Abstract
Parkinson’s disease (PD), as a neurodegenerative disorder, is characterized primarily by damage to the central nervous system, accompanied by astrocyte dysfunction and the activation of ferroptosis. Recent studies have shown that oligodendrocytes also exhibit functional abnormalities in the brains of PD patients and are involved in the ferroptotic process. However, it remains unclear whether there is an interaction between oligodendrocytes and astrocytes and how they induce neuronal ferroptosis. Here, we employed single-nucleus sequencing and spatial transcriptomics to characterize the intercellular communication network between oligodendrocytes and astrocytes in the PD environment. Among these, astrocytes are the primary recipients of signals sent by oligodendrocytes in the FGF (Fibroblast growth factors) signaling pathway. In PD, the communication intensity is weakened, involving FGF1 and FGF9 and their receptors FGFR1, FGFR2, and FGFR3. Subsequently, we further validated the significant activation of mitochondrial oxidative phosphorylation processes within oligodendrocytes and astrocytes in PD mice, and that astrocytes might also involve the interaction of Mt1 and Ca2+. Additionally, we demonstrated a significant reduction in the number of DA neurons in the SN region and a notable activation of ferroptosis, alongside a significant decrease in the antioxidant pathway NRF2/SLC7A11/GPX4. In summary, our data elucidate that ferroptosis in the midbrain SN region preferentially occurs in astrocytes under the dysregulation of oligodendrocytes, leading to ferroptosis in DA neurons. Thus, our study highlights the crucial role of oligodendrocyte-astrocyte crosstalk in driving neuronal inactivation and inflammatory expansion in PD.
Similar content being viewed by others
Introduction
Parkinson’s disease (PD) is the second most common neurodegenerative disorder worldwide. The pathological hallmarks include the death of dopaminergic (DA) neurons in the substantia nigra pars compacta (SNpc) and the misfolded aggregation of α-synuclein (α-Syn) into Lewy bodies (LB), leading to symptoms such as motor dysfunction, cognitive impairment, and emotional disturbances in patients1,2. Genetic, age-related, and environmental factors all contribute to the pathogenesis of PD3. However, due to the largely unknown mechanisms underlying the pathology and etiology of PD, current treatments exhibit short-term efficacy and significant side effects4. Therefore, it is crucial to further elucidate the fundamental causes of PD’s pathological changes and progression.
When considering the key factors in the pathological changes of PD, we became interested in the cascade reactions activated by ion dysregulation induced by the persistent inflammatory response in PD, rather than the broader direct result of glial cell activation and neuronal loss5. Ferroptosis has been identified as a key cause of neuronal loss in PD, which is associated with astrocyte dysfunction6,7. Astrocytes, as components of the blood-brain barrier, can mediate iron influx into the brain and regulate the transport of various types of iron, including transferrin-bound iron (Tf-Fe), non-transferrin-bound iron (NTBI), heme iron, and ceruloplasmin, among others. However, in PD, the dysfunction of astrocytes and iron transporters can lead to selective iron deposition in the SNpc. Subsequently, the inherent redox-active chemical properties of iron lay the foundation for its potential toxicity. The accumulated iron drives the expansion of mitochondrial reactive oxygen species (ROS) and lipid peroxidation through Fenton reactions, ultimately resulting in damage to DNA, lipids, and proteins8,9. To date, mutations in several genes have been characterized in PD and are related to ferroptosis, including ACSL4 (Acyl-CoA Synthetase Long-Chain Family Member 4), DJ-1 (Protein DJ-1), NRF2 (Nuclear Factor Erythroid 2-Related Factor 2), SLC7A11 (Solute Carrier Family 7 Member 11), and GPX4 (Glutathione Peroxidase 4)10.
