events led to a further doubling of the read depth in the next generation, consistent with them becoming homozygous (Supplementary Fig. 1E). These results support that the duplications are inherited in a Mendelian fashion. When the size of the duplications is compared between generations, the inherited duplications show the same distribution as those not observed to be inherited (Fig. 3C).
In light of TSK’s established role in DNA replication, we inferred that duplications may arise concurrently throughout the plant, as DNA replication occurs in all tissue types over the plant’s lifetime. To test this, samples were taken from multiple tissues (i.e., flowers, cauline leaves, rosette leaves, and roots) of the same second-generation tsk-4 plant (Fig. 3D). While the majority of duplications in a given sample were shared amongst the different tissues, tissue-specific events were also observed in all sampled tissue types (Fig. 3E, F). These results indicate that duplications are continually being generated throughout the life of the plant.
Tandem duplications can arise from a DNA repair pathway incorrectly resolving damaged DNA2,11. Since TSK participates in the HR DNA repair pathway12,13,15,17, we reasoned that another repair pathway may be acting on the damaged DNA in tsk-4 mutants. The classical nonhomologous end-joining pathway (c-NHEJ) is the dominant mechanism for repairing double-stranded DNA breaks in eukaryotic cells, with the Ku heterodimer (Ku70/Ku80) being a key player34,35,36. We generated both tsk-4 ku70-2 and tsk-4 ku80-7 double mutants, and observed comparable levels of duplications between these plants and tsk-4 plants grown alongside (Fig. 3G). Thus, c-NHEJ is not responsible for the tandem duplications in tsk-4 mutants.
We next tested the involvement of Pol θ, the key component of the theta-mediated end joining (TMEJ) repair pathway. Pol θ has long been associated with CAF-1 and TSK on account of similar developmental phenotypes when mutated37, and it plays a central role in maintaining the meristem when DNA replication is impaired38. We crossed tsk-4 to teb-5, a mutant in which Pol θ activity is lost37. Strikingly, we did not recover tsk-4 teb-5 double mutants from this cross, even after genotyping the progeny of tsk-4+/− teb-5 mutants (Supplementary Data 4). Looking at the siliques of tsk-4+/− teb-5 plants, we found a subset of seeds aborted (Fig. 3H), indicating that teb-5 is synthetic lethal with tsk-4.
The TMEJ pathway has been shown to cause characteristic small deletions by joining two sequences that share microhomology, or small templated insertions, which often match a sequence adjacent to the DNA break39. To assess whether TMEJ contributes to the formation of tandem duplications, we analyzed the sequence of the chimeric reads spanning the tandem duplication breakpoint junctions (Fig. 2A). Mutational signatures consistent with TMEJ were identified in 85.3% of the 204 tandem duplications assessed (Fig. 3I). Specifically, 61.8% of breakpoint junctions had small deletions with microhomologies at the intersection, 13.2% had insertions from which an identical template could be identified in adjacent sequences, and 10.3% had an insertion for which no template was immediately apparent. These results suggest that the majority of duplications in tsk-4 mutants are created by the activity of TMEJ, which acts as a backup repair pathway when TSK activity is lost. This conclusion is supported by a recent study in Caenorhabditis elegans, which demonstrates a key role for TMEJ in creating tandem duplications in the absence of the TSK orthologue TONSL40.
Tandem duplications preferentially arise in repetitive, late-replicating regions of the genome
Tandem duplications in tsk-4 mutants are widely distributed across the genome (Fig. 1A). To investigate their association with known genomic features, we assembled a large set of independent duplications >10 kb in size (n = 481; median span 199 kb; 84.4% >100 kb; Supplementary Data 5). The level of intersection among these duplications matched random expectation (Fig. 4A, B), as did their overlap with different chromatin states (Fig. 4C). However, a trend towards depletion in euchromatin (states E1-11) and enrichment in heterochromatic states (states H1-6) was present (Fig. 4C).
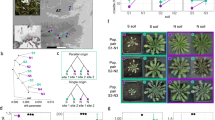
A Diagram of overlap quantification between genomic regions and the duplications. Created in BioRender. Thomson, G. (2025) https://BioRender.com/poypy51. B Overlap of 481 independent duplications on the genome (red), 10,000 random simulations plotted (gray), and their average (green). C Overlap of duplications with chromatin states45; constitutive heterochromatin (H1-6), facultative heterochromatin (F1-6), intergenic (I1-3), and euchromatin (E1-11). Blue arrows indicate observed overlap is in the bottom 2.5% of shuffled datasets. Line colors same as (B). D Diagram of the duplication breakends (+/−50 bp). Created in BioRender. Thomson, G. (2025) https://BioRender.com/poypy51. E Mean GC content of observed duplication breakends (red line) relative to the means of 10,000 random simulations (gray histogram). F Intersection of CNV breakends with relative timing of DNA replication during S phase44. E: Early S; EM: Early-Mid S; M: Mid S; ML: Mid-Late S; L: Late S. Line colors same as (B). Yellow and blue arrows indicate observed overlap is in the top or bottom 2.5% of shuffled datasets, respectively. G Intersection of duplication breakends with annotated protein coding genes97, or transposons122. Colors same as (B). H Mean number of breakend overlaps with S phase expressed genes116 in observed duplication breakends (red line) relative to the means of 10,000 random simulations (gray histogram). I Diagram of the border regions (+/−20 kb) around the duplication breakends. Created in BioRender. Thomson, G. (2025) https://BioRender.com/poypy51. J–N Mean intersection of genomic features with regions around duplication border regions. Observed levels (red lines) are plotted relative to 10,000 random simulations (gray histograms). These features are (J) tandem duplications in Arabidopsis accessions50,123, K the amount of annotated short tandem repeat sequence, L transposon sequence122, M chromatin boundaries63, and N T-DNA insertions67. In all graphs, the simulation mean histograms are scaled from zero to one. Pr is the probability (0 to 1) of obtaining a simulated set of duplications having a lower value relative to the observed. Source data are provided as a Source Data file.
Examination of the duplication breakends (+/- 50 bp of junctions with unduplicated genome; Fig. 4D) showed a modest 1.5% increase in mean GC content (Fig. 4E), though no clear pattern was seen across them (Supplementary Fig. 2A). Motif enrichment analysis found two overrepresented motifs, a poly(dA:dT) tract, and one with a guanine every 3 bp (Supplementary Fig. 2B). These motifs, which rarely co-occurred (Supplementary Fig. 2C), may reflect distinct sources of replication stress. Short tandem repeats such as these motifs have been demonstrated to cause replication stress via fork slippage41,42,43. We found that short tandem repeats are enriched at breakends (Supplementary Fig. 2D), and this is attributable to these two motifs (Supplementary Fig. 2E).
A trend towards constitutive heterochromatin was again observed at breakends (Supplementary Fig. 2F) with enrichment also present for late-replicating chromatin (Fig. 4F)44, and chromatin modifications associated with constitutive heterochromatin when analyzed on an individual basis (Supplementary Fig. 2G–M)45,46,47.
