Abstract
Urolithins are a class of bioactive metabolites derived from the metabolism of dietary ellagitannins by the human gut microbiota. In the gut, urolithins are dehydroxylated regioselectively based on microbiota composition and activity. A single 9-hydroxy urolithin dehydroxylase (ucd) operon in gut resident Enterocloster species has been described to date; however, most enzymes in the urolithin metabolic pathway remain uncharacterized. Here, we investigate urolithin cross-feeding between members of the gut microbiota and discover a novel urolithin dehydroxylase in a subset of Enterocloster species. We show that urolithin intermediates, released by gut resident Gordonibacter species during ellagic acid metabolism, are dehydroxylated at both the 9- and 10-positions by E. asparagiformis, E. citroniae, and E. pacaense, but not E. bolteae. Using untargeted proteomics, we uncover a 10-hydroxy urolithin dehydroxylase operon, termed uxd, responsible for these species-specific differences in urolithin metabolism. By inducing uxd expression with diverse urolithins, we show that 9-hydroxy urolithins are required for uxd transcription and 10-position dehydroxylation. Collectively, this study reveals some of the genes, proteins, and substrate features underlying differences in urolithin metabolism by the human gut microbiota.
Similar content being viewed by others
Introduction
The gut microbiota contributes to host homeostasis by regulating immune function, intestinal barrier integrity, and nutrient availability1. Due to the immense diversity of biosynthetic and metabolic gene clusters present in gut bacteria2, many ingested xenobiotics (food, drugs, pollutants) are metabolized by bacterial enzymes through reactions that the host alone cannot perform3,4,5. Urolithins (6H-dibenzo[b,d]pyran-6-ones) are a class of postbiotics derived from the metabolism of dietary ellagitannins by gut bacteria. The most common urolithin metabolite detected in host biological fluids following ellagitannin consumption is urolithin A (3,8-dihydroxy-urolithin, uroA)6,7. In the host, uroA can activate mitophagy to improve mitochondrial health and exercise capacity8,9,10. This mitophagy-promoting activity triggers immune responses in the context of adoptive T cell transfer by expanding T memory stem cells, a subset of minimally differentiated T cells that mediate potent anti-tumor responses11. UroA is also proposed to enhance gut barrier integrity by signaling through the aryl hydrocarbon receptor (AhR) pathway12 and has been shown to reduce Clostridioides difficile pathogenesis in murine models13. Based on the abovementioned activities, uroA is currently being evaluated in the clinic for indications in obesity (NCT05921266)14, cancer (NCT06022822), and aging (NCT06619457 and NCT06556706).
Dietary ellagitannins, the large and poorly absorbed precursors of urolithins, are plant secondary metabolites often consumed via berries, nuts, pomegranates, and aged alcoholic beverages15. This family of compounds is characterized by a central glucose (open or closed ring) linked to hexahydroxydiphenoyl (HHDP) and derivative groups. In the gut, the HHDP unit of ellagitannins is hydrolyzed and undergoes spontaneous lactonization, producing ellagic acid (EA)16. EA is decarboxylated and sequentially dehydroxylated by gut bacteria into diverse urolithins, depending on microbiota composition and function17,18. These individual differences in urolithin metabolism have been classified into 3 distinct metabotypes characterized by the terminal metabolites detected in host fluids and feces: 0 (no terminal metabolites), A (only uroA), and B (isouroA, uroB, and can include uroA)7. Like uroA, dietary sources of ellagitannins are in clinical trials for a variety of conditions; therefore, understanding the metabolism of this broad family of compounds is important since treatment responses may vary depending on urolithin metabotypes.
Although urolithin metabolism is prevalent in human populations6,19, few urolithin-metabolizing species have been described in the literature. The Eggerthellaceae family members Gordonibacter spp. and Ellagibacter isourolithinifaciens convert EA into urolithin M5 (3,4,8,9,10-pentahydroxy-urolithin, uroM5), urolithin M6 (3,8,9,10-tetrahydroxy-urolithin, uroM6), and urolithin C (3,8,9-trihydroxy-urolithin, uroC) via an EA decarboxylase, and 4- and 10-position dehydroxylases, respectively (Fig. 1a)20,21. E. isourolithinifaciens can further dehydroxylate uroC via an additional 8-position urolithin dehydroxylase, yielding isourolithin A (3,9-dihydroxy-urolithin, isouroA)22. A subset of gut resident Lachnospiraceae belonging to the Enterocloster genus (E. asparagiformis, E. bolteae, E. citroniae, and E. pacaense – also published as E. hominis23) can metabolize uroC into the bioactive terminal metabolite uroA through a 9-position urolithin C dehydroxylase encoded by the ucd operon24. We previously demonstrated that ucd-encoded proteins (UcdC, UcdF, and UcdO) belong to the xanthine dehydrogenase family and use NAD(P)H as an electron donor to dehydroxylate uroC and related 9-hydroxy urolithins, regardless of the presence or position of other hydroxyl groups on the urolithin scaffold. Intriguingly, a subset of uroC-metabolizing Enterocloster spp. can metabolize uroM6 into uroA by dehydroxylating both the 9- and 10-positions (Fig. 1a)25. The substrate specificity of the Ucd enzyme complex suggests that another gene cluster encodes a 10-position specific urolithin dehydroxylase in Enterocloster spp.
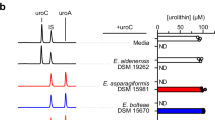
a Ellagic acid metabolism scheme. 9- and 10-position urolithin hydroxyl groups are highlighted in blue-grey and red circles, respectively. b Quantification of ellagic acid (EA) and urolithin concentrations in cell-free supernatants of G. massiliensis (Gm), G. urolithinfaciens (Gu), G. pamelaeae (Gp), or Gordonibacter sp. 28C (Gsp 28C) treated with EA (100 μM) in BHIrf media after 48 h. c Quantification of ellagic acid (EA) and urolithin concentrations in total cultures of Gordonibacter spp. treated with EA (100 μM) in BHIrf media after 48 h. Data from (b, c) are from matched cultures and are shown as means ± SEM (n = 2–3 biological replicates due to lack of growth for Gm and Gu in one replicate). d Aligned chromatograms (λ = 305 nm) of urolithins in dimethyl sulfoxide (DMSO, black lines) or EA-treated (100 μM, blue lines) cultures of Gu, E. bolteae (Eb), and E. asparagiformis (Ea) in BHI media after 5 days. e Aligned chromatograms (λ = 305 nm) of urolithins in uroM6-treated (100 μM, grey lines) cultures of Gu, Eb, and Ea in BHI media after 5 days. Data from (d, e) are from the same set of experiments. Retention times of urolithin standards are shown with grey dotted lines. Representative chromatograms from n = 1 of 3 biological replicates.
Here, we report the discovery of an operon, termed the urolithin 10-position (X) dehydroxylase (uxd), that encodes xanthine dehydrogenase family proteins responsible for urolithin dehydroxylation at the 10-position. We show that uroM6 is released by EA-metabolizing Gordonibacter spp., rendering this substrate available for metabolism by Enterocloster spp. By integrating untargeted proteomics, comparative genomics, and targeted gene expression analyses, we identify genes and proteins distinct from ucd that enable uroM6 dehydroxylation at the 10-position. Curiously, ucd and uxd operon expression is co-regulated, requiring a 9-hydroxy urolithin substrate for both 9- and 10-position dehydroxylation. Overall, this study uncovers an additional operon that regioselectively dehydroxylates urolithins and highlights how the regiochemistry of urolithins dictates dehydroxylase operon expression in Enterocloster spp.
Results
Urolithin M6 is released by Gordonibacter spp. and further metabolized by Enterocloster spp
Members of the Gordonibacter genus convert EA to intermediate urolithins uroM5, uroM6, and uroC (Fig. 1a)21. To determine whether 10-hydroxy urolithins (uroM5 and uroM6) were released by Gordonibacter spp. during EA metabolism, we incubated Gordonibacter pamelaeae, Gordonibacter massiliensis, Gordonibacter sp. 28C, and Gordonibacter urolithinfaciens with EA and monitored urolithin production in both whole cultures and supernatants by liquid chromatography-mass spectrometry (LC-MS). The products of EA metabolism were detected in cell-free supernatants after 2 days (Fig. 1b), albeit at lower concentrations than those observed in whole cultures (Fig. 1c). These data demonstrate that urolithin intermediates derived from EA metabolism are released by Gordonibacter spp. and become available to other urolithin-metabolizing bacteria.
To support this hypothesis, we performed co-culture experiments by incubating combinations of G. urolithinfaciens and Enterocloster spp. (E. bolteae or E. asparagiformis) with EA or uroM6. Based on the EA decarboxylase and urolithin dehydroxylase activities (4-, 9-, and 10-positions) of these 2-member communities, we hypothesized that uroA would be detected in co-cultures. As expected, G. urolithinfaciens alone metabolized EA to uroM5, uroM6, and uroC (Fig. 1d); however, uroM6 was not metabolized to uroC when supplied as the initial substrate (Fig. 1e). In co-cultures of G. urolithinfaciens and E. bolteae (9-position dehydroxylation only) treated with EA, we could detect the intermediate urolithin M7 (3,8,10-trihydroxy-urolithin, uroM7) and the terminal metabolite uroA (Fig. 1d). Direct supplementation with uroM6 yielded uroM7 but no uroA (Fig. 1e), demonstrating that only E. bolteae can dehydroxylate uroM6 under these conditions. In contrast, co-cultures of G. urolithinfaciens and E. asparagiformis (9- and 10-position dehydroxylation) treated with EA yielded uroA as a major product and a trihydroxy-urolithin metabolite (Fig. 1d). Since we did not have a standard for this metabolite, it was tentatively assigned as urolithin G (3,4,8-trihydroxy-urolithin, uroG), likely derived from the 9- and 10-position dehydroxylation of uroM5 released by G. urolithinfaciens. Direct supplementation with uroM6 yielded exclusively uroA (Fig. 1e), highlighting differences between activities of urolithin-metabolizing Enterocloster spp. Collectively, these data show that simple 2-member communities can metabolize EA to different urolithins based on dehydroxylase activity and that urolithin features (hydroxylation status and regiochemistry) dictate whether metabolism will occur in whole cells, likely via substrate-specific transporters or dehydroxylase gene induction by urolithin intermediates.
