Abstract
N6-Methyladenosine (m6A) reader proteins, which recognize m6A to regulate RNA metabolism, are important for plant adaptation to the changing environment. It remains unknown how the activities of plant m6A reader proteins are regulated in plant responses to stress. Here we show that the rice m6A reader protein EVOLUTIONARILY CONSERVED C-TERMINAL REGION 3 (OsECT3), required for rice tolerance to cold, is post-translationally modified by lysine acetylation, which reduces its m6A-binding activity. Under cold conditions, OsECT3 acetylation is reduced by cold-induced histone deacetylase HDA705 and low ACLA2-sourced acetyl-CoA levels, resulting in an increase in OsECT3 m6A-binding activity, the accumulation of cold-response-related mRNAs and improved tolerance of rice to cold stress. These results unravel a regulatory mechanism of an m6A reader protein to dynamically control m6A RNA levels under stress and suggest a link between lysine acetylation, metabolism and m6A pathways.
This is a preview of subscription content, access via your institution
Access options








Similar content being viewed by others
Data availability
The raw sequence data from m6A-seq and CLIP reported in this paper have been deposited in the Genome Sequence Archive at the National Genomics Data Center, Beijing Institute of Genomics, Chinese Academy of Sciences, under accession number PRJCA033583, which is publicly accessible at https://bigd.big.ac.cn/gsa. Source data are provided with this paper. All remaining data are available in the main text or in Supplementary Information.
References
Boccaletto, P. et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 46, D303–D307 (2018).
Shi, H., Wei, J. & He, C. Where, when, and how: context-dependent functions of RNA methylation writers, readers, and erasers. Mol. Cell 74, 640–650 (2019).
Xiao, W. et al. Nuclear m6A reader YTHDC1 regulates mRNA splicing. Mol. Cell 61, 507–519 (2016).
Roundtree, I. A. et al. YTHDC1 mediates nuclear export of N6-methyladenosine methylated mRNAs. eLife 6, e31311 (2017).
Shi, H. et al. YTHDF3 facilitates translation and decay of N6-methyladenosine-modified RNA. Cell Res. 27, 315–328 (2017).
Shima, H. et al. S-Adenosylmethionine synthesis is regulated by selective N6-adenosine methylation and mRNA degradation involving METTL16 and YTHDC1. Cell Rep. 21, 3354–3363 (2017).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Zaccara, S., Ries, R. J. & Jaffrey, S. R. Reading, writing and erasing mRNA methylation. Nat. Rev. Mol. Cell Biol. 20, 608–624 (2019).
Xu, C. et al. Structural basis for the discriminative recognition of N6-methyladenosine RNA by the human YT521-B homology domain family of proteins. J. Biol. Chem. 290, 24902–24913 (2015).
Arribas-Hernández, L. et al. An m6A-YTH module controls developmental timing and morphogenesis in Arabidopsis. Plant Cell 30, 952–967 (2018).
Scutenaire, J. et al. The YTH domain protein ECT2 is an m6A reader required for normal trichome branching in Arabidopsis. Plant Cell 30, 986–1005 (2018).
Wei, L. H. et al. The m6A reader ECT2 controls trichome morphology by affecting mRNA stability in Arabidopsis. Plant Cell 30, 968–985 (2018).
Hou, Y. et al. CPSF30-L-mediated recognition of mRNA m6A modification controls alternative polyadenylation of nitrate signaling-related gene transcripts in Arabidopsis. Mol. Plant 14, 688–699 (2021).
Song, P. et al. Arabidopsis N6-methyladenosine reader CPSF30-L recognizes FUE signals to control polyadenylation site choice in liquid-like nuclear bodies. Mol. Plant 14, 571–587 (2021).
Song, P. et al. m6A readers ECT2/ECT3/ECT4 enhance mRNA stability through direct recruitment of the poly(A) binding proteins in Arabidopsis. Genome Biol. 24, 103 (2023).
Guo, T. et al. The m6A reader MhYTP2 regulates MdMLO19 mRNA stability and antioxidant genes translation efficiency conferring powdery mildew resistance in apple. Plant Biotechnol. J. 20, 511–525 (2022).
