Abstract
Parkinson’s Disease (PD) is characterized by the loss of dopaminergic neurons in the substantia nigra pars compacta (SNpc), which is associated with changes in microglia function. While age remains the biggest risk factor, the underlying molecular cause of PD onset and its concurrent neuroinflammation are not well understood. Many identified PD risk genes have been directly linked to dopamine neuron impairment, while others are linked to immune cell function. In this study, we found that the PD risk gene FAM49B is critically expressed in microglia of the human SNpc and is downregulated with age and PD. We utilized human and murine microglia cells to demonstrate the role of FAM49B in regulating fundamental microglial functions such as cytoskeletal maintenance, migration, surface adherence, energy homeostasis, autophagy, and, importantly, inflammatory response. Downregulation of microglial FAM49B, as observed in the SNpc of aging individuals, led to significant alterations in these cellular functions, which are associated with increased microglial activation. Thus, our study highlights novel cell-type-specific roles of FAM49B and provides a potential mechanism for susceptibility to neuroinflammation, and reactive gliosis observed in both PD and normal aging.
Similar content being viewed by others
Introduction
Parkinson’s disease (PD) is the second most common neurodegenerative disorder, primarily characterized by the progressive degeneration of dopaminergic (DA) neurons in the substantia nigra pars compacta (SNpc), which leads to characteristic motor impairments1,2. Aging is the most relevant risk factor for PD, with its incidence increasing strongly after the age of 603. While rare monogenic forms of PD are caused by mutations in genes that impair DA neurons, such as SNCA, PRKN, PINK1, LRRK2, and PARK74, genes identified in genome-wide association studies (GWAS) suggest that cells of both the adaptive and innate immune system also play crucial roles in PD pathogenesis5,6,7,8,9. Accordingly, neuroinflammation, including microgliosis, has been described in PD for many decades10. A causal relationship between glial activity and PD has been proposed such that inflammaging, a low-grade inflammation resulting from an imbalance between pro- and anti-inflammatory signals and cellular senescence, can drive DA neuron loss in the SNpc11,12,13. Accordingly, recent studies indicate that microglial activation in PD is not a secondary response to neuronal loss but may actively contribute to neurodegeneration as a primary trigger. For example, α-synuclein, a protein central to PD pathology, can be taken up by microglia, triggering a TLR-Fyn-PKCδ-NFκB-mediated inflammatory response that exacerbates neuronal damage14,15.
While not completely understood, it has been shown that microglial phenotypes vary across brain regions and can shift in response to aging and disease states16. Relevant to PD, microglia in the vulnerable SNpc exhibit a transcriptional profile distinct from those in other CNS regions, with a predisposition toward a pro-inflammatory state17,18. This regional specificity has been proposed to contribute to the selective vulnerability of SNpc DA neurons in PD19. Interestingly, due to their close interaction, inflammatory signals from microglia can drive astrocytes toward a neurotoxic phenotype, further amplifying neuronal damage20.
Given the growing evidence linking microglial dysfunction to PD, targeting neuroinflammation has emerged as a potential therapeutic strategy for the disease. Anti-inflammatory treatments, such as PKCδ and NFκB inhibitors, have shown promise in preclinical PD models by reducing microglial activation and slowing disease progression15,21. Further understanding of the complex interplay between aging, microglia, neuroinflammation, and PD pathology may provide new insights into disease mechanisms and therapeutic targets for the disease. In the present study, we investigated the genetic risk factor FAM49B, which was previously identified by a GWAS22, and found that it regulates microglial activation and may contribute to processes associated with inflammaging in PD.
FAM49B (family with sequence similarity 49 member B; also known as CYRIB (CYFIP-Related Rac1 Interactor B)) has been characterized as a mitochondria-localized protein regulating mitochondrial function and cancer progression23 and inhibiting T cell activation by repressing Rac activity and modulating cytoskeleton reorganization24. While FAM49B is critical for T cell function25 and has been suggested to be a marker of homeostatic microglia26, neither its cellular function in microglia cells nor its role in brain inflammaging has been evaluated.
In brief, using computational analysis based on human brain gene expression data, we identified an age-related downregulation of FAM49B, specifically in microglia in the human SNpc. To further evaluate whether this age-related downregulation led to increased immunological response and microgliosis, we conducted functional experiments using microglial cell lines with FAM49B knockdown/knockout. Interestingly, the reduction of FAM49B induced microglia-relevant reactivity and significant metabolic changes, including dysfunctional oxidative phosphorylation and glycolysis, as well as modifications in cytoskeletal dynamics, migratory velocity, and adhesion capacity. Importantly, compared to controls, FAM49B-deficient microglia showed noticeably higher immunological activation. Our results imply that FAM49B is essential for regulating microglial immune responses. This informs the hypothesis that age-related downregulation of FAM49B in the SNpc may predispose microglia to an overactive gliosis phenotype, aggravating dopaminergic neuron loss and accelerating PD.
Crucially, the elevated immunological activity that we observed in our FAM49B-deficient microglial cells was successfully decreased by treatment with anti-inflammatory NFκB inhibitor treatment. Based on our results, FAM49B is a critical modulator of microglial immunological response. Consequently, its age-related downregulation could contribute to inflammaging and the subsequent pathophysiology of PD. Finally, anti-inflammatory treatment to target this pathway may present a viable strategy for reducing neuroinflammation and preserving dopaminergic neurons in PD.
Results
Downregulation of FAM49B in microglia with age in the human substantia nigra
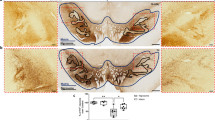
FAM49B has been identified as a genetic risk factor for PD27 (Fig. S1b); however, neither its role in PD pathology nor in which relevant cell type it is expressed is known. To investigate the cellular context of FAM49B expression, we analyzed bulk RNA sequencing data from 114 human SNpc samples obtained from the Genotype-Tissue Expression (GTEx) Project28. Cell-type proportions for each sample were computationally inferred by correlating bulk expression profiles with reference signatures of major brain cell types, including astrocytes, microglia, neurons, endothelial cells, and oligodendrocytes (see Methods). This analysis revealed that FAM49B expression was most significantly correlated with the estimated proportion of microglia (Wald test, p = 4.1 × 10−7), suggesting that FAM49B is primarily expressed in these resident immune cells of the SNpc (Fig. 1a, other cell types shown in Fig. S1a). Additionally, assessing further data sets from other brain areas confirmed that microglial FAM49B is downregulated with age in multiple brain regions (Fig. S1c). These findings were further supported by single-cell sequencing data from human SNpc tissue, which confirmed that FAM49B expression is highest in microglia (Fig. 1b, GSE14023129). Moreover, after correcting for potential confounders such as cell-type composition, we observed a significant negative correlation between FAM49B expression and donor age, indicating an age-associated downregulation of FAM49B in microglia (Wald test, p = 3.0 × 10−4, Fig. 1c). Since FAM49B was identified as PD risk factor, we sought to determine if FAM49B downregulation also occurs in patient microglia. To test this, we assessed a previously published single nuclei RNA-seq of human post-mortem in SNpc tissue from 15 sporadic PD cases and 14 age-matched controls30 and found a significant downregulation of FAM49B in the nigral microglia of PD cases (p = 3.1 × 10−8 and FDR 6.73 × 10−7, rank-based Wilcoxon test). In summary, our integrative transcriptomic analysis indicates that FAM49B is predominantly expressed in microglia within the human SNpc and is significantly downregulated with aging and in PD. To assess whether microglial Fam49b levels are also downregulated in the SNpc in aging mice, we analyzed the Fam49b expression in the SNpc of young (5 months) and middle-aged mice (13 months), which, based on reported data31, resembles the human age groups in Fig. 1c (young: <40, middle-aged: 40–60). We found a significant downregulation of Fam49b in the nigral microglia with advanced age (Fig. 1d, e).
a Correlation between FAM49B normalized expression and inferred relative microglia cell-type proportions across 114 human SNpc samples from the GTEx Project. FAM49B expression is most significantly associated with microglial abundance (p = 4.1 × 10⁻⁷, Wald test), consistent with microglia as the primary cell type expressing FAM49B in the SNpc. b Single-cell RNA sequencing data from human SNpc tissue (GSE140231) confirm that FAM49B expression is highest in microglia (SN = SNpc). c FAM49B expression negatively correlates with donor age in GTEx SNpc samples (p = 3.0 × 10⁻⁴, Wald test), indicating age-associated downregulation in microglia. d Bar graph shows fluorescence-based (CTCF) quantification of Fam49b expression levels in nigral microglia in mice (Nyoung = 3, nyoung = 110; Nold = 2, nold = 31; unpaired Mann–Whitney test: *p < 0.05). e Representative confocal images of young (5 months) and aged (13 months) midbrains are shown. Th: Tyrosine hydroxylase (DA neurons), Iba1: Allograft inflammatory factor 1 (microglia). f Bar graph of quantification of normalized counts showing decreased FAM49B expression in senescent (BrdU-treated) HMC3 cells (expression data from ref. 33) (Data was analyzed using an unpaired t-test. **p < 0.01, N = 3, 3).