A distinct dehydroxylase metabolizes 10-hydroxy urolithins
Based on the observation that urolithin intermediates are released by EA-metabolizing Gordonibacter spp. and that a subset of Enterocloster spp. can metabolize uroM6 to uroA, we sought to identify the genes and enzymes responsible for 10-position urolithin dehydroxylation, as previously done for the 9-position24. We reasoned that either the ucd-encoded UcdO (substrate-binding oxidoreductase) subunit of 9- and 10-position-metabolizing species like E. asparagiformis was promiscuous, acting at both the 9- and 10-positions of uroM6, or that a separate dehydroxylase was metabolizing the 10-position in these bacteria. Multiple sequence alignment of UcdO proteins from Enterocloster spp. (E. asparagiformis, E. citroniae, and E. pacaense) revealed that these sequences were >96.32% identical to the previously characterized E. bolteae UcdO (similarities >98.73%, Supplementary Fig. 1). Additionally, predicted urolithin binding site residues W345, Y375, F458, F464, Y536, Y624, and Y632 were conserved among all species (Supplementary Fig. 1), suggesting that substrate specificity should be identical in these bacteria. To validate that the ucd from E. asparagiformis encoded a 9-position-specific dehydroxylase, as observed in E. bolteae, we cloned and heterologously expressed the E. asparagiformis ucd operon in Rhodococcus erythropolis using a thiostrepton-inducible plasmid26 (pTipQC2-Ea_ucdCFO, Fig. 2a). In crude lysates, we could reliably detect the expression of UcdO in the insoluble fraction (Fig. 2b). Incubation of urolithins (uroM6, uroM7, uroC, or uroA) with ucd-expressing crude lysates confirmed that only 9-position hydroxyl groups, present on uroM6 and uroC, were dehydroxylated to uroM7 and uroA, respectively (Fig. 2c, d). We did not observe any 10-position dehydroxylation in uroM6 or uroM7-treated lysates, demonstrating that the E. asparagiformis ucd operon specifically dehydroxylates 9-hydroxy urolithins. These results suggest that a different, unidentified operon is responsible for 10-position dehydroxylation in some Enterocloster spp.
a Map of the pTipQC2-Ea_ucdCFO plasmid for the heterologous expression of the E. asparagiformis (Ea) ucd operon in R. erythropolis. Plasmid map generated in Benchling. ori: origin of replication (E. coli); bla beta-lactamase gene, tipA thiostrepton-inducible transcriptional activator, cmrA chloramphenicol efflux MFS transporter, tsr thiostrepton resistance gene, rep theta-type replicase (R. erythropolis). b Representative SDS-PAGE gel (10% bis-tris) of insoluble (I) and soluble (S) fractions from induced R. erythropolis pTipQC2-MCS and pTipQC2-Ea_ucdCFO crude lysates stained with colloidal Coomassie dye. Bands for UcdC, UcdF, and UcdO are expected at 31, 18, and 86 kDa, respectively. The uncropped gel is shown in Supplementary Fig. 9. c Aligned chromatograms (extracted ion intensities for [M-H]−: 259, 243, and 227) of DMSO or urolithin-treated crude lysates of R. erythropolis transformed with pTipQC2-Ea_ucdCFO. NADH (2.2 mM) was added to each lysate. Crude lysates were treated with DMSO (black), uroM6 (grey), uroM7 (red), or uroC (blue-grey, 200 μM each) for 48 h at room temperature anaerobically. Retention times of urolithin standards are shown with grey dotted lines. Data from n = 1 of 3 biological replicates are shown. d Quantification of urolithin extracted ion chromatogram (EIC) peak areas in crude lysates of R. erythropolis transformed with pTipQC2-MCS (multiple cloning site) or pTipQC2-Ea_ucdCFO plasmids. Data are shown as means ± SEM (n = 3 biological replicates) and are related to (c).
Ucd and Uxd dehydroxylases are induced by uroM6
Previous work from our lab has shown that dehydroxylation of urolithins by Enterocloster spp. is an inducible process triggered by substrate urolithins24. Thus, we hypothesized that, upon exposure to uroM6 (Fig. 1a), Enterocloster spp. would express Ucd and urolithin 10-position (X) dehydroxylase (Uxd) proteins. Therefore, we individually incubated E. asparagiformis, E. citroniae, and E. pacaense with uroM6 (or DMSO as a vehicle control), pooled isolates by treatment group, and performed reference-based proteomics on pooled lysates (Fig. 3a). To ensure that target enzymes were being expressed, we collected samples that were actively metabolizing uroM6. In all 3 tested isolates, uroA could be detected after a 4 h incubation with uroM6 (Fig. 3b). Filtering proteins by presence/absence revealed that 44 individual proteins were overrepresented in the uroM6 group (Fig. 3c). Among these, the most abundant proteins induced by uroM6 mapped to the ucd operon-encoded proteins UcdC and UcdO (Fig. 3d). However, we detected distinct xanthine dehydrogenase family proteins and FAD binding domain-containing proteins, which we annotated as putative Uxd proteins (Fig. 3d). The predicted FAD coenzyme-binding subunit (UxdC) was detected in all 3 species, while the substrate-binding oxidoreductase subunit (UxdO) was only detected in E. pacaense. Additionally, the molybdopterin cytosine dinucleotide (MCD) cofactor biosynthesis protein MogA (MOSC domain-containing protein) and the MCD cofactor chaperone protein XdhC (XdhC/CoxI family protein)27 were expressed, though to lesser extents than Ucd and Uxd proteins (Fig. 3d).
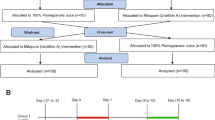
a Untargeted proteomics experimental design scheme. Treated isolates of E. asparagiformis (Ea), E. citroniae (Ec), and E. pacaense (Ep) were pooled by treatment group and lysed. Protein abundance was then determined by untargeted uHPLC-MS/MS. The scheme was generated using graphical elements from the NIAID NIH BIOART Source (bioart.niaid.nih.gov/bioart/42) and from BioRender. Castagner, B. (2025) https://BioRender.com/khbff31. b Aligned chromatograms (λ = 305 nm) of urolithins in DMSO or uroM6-treated (50 μM) mid-exponential phase isolates grown in mABB+H (4 h treatment with urolithins). Retention times of urolithin standards are shown with grey dotted lines. Data from n = 1 of 2 biological replicates are shown. c Venn diagram based on protein presence/absence in the untargeted proteomics dataset (from n = 2 biological replicates). d Heatmap of protein abundance (based on the average of total spectra from n = 2 biological replicates) for the 25 most abundant proteins detected in the uroM6 treatment group. Protein annotations from NCBI are shown on the right (along with the related taxon between brackets), and custom annotations are shown on the left. e Map of the pRG-Ea_uxdFOC plasmid for the heterologous expression of the E. asparagiformis (Ea) uxd operon in C. sporogenes. Plasmid map generated in Benchling. ori: origin of replication (E. coli); cat: chloramphenicol acetyltransferase; Pfdx-rbG: promoter of the C. sporogenes ferredoxin gene with riboswitch G. f Offset chromatograms (λ = 305 nm) of urolithins in uroM6-treated (100 μM) mid-exponential phase wild type (WT) and pRG-Ea_uxdFOC (uxd) C. sporogenes (Cs) cultures grown in BHI ± 30 μg/mL thiamphenicol (48 h treatment with uroM6). Retention times of urolithin standards are shown with grey dotted lines. Data from n = 1 biological replicate for each strain/transformant are shown. WT wild type; uxd #: Cs pRG-Ea_uxdFOC clone number. g Aligned chromatograms (λ = 305 nm) of urolithins in DMSO (black) or uroM7-treated (red, 100 μM) mid-exponential phase Cs WT, pRG-eGFP (eGFP), and uxd 1 cultures grown in BHI ± 30 μg/mL thiamphenicol (96 h treatment with DMSO or uroM7). Retention times of urolithin standards are shown with grey dotted lines. Data from n = 1 of 3 biological replicates for each strain/transformant are shown.
We next sought to validate the function of Uxd proteins through heterologous expression and noticed that, like Ucd, Uxd protein coding sequences were arranged in an operon with the following arrangement: uxdFOC. Therefore, we cloned the E. asparagiformis uxd operon into a shuttle plasmid to heterologously express UxdF, UxdO, and UxdC proteins in R. erythropolis (pTipQC2-Ea_uxdFOC). Unfortunately, electroporation of R. erythropolis with pTipQC2-Ea_uxdFOC did not yield any transformants capable of producing all three Uxd proteins, despite multiple attempts at transformation. We therefore searched the literature for alternative heterologous expression hosts that were anaerobic, genetically tractable, and ideally capable of xanthine dehydrogenase family protein expression. Recent studies demonstrated that Clostridium sporogenes, found in anaerobic soil and gut environments, is an appropriate host for the heterologous expression of xanthine dehydrogenase family proteins like 2,8-dioxopurine dehydrogenase (DOPDH)28,29. C. sporogenes encodes the molybdenum cofactor cytidylyltransferase MocA (WP_003494683.1) and the MCD cofactor chaperone XdhC (WP_033060855.1). These proteins, among others, participate in MCD biosynthesis and xanthine dehydrogenase family protein maturation30,31. In addition, the RiboCas series of plasmids, which utilize theophylline-inducible riboswitches to control Streptococcus pyogenes Cas9 expression, has been used to delete the spoIIE gene in multiple Clostridium spp., including C. sporogenes32. Based on the genotype and genetic tractability of C. sporogenes, we hypothesized that it would be an appropriate host for heterologous expression of Uxd proteins.
We therefore cloned the E. asparagiformis uxd operon and replaced the Cas9-encoding region of pRG-Cas2 to produce the pRG-Ea_uxdFOC shuttle plasmid (Fig. 3e). This plasmid was transformed into C. sporogenes via conjugation and resulting transformants were screened for protein expression and 10-position urolithin dehydroxylase activity. There were no obvious differences in protein expression between the wild type C. sporogenes and 10 randomly picked transformants (Supplementary Fig. 2). However, all 10 transformants were able to convert uroM6 to uroC (Fig. 3f). We further validated these findings by incubating uroM7, which lacks a catechol, with wild type C. sporogenes, an enhanced GFP (eGFP)-expressing control transformant, or a Uxd-expressing transformant. No background urolithins were detected in DMSO-treated media or cultures (Fig. 3g). In uroM7-treated cultures, uroA was detected solely in the Uxd-expressing C. sporogenes transformant, demonstrating the 10-position urolithin dehydroxylase activity conferred by the uxd operon (Fig. 3g). Collectively, these data demonstrate that a subset of Enterocloster spp. express distinct enzymes (Ucd and Uxd), along with their accessory proteins, that regioselectively dehydroxylate uroM6, uroM7, and uroC.
