Abstract
Plant cell surface pattern recognition receptors (PRRs) perceive non- or altered-self elicitors to induce immune responses. PRRs relay information across the plasma membrane and trigger downstream signalling via receptor-like cytoplasmic kinases such as BOTRYTIS-INDUCED KINASE 1 (BIK1). BIK1 associates with several PRRs and acts as a key executor of immune responses through the phosphorylation of substrate proteins. However, a comprehensive understanding of how BIK1 targets specific substrates and a full repertoire of these substrates are lacking. Here we defined the substrate specificity of BIK1 and used these data to predict candidate substrates in Arabidopsis. Using high-throughput biochemical and genetic screening of these candidates, we confirmed many as direct BIK1 substrates in vitro and novel regulators of plant immunity. Among the BIK1 substrates identified are MULTIPLE C2 DOMAIN AND TRANSMEMBRANE REGION PROTEIN 3, which we reveal regulates flagellin 22 (flg22)-induced plasmodesmata closure and immunity, and members of the largely uncharacterized CYCLIN-DEPENDENT KINASE-LIKE family, which we uncover as novel negative regulators of immunity. In parallel, we interrogated intracellular NUCLEOTIDE-BINDING LEUCINE-RICH REPEAT (NLR) immune receptors for potential BIK1 phosphorylation motifs and identified multiple NLRs as direct BIK1 substrates. We reveal that BIK1 phosphorylation regulates NLR oligomerization, thus controlling a key activation step for these immune receptors. Together, our unbiased biochemical screens shed light on the central role of BIK1 as a key kinase shaping multiple layers of plant immune signalling.
Similar content being viewed by others
Main
Plants possess a sophisticated, two-tiered immune system. At the cell surface, pattern recognition receptors (PRRs) of the receptor kinase or receptor protein families perceive diverse non- or altered-self elicitor molecules1,2,3. Perception of elicitors by PRRs activates cellular signalling that culminates in pattern-triggered immunity (PTI)4. Inside the cell, receptors of the nucleotide-binding and leucine-rich-repeat protein (NLR) family recognize pathogen-derived effector proteins to mediate effector-triggered immunity (ETI)5. Recent evidence indicates that PTI and ETI are interconnected6,7,8,9, although the molecular mechanisms underlying this crosstalk remain to be fully elucidated5.
PTI signalling downstream of activated PRR complexes is relayed by associated receptor-like cytoplasmic kinases (RLCKs), in particular members of the large RLCK-VII/AVRPPHB SUSCEPTIBLE 1 (PBS1)-LIKE (PBL) family4,10. In the model plant Arabidopsis thaliana (hereafter, Arabidopsis), the RLCK-VII/PBL family member BOTRYTIS-INDUCED KINASE 1 (BIK1) functions downstream of many PRRs and acts as a key mediator of immune signalling in response to diverse elicitors11,12,13,14. RLCKs such as BIK1 are generally thought to act via direct phosphorylation of substrate proteins4. The first BIK1 substrate identified was the NADPH oxidase RESPIRATORY BURST OXIDASE HOMOLOG D (RBOHD). Upon elicitor perception, RBOHD is phosphorylated by BIK1 at multiple regulatory sites within its cytosolic N-terminus that activate RBOHD-dependent production of reactive oxygen species (ROS)15,16, thereby triggering a key signalling output of PTI.
Substrate recognition by kinases is governed by several factors, including recognition of linear motifs immediately adjacent to a given phosphosite17,18,19. Each of the BIK1-targeted sites in RBOHD conforms to an S/T-X-X-L motif (where S/T is a given Ser/Thr phosphosite, X represents any amino acid and L indicates leucine)15. This observation subsequently aided the identification of additional BIK1 substrates based on the presence of S/T-X-X-L motif-conforming phosphosites20,21,22,23,24. However, the functional relevance of this +3 Leu motif was never directly assessed.
BIK1-mediated regulation of its substrates is motif dependent
The RBOHD regulatory phosphosites S39, S343 and S347 are targeted by BIK1, and each conforms to the S/T-X-X-L motif15. To test the importance of this motif in BIK1 transphosphorylation, we mutated these three phosphosites to non-phosphorylatable Ala (S39A/S343A/S347A, RBOHD3A) or each of the corresponding +3 Leu residues to Gly (L42G/L346G/L350G, RBOHD3G) and assessed phosphorylation by BIK1. Wildtype (WT) BIK1 but not a kinase-dead variant (K105A/K106A, BIK1*) phosphorylated the RBOHD N-terminus (RBOHD-N) but not RBOHD-N3A (Fig. 1a). Phosphorylation of RBOHD3G-N was severely reduced (Fig. 1a), indicating that the +3 Leu motif is required for targeting of these sites by BIK1. We confirmed dependence of BIK1 phosphorylation on the +3 Leu for the RBOHD S39 phosphosite using quantitative kinase assays (Fig. 1b).
a, In vitro kinase assay showing transphosphorylation of 6xHis-MBP-RBOHD-N variants (labelled MBP-RBOHD-N in figure) by BIK1. 6xHis-MBP-MBP (labelled MBP-MBP in figure) served as a negative control. CBB, Coomassie brilliant blue. b, Peptide kinase assay with RBOHD-Ser39 WT or mutant peptide substrates; n = 3 independent replicates. Mean values are shown with error bars ±s.d. RLU, relative light units. c, RBOHD activation assay in HEK293T cells; n = 5 biologically independent replicates. Mean values are shown with error bars ±s.d. d, flg22-induced ROS production in N. benthamiana expressing the indicated constructs; n = 24 leaf discs. Mean values are shown with error bars ±s.e.m. e, In vitro kinase assay showing transphosphorylation of 6xHis-MBP-PLL4-N variants by BIK1. 6xHis-MBP-MBP (labelled MBP-MBP in figure) served as a negative control. f, elf18-induced ROS production in N. benthamiana expressing the indicated constructs; n = 24 leaf discs. Mean values are shown with error bars ±s.e.m. g, AFM-generated model of BIK1 and bound RBOHD-Ser39 10mer motif. Residues that form the predicted +3 Leu-binding hydrophobic pocket on BIK1 are indicated. The Ser39 peptide is shown in yellow. BIK1 is shown in green with pocket residues highlighted in magenta (top) or coloured by electrostatic surface potential (red, negative; blue, positive; bottom). h, Peptide kinase assays using WT or mutant variants of BIK1 with RBOHD-Ser39 peptide substrate; n = 3 independent replicates. Mean values are shown with error bars ±s.d. i, 32P autoradiographic PSPA data generated using recombinant BIK1. j, Peptide kinase assays using RBOHD-Ser39 peptide substrate variants; n = 5 independent replicates. Mean values are shown with error bars ±s.d. k, BIK1 PSSM and derived substrate motif sequence logo. All experiments aside from AFM modelling and PSPA were performed at least twice with similar results. In b, c, h and j, different letters indicate statistical differences (one-way ANOVA (analysis of variance) followed by Tukey’s HSD (honestly significant difference), P < 0.05). Detailed statistical parameters are reported in Supplementary Table 6. In b, h and j, 6xHis-BIK1 is labelled BIK1 in the figure. In a, b, c and e, BIK1* indicates kinase-dead BIK1.
To assess the role of the +3 Leu motif in RBOHD activation, we heterologously co-expressed RBOHD variants with BIK1 in human embryonic kidney (HEK293T) cells (Extended Data Fig. 1a), an established method for quantitative measurement of RBOH activity25,26,27. As expected, BIK1 was able to activate WT RBOHD but not RBOHD3A (Fig. 1c). Activation of RBOHD3G was also severely attenuated compared to WT, indicating that the +3 Leu motif is required for full activation of RBOHD (Fig. 1c). To validate these findings in planta, we expressed RBOHD variants in Nicotiana benthamiana and measured ROS production induced by the elicitor flg22. Overexpression of WT RBOHD led to increased flg22-induced ROS production, while that of RBOHD3A or RBOHD3G did not (Fig. 1d).
The protein phosphatase POLTERGEIST-LIKE 4 (PLL4) is a BIK1 substrate that negatively regulates PTI signalling via inhibition of PRR activation21,28. A similar relevance of the +3 Leu motif for BIK1-dependent regulation was confirmed for PLL4 using PLL46G, a variant in which the +3 Leu/Ile residues of six BIK1-targeted phosphosites were mutated to Gly (I118G/L125G/I134G/L143G/I150G/L167G) and PLL46A, a phospho-null variant21 (Fig. 1e,f). Like PLL46A, PLL46G was not phosphorylated by BIK1, resulting in decreased PTI activation (as measured by ROS production) upon co-expression with the PRR ELONGATION FACTOR TU RECEPTOR (EFR) and treatment with its elicitor ligand, elf18 (ref. 29) (Fig. 1f).
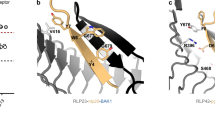
In silico structural modelling using AlphaFold Multimer (AFM)30 docked the RBOHD Ser39-substrate peptide into the BIK1 active site and placed the +3 Leu substrate residue into a peripheral BIK1 hydrophobic pocket (Fig. 1g). This predicted complex was also supported by structural alignment with a previously reported structure of BIK1 (ref. 31) and a phosphorylase kinase bound to a peptide substrate32 (Extended Data Fig. 1b). Mutation of BIK1 residues that comprise this putative hydrophobic pocket (V239A, Y243A or M251A) did not impair BIK1 catalytic activity (Extended Data Fig. 1c). However, mutation of these residues substantially decreased transphosphorylation of the RBOHD-S39 peptide (Fig. 1h), supporting a role for these hydrophobic residues in accommodating +3 Leu substrates.
Substrate specificity of BIK1
To more thoroughly define the sequence determinants of BIK1 substrate specificity, we performed a positional scanning peptide array (PSPA) analysis33,34,35, in which a combinatorial peptide library was used for 32P-labelled kinase assays with recombinant BIK1. PSPA data confirmed a clear preference for Leu at the +3 position (Fig. 1i).
To test the contribution of positions other than the +3 Leu to phosphorylation by BIK1, we individually mutated each of the −1, +1 or +2 residues within the RBOHD-S39 site motif to unfavourable residues based on the PSPA data (amino acid mutations F38A, G40L or P41A, respectively) and tested phosphorylation by BIK1 using quantitative peptide kinase assays. Transphosphorylation of F38A and G40L peptides was reduced to background levels, similar to S39A and L42G controls and a quadruple F38A/G40L/P41A/L42G mutant peptide (4xmut), while phosphorylation of P41A was significantly reduced compared to WT (Fig. 1j). This confirmed that selectivity at each of these positions contributes to substrate targeting by BIK1, at least in the context of this substrate.
Motif-based screening for candidate BIK1 substrates
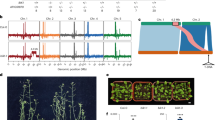
To identify candidate BIK1 substrates, we interrogated the predicted Arabidopsis proteome by assigning every Ser or Thr residue a score using a position-specific scoring matrix (PSSM) derived from the PSPA data (Fig. 1k). Several filters were applied to increase the likelihood that such proteins are involved in immunity and could co-localize with BIK1 (Methods). This yielded a list of 77 unique motif-containing proteins, which we termed candidate BIK1 substrate proteins (CBSPs) (Fig. 2a and Supplementary Table 1). Sixteen of these CBSP motifs were previously identified as in planta phosphosites36,37 (Supplementary Table 1). Modelling of CBSP motif interactions with BIK1 using AFM predicted that most substrates dock into the kinase active site with high confidence compared to corresponding scrambled CBSP peptides (Fig. 2b,c).
a, CBSP screen overview. b,c, AFM modelling of CBSP motifs. An overlaid model of all CBSPs with ipTM scores >0.75 (b) and the distribution of ipTM scores for WT or sequence-scrambled CBSP motifs (c). d, In vitro kinase assay using BIK1 and MBP-RBOHD-N. MBP-MBP served as a negative control. BIK1* indicates kinase-dead BIK1. Mean values are shown with error bars ±s.e.m.; P value from one-tailed t-test. a.u., arbitrary units. e, volcano plot of in vitro CBSP transphosphorylation by BIK1; n = 3 independent experimental replicates. Each point represents the mean ratio of autoradiographic signal for an MBP-CBSP protein versus a MBP control, as calculated in d. P values are derived from one-tailed t-tests. f–h, Volcano plots of infection assays using syringe-infiltrated Pst DC3000 (f), spray-inoculated Pst DC3000 Cor− (g) and drop-inoculated B. cinerea (h). Each point represents 1–3 independent experiments (n = 5–6 biologically independent plants per experimental replicate in f; n = 3–4 biologically independent plants per experimental replicate in g and h). Values are change in measured infection (either log10(c.f.u. cm−2 or lesion size; c.f.u., colony-forming units) of cbsp mutants versus corresponding Col-0 or Ler WT control. Each point represents the mean measured infection across all biological replicates for that genotype comparison. Linear mixed-effects modelling was used for statistical analysis. Detailed statistical parameters are reported in Supplementary Table 6. Models in f–h created with BioRender.com.