In cancer cells, transcriptional-replication collisions are major causes of replication fork collapse and large CNVs4. In tsk-4, most duplication breakends intersect with transcribed regions in protein-coding genes (49.9%) or transposons (TE) (24.7%) (Fig. 4G). We do not interpret this as evidence of transcriptional-replication collisions, as the high level of intersection with protein-coding genes is consistent with the Arabidopsis genome being gene dense (approximately one gene/4–5 kb)48. Furthermore, breakends intersecting protein-coding genes showed no bias on average towards transcriptional start (TSS) or termination (TTS) sites (Supplementary Fig. 2N), as would be expected if they stemmed from collisions with paused RNA polymerase II. Moreover, no association was detected with gene size (Supplementary Fig. 2O), and genes highly expressed during S phase were found to be depleted at breakends (Fig. 4H). Together, these findings suggest that in the absence of TSK activity, transcriptional-replication collisions are unlikely to be a common cause of the observed tandem duplications.
Replication fork collapse can lead to long resection tracts when HR is impaired49. We thus extended our analysis to the regions bordering the duplication breakends (+/−20 kb; Fig. 4I). We found that these border regions are enriched for single-nucleotide variants, indel variants and tandem duplications present in diverse Arabidopsis accessions (Fig. 4J, Supplementary Fig. 3A–C)50, consistent with elevated replication stress in some regions of the genome. These regions also showed a 67% increase on average in short tandem repeat tract length (Fig. 4K, Supplementary Fig. 3D, E). TE coverage was also increased by 22% in these regions (Fig. 4L, Supplementary Fig. 3F). LTR retrotransposons are particularly relevant, as they comprise approximately one-third of replication origins in Arabidopsis51,52. Border regions were enriched for origins of replication, as defined by short nascent strand sequencing (Supplementary Fig. 3G), in a TE-dependent manner (Supplementary Fig. 3H). LTR retrotransposons also commonly feature G-quadruplexes53, which are non-canonical DNA structures linked to both origins of replication (Supplementary Fig. 3I)54,55 and replication stress56,57,58,59. Border regions are enriched for G-quadruplexes present in TEs (Supplementary Fig. 3J–M). These findings suggest that replication origins and G-quadruplexes within LTRs are hotspots of replication stress, which leads to tandem duplications in tsk-4 mutants.
Like the origins, sites of replication termination are also stress-prone. In mammalian systems, termination sites align with topologically associated domain boundaries, which often show elevated mutation rates60,61,62. While Arabidopsis chromatin lacks these structures, transcription-linked chromatin boundaries are prevalent63,64,65. These boundaries align with replication timing edges (Supplementary Fig. 3N), and the binding of the negative transcription elongation factor BDR1 (Supplementary Fig. 3O). This indicates an organizational link between transcription, chromatin structure, and DNA replication. Consistent with the enrichment of breakends in heterochromatin, the border regions are also depleted of protein-coding genes (Supplementary Fig. 3P), have no association with R-loops (Supplementary Fig. 3Q)66, and are either depleted or unassociated with features of chromatin structure (Fig. 4M, Supplementary Fig. 3R–T).
Lastly, we observed that the border regions overlapped significantly with T-DNA insertion sites in Arabidopsis (Fig. 4N)67, which have been hypothesized to rely on the presence of double-stranded DNA breaks68. Overall, while tandem duplications occur across the genome in the absence of TSK, this analysis shows that late-replicating regions, and heterochromatic features, frequently co-occur with the tandem duplications.
The DNA damage response induces developmental defects in tsk-4 mutants
Arabidopsis tsk-4 mutants display many developmental phenotypes that vary stochastically between individual plants (Fig. 5Ai), all consistent with dysregulated meristematic tissue. These include nodes with double siliques (Fig. 5Aiii), thickened and split stems (Fig. 5Aiv), fasciation (Fig. 5Av), terminated meristems, increased branching, and small stature20,21,22,23,24. We hypothesized that there may be a link between these developmental phenotypes and the tandem duplications observed in tsk-4 mutants. To investigate this, we performed reciprocal crosses between fourth-generation tsk-4 mutants and Col-0. In the F2 generation, the tsk-4 mutation and tandem duplications are expected to segregate independently. As a representative tsk-4 phenotype, we quantified the frequency of nodes with double siliques (Fig. 5Aii-iii), and observed that this phenotype was mainly present in F2 plants homozygous for the tsk-4 mutation (Fig. 5B). This result suggests that the developmental phenotypes of tsk-4 plants are associated with the absence of TSK rather than the presence of tandem duplications within the genome.
A Variable phenotypes in tsk-4 plants: (i) representative images of tsk-4 sibling plants. Spiral phyllotaxis in (ii) Col-0 is disrupted in tsk-4 plants resulting in stochastic (iii) double silique nodes, (iv) fused stems and (v) fasciation. Scale bars are 5 cm. B Percentage of double silique nodes in F2 populations of Col-0 x tsk-4 crosses. Sample sizes are indicated below. Bars represent means with 95% confidence intervals. Lower case letters represent pairwise comparisons of a one-way ANOVA (F(6, 217) = 32.87, p = 4.83 × 10−28, ω² = 0.460) with Tukey’s Honestly Significant Difference post hoc testing. C Mean percentage of co-expressed SOG1-regulated gene groups (as defined by Bourbousse et al.71) that are either up- or down-regulated in tsk-4 plants relative to Col-0. Error bars represent standard errors from three independent plants. D Size of individual duplications in tsk-4 sog1-1 and tsk-4 wee1-2 (second-generation plants), and the single mutants grown alongside. Each point is an individual duplication. Region ≤100 kb shaded in dark gray, region ≤400 kb in light gray. PERMANOVA on pairwise 1-Wasserstein distances between plant-level log2 tandem duplication size distributions were performed for tsk-4 and tsk-4 sog1-1 (pseudo-F = 1.28, R2 = 0.176, adj. p = 0.607), and tsk-4 and tsk-4 wee1-2 (pseudo-F = 0.196, R2 = 0.047, adj. p = 0.667). E Percentage of double silique nodes in second-generation tsk-4 sog1-1 mutants. Sample sizes, bars and letters (F(13, 146) = 30.94, p = 1.65 × 10−35, ω²=0.71) as in (B). F Percentage of double silique nodes in second-generation tsk-4 atr-2 mutants. Sample sizes, bars and letters (F(9, 187) = 75.7, p = 1.43 × 10−57, ω²=0.77) as in (B). G Percentage of double silique nodes in second-generation tsk-4 wee1-2 mutants. Sample sizes, bars and letters (F(11, 204) = 71.94, p = 1.54 × 10−69, ω² = 0.76) as in (B). Source data are provided as a Source Data file.