The ucd and uxd operons co-localize in urolithin-metabolizing Enterocloster spp. genomes
Since ucd and uxd operons appeared to be restricted to a subset of Enterocloster spp., we wondered about the prevalence of these operons both between and within Enterocloster spp. Therefore, we downloaded and queried 329 Enterocloster spp. genomes from the National Center for Biotechnology Information (NCBI) genomes database for the presence of ucd and uxd at the nucleotide level. As anticipated, the ucd operon was detected in E. asparagiformis, E. bolteae, E. citroniae, and E. pacaense genomes, whereas the uxd operon was detected in E. asparagiformis, E. citroniae, and E. pacaense (Fig. 4a). There was no evidence of these operons in other Enterocloster spp. genomes, including metagenome-assembled genomes (MAGs). All genomes of urolithin-metabolizing taxa encoded uxd and/or ucd with the exception of one E. asparagiformis MAG (GCA_040307595.1) which had neither of the operons (Fig. 4a). By performing searches using the basic local alignment search tool (BLAST) for sequences similar to the E. asparagiformis uxd, we observed that E. asparagiformis, E. citroniae, and E. pacaense had the highest sequence identities (id) at the nucleotide level (>77% id and >98% coverage, Supplementary Fig. 3). In addition, these 3 taxa had high sequence id for UxdO (>93% id and 100% coverage, Supplementary Fig. 4) and UxdC (>75% id and >97% coverage, Supplementary Fig. 5) at the protein level. However, we identified one bacterium, named Lacrimispora sp. 210928-DFI.3.58 (GCA_020554865.1) in the NCBI, which had a nucleotide sequence with 66% id to uxd with 92% coverage (Supplementary Fig. 3) and proteins with 77% and 63% id to UxdO and UxdC, respectively (>97% coverage, Supplementary Fig. 3, 4). This taxon is annotated as Enterocloster sp900753815 in the Genome Taxonomy Database (GTDB) but was missed in the previous Enterocloster spp. genome search (Fig. 4a) due to its misclassification in the NCBI genomes database (Methods). All other protein hits from the BLAST searches had identities <50% and <57% compared to UxdO and UxdC, respectively. Altogether, ucd and uxd operons, including some homologs, are highly prevalent within a subset of Enterocloster spp.
a Heat map of ucd (blue-grey) and uxd (red) operon presence based on BLASTn matches (% identity >60 and length >300) in Enterocloster spp. genomes downloaded from the NCBI. Genomes (n = 329) are grouped according to Genome Taxonomy Database (release 10-RS226) nomenclature. b Comparisons between ucd and uxd genomic contexts in urolithin-metabolizing E. asparagiformis (Ea), E. citroniae (Ec), E. pacaense (Ep), and E. bolteae (Eb). Alignments were performed using the clinker tool using genome accessions in Supplementary Table 7. Protein sequences with identities ≥40% are connected by lines. Bold lines indicate xanthine dehydrogenase family proteins within and between ucd and uxd-encoded proteins. c Representative domain annotations for E. asparagiformis ucd and uxd-encoded proteins. InterPro accessions are denoted below each annotation. FAD flavin adenine dinucleotide, BD binding domain, XDH xanthine dehydrogenase, MoCo molybdenum cofactor, CO carbon monoxide, DH dehydrogenase. d AlphaFold3 (AF3)-predicted protein structures of the Ea Ucd (shades of blue-grey) and Uxd (shades of red) protein complexes. A single trimer of the predicted heterohexamer is shown for clarity. FAD ligands were predicted by AF3. The molybdopterin cytosine dinucleotide cofactor (MCD) and 2Fe-2S clusters were derived from the superposition of AF3 structures onto the Thauera aromatica 4-hydroxybenzoyl-CoA reductase (PDB 1RM6). The 4Fe-4S cluster near the FAD (shown in e) was omitted as it is not common to both Ucd and Uxd enzymes. e Electron transport chain ligands in the Thauera aromatica 4-hydroxybenxoyl-CoA dehydrogenase enzyme complex (PDB 1RM6).
Next, we compared proteins in the vicinity of the ucd and uxd operons in urolithin-metabolizing Enterocloster spp. to investigate their genomic contexts. The ucd and uxd operons were in close proximity (<5 kb between ucdO and uxdF) in 10-position metabolizing species (Fig. 4b). In these bacteria, Uxd proteins (UxdF, UxdO, and UxdC) were highly similar (>94, >96, and >83% protein similarity, respectively, Supplementary Fig. 6a, b). To our surprise, E. bolteae, which only metabolizes the 9-position of urolithins, possessed a uxd homolog near its ucd operon (Fig. 4b). In E. bolteae, the uxd operon homolog was more distant from the ucd (10.2 kb between ucdO and uxdF), and its Uxd proteins were less similar (<66% protein similarity) compared to those from 10-position metabolizing species (Supplementary Fig. 6a, b). Notably, the substrate-binding UxdO protein was >96% similar between E. asparagiformis, E. citroniae, and E. pacaense, but less than 63% similar (<44% protein id) when compared to E. bolteae (Supplementary Fig. 7). These different protein sequences could explain the lack of 10-position dehydroxylase activity observed in E. bolteae.
We next compared domain annotations between ucd and uxd-encoded proteins of E. asparagiformis, which serves as our reference in this study. Iron-sulfur cluster-binding proteins (UcdF and UxdF) were similar in length, and both possessed two 2Fe-2S cluster binding domains, characteristic of prokaryotic xanthine dehydrogenase family proteins, which use these proteins to shuttle electrons between FAD and the MCD cofactor (Fig. 4c)33. The MCD- and substrate-binding oxidoreductase subunits (UcdO and UxdO) both had functional domains typical of xanthine dehydrogenase family proteins: an XDH a/b hammerhead and molybdenum cofactor binding domains (Fig. 4c). The FAD-binding subunits (UcdC and UxdC) were more dissimilar, mostly due to the additional length of UxdC, which carried a 4Fe-4S cluster binding domain and an FAD/NAD(P) binding domain in addition to the FAD binding domain common to UcdC and UxdC (Fig. 4c). AlphaFold3 modeling34 of Ucd and Uxd complexes produced similar overall structures resembling other xanthine dehydrogenase family proteins (Fig. 4d)35. By superposing these predicted structures onto the crystal structure of the Thauera aromatica 4-hydroxybenzoyl-CoA reductase complex (dehydroxylase), we could place the MCD and 2Fe-2S clusters into UcdO and UxdO, and UcdF and UxdF, respectively. These ligands form an electron transport chain, likely enabling the dehydroxylation of substrates (e.g. urolithins) to their reduced products (Fig. 4e). In sum, ucd and uxd operons, which co-localize in urolithin-metabolizing Enterocloster spp. genomes, encode xanthine dehydrogenase proteins with similar predicted folds.
Uxd expression is regulated by 9-hydroxy urolithins
We next wondered whether uxd operon substrates (10-hydroxy urolithins) induce the expression of uxd (Fig. 5a). Therefore, E. asparagiformis, grown to the mid-exponential phase, was treated with DMSO, uroM6, uroM7, uroC, or uroA (structures in Fig. 1a), and total RNA was extracted for RT-qPCR analysis focusing on ucdO and uxdO gene expression. Curiously, uxd expression followed the same pattern as ucd (Fig. 5b); that is, only 9-hydroxy urolithins (uroM6 and uroC) induced uxd expression. UroM7, which bears a hydroxyl group at the 10-position, and uroA, which is dehydroxylated at both the 9- and 10-positions, did not induce the expression of either operon (Fig. 5b). When we quantified urolithin concentrations in matched cultures, where total RNA and urolithins were isolated from the same samples, we observed dehydroxylation of uroM6 at both the 9- and 10-positions and dehydroxylation of uroC at the 9-position; however, uroM7 was unchanged (Fig. 5c). To validate that 9-hydroxy urolithins are required for 10-position dehydroxylation, E. asparagiformis cultures were induced with 9-hydroxy urolithins lacking a 10-hydroxyl group: urolithin D (3,4,8,9-tetrahydroxy-urolithin, uroD), uroC, or isouroA (Fig. 5d). After a 2 h incubation, 9-position dehydroxylation was observed for uroD- and uroC-treated cultures, yielding uroG and uroA, respectively; however, no dehydroxylation was observed for isouroA at this timepoint since uroB was not detected in culture supernatants (Fig. 5e). Following this 2 h induction, E. asparagiformis cells were washed and resuspended in phosphate buffered saline (PBS) to minimize protein synthesis, then re-treated with urolithins to assess whether the 10-hydroxy urolithins could be dehydroxylated. In DMSO-induced resting cell suspensions, partial 9- and 10-position dehydroxylation was observed for uroM6 and uroC re-treated samples, suggesting basal activity or inducibility in these cell suspensions. Nevertheless, uroM7 remained unchanged (Fig. 5f), as previously observed in exponential phase cultures (Fig. 5c). In 9-hydroxy urolithin-induced resting cell suspensions, significant uroA production was observed for nearly all substrates (compared to DMSO-induced cultures). As anticipated, 10-position dehydroxylation of uroM7 to uroA was only observed in cell suspensions induced with 9-hydroxy urolithins (Fig. 5f), demonstrating the co-regulation of ucd and uxd operons in E. asparagiformis.
a Structure of urolithin M6 (uroM6). The 9- and 10-position hydroxyl groups are highlighted in grey and red, respectively. b RT-qPCR expression of E. asparagiformis (Ea) ucdO and uxdO target after treatment with DMSO or urolithins (100 μM for 2 h in mABB+H media, n = 3 biological replicates). Gene expression is displayed as log2FC (urolithin/DMSO); repeated-measures two-way ANOVA (matched by replicate and target gene) with Tukey’s multiple comparisons test. c Quantification of urolithin concentrations in matched samples from (b) determined by LC-MS (n = 3 biological replicates). d General urolithin structure with R groups showing hydroxyl group placement for urolithin D (uroD), urolithin C (uroC), and isourolithin A (isouroA). e Aligned chromatograms (λ = 305 nm) of DMSO or 9-hydroxy urolithin-induced (50 μM for 2 h) E. asparagiformis culture supernatants. Retention times of urolithin standards are shown with grey dotted lines. Data from n = 1 of 3 biological replicates are shown. f Quantification of urolithin concentrations in 9-hydroxy urolithin-induced (50 μM) E. asparagiformis cell suspensions re-treated with DMSO, uroM6, uroM7, or uroC (100 μM each, n = 3 biological replicates); two-way ANOVA (based on uroA concentration) with Dunnett’s multiple comparisons test (compared to re-treatment matched DMSO-induced samples). Bar colors are the same as in c. Data in (b, c, f) are shown as means ± SEM. **P < 0.01, ***P < 0.001, ****P < 0.0001.
In our previous study on the ucd operon, we performed RNA-seq experiments on DMSO and uroC-treated E. asparagiformis and E. bolteae24. In E. asparagiformis, uxd operon genes were substantially induced by uroC (log2FC(uroC/DMSO) Ea_uxdF: 4.9, Ea_uxdO: 5.8, Ea_uxdC: 6.0, Supplementary Fig. 8a); however, induction of the E. bolteae uxd homolog was less drastic (log2FC(uroC/DMSO) Eb_uxdF: 1.9, Eb_uxdO: 1.5, Eb_uxdC: 1.4, Supplementary Fig. 8b). These differences in operon induction could also contribute to the lack of 10-position dehydroxylase activity observed in E. bolteae.
Discussion
The human gut microbiota encodes millions of proteins with functions that remain largely unknown2,36. This is notably the case for polyphenol-metabolizing enzymes (also termed polyphenol utilization proteins37), which despite their widespread roles across ecosystems38,39, remain poorly characterized and misannotated in reference databases40. As urolithins can impact host health41, understanding how these metabolites are produced can inform therapeutic microbiota modulation strategies41,42. In this study, we used untargeted proteomics to uncover a 10-position urolithin dehydroxylase (Uxd) in a subset of urolithin-metabolizing Enterocloster spp. (E. asparagiformis, E. citroniae, and E. pacaense). Using diverse urolithins, which varied both in the number of hydroxyl groups and their positions on the parent structure, we characterized the regulation and substrate requirements for the induction and activity of urolithin dehydroxylases in E. asparagiformis.