To validate substrates, we purified recombinant full-length or soluble fragments of CBSPs and performed in vitro kinase assays with BIK1, using RBOHD-N and maltose-binding protein (MBP) as positive and negative controls, respectively (Fig. 2d). In total, 48 of 77 CBSPs could be successfully recombinantly purified, of which 32 were confirmed as phosphorylated by BIK1 (Fig. 2e).
BIK1 substrates regulate immunity
To test whether CBSPs are involved in immunity, we generated a collection of cbsp null mutants using published transfer-DNA or newly generated CRISPR/Cas9 (clustered regularly interspaced short palindromic repeats and CRISPR-associated protein 9) lines. We obtained and analysed 42 different single or higher-order mutants corresponding to 46 total CBSPs (Supplementary Table 2) and challenged them with three pathogens: virulent (DC3000) and partially disarmed (Cor−) strains of the hemi-biotrophic bacterium Pseudomonas syringae pv. tomato (Pst) and the necrotrophic fungus Botrytis cinerea. We identified 22 CBSPs as either positive or negative regulators of immunity (Fig. 2f–h), with the majority having no previously documented function in defence. The identification of known regulators of immunity (for example, MYELOCYTOMATOSIS 2 (MYC2)38, PLANT-SPECIFIC TFIIB-RELATED PROTEIN 1 (PBRP1)39, ENHANCED DISEASE RESISTANCE 4 (EDR4)40,41 and MITOGEN-ACTIVATED PROTEIN KINASE KINASE KINASE KINASES (MAP4Ks)42,43) highlights the power of our screening approach and reveals their previously undocumented potential regulation by BIK1.
BIK1 and MCTPs control flg22-induced plasmodesmata closure and immunity
MULTIPLE C2 DOMAIN AND TRANSMEMBRANE REGION PROTEIN 3 (MCTP3) was among the proteins confirmed as BIK1 substrates with no previously reported roles in immunity and features the highest-scoring BIK1 motif among the CBSP list (Supplementary Table 1). MCTPs possess N-terminal Ca2+-binding C2 domains, which mediate association with the plasma membrane, and C-terminal transmembrane domains anchoring MCTPs to the endoplasmic reticulum44,45,46. The predicted BIK1 motif, including S506, is found adjacent to the final C2 domain of the MCTP3 N-terminus. BIK1 phosphorylated MCTP3 in a site-specific manner (Fig. 3a and Extended Data Fig. 2a) and associated with MCTP3 specifically following flg22 treatment (Fig. 3b).
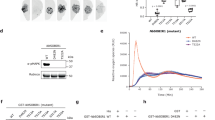
a, In vitro kinase assay between BIK1 and variants of an MCTP3 N-terminal region. b, Co-affinity purification (Co-AP) assay of native BIK1 using Col-0/35S::Lti6b-GFP or mctp3-2/pMCTP3::YFP-MCTP3 transgenic Arabidopsis seedlings, with or without 15 min treatment with 1 µM flg22. c,d, Plasmodesmata closure assay using particle bombardment and GFP diffusion. Representative images of epidermal cells (c) and quantification (d). In c, scale bar, 100 µm. In d, n ≥ 92 sites analysed per genotype/treatment from a minimum of six leaves from biologically independent plants. Statistical analysis in d was performed using a bootstrap analysis. Mean values are shown with error bars ±s.e.m. e,f, In vitro kinase assay between BIK1 and variants of CDKL5 (e) or CDKL6 (f) C-termini. In e, 2A indicates S523A/S610A. In f, 2A indicates S614A/S706A. g, Co-AP assay of native BIK1 using Col-0/35S::Lti6b-GFP or cdkl5 cdkl6/pCDKL5::CDKL5-GFP transgenic Arabidopsis seedlings. h, In planta phosphorylation of CDKL5-S610 as detected by affinity purification-mass spectrometry (AP-MS) using cdkl5 cdkl6/pCDKL5::CDKL5-GFP transgenic Arabidopsis seedlings. Normalized peptide abundance is shown with or without 1 μM flg22 treatment for 10 min. Peptide sequence with modification is shown above plot; error bars ±s.e.m.; P value was determined using unpaired two-tailed t-test; n = 3 independent biological replicates. a.u., arbitrary units. i, ROS production in adult Arabidopsis leaves following treatment with 100 nM flg22; n = 24. Mean values are shown with error bars ±s.e.m. j, Callose deposition in adult Arabidopsis leaves 24 h after treatment with 1 µM flg22; n = 31 to 32 biologically independent leaf discs. Mean values are shown with error bars ±s.e.m. k,l, Infection assays with Pst DC3000 by spray (k) or infiltration (l) inoculation, n = 8. Arabidopsis fls2 efr cerk1 (fec) mutants served as negative control in k. In l, WT or kinase-dead (*) pCDKL5::CDKL5-GFP lines in the cdkl5 cdkl6 background were used. Boxes show median values with 25th to 75th percentiles; error bars show min to max values. In j, k and l, statistical analysis was performed using a one-way ANOVA followed by Tukey’s HSD. Detailed statistical parameters are reported in Supplementary Table 6. All experiments were performed at least three times with similar results. In a, e and f, BIK1* indicates kinase-dead BIK1.
The MCTP3 S506-containing motif is conserved in several members of the family, including its close homologue MCTP4 (Extended Data Fig. 2b–d). MCTP3 and MCTP4 mediate plasma membrane–endoplasmic reticulum membrane tethering and thereby act as regulators of plasmodesmata aperture45,46,47. Plasmodesmata are cell–cell communication structures in plants and are dynamically closed in response to immune activation48,49. flg22-induced plasmodesmata closure, as measured by reduced cell-to-cell movement of green fluorescent protein (GFP), was lost in bik1 CRISPR knockout lines14 as well as mctp3 mctp4 (Fig. 3c,d), indicating that flg22-induced regulation of plasmodesmata is both BIK1 and MCTP3/4 dependent. In addition, mctp3 mctp4 showed increased susceptibility to all pathogens tested (Fig. 2f–h), suggesting that MCTPs are key BIK1 substrates positively regulating plasmodesmata closure and immunity. As MCTPs are evolutionarily conserved components of plasmodesmata50, our findings suggest they function in a core phosphorylation-dependent regulatory module of plasmodesmata.
CDKLs as BIK1 substrates and negative regulators of immunity
Among the CBSP list were multiple members of the CYCLIN-DEPENDENT KINASE-LIKE (CDKL) family. CDKLs are conserved and largely unstudied plant-specific kinases that are distinct from similarly named mammalian CDKLs (Extended Data Fig. 3a–c)51,52. Arabidopsis has 16 CDKLs, the majority of which contain at least one conserved BIK1 motif in their C-termini and several of which are transcriptionally regulated by elicitor treatment (Extended Data Fig. 3a–d). BIK1 was accordingly able to phosphorylate the C-termini of several CDKLs, including those found on our CBSP list (Extended Data Fig. 3e).
Several CDKLs, particularly CDKL5, interacted with BIK1 (Extended Data Fig. 4a–d). BIK1 phosphorylated the C-termini of CDKL5 and its close homologue CDKL6 in a motif-dependent manner (Fig. 3e,f) and associated with CDKL5 in Arabidopsis (Fig. 3g). We were able to confirm in planta phosphorylation of both BIK1-targeted sites on CDKL5 (S523 and S610; Supplementary Table 3), with both being induced by flg22 treatment (Fig. 3h and Extended Data Fig. 4e), albeit S523 being ambiguous depending on the peptide variant examined.
The cdkl5 cdkl6 double-knockout mutant showed increased responses to elicitors (Fig. 3i,j and Extended Data Fig. 4f–j) and resistance to Pst DC3000 (Figs. 2f and 3k and Extended Data Fig. 4k), as well as a dwarf developmental phenotype and elevated PR1 expression (Extended Data Fig. 5a,b), suggesting that CDKL5 and 6 are negative regulators of defence. However, we cannot exclude the possibility that these CDKLs are guarded by NLR(s), with their absence thus triggering NLR-dependent autoimmunity in a mechanism unrelated to their signalling function3. To clarify the function of these CDKLs in immunity, we compared complementation of cdkl5 cdkl6 by WT or kinase dead (CDKL5*, K166A; Extended Data Fig. 5c) variants. While CDKL5* rescued the growth defect and elevated PR1 seen in cdkl5 cdkl6 (Extended Data Fig. 5d,e), it was unable to rescue increased disease resistance (Fig. 3l), indicating that CDKL5-mediated signalling negatively regulates PTI in a kinase activity-dependent manner.
NLRs as BIK1 substrates
Among our initial CBSP list were two NLRs (Supplementary Table 1). Plant NLRs are mainly classified based on their N-terminal domains: coiled-coil (CC)-NLRs (CNLs), Toll/interleukin-1 receptor-NLRs (TNLs) and RESISTANCE TO POWDERY MILDEW 8 (RPW8)-like-NLRs (RNLs)53. Effector perception is primarily achieved by ‘sensor’ CNLs and TNLs, while RNLs function as immune signalling ‘helpers’54,55. Emerging evidence indicates that PTI and ETI are tightly interconnected8,9,56,57; however, the molecular mechanism(s) coordinating such interplay remain largely unknown. In addition, recent studies suggest that NLRs can be regulated by phosphorylation58,59. To systematically determine whether NLRs may be regulated by BIK1, we performed a targeted screen of all known Arabidopsis NLRs for potential BIK1 motifs, in parallel to our previous, proteome-wide CBSP screen. Using a relaxed cut-off score of 2.5 (Supplementary Table 4) and filtering for sensor NLRs with known cognate bacterial effectors (to ease subsequent functional characterization), we identified five NLRs as potential BIK1 substrates (Extended Data Fig. 6a), including the TNLs RESISTANCE TO RALSTONIA SOLANACEARUM-SUSCEPTIBLE (RRS1-S), RESISTANT TO PSEUDOMONAS SYRINGAE 4B (RPS4B) and HOPB-ACTIVATED RESISTANCE 1 (BAR1) and the CNLs RESISTANT TO PSEUDOMONAS SYRINGAE 5 (RPS5) and RESISTANT TO P. SYRINGAE PV MACULICOLA 1 (RPM1). The predicted motifs were found across NLRs domains and were conserved across Arabidopsis accessions (Extended Data Fig. 6b).