The inability to repair damaged DNA in the absence of TSK is a source of cellular stress22,23. Thus, we considered whether the developmental defects observed in tsk-4 mutants result from the DNA damage response (DDR) pathway. This is a dose-dependent stress response progressively activating DNA repair, cell-cycle arrest and, if required, programmed cell death69. Cell-cycle disruption and stem cell death have previously been described in tsk-4 mutants22,23. The NAC transcription factor SOG1 is a central component of the plant DDR, particularly in regulating the G2/M cell-cycle transition69,70,71. At the transcriptional level, SOG1 regulates ~60% of the genes in the DDR pathway71. Analysis of our RNA-seq datasets indicated transcriptional modulation of SOG1-dependent genes in tsk-4 plants, which is consistent with DDR activation through SOG1. This includes upregulation of DNA repair genes (groups 1–3), and down regulation of cell-cycle associated genes (groups 9–11) (Fig. 5C).
To directly test the involvement of the DDR, we first introduced the sog1-1 mutant allele into the tsk-4 background and observed that the second-generation double mutants continued to generate tandem duplications (Fig. 5D). However, the developmental phenotypes associated with tsk-4 mutants were partially suppressed. Specifically, the fasciation, thickened and split stems, and frequent double silique nodes were eliminated (Fig. 5E, Supplementary Fig. 4A). However, the irregular phyllotaxy, reduced bolt height, and increased branching phenotypes were still in evidence within the populations (Supplementary Fig. 4A–C). Across all lines, 32% of plants resembled wild type, and a further 26% only showed reduced stature (Supplementary Fig. 4A–C). In the subsequent generation of tsk-4 sog1-1 mutants, the majority of plants exhibited growth reduction and increased branching (Supplementary Fig. 4D).
Since TSK functions at nascent chromatin following DNA replication12,13,14,15, we next interrogated the DDR components that activate as a result of replication-associated genome stress. ATR is a kinase that initiates signaling in response to persistent single-stranded DNA, which is commonly found at stalled replication forks69,72,73. This triggers CDK inhibition through the conserved checkpoint kinase WEE1, which rapidly delays S phase progression74,75,76. This mechanism is reinforced by ATR also activating transcription factors, such as SOG1, which further increase WEE1 expression77,78. Looking at tsk-4 double mutants with either wee1-2 or atr-2, we observed near-complete suppression of all developmental phenotypes present in tsk-4 single mutants (Fig. 5F, G, Supplementary Figs. 5, 6). In these two double mutant backgrounds, a modest reduction in growth was observed in the third generation (Supplementary Figs. 5E–G, 6E–G), and increased branching was present specifically in some lineages of tsk-4 wee1-2 plants (Supplementary Fig. 6E–G). However, this phenotype was present to a much lesser extent than in third-generation tsk-4 sog1-1 lines. In the third generation of tsk-4 atr-2 plants, 71% of plants across all lines resembled wild type, and a further 22% showed only a reduced stature; in tsk-4 wee1-2, these proportions were 40% and 40%, respectively. In addition, fasciation and split stems were not observed in tsk-4 plants lacking WEE1 or ATR. Finally, we analyzed tandem duplication levels in tsk-4 wee1-2 plants, and observed similar results as in tsk-4 sog1-1 mutants, in that inactivating WEE1 does not affect the generation of duplications (Fig. 5D). Together, these results demonstrate that the developmental phenotypes in tsk-4 plants are a consequence of the DDR pathway, and a parallel outcome of the DNA damage which leads to tandem duplications.
Discussion
This study demonstrates the importance of TSK-mediated resolution of replication stress in maintaining genome stability and normal plant development. In tsk-4 mutants, appreciable numbers of large tandem duplications are generated and distributed throughout the genome. They are dependent on TMEJ and arise throughout the life of the plant (5-15 heritable events per generation). As a result, the genomes of closely related individual plants lacking TSK activity can diverge rapidly, on a scale which approximates the divergence of Arabidopsis plants sampled from opposite sides of the world79. Modulating TSK activity may therefore be a useful tool in plant breeding to create targeted tandem duplications that increase gene expression in a non-GMO manner for the purpose of engineering new traits.
The creation of tandem duplications in tsk-4 mutants reveals the elusive source of DNA damage long reported in CAF-1 mutants of different organisms26,76,80,81,82. Since CAF-1, replication-dependent H3 variants, and TSK/TONSL are all highly conserved across multicellular eukaryotes, their roles in ameliorating replication stress are likely conserved in humans, and may be important for disease prevention. The presence of a large number of tandem duplications distributed throughout the genome is a characteristic of many cancers. Such cancers are categorized into three different groups based on the size of the duplications11. The Group 2 tandem duplicator phenotype, common in cancers driven by CCNE1 pathway activation or CDK12 loss5, is defined by oncogene-enriched duplications with a median size of 231 kb, which is closely matching the median size of duplications (199 kb) observed in tsk-4 mutants (Fig. 6A). Such cancers are associated with replication stress, and a dependence on DDR activation via the signaling kinase module ATR-WEE14,83. It is therefore possible that these cancers involve disruption of the TONSL pathway, thus interfering with proper resolution of replication stress.
A Distribution of tandem duplication sizes in tumors exhibiting a tandem duplicator phenotype5 plotted above the size distribution of tandem duplications observed in tsk-4 plants. Constructed from Supplementary Data 5. B Model of the role of the CAF-1-H3.1-TSK pathway in DNA damage and DDR. CAF-1 loads H3.1 and TSK onto newly replicated DNA, which, when DNA damage is encountered, facilitates DNA repair. This minimizes replication stress and DDR activation, which also depend on TOP1α relieving torsional stress. When TSK function is lost, replication stress increases as damage either goes unrepaired or becomes more severe. This in turn increases ATR-WEE1 DDR pathway activation to a level that, when damaged DNA is repaired, results in tandem duplications. Sufficiently high DDR activation can induce programmed cell death and give rise to developmental phenotypes. Panel created in BioRender. Thomson, G. (2025) https://BioRender.com/l4mxike.
In addition to producing tandem duplications, the replicative stress that occurs in the absence of TSK in plants leads to substantial DDR pathway activation. Specifically, ATR–WEE1 checkpoint signaling, and to a lesser extent SOG1 activity, disrupts the cell cycle and generates the stochastic developmental phenotypes observed in tsk-4 mutants. While some cells carrying tandem duplications continue to divide, others undergo arrest and cell death, which likely triggers irregular proliferation of adjacent cells (Fig. 6B)84,85. This connection between endogenous genome instability in individual cells and developmental disruption at the tissue level is reminiscent of the influence of SOG1 on root development in response to genotoxic stress86. Our results demonstrate that a similar process occurs in aerial organs. We conclude that TSK is a central node in an organism-wide axis of communication between genome stability, tissue-level patterning, and organism development.