The metabolism of dietary ellagitannins by the gut microbiota requires a diverse community of microorganisms harboring specialized enzymes (hydrolases, decarboxylases, and regioselective dehydroxylases)43. We showed that EA-treated Gordonibacter spp. release urolithin intermediates in culture media, either through diffusion, leakage, or cell lysis44, that become substrates for urolithin-metabolizing Enterocloster spp. In simple 2-member communities, G. urolithinfaciens paired with E. bolteae or E. asparagiformis could cooperate to metabolize EA to the terminal metabolite uroA (and to the intermediates uroM7 and uroG, respectively). These experiments also highlighted that G. urolithinfaciens cannot metabolize exogenous uroM6 (supplied directly), despite its ability to convert EA or uroM5 to uroC via uroM6 (Fig. 1a)45. This observation could be explained by a lack of uroM6 transport or by a failure to induce Eadh2 (10-position dehydroxylase) in the ellagic acid metabolism (eam) gene cluster46. Additionally, different urolithin intermediates accumulated in co-cultures depending on both their hydroxylation status upon release by G. urolithinfaciens and the dehydroxylases present in each Enterocloster spp. Indeed, along with uroA, substantial accumulation of uroM7 was observed with E. bolteae, whereas only some minor amounts of uroG were observed with E. asparagiformis. The abovementioned results indicate that urolithins could undergo substrate shunting whereby intermediates get trapped in alternative pathways, unable to be further dehydroxylated due to issues with transport or dehydroxylase induction. Additional studies are required to determine how the exchange of urolithin intermediates (cross-feeding) affects the fitness of urolithin-metabolizing species47 and whether any species or strain level differences could alter the metabolic fate of urolithins in microbial communities.
Despite belonging to different phyla, both Gordonibacter spp. (Actinomycetota phylum) and a subset of urolithin-metabolizing Enterocloster spp. (Bacillota phylum) have evolved distinct molybdoenzymes to dehydroxylate the 10-position of urolithins (convergent evolution). The Eadh2 enzyme, part of the recently described eam gene cluster in G. urolithinfaciens, belongs to the DMSO reductase family46, while the Uxd complex belongs to the xanthine dehydrogenase family of enzymes. Most characterized polyphenol dehydroxylases belong to the DMSO reductase family48. Examples of these enzymes include the Eggerthella spp. hydrocaffeic acid dehydroxylase (Hcdh), dopamine dehydroxylase (Dadh), catechin dehydroxylase (Cadh) and epicatechin dehydroxylase (eCadh), and Gordonibacter spp. hydrocaffeic acid dehydroxylase (Hcdh) and 3,4-dihydroxyphenylacetic acid (DOPAC) dehydroxylase (Dodh)48,49,50. Interestingly, DMSO reductase family dehydroxylases require a catechol with a para-hydroxyl group for dehydroxylation48,49, whereas xanthine dehydrogenase family dehydroxylases, like Ucd, Uxd, and 4-HBCR, do not require this structural feature for enzyme activity24,51. Indeed, isouroA, uroM7, and 4-hydroxybenzoyl-CoA are dehydroxylated at positions lacking a catechol motif by the abovementioned enzymes, respectively. These differences in molybdoenzyme family and substrate features suggest a different mechanism of uroM6 dehydroxylation for Eadh2 and Uxd46.
Our comparative genomics analyses highlighted that a subset of urolithin-metabolizing Enterocloster spp. carry a uxd operon located near their ucd operon. Surprisingly, E. bolteae possessed a uxd operon homolog in its genome, though the resulting Uxd protein sequences were dissimilar (<66% protein similarity) to those from 10-position urolithin metabolizing species. The endogenous substrate and function of this uxd homolog is currently unknown, though it is unlikely to dehydroxylate urolithins since E. bolteae only dehydroxylates urolithins at the 9-position via ucd. The evolutionary origins of the ucd and uxd operons are rather puzzling as both operons are co-regulated and encode xanthine dehydrogenase family proteins that dehydroxylate urolithins regioselectively, yet their genetic arrangement (ucdCFO vs. uxdFOC) and coenzyme-binding subunits differ substantially. These features suggest that both operons may have undergone extensive genetic rearrangements throughout their evolution. Nevertheless, both AlphaFold3-predicted enzyme complexes adopted similar folds and subunit arrangements. Intriguingly, the UxdC coenzyme-binding subunit was predicted to bind 2 FAD molecules, at opposite ends, in agreement with the InterPro protein domain annotations. It is likely that UxdC utilizes 4Fe-4S clusters to shuttle electrons between the distal FAD towards the proximal FAD (next to UxdF), analogous to the role of the 4Fe-4S cluster in the T. aromatica 4-HBCR35.
In this study, we focused on Gordonibacter and Enterocloster spp. since these gut-associated genera are both prevalent in human populations52 and have been shown to cooperatively metabolize EA in vivo18. It is worth noting that a growing list of bacterial strains has been reported to either partially or fully metabolize EA to terminal urolithin metabolites like uroA, isouroA, and uroB. These include Ellagibacter isourolithinifaciens20, Streptococcus thermophilus FUA32953, Lactococcus garvieae FUA00954, Enterococcus faecium FUA02755, Limosilactobacillus fermentum FUA03356, and Bifidobacterium pseudocatenulatum INIA P81557. Unfortunately, with the exception of E. isourolithinifaciens46, many of these strains lack publicly accessible genomes and are generally not commercially available, making polyphenol-metabolizing enzyme discovery and validation difficult. Future enzyme discovery efforts using these strains, and others yet to be discovered, would enable a better understanding of the diversity of enzymes involved in EA and urolithin metabolism across taxa in the gut microbiota.
The transcription of gut bacterial xenobiotic metabolizing enzymes is often an inducible process triggered upon exposure to their substrates58,59. In urolithin-metabolizing Enterocloster spp., the ucd operon (9-position dehydroxylase) is induced upon treatment with 9-hydroxy urolithins like uroM6, uroC, and isouroA24. Surprisingly, uxd transcription was not triggered by the Uxd substrate uroM7 (3,8,10-trihydroxy-urolithin) and no 10-position dehydroxylation was observed when this compound was supplied as an initial substrate. Instead, ucd induction by its substrates was required for uxd transcription and for the conversion of uroM7 to uroA. These findings suggest that the ucd and uxd operons, which are separated by less than 3 kb in E. asparagiformis genome, share regulatory proteins that coordinate their transcription, along with accessory proteins for MCD cofactor biosynthesis, after sensing 9-hydroxy urolithins. This regulatory mechanism is consistent with the urolithin metabolites that are released by EA-metabolizing Gordonibacter spp. (uroM5, uroM6, and uroC), which all feature a 9-position hydroxyl group. Thus, Gordonibacter spp. and Enterocloster spp. may be particularly well adapted to cooperatively metabolize EA in the gut, as reflected by their complementary urolithin-metabolizing enzymes.
In sum, our study highlights the genes, proteins, and substrate features underlying urolithin metabolism in a subset of Enterocloster spp. Dietary polyphenols and their postbiotic metabolites are gaining widespread attention for their roles in host health60, though many of molecular mechanisms underlying their production by gut bacteria remain unknown. Enzyme discovery efforts described here and elsewhere48,50,61,62 not only contribute to our basic understanding of polyphenol-bacteria ecology but also explain some of the inter-individual differences in polyphenol metabolism (metabotypes7) observed in human populations.
Methods
General reagents
A complete list of reagents and chemicals used in this study is provided in Supplementary Table 1. Reference numbers for commercial media, kits, and master mixes are provided in relevant sections below.
Anaerobic bacterial strains and culturing conditions
Strains of bacteria used in this study are listed in Supplementary Table 2. Enterocloster spp. and Gordonibacter spp. were grown from glycerol stocks on mABB+H (recipe below) or Brain Heart Infusion (BHI, Difco #237500) agar plates for 48–72 h at 37 °C in a Coy vinyl anaerobic chamber, which was maintained with a gas mixture of 3% H2, 10% CO2, 87% N2. To make overnight cultures of bacteria, a single isolated colony was inoculated into 5 mL of medium. For Enterocloster spp., media were mABB+H or BHI. For Gordonibacter spp., media were BHI or BHIrf (BHI + 1% w/v arginine + 10 mM sodium formate)50. Cultures were incubated at 37 °C between 16 and 24 h for Enterocloster spp. and 48 h for Gordonibacter spp.
Modified anaerobe basal broth with hemin (mABB+H)
For 1 L of modified anaerobe basal broth (mABB, based on Oxoid Anaerobe Basal Broth), the following components were dissolved in MilliQ water, then autoclaved: 16 g peptone, 7 g yeast extract, 5 g sodium chloride, 1 g starch, 1 g D-glucose monohydrate, 1 g sodium pyruvate, 0.5 g sodium succinate, 1 g sodium thioglycolate, 15 g agar (for plates). The autoclaved solution was allowed to cool, then the following filter-sterilized solutions were added aseptically: 10 mL of 100 mg/mL L-arginine-HCl, 10 mL of 50 mg/mL L-cysteine, 8 mL of 50 mg/mL sodium bicarbonate, 50 μL of 10 mg/mL vitamin K1, 20 mL of 50 mg/mL dithiothreitol, and, for mABB+H, 10 mL of 0.5 mg/mL haemin. The media was then placed in the anaerobic chamber and allowed to reduce for at least 24 h prior to its use in experiments.
Genomic DNA extraction of isolates and identity validation
Genomic DNA was extracted from 0.5 to 1 mL of overnight culture using the One-4-All Genomic DNA Miniprep Kit (BioBasic #BS88503) according to the manufacturer’s instructions. The identities of all bacteria in this study were validated by full-length (V1-V9) 16S rRNA sequencing using primers 16S_V1_27_f and 16S_V9_1492_r listed in Supplementary Table 3. The purified genomic DNA was used as a template for PCR reactions using the Q5 High-Fidelity polymerase (NEB #M0491). PCR tubes were placed in a thermal cycler and targets ( ~1.5 kb) were amplified according to the following cycling conditions: 30 s at 98 °C, 30 cycles (10 s at 98 °C, 20 s at 60 °C, 45 s at 72 °C), 2 min at 72 °C, and hold at 10 °C. 5 μL of the reaction was mixed with 6X Gel Loading Dye (NEB #B7024A) and loaded onto a 1% agarose gel (made with 1X TAE buffer) containing FroggaStain (FroggaBio #FB555). PCR product sizes were compared to the Quick-Load Purple 1 kb Plus DNA Ladder (NEB #N0550).
PCR products ( ~1.5 kb) were purified using the Monarch PCR & DNA Cleanup Kit (NEB #T1030) according to the manufacturer’s instructions for products <2 kb. Purified 16S PCR products were eluted in nuclease-free water, quantified using the Qubit dsDNA HS assay kit (Invitrogen #Q32851), and adjusted to 30 ng/μL. Linear/PCR product sequencing was performed by Plasmidsaurus using Oxford Nanopore Technology with custom analysis and annotation.
Gordonibacter spp. ellagic acid metabolism assays
Overnight cultures of Gordonibacter spp. (Supplementary Table 2) were diluted 1/50 in fresh BHIrf media. Ellagic acid (10 mM stock in DMSO) or an equal volume of DMSO were added to a final concentration of 100 μM. Cultures were incubated at 37 °C under anaerobic conditions, and samples were collected after 2 days. For urolithin detection in supernatants, 300 μL of culture was centrifuged at 5000 x g for 3 min, and the supernatant was collected. Metabolites in cultures and supernatants were extracted following Extraction Method C.