One hallmark of brain aging is the accumulation of senescent cells, including senescent glial cells, which contribute to neuroinflammation and inflammaging in the midbrain. Pro-inflammatory senescent brain cells have also been associated with PD12,32. Hence, we sought to determine if senescence in human microglia could alter the expression of FAM49B and utilized our previously reported DNA damage-induced senescent HMC3 cells as an in vitro model of senescence33. The senescence phenotype was validated by a battery of molecular methods (see detailed characterization:33). Importantly, these senescent microglia showed significant downregulation of FAM49B (Fig. 1f), indicating a possible role of senescence in the inflammaging phenotype of SNpc microglia.
Reduced expression of FAM49B in microglia results in metabolic changes
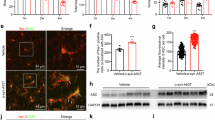
To determine the function of FAM49B in human microglia, we employed the commonly used human HMC3 cell line34 and generated a CRISPR/Cas9-mediated knockout clone. After confirming the knockout, bulk RNA sequencing was performed to compare the gene expression of wild-type (WT) and knockout (KO) HMC3 cells (Fig. 2a). Assessment of the gene expression profiles revealed significant global expression changes (Fig. 2b) and KEGG pathway analysis revealed several interesting hits, including PD (top hit), Alzheimer’s, Huntington’s, ALS, but also oxidative phosphorylation, cellular senescence, and endocytosis (Fig. S2h, i). Interestingly, contributing to the identification of the KEGG pathway oxidative phosphorylation, substantial alterations in mitochondrial genes have been observed (Figs. 2c, S2a, b). These changes are of particular interest since FAM49B has recently been reported to be involved in the regulation of mitochondrial function and integrity in pancreatic ductal adenocarcinoma cells23. However, the role of FAM49B in microglia in the context of PD has yet to be characterized. Thus, we applied diverse methods to investigate the metabolic changes that occur in FAM49B-KO microglia. A Seahorse real-time cell metabolic analysis (XF Mito Stress Test) revealed that the KO of FAM49B in HMC3 microglia caused a significant decrease in basal and maximal respiration, ATP production, and spare respiratory capacity (Fig. 2d–h) while not affecting non-mitochondrial oxygen consumption and coupling efficiency (Fig. S2c, d). In addition, the overall energy production was impaired as suggested by a significant change in glycolysis gene expression (Fig. S2g). To test whether glycolysis function was impaired, we performed a 2-deoxyglucose assay, which leads to an increase in cell death if cells utilize glycolysis. Interestingly, the FAM49B-KO microglia cells were less susceptible as compared to the WT microglia (which partially depend on glycolysis35) in this assay, further suggesting a partial impairment of the glycolytic pathway (Fig. 2i). To confirm these findings in an additional microglia model, mouse BV2 cells were used. An siRNA-mediated knockdown (KD) of Fam49b (Figs. 2j, S2k) significantly reduced the membrane potential of mitochondria in these cells, while the overall organelle number was not affected (Figs. 2k, S2e). Using immunogold labeling in combination with transmission electron microscopy revealed that Fam49b localized to mitochondria (~50% of all Fam49b molecules were localized to mitochondria). Interestingly, triggering an immune response in BV2 cells by lipopolysaccharide (LPS) treatment resulted in a reduction of Fam49b globally and specifically in mitochondria (Figs. 2l, S2k). We also found a significant upregulation of free radicals produced in Fam49b-KD cells (Fig. S2f), an additional indication of mitochondrial impairment.
a Western blot analysis confirming ablation of FAM49B in HMC3 cells (n = 6, 6). b Volcano plot shows global gene expression changes in WT vs. FAM49B-KO HMC3 cells (numbers indicate significantly up- or downregulated genes). c Heatmap of oxidative phosphorylation genes significantly altered in FAM49B-KO cells. d Seahorse graph showing a significant reduction in oxidative phosphorylation in FAM49B-KO cells (n = 18 for each condition, Student’s t-test per time point, **p < 0.01, ***p < 0.001. Data are presented as mean ± SEM). Bar graphs show a significant reduction in basal respiration (e), maximal respiration (f), ATP production (g), and spare respiratory capacity (h). (n = 18 for each condition, Student’s t-test per time point, **p < 0.01, ***p < 0.001. Data are presented as mean ± SEM. i Bar graph shows cell viability in HMC3 WT and FAM49BKO cells treated with 2-Deoxyglucose. (n = 4 (WT)/4 (KO)), **p < 0.01, 2-way ANOVA). j Western blot analysis confirming reduction of Fam49b in BV2 cells by siRNA (n = 3, 3). k Representative confocal images of BV2 cells stained with functional dye MitoTracker Red CMXRos 48 h post siRNA transfection. Analysis shows reduced mitochondrial membrane potential measured by average fluorescence intensity (n, ctrl = 348, siFam49b = 598). l Representative TEM images using immunogold labeling for Fam49b in BV2 cells. Bar graph shows quantification of the percentage of Fam49b located in mitochondria under control or LPS treatment (N = 6, 4). Scale bar 500 nm. Data are presented as mean ± SEM. Data for (a, e–j, and l were analyzed using an unpaired Mann–Whitney test. Data for (k) was analyzed using an unpaired t-test with Welch’s correction. *p < 0.05, **p < 0.01, ***p < 0.001.
In summary, the reduction of FAM49B significantly impaired the metabolism of both mouse and human microglia cells. Interestingly, this metabolic change partially resembles microglia that face pro-inflammatory stimuli and prepare to respond36.
FAM49B-KO and KD microglia show alterations in cytoskeletal dynamics and migration
Microglia exhibit significant mobility and are considered the most dynamic cells of the healthy mature brain37,38. They are constantly active and survey their environment by extending and retracting their processes to detect changes, such as neuronal damage, infection, or inflammation. When responding to inflammatory cues, microglia can adopt an ameboid morphology, increase migratory and endocytic capacities, and move toward areas of damage or pathology37. Microglia are responsible for the elimination of microbes, dead cells/cell debris, excessive synapses, protein aggregates, and other antigens that may endanger the CNS39. Recently, it has been reported that microglia can use tunneling nanotubes to interact with other microglia or neurons to protect them from aggregate-induced cellular dysfunction and death40. In order to complete these tasks, cytoskeletal remodeling is essential, as microglia have to extend their processes, or nanotubes, and migrate toward the cellular debris to phagocytose it41. Given these critical functions and the previously published findings that FAM49B is a regulator of actin dynamics24 and directly interacts with activated Rac Family Small GTPase 1 (RAC1), known to regulate cytoskeleton organization42, we investigated the impact of FAM49B loss on cytoskeletal dynamics and migration in our microglial cell lines. Indeed, gene expression profiling of genes related to cytoskeletal dynamics revealed striking rearrangements, indicating an effect on cell migration, adhesion, and other cytoskeletal functions, such as endocytosis (Figs. 3a, S3a, c). Interestingly, while WT microglia displayed a ramified morphology consistent with a surveillant state, FAM49B-KO induced a transition to an ameboid (Fig. 3b), reactive phenotype, additionally marked by upregulation of IL-8 (Fig. 5b). In line with this finding, it was previously reported that FAM49B-KO in human melanoma cells induced a round shape and increased migration speed43. Employing a trans-well assay, we assessed the migration across a porous mesh. Interestingly, the HMC3 FAM49B-KO and the BV2 Fam49b-KD cells migrated more efficiently than their respective controls, suggesting an increase in migration activity (Fig. 3c). Based on respective gene expression changes (adhesion genes heatmap: Fig. S3a), we aimed to evaluate the differences in adhesion capabilities. To assess adhesion differences, we utilized two experimental approaches. First, we employed an impedance-based assay. BV2 and HMC3 cells were plated in equal numbers onto an impedance plate, which measures the cell index based on electrical impedance, a direct indicator of adhesion. We tracked impedance changes from the moment cells were plated until they reached confluency. Both Fam49b-KD BV2 cells and FAM49B-KO HMC3 cells adhered at a significantly slower rate than control cells (Fig. 3d, e). A similar trend was confirmed with a counting assay performed in the HMC3 cells. Equal numbers of cells were plated overnight, and the following day, attached cells were counted and compared to live cells remaining unattached in the supernatant. FAM49B-KO cells displayed delayed adhesion to the culture plate surface (Fig. S3b). Next, we examined microglial endocytosis, a process that requires significant cytoskeletal rearrangement. As the primary macrophage of the CNS, endocytosis is a crucial function of microglia44,45. Since we observed that endocytosis-related genes were differentially expressed in FAM49B-KO HMC3 cells, we hypothesized that their endocytic capacity might be changed (endocytosis genes heatmap: Fig. S3c). Applying fluorescence-labeled dextran and microscopy-based analysis, we determined that intracellular dextran levels were significantly lower in Fam49b-KD and FAM49B-KO microglia (Fig. 3f, g). To test whether this outcome is due to reduced endocytosis or enhanced clearance by autophagy, we performed an autophagy assay. We found that the autophagic clearance is faster in FAM49B-KO cells, suggesting an increased autophagic flux, which is in line with the upregulation of multiple autophagy regulatory genes (Fig. 3h, i) (proteolysis/autophagy genes heatmap: Fig. S3d,e). In summary, our findings indicate that microglia lacking FAM49B exhibit significantly impaired adhesion and increased autophagy, both critical hallmarks of microglial function. Additionally, we observed that KO and KD cells migrated faster than WT cells, which is in line with an activation phenotype and their reduced adhesion, which may lower resistance to movement.