Methods
Plant materials
Arabidopsis seeds were stratified at 4༠C for 2-3 days and germinated on ½ MS agar plates before being transferred to soil (Pro-Mix BX Mycorrhizae Growing Mix; Premier Horticulture Inc Quakertown, PA, USA). Plants were grown under cool-white fluorescent lights (~100 μmol m−2 s−1) in long-day conditions (16 h light/8 h dark). Multiple SALK T-DNA lines were utilized in this work, from the Col-0 background87; tsk-4 (aliases mgo3-4, bru1-4; At3g18730, SALK_034207)20, ku70-2 (At1g16970, SALK_123114)88, ku80-7 (At1g48050, SALK_112921)89, teb-5 (At4g32700, SALK_018851)37, atr-2 (At5g40820, SALK_032641)73 and wee1-2 (At1g02970, SALK_147968)74. The sog1-1 mutation is a GGGA > RAGA missense mutation at codon 155 in the Ler-0 background, which has since been introgressed in Col-070,90.
The mgo1CRISPR background was created using plasmids (described below) transformed using floral dip with Agrobacterium tumefaciens GV3101 (pMP90)91.
Plasmid construction
The binary vector containing CRISPR/Cas9 reagents used to generate mgo1CRISPR was created with the modular cloning (MoClo) toolkit92,93. Annealed oligonucleotides comprising the spacer sequence targeting MGO1 (At5g55300) (5‘-attgATCATCAGTTCCATCTTCTC-3‘, 5‘-aaacGAGAAGATGGAACTGATGAT-3‘) were cloned using the Esp3I restriction enzyme into the previously described Level1_blank_sgRNAv2_Sp_AtU6_(Position 3) vector94. It was then combined into the Level 2 backbone pAGM4723 with Level 1 vectors containing the following cassettes: nos::BAR::nosT, RPS5a::zCas9i::rbcS-E995 and a pFAST-R selection cassette.
Plant phenotyping
The number of nodes with double siliques was used as a representative phenotype of tsk plants. Such nodes were defined as two siliques being +/−2 mm of each other. The fraction of such nodes per plant was assessed in five-week-old plants by counting them on primary bolts excluding those within 1 cm of the top of the bolt. Stem length was measured in centimeters from rosette to tip.
The degree of “bushyness” in tsk-4 sog1-1 plants was a qualitative categorization, with representative photos given in Supplementary Fig. 4A.
Nucleic acid extraction and sequencing
Genomic DNA was extracted from leaf tissue (unless specifically stated otherwise) using the CTAB protocol96. DNA sequencing libraries were prepared at the Yale Center for Genome Analysis (YCGA). Genomic DNA was sonicated to an average fragment size of 350 bp using a Covaris E220 instrument (Covaris, Woburn, MA) and libraries were generated using the xGen Prism library prep kit for NGS (Integrated DNA Technologies, Coralville, IA). Paired-end 150 bp sequencing was performed on an Illumina NovaSeq 6000 using the S4 XP workflow (Illumina, San Diego, CA).
RNA was extracted from three-week-old leaf tissue using the RNeasy Plant Mini Kit (Qiagen, Hilden, Germany). RNA quality was verified using the Agilent 2100 Bioanalyzer Nano RNA Assay. Libraries were prepared at the YCGA with 1 μg of total RNA using Illumina’s TruSeq Stranded Total RNA with Ribo-Zero Plant (Illumina). The libraries were amplified with eight PCR cycles, validated using Agilent Bioanalyzer 2100 High sensitivity DNA assay and quantified using the KAPA Library Quantification Kit (Illumina® Platforms). Sequencing was done on an Illumina NovaSeq 6000 using the S4 XP workflow.
WGS analysis
Analysis of all sequencing datasets utilized the TAIR10 genome97,98. DNA sequencing reads were processed by Fastp (v0.23.2; -q 20 -l 20)99 and aligned using BWA-MEM (v0.7.17)100. Duplicate reads were removed from alignments using Samtools (v1.16)101. Alignment statistics of genomic datasets produced in this study are listed in Supplementary Data 6.
Local read realignment was then performed using Abra2 (v2.23)102. Analysis of single-nucleotide variants and small insertions/deletions was carried out using the Strelka2 (v2.9.10) germline pipeline103. CNVs were defined using sequencing depth identified using GATK (v4.2.0.0/v4.4.0.0)104 to segment the genome. This followed tutorials #11682 and #11683, omitting allele specific steps, a strategy which agreed with manual inspections. This pipeline also used Picard (v2.25.6)105 to add read group (‘RG’) tags. Genome segments with coverage above Col-0 were merged (+/−100 bp) into contiguous amplified regions, which we term CNVs. The coverage of the segment with the highest level of coverage assigned as the CNV coverage. This was achieved using R (v4.1.2)106, Bedtools (v2.30.0)107 and GNU Awk (v5.1.0)108. CNVs less than 1.5x Col-0 coverage were removed to avoid false positives. This was augmented by manual inspection in the analysis of tissue-specific CNVs.
Identification of chimeric reads and characterization of junctions was achieved by filtering SA-tagged reads using Samtools, and using Pysam (v0.19.0)101,109 to analyze alignments. In the experiments which tracked CNVs over generations, or between tissues, identical CNVs were defined as those starting and ending +/−200 bp of each other, These were identified using the R fuzzyjoin package110.
The Deeptools (v3.5.1)111 and pyGenomeTracks (v3.8)112 programs were used to visualize CNVs.
Extraction of ultra-high molecular weight (uHMW) DNA and optical mapping
Optical mapping was performed by The French Plant Genomic Resources Center (CNRGV), INRAE. uHMW DNA was purified from 0.8 g of young frozen pooled leaves (6 individual plants, progeny of tsk_4_Gen3_Fam2_plant2) combining the Bionano Prep Plant Tissue DNA Isolation Base Protocol (30068; Bionano Genomics, San Diego, CA) and a CTAB DNA purification protocol113. Briefly, frozen leaves were ground into 2 mm pieces and then disrupted with a rotor stator in homogenization buffer containing spermine, spermidine and 2- mercaptoethanol. Nuclei were washed in homogenization buffer, TC buffer (50 mM Tris-HCl pH 7.5, 75 mM NaCl, 6 mM MgCl2, 0.1 mM CaCl2) and finally resuspended in 2 ml of TC. The nuclei suspension was mixed with an equal volume of 2X CTAB buffer (1.4 M NaCl, 100 mM Tris-HCl pH 8.0, 2% CTAB, 20 mM EDTA, 0.5% (w/v) Na2S2O5, 2% (v/v) 2-mercaptoethanol) and incubated at 50 °C for 40 min under gentle homogenization. After incubation, the nuclei were isolated with choloform:isoamylalcohol (24:1) and precipitated with 0.7 volume of isopropanol. Following 70% ethanol washes, the DNA medusa was finally resuspended in TE buffer. After complete resuspension, the DNA was quantified using the Qubit dsDNA BR Assay (Invitrogen, Thermo Fisher Scientific, Waltham, MA). The presence of mega base size DNA was visualized by pulsed field gel electrophoresis (PFGE).