Upon transient expression in N. benthamiana, RRS1-S, RPS4B, BAR1, RPS5 and RPM1, but not the negative control RNL N REQUIREMENT GENE 1.1 (NRG1.1) associated with BIK1 (Extended Data Fig. 6c). RPM1 was not included in our analysis due to its strong induction of cell death in this system60,61. The BIK1 motif-containing domains of RPS5, RPS4B and RRS1-S could be purified and were phosphorylated by BIK1 in vitro (Fig. 4a,b and Extended Data Fig. 6d), whereas NRG1.1 was not (Extended Data Fig. 6e), suggesting our screen could identify true NLR substrates of BIK1. Based on these findings, we selected RPS4B and RPS5 as TNL- and CNL-representative BIK1 substrates, respectively, for further analysis. We then confirmed their association with BIK1 in Arabidopsis (Fig. 4c,d) and site-specific phosphorylation by BIK1 (Extended Data Fig. 6f–i).
a,b, In vitro kinase assays of 6xHis-BIK1 with MBP-RPS5-CC domain (RPS5 in the figure) (a) or MBP-RPS4B NBARC (RPS4B in the figure) (b). MBP-MBP served as negative control. BIK1* indicates kinase-dead BIK1. c, Co-AP assay using Col-0 protoplasts transfected with plasmids expressing BIK1-HA and RPS5-FLAG or NRG1.1-FLAG. NRG1.1-FLAG served as a negative control. d, Co-AP assay using protoplasts from Col-0 or RPS4B-FLAG transgenic plants. Protoplasts were transfected with plasmid expressing BIK1-HA. e,f, BIK1-motif phospho-mimetic mutation in RPS5 (e) or RPS4B (f) compromises ETI-triggered cell death. In e, RPS5-MYC variants and PBS1-HA with or without AvrPphB-FLAG were co-expressed in N. benthamiana leaves. In f, RPS4B-FLAG variants and RRS1B-MYC with or without AvrRps4 were co-expressed in N. benthamiana. Cell death was photographed 2 d.p.i. for RPS5 (e), and 4–5 d.p.i. for RPS4B (f). g,h, Cell death after infiltration of Pst DC3000D36E expressing AvrPphB (g) or AvrRps4 (h) into Arabidopsis RPS5-MYC (g) or RPS4B-FLAG (h) complementation lines. Cell death was scored 12 h.p.i. (hours post infiltration) (g) or 24 h.p.i. (h), and numbers indicate the number of leaves with cell death relative to the total number of infiltrated leaves; n = 16. i,j, Growth of Pst DC3000 expressing AvrPphB (i) or AvrRps4 (j) was assessed in Arabidopsis RPS5-MYC or RPS4B-FLAG complementation lines. Bacterial colonies were counted 3 days after spray inoculation, and statistical significance was tested by one-way ANOVA followed by Tukey’s HSD (different letters indicate P < 0.05). Detailed statistical parameters are reported in Supplementary Table 6. Mean values are shown with error bars ±s.e.m.; n = 3 biologically independent plants). k, The RPS5-S19D mutation compromises RPS5 oligomerization upon AvrPphB recognition following transient expression in N. benthamiana. Samples were extracted 2 d.p.i. for BN–PAGE and SDS–PAGE. l, The RPS4B-S520D mutation compromises RRS1B-RPS4B oligomerization following transient expression in N. benthamiana. Samples were collected 2 d.p.i. for BN–PAGE and SDS–PAGE. The expected size of RPS4B-MYC is indicated by an open arrowhead. m,n, BIK1 dissociates from RPS5 (m) and RPS4B (n) upon flg22 treatment in Arabidopsis transgenic protoplasts. Co-AP assay was conducted using protoplasts isolated from RPS5-MYC (m) or RPS4B-FLAG (n) transgenic Arabidopsis lines transfected with plasmid expressing BIK1-HA, with or without 100 nM flg22 treatment for 10 min. All experiments were conducted at least three times with similar results.
NLR phosphorylation suppresses ETI activation
Activation of RPS4B and RPS5 by their respective cognate effectors, AvrRps4 and AvrPphB, can be reconstituted upon heterologous expression in Nicotiana spp.62,63, which possess functional homologues of Arabidopsis BIK1 (refs. 64,65). RPS5 is activated upon cleavage of the host target protein PBS1 by the cysteine protease effector AvrPphB66. Co-expression of RPS5, PBS1 and AvrPphB in N. benthamiana triggers rapid cell death62, which was greatly attenuated when expressing the RPS5 phospho-mimetic (S19D) mutant (Fig. 4e). Neither differential protein accumulation nor cleavage of PBS1 and AvrPphB could explain the observed reduction in cell death (Extended Data Fig. 7a), indicating that the S19D mutation compromised RPS5 activation. RPS4B requires its paired TNL RRS1B for AvrRps4 recognition, similar to the well-studied paralogue RPS4–RRS1 pair63. Co-expression of RPS4B, RRS1B and AvrRps4 induced cell death, which was abolished by the BIK1-motif phospho-mimetic mutation (S520D) in RPS4B, whereas S520A accelerated cell death (Fig. 4f). As with RPS5, differential accumulation did not explain the loss of cell death induced by RPS4B-S520D (Extended Data Fig. 7b).
Consistently, RPS5-S19D expression attenuated cell death in Arabidopsis upon Pst DC3000D36E(AvrPphB) infiltration, while RPS5-S19A expression did not (Fig. 4g and Extended Data Fig. 7c,d). RPS4B-S520D complementation lines were completely non-responsive to AvrRps4 in our cell death assays, similar to results in N. benthamiana (Fig. 4h and Extended Data Fig. 7e,f). Expression of RPS4B-S520A led to a stunted developmental phenotype, indicative of constitutive immune induction, although PR1 levels were not elevated in these lines (Extended Data Fig. 7g,h).
AvrPphB-induced Pst DC3000 growth restriction was not affected by RPS5-S19A and -S19D mutations (Fig. 4i), suggesting that the phosphorylation on S19 only compromises the early onset of cell death, not all ETI responses. Conversely, AvrRps4-induced Pst DC3000 growth restriction correlated with AvrRps4-induced cell death phenotype (Fig. 4j), suggesting that for RPS4B, its phosphorylation by BIK1 affects both cell death induction and bacterial growth restriction.
We also generated WT, phospho-ablative or -mimetic complementation lines for RRS1-S and BAR1. While the RRS1 BIK1-motif phospho-mimetic mutation did not affect AvrRps4 recognition, the BAR1 phospho-mimetic mutant (T599D/T626D) showed compromised growth restriction after recognition of its cognate effector HopB1d67 (Extended Data Fig. 8). Our data therefore indicate that BIK1-mediated phosphorylation inhibits the activation of multiple NLRs.
BIK1-mediated phosphorylation inhibits NLR oligomerization
We next sought to investigate the mechanism by which BIK1-mediated NLR phosphorylation suppresses their activation. Localization of NLRs has been reported to be critical for their function68,69,70. However, phospho-mimetic and -ablative NLR variants localized similarly to their respective WT proteins (Extended Data Fig. 9a). We then investigated whether BIK1-mediated phosphorylation affects NLR biochemical properties. RPS5 interacts with PBS1, and this interaction is important for RPS5 activation62, while RPS4B requires interaction with its paired partner RRS1B for activation63. However, BIK1 phosphosite mutations did not affect these interactions (Extended Data Fig. 9b,c).
Appropriate NLR oligomerization is critical for their function71,72. RPS5 WT and S19A, but not S19D, oligomerized in the presence of AvrPphB and PBS1 in N. benthamiana, suggesting that phosphorylation by BIK1 compromises AvrPphB-induced RPS5 oligomerization (Fig. 4k). Recently, the TNL pair CHS3-CSA1 and RPS4-RRS1 were reported to form constitutive oligomers in planta73,74. Co-expression of RRS1B with RPS4B but not RPS4B-S520D resulted in oligomerization (Fig. 4l and Extended Data Fig. 9d), indicating that phosphorylation by BIK1 suppresses RPS4B activation by compromising RPS4B-RRS1B oligomer pre-formation. Inhibition of oligomerization by BIK1-dependent phosphorylation is thus a conserved mechanism across different NLRs. BIK1 dissociated from RPS4B and RPS5 upon flg22 treatment (Fig. 4m,n and Extended Data Fig. 10), suggesting that PTI may alleviate the constitutive BIK1-mediated phosphorylation-dependent inhibition of NLR activation.
Recent studies suggest that PTI activation may be directly involved in regulating NLR oligomerization. Both PTI and ETI activation lead to the oligomerization of the RNL NRG1.2, and the common PRR co-receptor BAK1/BKK1 suppresses the oligomerization of the TNL pair CHS3–CSA1 and subsequent cell death activation57,73. Inhibition of NLR oligomerization by PRR complex component in naive cells might thus be a common strategy for NLR regulation, facilitating effective ETI potentiation upon PTI activation. Recent studies have also implicated members of other kinase families in NLR phosphorylation and regulation58,59. Future research will be needed to delineate how such phosphorylation events may contribute to the overall regulation of NLRs pre- and post-immune elicitation.
Conclusion
The BIK1 substrates identified in our study reveal how BIK1 controls elicitor-induced plasmodesmata closure, identify previously uncharacterized plant-specific kinases as immune regulators and provide mechanistic insights into PTI–ETI interplay via BIK1-mediated NLR phosphorylation. Our motif-based strategy to screen for BIK1 substrates bypasses limitations associated with both protein–protein interaction and phosphoproteomic-based approaches to identify kinase substrates, allowing us to identify novel components and regulatory mechanisms in immunity that may not have been identified by other means. However, phosphorylation motifs correspond to linear interactions at the kinase active site and constitute only one mechanism controlling kinase–substrate targeting17, and thus BIK1 may target substrate proteins that would not be identified based solely on their motifs. While many putative substrates were phosphorylated by BIK1 in vitro in our study, further work may be required to conclusively demonstrate a direct kinase–substrate relationship in planta for all the putative targets we have identified.
BIK1 is one member of the 46-member RLCK-VII/PBL family75, several of which function in immunity or other receptor-kinase-mediated pathways21,75,76,77,78,79,80. Despite apparent pathway-specific functions, other PBLs have been found to phosphorylate similar substrates to BIK1 (refs. 21,25). Our approach, in addition to significantly increasing our knowledge of plant immune signalling and its regulation, provides a blueprint that can be applied to identify RLCK (and potentially other kinase) substrates and therefore clarify the molecular basis of specificity within and across plant signalling pathways.
Methods
Plant growth conditions
Arabidopsis plants were grown on soil in a controlled environment growth chamber at 120 µmol light intensity, 50% relative humidity and 20 °C in a 10 h light/14 h dark cycle. Plants grown for bombardment assays were grown at 80 µmol light intensity and 22 °C in a 10 h light/14 h dark cycle. Arabidopsis seedlings were grown in vitro on half-strength Murashige and Skoog media supplemented with vitamins, 1% sucrose, 0.9% agar, stratified for 2–3 days in the dark at 4 °C and moved to a growth chamber set to 22/18 °C, 60% relative humidity and a 16 h light/8 h dark cycle.
N. benthamiana plants were grown in a controlled environment chamber at 150 µmol light intensity, 45–60% relative humidity and 25 °C in a 16 h light/8 h dark cycle.
Molecular cloning
Constructs for expression in Escherichia coli were generated by cloning PCR fragments into pOPINM using InFusion (Takara), Golden Gate assembly into pOPINM (TSL Synbio) or pET28a(+)-GG, or using restriction cloning into pGEX-4T1 or pET28(+)-TEV.
For Gateway cloning, PCR products were cloned into pDONR vectors using BP Clonase (Invitrogen) and subsequently into pDEST using LR Clonase (Invitrogen). Constructs for expression in HEK293T cell were generated by GreenGate cloning81. For generation of CDKL5-GFP transgenic lines, the CDKL5 coding sequence and pCDKL5 promoter sequence were amplified from Arabidopsis complementary DNA and genomic DNA, respectively. The CDKL5 coding DNA sequence (CDS) was cloned into pGGC000 using GreenGate cloning, and the pCDKL5 promoter was added by InFusion cloning.
For CRISPR/Cas9 cloning, guide RNAs (gRNAs) were designed using the CHOPCHOP (v3) tool82, and constructs were generated using a golden gate cloning system83, unless otherwise stated. One primer pair for each gRNA was designed including a BsaI site (Supplementary Table 5). Either annealed oligomers of gRNA primers were assembled into pAGM55261 (ref. 84), or the forward primer containing the Cas9-target sequence was used in a PCR with the forward gRNA primer and the reverse primer on the guide backbone BCJJ458A. The product was cloned together with the pU6 promoter from pICSL90002 into level 1 vectors. For each gene, two level 1 vectors were generated containing the pU6 promoter and the gRNA. gRNAs targeting CDKL5 were cloned into pICH47761 and pICH47772. The gRNAs targeting CDKL6 were cloned into pICH47781 and pICH47791. All four level 1 vectors were cloned into the level M vector pAGM8067 with endlinker pICH50932 using BpiI. The resulting level M vector was cloned with BCJJ369B and endlinker pICH79311 into level P vector pICSL4723 using BsaI. The level P vector served as a binary vector for plant expression. For mutation of CNGC19 in the cngc20-2 background, a single-guide RNA (sgRNA) targeting CYCLIC-NUCLEOTIDE-GATED CHANNEL 19 (CNGC19) (CNGC19/Cas9-Forward/Reverse) was designed using the CRISPR-GE tool85 and cloned into the AtU626-sgRNA-SK86. The cassettes were obtained by digestion with SpeI and NheI and inserted into the pCAMBIA1300-pYAO:Cas9 (ref. 86). For mutation of MAP4K6 in the map4k5-1 background, a single sgRNA was cloned into the pKIR1.1 binary vector87.