Enterocloster spp. treatments with urolithins
Urolithins used in this study (Supplementary Table 1) were dissolved in DMSO to a concentration of 10 mM. Overnight cultures of Enterocloster spp. (Supplementary Table 2) were diluted 1/50 into fresh media and incubated at 37 °C in an anaerobic chamber. After 5–7 hours (5 h for E. asparagiformis and E. pacaense, and 7 h for E. citroniae) of incubation, 10 mM urolithins (or an equivalent volume of DMSO) were added to the growing cultures at a final concentration of 50 or 100 μM for protein expression or RNA expression, respectively. For protein expression analyses, the cultures were incubated for an additional 4 h. For RNA expression analyses and inducibility tests, the cultures were incubated for an additional 2 h. Metabolites in cultures were extracted following Extraction Method A.
Co-cultures of G. urolithinfaciens and Enterocloster spp
Overnight cultures of G. urolithinfaciens (Gu), E. bolteae (Eb), and E. asparagiformis (Ea) were diluted 1/100 (each) into fresh BHI media (5 mL) according to the following four treatment groups: BHI (sterile), Gu, Gu + Eb, Gu + Ea. Each treatment group was split into 3 × 500 µL aliquots and treated with DMSO, ellagic acid (10 mM stock in DMSO), or urolithin M6 (10 mM stock in DMSO) to a final concentration of 100 μM. Samples were incubated at 37 °C under anaerobic conditions, and 200 μL samples were taken after 5 days and stored at −70 °C until metabolite extraction. Metabolites in cultures were extracted following Extraction Method A.
Urolithin extraction from bacterial cultures
Frozen (−70 °C) bacterial cultures were thawed at room temperature. For quantification of urolithin concentrations, urolithin standards (stock 10 mM in DMSO) were spiked into separate media aliquots immediately before extraction.
Extraction Method A: This method was used for cultures and their supernatants. Samples were diluted with an equal volume of MeOH, vortexed briefly, and incubated at 50 °C for 10 min. Samples were centrifuged at 20,000 x g for 5 min to pellet insoluble material, then transferred to LC-MS vials. Urolithins were then analyzed by LC-MS.
Extraction Method B: This method was used to extract urolithins from crude bacterial lysates. Lysates were diluted with 3 volumes of MeOH, vortexed briefly, and incubated at room temperature for 10 min. Samples were centrifuged at 20,000 x g for 5 min to pellet insoluble material, then transferred to LC-MS vials. Urolithins were then analyzed by LC-MS.
Extraction Method C: This method was used for cultures and supernatants. Three volumes of ethyl acetate with 0.1% formic acid were added to the sample and vortexed briefly. Samples were centrifuged at 10,000 x g for 5 min to separate the organic and inorganic layers. The organic layer was collected and dried using a rotary evaporator. Concentrated samples were dissolved in 50% methanol, centrifuged at 20,000 x g for 5 min to pellet insoluble material, and then transferred to LC-MS vials. Urolithins were then analyzed by LC-MS.
LC-MS method to quantify urolithins
Samples were injected (10 μL) into a 1260 Infinity II Single Quadrupole LC/MS system (Agilent) fitted with a Poroshell 120 EC-C18 4.6 × 50 mm, 2.7 μm column (Agilent #699975-902) with a compatible guard column (Agilent #820750-911) incubated at 30 °C. The mobile phase was composed of MilliQ water + 0.1% formic acid (solvent A) and acetonitrile + 0.1% formic acid (solvent B). The flow rate was set to 0.7 mL/min. The gradient was as follows: 0-8 min: 10-30% B, 8–10 min: 30–100% B, 10–13.5 min: 100% B isocratic, 13.5–13.6 min: 100–10% B, then 13.6–15.5 min: 10% B. The multiple wavelength detector was set to monitor absorbance at 305 nm. The mass spectrometer (API-ES) was run in negative mode in both selected ion monitoring (SIM) and scan (100–1000 m/z) modes to validate peak identities. Capillary voltage was set to 3000 V, drying gas to 10.0 L/min, nebulizer pressure to 30 psig, and drying gas temperature to 350 °C. Peaks were validated based on retention times compared to spike-in standards and mass-to-charge ratios. To quantify urolithins, integrated peak areas (derived from Agilent ChemStation software) for the compounds of interest were compared to spike-in standards of known concentrations. When standards were not available or overlapped with other peaks, the extracted ion chromatogram peak area was used: ellagic acid [M-H]−: 301, urolithin M5 [M-H]−: 275, urolithin E [M-H]−: 259, urolithin D [M-H]−: 259, urolithin M7 [M-H]−: 243, urolithin M6 [M-H]−: 259, urolithin C [M-H]−: 243, urolithin G [M-H]−: 243, urolithin A [M-H]−: 227, isourolithin A [M-H]−: 227, urolithin B [M-H]−: 211. Blank runs of 50% MeOH were included at the beginning and end of LC-MS sequences to ensure proper column washing.
Plasmid construction in E. coli
All plasmids used in this study were constructed through DNA assembly using the NEBuilder HiFi DNA Assembly Master Mix (NEB #E2621). Linearized pTipQC2 plasmid DNA (Hokkaido Systems Science Co. #RE-0006) was generated through overnight ( ~16 h) restriction digestion with NdeI (NEB #R0111) and XhoI (NEB #R0146) in rCutSmart buffer at 37 °C. Linearized pRG_BB (backbone) plasmid DNA was generated through PCR amplification of the pRG-Cas2 plasmid (RiboCas plasmid series32) using the Q5 DNA polymerase (NEB # M0491). To reduce contamination from the pRG-Cas2 template, DpnI (NEB #R0176) was added to the pRG_BB PCR reaction after thermal cycling and incubated for 1 h at 37 °C. The Ea_ucdCFO and Ea_uxdFOC DNA inserts were generated through PCR amplification of ucd and uxd from E. asparagiformis genomic DNA using the Q5 DNA polymerase. The enhanced green fluorescent protein (eGFP) insert was generated through PCR amplification of eGFP from pACBB-eGFP plasmid DNA using the Q5 DNA polymerase. pACBB-eGFP was a gift from Claudia Schmidt-Dannert (Addgene plasmid #32551, http://n2t.net/addgene:32551, RRID: Addgene_32551).
In all cases, Q5 DNA polymerase PCR cycling was as follows: 30 s at 98 °C, 30 cycles (10 s at 98 °C, 20 s at Ta, reaction-specific extension time at 72 °C), 2 min at 72 °C. Primer sequences are listed in Supplementary Table 3 and specific PCR conditions (primers, Ta, and extension time) are listed in Supplementary Table 4. Restriction digest products and PCR amplicons were loaded into 0.6–1% w/v agarose gels (in 1X TAE) and purified using the Monarch DNA Gel Extraction Kit (NEB #T1020) according to the manufacturer’s instructions for different DNA sizes. Purified DNA concentrations were determined using the Qubit dsDNA HS assay kit (Invitrogen #Q32851), and fragments were mixed for HiFi DNA assembly at 50 °C for 1 h according to the manufacturer’s instructions. HiFi DNA assembly constructs (plasmid backbones and inserts) are provided in Supplementary Table 5.
HiFi DNA assembly reactions were diluted ¼ in nuclease-free water and 2 μL were transformed into homemade CaCl2 competent E. coli NEB10β (NEB #C3019) via heat shock. Cells were spread onto selective LB + 100 μg/mL ampicillin or LB + 30 μg/mL chloramphenicol agar plates, depending on the plasmid backbone, and incubated at 37 °C overnight. The next day, colonies were picked and grown in selective LB media. Plasmids were purified using the Plasmid DNA Miniprep Kit (BioBasic #BS414) and size was confirmed with a diagnostic restriction digest. Whole plasmid sequencing was performed by Plasmidsaurus using Oxford Nanopore Technology with custom analysis and annotation. GenBank accessions for sequenced plasmids are listed in Supplementary Table 3. We observed an unintentional mutation in the pRG-eGFP plasmid in the eGFP coding region (Y146F); however, this does not affect the intended use as a protein expression control. Plasmid maps were generated in Benchling (2025): https://benchling.com.
pTipQC2 plasmid transformation into R. erythropolis
Electrocompetent R. erythropolis were prepared by inoculating 50 mL NBYE (0.8% w/v nutrient broth, 0.5% yeast extract) + 0.05% Tween-80 media with 1 mL of a stationary phase (48–72 h growth from a single colony) R. erythropolis culture and grown aerobically for 16 h at 30 °C with shaking at 200 RPM. After this incubation, cells were pelleted at 5000 x g for 10 min at 4 °C and washed according to the following sequence: 2 washes of (10 mL of ice cold sterile MilliQ water), 10 mL of ice cold sterile 10% glycerol. The final pellet was resuspended in 2.5 mL of ice cold sterile 10% glycerol. The resuspended electrocompetent R. erythropolis were aliquoted (50 μL/aliquot), flash-frozen in liquid nitrogen, and stored at −80 °C long-term. For transformations, frozen electrocompetent cells were diluted in sterile MilliQ water, then 3 μL ( ~ 0.5-1 μg) of various plasmid DNA (isolated from E. coli) were added to appropriate tubes. Cells with plasmid DNA were transferred to 0.1 cm gap cuvettes (Bio-Rad #165-2089) and electroporated (1.8 kV, 25 μF, 200 Ω). Time constants were between 4.0 and 4.4 ms. The cuvette was immediately filled with 1 mL LB post shock and cells were allowed to recover at 30 °C for 3 h before plating 100 μL of undiluted culture and of the remaining concentrated recovery culture on LB + 30 μg/mL chloramphenicol plates at 30 °C. After 3–4 days of incubation, colonies were picked and grown in selective liquid LB + 30 μg/mL chloramphenicol at 30 °C with shaking at 200 RPM. Ucd protein-producing colonies were verified by inducing protein expression and assaying proteins by SDS-PAGE (described below).
Heterologous expression of Ucd proteins in R. erythropolis
All growth steps below were performed in selective media (LB + 30 μg/mL chloramphenicol) in aerobic conditions at 30 °C with shaking at 220 RPM. Single colonies of R. erythropolis transformed with pTipQC2-MCS (multiple cloning site) or pTipQC2-Ea_ucdCFO were inoculated into 15 mL selective media and grown for 72 h to produce overnight cultures. Overnight cultures were then thoroughly resuspended and diluted 1:10 into 25 mL fresh selective media and grown for 7-8 h until OD600 values reached ~0.6. Thiostrepton (5 mg/mL in DMSO) was added to a final concentration of 0.5 μg/mL, and cultures were incubated aerobically for 16 h at 30 °C to induce protein expression. The next morning, 20 mL from each culture were pelleted and resuspended in 4 mL of lysis buffer: 1 X PBS, 1 mM dithiothreitol (DTT), 1% Triton X-100, 2 mg/mL egg white lysozyme, and 1 tablet/10 mL cOmplete Mini, EDTA-free (Roche #11836170001). The resuspended cells in lysis buffer were incubated on ice for 1 h with shaking, then sonicated on ice using a Misonix Sonicator 3000 set to power level 3/10 according to the following sequence (aerobically, in a cold room): 20 s ON, 40 s OFF, for a total of 4 min ON. To evaluate dehydroxylase activity, crude lysates (100 μL) were treated with 2 μL of 10 mM urolithin solution (or DMSO vehicle) and with 8 μL of 30 mM NADH (dissolved in lysis buffer), then incubated anaerobically at chamber temperature (~28 °C) for 48 h. Metabolites in lysates were extracted following Extraction Method B.