Luo, W. et al. The m6A reader SiYTH1 enhances drought tolerance by affecting the mRNA stability of genes related to stomatal closure and ROS scavenging in Setaria italica. J. Integr. Plant Biol. 65, 2569–2586 (2023).
Wang, S. et al. m6A mRNA modification promotes chilling tolerance and modulates gene translation efficiency in Arabidopsis. Plant Physiol. 192, 1466–1482 (2023).
Vicente, A. M. et al. The plant cytosolic m6A RNA methylome stabilizes photosynthesis in the cold. Plant Commun. 4, 100634 (2023).
Yang, D. et al. RNA N6-methyladenosine responds to low-temperature stress in tomato anthers. Front. Plant Sci. 12, 687826 (2021).
Lesk, C., Rowhani, P. & Ramankutty, N. Influence of extreme weather disasters on global crop production. Nature 529, 84–87 (2016).
Ding, Y. & Yang, S. Surviving and thriving: how plants perceive and respond to temperature stress. Dev. Cell 57, 947–958 (2022).
Guo, X., Liu, D. & Chong, K. Cold signaling in plants: insights into mechanisms and regulation. J. Integr. Plant Biol. 60, 745–756 (2018).
Ma, Y. et al. COLD1 confers chilling tolerance in rice. Cell 160, 1209–1221 (2015).
Ke, Y. G. et al. Characterization of OsDREB6 responsive to osmotic and cold stresses in rice. J. Plant Biol. 57, 150–161 (2014).
Ma, Q. et al. Enhanced tolerance to chilling stress in OsMYB3R-2 transgenic rice is mediated by alteration in cell cycle and ectopic expression of stress genes. Plant Physiol. 150, 244–256 (2009).
Liu, K. et al. Overexpression of OsCOIN, a putative cold inducible zinc finger protein, increased tolerance to chilling, salt and drought, and enhanced proline level in rice. Planta 226, 1007–1016 (2007).
Shvedunova, M. & Akhtar, A. Modulation of cellular processes by histone and non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 23, 329–349 (2022).
Xu, Q. et al. ROS-stimulated protein lysine acetylation is required for crown root development in rice. J. Adv. Res. 48, 33–46 (2023).
Xu, Q. et al. Histone deacetylases control lysine acetylation of ribosomal proteins in rice. Nucleic Acids Res. 49, 4613–4628 (2021).
Yu, Y., Zhao, F., Yue, Y., Zhao, Y. & Zhou, D. X. Lysine acetylation of histone acetyltransferase adaptor protein ADA2 is a mechanism of metabolic control of chromatin modification in plants. Nat. Plants 10, 439–452 (2024).
Zhang, H., Zhao, Y. & Zhou, D. Rice NAD+-dependent histone deacetylase OsSRT1 represses glycolysis and regulates the moonlighting function of GAPDH as a transcriptional activator of glycolytic genes. Nucleic Acids Res. 45, 12241–12255 (2017).
Li, Y. et al. METTL3 acetylation impedes cancer metastasis via fine-tuning its nuclear and cytosolic functions. Nat. Commun. 13, 6350 (2022).
Zhang, X. L. et al. K235 acetylation couples with PSPC1 to regulate the m6A demethylation activity of ALKBH5 and tumorigenesis. Nat. Commun. 14, 3815 (2023).
Narita, T., Weinert, B. T. & Choudhary, C. Functions and mechanisms of non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 20, 156–174 (2019).
Zheng, Y. et al. Histone deacetylase HDA9 and transcription factor WRKY53 are mutual antagonists in regulation of plant stress response. Mol. Plant 13, 598–611 (2020).
Ma, X. et al. An enhanced network of energy metabolism, lysine acetylation, and growth-promoting protein accumulation is associated with heterosis in elite hybrid rice. Plant Commun. 4, 100560 (2023).
Choudhary, C. et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325, 834–840 (2009).
Okada, A. K. et al. Lysine acetylation regulates the interaction between proteins and membranes. Nat. Commun. 12, 6466 (2021).
Márquez, I. et al. Structural and functional insights into lysine acetylation of cytochrome c using mimetic point mutants. FEBS Open Bio 11, 3304–3323 (2021).
Jin, D. et al. m6A demethylase ALKBH5 inhibits tumor growth and metastasis by reducing YTHDFs-mediated YAP expression and inhibiting miR-107/LATS2-mediated YAP activity in NSCLC. Mol. Cancer 19, 40 (2020).