Labeling and staining of the uHMW DNA were performed according to the Bionano Prep Direct Label and Stain (DLS) protocol (CG-30553-1; Bionano Genomics). Briefly, labeling was performed by incubating 750 ng genomic DNA with DLE-1 enzyme (recognizing the site CTTAAG) in the presence of DL-Green dye. Following proteinase K digestion and DL-Green cleanup by membrane adsorption, the DNA backbone was stained by mixing the labeled DNA with DNA Stain solution and incubated overnight at room temperature. The DLS DNA concentration was assessed with the Qubit dsDNA HS Assay (Invitrogen).
Labeled and stained DNA was loaded on the Saphyr chip. Loading of the chip and running of the Bionano Genomics Saphyr System were performed according to the Saphyr System User Guide (30247; Bionano Genomics). Data processing was performed using the Bionano Access and Solve v.3.8 software (https://bionano.com/software-and-data-analysis-support/).
Optical mapping analysis
A pseudo-reference genome was created by inserting tandem duplications into the TAIR10 genome, based on 11 tandem duplications likely to be inherited from the tsk_4_Gen3_Fam2_plant2. A corresponding BED file documenting the coordinates of the tandem duplications was also created alongside. A reference CMAP file for this pseudo-reference genome was constructed using the OMTools (v.1.4a)114 FastaToOM (--enzymestring CTTAAG) command. The OMTools DataTools (--minsize 100000 --minsig 9) command was used to filter the raw mapping data, which was then mapped to the pseudo-reference genome using OMTools OMBlastMapper (--filtermode 1 --alignmentjoinmode 1 --minscore 0.3 --closeref 500000 --closefrag 50000). Molecules with multiple alignment segments that were non-overlapping and colinear on both the reference and the molecule were grouped into “chained” molecules and analyzed separately.
Molecules with alignments intersecting one of the 11 inherited tandem duplications were classified into three groups; 1) those that span a breakpoint junction, 2) those that span a breakpoint junction and one breakend, or 3) those that span a breakpoint junction and both breakends. All molecules were required to have ≥2 DLE sites either side of a junction.
Association of tsk CNVs with annotated genomic features
Simulation testing was used to assess the overlap between CNVs observed in tsk plants. Unique CNVs >10 kb were compiled and bedtools intersect (-wao) was used with individual features. Simulated CNV sets were implemented using bedtools shuffle to randomize the position of the CNVs before again using bedtools intersect (-wao) to ascertain the level of overlap controlling for CNV size. This was run 10,000 times. The resulting output was then summarized using R. The same strategy was implemented to look at the overlap between CNVs and previously annotated features, or at the overlap between features themselves. When looking at CNV breakends, an R script was used to create the BED file of breakends for each new set of randomized CNVs. When summarizing previously annotated features, counts were used for discrete features, and number of bp covered was used for continuous features.
We report the probability (Pr) that a randomized mean level of overlap is less than what is observed. Values > 0.975 are considered enriched, while those less than 0.025 are considered depleted. GC overlap was calculated by intersecting a bedgraph file of GC content at 10 bp resolution. Short tandem repeats were annotated using Tandem Repeats Finder115. The position of other features was carried out using published analyses, with the exception of S phase genes116, replication timings44 and T-DNA insertions67 for which raw data were used to replicate the published analysis. We termed the 6548 genes with normalized counts ≥1 in S phase shoot scRNA-Seq (replicates merged) to be S phase genes.
Reanalysis of ChIP-seq data assaying chromatin modifications45,46,47 was also carried out. Sequencing reads were processed by Fastp (-q 20 -l 20) and aligned using BWA-MEM. Duplicate reads were removed from alignments using Samtools. MACS2 (v2.2.9)117 was used to call peaks with the --broad parameter used for H3K27me1, H3K9me2, H3K27me3, H2AK121ub, and H3K36me3. All peaks were filtered out for q < 0.05 and, if multiple datasets were available, those present in ≥50% of the datasets.
Motif enrichment utilized the XSTREME tool118. A list of previously published datasets utilized44,45,46,47,50,52,63,66,67,97,116,119,120,121,122,123 are listed in Supplementary Data 7.
RNA-seq analysis
RNA sequencing reads were processed using Fastp (-q 20 -l 20). Transcript abundances were quantified with Salmon (v1.4.0)124 using the Araport11 genome annotation97. Features were filtered to have counts of at least 10. Global trends in transcriptional perturbation were calculated by taking the ratio of abundances, normalized to transcripts per million, in tsk-4 plants relative to those of Col-0 sequenced alongside.
Statistical analysis
Testing the association of tandem duplications with genomic features made use of simulation testing outlined above. For comparisons of plant phenotypes, the differences in means were tested using one-way ANOVA followed by Tukey’s Honestly Significant Difference post hoc tests.
Wasserstein distances were computed between empirical distributions of log2 tandem duplication size within each plant, and genotype effects were assessed by permutation (upper-tail) of the PERMANOVA pseudo-F statistic with labels at the plant level. The p-values were adjusted across contrasts using the Bonferroni–Holm method.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Source data is provided with this paper sufficient to recreate all presented results. Sequencing data (DNA-seq and RNA-seq datasets) generated for this study are available from the NCBI SRA under BioProject ID PRJNA1332561. Individual accession numbers are listed in Supplementary Data 6. Optical mapping data has been deposited in the EMBL-EBI ENA with the accession number PRJEB106903. Previously published data plotted in Fig. 2D has the SRA identifiers ERR3852258 and BioProject ID PRJEB3583226, SRR5818181 and BioProject ID PRJNA39389233 and SRR24877660 and BioProject ID PRJNA97893832. For the purposes of testing the association of tandem duplications with genomic features in Fig. 4 and Supplementary Figs. 2, 3 several datasets were reanalyzed. These are Repli-Seq data with the BioProject ID PRJNA33054744, shoot scRNA-Seq data with the GSA ID: CRX125603 and CRX125604, BioProject ID PRJCA003094116 and T-DNA integration sites with the BioProject ID PRJNA39361367. Previously published ChIP-seq data for assaying chromatin modifications plotted in Supplementary Fig. 2G–M were sourced for H3K27me1, H3K9me2, H3K4me3, H3K36me3 with the BioProject ID PRJNA96476345, H3K27me3, H2AK121ub with BioProject ID PRJNA35187047 and H3K9Ac with BioProject ID PRJNA35312146. Source data are provided with this paper.
References
Macintyre, G. et al. Copy number signatures and mutational processes in ovarian carcinoma. Nat. Genet. 50, 1262–1270 (2018).
Li, Y. et al. Patterns of somatic structural variation in human cancer genomes. Nature 578, 112–121 (2020).
Steele, C. D. et al. Signatures of copy number alterations in human cancer. Nature 606, 984–991 (2022).
Yang, Y. et al. Transcription and DNA replication collisions lead to large tandem duplications and expose targetable therapeutic vulnerabilities in cancer. Nat. Cancer 5, 1885–1901 (2024).