a Heatmap of cytoskeletal genes that were significantly altered in FAM49B-KO cells. b Representative brightfield images of HMC3 WT and FAM49B-KO cells. Dot plot shows quantification of roundness comparing WT vs. KO (scale bar 50 μm, N = 3, n = 38 (WT)/67 (KO), **p < 0.01, Student’s unpaired t-test). c Analysis of trans-well migration assays performed in the HMC3 cells (N = 3, 3) and BV2 cells (N = 8, 8) showing increased migration in FAM49B-KO and KD cells. d Cell adhesion assay measured using the XCELLigence Agilent impedance device, showing impaired adhesion in Fam49b-KD BV2 cells (N = 8, 8) and e FAM49B-KO HMC3 cells (n = 4, 4). Data shown as mean (±SEM) analyzed using multiple t-test. ***p < 0.001. f Endocytosis assay performed using FITC-Dextran quantification in BV2 cells (siCtrl: N = 3, n = 260; siFam49b: N = 3, n = 204) and g HMC3 cells (WT: N = 3, n = 701, KO: N = 3, n = 940). Data was analyzed using Welch’s t-test. **p < 0.01 ***p < 0.001. h Autophagy assay, measuring GFP fluorescence over time after DAPGreen uptake in HMC3 WT and FAM49B-KO cells. Data shown as mean fluorescence ± SEM, two-way repeated measures ANOVA (n = 5 wells for each condition, p < 0.001, ***). i Representative images (GFP/brightfield) of cells at times 5 min and 55 min (note cell movement over time in ROIs), showing pronounced decline in fluorescence in KO cells (scale bar 100 µm).
Exposure of FAM49B-KO microglia to DA neuron cultures triggers a global immune response
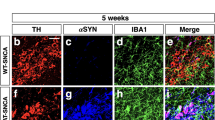
In the midbrain, activated microglia release molecules that can activate a response in surrounding cells. Activation of astrocytes by microglia, for example, can exacerbate neuroinflammation20. In PD models, it has been suggested that α-synuclein enhances microglial motility and that microglia-derived TNF-α induces astrocyte migration46,47. Since we observed a significant increase in migratory behavior in FAM49B-KO cells (Fig. 3c), we sought to investigate their impact on the midbrain environment, including DA neurons, glial cells and fibroblasts. Using human stem cell-derived DA neuron cultures that include glial cells, epithelial cells and perivascular fibroblasts48, we transferred either WT or FAM49B-KO HMC3 cells onto these cultures and monitored overall migratory behavior. Interestingly, we observed a significant increase in motility when FAM49B-KO cells were plated onto the cultures as measured by live imaging for 36 h (Fig. 4). Additionally, we have previously found that depletion of the PD risk gene SATB1 induced a senescence phenotype in DA neurons, leading to a significant microglial immune response in the midbrain12, thus, we sought to determine the effect of exposure of senescent SATB1-KO neuron cultures to FAM49B-KO microglia as compared to WT neuron cultures. As expected, plating of FAM49B-KO microglia onto senescent DA neuron cultures resulted in an even higher reaction, indicated by a significant increase in overall motility as measured per cell by velocity (Fig. 4a, b) and distance traveled (Fig. S4a). It has been shown that PAI-1 (SERPINE1) acts as a hub both as a target and initiator of various pathways that regulate cellular motility via cues, including urokinase-type plasminogen activator (uPA (PLAU)) and growth factors such as transforming growth factor β1 (TGF-β1 (TGFB1))49. Interestingly, these migratory signaling molecules were found to be upregulated in FAM49B-KO microglia (Fig. 4c), suggesting that these molecules stimulate an overall migratory response in the human stem cell-derived midbrain cultures. Interestingly, comparing DA neuron survival using a live/dead assay did not reveal any significant differences in survival after 24 h of exposure to WT or FAM49B-KO microglia (Fig. S4b), suggesting that a longer exposure (as in aged human patients) is required to impact neurodegeneration.
a Scatterplot of migration velocity of global cell movement induced by plating WT and FAM49B-KO microglia (HMC3) on WT and SATB1-KO dopamine neuron cultures (DA) (HMC3WT/DAWT; HMC3WT/DASATB1-KO; HMC3FAM49B-KO/DASATB1-KO). Mean ± SD is shown (n > 4.200, N = 3, 3 per condition), data were analyzed using One-way ANOVA. **p < 0.01 ****p < 0.001. b Representative images showing individual cell migration tracks over 36 h of tracking to compare different plating conditions (FIJI, tracking). c Marker gene expression of migratory cues in HMC3 cells. (Data are presented as mean ± SEM and were analyzed using an unpaired Mann–Whitney t-test. *p < 0.05, **p < 0.01).
Taken together, these results suggest that the presence of FAM49B-reduced microglia (as found in the aged SNpc) may contribute to increased immune activation in surrounding midbrain cell types, potentially leading to inflammaging. This could eventually contribute to the age-related vulnerability of the SNpc and the age-driven presence of inflammation markers.
FAM49B-KO in microglia increased immune response which is ameliorated by NFκB inhibition
Microglia, the resident macrophages of the central nervous system, play a crucial role in immune surveillance and inflammatory responses. Upon encountering inflammatory stimuli, microglia can polarize toward a pro-inflammatory state and secrete cytokines such as interleukin-8 (IL-8), which mediates neuroinflammation39. Interestingly, it has been reported that the CSF levels of IL-8 are among the most important pro-inflammatory mediators associated with motor and cognitive decline, as well as with neurodegenerative/PD-specific biomarkers50. Given their central role in neuroinflammation, dysregulation of microglial activity has been implicated in aging and neurodegenerative diseases, including PD51,52.