Recent studies have shown that white matter and oligodendrocyte damage are common phenomena in many neurodegenerative diseases, and oligodendrocyte dysfunction may occur early in the disease progression11. In the brain, oligodendrocytes contain the majority of iron, as myelination and metabolic enzyme activity require iron12,13. However, the process of myelination in oligodendrocytes also requires significant consumption of oxygen and ATP, leading to the formation of ROS, and the low GSH content within oligodendrocytes further exacerbates this effect14. Therefore, oligodendrocytes may be involved in the ferroptosis of DA neurons in PD. Quantitative analysis has shown a significant increase in iron accumulation in DA neurons and glial cells in the brains of post-mortem PD patients. Specifically, the iron content in Olig2-positive oligodendrocytes increased by 2.5 times12. Moreover, current studies have found oligodendrocyte demyelination in PD15, Alzheimer’s disease (AD)11, and multiple sclerosis (MS)13. This observation suggests that demyelinated oligodendrocytes may release large amounts of iron into the neuronal milieu, thereby exacerbating neuronal ferroptosis.
Currently, glial pathology is suspected to drive the neuroinflammatory processes throughout the brain16. The relationship between oligodendrocytes and astrocytes is particularly intriguing, and it is worth investigating how they act as “partners in crime” to contribute to neuronal ferroptosis in PD. Therefore, we used single-nucleus RNA sequencing (snRNA-seq) to obtain the cellular composition of brain tissue in MPTP (1-Methyl-4-phenyl-1,2,3,6-tetrahydropyridine) model mice and employed spatial transcriptomics to obtain a global view of PD-related changes. The aim is to uncover the specific pathways through which oligodendrocytes and astrocytes induce neuronal inactivation in PD.
Results
PD Model
Here, we first conducted PD-related assessments on the MPTP model mice used in this study. Following MPTP toxin injection, the PD mice exhibited a significant decrease in body weight (Supplementary Fig. 1A). In terms of behavioral performance, compared to the Ctrl group, the PD group showed a marked reduction in the time spent on the rotarod (Supplementary Fig. 1B). In the open field test, the PD mice exhibited a significant decrease in both the total distance and average speed, along with a notable increase in rest time (Supplementary Fig. 1C). Additionally, immunohistochemical analysis of tyrosine hydroxylase (TH) in the substantia nigra (SN) region of the brain showed a significant reduction in the expression of TH, a specific marker of DA neurons, in the SN of PD mice (Supplementary Fig. 1D). Therefore, based on the behavioral assessments and TH staining results from both groups of mice, the MPTP-induced PD model mice in this study exhibit pathophysiological features consistent with PD.
Single-nucleus analysis based on PD
To comprehensively characterize the cellular composition in PD, we further filtered out low-quality data, including multicellular, doublet, or unbound cells, based on initial quality control from Cell Ranger. Ultimately, we quantified 6255 cells in the Ctrl group and 8202 cells in the PD group. UMAP visualization results identified 19 initial clusters in both the Ctrl and PD groups (Fig. 1A, B). To identify the major cell types, we manually annotated 12 cell types based on specific marker genes within each cluster and the similarity matrix of the cell clusters (Fig. 1C, D). Three clusters were annotated as astrocytes due to their high expression levels of astrocyte-specific genes (Gli3, Tnc, ATP13A4). One cluster was annotated as microglia (Ikzf1, Siglech, Ly86). Two clusters expressed oligodendrocyte-specific marker genes characteristic of different stages; thus, they were annotated as oligodendrocyte precursor cells (Neu4, Cspg4, Rlbp1, Susd5) and oligodendrocytes (Plp1 and PLXNB3), respectively (Fig. 1E, F). The remaining clusters were annotated as various types of neurons, epithelial cells, smooth muscle cells, and endothelial cells (Fig. 1E, F). Here, we mainly focused on oligodendrocytes and astrocytes. Oligodendrocytes show a notable increasing trend in PD, and there is also an increase in the number of astrocytes (Fig. 1D). However, given the close physical interactions between these two types of glial cells, snRNA-seq data at the transcriptomic level alone may not be sufficient to provide insights into the changes in cell-to-cell crosstalk and spatial characteristics.
A Schematic diagram of the study design. Twenty mice were randomly divided into two groups, with ten mice per group. The PD group received MPTP toxin injections, followed by behavioral assessments and TH immunohistochemical tests. Subsequently, single-nucleus RNA sequencing and spatial transcriptomics were performed. Results were validated using multiplex IHC imaging, Elisa, qRT-PCR, and Western Blot experiments. B UMAP plot of the single-nucleus RNA sequencing, showing the distribution of 19 original clusters in the brain tissue. The results after annotating the different clusters are encoded with numbers. C Heatmap of similarity among cell groups, where the numbers represent Pearson correlation coefficients, indicating the degree of correlation between cell groups. D Characteristics of the quantity changes in 12 cell types between the Ctrl and PD groups. E Feature plot of classic marker genes for the 12 cell types. F Classic marker gene expression defines the 12 cell types.