Menghi, F. et al. The tandem duplicator phenotype is a prevalent genome-wide cancer configuration driven by distinct gene mutations. Cancer Cell 34, 197–210.e5 (2018).
Raynaud, F., Mina, M., Tavernari, D. & Ciriello, G. Pan-cancer inference of intra-tumor heterogeneity reveals associations with different forms of genomic instability. PLoS Genet. 14, e1007669 (2018).
Baslan, T. et al. Novel insights into breast cancer copy number genetic heterogeneity revealed by single-cell genome sequencing. eLife 9, e51480 (2020).
van Dijk, E. et al. Chromosomal copy number heterogeneity predicts survival rates across cancers. Nat. Commun. 12, 3188 (2021).
Smith, J. C. & Sheltzer, J. M. Genome-wide identification and analysis of prognostic features in human cancers. Cell Rep. 38, 110569 (2022).
Menghi, F. et al. The tandem duplicator phenotype as a distinct genomic configuration in cancer. Proc. Natl. Acad. Sci. USA 113, E2373–E2382 (2016).
Scully, R., Glodzik, D., Menghi, F., Liu, E. T. & Zhang, C.-Z. Mechanisms of tandem duplication in the cancer genome. DNA Repair 145, 103802 (2025).
Duro, E. et al. Identification of the MMS22L-TONSL complex that promotes homologous recombination. Mol. Cell 40, 632–644 (2010).
O’Donnell, L. et al. The MMS22L-TONSL complex mediates recovery from replication stress and homologous recombination. Mol. Cell 40, 619–631 (2010).
Saredi, G. et al. H4K20me0 marks post-replicative chromatin and recruits the TONSL–MMS22L DNA repair complex. Nature 534, 714–718 (2016).
Davarinejad, H. et al. The histone H3.1 variant regulates TONSOKU-mediated DNA repair during replication. Science 375, 1281–1286 (2022).
Yuan, W. et al. H3.1K27M-induced misregulation of the TONSOKU-H3.1 pathway causes genomic instability. Nat. Commun. 16, 3547 (2025).
Piwko, W. et al. The MMS22L-TONSL heterodimer directly promotes RAD51-dependent recombination upon replication stress. EMBO J. 35, 2584–2601 (2016).
Burrage, L. C. et al. Bi-allelic Variants in TONSL Cause SPONASTRIME Dysplasia and a Spectrum of Skeletal Dysplasia Phenotypes. Am. J. Hum. Genet. 104, 422–438 (2019).
Chang, H. R. et al. Hypomorphic mutations in TONSL cause SPONASTRIME dysplasia. Am. J. Hum. Genet. 104, 439–453 (2019).
Guyomarc’h, S., Vernoux, T., Traas, J., Zhou, D.-X. & Delarue, M. MGOUN3, an Arabidopsis gene with TetratricoPeptide-Repeat-related motifs, regulates meristem cellular organization. J. Exp. Bot. 55, 673–684 (2004).
Takeda, S. et al. BRU1, a novel link between responses to DNA damage and epigenetic gene silencing in Arabidopsis. Genes Dev. 18, 782–793 (2004).
Suzuki, T. et al. A novel Arabidopsis gene TONSOKU is required for proper cell arrangement in root and shoot apical meristems. Plant J. 38, 673–684 (2004).
Suzuki, T. et al. TONSOKU is expressed in S phase of the cell cycle and its defect delays cell cycle progression in Arabidopsis. Plant Cell Physiol. 46, 736–742 (2005).
Ohno, Y. et al. Ectopic gene expression and organogenesis in Arabidopsis mutants missing BRU1 required for genome maintenance. Genetics 189, 83–95 (2011).
Loehlin, D. W. & Carroll, S. B. Expression of tandem gene duplicates is often greater than twofold. Proc. Natl. Acad. Sci. USA 113, 5988–5992 (2016).
Picart-Picolo, A. et al. Large tandem duplications affect gene expression, 3D organization, and plant-pathogen response. Genome Res. 30, 1583–1592 (2020).
Yang, Q.-L., Xie, Y., Qiao, K., Lim, J. Y. S. & Wu, S. Modern biology of extrachromosomal DNA: A decade-long voyage of discovery. Cell Res. 35, 11–22 (2025).
Laufs, P., Dockx, J., Kronenberger, J. & Traas, J. MGOUN1 and MGOUN2: two genes required for primordium initiation at the shoot apical and floral meristems in Arabidopsis thaliana. Development 125, 1253–1260 (1998).
Graf, P. et al. MGOUN1 encodes an Arabidopsis type IB DNA topoisomerase required in stem cell regulation and to maintain developmentally regulated gene silencing. Plant Cell 22, 716–728 (2010).
Tagami, H., Ray-Gallet, D., Almouzni, G. & Nakatani, Y. Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 116, 51–61 (2004).
Otero, S., Desvoyes, B., Peiró, R. & Gutierrez, C. Histone H3 Dynamics Reveal Domains with Distinct Proliferation Potential in the Arabidopsis Root. Plant Cell 28, 1361–1371 (2016).
Wang, L., Xue, M., Zhang, H., Ma, L. & Jiang, D. TONSOKU is required for the maintenance of repressive chromatin modifications in Arabidopsis. Cell Rep. 42, 112738 (2023).
Jiang, D. & Berger, F. DNA replication-coupled histone modification maintains Polycomb gene silencing in plants. Science 357, 1146–1149 (2017).
Bundock, P., van Attikum, H. & Hooykaas, P. Increased telomere length and hypersensitivity to DNA damaging agents in an Arabidopsis KU70 mutant. Nucleic Acids Res. 30, 3395–3400 (2002).
West, C. E. et al. Disruption of the Arabidopsis AtKu80 gene demonstrates an essential role for AtKu80 protein in efficient repair of DNA double-strand breaks in vivo. Plant J. 31, 517–528 (2002).
Tamura, K., Adachi, Y., Chiba, K., Oguchi, K. & Takahashi, H. Identification of Ku70 and Ku80 homologues in Arabidopsis thaliana: evidence for a role in the repair of DNA double-strand breaks. Plant J. 29, 771–781 (2002).
Inagaki, S. et al. Arabidopsis TEBICHI, with helicase and DNA polymerase domains, is required for regulated cell division and differentiation in meristems. Plant Cell 18, 879–892 (2006).
Nisa, M. et al. The plant DNA polymerase theta is essential for the repair of replication-associated DNA damage. Plant J. 106, 1197–1207 (2021).
Schimmel, J., van Schendel, R., den Dunnen, J. T. & Tijsterman, M. Templated Insertions: A Smoking Gun for Polymerase Theta-Mediated End Joining. Trends Genet. 35, 632–644 (2019).
van Schendel, R., Romeijn, R., Kralemann, L. E. M., Hooykaas, P. J. J. & Tijsterman, M. TONSL supresses tandem duplications across kingdoms through chromatin-guided repair. Nat. Commun. (2026, In press).