Toll-like receptors (TLRs) expressed in microglia are elevated in individuals with PD, contributing to neuroinflammation and neurodegeneration. TLR-mediated inflammation has been suggested to contribute to the loss of DA neurons in PD53, and TLR2/6 and TLR5 have been shown to bind α-synuclein54. Interestingly, loss of FAM49B significantly led to a signification upregulation of TLRs, including TLR5 and 6 (Fig. 5a). TLRs are known to regulate immune responses via NFκB activation. In addition to TLRs, it has been found that both LPS and α-synuclein (specifically in microglia55) can bind to the receptor for advanced glycation end products (RAGE) and induce the non-canonical inflammasome driven by CASP456. Accordingly, we found both CASP4 and the inflammasome gene NLPR1 upregulated in FAM49B-KO cells (Fig. 5a). To further investigate the role of FAM49B in microglial immune responses, we stimulated WT and FAM49B-KO or -KD cells with inflammatory triggers, including tumor necrosis factor-alpha (TNF-α, in HMC3 cells) and lipopolysaccharides (LPS, in BV2 and HMC3 cells). Notably, the absence or reduction of FAM49B led to a significantly heightened pro-inflammatory response. FAM49B-KO in HMC3 cells caused a significant upregulation of IL-8 even without TNF-α stimulation (Fig. 5b). Importantly, both FAM49B-KO and Fam49b-KD cells exhibited significantly increased expression of multiple immune response genes, following TNF-α or LPS treatment, compared to their WT or control counterparts (Figs. 5c, S5a (HMC3) and Fig. S5b (BV2)). It is well-established that activation of c-Jun serves as a central regulatory mechanism of microglial inflammatory responses57. Hence, we performed western blotting to determine the level of Jun activation and found that even without inflammation induction, FAM49B-KO cells showed significantly higher levels of phospho-Jun compared to WT (Fig. 5e). Consistent with this overall increase in reactivity, transcriptomic analysis revealed the activation of several pro-inflammatory pathways, including mTOR, AMPK, MAPK, and TNF signaling (Fig. 5f).
a TLR genes, JUN and NFKB1 expression in HMC3 cells. b IL8 expression in HMC3 cells, c IL8 expression is triggered by TNFα stimulation. FAM49B-KO cells react significantly more to this stimulation. d The expression of IL8 is significantly ameliorated by NFkB inhibitor IV treatment in HMC3 FAM49B-KO cells. e Western blot and quantification of WT vs. FAM49B-KO HMC3 cells showing upregulation of Jun phosphorylation. f Circos plot correlating the top 15 contributing differentially expressed genes to common inflammatory pathways (data are presented as mean ± SEM and were analyzed using multiple t-tests in (a) and unpaired t-test in (b, c, e), and two-way ANOVA (d). *p < 0.05, ***p < 0.001).
These findings suggest that the reduction of FAM49B primes microglia toward an exaggerated inflammatory response, potentially contributing to neuroinflammation observed in aging and neurodegenerative disorders.
To explore whether pharmacological intervention could mitigate this excessive immune response, we treated both WT and FAM49B-KO HMC3 microglia with NFκB inhibitor IV. Quantitative PCR analysis revealed that the inhibitor significantly reduced IL-8 expression (Fig. 5d), suggesting that anti-inflammatory treatments may help ameliorate the detrimental hyperactivation of microglia associated with aging and PD. These findings are consistent with previous studies demonstrating that anti-inflammatory interventions can improve neuroinflammatory pathology in models of PD58,59. Moreover, systemic inflammation, as modeled by LPS administration, has been shown to induce microglial overactivation and contribute to DA neuron loss in the SNpc, further establishing a link between chronic inflammation and neurodegeneration60. It has been reported that stimulation of microglia with LPS or α-SYN induced proliferation61,62; thus, we tested the impact of FAM49B-KO on this response. Interestingly, while both LPS and α-SYN pre-formed fibrils (PFF) stimulation induced proliferation in WT cells, FAM49B-KO did not increase proliferation (Fig. S5c).
Taken together, our results suggest that FAM49B serves as a key regulator of microglial activation, and its downregulation with age could be associated with increased reactivity, inflammaging, and, ultimately, neurodegenerative diseases.
Discussion
Given the discovery of disease- and age-related downregulation of the PD risk factor FAM49B in human nigral microglia, this study provides critical new insights into how aging and PD converge on the innate immune system, particularly on the resident immune cell of the brain, to contribute to neurodegeneration. While the intersection of age-related immune dysfunction and PD pathogenesis remains poorly understood, growing evidence suggests that chronic glial activation, impaired immune homeostasis, and dysregulated phagocytic clearance contribute to disease. Aging is the strongest risk factor for idiopathic PD3. Using human gene expression data from different age groups, our results suggest how aging alters the molecular and functional landscape of microglia, driven by the reduction of FAM49B. This decline leads to increased expression of inflammatory mediators, loss of homeostatic signatures, and dysregulated endocytic and autophagic capacity. These observations support the view that age-associated dysfunction of microglia may contribute to vulnerability of the SNpc and a permissive environment for the development of PD. Additionally, we found that FAM49B expression is significantly downregulated in microglia of PD patients compared to age-matched controls. The key finding of our study is the identification of FAM49B as a molecular link between aging, immune dysregulation, and PD risk. A GWAS has implicated FAM49B as a PD risk gene27, while its role in glial function and inflammaging remained elusive. Here, we show that FAM49B expression is downregulated in microglia of the SNpc with age as well as with disease, suggesting a critical role in maintaining immune homeostasis in the brain. Our functional studies revealed that FAM49B-KO in microglia results in aberrant activation, with upregulation of pro-inflammatory cytokines, consistent with a transition toward a disease-associated-like or responding state. However, this activation was not accompanied by functional upregulation of microglial proliferation mechanisms. Instead, FAM49B-deficient microglia showed impaired proliferative response to LPS or a-SYN stimuli, indicating a separation of their inflammatory response from proliferative ability. This is highly relevant for PD, where a hallmark of pathology is the accumulation of activated and ameboid microglia in the SNpc63. We suggest that FAM49B might play a dual role, on the one hand, regulating inflammatory responses, and on the other hand, coordinating cytoskeletal function. Thus, it is plausible that loss of FAM49B may tip the homeostatic balance toward a maladaptive microglial phenotype that is neuroinflammatory, which could contribute to midbrain inflammaging. This aligns with age-related microglial dysfunction and thereby contributes to midbrain inflammaging.
Our findings support previous research that identifies senescent or dystrophic microglia in aged and diseased brains64, suggesting that immunosenescence in the brain contributes to neurodegenerative vulnerability. More concretely, our data confirm the outcome of single-cell studies showing age-associated occurrence of dysfunctional glial states with pro-inflammatory but inefficient immune profiles18,65. Despite the ubiquitous gene expression of PD-associated factors, the SNpc is most vulnerable to degeneration, suggesting that, in addition to the intrinsic vulnerability of DA neurons, local cellular context, including the density and state of glial cells, are critical determinants of susceptibility. Importantly, we observed that SNpc expression of FAM49B was highest in microglia, further supporting its relevance to nigral immune health. Reduction of FAM49B in microglia, which occurs with age in this vulnerable brain region, is associated with a shift from surveillance to increased immune activation.
While our study, based on human midbrain-derived sequencing data as well as mouse expression data and modeled in microglia-like cell lines, showed highly reproducible results, it has some limitations. We chose cell lines (which were thoroughly characterized34,66) to ensure high reproducibility. We are aware that the cell line-based results could be reproduced in animal models of neurodegenerative disease and/or human primary microglia. However, these approaches have their own caveats, such as inter-individual microglia differences and variation in microglia states due to isolation procedures, which could influence FAM49B expression and thus the outcome and interpretation of such functional analyses.
In summary, our results establish the PD risk factor FAM49B as a novel regulator of microglial state and function with a high relevance to both PD pathogenesis and brain aging. Its downregulation with age could lead to a “second hit” cascade in PD that exacerbates inflammaging (due to basal upregulation of interleukin expression) while reducing protective clearance mechanisms (reduced reactive proliferation, dysregulated endocytosis/autophagy), a combination that may increase vulnerability to DA neuron loss. Our findings suggest a mechanistic rationale for why certain individuals, particularly older adults with a genetic predisposition, may develop PD in response to otherwise subthreshold stressors. Potentially, our data suggest FAM49B as a novel drug target, where boosting its function may offer a strategy to simultaneously suppress neurotoxic inflammation. Interestingly, in addition to the direct role of FAM49B in regulating microglial function, upstream pathways that modulate FAM49B may offer translational opportunities. For example, LANCL1 has been described as a regulator of FAM49B and is critical for CNS oxidative stress homeostasis and neuronal survival67. Since abscisic acid (ABA) binds to LANCL1 and exerts anti-inflammatory effects in PD models68,69, future studies could evaluate whether the ABA-LANCL1-FAM49B axis represents a viable therapeutic strategy to restore microglial homeostasis and reduce neuroinflammation. Additionally, we found that FAM49B-KO cells upregulate the expression of TLRs, which have been indicated to increase in the context of PD and mediate inflammatory responses in microglia by activation of NFκB. This process of chronic activation of TLRs and neuroinflammation was suggested to lead to neurodegeneration in PD53. Both toxin-induced (1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine, MPTP) and pre-formed fibrils of αSyn-induced PD mouse models were ameliorated by NFκB inhibition70,71.