Spatial transcriptomics shows spatial localization characteristics of PD glial cell marker genes
snRNA-seq provides a transcriptomic map of highly heterogeneous cell populations but lacks information on the spatial interactions between cells. Therefore, to better understand the spatial characteristics and potential communication roles of PD neuroglial cells, we utilized 10× Visium CytAssist technology to obtain the in situ gene expression profiles from sample groups, Ctrl and PD, capturing 6647 spots (3697 in the Ctrl group and 2950 in the PD group) and identifying 15 clusters through quality control (Fig. 2A, B). Based on this, we conducted a detailed investigation of the spatial characteristic genes identified in the clusters and spatially visualized the neuroglial cell-specific marker genes in UMAP. Among these, Gfap, Tnc, and Gldc are classic markers for astrocytes, all found in cluster 4, primarily located in the hippocampus and striatum, with increased expression levels in PD, consistent with the snRNA-seq findings of increased astrocytes in PD (Figs. 1D and 2C). Interestingly, Tnc and Gldc are predominantly distributed at the periphery of the brain in the in situ gene expression profiles, while Gfap shows a significant distribution in the midbrain, especially in the SN. As a classic marker of reactive astrocytes, this suggests that the pro-inflammatory response in the SN might be more contributed by Gfap-positive astrocytes (Fig. 2C). The oligodendrocyte-specific marker genes Enpp6 and Lpar1 are present in cluster 9, mainly located around the hippocampus and SN, with decreased expression levels in PD. Particularly in the SN, this is contrary to the snRNA-seq findings of increased oligodendrocytes in PD, and we did not detect specific marker genes for microglia in the spatial transcriptomics, which may be attributed to the limitations of spatial transcriptomics sequencing (Fig. 2C). Additionally, we identified the distribution of TH in the in situ gene expression profiles, located in cluster 8, primarily in the SN, with decreased expression levels in PD, consistent with the pathological changes in DA neurons in PD and previous TH immunohistochemistry results (Fig. 2C). Overall, we used spatial transcriptomics to map the in situ gene expression profiles of the hemispheres of PD mice. Our findings suggest that both oligodendrocytes and astrocytes might be involved in the PD pathological process, but spatial characteristic genes alone do not directly represent the cell type. Thus, the spatial distribution characteristics of PD neuroglial cells cannot rely solely on spatial transcriptomics.
A The UMAP plot shows the spots across the entire hemisphere region, with a total of 15 clusters co-localized and color-coded by different regions. B For the Spot groups obtained from clustering, their proportions within the samples were calculated and displayed in a bar chart. C The spatial visualization of spatial feature genes in the UMAP clustering results. The diagram shows the spatial distribution characteristics of in situ gene expression profiles for TH (dopaminergic neurons), Enpp6 and Lpar1 (oligodendrocytes), and Gfap, Tnc, and Gldc (astrocytes).
Spatial transcriptomics deconvolution analysis reveals spatial distribution characteristics of glial cells in PD
Using the cell type deconvolution method RCTD, we mapped the cell types obtained from this snRNA-seq onto the spatial transcriptome. Here, we first calculated the weight of each cell type generated by snRNA-seq in each spot (Fig. 3A). Based on this, we generated the relative binding abundance percentage of the 12 cell types measured by snRNA-seq within the 15 clusters of the spatial transcriptome. Among them, astrocytes exhibited relatively high binding abundance in cluster 4 of the spatial transcriptome, oligodendrocytes showed relatively high binding abundance in cluster 9 of the spatial transcriptome, while microglia and oligodendrocyte precursor cells did not exhibit significant specificity in any clusters of the spatial transcriptome (Fig. 3B). In summary, these results suggest that the percentages of astrocytes and oligodendrocytes in the spatial transcriptome are largely consistent with those in snRNA-seq, with some minor differences possibly attributable to the different regions covered by the sequencing.