For Arabidopsis protoplast expression, PCR fragments amplified from Arabidopsis cDNA were cloned into p641, then golden gate assembled into a modified protoplast expression vector (HBT) with pICSL50007 (C-terminal 3xFLAG). For N. benthamiana transient expression constructs under 35S promoter, PCR fragments were amplified from Arabidopsis cDNA, gDNA, cloned effector constructs or synthesized genes and subcloned into p641-Esp3I, then golden gate assembled into pICSL86977OD vector with appropriate epitope tags (pICSL50007 (C-terminal 3× FLAG), pICSL50009 (C-terminal 6xHA), pICSL50010 (C-terminal 4xMYC) or pICSL50044 (C-terminal mEGFP) (TSL Synbio). For stable transgenic line generation, PCR products for coding sequences and promoter sequences were amplified from Arabidopsis gDNA and subcloned into p641-Esp3I. Coding sequences and promoter constructs were golden gate assembled into pICSL86955OD with pICH41421 (NOS terminator) and appropriate C-terminal tag construct. For HopB1d cloning into a broad-range expression plasmid (pVSP61-GG), the coding sequence and promoter sequence was synthesized (Genescript) and subcloned into pUC19B using SmaI, and golden gate assembled into pVSP61-GG with pICSL50009.
Site-directed point mutations were generated using DpnI-mediated overlapping extension PCR methods, Golden-Gate cloning or gene synthesis (Genescript). All primers used in this study are listed in Supplementary Table 5.
Plant transformation and screening
Binary vectors were transformed into Agrobacterium tumefaciens GV3101, and Arabidopsis was transformed by floral dip. For transgenic Arabidopsis lines, seedlings were selected with BASTA spray on soil. Experiments with transgenic NLR lines were conducted in BASTA-selected T2 lines. CDKL5-GFP transgenic lines were selected with hygromycin. For CRISPR/Cas9 seed selection, T1 seeds were screened for FastRed fluorescence using a Leica MZ FLIII microscope. PCR and sequencing across sgRNA sites in target genes were performed in T1 plants, and mutation-positive plants were used for seed propagation. T2 seeds were counter-selected for FastRed to isolate lines without the CRISPR/Cas9 transgene cassette. Homozygous plants were isolated based on PCR (C19-Cas9-PCR-F/R) and sequencing.
Arabidopsis protoplast isolation, transfection and PAMP treatment
Arabidopsis protoplast isolation was conducted as previously reported88, with modifications. Briefly, leaves from 5-week-old Arabidopsis were cut, and the epidermis was removed using 3M tape. Prepared leaves were immersed with enzyme solution (1.5% cellulase R10, 0.4% macerozyme, 0.4 M mannitol, 20 mM MES pH 5.7, 20 mM KCl, 10 mM CaCl2, 0.1% BSA) for 90 min, and the isolated mesophyll protoplasts werecollected using centrifugation at 100 × g for 2 min. After two washes with W5 buffer (154 mM NaCl, 125 mM CaCl2, 5 mM KCl, 2 mM MES pH 5.7), protoplasts were resuspended in MMg buffer (0.4 M mannitol, 15 mM MgCl2, 4 mM MES pH 5.7) at a density of 2 × 105 ml−1. Fifty micrograms of plasmid constructs was transfected in 500 μl of protoplasts with equal volume of PEG solution (40% PEG4000, 200 mM mannitol, 100 mM CaCl2) for 15 min. Transfection was stopped by washing once with W5 buffer. After 12–16 h of incubation in W5 buffer, the transfected protoplasts were treated with 100 nM flg22 for times as indicated in figure captions and collected using centrifugation at 100 × g for 5 min.
Protein extraction, co-affinity purification and immunoblotting
Assays were performed as previously described21 with modifications. Briefly, for the study of NLRs in N. benthamiana, 1 g of Agrobacteria-infiltrated leaves were collected 2 days post infiltration (d.p.i.). Samples were ground in liquid nitrogen, and proteins were extracted in 2 ml protein extraction buffer (10% glycerol, 25 mM Tris pH 7.5, 1 mM EDTA, 150 mM NaCl, 1 mM Na3VO4, 1× protease inhibitor cocktail, 1 mM PMSF, 0.5% IGEPAL CA-630, 5 mM DTT). Protein extract was mixed with 2× SDS loading buffer and boiled at 90 °C for 10 min. For Arabidopsis leaf samples, four 8 mm leaf discs were ground in liquid nitrogen and boiled in 2× SDS loading buffer at 90 °C. For Arabidopsis seedlings, 48 seedlings per condition were ground in liquid nitrogen following treatment, and proteins were extracted in protein extraction buffer (10% glycerol, 50 mM Tris pH 7.5, 2 mM EDTA, 150 mM NaCl, 1 mM Na3VO4, 1× protease inhibitor cocktail, 1 mM PMSF, 0.5% IGEPAL CA-630, 5 mM DTT) at a 2:1 powder/buffer ratio. One hundred microlitres of extract were used for input sample, and the rest was incubated with 50 µl of α-FLAG M2 affinity gel (Sigma) for 120 min. The resin was washed four times with washing buffer (10% glycerol, 50 mM Tris pH 7.5, 2 mM EDTA, 150 mM NaCl, 0.5% IGEPAL CA-630), and proteins were eluted by boiling at 90 °C for 10 min in SDS loading buffer. For Arabidopsis protoplasts, collected protoplasts were vortexed in 400 µl of protein extraction buffer. Forty microlitres was taken for input samples and the rest co-incubated with 20 µl of α-MYC EZview resin (Sigma) or α-FLAG M2 affinity gel (Sigma) for 90 min. The resins were washed four times with washing buffer (10% glycerol, 25 mM Tris pH 7.5, 1 mM EDTA, 150 mM NaCl, 0.5% IGEPAL CA-630) and boiled at 90 °C in SDS loading buffer.
Proteins were separated by SDS–PAGE and immunoblotted using α-MYC (Cell Signaling), α-mouse-HRP (Sigma), α-BIK1 (Agrisera), α-rabbit-HRP (Sigma), α-FLAG-HRP (Sigma) or α-HA-HRP (Rosche).
Blue-native PAGE
Blue-native PAGE (BN–PAGE) was conducted as previously published with the following modifications89,90. Briefly, four 4 mm diameter leaf discs from the agroinfiltrated N. benthamiana leaves were collected and ground in liquid nitrogen. Four hundred microlitres of 1× Native sample buffer (Invitrogen) with 0.2% n-dodecyl-beta-maltoside (DDM) was added in the ground tissue sample. Twenty microlitres of protein sample was mixed with 0.5 µl of Coomassie blue additive (Invitrogen). Protein samples were separated by BN–PAGE according to the manufacturer’s protocol. The proteins were transferred to PVDF membranes and immunoblotted using appropriate antibodies.
Recombinant protein expression and purification
All proteins were expressed in E. coli strain BL21(DE3) Rosetta pLysS unless otherwise noted. All proteins were purified using HisPur cobalt resin (Thermo), PureCube Co-NTA agarose (Cube Biotech) or GST-Bind Resin (Millipore).
PSPA
Recombinant BIK1 was added to a 384-well plate containing peptide substrate library mixtures in solution phase at 50 μM (Anaspec, AS-62017-1 and AS-62335) in kinase buffer (25 mM Tris–Cl pH 7.5, 5 mM MnCl2, 5 mM MgCl2, 1 mM dithiothreitol). The reaction was initiated with the addition of 50 μM ATP (50 μCi ml−1 γ-32P-ATP, Perkin-Elmer) and incubated for 90 min. After completion of the reaction, the solutions were spotted onto streptavidin-conjugated membranes (Promega, V2861), where the peptides tightly associated through their C-terminal biotinylation. The membranes were rinsed and then imaged using the Typhoon FLA 7000 phosphorimager (GE) to measure the extent of peptide phosphorylation. Raw data (GEL file) were quantified using ImageQuant (GE) to generate densitometry matrices. Raw BIK1 PSPA data are provided in Supplementary Table 6.
PSSM scoring and in silico CBSP screen
The densitometry matrices were processed, first by normalizing all amino acid columns against nonselective positions (that is, the −5 position for N-terminal residues and +4 position for C-terminal residues). All values were log2-scaled to generate the BIK1 PSSM (Supplementary Table 6). Individual motif scores were derived by the sum of the log-transformed matrix values. Scoring was applied to all Ser and Thr sites within the predicted Arabidopsis proteome, based on the reference genome Araport11 (ref. 91). The resulting scores were sorted by their values. We then set a cut-off score of 3.0 for motifs to examine further based on the known RBOHD-S39 site (score ~3.2). This produced a list of motifs corresponding to approximately 900 unique proteins, which we subsequently filtered by transcriptional regulation (log2 fold change >1 or less than −1) of corresponding genes by the elicitors flg22, elf18 and AtPep1 (ref. 92) and predicted exposure of the motif to the cytosol and/or nucleus, where BIK1 would be potentially active as a PM-associated cytoplasmic protein with reported partial nuclear localization31,93.
AFM structural modelling
Protein complex predictions for full-length BIK1 and 10-mer peptide of RBOHD S39 was performed using the ColabFold platform (v1.5.2)30,94. Then 7-mer CBSP and CBSP-scrambled motifs were performed using the ColabFold platform (v1.5.5)30,94. Peptides were scrambled with Peptide Nexus, positioning the phospho-accepting residue at position 0. The process involved three independent queries, without the use of template structures, no relaxation steps, and MMseqs2 UniRef + Environmental for multiple sequence alignment (MSA) mode. The AlphaFold2 Multimer v3 model was used to generate five models per query, with each model undergoing three recycling steps and one ensemble calculation. The models were ranked based on their multimer scores, and the model with the highest score (0.8 × interface predicted Template Modeling (ipTM) score + 0.2 × predicted Template Modelling score) was chosen for BIK1 substrate interaction analysis.
In vitro kinase assays and autoradiography quantification
Assays were performed as described previously21 with modifications. Briefly, equal amounts (1 µg) of kinase and substrate protein were mixed in kinase buffer (25 mM Tris–Cl pH 7.4, 5 mM MnCl2, 5 mM MgCl2, 1 mM dithiothreitol). Reactions were initiated by addition of 5 µM ATP supplemented with 0.5 µCi 32P-γ-ATP in a final reaction volume of 30 µl. Reactions were carried out at 25 °C for 30 min and stopped by addition of SDS-loading dye and heating at 70 °C for 10 min. Proteins were resolved by SDS–PAGE, transferred to PVDF membrane and stained with Coomassie brilliant blue G-250. Autoradiographs were imaged using an Amersham Typhoon Phosphorimager (GE Healthcare). For semi-quantitative assays, autoradiographs were quantified using ImageQuant TL (Amersham). Areas of the same size were drawn around bands corresponding to recombinant proteins to determine the autoradiography volume. These quantified volumes were normalized for background noise by subtracting the median detected autoradiographic volume within the defined area. For peptide-based assays, BIK1 and substrate peptide (sequences found in Supplementary Table 7) were mixed (final concentration 1 µM and 100 µM, respectively) in kinase buffer. Reactions were initiated with the addition of 100 µM ATP supplemented with 0.1 µCi 32P-γ-ATP in a final reaction volume of 25 µl and carried out at room temperature for 20 min. Twenty microlitres of each reaction was transferred to a corresponding pre-cut 1 cm2 piece of cation exchange paper (SVI-P). Samples were washed three times in 0.5% phosphoric acid, and 32P signal was measured as counts per minute (c.p.m.) on a scintillation counter.
In vitro identification of BIK1-targeted phosphorylation sites on RPS4B and RPS5
Kinase reactions were performed as described above without addition of 32P-γ-ATP. Kinase reactions (2 µg of total protein) were mixed with an equal volume (25 µl) of reduction/alkylation buffer containing 7.4 M urea, 10 mM tris(2-carboxyethyl)phosphine-HCl and 30 mM 2-chloroacetamide followed by incubation for 60 min at 60 °C in the dark with shaking at 500 r.p.m. Samples were then diluted to a final volume of 250 µl with 50 mM ammonium bicarbonate (ABC). Reactions containing RPS4B as the substrate protein were incubated overnight at 37 °C with 100 ng of trypsin (Promega). Reactions containing RPS5 as the substrate protein were first incubated overnight at 37 °C with 100 ng of trypsin followed by overnight incubation at 37 °C with 100 ng of Asp-N (Promega). Afterwards, digests were acidified by the addition of 20 µl of 10% trifluoroacetic acid (TFA). Peptides were cleaned up using the StageTip method95, dried down, redissolved in 20 µl of MS (mass spectrometry) buffer (3% acetonitrile (AcN), 0.1% formic acid) and loaded into MS sample vials.