Interestingly, binding of α-synuclein monomers or oligomers to TLR5 efficiently activated the NOD-like receptor pyrin domain containing 3 (NLRP3) inflammasome72 and TLR6 (which binds α-synuclein as TLR2/6 heterodimer54) induces expression of inflammatory cytokines by activating NFκB73. This is in line with our findings that TLR5 and 6 were significantly upregulated in FAM49B-KO cells, and the pro-inflammatory activity in FAM49B-KO microglia was ameliorated by NFκB inhibition. Additionally, it has been reported that RAGE receptors can mediate uptake of LPS or α-SYN, leading to non-canonical inflammasome activation driven by CASP456. Indeed, we found both CASP4 and the inflammasome gene NLPR1 upregulated in FAM49B-KO cells, suggesting an additional, TLR-independent way of activation. The previously published findings, in combination with our results, suggest that microglia activity (partially mediated by NFκB) drives the neuroinflammatory and neurodegenerative pathology in diverse PD models. With future studies, it would be interesting to determine if FAM49B levels were reduced in these animals, possibly connecting detrimental microglia activity to genetic predisposition and age-related susceptibility.
Our research establishes the PD risk gene FAM49B as a key molecular regulator of microglial activity during aging and PD, thus providing insights into genetic vulnerability and immune function failure that may contribute to neurodegeneration. Hence, research focused on immune homeostasis pathways of the aging brain shows promise as a method to potentially prevent or delay PD and other age-related neurodegenerative disorders.
Methods
Age-associated differential gene expression analysis of the human SNpc
To deconvolute and estimate the relative proportions of major brain cell types in each bulk RNA-seq sample, we applied the methodology as described by Chikina et al.74. This method uses marker gene sets to infer latent variables representing cell-type abundance, enabling adjustment for cellular heterogeneity in bulk tissue expression data. Marker gene sets for neurons, astrocytes, microglia, oligodendrocytes, and endothelial cells were obtained from Zhang et al.75, who identified these markers from purified brain cell populations. Using these marker sets, we computed relative cell-type proportion estimates for each GTEx SNpc sample using normalized gene expression values provided by the GTEx consortium (version 8). These estimates were then used as covariates in a linear model framework to assess gene expression associations with donor age while controlling for differences in cell-type composition across samples. FAM49B expression was specifically evaluated for its correlation with inferred cell-type proportions and for age-associated differential expression after covariate adjustment. Significance was assessed using linear regression models, and p-values were corrected for multiple testing. We further confirmed that the age-associated change in FAM49B expression could be specifically attributed to microglia expression through an F-test, allowing for the identification of cell-type-specific expression changes. All code to replicate the analysis is available in the Supplementary Material.
HMC3 cell culture
HMC3 cells were cultured according to the vendor’s instructions. Cells were grown at 37 °C in 5% CO2. The medium used was Corning Minimum Essential Medium Eagle with Earle’s salts and L-glutamine (#10-010-CV) with 1% Pen-Strep, 1% Sodium pyruvate, 1% MEM Non-essential Amino Acid Solution (#M7145-100 mL, SIGMA) and 10% Fetal Bovine Serum (FBS). Cells were passaged every 2–3 days with Trypsin-EDTA (ThermoFisher #25200072). HMC3 cells were stimulated with 0.3 μg/mL TNFα (#TNA-H4211, Acro Biosystems), LPS (Sigma), and a-synuclein PFFs (StressMarq).
Immunohistochemistry in mice
Five-month-old animals used were C57/BL6J, and 13-month-old animals were Drd-1a-tdtomato. Mice were anesthetized via isoflurane inhalation prior to being transcardially perfused first using PBS, followed by 4% paraformaldehyde in PBS. Brains were extracted and fixed for 18 h at 4 °C, and then cryoprotected in 30% sucrose in PBS for 48 h at 4° C. Fixed brains were then frozen in O.C.T. compound (ThermoFisher #4585) before being cryo-sectioned (Leica Biosystems) at −21 °C. Free-floating coronal sections (40 μM) were collected between −3.9 mm and −2.5 bregma containing the substantia nigra. Sections were blocked for 3 h in 5% NDS (Sigma #D9663) and 0.2% Triton-X-100 (Sigma #9036-19-5) in PBS, then incubated overnight in 1% NDS and 0.2% Triton-X-100 in PBS containing primary antibodies for 24 h. Sections were washed 3X in 1% NDS and 0.2% Triton-X-100 in PBS for 10 min each before being incubated at room temperature for 4 h in 1% NDS and 0.2% Triton-X-100 in PBS containing fluorescent secondary antibodies, and washed as above (3 × 10 min). Sections were then mounted onto microscope slides (Epredia #4951), and cover-slipped (VWR) using DAPI-Fluoromount-G (Southern Biotech #0100-20). The primary antibodies were the following: anti-Tyrosine-Hydroxylase (1:500, #ab9702), anti-FAM49B antibody (1:500, SCT #sc-390478), and anti-IBA1 antibody (1:500, Wako-#019-19741). The secondary antibodies were the following: donkey anti-chicken Alexa Fluor 546 (1:1000, Invitrogen #A11041), donkey anti-mouse Alexa Fluor 488 (1:1000, Invitrogen #A21202), donkey anti-rabbit Alexa Fluor 697 (1:1000, Invitrogen #A31573), donkey anti-rabbit Alexa Fluor 405 (1:1000, Invitrogen #A48258). Images were obtained using an Olympus confocal microscope and processed using Fiji. Images were first Z-projected to obtain the Z_SUM, then thresholded in the IBA1 channel to create ROIs, and measurements were taken in the FAM49B-488 channel to calculate the corrected total cell fluorescence (CTCF).
BV2 cell culture
BV2 cells were cultured according to the vendor’s instructions. Cells were grown at 37 °C in 5% CO2. The medium used was RPMI (#11875-093, Gibco) with 1% Pen-Strep and 10% Fetal Bovine Serum (FBS). Cells were passaged every 2–3 days with Trypsin-EDTA (#25200072, ThermoFisher).
siRNA transfection
BV2 cells were seeded at 2.5 × 105 per well on a 6-well plate using RPMI medium and left overnight. Cell medium was changed 30 min prior to transfection, which was performed using Lipofectamine RNAiMAX (ThermoFisher #13778030), OptiMEM (Gibco #31985062), 15 pmol Silencer Select siFAM49B siRNA (ThermoFisher #4390771), and 15 pmol Silencer Select Negative Control (ThermoFisher #4390843) in accordance with the ThermoFisher RNAiMAX protocol. Twenty-four hours following transfection, cells were passaged onto the respective vessels required for each experiment.
Trans-well migration assay
HMC3: WT and FAM49B KO HMC3 cells were cultured in separate T25 flasks using EMEM medium supplemented with 10% FBS and 1% penicillin/streptomycin and incubated at 37 °C. Upon reaching 90% confluency, the cell medium was removed and replaced with starvation medium (EMEM supplemented with 1% FBS and 1% penicillin/streptomycin), and cells were incubated at 37 °C for 24 h. Flasks were rinsed with 2 mL pre-warmed D-PBS and then passaged with 1 mL trypsin and neutralized with 2 mL starvation medium. Cells were seeded at 5.0 × 104 per well in 250 µL onto 12.0 µm trans-well inserts (Millipore #PIXP01250) coated with 0.1% gelatin. 500 µL normal (not starvation) HMC3 media was placed in the bottom chamber of each well, and cells were incubated at 37 °C for 48 h. Inserts were rinsed with 250 µL PBS, and wells were rinsed with 500 µL PBS and then aspirated. 500 µL 2.8 µM Calcein AM (in 0.5 M EDTA) was placed in each well, and cells were incubated at 37 °C for 1 h. Wells were scraped, and the solution was transferred into a black 96-well plate (100 μL/well). Samples were excited using a wavelength of 485 nm, and fluorescence at 535 nm was measured using a SpectraMax iD3 plate reader (Molecular Devices).