A The composition of cell types at each spot in the spatial slice after deconvolution, with different colors representing different cell types. B The distribution of 12 cell types, identified by snRNA-seq, across the clusters in the spatial transcriptomics clustering after deconvolution. The horizontal axis represents clusters, the vertical axis represents the average proportion of specific cell types within each cluster, and different colored lines represent different cell types. C The distribution of astrocytes, oligodendrocytes, oligodendrocyte precursor cells, and microglia in the spatial slice after deconvolution. The diagram shows a notable colocalization of glial cells in the SN region. D The chord diagram illustrates the co-occurrence of specific pairs of cell types at all captured spots in the sample spatial slice. Thicker lines indicate a higher number of spots containing a particular pair of cell types. The diagram reveals significant colocalization among astrocytes, oligodendrocytes, and microglia within glial cells, predominantly between astrocytes and oligodendrocytes.
Focusing on astrocytes and oligodendrocytes, our results display that the mapping results of astrocytes in each spot are consistent with the variations in the spatial feature gene Gfap. Interestingly, the distribution of astrocytes in the SN is reduced in PD (Fig. 3C, the red rectangle indicates the SN area). On the other hand, oligodendrocytes exhibit a notable regional distribution in the midbrain, particularly in the SN, and their distribution in the SN increases under PD conditions (Fig. 3C). This potentially suggests that oligodendrocytes also participate in the pathological process of PD, even though the number of oligodendrocytes in the SN has not shown a decreasing trend at the current stage. This is consistent with recent studies evaluating their role15. Moreover, there is a good spatial consistency between astrocytes and oligodendrocytes in the SN, indicating that certain interactions between these cells may prominently contribute to the pathological changes in PD, collectively promoting neuronal inactivation, inflammation, and immune transformation in the SN (Fig. 3D).
Communication between astrocytes and oligodendrocytes in PD
Given the potential complexity of intercellular communication changes between astrocytes and oligodendrocytes in PD brain tissue, we employed the CellChat tool to quantitatively infer cell-cell communication based on snRNA-seq data. Our aim was to identify the patterns of communication and crosstalk between astrocytes and oligodendrocytes in a PD context. First, we characterized that both the number and intensity of communication among 12 cell types decreased in the PD environment (Fig. 4A, B). Additionally, our results emphasize that the primary communication in the PD environment occurs between oligodendrocytes and astrocytes, which aligns with our previous spatial transcriptome observations showing prominent distributions of astrocytes and oligodendrocytes in the SN in PD (Fig. 3C).
A Interaction strength and number of interactions among the 12 annotated cell types in the PD group. Red lines indicate increased interaction times or strength, while blue lines indicate decreased interaction times or strength. B In the FGF signaling pathway, the strength of incoming and outgoing interactions among the 12 annotated cell types in Ctrl and PD groups. C The interactions of ligand-receptor pairs FGF1-FGFR1, FGF1-FGFR2, FGF1-FGFR3, and FGF9-FGFR2 in the FGF signaling pathway in Ctrl and PD groups. D Violin plots illustrating the expression levels of ligands FGF1 and FGF9, and receptors FGFR1, FGFR2, and FGFR3 in various cell types within the FGF signaling pathway. E Spatial distribution maps of ligands FGF1 and FGF9, and receptors FGFR1, FGFR2, and FGFR3 in Ctrl and PD group slices, as shown by spatial transcriptomics. F Spatial colocalization of interactions captured by multiplex IHC for FGF1-FGFR1, FGF1-FGFR2, FGF1-FGFR3, and FGF9-FGFR2 in Ctrl and PD groups (The quantified area is within the white rectangle). Red indicates ligands, green indicates receptors, and blue indicates DAPI. This experiment was repeated once with similar results. Scale bar represents 400 μm.