Liquid chromatography–tandem mass spectrometry (LC–MS/MS) was performed using an M-Class HPLC (Waters) coupled to an Exploris 480 orbitrap mass spectrometer (Thermo Fisher Scientific). Four microlitres of each peptide sample was loaded onto a nanoEase M/Z Symmetry C18 100 Å, 5 µm, 180 µm × 20 mm trap column at a flow rate of 15 µl min−1 in buffer A (0.1% formic acid in water) for 2 min. Peptide separation was performed on a nanoEase M/Z HSS C18 T3 Col 100 Å, 1.8 µm, 75 µm × 250 mm analytical column at a flow rate of 300 nl min−1 using a 70 min gradient from 5% to 95% buffer B (buffer B = 0.1% formic acid in AcN). Peptides were ionized at a spray voltage of 2.4 kV and a capillary temperature of 270 °C. The mass spectrometer was operated in data-dependent mode with 3 s between master scans. Full-scan MS spectra (350–1,200 m/z) were collected with a maximum injection time of 45 ms at a resolution of 120,000 and a normalized automatic gain control (AGC) target of 300%. Dynamic exclusion was set to 15 s. High-resolution MS/MS spectra were collected in the orbitrap with a maximum injection time of 150 ms at 60,000 resolution (isolation window 1.2 m/z), a normalized AGC target of 100% and a normalized collision energy of 30%. Only precursors with charge states from 2 to 7 were selected for fragmentation.
The MS proteomics data were handled using the local laboratory information management system96.
Raw files were processed using MSFragger97,98 (version 4.2) implemented in fragpipe (version 23) using default closed-search settings unless otherwise specified. MS/MS spectra were searched against an E. coli database (Uniprot ID UP000000625, downloaded on 4 June 2024) with sequences for His-BIK1, HisMBP-RPS4B and HisMBP-RPS5 added. Trypsin of Trypsin + Asp-N was set as protease, allowing for up to two missed cleavages. Carbamidomethylation of cysteines was set as a fixed modification with methionine oxidation, N-terminal acetylation and phosphorylation of serine, threonine and tyrosine set as variable modifications. Label-free quantification was performed with IonQuant99 (version 1.11.19) using default settings implemented in fragpipe with the match between runs feature enabled. Peptide-spectrum match data are provided in Supplementary Table 8.
Identification and quantification of in vivo CDKL5 phosphosites by affinity purification-mass spectrometry (AP-MS)
For the MS-based identification of in vivo CDKL5 p-sites, 14-day-old cdkl5 cdkl6/pCDKL5:CDKL5-GFP seedlings (60 seedlings per treatment, grown in liquid 1× MS) were vacuum-infiltrated with MS (mock) or MS supplemented with 1 μM flg22 for 10 min. Protein extraction and GFP-trap AP were performed as described above. Samples were loaded on a 4–12% NuPAGE gel (Invitrogen) and run 5 cm into the gel. The gel was stained for 10 min with InstantBlue (Abcam). The CDKL5-GFP sample was excised from the gel and cut into fragments for processing. The excised gel portions were cut into smaller pieces and washed three times with 50% (v/v) AcN, 50 mM ABC (50% v/v AcN/ABC), 30 min each, followed by dehydration in AcN, 10 min. Gel pieces were then reduced with 10 mM DTT for 30 min at 45 °C followed by alkylation with 55 mM chloroacetamide for 20 min at room temperature and a further three washes with 50% (v/v) AcN/ABC, 30 min each. Gel pieces were dehydrated again with AcN before rehydration with 40 µl of trypsin (Pierce Trypsin Protease, MS grade, catalogue number 90058) working solution (100 ng of trypsin in 50 mM ABC, 5% (v/v) AcN). Where required, gel pieces were covered with 50 mM ABC to a final volume before incubation at 37 °C overnight. Tryptic peptides were extracted from the gel pieces three times in an equal volume of 50% (v/v) AcN, 5% (v/v) formic acid (Pierce LC–MS grade, catalogue number 85178), 30 min each. Extracted peptides were dried in a speed-vac and resuspended in 2% (v/v) AcN/0.2% (v/v) TFA (Merck, catalogue number 302031). A total of three biological replicates for each sample type were submitted.
Approximately 35% of each sample was analysed using an Orbitrap Fusion Tribrid Mass Spectrometer (Thermo Fisher Scientific) coupled to a U3000 nano-UPLC (Thermo Fisher Scientific). The dissolved peptides were injected onto a reverse phase trap column NanoEase m/z Symmetry C18, beads diameter 5 μm, inner diameter 180 μm × 20 mm length (Waters). Trap column flow rate was 20 μl min−1 in 2% (v/v) AcN, 0.05% (v/v) TFA. Peptides were eluted from trap column onto the analytical column NanoEase m/z HSS C18 T3 Column, beads diameter 1.8 μm, inner diameter 75 μm × 250 mm length (Waters). The column was equilibrated with 3% buffer B (buffer B, 80% (v/v) AcN in 0.05% (v/v) formic acid; buffer A, 0.1% (v/v) formic acid) before subsequent elution with the following steps of a linear gradient: 2.5 min 3% buffer B, 5 min 6.3% buffer B, 13 min 12.5% buffer B, 50 min 42.5% buffer B, 58 min 50% buffer B, 61 min 65% buffer B, 63 min 99% buffer B, 66 min 99% buffer B, 67 min 3% buffer B, 90 min 3% B. The flow rate was set to 200 nl min−1. The mass spectrometer was operated in positive ion mode with nano-electrospray ion source. Molecular ions were generated by applying voltage +2.2 kV to a conductive union coupling the column outlet with fused silica PicoTip emitter, ID 10 μm (New Objective), and the ion transfer capillary temperature was set to 275 °C. The mass spectrometer was operated in data-dependent mode using a full scan, m/z range 300–1,800, nominal resolution of 120,000, target value 1 × 106, followed by MS/MS scans of the 40 most abundant ions. MS/MS spectra were acquired using normalized collision energy of 30%, isolation width of 1.6 m/z, resolution of 120,000 and a target value set to 1 × 105. Precursor ions with charge states 2–7 were selected for fragmentation and put on a dynamic exclusion list for 30 s. To improve detection of phosphorylation, multistage activation was applied for detection of −98, −49 or −32.7 from the precursor (corresponding to the neutral loss of phosphoric acid from +1, +2 and +3 charge states, respectively) during any of the MS/MS scans. The minimum automatic gain control target was set to 5 × 103, and the intensity threshold was calculated to be 4.8 × 104. The peptide match feature was set to the preferred mode, and the feature to exclude isotopes was enabled.
Peak lists in the form of Mascot generic files were prepared from raw data files using MS Convert (Proteowizard) and sent to a peptide search on Mascot server v.3.1 using Mascot Daemon (Matrix Science) against a contaminants database and the Araport11 protein database. Tryptic peptides with up to two possible mis-cleavages and charge states +2, +3 and +4 were allowed in the search. The following peptide modifications were included in the search: carbamidomethylated cysteine (fixed), oxidized methionine (variable) and phosphorylated serine, threonine and tyrosine (variable). Data were searched with a monoisotopic precursor and fragment ion mass tolerance 10 p.p.m. and 0.8 Da, respectively. Decoy database was used to validate peptide sequence matches.
Mascot search files (.dat) were used to construct a spectral library with Skyline software (v22.2)100 in a DDA and MS1 filtering workflow. CDKL5 (AT5G44290) protein was added to the document target list and the corresponding Thermo.raw files then imported via results. Automatic peak area selection was used with minimal manual correction. A selection of 11 control peptides based on library rank and of similar intensity were used to determine relative CDKL5 levels in each sample. All peptide variants carrying S523 and S610 phosphorylation were included. Data were exported to Excel and each phospho-peptide intensity normalized to relative CDKL5 level (summed control peptide intensities) per sample.
In vitro pull down
Assays were performed as previously described21.
Split luciferase assay
Assays were performed as previously described101 with minor modifications. Briefly, vector combinations were infiltrated into 4- to 5-week-old N. benthamiana leaves. After 2 d.p.i., 4 mm leaf discs were placed in 96-well plates in double-distilled H2O. After equilibration for 6 h, the water was replaced with 1 mM luciferin, and the leaf discs were kept in darkness for 1 h. Luminescence was then measured using a Tecan Spark.
RBOHD activation assay in HEK293T cells
Assays were performed as previously described25. In brief, HEK293T cells were seeded into white, tissue -culture coated 96-well plates (Greiner) and incubated at 37 °C and 5% CO2. After 24 h, cells were transfected with the PEI-MAX transfection reagent (Polysciences), using either 50 ng RBOHD plasmid or 50 ng RBOHD and 15 ng BIK1 plasmids per well. Forty-eight hours after transfection, cells were washed with Hank’s balanced salt solution (Gibco). One hundred microlitres of measurement buffer (Hank’s balanced salt solution with 62 μM L-012 and 60 μg ml−1 HRP) was added to each well, and luminescence was recorded with a Tecan Spark over a time span of 30 min. For each sample, five wells were transfected and assayed in parallel.
Confocal microscopy
Confocal microscopy was performed with Agrobacterium-infiltrated N. benthamiana leaves at 3 d.p.i. using a Leica Stellaris/SP5 system. GFP was excited at 488 nm, and emission was collected between 494 nm and 560 nm. Images were processed identically between mutant variants using Fiji software (v2.9.0)102.
P. syringae infection and bacterial growth restriction assay
Pst DC3000 carrying AvrRps4, AvrPphB or HopB1d were generated by triparental mating with E. coli HB101 strain as described previously61. P. syringae colonies were plated on King’s B agar plates supplemented with rifampin and grown at 28 °C for 1–2 days. For the study of CBSPs, 4-week-old Arabidopsis plants were inoculated using syringe infiltration or spray inoculation. For syringe infiltration, Pst DC3000 was diluted in 10 mM MgCl2 to an optical density at 600 nm (OD600) of 0.001. For spray inoculation, Pst DC3000 Cor− was diluted in 10 mM MgCl2 with 0.04% Silwet L-77 to an OD600 of 0.2. Plants were maintained under high humidity conditions for 1 day, followed by 2 days under normal growth chamber conditions. Leaf disks were taken from infected plant tissue and homogenized in 200 μl of 10 mM MgCl2. Serial dilutions of the homogenate were plated on King’s B agar plates with rifampin. After 2 days of incubation, colonies were counted. For the study of NLRs, bacteria were spray inoculated at OD600 of 0.1 (AvrRps4 and AvrPphB) or OD600 of 1.0 (HopB1d) in 10 mM MgCl2 to 5-week-old Arabidopsis plants. At 3 d.p.i., leaf discs were collected and ground in 500 μl of 10 mM MgCl2. Each sample was serial diluted and plated on King’s B agar plate supplemented with appropriate antibiotics. Bacterial colony counts were assessed after 2 d.
B. cinerea infection
Spores of B. cinerea BMM strain were grown in the dark on potato dextrose broth 1.5% agar for 10 days. Cut leaves of 4-week-old Arabidopsis plants were transferred onto H2O agar plates. Spores were resuspended in 1/2 potato dextrose broth to a concentration of 5 × 105 spores per ml, and 5 μl of this suspension was drop-inoculated onto the leaves. Plates were sealed and incubated for 2–3 days at 22 °C under dim light. Pictures of the infected leaves were taken, and the areas of lesions were quantified using ImageJ (v1.53t).
Callose deposition
Four-millimetre leaf discs of 4-week-old Arabidopsis plants were used for the assays. Assays were performed as described previously103, with the following modifications. Washing was accomplished with 100% ethanol, and an extra overnight washing step was included. The subsequent aniline blue staining was also performed overnight. Callose depositions were imaged using a Leica Thunder M205 FCA.
ROS measurement
Assays were performed as previously described15.
Plasmodesmal flux assays
Microprojectile bombardment of pB7WG2.0.RFPER and pB7WG2.0.GFP was performed on 4- to 6-week-old Arabidopsis plants as previously described104. Two hours after bombardment, leaves were infiltrated with water (mock) or 100 nM flg22. Z-stack images were collected the next day on a LSM Zeiss 800 microscope; red fluorescent protein was excited at 561 nm with a DPSS laser and emission collected between 600 nm and 620 nm, while GFP was excited at 488 nm with an argon laser and emission collected between 505 nm and 530 nm using a ×20 water dipping objective (W N-Achroplan ×20/0.5). The relative spread of GFP was calculated as the number of cells showing GFP normalized to the mean of the given genotype mock treated data, and statistical significance between flg22 and mock treatment within a genotype was analysed using the medianBootstrap method105.