SDS-PAGE analysis of Ucd proteins
Crude lysates described above were centrifuged for 2 min at 20,000 x g. The insoluble pellet was separated from the soluble supernatant. The insoluble pellet (from 100 μL of crude lysate) was resuspended in 100 μL of 1X reducing loading dye (62.5 mM Tris-HCl (pH 6.8), 2% w/v SDS, 10% glycerol, 0.01% w/v bromophenol blue, 38 mM DTT). The soluble fraction was diluted with 4X reducing loading dye to a final concentration of 1X. All samples were heated at 95 °C for 5 min, then 15 μL were loaded onto a 10% bis-tris polyacrylamide protein gel. Gels were fixed and stained with 10X colloidal Coomassie solution (0.4% w/v Coomassie Brilliant Blue R250, 10% w/v citric acid, 8% w/v ammonium sulfate, and 20% v/v methanol).
Protein extraction from uroM6-treated Enterocloster spp
All steps other than sonication were carried out under anaerobic conditions. To extract proteins, 10 mL of treated (50 μM urolithin M6 or DMSO for 4 h) Enterocloster spp. culture (see Enterocloster spp. treatments with urolithins) were pelleted (6500 x g for 3 min) and the supernatant was kept for LC-MS analysis of urolithins. The pellets of Ea, Ec, and Ep cultures treated with either uroM6 or DMSO were pooled in 10 mL of pre-reduced PBS, pelleted again, and resuspended in 0.6 mL of pre-reduced lysis buffer: 20 mM Tris, pH 7.5, 500 mM NaCl, 10 mM MgSO4, 10 mM CaCl2, and 1 tablet/10 mL cOmplete Mini, EDTA-free (Roche #11836170001). The resuspended pellet was then sonicated on ice using a Misonix Sonicator 3000 set to power level 2/10 according to the following sequence (aerobically, in a cold room): 20 s ON, 40 s OFF, for a total of 2 min ON. Tubes were centrifuged at 20,000 x g for 2 min to pellet insoluble particles, and the supernatant was kept for downstream proteomics analyses.
Proteomics analysis of uroC-treated Enterocloster spp
Extracted proteins were submitted for proteomic analysis at the Research Institute of the McGill University Health Centre (RI-MUHC). For each sample, protein lysates were loaded onto a single stacking gel band to remove lipids, detergents, and salts. The gel band was reduced with DTT, alkylated with iodoacetic acid, and digested with trypsin. Extracted peptides were re-solubilized in 0.1% aqueous formic acid and loaded onto a Thermo Acclaim Pepmap (Thermo, 75 μm ID X 2 cm C18 3 μm beads) precolumn and then onto an Acclaim Pepmap Easyspray (Thermo, 75 μm ID X 15 cm with 2 μm C18 beads) analytical column separation using a Dionex Ultimate 3000 uHPLC at 250 nL/min with a gradient of 2–35% organic (0.1% formic acid in acetonitrile) over 3 hours. Peptides were analyzed using a Thermo Orbitrap Fusion mass spectrometer operating at 120,000 resolution (FWHM in MS1) with HCD sequencing (15,000 resolution) at top speed for all peptides with a charge of 2+ or greater. The raw data were converted into *.mgf format (Mascot generic format) for searching using the Mascot 2.6.2 search engine (Matrix Science) against a custom Enterocloster spp. protein database composed of Enterocloster asparagiformis (GenBank Assembly GCA_025149125.1), Enterocloster citroniae (GenBank Assembly GCA_001078435.1), Enterocloster pacaense (GenBank Assembly GCA_900566185.1), and a database of common contaminant proteins. Mascot was searched with a fragment ion mass tolerance of 0.100 Da and a parent ion tolerance of 5.0 ppm. O-63 of pyrrolysine, carboxymethyl of cysteine, and j + 66 of leucine/isoleucine indecision were specified in Mascot as fixed modifications. Deamidation of asparagine and glutamine and oxidation of methionine were specified in Mascot as variable modifications.
The database search results were loaded into Scaffold Q+ Scaffold_5.0.1 (Proteome Sciences) for statistical treatment and data visualization. Scaffold (v5.3.0) was used to validate MS/MS-based peptide and protein identifications. Peptide identifications were accepted if they could be established at greater than 95.0% probability by the Peptide Prophet algorithm63 with Scaffold delta-mass correction. Protein identifications were accepted if they could be established at greater than 99.0% probability and contained at least 2 identified peptides. Protein probabilities were assigned by the Protein Prophet algorithm64. Proteins that contained similar peptides and could not be differentiated based on MS/MS analysis alone were grouped to satisfy the principles of parsimony. Proteins sharing significant peptide evidence were grouped into clusters. Protein quantification and differential expression were determined in Scaffold using the following parameters: Quantitative method was set to total spectra, and the minimum value was set to 0.5 in case proteins were not detected in one condition.
pRG plasmid conjugation into C. sporogenes
Homemade CaCl2 competent E. coli CA434 conjugal donor cells (carrying the R702 plasmid)65 were transformed with the pRG-Ea_uxdFOC plasmid via heat shock, allowed to recover in SOC medium (BioBasic #SD7009), and plated onto LB + 30 μg/mL chloramphenicol (for the pRG plasmids) + 15 μg/mL tetracycline (for the R702 plasmid). Plates were incubated at 30 °C aerobically.
The day before conjugation, single colonies of CA434 donor cells carrying pRG-Ea_uxdFOC were inoculated into 5 mL of LB + 30 μg/mL chloramphenicol + 15 μg/mL tetracycline and grown overnight at 30 °C with shaking (aerobically). A single colony of the recipient C. sporogenes was inoculated into 5 mL of BHI and grown overnight at 37 °C (anaerobically). The next day, 1.5 mL of CA434 donor was pelleted at 6500 x g for 3 min, washed with 1 mL of sterile PBS, re-pelleted, placed on ice, and brought to the anaerobic chamber. An aliquot from the C. sporogenes overnight culture (in BHI) was taken out of the anaerobic chamber and heat shocked at 50 °C for 5 min to increase transformation efficiency66. Afterwards, 300 μL of heat shocked C. sporogenes recipient culture was mixed with the pellets from appropriate donors. The resulting mixture was spotted (3 × 20 μL) onto pre-reduced BHI plates without antibiotics and incubated for 24 h at 37 °C (anaerobically). The following day, all 3 spots were scraped with a 10 μL sterile loop and resuspended in 300 μL of PBS. 100 μL of this suspension was spread onto a BHI agar plate supplemented with 250 μg/mL D-cycloserine (against CA434) + 30 μg/mL thiamphenicol (for pRG-Ea_uxdFOC) and grown for 24 h at 37 °C. Individual transformants were picked and re-streaked onto BHI + 250 μg/mL D-cycloserine + 30 μg/mL thiamphenicol. Plasmid-positive C. sporogenes colonies were identified by colony PCR (cPCR) using the OneTaq 2X Quick-Load Master Mix (NEB #M0486) with cat_cPCR primers and Ea_uxdC_cPCR_f/RepOri_cPCR_r and conditions detailed in Supplementary Tables 3, 4. In all cases, PCR cycling was as follows: 5 min at 94 °C, 35 cycles (20 s at 94 °C, 30 s at Ta, reaction-specific extension time at 68 °C), 5 min at 68 °C. Cultures from pRG-Ea_uxdFOC-positive colonies, grown in BHI + 30 μg/mL thiamphenicol, were mixed equally with 50% glycerol and stored at −70 °C.
Heterologous expression of Uxd proteins and dehydroxylase activity assay in C. sporogenes
All growth steps below were performed in selective media (BHI + 30 μg/mL thiamphenicol) in anaerobic conditions at 37 °C. Single colonies of C. sporogenes transformed with pRG-eGFP or pRG-Ea_uxdFOC, grown on BHI + 30 μg/mL thiamphenicol agar plates, were inoculated into 5 mL selective media and grown overnight. Overnight cultures were then diluted 1/100 into 10 mL fresh selective media and grown for 6 h. Theophylline (100 mM in 0.1 M NaOH) was added to a final concentration of 2 mM to induce protein expression. For the initial screen with uroM6, the media (BHI) and cultures (WT and pRG-Ea_uxdFOC) were split into 200 μL aliquots and treated with 100 μM uroM6. Whole cell cultures were incubated for 48 h at 37 °C (anaerobically). For the latter experiment with uroM7, the media (BHI) and cultures (WT, pRG-eGFP, and pRG-Ea_uxdFOC) were split into 200 μL aliquots and treated with either DMSO or 100 μM uroM7. Whole cell cultures were incubated for 96 h at 37 °C (anaerobically) to ensure complete urolithin conversion. To evaluate dehydroxylase activity for both sets of experiments, urolithin metabolites were extracted from whole cultures following Extraction Method A.
SDS-PAGE analysis of Uxd proteins
C. sporogenes pRG-Ea_uxdFOC cultures induced with 2 mM theophylline for 24 h were pelleted at 5000 g for 5 min, and the pellets were resuspended in 1.5 mL of PBS + 2 mg/mL lysozyme. The cell suspensions were sonicated on ice using a Misonix Sonicator 3000 set to power level 2/10 according to the following sequence (aerobically, in a cold room): 20 s ON, 40 s OFF, for a total of 3 min ON. The crude lysate was diluted with 4X reducing loading dye to a final concentration of 1X: 62.5 mM Tris-HCl pH 6.8, 2% w/v SDS, 10% glycerol, 0.01% w/v bromophenol blue, 38 mM DTT. All samples were heated at 95 °C for 5 min, then 10 μL were loaded onto a 10% bis-tris polyacrylamide protein gel. Gels were fixed and stained with 10X colloidal Coomassie solution according to the European Molecular Biology Laboratory Proteomics Core Facility recipe: 0.4% w/v Coomassie Brilliant Blue R250, 10% w/v citric acid, 8% w/v ammonium sulfate, and 20% v/v methanol.
Comparative genomics
The analysis of ucd and uxd operon presence in Enterocloster spp. genomes was done by first downloading all available (n = 452) Enterocloster spp. genome FASTA sequences from the NCBI genomes database (https://www.ncbi.nlm.nih.gov/datasets/genome/?taxon=2719313&typical_only=true Accessed 01-08-2025). Atypical genomes were excluded. All FASTA files with GenBank (GCA) accessions were retained for further analysis (n = 452). The Genome Taxonomy Database (GTDB) table for Enterocloster spp. (release 10-RS226 (16-04-2025)) was downloaded from (https://gtdb.ecogenomic.org/searches?s=al&q=Enterocloster Accessed 02-08-2025). Each genome was queried using rBLAST (v1.2.0, DOI: 10.18129/B9.bioc.rBLAST) for matches to the E. bolteae ucd operon and E. asparagiformis uxd operon nucleotide sequences with criteria: -perc_identity 60 -max_target_seqs 5 -subject_besthit. Matches were further filtered for length >300 bases to exclude small fragments. NCBI GenBank accessions for all queries without GTDB taxonomy or the “Undefined (Failed Quality Check)” flag were removed from further analyses, yielding a total of n = 329 genomes (accessions listed in Supplementary Table 6). We note that E. pacaense is labeled as s__Enterocloster sp000155435 in GTDB release 10-RS226 (current release on 15-09-2025) but was labeled as s__Enterocloster pacaense in release 09-RS220. Presence/absence of each operon was plotted in ggplot2 (v.3.5.2). All code, tables, and resulting RData files from this analysis were deposited in Zenodo (https://doi.org/10.5281/zenodo.16740616).