Next, we calculated the signaling pathways involved in cell-cell communication in brain tissue, focusing on the communication changes between oligodendrocytes and astrocytes in the PD environment (Fig. 4A and Supplementary Fig. 2A, B, C). Notably, the FGF (Fibroblast growth factors) signaling pathway was identified as the primary signaling route between oligodendrocytes and astrocytes, with an overall reduction in weight in the PD environment (Fig. 4B). In the FGF signaling pathway, compared to the Ctrl group, oligodendrocytes in the PD environment increased their communication with astrocytes, with strong communication primarily occurring between ligands, including FGF1 and FGF9, and receptors, including FGFR1, FGFR2, and FGFR3 (Fig. 4C). These ligands and receptors mediate active signal transduction between astrocytes and oligodendrocytes. Further sequencing results showed that in the PD environment, oligodendrocytes significantly increased the expression of FGF1 among the ligands in the FGF signaling pathway. The expression levels of receptors FGFR1, FGFR2, and FGFR3 in astrocytes were reduced (Fig. 4D), indicating that the increased communication between oligodendrocytes and astrocytes may be facilitated by the additional production of FGF1 by oligodendrocytes.
We compared the contributions of FGF1 and its receptors to the FGF signaling pathway. Interestingly, the contribution of FGF1-FGFR2 to the FGF signaling pathway remained largely unchanged in the PD environment, while the contributions of FGF1-FGFR1 and FGF1-FGFR3 were reduced (Fig. 4C). Mechanistically, the inability of FGF1 to bind to its receptors could weaken its neuroprotective abilities and its role in promoting oligodendrocyte development and myelin regeneration, particularly its inhibition of the p53 ferroptosis pathway upstream of neuronal mitochondrial events17. On the other hand, the significant increase in FGF1 could induce the production of the chemokine CXCL8 in astrocytes, representing another pro-inflammatory pattern based on oligodendrocytes and astrocytes. It is noteworthy that while examining the spatial transcriptomic in situ gene expression profiles of FGF1 and its receptors, we found a decreasing trend in the distribution of FGF1-FGFR2 in PD. This discrepancy might be attributed to interference from cell types outside the SN.
Importantly, we demonstrated a significant reduction in the co-localization of FGF1-FGFR1, FGF1-FGFR2, and FGF1-FGFR3 in the SN region of brain tissue in the PD environment (Fig. 4E, F, the white rectangle indicates the SN area, detailed analysis data can be found in Supplementary Fig. 3). It should be noted that our spatial transcriptomic mapping results also showed the distribution of FGF9 in the SN. Given that among the receptors, FGFR1, FGFR2, and FGFR3, only FGFR2 is expressed in both oligodendrocytes and astrocytes, we also performed immunofluorescence labeling of FGF9-FGFR2 in brain tissue. The results showed a significant reduction in the co-localization of FGF9-FGFR2 in the PD SN, which may be closely related to inflammation and neuronal oxidative damage in the brain18.
Differential genes in PD oligodendrocytes and astrocytes
To investigate the potential pro-inflammatory interaction between oligodendrocytes and astrocytes in PD based on the FGF signaling pathway, we further characterized the differentially expressed genes in oligodendrocytes and astrocytes to assess their specific functional roles in PD (Figs. 5A and 6A). First, in oligodendrocytes, the results of GO functional enrichment analysis showed that Atpif1, Cend1, Prdx5, Srebf2, Atp5g1 and Stard13 were enriched in mitochondrial functions (Fig. 5B). Additionally, Slc25a4, Ndufb10, Atp5g1, Cox7a2, Ndufa4, and Ndufs6 were enriched in mitochondrial inner membrane functions (Fig. 5B), indicating that oligodendrocytes in PD may be in a state of abnormal mitochondrial activation. Furthermore, beyond its role in the import of ADP and export of ATP in mitochondria, Slc25a4 is also involved in the influx of Ca2+ into mitochondria19, prompting us to re-evaluate the role of Ca2+ in oligodendrocytes. Specifically, as a crucial contributor to mitochondrial energy supply, Ca2+ is involved in maintaining mitochondrial membrane potential, the activity of respiratory chain complexes, and communication between mitochondria and the nucleus. Therefore, the abnormal mitochondrial state in PD may also be influenced by Ca2+ 20. Notably, KEGG functional enrichment revealed that ferroptosis, PD, oxidative phosphorylation, the tricarboxylic acid cycle, and DA synapses were all enriched in oligodendrocytes (Fig. 5C). Specifically, Fth1, Vdac3, Vdac2, Acsl1, and Acsl3 were enriched in ferroptosis, suggesting the activation of ferroptosis in oligodendrocytes in PD. Furthermore, Gnao1, Mapk10, Ppp2r2a, Gng12, and Ppp2r1b, which are involved in neuronal growth and development in the brain, were found to be reduced in oligodendrocytes and enriched in DA synapses (Fig. 5C), indicating potential functional abnormalities in oligodendrocytes.