N. benthamiana cell death assays
Five-week-old N. benthamiana plants were used for transient gene expression assays. For RPS5 cell death assay, Agrobacteria carrying RPS5-MYC, PBS1-HA and AvrPphB-FLAG were mixed at OD600 = 0.1, 0.4 and 0.1, respectively, in agroinfiltration buffer (10 mM MgCl2, 10 mM MES-KOH pH 5.7). For RPS4B-RRS1B, Agrobacteria carrying RPS4B-MYC, RRS1B-MYC and AvrRps4-HA were mixed at OD600 = 0.4 each in agroinfiltration buffer. Indicated Agrobacteria mixtures were infiltrated into N. benthamiana leaves. Cell death was scored at 2 d.p.i. for RPS5 and 4–5 d.p.i. for RPS4B-RRS1B using Fusion FX camera (Vilber Lourmat) using the published setting106 or UV transilluminator.
Arabidopsis cell death and electrolyte leakage assays
Pst DC3000D36E carrying AvrRps4 or AvrPphB were infiltrated at OD600 = 0.1 in 10 mM MgCl2 into the fully expanded leaves of 5-week-old Arabidopsis plants. Cell death was scored visually at the indicated time points. For electrolyte leakage assays, four 4-mm-diameter leaf discs per condition were sampled in 3 ml deionized H2O. For each condition, 3–4 replicates were sampled. After 1 h incubation, electrolyte leakage was measured using Horiba B-174 Twin conductivity meter.
Statistics
Statistical analyses were performed using GraphPad Prism (v9.5.1) or R (v4.4.1; packages used included ‘lme4’ for fitting linear mixed-effects models and ‘lmerTest’ for obtaining P values)107,108. All statistical data and exact P values are displayed in figures or in Supplementary Table 9.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw proteomics data for the identification of phosphorylation sites on recombinant RPS4B and RPS5 are available via ProteomeXchange with identifier PXD065397. CDKL5-GFP AP-MS data are available on PanoramaPublic and ProteomeXchange with identifier PXD066643. All other data supporting the findings of this research are presented in the main text, figures and Supplementary Information. Additional data related to this study are available from the corresponding authors upon request. Source data are provided with this paper.
Change history
03 March 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41477-026-02255-2
References
Albert, I., Hua, C., Nürnberger, T., Pruitt, R. N. & Zhang, L. Surface sensor systems in plant immunity. Plant Physiol. 182, 1582–1596 (2020).
Boutrot, F. & Zipfel, C. Function, discovery, and exploitation of plant pattern recognition receptors for broad-spectrum disease resistance. Annu. Rev. Phytopathol. 55, 257–286 (2017).
Jones, J. D. G., Staskawicz, B. J. & Dangl, J. L. The plant immune system: from discovery to deployment. Cell 187, 2095–2116 (2024).
DeFalco, T. A. & Zipfel, C. Molecular mechanisms of early plant pattern-triggered immune signaling. Mol. Cell 81, 3449–3467 (2021).
Ngou, B. P. M., Ding, P. & Jones, J. D. G. Thirty years of resistance: zig-zag through the plant immune system. Plant Cell 34, 1447–1478 (2022).
Ngou, B. P. M., Ahn, H. K., Ding, P. & Jones, J. D. G. Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature 592, 110–115 (2021).
Yuan, M. et al. Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature 592, 105–109 (2021).
Pruitt, R. N. et al. The EDS1–PAD4–ADR1 node mediates Arabidopsis pattern-triggered immunity. Nature 598, 495–499 (2021).
Tian, H. et al. Activation of TIR signalling boosts pattern-triggered immunity. Nature 598, 500–503 (2021).
Liang, X. & Zhou, J. M. Receptor-like cytoplasmic kinases: central players in plant receptor kinase-mediated signaling. Annu. Rev. Plant Biol. 69, 267–299 (2018).
Lu, D. et al. A receptor-like cytoplasmic kinase, BIK1, associates with a flagellin receptor complex to initiate plant innate immunity. Proc. Natl Acad. Sci. USA 107, 496–501 (2010).
Zhang, J. et al. Receptor-like cytoplasmic kinases integrate signaling from multiple plant immune receptors and are targeted by a Pseudomonas syringae effector. Cell Host Microbe 7, 290–301 (2010).
Liu, Z. et al. BIK1 interacts with PEPRs to mediate ethylene-induced immunity. Proc. Natl Acad. Sci. USA 110, 6205–6210 (2013).
Song, B. et al. New alleles of Arabidopsis BIK1 reinforce its predominant role in pattern-triggered immunity and caution interpretations of other reported functions. Nat. Plants https://doi.org/10.1038/s41477-025-02187-3 (2026).
Kadota, Y. et al. Direct regulation of the NADPH oxidase RBOHD by the PRR-associated kinase BIK1 during plant immunity. Mol. Cell 54, 43–55 (2014).
Li, L. et al. The FLS2-associated kinase BIK1 directly phosphorylates the NADPH oxidase RbohD to control plant immunity. Cell Host Microbe 15, 329–338 (2014).
Miller, C. J. & Turk, B. E. Homing in: mechanisms of substrate targeting by protein kinases. Trends Biochem. Sci. 43, 380–394 (2018).
Ubersax, J. A. & Ferrell, J. E. Mechanisms of specificity in protein phosphorylation. Nat. Rev. Mol. Cell Biol. 8, 530–541 (2007).
Pinna, L. A. & Ruzzene, M. How do protein kinases recognize their substrates?. Biochim. Biophys. Acta 1314, 191–225 (1996).
Thor, K. et al. The calcium-permeable channel OSCA1.3 regulates plant stomatal immunity. Nature 585, 569–573 (2020).
DeFalco, T. A. et al. A conserved module regulates receptor kinase signalling in immunity and development. Nat. Plants 8, 356–365 (2022).
Wang, W. et al. SHOU4/4L link cell wall cellulose synthesis to pattern-triggered immunity. New Phytol. 238, 1620–1635 (2023).
Kong, L. et al. Dual phosphorylation of DGK5-mediated PA burst regulates ROS in plant immunity. Cell 187, 609–623.e21 (2024).
Dindas, J. et al. Direct inhibition of phosphate transport by immune signaling in Arabidopsis. Curr. Biol. 32, 488–495.e5 (2022).
Fujita, S. et al. SCHENGEN receptor module drives localized ROS production and lignification in plant roots. EMBO J. 39, e103894 (2020).
Chu, J. et al. Conservation of the PBL-RBOH immune module in land plants. Curr. Biol. 33, 1130–1137.e5 (2023).
Köster, P. et al. A bi-kinase module sensitizes and potentiates plant immune signaling. Sci. Adv. 11, eadt9804 (2025).
Holton, N., Nekrasov, V., Ronald, P. C. & Zipfel, C. The phylogenetically-related pattern recognition receptors EFR and XA21 recruit similar immune signaling components in monocots and dicots. PLoS Pathog. 11, e1004602 (2015).
Zipfel, C. et al. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125, 749–760 (2006).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. Preprint at bioRxiv https://doi.org/10.1101/2021.10.04.463034 (2022).
Lal, N. K. et al. The receptor-like cytoplasmic kinase BIK1 localizes to the nucleus and regulates defense hormone expression during plant innate immunity. Cell Host Microbe 23, 485–497 e5 (2018).
Lowe, E. D. et al. The crystal structure of a phosphorylase kinase peptide substrate complex: kinase substrate recognition. EMBO J. 16, 6646–6658 (1997).
Hutti, J. E. et al. A rapid method for determining protein kinase phosphorylation specificity. Nat. Methods 1, 27–29 (2004).
Johnson, J. L. et al. An atlas of substrate specificities for the human serine/threonine kinome. Nature 613, 759–766 (2023).
Yaron-Barir, T. M. et al. The intrinsic substrate specificity of the human tyrosine kinome. Nature 629, 1174–1181 (2024).
Willems, P. et al. The Plant PTM Viewer, a central resource for exploring plant protein modifications. Plant J. 99, 752–762 (2019).
Heazlewood, J. L. et al. PhosPhAt: a database of phosphorylation sites in Arabidopsis thaliana and a plant-specific phosphorylation site predictor. Nucleic Acids Res. 36, D1015–D1021 (2008).
Aerts, N., Pereira Mendes, M. & Van Wees, S. C. M. Multiple levels of crosstalk in hormone networks regulating plant defense. Plant J. 105, 489–504 (2021).
Xu, B., Fan, B. & Chen, Z. A critical role of a plant-specific TFIIB-related protein, BRP1, in salicylic acid-mediated immune response. Front. Plant Sci. 15, 1427916 (2024).
Wu, G. et al. ENHANCED DISEASE RESISTANCE4 associates with CLATHRIN HEAVY CHAIN2 and modulates plant immunity by regulating relocation of EDR1 in Arabidopsis. Plant Cell 27, 857–873 (2015).
Zhao, Y., Wu, G., Shi, H. & Tang, D. RECEPTOR-LIKE KINASE 902 associates with and phosphorylates BRASSINOSTEROID-SIGNALING KINASE1 to regulate plant immunity. Mol. Plant 12, 59–70 (2019).
Zhang, M. et al. The MAP4 kinase SIK1 ensures robust extracellular ROS burst and antibacterial immunity in plants. Cell Host Microbe 24, 379–391.e5 (2018).
Jiang, Y. et al. MAP4K4 associates with BIK1 to regulate plant innate immunity. EMBO Rep. 20, e47965 (2019).
Liu, L., Li, C., Liang, Z. & Yu, H. Characterization of multiple C2 domain and transmembrane region proteins in Arabidopsis. Plant Physiol. 176, 2119–2132 (2018).
Brault, M. L. et al. Multiple C2 domains and transmembrane region proteins (MCTPs) tether membranes at plasmodesmata. EMBO Rep. 20, e47182 (2019).
Li, Z. P. et al. Plant plasmodesmata bridges form through ER-dependent incomplete cytokinesis. Science 386, 538–545 (2024).
Pérez-Sancho, J. et al. Plasmodesmata act as unconventional membrane contact sites regulating intercellular molecular exchange in plants. Cell 188, 958–977.e23 (2025).
Cheval, C. et al. Chitin perception in plasmodesmata characterizes submembrane immune-signaling specificity in plants. Proc. Natl Acad. Sci. USA 117, 9621–9629 (2020).
Tee, E. E. & Faulkner, C. Plasmodesmata and intercellular molecular traffic control. New Phytol. 243, 32–47 (2024).
Johnston, M. G. et al. Comparative phyloproteomics identifies conserved plasmodesmal proteins. J. Exp. Bot. 74, 1821–1835 (2023).
Menges, M., de Jager, S. M., Gruissem, W. & Murray, J. A. H. Global analysis of the core cell cycle regulators of Arabidopsis identifies novel genes, reveals multiple and highly specific profiles of expression and provides a coherent model for plant cell cycle control. Plant J. 41, 546–566 (2005).
Ivanov, S. & Harrison, M. J. Receptor-associated kinases control the lipid provisioning program in plant-fungal symbiosis. Science 383, 443–448 (2024).
Chai, J., Song, W. & Parker, J. E. New biochemical principles for NLR immunity in plants. Mol. Plant Microbe Interact. 36, 468–475 (2023).
Feehan, J. M., Castel, B., Bentham, A. R & Jones, J. D. Plant NLRs get by with a little help from their friends. Curr. Opin. Plant Biol. 56, 99–108 (2020).
Jones, J. D. G., Vance, R. E. & Dangl, J. L. Intracellular innate immune surveillance devices in plants and animals. Science https://doi.org/10.1126/science.aaf6395 (2016).
Kourelis, J. et al. The helper NLR immune protein NRC3 mediates the hypersensitive cell death caused by the cell-surface receptor Cf-4. PLoS Genet. 18, e1010414 (2022).
Feehan, J. M. et al. Oligomerization of a plant helper NLR requires cell-surface and intracellular immune receptor activation. Proc. Natl Acad. Sci. USA 120, e2210406120 (2023).
Li, J. et al. TIR immune signalling is blocked by phosphorylation to maintain plant growth. Nat. Plants 11, 1193–1204 (2025).
Zhong, C. et al. A cell wall-associated kinase phosphorylates NLR immune receptor to negatively regulate resistosome formation. Nat. Plants 11, 561–579 (2025).
Chung, E. H. et al. Specific threonine phosphorylation of a host target by two unrelated type III effectors activates a host innate immune receptor in plants. Cell Host Microbe 9, 125–136 (2011).