Comparisons between ucd and uxd operon genomic contexts were done using CAGECAT67 (v1.0, https://cagecat.bioinformatics.nl/) using the clinker tool for visualization. GenBank files of ucd and uxd operon genomic contexts were downloaded from the NCBI Sequence Viewer from genome accessions in Supplementary Table 7. The identity threshold was set to 40% and proteins were manually recolored to highlight conserved features. Protein similarity matrices were generated from clinker alignment.txt output files.
Multiple sequence alignment of UcdO and UxdO proteins from E. asparagiformis, E. bolteae, E. citroniae, and E. pacaense were performed in Benchling using the Clustal Omega algorithm using protein sequences provided in Supplementary Table 8.
Searches for sequences similar to the E. asparagiformis DSM 15981 uxd operon (nucleotide) sequence were conducted using the blastn algorithm against refseq_genomes limited to Bacteria (taxid: 2). Searches for sequences similar to the E. asparagiformis DSM 15981 UxdO and UxdC (protein) sequences (Supplementary Table 8) were conducted using the blastp algorithm against refseq_protein limited to Bacteria (taxid: 2). To generate figures, the NCBI multiple sequence alignment viewer (v1.26.0) pages were sorted by identity and coverage and exported.
Protein domain annotations
Domain annotations for E. asparagiformis Ucd proteins (UcdC, UcdF, and UcdO) and Uxd (UxdF, UxdO, and UxdC) proteins were obtained from InterPro68 via UniProt accessions listed in Supplementary Table 8.
Protein structures and homology modeling
Protein structures were generated using AlphaFold3 (AF3, https://alphafoldserver.com/) with protein sequences from E. asparagiformis (Ea) listed in Supplementary Table 8. For the Ea Ucd complex, a single copy of UcdC, UcdF, and UcdO protein sequences and a single FAD ligand were used as input. For the Ea Uxd complex, a single copy of UxdC, UxdF, and UxdO protein sequences and 2 FAD ligands were used as input. AF3 output model_0.cif (crystallographic information file) files were opened in PyMOL (v2.4.1, Schrodinger LLC). For homology modeling, FoldSeek69, integrated into the AlphaFold Protein Structure Database (https://alphafold.ebi.ac.uk/), was used to search the RCSB Protein Data Bank (RCSB PDB, https://www.rcsb.org/) for proteins that had similar folds to the Ea UcdO and UxdO subunits. The crystal structure of the Thauera aromatica 4-hydroxybenzoyl-CoA reductase (PDB 1RM6)35 was used as a reference to position (using the “super” command in PyMOL to align oxidoreductase subunits) the molybdopterin cytosine dinucleotide (MCD) cofactor and 2Fe-2S clusters in UcdO/UxdO and UcdF/UxdF, respectively.
RNA extraction and RT-qPCR
A volume of 1 mL of treated (DMSO or 100 μM uroM6, uroM7, uroC, or uroA for 2 h) E. asparagiformis culture was pelleted (6,500 g for 3 min), and the supernatant was removed for later LC-MS analysis. The pellet was then mixed with 900 μL TRI reagent (Zymo Research #R2050-1-50) and transferred to a ZR BashingBead lysis tube (Zymo Research #S6012-50). Samples were lysed in a Mini Beadbeater 16 (Biospec #607) according to the following sequence: 1 min ON, 5 min OFF. Bead beating was done for a total of 2 min ON to preserve longer transcripts. RNA isolation was then performed using the Direct-zol RNA Miniprep Kit (Zymo Research # R2051) according to the manufacturer’s instructions (including an on-column DNase digestion). To ensure complete DNA removal, an additional DNA digestion step was performed on the isolated RNA using the Ambion DNA-free DNA Removal Kit (Invitrogen #AM1906) according to the manufacturer’s instructions. RNA concentration and quality were verified by NanoDrop and 1% (w/v) agarose gel electrophoresis.
Isolated RNA samples (500 ng) were reverse transcribed using the iScript Reverse Transcription Supermix (Bio-Rad #1708840) in a reaction volume of 10 μL. The iScript No-RT Control Supermix was used as a no-enzyme control for reverse transcription (-RT). The reaction mixtures were incubated in a thermal cycler: 5 min at 25 °C, 20 min at 48 °C, and 1 min at 95 °C. Both cDNA and -RT controls were diluted 1/20 in nuclease-free water before use. qPCR reactions were conducted using the Luna Universal qPCR Master Mix kit (NEB #M3003). The Ea_ucdO_qPCR, Ea_uxdO_qPCR, and Ea_CTP_Ref primer pairs (Supplementary Table 3) were added to their respective master mixes (final primer concentration of 250 nM) and 7 μL of diluted template (cDNA, -RT, no template) were added to 28 μL of master mix for technical triplicates. Replicate mixes were pipetted (10 μL/well) into a MicroAmp Fast 96-Well Reaction Plate (Applied Biosystems #4346907) and the plates were sealed (PlateSeal #PS-PETSTXL-100), then spun down for 2 min to eliminate air bubbles. The qPCR detection parameters on the Viia7 qPCR machine (Applied Biosystems #4346907) were as follows: SYBR Green detection, ROX reference dye, 10 μL reaction volume. The thermal cycling conditions were: 1 min at 95 °C, 40 cycles (15 s at 95 °C, 30 s at 60 °C), and melt analysis (60–95 °C). Data were analyzed according to the 2−ΔΔCt method with the E. asparagiformis CTP synthase (Ea_CTP) gene serving as the reference gene due to its stable expression upon treatment with uroC24.
E. asparagiformis cell suspension assays to test dehydroxylase inducibility
Overnight cultures of E. asparagiformis (Ea) were diluted 1/50 into fresh mABB+H media (25 mL) and grown for 5 h at 37 °C in anaerobic conditions. Each culture was then split into 4 × 5 mL aliquots, and each aliquot was treated with DMSO or 9-hydroxy urolithins (uroD, uroC, or isouroA, 50 μM final concentration for each compound) for 2 h at 37 °C to induce protein expression. Cultures were then pelleted at 6500 x g for 3 min, and supernatants were collected for downstream extraction and LC-MS analysis using Extraction Method A. The pellets were washed with 5 mL of pre-reduced PBS, pelleted once again, and resuspended in 1 mL of PBS. This cell suspension was split into 4 × 200 μL aliquots, and each aliquot was treated with either DMSO, uroM6, uroM7, or uroC (100 μM final concentration). Samples were briefly vortexed and incubated at room temperature in the anaerobic chamber for 20 h. Metabolites in cultures were extracted following Extraction Method A.
Statistical analyses and graphing
Statistical methods were not used to determine sample sizes, experiments were not randomized, and the investigators were not blinded. Data points were assumed to be normally distributed, though this was not formally tested. Details related to each statistical test performed are supplied in the Figure legends, and statistical test results are provided in the Source data file for each figure panel, where applicable. In all cases, α = 0.05 and tests were two-tailed. Data were plotted in GraphPad Prism (v10.4.0) or ggplot2 (v3.5.2). Figures were assembled in Affinity Designer 2 (v2.6.3.332).
Data availability
Untargeted proteomics datasets generated in this study were deposited in the PRIDE repository under accession PXD066865 and DOI 10.6019/PXD066865. DNA sequences of plasmids generated in this study were deposited in GenBank: pTipQC2-Ea_ucdCFO (PX095335.1), pTipQC2-Ea_uxdFOC (PX095336.1), pRG-eGFP (PX095337.1), and pRG-Ea_uxdFOC (PX095338.1). RNA-sequencing data for uroC-treated E. bolteae and E. asparagiformis are available at NCBI SRA BioProject ID PRJNA996126 under BioSample accession codes SAMN36514640 and SAMN36514641, respectively.24 The T. aromatica 4-HBCR X-ray crystal structure is available in the Protein Data Bank (PDB) under accession 1RM6.35
Code availability
All code, tables, and RData files obtained from the analysis of Enterocloster spp. genomes were deposited in Zenodo (https://doi.org/10.5281/zenodo.16740616).
References
Sanz, Y. et al. The gut microbiome connects nutrition and human health. Nat. Rev. Gastroenterol. Hepatol. 22, 534–555 (2025).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. 39, 105–114 (2021).
Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. & Goodman, A. L. Mapping human microbiome drug metabolism by gut bacteria and their genes. Nature 570, 462–467 (2019).
Lindell, A. E., Zimmermann-Kogadeeva, M. & Patil, K. R. Multimodal interactions of drugs, natural compounds and pollutants with the gut microbiota. Nat. Rev. Microbiol 20, 431–443 (2022).
Javdan, B. et al. Personalized mapping of drug metabolism by the human gut microbiome. Cell 181, 1661–1679.e1622 (2020).
Singh, A. et al. Direct supplementation with Urolithin A overcomes limitations of dietary exposure and gut microbiome variability in healthy adults to achieve consistent levels across the population. Eur. J. Clin. Nutr. 76, 297–308 (2022).
Garcia-Villalba, R. et al. Urolithins: a comprehensive update on their metabolism, bioactivity, and associated gut microbiota. Mol. Nutr. Food Res. 66, e2101019 (2022).
Ryu, D. et al. Urolithin A induces mitophagy and prolongs lifespan in C. elegans and increases muscle function in rodents. Nat. Med. 22, 879–888 (2016).
Liu, S. et al. Effect of urolithin A supplementation on muscle endurance and mitochondrial health in older adults: a randomized clinical trial. JAMA Netw. Open 5, e2144279 (2022).
Faitg, J., D’Amico, D., Rinsch, C. & Singh, A. Mitophagy activation by urolithin A to target muscle aging. Calcif. Tissue Int. 114, 53–59 (2024).
Denk, D. et al. Expansion of T memory stem cells with superior anti-tumor immunity by urolithin A-induced mitophagy. Immunity 55, 2059–2073.e2058 (2022).
Singh, R. et al. Enhancement of the gut barrier integrity by a microbial metabolite through the Nrf2 pathway. Nat. Commun. 10, 89 (2019).
Ghosh, S., Erickson, D., Chua, M. J., Collins, J. & Jala, V. R. The microbial metabolite urolithin A reduces Clostridioides difficile toxin expression and toxin-induced epithelial damage. mSystems 9, e0125523 (2024).
Pinaffi-Langley, A. C.dC. et al. Urolithin A supplementation to improve endothelial and cerebrovascular function in middle-aged adults with obesity: study protocol for a double-blind placebo-controlled randomized study. Clin. Nutr. Open Sci. 61, 217–230 (2025).
Landete, J. M. Ellagitannins, ellagic acid and their derived metabolites: a review about source, metabolism, functions and health. Food Res. Int. 44, 1150–1160 (2011).
Aguilera-Carbo, A., Augur, C., Prado-Barragan, L. A., Favela-Torres, E. & Aguilar, C. N. Microbial production of ellagic acid and biodegradation of ellagitannins. Appl. Microbiol. Biotechnol. 78, 189–199 (2008).
Garcia-Villalba, R. et al. Metabolism of different dietary phenolic compounds by the urolithin-producing human-gut bacteria Gordonibacter urolithinfaciens and Ellagibacter isourolithinifaciens. Food Funct. 11, 7012–7022 (2020).