A The volcano plot shows the Top 20 significantly upregulated/downregulated genes in astrocytes in the PD group compared to the Ctrl group, with red representing upregulated genes and blue representing downregulated genes. B Results of the GO enrichment analysis. The results indicate that multiple mitochondrial functions and ATP generation-related processes are significantly enriched in astrocytes. C Results of the KEGG enrichment analysis. The results show that PD-related genes are significantly enriched in astrocytes, and oxidative phosphorylation is functionally enriched in astrocytes.
A The volcano plot shows the Top 20 significantly upregulated/downregulated genes in oligodendrocytes in the PD group compared to the Ctrl group, with red representing upregulated genes and blue representing downregulated genes. B Results of the GO enrichment analysis. The results indicate that oligodendrocytes in PD also exhibit mitochondrial function enrichment similar to that observed in astrocytes. C Results of the KEGG enrichment analysis. In addition to the oxidative phosphorylation functional enrichment observed in astrocytes, oligodendrocytes also show functional enrichment of ferroptosis-related genes, indicating the presence of ferroptosis processes in oligodendrocytes in PD.
In astrocytes, we identified several genes associated with the mitochondrial respiratory chain, including Cox8a, Cox4i1, Ndufa4, and Cox6c, which were upregulated (Fig. 6B). Consequently, gene sets related to the mitochondrial inner membrane and ATP production were enriched in astrocytes (Fig. 6B). Additionally, Ndufa4, Ndufa13, Ndufb9, Ndufb8, Ndufc2, Ndufb6, and Ndufs7, which are subunits of mitochondrial complex I (NADH dehydrogenase)21, were elevated in astrocytes, indicating their enrichment in respiratory functions (Fig. 6B). These results suggest that astrocytes in PD may be in a state of high metabolic activity, but this also implies a potential increase in ROS production. Moreover, Mt1 and Mt2, which are receptors for melatonin primarily distributed in the central nervous system, are involved in synaptic remodeling, immune regulation, and intracellular Ca2+ signaling in the SN22. Notably, Mt1 participates in the suppression of the NRF2/GPX4-mediated ferroptosis pathway in the brain23, but these genes were significantly downregulated in astrocytes. Therefore, two hypotheses can be proposed: (1) Astrocytes in PD may induce the activation of the NRF2/GPX4-mediated ferroptosis process through this pathway, even though the number of astrocytes does not decrease. (2) There may be a significant increase in Ca2+ content in astrocytes, promoting mitochondrial oxidative phosphorylation and ROS release. KEGG functional enrichment analysis revealed the activation of various pathways in astrocytes. Among these, Ubb, Uchl1, and Slc25a4 were mainly functionally enriched in PD, indicating astrocytes’ involvement in the PD pathological process (Fig. 6C). The high levels of Atp6v0c, which are enriched in mitochondrial oxidative phosphorylation functions, further suggest a high metabolic state and high ROS release in astrocytes in PD.
In summary, based on the results of differentially expressed genes, oligodendrocytes and astrocytes exhibit a similar enrichment of mitochondrial-related gene sets, indicating a similar state of mitochondrial activity in these cells, thereby accelerating ROS production. Additionally, the significant enrichment of ferroptosis-related genes in oligodendrocytes suggests the activation of the ferroptosis process in oligodendrocytes in PD, which may be a critical factor leading to the inactivation of DA neurons in PD. Specifically, ferroptosis occurring in oligodendrocytes may be transmitted to astrocytes, further resulting in ferroptosis in DA neurons.
