Abstract
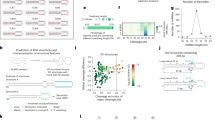
Small RNAs, including microRNA, small interfering RNA (siRNA) and PIWI-interacting RNA, are regulatory RNAs that play critical roles in gene regulation, development, viral defence and environmental response1. The biogenesis of microRNA and siRNA relies on the Dicer family ribonucleases to capture, measure and cleave their double-stranded RNA substrates2,3. In Arabidopsis, DICER-LIKE 4 (DCL4) produces 21-nucleotide siRNA in association with Double-Stranded RNA-Binding Protein 4 (DRB4) for post-transcriptional gene silencing4,5,6,7,8,9,10,11. Here we determined the structures of the DCL4–RNA complex in a dicing-competent conformation and the DCL4–DRB4–RNA complex in a pre-dicing conformation. DCL4 measures 21 nucleotides along RNA between its PAZ and RNase III domains to determine the product siRNA length. A DCL4-specific loop locates the second double-stranded RNA binding domain of DCL4 and DRB4 to a distal position of the substrate RNA, yielding a preference for long RNA substrates. Our studies demonstrate the molecular basis of substrate recognition, length measurement and long RNA preference by the DCL4–DRB4 complex for 21-nucleotide siRNA biogenesis in plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The structures have been deposited in the PDB with the accession codes 9KBY and 9KBZ. The cryo-EM maps have been deposited in the Electron Microscopy Data Bank with accession codes EMD-62234 and EMD-62235. Source data are provided with this paper.
References
Yu, Y., Zhang, Y., Chen, X. & Chen, Y. Plant noncoding RNAs: hidden players in development and stress responses. Annu. Rev. Cell Dev. Biol. 35, 407–431 (2019).
Vergani-Junior, C. A., Tonon-da-Silva, G., Inan, M. D. & Mori, M. A. DICER: structure, function, and regulation. Biophys. Rev. 13, 1081–1090 (2021).
Xu, Y. & Chen, X. microRNA biogenesis and stabilization in plants. Fundam. Res. 3, 707–717 (2023).
Fukudome, A. & Fukuhara, T. Plant dicer-like proteins: double-stranded RNA-cleaving enzymes for small RNA biogenesis. J. Plant Res. 130, 33–44 (2017).
Xie, Z., Allen, E., Wilken, A. & Carrington, J. C. DICER-LIKE 4 functions in trans-acting small interfering RNA biogenesis and vegetative phase change in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 102, 12984–12989 (2005).
Gasciolli, V., Mallory, A. C., Bartel, D. P. & Vaucheret, H. Partially redundant functions of Arabidopsis DICER-like enzymes and a role for DCL4 in producing trans-acting siRNAs. Curr. Biol. 15, 1494–1500 (2005).
Dunoyer, P., Himber, C. & Voinnet, O. DICER-LIKE 4 is required for RNA interference and produces the 21-nucleotide small interfering RNA component of the plant cell-to-cell silencing signal. Nat. Genet. 37, 1356–1360 (2005).
Yoshikawa, M., Peragine, A., Park, M. Y. & Poethig, R. S. A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis. Genes Dev. 19, 2164–2175 (2005).
Nakazawa, Y., Hiraguri, A., Moriyama, H. & Fukuhara, T. The dsRNA-binding protein DRB4 interacts with the Dicer-like protein DCL4 in vivo and functions in the trans-acting siRNA pathway. Plant Mol. Biol. 63, 777–785 (2007).
Hiraguri, A. et al. Specific interactions between Dicer-like proteins and HYL1/DRB-family dsRNA-binding proteins in Arabidopsis thaliana. Plant Mol. Biol. 57, 173–188 (2005).
Adenot, X. et al. DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr. Biol. 16, 927–932 (2006).
Yang, Z. et al. Crop antiviral defense: past and future perspective. Sci. China Life Sci. 67, 2617–2634 (2024).
Blevins, T. et al. Identification of Pol IV and RDR2-dependent precursors of 24 nt siRNAs guiding de novo DNA methylation in Arabidopsis. eLife 4, e09591 (2015).
Wang, Q. et al. Mechanism of siRNA production by a plant Dicer–RNA complex in dicing-competent conformation. Science 374, 1152–1157 (2021).