Shi, L. & Tu, B. P. Acetyl-CoA and the regulation of metabolism: mechanisms and consequences. Curr. Opin. Cell Biol. 33, 125–131 (2015).
Wellen, K. E. et al. ATP–citrate lyase links cellular metabolism to histone acetylation. Science 324, 1076–1080 (2009).
Xu, Q. et al. ACL and HAT1 form a nuclear module to acetylate histone H4K5 and promote cell proliferation. Nat. Commun. 14, 3265 (2023).
Du, Y. et al. SUMOylation of the m6A-RNA methyltransferase METTL3 modulates its function. Nucleic Acids Res. 46, 5195–5208 (2018).
Hou, G. et al. SUMOylation of YTHDF2 promotes mRNA degradation and cancer progression by increasing its binding affinity with m6A-modified mRNAs. Nucleic Acids Res. 49, 2859–2877 (2021).
Liao, Y. et al. HSP90β impedes STUB1-induced ubiquitination of YTHDF2 to drive sorafenib resistance in hepatocellular carcinoma. Adv. Sci. 10, e2302025 (2023).
Li, J. et al. O-GlcNAcylation promotes the cytosolic localization of the m6A reader YTHDF1 and colorectal cancer tumorigenesis. J. Biol. Chem. 299, 104738 (2023).
Hofmann, S., Kedersha, N., Anderson, P. & Ivanov, P. Molecular mechanisms of stress granule assembly and disassembly. Biochim. Biophys. Acta Mol. Cell. Res. 1868, 118876 (2021).
Lee, K. P. et al. The m6A reader ECT1 drives mRNA sequestration to dampen salicylic acid-dependent stress responses in Arabidopsis. Plant Cell 36, 746–763 (2024).
Wu, X. et al. N6-Methyladenosine-mediated feedback regulation of abscisic acid perception via phase-separated ECT8 condensates in Arabidopsis. Nat. Plants 10, 469–482 (2024).
Hu, Y., Lu, Y., Zhao, Y. & Zhou, D. X. Histone acetylation dynamics integrates metabolic activity to regulate plant response to stress. Front. Plant Sci. 10, 1236 (2019).
Lu, Y. et al. Metabolic regulation of the plant epigenome. Plant J. 114, 1001–1013 (2023).
Fu, W., Wu, K. & Duan, J. Sequence and expression analysis of histone deacetylases in rice. Biochem. Biophys. Res. Commun. 356, 843–850 (2007).
Kim, J. M., To, T. K. & Seki, M. An epigenetic integrator: new insights into genome regulation, environmental stress responses and developmental controls by HISTONE DEACETYLASE 6. Plant Cell Physiol. 53, 794–800 (2012).
He, Y. et al. Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol. Plant 11, 1210–1213 (2018).
Liu, W. et al. DSDecode: a web-based tool for decoding of sequencing chromatograms for genotyping of targeted mutations. Mol. Plant 8, 1431–1433 (2015).
Lv, Y. et al. New insights into the genetic basis of natural chilling and cold shock tolerance in rice by genome-wide association analysis. Plant Cell Environ. 39, 556–570 (2016).
Huang, D. et al. Nanoscale zero-valent iron assisted phytoremediation of Pb in sediment: impacts on metal accumulation and antioxidative system of Lolium perenne. Ecotoxicol. Environ. Saf. 153, 229–237 (2018).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, 252–258 (2014).
Zhao, H. et al. PaACL silencing accelerates flower senescence and changes the proteome to maintain metabolic homeostasis in Petunia hybrida. J. Exp. Bot. 71, 4858–4876 (2020).
Wang, L. et al. The ATR–WEE1 kinase module inhibits the MAC complex to regulate replication stress response. Nucleic Acids Res. 49, 1411–1425 (2021).
Song, P. et al. Methyltransferase ATMETTL5 writes m6A on 18S ribosomal RNA to regulate translation in Arabidopsis. New Phytol. 244, 571–587 (2024).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Cui, S. et al. The RNA binding protein EHD6 recruits the m6A reader YTH07 and sequesters OsCOL4 mRNA into phase-separated ribonucleoprotein condensates to promote rice flowering. Mol. Plant 17, 935–954 (2024).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Picard. Broad Institute https://broadinstitute.github.io/picard/ (2019).