Choi, S. et al. Direct acetylation of a conserved threonine of RIN4 by the bacterial effector HopZ5 or AvrBsT activates RPM1-dependent immunity in Arabidopsis. Mol. Plant 14, 1951–1960 (2021).
Pottinger, S. E. & Innes, R. W. RPS5-mediated disease resistance: fundamental insights and translational applications. Annu. Rev. Phytopathol. https://doi.org/10.1146/annurev-phyto-010820-012733 (2020).
Saucet, S. B. et al. Two linked pairs of Arabidopsis TNL resistance genes independently confer recognition of bacterial effector AvrRps4. Nat. Commun. 6, 6338 (2015).
Zhang, S., Li, H., Zhang, M. & An, Y. The MAP4 kinase NbM4K3 regulates immune responses in Nicotiana benthamiana. Phytopathol. Res. 6, 44 (2024).
Huang, W. R. H. et al. Receptor-like cytoplasmic kinases of different subfamilies differentially regulate SOBIR1/BAK1-mediated immune responses in Nicotiana benthamiana. Nat. Commun. 15, 4339 (2024).
Ade, J., DeYoung, B. J., Golstein, C. & Innes, R. W. Indirect activation of a plant nucleotide binding site–leucine-rich repeat protein by a bacterial protease. Proc. Natl Acad. Sci. USA 104, 2531–2536 (2007).
Laflamme, B. et al. The pan-genome effector-triggered immunity landscape of a host-pathogen interaction. Science https://doi.org/10.1126/science.aax4079 (2021).
Shepherd, S., Yuen, E. L. H., Carella, P. & Bozkurt, T. O. The wheels of destruction: plant NLR immune receptors are mobile and structurally dynamic disease resistance proteins. Curr. Opin. Plant Biol. 74, 102372 (2023).
Saile, S. C. et al. Arabidopsis ADR1 helper NLR immune receptors localize and function at the plasma membrane in a phospholipid dependent manner. New Phytol. 232, 2440–2456 (2021).
Jacob, P. et al. Plant ‘helper’ immune receptors are Ca2+-permeable nonselective cation channels. Science https://doi.org/10.1126/science.abg7917 (2021).
Burdett, H. et al. The plant “resistosome”: structural insights into immune signaling. Cell Host Microbe 26, 193–201 (2019).
Bi, G. & Zhou, J.-M. Regulation of cell death and signaling by pore-forming resistosomes. Annu. Rev. Phytopathol. 59, 239–263 (2021).
Yang, Y. et al. Paired plant immune CHS3-CSA1 receptor alleles form distinct hetero-oligomeric complexes. Science 383, 719–730 (2024).
Ahn, H.-K. et al. Recognition-dependent activation of the RRS1-R/RPS4 immune receptor complex. Preprint at bioRxiv https://doi.org/10.1101/2025.04.11.646618 (2025).
Rao, S. et al. Roles of receptor-like cytoplasmic kinase VII members in pattern-triggered immune signaling. Plant Physiol. 177, 1679–1690 (2018).
Li, P. et al. The receptor-like cytoplasmic kinase RIPK regulates broad-spectrum ROS signaling in multiple layers of plant immune system. Mol. Plant 14, 1652–1667 (2021).
Luo, X. et al. Tyrosine phosphorylation of the lectin receptor-like kinase LORE regulates plant immunity. EMBO J. 39, e102856 (2020).
Lee, D. et al. Regulation of reactive oxygen species during plant immunity through phosphorylation and ubiquitination of RBOHD. Nat. Commun. 11, 1838 (2020).
Alassimone, J. et al. Polarly localized kinase SGN1 is required for Casparian strip integrity and positioning. Nat. Plants 2, 16113 (2016).
Wang, W. et al. Receptor-like cytoplasmic kinases PBL34/35/36 are required for CLE peptide-mediated signaling to maintain shoot apical meristem and root apical meristem homeostasis in Arabidopsis. Plant Cell 34, 1289–1307 (2022).
Lampropoulos, A. et al. GreenGate–a novel, versatile, and efficient cloning system for plant transgenesis. PLoS ONE 8, e83043 (2013).
Labun, K. et al. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res. 47, W171–W174 (2019).
Castel, B., Tomlinson, L., Locci, F., Yang, Y. & Jones, J. D. G. Optimization of T-DNA architecture for Cas9-mediated mutagenesis in Arabidopsis. PLoS ONE 14, e0204778 (2019).
Grützner, R. et al. High-efficiency genome editing in plants mediated by a Cas9 gene containing multiple introns. Plant Commun. 2, 100135 (2021).
Xie, X. et al. CRISPR-GE: a convenient software toolkit for CRISPR-based genome editing. Mol. Plant 10, 1246–1249 (2017).
Yan, L. et al. High-efficiency genome editing in Arabidopsis using YAO promoter-driven CRISPR/Cas9 system. Mol. Plant 8, 1820–1823 (2015).
Tsutsui, H. & Higashiyama, T. pKAMA-ITACHI vectors for highly efficient CRISPR/Cas9-mediated gene knockout in Arabidopsis thaliana. Plant Cell Physiol. 58, 46–56 (2017).
Wu, F. H. et al. Tape-Arabidopsis Sandwich - a simpler Arabidopsis protoplast isolation method. Plant Methods 5, 16 (2009).
Hu, M., Qi, J., Bi, G. & Zhou, J. M. Bacterial effectors induce oligomerization of immune receptor ZAR1 in vivo. Mol. Plant 13, 793–801 (2020).
Ahn, H.-K. & Jones, J. Blue native-PAGE of protein complexes in plant cells. protocols.io https://doi.org/10.17504/protocols.io.3byl4jyd8lo5/v1 (2023).
Cheng, C.-Y. et al. Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant J. 89, 789–804 (2017).
Bjornson, M., Pimprikar, P., Nürnberger, T. & Zipfel, C. The transcriptional landscape of Arabidopsis thaliana pattern-triggered immunity. Nat. Plants 7, 579–586 (2021).
Veronese, P. et al. The membrane-anchored BOTRYTIS-INDUCED KINASE1 plays distinct roles in Arabidopsis resistance to necrotrophic and biotrophic pathogens. Plant Cell 18, 257–273 (2006).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Türker, C. et al. B-Fabric: the Swiss army knife for life sciences. In Proceedings of the 13th International Conference on Extending Database Technology 717–720 (Association for Computing Machinery, 2010); https://doi.org/10.1145/1739041.1739135
Kong, A. T., Leprevost, F. V., Avtonomov, D. M., Mellacheruvu, D. & Nesvizhskii, A. I. MSFragger: ultrafast and comprehensive peptide identification in mass spectrometry–based proteomics. Nat. Methods 14, 513–520 (2017).
Yu, F. et al. Fast quantitative analysis of timsTOF PASEF data with MSFragger and IonQuant. Mol. Cell. Proteom. 19, 1575–1585 (2020).
Yu, F., Haynes, S. E. & Nesvizhskii, A. I. IonQuant enables accurate and sensitive label-free quantification with FDR-controlled match-between-runs. Mol. Cell. Proteom. 20, 100077 (2021).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Chen, H. et al. Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol. 146, 368–376 (2007).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Mason, K. N., Ekanayake, G. & Heese, A. Staining and automated image quantification of callose in Arabidopsis cotyledons and leaves. Methods Cell. Biol. 160, 181–199 (2020).
Tee, E. E., Samwald, S. & Faulkner, C. Quantification of cell-to-cell connectivity using particle bombardment. in Plasmodesmata: Methods and Protocols (eds. Benitez-Alfonso, Y. & Heinlein, M.) 263–272 https://doi.org/10.1007/978-1-0716-2132-5_17 (Springer, 2022).
Johnston, M. G. & Faulkner, C. A bootstrap approach is a superior statistical method for the comparison of non-normal data with differing variances. New Phytol. 230, 23–26 (2021).
Bourras, S. et al. The AvrPm3-Pm3 effector-NLR interactions control both race-specific resistance and host-specificity of cereal mildews on wheat. Nat. Commun. 10, 2292 (2019).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest package: tests in linear mixed effects models. J. Stat. Softw. 82, 1–26 (2017).
Acknowledgements
The authors thank all members of the Zipfel and DeFalco labs for discussion and feedback. This work was supported by a Leukemia & Lymphoma Society Award (to J.L.J. and L.C.C.); the Claudia Adams Barr Program for Cancer Research Award (to J.L.J.); the Gatsby Charitable Foundation (H.-K.A., J.D.G.J. and C.Z.); the University of Zürich (C.Z.); the Natural Science Foundation of China (32120103004 to J.-M.Z.); the National Institute of Health (grants P01-CA120964, R35-CA197588 and P01-CA117969 to L.C.C.); the Biotechnology and Biological Research Council (BB/X007685/1, BB/Y008782/1, BB/X016056/1, BBS/E/JI/230001A to C.F.); the European Research Council (under grant agreement numbers 309858 and 773153 and grants ‘PHOSPHOinnATE’ and ‘IMMUNO-PEPTALK’ to C.Z.); the Swiss National Science Foundation (grant numbers 31003A_182625 and 310030_212382 to C.Z.); the Natural Sciences and Engineering Council of Canada (PDF-532561-2019 and Discovery Grant RGPIN-2023-04222 to T.A.D.); the Canada Foundation for Innovation and Ontario Research Fund (CFI-JELF project number 44759 to T.A.D.); and Western University (Western Advancing Research Chair to T.A.D.). L.H. was funded by a Zurich-Basel Plant Science Center-Syngenta Fellowship. P.K. and M.B. were supported by the European Union’s Horizon 2020 Research and Innovation Program under Marie Skłodowska-Curie Actions (grant agreement numbers 892398 and 703954, respectively). J.D., Á.D.F.-F. and T.A.D. were supported by postdoctoral fellowships from the European Molecular Biology Organization (EMBO LTF numbers 683-2018, 580-2022 and 100-2017, respectively).
Author information
Authors and Affiliations
Contributions
C.Z. and T.A.D. conceived the project. R.T., S.C., M.L.N.V., F.S., K.W.B., C.Z. and T.A.D. designed experiments and analysed data. R.T., S.C., M.L.N.V., F.S., E.K., K.W.B. and T.A.D. performed biochemical experiments. J.L.J., T.M.Y.B. and E.M.H. performed PSPA and PSSM experiments. R.T., M.B. and T.A.D. performed CBSP filtering. R.T., J.L.J. and T.A.D. performed structural modelling. P.K. performed HEK cell assays. R.T., S.C., M.L.N.V., F.S., E.E.T. and T.A.D. performed immune assays. S.C. and Á.D.F.F. performed confocal imaging. P.D. and K.W.B. performed phosphoproteomics. R.T., S.C., M.L.N.V., F.S., E.K., A.C., P.K., L.H., Y.L., H.K.A., J.D., B.S., J.R., W.W., M.S., K.W.B. and T.A.D. generated DNA constructs and/or transgenic plant lines. J.L.J., E.M.B., J.M.Z., J.D.G.J., F.L.H.M., L.C.C., C.F., C.Z. and T.A.D. acquired research funding and supervised research. R.T., S.C., M.L.N.V., F.S. and T.A.D. generated figures. R.T., S.C., M.L.N.V., C.Z. and T.A.D. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
L.C.C. is a founder and member of the board of directors of Agios Pharmaceuticals; is a founder of and receives research support from Petra Pharmaceuticals; is listed as an inventor on a patent (WO2019232403A1, Weill Cornell Medicine) for combination therapy for phosphoinositide 3-kinase (PI3K)-associated disease or disorder and the identification of therapeutic interventions to improve response to PI3K inhibitors for cancer treatment; is a co-founder of and shareholder in Faeth Therapeutics; has equity in and consults for Cell Signaling Technologies, Volastra, Larkspur and 1 Base Pharmaceuticals; and consults for Loxo-Lilly. T.M.Y. is a co-founder of DeStroke. J.L.J. has received consulting fees from Scorpion Therapeutics and Volastra Therapeutics.
Peer review
Peer review information
Nature Plants thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 HEK293T expression of RBOHD variants and substrate-accommodating hydrophobic pocket residues of BIK1.
a, Western blot showing expression of 2xStrepII tagged RBOHD and BIK1 variants in HEK293T cells. BIK1* indicates kinase-dead BIK1. Experiment was performed twice with similar results. b, Alignment-based structural model of BIK1 (5tos) and PHK (2phk) with substrate peptide bound. Residues comprising the putative +3 Leu-binding pocket of BIK1 are highlighted. c, In vitro kinase assay showing autophosphorylation activities of BIK1 mutants. Assay was performed twice with similar results.