Zhai, J. et al. A one precursor one siRNA model for Pol IV-dependent siRNA biogenesis. Cell 163, 445–455 (2015).
Singh, J., Mishra, V., Wang, F., Huang, H. Y. & Pikaard, C. S. Reaction mechanisms of Pol IV, RDR2, and DCL3 drive RNA channeling in the siRNA-directed DNA methylation pathway. Mol. Cell 75, 576–589.e5 (2019).
Xie, G., Du, X., Hu, H. & Du, J. Molecular mechanisms underlying the establishment, maintenance, and removal of DNA methylation in plants. Annu. Rev. Plant Biol. 76, 143–170 (2025).
Cao, M. et al. Virus infection triggers widespread silencing of host genes by a distinct class of endogenous siRNAs in Arabidopsis. Proc. Natl Acad. Sci. USA 111, 14613–14618 (2014).
Du, X. The cellular RNA-dependent RNA polymerases in plants. New Phytol. 244, 2150–2155 (2024).
Kumakura, N. et al. SGS3 and RDR6 interact and colocalize in cytoplasmic SGS3/RDR6-bodies. FEBS Lett. 583, 1261–1266 (2009).
Nagano, H., Fukudome, A., Hiraguri, A., Moriyama, H. & Fukuhara, T. Distinct substrate specificities of Arabidopsis DCL3 and DCL4. Nucleic Acids Res. 42, 1845–1856 (2014).
Fukudome, A. et al. Specific requirement of DRB4, a dsRNA-binding protein, for the in vitro dsRNA-cleaving activity of Arabidopsis Dicer-like 4. RNA 17, 750–760 (2011).
Kakiyama, S., Tabara, M., Nishibori, Y., Moriyama, H. & Fukuhara, T. Long DCL4-substrate dsRNAs efficiently induce RNA interference in plant cells. Sci. Rep. 9, 6920 (2019).
Zhan, J. & Meyers, B. C. Plant small RNAs: their biogenesis, regulatory roles, and functions. Annu. Rev. Plant Biol. 74, 21–51 (2023).
Allen, E., Xie, Z., Gustafson, A. M. & Carrington, J. C. MicroRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121, 207–221 (2005).
Chiliveri, S. C., Aute, R., Rai, U. & Deshmukh, M. V. DRB4 dsRBD1 drives dsRNA recognition in Arabidopsis thaliana tasi/siRNA pathway. Nucleic Acids Res. 45, 8551–8563 (2017).
Liu, Z. et al. Cryo-EM structure of human Dicer and its complexes with a pre-miRNA substrate. Cell 173, 1191–1203.e12 (2018).
Jouravleva, K. et al. Structural basis of microRNA biogenesis by Dicer-1 and its partner protein Loqs-PB. Mol. Cell 82, 4049–4063.e6 (2022).
Su, S. et al. Structural insights into dsRNA processing by Drosophila Dicer-2–Loqs-PD. Nature 607, 399–406 (2022).
Zapletal, D. et al. Structural and functional basis of mammalian microRNA biogenesis by Dicer. Mol. Cell 82, 4064–4079.e13 (2022).
Yamaguchi, S. et al. Structure of the Dicer-2–R2D2 heterodimer bound to a small RNA duplex. Nature 607, 393–398 (2022).
Wei, X. et al. Structural basis of microRNA processing by Dicer-like 1. Nat. Plants 7, 1389–1396 (2021).
Lee, Y. Y., Lee, H., Kim, H., Kim, V. N. & Roh, S. H. Structure of the human DICER–pre-miRNA complex in a dicing state. Nature 615, 331–338 (2023).
Liu, Y. et al. Molecular basis of plant DCL4 action that outcompetes DCL2. Nat. Plants https://doi.org/10.1038/s41477-026-02243-6 (2026).
Curaba, J. & Chen, X. Biochemical activities of Arabidopsis RNA-dependent RNA polymerase 6. J. Biol. Chem. 283, 3059–3066 (2008).
Rajeswaran, R. et al. Sequencing of RDR6-dependent double-stranded RNAs reveals novel features of plant siRNA biogenesis. Nucleic Acids Res. 40, 6241–6254 (2012).
Bouché, N., Lauressergues, D., Gasciolli, V. & Vaucheret, H. An antagonistic function for Arabidopsis DCL2 in development and a new function for DCL4 in generating viral siRNAs. EMBO J. 25, 3347–3356 (2006).