Shastri, N. et al. Genome-wide Identification of Structure-Forming Repeats as Principal Sites of Fork Collapse upon ATR Inhibition. Mol. Cell 72, 222–238.e11 (2018).
Gadgil, R. Y. et al. Replication stress at microsatellites causes DNA double-strand breaks and break-induced replication. J. Biol. Chem. 295, 15378–15397 (2020).
Tubbs, A. et al. Dual roles of poly(da:dt) tracts in replication initiation and fork collapse. Cell 174, 1127–1142.e19 (2018).
Concia, L. et al. Genome-Wide Analysis of the Arabidopsis Replication Timing Program. Plant Physiol. 176, 2166–2185 (2018).
Jamge, B. et al. Histone variants shape chromatin states in Arabidopsis. eLife 12, RP87714 (2023).
Kim, Y. J. et al. POWERDRESS and HDA9 interact and promote histone H3 deacetylation at specific genomic sites in Arabidopsis. Proc. Natl. Acad. Sci. USA 113, 14858–14863 (2016).
Zhou, Y., Romero-Campero, F. J., Gómez-Zambrano, Á, Turck, F. & Calonje, M. H2A monoubiquitination in Arabidopsis thaliana is generally independent of LHP1 and PRC2 activity. Genome Biol. 18, 69 (2017).
Arabidopsis Genome Initiative Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000).
Pavani, R. et al. Structure and repair of replication-coupled DNA breaks. Science 385, eado3867 (2024).
Liu, D.-X. et al. Calling large indels in 1047 Arabidopsis with IndelEnsembler. Nucleic Acids Res. 49, 10879–10894 (2021).
Vergara, Z. et al. Retrotransposons are specified as DNA replication origins in the gene-poor regions of Arabidopsis heterochromatin. Nucleic Acids Res. 45, 8358–8368 (2017).
Sequeira-Mendes, J. et al. Differences in firing efficiency, chromatin, and transcription underlie the developmental plasticity of the Arabidopsis DNA replication origins. Genome Res. 29, 784–797 (2019).
Lexa, M. et al. Quadruplex-forming sequences occupy discrete regions inside plant LTR retrotransposons. Nucleic Acids Res. 42, 968–978 (2014).
Prorok, P. et al. Involvement of G-quadruplex regions in mammalian replication origin activity. Nat. Commun. 10, 3274 (2019).
Bastolla, U. et al. The topography of DNA replication origins in Eukarya: GGN clusters, landmark nucleosomes, CDC6 and G4 structures. BioRxiv, https://doi.org/10.1101/2024.01.04.574144 (2024).
Lopes, J. et al. G-quadruplex-induced instability during leading-strand replication. EMBO J. 30, 4033–4046 (2011).
Lemmens, B., van Schendel, R. & Tijsterman, M. Mutagenic consequences of a single G-quadruplex demonstrate mitotic inheritance of DNA replication fork barriers. Nat. Commun. 6, 8909 (2015).
Koole, W. et al. A Polymerase Theta-dependent repair pathway suppresses extensive genomic instability at endogenous G4 DNA sites. Nat. Commun. 5, 3216 (2014).
Williams, S. L. et al. Replication-induced DNA secondary structures drive fork uncoupling and breakage. EMBO J. 42, e114334 (2023).
Sarni, D. et al. 3D genome organization contributes to genome instability at fragile sites. Nat. Commun. 11, 3613 (2020).
Sidiropoulos, N. et al. Somatic structural variant formation is guided by and influences genome architecture. Genome Res. 32, 643–655 (2022).
Faseela, E. E., Notani, D. & Sabarinathan, R. Replication stress underlies genomic instability at CTCF/cohesin-binding sites in cancer. BioRxiv https://doi.org/10.1101/2023.10.24.563697 (2023).
Sun, L. et al. Mapping nucleosome-resolution chromatin organization and enhancer-promoter loops in plants using Micro-C-XL. Nat. Commun. 15, 35 (2024).
Lee, H. & Seo, P. J. Accessible gene borders establish a core structural unit for chromatin architecture in Arabidopsis. Nucleic Acids Res. 51, 10261–10277 (2023).
Wang, C. et al. Genome-wide analysis of local chromatin packing in Arabidopsis thaliana. Genome Res. 25, 246–256 (2015).
Xu, W. et al. The R-Loop Atlas of Arabidopsis Development and Responses to Environmental Stimuli. Plant Cell 32, 888–903 (2020).
Shilo, S. et al. T-DNA-genome junctions form early after infection and are influenced by the chromatin state of the host genome. PLoS Genet. 13, e1006875 (2017).
Thomson, G., Dickinson, L. & Jacob, Y. Genomic consequences associated with Agrobacterium-mediated transformation of plants. Plant J. 117, 342–363 (2024).
Pedroza-Garcia, J. A., Xiang, Y. & De Veylder, L. Cell cycle checkpoint control in response to DNA damage by environmental stresses. Plant J. 109, 490–507 (2022).
Yoshiyama, K., Conklin, P. A., Huefner, N. D. & Britt, A. B. Suppressor of gamma response 1 (SOG1) encodes a putative transcription factor governing multiple responses to DNA damage. Proc. Natl. Acad. Sci. USA 106, 12843–12848 (2009).
Bourbousse, C., Vegesna, N. & Law, J. A. SOG1 activator and MYB3R repressors regulate a complex DNA damage network in Arabidopsis. Proc. Natl. Acad. Sci. USA 115, E12453–E12462 (2018).
Saldivar, J. C., Cortez, D. & Cimprich, K. A. The essential kinase ATR: ensuring faithful duplication of a challenging genome. Nat. Rev. Mol. Cell Biol. 18, 622–636 (2017).
Culligan, K., Tissier, A. & Britt, A. ATR regulates a G2-phase cell-cycle checkpoint in Arabidopsis thaliana. Plant Cell 16, 1091–1104 (2004).
De Schutter, K. et al. Arabidopsis WEE1 kinase controls cell cycle arrest in response to activation of the DNA integrity checkpoint. Plant Cell 19, 211–225 (2007).
Cools, T. et al. The Arabidopsis thaliana checkpoint kinase WEE1 protects against premature vascular differentiation during replication stress. Plant Cell 23, 1435–1448 (2011).
Eekhout, T. et al. G2/M-checkpoint activation in fasciata1 rescues an aberrant S-phase checkpoint but causes genome instability. Plant Physiol. 186, 1893–1907 (2021).
Ogita, N. et al. Identifying the target genes of SUPPRESSOR OF GAMMA RESPONSE 1, a master transcription factor controlling DNA damage response in Arabidopsis. Plant J. 94, 439–453 (2018).
Nisa, M. et al. Distinctive and complementary roles of E2F transcription factors during plant replication stress responses. Mol. Plant 16, 1269–1282 (2023).
Jiao, W.-B. & Schneeberger, K. Chromosome-level assemblies of multiple Arabidopsis genomes reveal hotspots of rearrangements with altered evolutionary dynamics. Nat. Commun. 11, 989 (2020).