BV2: Transfected BV2 cells were seeded at 5.0 × 105 per well on a 6-well plate using DMEM medium supplemented with 10% FBS and 1% pen/strep and incubated at 37 °C overnight. Cell medium was removed and replaced with starvation medium (DMEM supplemented with 1% FBS and 1% penicillin/streptomycin) 30 min prior to siRNA transfection. Twenty-four hours following transfection, cells were passaged in 500 μL trypsin and neutralized with starvation medium. Cells were seeded at 5.0 × 104 per well in 250 μL onto 8.0 μm trans-well inserts (VWR #734-2748) coated with 0.1% gelatin (Sigma #G1393). 500 μL normal (not starvation) BV2 media was placed in the bottom chamber of each well, and cells were incubated at 37 °C for 24 h. Inserts and wells were rinsed with PBS, cells were stained with Calcein AM, and fluorescence was measured as described above.
Roundness assay
To determine changes in the shape of the microglia, we applied a measurement of roundness as previously published12. In brief, brightfield images of living microglia cultures were taken using an Accu-Scope EXI-410 microscope with a Skye2 camera. Images were analyzed using the roundness measure in FIJI. While typically, ramified (branched) microglia indicate a surveillance state, more ameboid cells suggest activation.
Microglia co-culture with embryonic stem cells and cell tracking
Human embryonic stem cells [ESCs; wild-type H9 (WA-09), SATB1KO 12] were differentiated into midbrain dopamine (mDA) neurons according to an optimized version of a previously established protocol12.WT and SATB1-KO ESCs were differentiated at a density of 60k/well on a 96-well plate (µ-Plate 96 well square; Ibidi). On day 50, either HMC3-WT and HMC3-KO cells were fluorescently labeled (Cell tracker Orange CMTMR; Invitrogen) according to the manufacturer’s instructions and plated on top of either WT or SATB1-KO differentiated mDA neurons. Co-cultures are fluorescently imaged (586,647) every 30 min for 66 h using the Agilent BioTek Lionheart FX live imaging system. Note that cell labeling is not exclusive to HMC3 cells, and ESCs become fluorescently labeled within 30 min of co-culture. Cell tracking analysis was completed using TrackMate76. Statistical analysis was completed using Ordinary one-way ANOVA (prism).
Western blot
Cells were lysed for protein using RIPA buffer (ThermoFisher #89900) and protease and phosphatase inhibitors (#11836170001, Roche). Protein concentrations for each sample were determined using a BCA assay (ThermoFisher #23225) with a SpectraMax iD3 plate reader (Molecular Devices). Protein samples were boiled in 2X Tris-Glycine-SDS sample buffer (Novex #LC2676) at 95 °C for 5 min before being separated with electrophoresis using 10% Tris-glycine (Invitrogen #XP04200). Samples were then electro-transferred to nitrocellulose membranes (BioRad) and blocked using 5% BSA in TBS-Tween for 60 min, before undergoing overnight incubation with the required primary antibody in blocking solution. The primary antibodies used are listed as follows: mouse monoclonal anti-FAM49B antibody (SCT #sc-390478; ThermoFisher #PA5-52647), anti-TOM20 antibody (SCT #11415). To visualize and quantify protein bands, a ChemiDocXRS+ (BioRad) was used.
RNA isolation, qPCR, and RNA-seq
RNA was isolated from cells using the RNeasy Plus Mini Kit (#74136, QIAGEN) and QIAshredder (QIAGEN, #7956). Cells were lysed using 2-Mercaptoethanol. 200 ng of RNA was reverse transcribed using the Applied Biosystems High-Capacity cDNA Reverse Transcription Kit (ThermoFisher # 4368814), creating an output volume of 20 µL. Real-time qPCR was performed using a Quant Studio 3 (Applied Biosystems) and the Applied Biosystems Power SYBR Green PCR Master Mix (ThermoFisher #4367659). Reaction specificity was confirmed via melt curve analysis. Previously described in Russo et al.33, for RNA-seq, 500 ng of RNA for each sample was sent to Azenta for bulk RNAseq. In brief, sample quality control and determination of concentration were performed using TapeStation Analysis by Azenta, followed by library preparation and sequencing. Computational analysis, included in their standard data analysis package, was used for data interpretation. The gene expression data of the HMC3 RNA-seq are accessible on ArrayExpress (Accession number E-MTAB-15277).
Transmission electron microscopy
For transmission electron microscopy (TEM), cells were grown on Aclar film and fixed for 30 min in a mixture of cold 2% paraformaldehyde and 0.1% glutaraldehyde in 0.1 M phosphate buffer (PB), pH 7.4. After washing, cells were post-fixed with 1% osmium tetroxide for 30 min, and stained with 1% uranyl acetate for 30 min, dehydrated through an ascending series of ethanol, and embedded in Durcupan resin for 48 h at 60 °C. Ultrathin sections (60–90 nm) were cut on an ultramicrotome (Reichert Ultracut E) and placed on formvar-coated nickel slot grids. Sections were post-embedding immunogold labeled for rabbit FAM49b within 24 h of sectioning using a modification of the protocol of Phend et al.77. In brief, grids were rinsed in TBS containing 0.005% Tergitol NP-10, pH 7.6 (hereafter referred to as Tris-tergitol pH 7.6), incubated in saturated (10%) sodium meta-periodate for 5 s, rinsed, incubated in 1% sodium borohydride for 1 min, rinsed, and incubated in primary antibody (rabbit anti-FAM49B 1: 100) overnight at room temperature. The next day, grids were rinsed in Tris-tergitol pH 7.6, followed by Tris-tergitol pH 8.2, incubated in secondary antibody (goat anti-rabbit IgG conjugated to 18 nm gold particles) (Jackson Immuno Research) 1:25 in Tris-tergitol pH 8.2 for 1 h and rinsed. Grids were then stained with 1% methanolic uranyl acetate and 0.3% aqueous lead citrate and imaged using a JEOL 1200EX transmission electron microscope.
Endocytosis assay
Cells were treated with 1 mg/mL of 75 kDa Fluorescein isothiocyanate-Dextran (Sigma Aldrich #60842-46-8) in their respective medium for 15 min at 37 °C. Cells were then washed two times with cold PBS before being fixed in 4% PFA in PBS for 15 min at room temperature. Following fixation, cells were incubated with Phalloidin-647 (ThermoFisher #A22287) in PBS for 1 h at room temperature. Cells were then washed 1x in PBS, mounted using DAPI Fluoromount-G (Southern Biotech #0100-20) and imaged using an Olympus confocal microscope.
Live imaging autophagy assay
To test the impact of FAM49B KO on autophagy impairment, we used the autophagy detection dye DAPGreen (Dojindo, #D676). Wild-type control HMC3 cells as well as FAM49B KO HMC3 cells were treated with 0.1 µM DAPGreen (in DMSO) for 45 min. Cells were washed twice with normal medium and then treated with media containing 1 µL/mL DMSO for the duration of the live imaging protocol. Cells were immediately placed in the Lionheart FX live imaging microscope, and GFP fluorescence measurements were recorded every 30 s for 5 h. Fluorescence intensity was first normalized by cell number for each ROI and then normalized to the first time point for each condition.
Cell adhesion assays
Cell adhesion assay was performed using the XCELLigence Agilent Impedance device. Prior to seeding of cells, the 16-well E-plate was coated using 0.1% gelatin in D-PBS. HMC3 cells and BV2 cells were seeded at 1.5 × 104 cells per well. Cell index readings were taken every 1 min for a total of 60 min (BV2) and every 2 min for a total of 24 h (HMC3), or until cells reached confluency. An additional cell adhesion test was performed in the HMC3 cells using a counting assay. Equal numbers of cells were plated overnight, and the following day, attached cells were counted and compared to live cells remaining unattached in the supernatant. FAM49B-KO cells displayed delayed adhesion to the culture plate surface.