Creasey, K. M. et al. miRNAs trigger widespread epigenetically activated siRNAs from transposons in Arabidopsis. Nature 508, 411–415 (2014).
Zhang, X. et al. Suppression of endogenous gene silencing by bidirectional cytoplasmic RNA decay in Arabidopsis. Science 348, 120–123 (2015).
Martínez de Alba, A. E. et al. In plants, decapping prevents RDR6-dependent production of small interfering RNAs from endogenous mRNAs. Nucleic Acids Res. 43, 2902–2913 (2015).
Wang, Q. et al. A SYBR Gold-based label-free in vitro dicing assay. Bio Protoc. 12, e4382 (2022).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Qin, H. et al. Structure of the Arabidopsis thaliana DCL4 DUF283 domain reveals a noncanonical double-stranded RNA-binding fold for protein–protein interaction. RNA 16, 474–481 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Acknowledgements
We thank the staff members at SUSTech Cryo-EM Center and PKU-IAAS Cryo-EM Center for cryo-EM data collection. This work was supported by the National Natural Science Foundation of China (grant nos 92581101 to J.D., 32325008 to J.D., 32261160572 to H.G. and 32301080 to Q.W.), the Guangdong Basic and Applied Basic Research Foundation (grant no. 2023B1515120048 to J.D.), the Shenzhen Science and Technology Program (grant nos RCJC20221008092720004 to J.D., 20231120201445001 to J.D. and ZDSYS20230626091659010 to H.G. and J.D.) and the New Cornerstone Science Foundation (grant no. NCI202235 to H.G.). J.D. is an investigator at the SUSTech Institute for Biological Electron Microscopy.
Author information
Authors and Affiliations
Contributions
C.W., C.C., J. Zhao and Q.W. performed the structural studies and biochemical assays. N.W., Z.Z., K.J., Y. Li and Y.X. contributed to data analysis and discussion. Y. Liu, P.W., J. Zhai and H.G. provided functional insights. J.D. supervised the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Sounghun Roh, Xiaobin Wei and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Protein purification and in vitro activity assay.
a, The purification of Arabidopsis DCL4 (left) and DCL4-DRB4 complex (right). b, In vitro dicing assay of DCL4 and DCL4-DRB4 complex showing that DCL4-DRB4 has a stronger activity than DCL4 alone. The in vitro assay was repeated at least three times independently with similar results.
Extended Data Fig. 2 Cryo-EM image processing workflow of DCL4-hairpin RNA complex.
a, A representative cryo-EM image of DCL4-hairpin RNA complex. b, Flowchart of cryo-EM data processing. c, Typical 2D class averages of DCL4-dsRNA complex. d, Gold-standard Fourier shell correlation (GSFSC) of the final map of DCL4-hairpin RNA complex in dicing state. e, Angular distribution heat map calculated in cryoSPARC for particle projections contributing to the local resolution of DCL4-hairpin RNA complex.
Extended Data Fig. 3 The comparison of DCL4 and DCL3 reveals potential molecular mechanisms for measuring RNA length by Dicers.
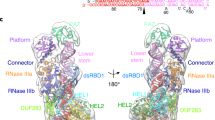
a, Superimpositions of DCL4 (colored) and DCL3 (PDB code: 7VG2, in silver) based on the positions of their RIIIa/b domains. The directions of the conformational changes of platform-PAZ-connector cassette around the RNA duplex from DCL4 to DCL3 are shown by arrows. The red and black boxes denote the guide strand 5′-end and complementary strand 3′-end recognition regions, which are detailed in panel b-c and d-e, respectively. b-c, Recognition of the guide strand 5′-end first nucleotides by DCL4 (b) and DCL3 (c). Interacting residues and hydrogen bonds are highlighted in sticks and dashed lines, respectively. d-e, Recognition of 3′-end of the complementary strand by DCL4 (d) and DCL3 (e). f, The dsRBD1 domain uses a positively charged surface to bind to the RNA backbone. g, Structural comparison of DCL4-DRB4-RNA complex (colored) and DCL3-RNA complex (PDB code: 7VG2, in sliver) shows the different orientations of their second dsRBDs.
Extended Data Fig. 4 Multiple sequence alignment of the Platform-PAZ domains of DCLs.