Meng, J., Cui, X., Rao, M. K., Chen, Y. & Huang, Y. Exome-based analysis for RNA epigenome sequencing data. Bioinformatics 29, 1565–1567 (2013).
Wei, J. et al. FTO mediates LINE1 m6A demethylation and chromatin regulation in mESCs and mouse development. Science 376, 968–973 (2022).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Ge, S. X., Jung, D. & Yao, R. ShinyGO: a graphical gene-set enrichment tool for animals and plants. Bioinformatics 36, 2628–2629 (2020).
Acknowledgements
We thank T. Hu for the construction of transgenic plants. We thank X. Li for management and H. Song for help in confocal microscopy. We also thank the National Center for Protein Sciences at Peking University in Beijing, China, for assistance with sequencing, and G. Li and X. Zhang for help with the MGI2000 experiment. This work was supported by grants from the National Key R&D Program of China (nos 2024YFF1000302 and 2023ZD04073), the National Natural Science Foundation of China (nos 32070563, 22225704 and 32470307), the Fundamental Research Funds for the Central Universities (no. 2662023SKPY002) and the earmarked fund for China Agriculture Research System (no. CARS-01). The Fundamental Research Program of Hubei Province (2024AFE001).
Author information
Authors and Affiliations
Contributions
Y.Z. conceived and designed the research, and D.-X.Z. supervised the research. N.M. performed the experiments and wrote the initial draft. P.S. performed the m6A-seq, FA-CLIP, m6A-IP–qPCR and FA-RIP–qPCR experiments. Z.C., Y.L. and P.S. performed the data analysis. Z.L., M.D. and T.L. participated in the experiments. X.M., Q.X. and Y.Y. joined in the transgenic plant regeneration. Y.Z. and D.-X.Z. analysed the data. Y.Z., D.-X.Z. and G.J. revised the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Hunseung Kang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
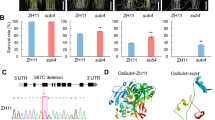
Extended Data Fig. 1 Relative expression levels of YTH domain-containing genes in rice.
RT-qPCR analysis of YTH domain family gene expression levels in 14-day-old wild-type rice seedlings. Actin was used as a reference gene. Data represent the means ± s.d. (three separate experiments per sample). The significance was calculated using one-way ANOVA with Tukey’s multiple comparison tests. Different lowercase letters over the bars indicate significant differences (p < 0.05).
Extended Data Fig. 2 Sequence alignment of the YTH domains from different species.
Os, Oryza sativa; At, Arabidopsis thaliana; Hs, Homo sapiens. Tryptophan (W), which forms the methyl-interacting aromatic cage, is highlighted in cyan. Lysine (K), the acetylation site shown in Fig. 1a, is highlighted in magenta.
Extended Data Fig. 3 Production and detection of OsECT3 transgenic plants.
a, Production of rice OsECT3 CRISPR-Cas9 mutants. The positions of single guide RNA (sgRNA) in the gene (indicated by black arrows) and decoded sequence mutations of the OsECT3 gene are shown. b, c, Detection of relative mRNA and protein levels in WT and COM-E3-F (OsECT3pro::OsECT3-Flag/osect3) plants by RT-qPCR (b) and immunoblotting with anti-Flag antibody (c), respectively. d, Determination of relative OsECT3 mRNA levels in WT and COM-E3-G (OsECT3pro::OsECT3-GFP/osect3), COM-E3K471R (OsECT3pro::OsECT3K471R-GFP/osect3), and COM-E3K471Q (OsECT3pro::OsECT3K471Q-GFP/osect3) plants by RT-qPCR. e-g, Detection of GFP-targeted OsECT3 (e), OsECT3(K471R/Q) (f, g) protein levels using anti-GFP antibodies in the respective complementary transgenic plants. For all data, bars indicate the mean ± s.d. of three replicates (one-way ANOVA with Tukey’s multiple comparison tests, the significance level was p < 0.05). RT-qPCR used ACTIN as a reference gene, each sample are three biological replicates. Immunoblot analyses show the indicated protein expression of the transgenes in T2 seedlings 14 days after germination. Four independent lines are shown for each construct, anti-actin used as a control.