Extended Data Fig. 2 MCTP3 is an elicitor-induced BIK1 substrate.
a, Peptide kinase assay using RBOHD-Ser39 or MCTP3-Ser506 peptide variants as substrates for WT or kinase-dead (BIK1*) BIK1 variants; n = 3 independent replicates. Mean values are shown with error bars ±SD. Different letters indicate statistical differences (One-way ANOVA with Tukey’s multiple comparisons test). Detailed statistical parameters are reported in Supplemental Table 6. Assay was performed twice with similar results. b, Phylogenetic tree of the Arabidopsis MCTP family. Members with a predicted BIK1 motif are indicated with an asterisk. Identifiers for protein sequences used in alignment are provided in Supplementary Table 10. c, Alignment of the MCTP3-Ser506 BIK1 motif across the Arabidopsis MCTP family. d, Heatmap of MCTP expression following treatment with the indicated elicitors. Numbers indicate time in min following elicitor treatment. Data are taken from Bjornson et al.92. Values are shown as log2-fold change.
Extended Data Fig. 3 The plant CDKL family.
a, Phylogenetic tree of human and Arabidopsis CDK/CDKL families, based on alignment of kinase domains. At = Arabidopsis thaliana, Hs = Homo sapiens. b, Alignment of BIK1 motifs identified in CDKL C-termini and phylogenetic tree of the Arabidopsis CDKL family. Members with a predicted BIK1 motif are indicated with an asterisk. Numbers above alignment indicate the amino acid position of BIK1-targeted sites in CDKL5 and 6. c, Phylogeny of the kinase domains of CDKLs from 56 different species across Embryophyta. Arabidopsis CDKs are an outgroup. Identifiers for protein sequences used in alignments are provided in Supplementary Table 10. d, Heatmap of CDKL expression following treatment with the indicated elicitors. Numbers indicated time in min following elicitor treatment. Data are taken from Bjornson et al.92. Values are shown as log2-fold change. e, In vitro kinase assay showing transphosphorylation of CDKL C-termini by BIK1. GST served as a negative control. Assay was performed three times with similar results.
Extended Data Fig. 4 CDKL5 is a BIK1 substrate and negative regulator of immunity.
a, Split luciferase assay showing interaction between CDKLs and BIK1 in N. benthamiana; n = 12. Mean values are shown with error bars ±SEM. Statistical significance was tested by one-way ANOVA followed by Tukey’s HSD (different letters indicate p < 0.05). b, Western blot of protein expression of nLUC and cLUC fusion proteins. c, Co-AP of BIK1-HA with CDKL5/6-GFP in N. benthamiana. Lti6b-GFP served as a negative control. d, Pull-down assay between BIK1 and full-length CDKL5 and 6. MBP served as negative control. Open and closed arrowheads indicate the expected sizes of free MBP and MBP-CDKL5/6, respectively. Assays in a-d were performed at least twice with similar results. e, Relative peptide abundance for Ser523 phosphopeptides identified in CDKL5-GFP AP-MS experiments. Normalized phosphopeptide abundance is shown for two Ser523-containing peptide variants, with or without 1 μM flg22 for 10 min. Peptide sequence and modifications are shown above plots; p-values were determined using unpaired two-tailed t-tests; n = 3 biologically independent samples; Mean values are shown with error bars ±SEM. f, ROS production in adult Arabidopsis leaves following treatment with 100 nM elf18; n = 16 biologically independent leaf discs. g, ROS production in adult Arabidopsis leaves following treatment with 100 nM pep1; n = 24 biologically independent leaf discs. h, ROS production in adult Arabidopsis leaves following treatment with 100 nM nlp20; n = 24 biologically independent leaf discs. i, ROS production in adult Arabidopsis leaves following treatment with 100 µg/ml colloidal chitin; n = 24 biologically independent leaf discs. In panels f-i, error bars ±SEM. j, Callose deposition in adult Arabidopsis leaves 24 h after treatment with 1 µM elf18; n = 31 to 32 biologically independent leaf discs. Statistical significance was tested by one-way ANOVA followed by Tukey’s HSD (different letters indicate p < 0.05). In e-j, mean values are shown with error bars ±SEM. k, Infection assay with Pst DC3000 by infiltration; n = 8 individual plants. Arabidopsis fec mutants served as negative control. Statistical analysis was performed using a one-way ANOVA followed by Tukey’s test. Boxes show median values with 25th to 75th percentiles; error bars show min to max values. Detailed statistical parameters are reported in Supplemental Table 6. All assays in f-k were performed at least three times with similar results.
Extended Data Fig. 5 Complementation of cdkl5 cdkl6 by WT or kinase-dead CDKL5.
a, Growth phenotype of adult cdkl5 cdkl6 mutant plants. b, Western blot showing elevated PR1 protein levels in cdkl5 cdkl6 mutants. Four independent replicates are shown. c, Kinase assay showing autophosphorylation of WT or kinase-dead variants of CDKL5 or 6 (K168A or K242A, respectively). Assay was performed twice with similar results. d, Western blot showing PR1 levels in WT or kinase-dead (KD, K168A) cdkl5 cdkl6/pCDKL5::CDKL5-GFP lines. Assay was performed twice with similar results. e, Growth phenotype of adult cdkl5 cdkl6 complementation lines.
Extended Data Fig. 6 Identification of sensor NLRs as BIK1 substrates.
a, BIK1 motif distribution on candidate sNLRs. NLR domains are labeled and positions of predicted BIK1 motif phosphosites are indicated. LRR, leucine rich repeat domain; WRKY, WRKY domain. b, Predicted BIK1 motifs are conserved across Arabidopsis accessions. Genomic sequences of NLRs from different Arabidopsis accessions were extracted from 1001genomes.org and filtered for presence of the NLR sequences. Sequence logos were generated based on BIK1 motif sequence alignment of 877 (RPS4B), 560 (RPS5), 912 (RRS1), 967 (BAR1) and 693 (RPM1) accession sequences using Geneious. c, Co-AP assay showing BIK1 association with sensor NLR substrate candidates in N. benthamiana. NRG1.1 served as a negative control. d, In vitro kinase assay using 6xHis-BIK1 (WT or kinase-dead, BIK1*) and 6xHis-MBP-tagged RRS1-S-C-JID domain (labeled MBP-RRS1 in figure). e, BIK1 does not phosphorylate NRG1.1 in vitro. In vitro kinase assay using 6xHis-BIK1 and 6xHis-MBP-NRG1.1-CC domain. 6xHis-MBP-tagged RPS5-CC domain was used as a positive control. All proteins were tested and blotted on the same membrane but are displayed as separate images for visualization purposes. f and g, BIK1 phosphorylates RPS4B (f) and RPS5 (g) in vitro. In vitro kinase assay using 6xHis-BIK1 (WT or kinase-dead, BIK1*) and 6xHis-MBP-tagged RPS4B-NBARC domain (f, labeled MBP-RPS4B in figure) or 6xHis-MBP-tagged RPS5-CC domain (g, labeled MBP-RPS5 in figure). In d-g, 6xHis-MBP-MBP was used as a negative control (labeled MBP-MBP in figure). h, Summary of phosphorylated residues identified on 6xHis-MBP-RPS4B-NBARC in vitro. Boldface indicates phosphorylated residues. i, Summary of phosphorylated residues identified on 6xHis-MBP-RPS5-CC in vitro. Boldface indicates phosphorylated residues. Data from three independent replicates are shown. All experiments in a-g were conducted at least three times with similar results.
Extended Data Fig. 7 Expression of RPS5 and RPS4B variants in N. benthamiana and complementation lines.
a and b, Western blots showing protein expression level corresponding to Fig. 4e (a) or Fig. 4f (b). The experiments were conducted three times with similar results. c, Electrolyte leakage measurement 12 hpi of Pst DC3000D36E(AvrPphB) into 5-week-old Arabidopsis lines. Statistical significance was tested by one-way ANOVA followed by Tukey’s HSD (different letters indicate p < 0.05). Mean values are shown with error bars ±SEM. N = 4 biologically independent independent plants. d, Western blot showing protein expression level of transgenic lines corresponding to Fig. 4g. e, Electrolyte leakage measurement 24 hpi Pst DC3000D36E(AvrRps4) into 5-week-old Arabidopsis lines. Statistical significance was tested by one-way ANOVA followed by Tukey’s HSD (different letters indicate p < 0.05). Mean values are shown with error bars ±SEM. N = 4 biologically independent plants. f, Western blot showing protein expression level of transgenic lines corresponding to Fig. 4h. g, Growth phenotype of RPS4B complementation lines. Plants were photographed 7 weeks after germination. Plants were selected by Basta spraying. Experiments were conducted more than three times with similar results. h, Western blot showing PR1 accumulation in 5-week-old plants. For each sample, 4 leaf discs were collected from 2 leaves from 2 independent plants each. Detailed statistical parameters are reported in Supplemental Table 6. Experiments were conducted three times with similar results.
Extended Data Fig. 8 RRS1 and BAR1 complementation lines.
a, Cell death phenotypes after infiltration of Pst DC3000D36E(AvrRps4). Images show leaves 1 dpi. Numbers indicate the number of leaves with cell death relative to the total number of infiltrated leaves. b, Pst DC3000 expressing AvrRps4 was sprayed on 5-week-old RRS1-S mutant complementation lines and bacterial colonies were counted at 3 dpi. c, Protein expression level of RRS1 complementation lines. d, Pst DC3000 expressing HopB1d was sprayed on five-week-old RRS1-S complementation lines and bacterial colonies were counted at 3 dpi. In b and d, statistical significance was tested by one-way ANOVA followed by Tukey’s HSD (different letters indicate p < 0.05). Mean values are shown with error bars ±SEM; n = 3 biologically-independent plants. e, Western blot showing protein expression level of BAR1 mutant complementation lines. Detailed statistical parameters are reported in Supplemental Table 6. All experiments were conducted at least three times with similar results.
Extended Data Fig. 9 Effect of phosphorylation on NLR subcellular localization and interactions.
a, Confocal microscopy (GFP and bright field) 3 dpi with Agrobacterium constructs as indicated. For RPS4B-GFP, RRS1B-FLAG was co-expressed at 1:1 ratio due to low expression level when expressed alone. All constructs were co-infiltrated with a P19 construct. Scale bar represents 20 µm. b, Co-AP assay showing association between PBS1-HA and RPS5-MYC variants in N. benthamiana. G2Citrine-HA was used as a negative control. c, Co-AP showing association between RRS1B-FLAG and RPS4B-MYC in N. benthamiana. RPS4-MYC was used as negative control. All experiments were conducted at least three times with similar results. d, AvrRps4 does not trigger RPS4/RRS1 and RPS4B/RRS1B oligomerization in N. benthamiana. All experiments were conducted at least three times with similar results.
Extended Data Fig. 10 flg22-induced dissociation of RPS4B and BIK1 in Arabidopsis.
Co-AP using 3-week-old transgenic Arabidopsis seedlings with or without 1 µM flg22 treatment (10 min). The NRG1.1-FLAG line was used as negative control. Experiment was repeated three times with similar results.
Supplementary information
Supplementary Table 1 (download XLSX )
The CBSP list.
Supplementary Table 2 (download XLSX )
Genotypes used in this study.
Supplementary Table 3 (download XLSX )
CDKL5-GFP AP-MS phosphopeptide abundance.
Supplementary Table 4 (download XLSX )
Arabidopsis NLRs with BIK1 motif scores >2.5.
Supplementary Table 5 (download XLSX )
Primers used in this study.
Supplementary Table 6 (download XLSX )
BIK1 PSPA data and PSSM.
Supplementary Table 7 (download XLSX )
Peptides used in this study.
Supplementary Table 8 (download XLSX )
Peptide-spectrum match data for RPS4B and RPS5 LC–MS/MS.
Supplementary Table 9 (download XLSX )
Statistical analyses.
Supplementary Table 10 (download XLSX )
Identifiers for proteins used in sequence alignments.
Source data
Source Data Figs. 1–4 and Extended Data Figs. 1 and 3–10 (download PDF )
Unprocessed western blots and/or gels.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Toth, R., Choi, S., Le Naour--Vernet, M. et al. Motif-based substrate mapping of the receptor-like cytoplasmic kinase BIK1 reveals novel components and regulatory nodes of plant immunity. Nat. Plants 12, 465–480 (2026). https://doi.org/10.1038/s41477-025-02218-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-025-02218-z