Hisanaga, T. et al. The ATM-dependent DNA damage response acts as an upstream trigger for compensation in the fas1 mutation during Arabidopsis leaf development. Plant Physiol. 162, 831–841 (2013).
Muchová, V. et al. Homology-dependent repair is involved in 45S rDNA loss in plant CAF-1 mutants. Plant J. 81, 198–209 (2015).
Endo, M. et al. Increased frequency of homologous recombination and T-DNA integration in Arabidopsis CAF-1 mutants. EMBO J. 25, 5579–5590 (2006).
Xu, H. et al. CCNE1 copy number is a biomarker for response to combination WEE1-ATR inhibition in ovarian and endometrial cancer models. Cell Rep. Med. 2, 100394 (2021).
Cruz-Ramírez, A. et al. A SCARECROW-RETINOBLASTOMA protein network controls protective quiescence in the Arabidopsis root stem cell organizer. PLoS Biol. 11, e1001724 (2013).
Takahashi, N. et al. DNA double-strand breaks enhance brassinosteroid signaling to activate quiescent center cell division in Arabidopsis. J. Exp. Bot. 75, 1364–1375 (2024).
Johnson, R. A. et al. SUPPRESSOR OF GAMMA RESPONSE1 links DNA damage response to organ regeneration. Plant Physiol. 176, 1665–1675 (2018).
Alonso, J. M. et al. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657 (2003).
Kannan, K., Nelson, A. D. L. & Shippen, D. E. Dyskerin is a component of the Arabidopsis telomerase RNP required for telomere maintenance. Mol. Cell. Biol. 28, 2332–2341 (2008).
Dumont, M., Massot, S., Doutriaux, M.-P. & Gratias, A. Characterization of Brca2-deficient plants excludes the role of NHEJ and SSA in the meiotic chromosomal defect phenotype. PLoS ONE 6, e26696 (2011).
Preuss, S. B. & Britt, A. B. A DNA-damage-induced cell cycle checkpoint in Arabidopsis. Genetics 164, 323–334 (2003).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Weber, E., Engler, C., Gruetzner, R., Werner, S. & Marillonnet, S. A modular cloning system for standardized assembly of multigene constructs. PLoS ONE 6, e16765 (2011).
Engler, C. et al. A golden gate modular cloning toolbox for plants. ACS Synth. Biol. 3, 839–843 (2014).
Sagawa, C. H. D. et al. An efficient multiplex approach to CRISPR/Cas9 gene editing in citrus. Plant Methods 20, 148 (2024).
Grützner, R. et al. High-efficiency genome editing in plants mediated by a Cas9 gene containing multiple introns. Plant Commun. 2, 100135 (2021).
Weigel, D. & Glazebrook, J. Arabidopsis: A Laboratory Manual (CSHL Press, 2002).
Cheng, C.-Y. et al. Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant J. 89, 789–804 (2017).
Reiser, L. et al. The Arabidopsis Information Resource in 2024. Genetics 227, iyae027 (2024).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv, https://doi.org/10.48550/arxiv.1303.3997 (2013).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Mose, L. E., Perou, C. M. & Parker, J. S. Improved indel detection in DNA and RNA via realignment with ABRA2. Bioinformatics 35, 2966–2973 (2019).
Kim, S. et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat. Methods 15, 591–594 (2018).
Auwera, G. van der & O’Connor, B. D. Genomics in the cloud: Using Docker, GATK, and WDL in Terra (O’Reilly Media, 2020).
Broad Institute. Picard Toolkit. https://github.com/broadinstitute/picard/ (2019).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2021).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Free Software Foundation. GNU Awk (Free Software Foundation, 1989).
Heger, A., Jacobs, K. & Contributors. Pysam: htslib interface for Python (Github, 2022).
Robinson, D. fuzzyjoin: Join Tables Together on Inexact Matching (CRAN, 2020).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Lopez-Delisle, L. et al. pyGenomeTracks: reproducible plots for multivariate genomic datasets. Bioinformatics 37, 422–423 (2021).
Vondrak, T. et al. Characterization of repeat arrays in ultra-long nanopore reads reveals frequent origin of satellite DNA from retrotransposon-derived tandem repeats. Plant J. 101, 484–500 (2020).
Leung, A. K.-Y., Jin, N., Yip, K. Y. & Chan, T.-F. OMTools: a software package for visualizing and processing optical mapping data. Bioinformatics 33, 2933–2935 (2017).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Zhang, T.-Q., Chen, Y. & Wang, J.-W. A single-cell analysis of the Arabidopsis vegetative shoot apex. Dev. Cell 56, 1056–1074.e8 (2021).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Grant, C. E. & Bailey, T. L. XSTREME: comprehensive motif analysis of biological sequence datasets. BioRxiv, https://doi.org/10.1101/2021.09.02.458722 (2021).
Marsico, G. et al. Whole genome experimental maps of DNA G-quadruplexes in multiple species. Nucleic Acids Res. 47, 3862–3874 (2019).
Yu, X., Martin, P. G. P. & Michaels, S. D. BORDER proteins protect expression of neighboring genes by promoting 3’ Pol II pausing in plants. Nat. Commun. 10, 4359 (2019).
Göbel, A.-M. et al. Mutations of PDS5 genes enhance TAD-like domain formation Arabidopsis thaliana. Nat. Commun. 15, 9308 (2024).
Panda, K. & Slotkin, R. K. Long-read cDNA sequencing enables a “gene-like” transcript annotation of transposable elements. Plant Cell 32, 2687–2698 (2020).
The 1001 Genomes Consortium 1,135 Genomes reveal the global pattern of polymorphism in Arabidopsis thaliana. Cell 166, 481–491 (2016).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Acknowledgements
We would like to thank Anne Britt and Julie Law for sending us sog1-1 seeds. We also want to acknowledge Christopher Bolick and Nathan Guzzo from Yale University for help with plant maintenance. We also thank members of the Jacob laboratory for comments and discussion. This project was made possible by a grant (R35GM128661) from the National Institutes of Health (NIH), a Yale Cancer Center Pilot Award to Y.J. (funded by NIH P30CA016359), and an equipment support grant to the Yale Center for Genomic Analysis (NIH 1S10OD030363-01A1).
Author information
Authors and Affiliations
Contributions
Y.J. obtained funding and supervised the study. Y.-C.H., G.T., C.L. and Y.J. grew plants and extracted DNA. G.T. and A.P. performed bioinformatic analysis. H.-S.L. constructed and generated the MGO1 CRISPR lines. G.T. and C.L. created the figures. G.T., Y.J. and C.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Frederic Pontvianne, Shunping Yan, who co-reviewed with Yan Zhao, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Thomson, G., Poulet, A., Huang, YC. et al. TONSOKU prevents the formation of large tandem duplications and restrains ATR–WEE1 checkpoint activation. Nat Commun 17, 2874 (2026). https://doi.org/10.1038/s41467-026-70906-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-026-70906-1