Oligodendrocytes and astrocytes may induce neuronal ferroptosis through the NRF2/SLC7A11/GPX4 pathway
The aforementioned results indicate that there are significant changes in oligodendrocytes and astrocytes under PD conditions based on the FGF signaling pathway. This involves a notable decrease in the colocalization of FGF1 and FGF9 and their receptors FGFR1, FGFR2, and FGFR3. Additionally, the significantly differentially expressed genes in oligodendrocytes and astrocytes suggest high mitochondrial respiratory activity within these cells. The significant activation of oxidative phosphorylation processes in astrocytes may be influenced by Mt1-mobilized Ca2+. Thus, we first demonstrated that the protein expression levels of mitochondrial oxidative phosphorylation-related Slc25a4, Ndufa4, and Atp6v0c in oligodendrocytes and astrocytes significantly increase under PD conditions (Fig. 7A).
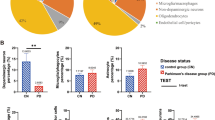
A The results of the Western Blot show that the protein expression levels of Slc25a4, Ndufa4, and Atp6v0c, which are related to mitochondrial oxidative phosphorylation, are significantly increased in astrocytes and oligodendrocytes under PD conditions. Additionally, in PD astrocytes, there is a significant decrease in the expression of the Mt1 protein. B The results of the ELISA kit show that the Ca2+ content in astrocytes of PD mice is significantly increased. At the same time, the iron ion content in the SN of mice is significantly elevated, while the GSH content, total SOD activity, and MDA content are significantly decreased, indicating the presence of high levels of ROS and the activation of ferroptosis processes in the SN of PD mice.
Next, we examined the protein expression levels of Mt1 and the Ca2+ content in astrocytes. As we anticipated, the protein expression level of Mt1 in astrocytes was significantly reduced in PD, and the Ca2+ content significantly increased (Fig. 7A, B). This reinforces our previous prediction that the decreased mRNA expression level of Mt1 in astrocytes facilitates Ca2+ entry into mitochondria, accelerating mitochondrial oxidative phosphorylation. It is noteworthy that although the significant enrichment of ferroptosis-related genes in oligodendrocytes and the changes in Mt1 and Ca2+ content in astrocytes suggest the activation of the ferroptosis process in both cell types, our prior snRNA-seq data indicated a significant increase in the number of oligodendrocytes and astrocytes under PD conditions. We propose that this may be due to the protective effects of FGF1 or other processes. However, as the primary culprits of oxidative damage, the activation of ferroptosis in oligodendrocytes and astrocytes may first target the more vulnerable neurons in the SN DA neurons. Therefore, we investigated the occurrence of ferroptosis in the SN. Our results showed that the iron content in the SN of PD mice was significantly elevated (Fig. 7B). At the same time, there was a marked decrease in GSH levels, total SOD activity, and MDA (Fig. 7B). Additionally, there were observations of mitochondrial cristae shrinkage and darkening, reduced cristae, nuclear membrane shrinkage, and unclear nucleoli, indicating high levels of ROS and activation of the ferroptosis process within the SN of PD mice (Supplementary Fig. 4, The green arrows indicate normal mitochondria in the Ctrl group, while the red arrows denote abnormal mitochondria in the PD condition).
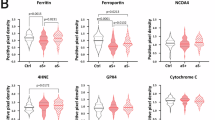
To further explore the specific pathways leading to the activation of ferroptosis in the SN of PD mice, we first referenced the Mt1-related NRF2/GPX4 antioxidant pathway in astrocytes. Using dual-label IHC experiments for NRF2/GPX4, we demonstrated that the numbers of NRF2 and GPX4 in the SN of PD mice were significantly reduced (Fig. 8A). Based on this, we examined SLC7A11, a cystine/glutamate transporter in the NRF2/GPX4 antioxidant axis that regulates GSH production and is regulated by NRF2. We included SLC7A11 as an additional antioxidant factor in the NRF2/GPX4 axis for examination. Using qRT-PCR results, we confirmed that the mRNA levels of NRF2, SLC7A11, and GPX4 were significantly decreased in the SN of PD mice (Fig. 8B). Building on this, we validated the significant decline in the protein levels of NRF2, SLC7A11, and GPX4 in the midbrain of PD mice through Western blot experiments, suggesting that the activation of the ferroptosis process in the SN of PD mice may be related to this pathway (Fig. 8C). Furthermore, to validate our previous hypothesis, we also demonstrated that the gene content and protein levels of TH in the SN significantly decreased under PD conditions. In summary, these results roughly corroborate the unique characteristics of neuron damage due to crosstalk between oligodendrocytes and astrocytes in the SN of PD mice. Specifically, the secretion of FGF1 by oligodendrocytes and the interaction between FGF within the SN and its receptors FGFR1, FGFR2, and FGFR3 are weakened in a PD environment, which could impair their ability to protect neurons from ferroptosis. Concurrently, oligodendrocytes transfer the load of ferroptosis to astrocytes, accompanied by an overactivation of the oxidative phosphorylation process (Slc25a4, Ndufa4, and Atp6v0c), potentially leading to ferroptosis in some astrocytes and their reduced numbers within the SN. Additionally, the significant weakening of the NRF2/SLC7A11/GPX4 antioxidant pathway results in ferroptosis of DA neurons in the SN (Fig. 9).