Iglesias-Aguirre, C. E. et al. In vivo administration of gut bacterial consortia replicates urolithin metabotypes A and B in a non-urolithin-producing rat model. Food Funct. 14, 2657–2667 (2023).
Cortes-Martin, A. et al. The gut microbiota urolithin metabotypes revisited: the human metabolism of ellagic acid is mainly determined by aging. Food Funct. 9, 4100–4106 (2018).
Beltran, D., Romo-Vaquero, M., Espin, J. C., Tomas-Barberan, F. A. & Selma, M. V. Ellagibacter isourolithinifaciens gen. nov., sp. nov., a new member of the family Eggerthellaceae, isolated from human gut. Int J. Syst. Evol. Microbiol 68, 1707–1712 (2018).
Selma, M. V., Beltran, D., Garcia-Villalba, R., Espin, J. C. & Tomas-Barberan, F. A. Description of urolithin production capacity from ellagic acid of two human intestinal Gordonibacter species. Food Funct. 5, 1779–1784 (2014).
Selma, M. V. et al. Isolation of human intestinal bacteria capable of producing the bioactive metabolite isourolithin a from ellagic acid. Front Microbiol. 8, 1521 (2017).
Hitch, T. C. A. et al. HiBC: a publicly available collection of bacterial strains isolated from the human gut. Nat. Commun. 16, 4203 (2025).
Pidgeon, R. et al. Diet-derived urolithin A is produced by a dehydroxylase encoded by human gut Enterocloster species. Nat. Commun. 16, 999 (2025).
Iglesias-Aguirre, C. E. et al. Gut bacteria involved in ellagic acid metabolism to yield human urolithin metabotypes revealed. J. Agric Food Chem. 71, 4029–4035 (2023).
Nakashima, N. & Tamura, T. Isolation and characterization of a rolling-circle-type plasmid from Rhodococcus erythropolis and application of the plasmid to multiple-recombinant-protein expression. Appl Environ. Microbiol. 70, 5557–5568 (2004).
Neumann, M. & Leimkuhler, S. The role of system-specific molecular chaperones in the maturation of molybdoenzymes in bacteria. Biochem Res Int. 2011, 850924 (2011).
Liu, Y. et al. A widely distributed gene cluster compensates for uricase loss in hominids. Cell 186, 3400–3413.e3420 (2023).
Liu, Y. et al. Gut bacteria degrade purines via the 2,8-dioxopurine pathway. Nat. Microbiol 10, 2291–2305 (2025).
Neumann, M., Schulte, M., Junemann, N., Stocklein, W. & Leimkuhler, S. Rhodobacter capsulatus XdhC is involved in molybdenum cofactor binding and insertion into xanthine dehydrogenase. J. Biol. Chem. 281, 15701–15708 (2006).
Neumann, M., Mittelstadt, G., Seduk, F., Iobbi-Nivol, C. & Leimkuhler, S. MocA is a specific cytidylyltransferase involved in molybdopterin cytosine dinucleotide biosynthesis in Escherichia coli. J. Biol. Chem. 284, 21891–21898 (2009).
Canadas, I. C., Groothuis, D., Zygouropoulou, M., Rodrigues, R. & Minton, N. P. RiboCas: a universal CRISPR-based editing tool for Clostridium. ACS Synth. Biol. 8, 1379–1390 (2019).
Hille, R., Hall, J. & Basu, P. The mononuclear molybdenum enzymes. Chem. Rev. 114, 3963–4038 (2014).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Unciuleac, M., Warkentin, E., Page, C. C., Boll, M. & Ermler, U. Structure of a xanthine oxidase-related 4-hydroxybenzoyl-CoA reductase with an additional [4Fe-4S] cluster and an inverted electron flow. Structure 12, 2249–2256 (2004).
Spanogiannopoulos, P., Bess, E. N., Carmody, R. N. & Turnbaugh, P. J. The microbial pharmacists within us: a metagenomic view of xenobiotic metabolism. Nat. Rev. Microbiol. 14, 273–287 (2016).
Zheng, B., He, Y., Zhang, P., Huo, Y. X. & Yin, Y. Polyphenol utilization proteins in the human gut microbiome. Appl Environ. Microbiol. 88, e0185121 (2022).
McGivern, B. B. et al. Microbial polyphenol metabolism is part of the thawing permafrost carbon cycle. Nat. Microbiol 9, 1454–1466 (2024).
Rodriguez-Daza, M. C. et al. Polyphenol-mediated gut microbiota modulation: toward prebiotics and further. Front Nutr. 8, 689456 (2021).
McGivern, B. B., Woyda, R., Flynn, R. M. & Wrighton, K. C. CAMPER: curated annotations for profiling microbial polyphenol metabolic potential. bioRxiv https://doi.org/10.1101/2023.09.24.559193 (2023).
D’Amico, D. et al. Impact of the natural compound urolithin a on health, disease, and aging. Trends Mol. Med. 27, 687–699 (2021).
Kok, C. R., Rose, D. & Hutkins, R. Predicting personalized responses to dietary fiber interventions: opportunities for modulation of the gut microbiome to improve health. Annu Rev. Food Sci. Technol. 14, 157–182 (2023).
Hu, J., Mesnage, R., Tuohy, K., Heiss, C. & Rodriguez-Mateos, A. Poly)phenol-related gut metabotypes and human health: an update. Food Funct. 15, 2814–2835 (2024).
McKinlay, J. B. Are bacteria leaky? Mechanisms of metabolite externalization in bacterial cross-feeding. Annu Rev. Microbiol. 77, 277–297 (2023).
Watanabe, H., Kishino, S., Kudoh, M., Yamamoto, H. & Ogawa, J. Evaluation of electron-transferring cofactor mediating enzyme systems involved in urolithin dehydroxylation in Gordonibacter urolithinfaciens DSM 27213. J. Biosci. Bioeng. 129, 552–557 (2020).
Bae, M. et al. Distinct classes of gut bacterial molybdenum-dependent enzymes produce urolithins. bioRxiv https://doi.org/10.1101/2025.01.17.632430 (2025).
Culp, E. J. & Goodman, A. L. Cross-feeding in the gut microbiome: ecology and mechanisms. Cell Host Microbe 31, 485–499 (2023).
Dong, X., Bae, M., Le, C., Aguilar Ramos, M. A. & Balskus, E. P. Enantiocomplementary gut bacterial enzymes metabolize dietary polyphenols. J. Am. Chem. Soc. 147, 7231–7244 (2025).
Maini Rekdal, V. et al. A widely distributed metalloenzyme class enables gut microbial metabolism of host- and diet-derived catechols. Elife 9, https://doi.org/10.7554/eLife.50845 (2020).
Bae, M. et al. Metatranscriptomics-guided discovery and characterization of a polyphenol-metabolizing gut microbial enzyme. Cell Host Microbe 32, 1887–1896.e1888 (2024).
Boll, M. et al. Redox centers of 4-hydroxybenzoyl-CoA reductase, a member of the xanthine oxidase family of molybdenum-containing enzymes. J. Biol. Chem. 276, 47853–47862 (2001).
Goel, A. et al. Toward a health-associated core keystone index for the human gut microbiome. Cell Rep. 44, 115378 (2025).
Xia, M. et al. Improvement of urolithin a yield by in vitro cofermentation of Streptococcus thermophilus FUA329 with human gut microbiota from different urolithin metabotypes. J. Agric Food Chem. 72, 3008–3016 (2024).
Mi, H. et al. Lactococcus garvieae FUA009, a novel intestinal bacterium capable of producing the bioactive metabolite urolithin a from ellagic acid. Foods 11, 2621 (2022).
Zhang, X. et al. Isolation and characterization of a novel human intestinal Enterococcus faecium FUA027 capable of producing urolithin A from ellagic acid. Front Nutr. 9, 1039697 (2022).
Wang, Z. et al. Function identification of a lactonase gene lac1563 involved in producing urolithin a of Limosilactobacillus fermentum FUA033. J. Agric Food Chem. 73, 6795–6806 (2025).
Gaya, P., Peirotén, A., Medina, M., Alvarez, I. & Landete, J. M. Bifidobacterium pseudocatenulatum INIA P815: the first bacterium able to produce urolithins A and B from ellagic acid. J. Funct. Foods 45, 95–99 (2018).
Maurice, C. F., Haiser, H. J. & Turnbaugh, P. J. Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 152, 39–50 (2013).
Haiser, H. J. et al. Predicting and manipulating cardiac drug inactivation by the human gut bacterium Eggerthella lenta. Science 341, 295–298 (2013).
Fraga, C. G., Croft, K. D., Kennedy, D. O. & Tomas-Barberan, F. A. The effects of polyphenols and other bioactives on human health. Food Funct. 10, 514–528 (2019).
Dong, Z. et al. Discovery of an ene-reductase initiating resveratrol catabolism in gut microbiota and its application in disease treatment. Cell Rep. 44, 115517 (2025).
Bess, E. N. et al. Genetic basis for the cooperative bioactivation of plant lignans by Eggerthella lenta and other human gut bacteria. Nat. Microbiol 5, 56–66 (2020).
Keller, A., Nesvizhskii, A. I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Nesvizhskii, A. I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003).
Purdy, D. et al. Conjugative transfer of clostridial shuttle vectors from Escherichia coli to Clostridium difficile through circumvention of the restriction barrier. Mol. Microbiol 46, 439–452 (2002).
Kirk, J. A. & Fagan, R. P. Heat shock increases conjugation efficiency in Clostridium difficile. Anaerobe 42, 1–5 (2016).
van den Belt, M. et al. CAGECAT: the CompArative GEne Cluster Analysis Toolbox for rapid search and visualisation of homologous gene clusters. BMC Bioinforma. 24, 181 (2023).
Blum, M. et al. InterPro: the protein sequence classification resource in 2025. Nucleic Acids Res 53, D444–D456 (2025).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Acknowledgements
This research was funded by the Canadian Institutes of Health Research (CIHR) grant PJT-437944, a Weston Family Microbiome Initiative’s 2021 Transformational Research program, and a Weston Family Foundation 2024 Proof-of-Principle grant to B.C. B.C. held a tier II Canada Research Chair (CRC) in Therapeutic Chemistry (2015-2025). R.P. is supported by the CIHR Canada Graduate Scholarship-Doctoral (#493808 for 2023-2026) and by the Fonds de Recherche du Québec-Santé : Bourse de formation au doctorat (#316063 for 2022-2023). A.G. is supported by the CIHR Canada Graduate Scholarship-Masters for 2024-2025. Untargeted proteomics sample preparation and analysis were performed by the Research Institute of the McGill University Health Centre (RI-MUHC) Proteomics and Molecular Analysis Platform. Molecular biology experiments were enabled by the McGill University Imaging and Molecular Biology Platform (IMBP).
Author information
Authors and Affiliations
Contributions
R.P. designed the study, performed experiments, analyzed data, created figures, and wrote the initial manuscript. A.G. performed experiments, analyzed data, and created figures. L.D. consulted on experimental design and methodology. B.C. designed the study, supervised the research, obtained research funding, and wrote the initial manuscript with R.P. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pidgeon, R., Giurleo, A., Dridi, L. et al. The presence and induction of regioselective dehydroxylases dictate urolithin metabolism by Enterocloster species. npj Biofilms Microbiomes 11, 220 (2025). https://doi.org/10.1038/s41522-025-00849-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41522-025-00849-w