Extended Data Fig. 4 Analysis of OsECT3/OsECT33WA RNA-binding activities and production of OsECT33WA transgenic plants.
a-b, Tests of OsECT3- (a) and OsECT33WA- (b) binding activities to RNA probes containing the RRACH motif. For the RNA-EMSA, 4 nmol of the RNA probe were used. RNA probe sequence is 5′-FAM-UCUUUGGXCUGACUUGGACUCUUUA-3′ (X = A/m6A). Protein concentrations were indicated. Two repetitions were performed. c-d, Detection of OsECT3 mRNA and protein levels in WT and COM-E33WA (OsECT3pro::OsECT33WA-Flag/osect3) plants by RT-qPCR (c) and immunoblotting with anti-Flag antibody (d). Bars indicate the mean ± s.d. of three replicates (one-way ANOVA with Tukey’s multiple comparison tests, the significance level was p < 0.05).
Extended Data Fig. 5 Acetylation of the lysine-471 residue of OsECT3 affects its m6A binding activity, but not its subcellular localization, protein level or stability.
a, Computer modeling of m6A (colored in magenta) binding affinity of OsECT3(K471Q) and OsECT3(K471R) (colored in orange) proteins. b, OsECT3 and OsECT3(K471R/Q) mutants localized in the cytoplasm under normal and cold conditions. Confocal images showing cytoplasmic localization of GFP in root tips of OsECT3pro::OsECT3-GFP/osect3 and OsECT3pro::OsECT3K471R/Q-GFP/osect3. Scale bar = 5 μm. c, Analysis of OsECT3 protein levels in 14-day-old seedlings in complementation plants with WT OsECT3 or its mutant variants. OsECT3 was detected using an anti-GFP antibody, with anti-actin used as a control. d, Recombinant OsECT3 and its mutant proteins (OsECT3(K471R/Q)) were incubated with rice cell extracts for semi-in vitro protein degradation assays. The degradation rate of OsECT3 and OsECT3(K471R/Q) were not significantly different. CBB (Coomassie Blue) staining was used as a loading control. Two independent experiments were performed in b-d.
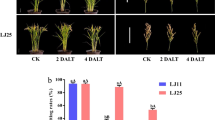
Extended Data Fig. 6 Phenotypes of osect3 mutants in response to heat, salt, and PEG6000 stress.
Phenotypes of 14-day-old wild-type, osect3 seedlings before and after 42 °C treatment (a), 20% PEG6000 treatment (b), 180 mM NaCl treatment (c) for 3 days and subsequent recovery for 14 days. Bars = 4 cm. Mean ± s.d. of 5 biological replicates (n = 36 plants for each replicate) are shown in a-c. P values from two-tailed t-test.
Extended Data Fig. 7 HDA705 transgenic plant production and stress-induced expression analysis.
a, Generation of hda705 knockout mutants using CRISPR/Cas9 technology. Two positions (black arrow indicated) of the designed sgRNA and the decoded mutations of the HDA705 gene are shown. b, Detection of HDA705 in hda705 complementation plants (HDA705pro::HDA705-GFP/hda705) by immunoblotting with anti-GFP, and anti-Actin used as a control. Two independent experiments were performed. c-d, Relative expression levels of HDA705 in wild-type seedlings under 20% PEG6000 (c) and 180 mM NaCl (d) treatments. Actin was used as the internal control gene. Data are presented as mean ± s.d. of three replicates. p values indicate significant differences compared to normal at each time point, as determined by two-tailed, unpaired t-test.
Extended Data Fig. 8 HDA705 deacetylase activity towards OsECT3 in the HDA705 overexpression plants under cold stress.