Mitochondrial assays
Cell respiration was measured using the Agilent Seahorse XFe96 Analyzer and the Mito Stress Test assay (Agilent #103015-100), which delivers a series of three drugs (Oligomycin, FCCP, Rotenone+Antimycin A) while the oxygen consumption rate (OCR) is recorded. OCR is used to calculate basal respiration, maximal respiration, proton leak, non-mitochondrial oxygen consumption, ATP production, spare respiratory capacity, and coupling efficiency. A BCA assay was performed in order to normalize the results based on total cell protein content. To measure mitochondrial membrane potential, a functional dye MitoTracker Red CMXRos (ThermoFisher #M7513) was diluted to 200 nM in D-PBS and incubated with cells for 15 min at 37 °C. Cells were then washed with PBS 3X and fixed before being mounted and imaged using an Olympus Confocal microscope. CTCF values were measured using Fiji. To measure mitochondrial ROS levels, MitoSOX Red superoxide indicator (ThermoFisher #M36007) was diluted in HBSS and incubated onto cells for 15 min at 37 °C. Cells were then imaged using the Agilent BioTek LionHeartFX live imaging system.
qPCR primers used
Target | Forward | Reverse |
|---|---|---|
B-Actin | AGCGAGCATCCCCCAAAGTT | GGGCACGAAGGCTCATCATT |
IL878 | ACTGAGAGTGATTGAGAGTGGAC | AACCCTCTGCACCCAGTTTTC |
CD11B78 | ACTTGCAGTGAGAACACGTATG | TCATCCGCCGAAAGTCATGTG |
TMEM11978 | CGGCCTATTACCCATCGTCC | CTGGGCTAACAAGAGAGACCC |
CYRIB-m25 | AGGAGCTGGCCACGAAATAC | GGCGTACTAGTCAAGGCTCC |
IL7-m79 | CTGATGATCAGCATCGATGAATTGG | GCAGCACGATTTAGAAAAGCAGCTT |
IL6-m (Origene #MP206798) | TACCACTTCACAAGTCGGAGGC | CTGCAAGTGCATCATCGTTGTTC |
TMEM119-m (Origene #MP217346) | ACTACCCATCCTCGTTCCCTGA | TAGCAGCCAGAATGTCAGCCTG |
Cd11b-m80 | TCATTCGCTACGTAATTGGG | GATGGTGTCGAGCTCTCTGC |
Data availability
The gene expression data of the HMC3 RNA-seq is accessible on ArrayExpress (Accession number E-MTAB-15277).
References
Moore, D. J., West, A. B., Dawson, V. L. & Dawson, T. M. Molecular pathophysiology of Parkinson’s disease. Annu. Rev. Neurosci. 28, 57–87 (2005).
Dauer, W. & Przedborski, S. Parkinson’s disease: mechanisms and models. Neuron 39, 889–909 (2003).
De Lau, L. M. & Breteler, M. M. Epidemiology of Parkinson’s disease. Lancet Neurol. 5, 525–535 (2006).
Martin, I., Dawson, V. L. & Dawson, T. M. Recent advances in the genetics of Parkinson’s disease. Annu. Rev. Genomics Hum. Genet. 12, 301–325 (2011).
Chang, D. et al. A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci. Nat. Genet. 49, 1511–1516 (2017).
Hamza, T. H. et al. Common genetic variation in the HLA region is associated with late-onset sporadic Parkinson’s disease. Nat. Genet. 42, 781–785 (2010).
Holmans, P. et al. A pathway-based analysis provides additional support for an immune-related genetic susceptibility to Parkinson’s disease. Hum. Mol. Genet. 22, 1039–1049 (2013).
Pierce, S. & Coetzee, G. A. Parkinson’s disease-associated genetic variation is linked to quantitative expression of inflammatory genes. PLoS ONE 12, e0175882 (2017).
Kam, T.-I., Hinkle, J. T., Dawson, T. M. & Dawson, V. L. Microglia and astrocyte dysfunction in Parkinson’s disease. Neurobiol. Dis. 144, 105028 (2020).
McGeer, P. L., Itagaki, S., Boyes, B. E. & McGeer, E. Reactive microglia are positive for HLA-DR in the substantia nigra of Parkinson’s and Alzheimer’s disease brains. Neurology 38, 1285–1285 (1988).
Calabrese, V. et al. Aging and Parkinson’s disease: inflammaging, neuroinflammation and biological remodeling as key factors in pathogenesis. Free Radic. Biol. Med. 115, 80–91 (2018).
Riessland, M. et al. Loss of SATB1 induces p21-dependent cellular senescence in post-mitotic dopaminergic neurons. Cell Stem Cell. https://doi.org/10.1016/j.stem.2019.08.013 (2019).
Russo, T. & Riessland, M. Age-related midbrain inflammation and senescence in Parkinson’s disease. Front. Aging Neurosci. 14, 917797. https://doi.org/10.3389/fnagi.2022.917797 (2022).
Panicker, N. et al. Fyn kinase regulates misfolded α-synuclein uptake and NLRP3 inflammasome activation in microglia. J. Exp. Med. 216, 1411–1430 (2019).
Gordon, R. et al. Protein kinase Cδ upregulation in microglia drives neuroinflammatory responses and dopaminergic neurodegeneration in experimental models of Parkinson’s disease. Neurobiol. Dis. 93, 96–114 (2016).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271.e256 (2019).
De Biase, L. M. et al. Local cues establish and maintain region-specific phenotypes of basal ganglia microglia. Neuron 95, 341–356.e346 (2017).
Olah, M. et al. A transcriptomic atlas of aged human microglia. Nat. Commun. 9, 539 (2018).
Surmeier, D. J., Obeso, J. A. & Halliday, G. M. Selective neuronal vulnerability in Parkinson disease. Nat. Rev. Neurosci. 18, 101–113 (2017).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Singh, S. S. et al. NF-κB-mediated neuroinflammation in Parkinson’s disease and potential therapeutic effect of polyphenols. Neurotox. Res. 37, 491–507 (2020).
Nalls, M. A. et al. Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease. Nat. Genet. 46, 989–993 (2014).
Chattaragada, M. S. et al. FAM49B, a novel regulator of mitochondrial function and integrity that suppresses tumor metastasis. Oncogene 37, 697–709 (2018).
Shang, W. et al. Genome-wide CRISPR screen identifies FAM49B as a key regulator of actin dynamics and T cell activation. Proc. Natl. Acad. Sci. USA 115, E4051–E4060 (2018).
Park, C.-S. et al. Fam49b dampens TCR signal strength to regulate survival of positively selected thymocytes and peripheral T cells. eLife 13, e76940 (2024).
Trainor, A. R., MacDonald, D. S. & Penney, J. Microglia: roles and genetic risk in Parkinson’s disease. Front. Neurosci. 18, 1506358 (2024).
Nalls, M. A. et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol. 18, 1091–1102 (2019).
Consortium, T. G. et al. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Agarwal, D. et al. A single-cell atlas of the human substantia nigra reveals cell-specific pathways associated with neurological disorders. Nat. Commun. 11, 4183 (2020).
Martirosyan, A. et al. Unravelling cell type-specific responses to Parkinson’s disease at single cell resolution. Mol. Neurodegener. 19, 7 (2024).
Graff, B. J., Payne, S. J. & El-Bouri, W. K. The ageing brain: investigating the role of age in changes to the human cerebral microvasculature with an in silico model. Front. Aging Neurosci. 13, 2021 (2021).
Chinta, S. J. et al. Cellular senescence is induced by the environmental neurotoxin paraquat and contributes to neuropathology linked to Parkinson’s disease. Cell Rep. 22, 930–940 (2018).
Russo, T., Plessis-Belair, J., Sher, R. & Riessland, M. Regulatory network inference of induced senescent midbrain cell types reveals cell type-specific senescence-associated transcriptional regulators. Preprint at bioRxiv https://doi.org/10.1101/2025.02.06.636893 (2025).
Dello Russo, C. et al. The human microglial HMC3 cell line: where do we stand? A systematic literature review. J. Neuroinflamm. 15, 259 (2018).
Sangineto, M. et al. Metabolic reprogramming in inflammatory microglia indicates a potential way of targeting inflammation in Alzheimer’s disease. Redox Biol. 66, 102846 (2023).
Yang, S. et al. Microglia reprogram metabolic profiles for phenotype and function changes in central nervous system. Neurobiol. Dis. 152, 105290 (2021).
Paolicelli, R. C. et al. Microglia states and nomenclature: a field at its crossroads. Neuron 110, 3458–3483 (2022).
Tremblay, M.-È. The role of microglia at synapses in the healthy CNS: novel insights from recent imaging studies. Neuron Glia Biol. 7, 67–76 (2011).
Colonna, M. & Butovsky, O. Microglia function in the central nervous system during health and neurodegeneration. Annu. Rev. Immunol. 35, 441–468 (2017).
Scheiblich, H. et al. Microglia rescue neurons from aggregate-induced neuronal dysfunction and death through tunneling nanotubes. Neuron 112, 3106–3125.e3108 (2024).
Smolders, S. M.-T. et al. Microglia: brain cells on the move. Prog. Neurobiol. 178, 101612 (2019).