Structure-based sequence alignment of the Platform-PAZ domains of DCL4 from multiple species, along with DCL1, DCL2, and DCL3. The key residues involved in the recognition of the 5′-end of the guide strand RNA, including the charged interactions with the phosphate group and the stacking interaction with the first base, are all conserved in DCL4 and are highlighted by red triangles. The secondary structure diagram of DCL4 is shown on top.
Extended Data Fig. 5 Multiple sequence alignment of the RNase IIIa, and RNase IIIb of DCLs.
Structure-based sequence alignment of DCL4 from multiple species, along with DCL1, DCL2, and DCL3, reveals that the R1135, Y1339, and S1343 of Arabidopsis DCL4, which are important residues of the RIII domains involved in hydrogen bonding interaction with the LinkerRBDs, are conserved in the DCL4 proteins. The secondary structure diagram of DCL4 is shown on top.
Extended Data Fig. 6 Multiple sequence alignment of the dsRBDs and LinkerRBDs of DCLs.
Structure-based sequence alignment of DCL4 from multiple species, along with DCL1, DCL2, and DCL3, reveals that only the DCL4 clade DCL proteins have the LinkerRBDs long enough to extend dsRBD2 to a distal position on RNA. The secondary structure diagram for DCL4 is shown on top.
Extended Data Fig. 7 Cryo-EM image processing workflow of DCL4-DRB4CTD-dsRNA complex.
a, A representative cryo-EM image of DCL4-DRB4CTD-dsRNA complex. b, Flowchart of cryo-EM data processing. c, 2D class averages. d, Gold-standard Fourier shell correlation (GSFSC) of the final map. e, Angular distribution heat map calculated in cryoSPARC for particle projections contributing to the local resolution. f, Cryo-EM maps showing the fitting of the representative protein and RNA regions.
Extended Data Fig. 8 A working model for the long RNA processing and 21-nt siRNA biogenesis by the DCL4-DRB4 complex.
a, A proposed model illustrating DRB4’s role in assisting DCL4 with long dsRNA processing. The DRB4CTD-DCL4dsRBD2 interaction may position the DRB4dsRBD1/2 (modeled from PDB: 2N3F) to a distal position of the dsRNA, achieving the long RNA substrate preference of the DCL4-DRB4 complex. b, A working model of the long RNA processing by the DCL4-DRB4 complex to produce phasiRNA by continuously dicing. The red arrow and stars indicate the direction of RNA translocation and the cleavage site, respectively. The DRB4dsRBD1/2 may help DCL4 search and capture long RNA substrates. Once the RNA is bound by the DCL4-DRB4 complex, DCL4 helicase domain could pump the RNA towards the platform-PAZ domain cassette, resulting in RNA bending. Upon the platform-PAZ domain cassette capturing the RNA termini, the pumping may push the RNA to further bend to approach the catalytic center of DCL4 RIIIs for cleavage. After cleavage, the 21-nt siRNA duplex is released, and the remaining RNA fragment, which is held by the helicase domain, could further be pumped by the helicase to start the next dicing cycle.
Extended Data Fig. 9 In vitro dicing assay of DCL4 and DCL4-DRB4 complex towards RNA substrates with different lengths.
a, The sequences of three RNA duplexes used in this assay. b, In vitro dicing assay of DCL4 and DCL4-DRB4 complex to process RNAs with different lengths (30/32-nt, 55/57-nt, and 107/109-nt). The schematic view of the RNA length is shown on the right. The cutting sites on RNA are denoted by triangles. The in vitro assay was repeated at least three times independently with similar results.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1.
Source data
Source Data Fig. 1h (download PDF )
Unmodified gels for the three repeats of Fig. 1h.
Source Data Extended Data Fig. 1 (download PDF )
Unmodified gels for Extended Data Fig. 1a and four repeats of Extended Data Fig. 1b.
Source Data Extended Data Fig. 9b (download PDF )
Unmodified gels for the three repeats of Extended Data Fig. 9b.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, C., Chi, C., Liu, Y. et al. Molecular basis of DRB4-assisted long RNA processing and 21-nucleotide siRNA biogenesis by DCL4 in plants. Nat. Plants 12, 512–519 (2026). https://doi.org/10.1038/s41477-026-02236-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41477-026-02236-5