a, Detection of relative HDA705 mRNA levels in WT and HDA705 overexpression (OE-HDA705) plants under normal condition. b, Relative expression levels of HDA705 under cold conditions for 0, 6, 12, and 24 h in the OE-HDA705 plants. ACTIN was used as a reference gene, data are presented as mean ± s.d. of three replicates. The significance was calculated using one-way ANOVA with Tukey’s multiple comparison tests in a-b. c, Protein levels of HDA705 treated by cold for 0, 12 h in the OE-HDA705 plants. HDA705-Flag protein was detected with anti-Flag antibody and anti-Actin antibody was used as a loading control. d, Detection of deacetylation level of OsECT3 treated by cold for 0, 12 h in the OE-HDA705 plants. OsECT3 protein was immunoprecipitated with anti-OsECT3 antibody, followed by immunoblotting with anti-OsECT3 or anti-ac-lys antibodies. e, Tests of HDA705 deacetylase activity in 14-day-old WT seedlings under normal and cold conditions. HDA705 protein was isolated by immunoprecipitation using anti-HDA705. HDA705 activities were shown relative to normal conditions (set at 100 %). Error bars are mean ± s.d. from three biological replicates. Significant difference was calculated by the two-tailed, unpaired t-test, the significance level was p < 0.05. Two replicates are shown.
Extended Data Fig. 9 OsECT3 binds to mRNA 3’UTR regions and m6A levels quantification under normal and cold conditions.
a, Go enrichment analysis of known genes bond by OsECT3 under cold stress. b, Motifs identified by HOMER software based on the OsECT3 binding and m6A sites under normal and cold stress conditions. c, Distribution of the distance of OsECT3 binding and m6A sites under cold stress conditions compared to those under normal conditions. d, Boxplot showing m6A levels of 1,701 common OsECT3- and m6A-targeted genes under both conditions. Results were calibrated with m6A spike-ins to diminish the difference in efficiency during immunoprecipitation in m6A-seq. The medians (horizontal lines), interquartile ranges (boxes), whiskers (± 1.5× interquartile range) and outliers of the data are shown (n = 1,701 common OsECT3- and m6A-targeted genes from two biological replicates). P-values were calculated using two-sided Wilcoxon test. e, LC-MS/MS quantification of the m6A/A ratio in polyadenylated RNA isolated from 14-day-old WT seedlings under both conditions. Data are presented as mean ± s.d., n = 6 independent experiments with 2 technical replicates each. ns, not significant by unpaired two-tailed t-test. f, Venn diagrams showing the overlap of OsECT3 and OsECT3(K471Q/R) binding sites detected by FA-CLIP with m6A peaks detected by m6A-seq under cold condition. g, Go enrichment analysis of genes bond by OsECT3, but not by OsECT3(K471Q) under cold stress.
Extended Data Fig. 10 mRNA m6A enrichment and stability of 4 OsETC3 target genes under normal and cold conditions.
a. Relative mRNA m6A levels of the 4 cold-repsonsive mark genes by m6A-IP-qPCR in WT, osect3, COM-E3-G, COM-E33WA, COM-E3K471R, and COM-E3K471Q seedlings under normal and cold conditions. External m6A spike-in was used for calibration. Data are presented as mean ± SE, n = 3 independent experiments. The significance was calculated using one-way ANOVA with Tukey’s multiple comparison tests. b-c, The mRNA half-lives of the 4 genes in WT and osect3 seedlings under normal conditions (b) and COM-E3K471Q, COM-E3K471R seedlings under cold conditions (c). Data are presented as mean ± s.d., n = 2 independent experiments, each with 2 technical replicates.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2.
Supplementary Dataset 1 (download XLSX )
Statistics of OsECT3 and m6A targets under normal conditions.
Supplementary Dataset 2 (download XLSX )
Statistics of OsECT3 and m6A targets under cold conditions.
Supplementary Dataset 3 (download XLSX )
Statistics of OsECT3/OsECT3(K471Q), OsECT3(K471R) and m6A targets under cold conditions.
Source data
Source Data Figs. 1–3 and 5 (download PDF )
Unprocessed western blots and/or gels.
Source Data Figs. 2 and 4–8 (download XLSX )
Statistical source data.
Source Data Extended Data Figs. 1, 3, 4, 6–8 and 10 (download XLSX )
Statistical source data.
Source Data Extended Data Figs. 3–5, 7 and 8 (download PDF )
Unprocessed western blots and/or gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ma, N., Song, P., Liu, Z. et al. Regulation of m6A RNA reader protein OsECT3 activity by lysine acetylation in the cold stress response in rice. Nat. Plants 11, 1165–1180 (2025). https://doi.org/10.1038/s41477-025-02013-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02013-w
This article is cited by
-
Reciprocal control of metabolic and chromatin regulators improves rice tolerance to heat
Nature Communications (2025)