Bosco, E. E., Mulloy, J. C. & Zheng, Y. Rac1 GTPase: a “Rac” of all trades. Cell Mol. Life Sci. 66, 370–374 (2009).
Fort, L. et al. Fam49/CYRI interacts with Rac1 and locally suppresses protrusions. Nat. Cell Biol. 20, 1159–1171 (2018).
Li, Q. & Barres, B. A. Microglia and macrophages in brain homeostasis and disease. Nat. Rev. Immunol. 18, 225–242 (2018).
Solé-Domènech, S., Cruz, D. L., Capetillo-Zarate, E. & Maxfield, F. R. The endocytic pathway in microglia during health, aging and Alzheimer’s disease. Ageing Res. Rev. 32, 89–103 (2016).
Burda, J. E. & Sofroniew, M. V. Reactive gliosis and the multicellular response to CNS damage and disease. Neuron 81, 229–248 (2014).
Campolo, M. et al. TLR4 absence reduces neuroinflammation and inflammasome activation in Parkinson’s diseases in vivo model. Brain Behav. Immun. 76, 236–247 (2019).
Kim, T. W. et al. Biphasic activation of WNT signaling facilitates the derivation of midbrain dopamine neurons from hESCs for translational use. Cell Stem Cell 28, 343–355.e345 (2021).
Czekay, R. P. et al. PAI-1: an integrator of cell signaling and migration. Int. J. Cell Biol. 2011, 562481 (2011).
Lerche, S. et al. CSF and serum levels of inflammatory markers in PD: sparse correlation, sex differences and association with neurodegenerative biomarkers. Front. Neurol. 13, 834580 (2022).
Tansey, M. G. & Goldberg, M. S. Neuroinflammation in Parkinson’s disease: its role in neuronal death and implications for therapeutic intervention. Neurobiol. Dis. 37, 510–518 (2010).
Heneka, M. T., Kummer, M. P. & Latz, E. Innate immune activation in neurodegenerative disease. Nat. Rev. Immunol. 14, 463–477 (2014).
Heidari, A., Yazdanpanah, N. & Rezaei, N. The role of Toll-like receptors and neuroinflammation in Parkinson’s disease. J. Neuroinflamm. 19, 135 (2022).
Zhang, X., Yu, H. & Feng, J. Emerging role of microglia in inter-cellular transmission of α-synuclein in Parkinson’s disease. Front. Aging Neurosci. 16, 2024 (2024).
Long, H. et al. Interaction of RAGE with α-synuclein fibrils mediates inflammatory response of microglia. Cell Rep. 40, 111401 (2022).
Yi, Y. S. Functional interplay between non-canonical inflammasomes and autophagy in inflammatory responses and diseases. Korean J. Physiol. Pharm. 29, 129–138 (2025).
Waetzig, V. et al. c-Jun N-terminal kinases (JNKs) mediate pro-inflammatory actions of microglia. Glia 50, 235–246 (2005).
Caggiu, E. et al. Inflammation, infectious triggers, and Parkinson’s disease. Front. Neurol. 10, 122 (2019).
Chen, H. et al. Nonsteroidal anti-inflammatory drugs and the risk of Parkinson disease. Arch. Neurol. 60, 1059–1064 (2003).
Qin, L. et al. Systemic LPS causes chronic neuroinflammation and progressive neurodegeneration. Glia 55, 453–462 (2007).
Jung, H. et al. LPS induces microglial activation and GABAergic synaptic deficits in the hippocampus accompanied by prolonged cognitive impairment. Sci. Rep. 13, 6547 (2023).
Yin, S. et al. TREM2 signaling in Parkinson’s disease: regulation of microglial function and α-synuclein pathology. Int. Immunopharmacol. 143, 113446 (2024).
Badanjak, K., Fixemer, S., Smajić, S., Skupin, A. & Grünewald, A. The contribution of microglia to neuroinflammation in Parkinson’s disease. Int. J. Mol. Sci. https://doi.org/10.3390/ijms22094676 (2021).
Streit, W. J., Sammons, N. W., Kuhns, A. J. & Sparks, D. L. Dystrophic microglia in the aging human brain. Glia 45, 208–212 (2004).
Clarke, L. E. et al. Normal aging induces A1-like astrocyte reactivity. Proc. Natl. Acad. Sci. USA 115, E1896–E1905 (2018).
Henn, A. et al. The suitability of BV2 cells as alternative model system for primary microglia cultures or for animal experiments examining brain inflammation. Altex 26, 83–94 (2009).
Huang, H. et al. LANCL1, a cell surface protein, promotes liver tumor initiation through FAM49B-Rac1 axis to suppress oxidative stress. Hepatology 79, 323–340 (2024).
Spinelli, S. et al. LANCL1 binds abscisic acid and stimulates glucose transport and mitochondrial respiration in muscle cells via the AMPK/PGC-1α/Sirt1 pathway. Mol. Metab. 53, 101263 (2021).
Yang, Z., Liu, T., Kong, X. & Wei, J. Neuroprotective effect of abscisic acid on MPTP-induced Parkinson’s disease in mice. Mol. Nutr. Food Res. 69, e70111 (2025).
Ghosh, A. et al. Selective inhibition of NF-κB activation prevents dopaminergic neuronal loss in a mouse model of Parkinson’s disease. Proc. Natl. Acad. Sci. USA 104, 18754–18759 (2007).
Dutta, D. et al. Selective targeting of the TLR2/MyD88/NF-κB pathway reduces α-synuclein spreading in vitro and in vivo. Nat. Commun. 12, 5382 (2021).
Scheiblich, H. et al. Microglial NLRP3 inflammasome activation upon TLR2 and TLR5 ligation by distinct α-synuclein assemblies. J. Immunol. 207, 2143–2154 (2021).
Akira, S. & Takeda, K. Toll-like receptor signalling. Nat. Rev. Immunol. 4, 499–511 (2004).
Chikina, M., Zaslavsky, E. & Sealfon, S. C. CellCODE: a robust latent variable approach to differential expression analysis for heterogeneous cell populations. Bioinformatics 31, 1584–1591 (2015).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Ershov, D. et al. TrackMate 7: integrating state-of-the-art segmentation algorithms into tracking pipelines. Nat. Methods 19, 829–832 (2022).
Phend, K. D., Rustioni, A. & Weinberg, R. J. An osmium-free method of epon embedment that preserves both ultrastructure and antigenicity for post-embedding immunocytochemistry. J. Histochem. Cytochem. 43, 283–292 (1995).
Wang, Y.-J., Monteagudo, A., Downey, M. A., Ashton-Rickardt, P. G. & Elmaleh, D. R. Cromolyn inhibits the secretion of inflammatory cytokines by human microglia (HMC3). Sci. Rep. 11, 8054 (2021).
Martin, C. E. et al. Interleukin-7 availability is maintained by a hematopoietic cytokine sink comprising innate lymphoid cells and T cells. Immunity 47, 171–182.e174 (2017).
Skelly, D. T., Hennessy, E., Dansereau, M.-A. & Cunningham, C. A systematic analysis of the peripheral and CNS effects of systemic LPS, IL-1Β, TNF-α and IL-6 challenges in C57BL/6 mice. PLoS ONE 8, e69123 (2013).
Acknowledgements
This work was supported in part through NINDS grant 1R01NS124735 (M.R.) and a Starter Grant by the Thomas Hartman Foundation (M.R., J.L.P.). J.M. was partially supported by a Dr. W. Burghardt Turner Fellowship. Opinions, interpretations, conclusions, and recommendations are those of the author and are not necessarily endorsed by the sponsors.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.M., G.-J.L., C.Y.P., M.R. Methodology/Investigation: J.M., G.-J.L., C.Y.P., C.W., T.V.B., S.A., S.L., M.D., T.R., W.A., Z.B.H., M.W., B.K., M.R. Visualization: J.M., G.-J.L., C.Y.P., M.R. Supervision: M.R., O.G.T., J.L.P. Writing—original draft: M.R., J.M., G.-J.L., C.Y.P. Writing—review & editing: J.M., G.-J.L., C.Y.P., C.W., T.V.B., S.A., S.L., M.D., T.R., W.A., M.W., B.K., J.L.P., O.G.T., M.R.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Martin, J., Lai, GJ., Park, C.Y. et al. Age-related nigral downregulation of the Parkinson’s risk factor FAM49B primes human microglia for inflammaging. npj Aging 12, 1 (2026). https://doi.org/10.1038/s41514-025-00296-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41514-025-00296-z