A Using multiplex IHC, we captured the spatial distribution and quantitative changes of NRF2/GPX4 in the slices. Red represents NRF2, green represents GPX4, blue represents DAPI, and the white arrows indicate representative images showing the quantitative changes of NRF2/GPX4 in the SN region. This experiment was repeated once, with similar results. The scale bar is 400 μm. B, C The results of qRT-PCR and Western Blot indicate that the significant reduction in the number of dopaminergic neurons in the SN of PD mice may be caused by ferroptosis due to the inhibition of the NRF2/SLC7A11/GPX4 antioxidant pathway.
In the PD environment, the interaction between FGF1 secreted by oligodendrocytes and FGF9 in the SN with FGFR1, FGFR2, and FGFR3 receptors on astrocytes is weakened, which may reduce their ability to protect neurons from oxidative damage. Therefore, ferroptosis in the midbrain SN primarily occurs in astrocytes under the dysregulation of oligodendrocytes, subsequently leading to neuronal ferroptosis. The significant accumulation of ROS released by astrocytes and oligodendrocytes, the substantial buildup of iron ions in the SN, and the marked reduction of antioxidant factors NRF2/SLC7A11/GPX4 may be specific pathways leading to the substantial decrease in SN dopaminergic neurons.
Discussion
As a multi-system disorder, PD is characterized by widespread ion imbalances, inflammation, and immune shifts, which are key factors leading to neuronal inactivation. In this study, we used single-cell nucleus sequencing combined with spatial transcriptomics to examine the colocalization of glial cells in the SN at the spatial level. We thoroughly elucidated the involvement of the FGF signaling pathway between oligodendrocytes and astrocytes in PD, primarily involving FGF1, FGF9, and their receptors FGFR1, FGFR2, and FGFR3. Both oligodendrocytes and astrocytes showed significant oxidative phosphorylation processes, with a notable enrichment of ferroptosis-related genes in oligodendrocytes, which might be a crucial reason for the inactivation of DA neurons in PD mice. Additionally, we demonstrated the highly active oxidative phosphorylation processes in oligodendrocytes and astrocytes by measuring the protein expression levels of Slc25a4, Atp6v0c, and Ndufa4. We roughly confirmed the involvement of Ca2+ in activating the ferroptosis process by accelerating mitochondrial oxidative phosphorylation through the protein expression level of Mt1 and the Ca2+ content in astrocytes. Building on this, we verified the presence of significant ferroptosis in the midbrain of PD mice and a marked weakening of the NRF2/SLC7A11/GPX4 antioxidant pathway, which could be one of the main pathways leading to neuronal ferroptosis.
Glial cell activation is involved in the onset and progression of PD through various pathways, with the primary evidence for immune involvement being the activation of microglia and astrocytes. However, the role of neuroinflammation in PD pathogenesis remains undetermined24. For a long time, PD neuroinflammation has been considered a downstream response to neuronal damage. However, as research advances, changes in the physiological functions of glial cells are increasingly recognized as causally related to brain diseases25,26. Thus, we have expanded the scope of discussion to not only focus on the most abundant cell types in the PD SN but also to characterize PD-related glial cells and disease-specific molecular features throughout the midbrain. As previously described, our results characterized a significant increase in the colocalization and number of microglia and astrocytes in the SN of PD mice, indicating a classic pro-inflammatory pattern in the PD midbrain27. On this basis, we found that oligodendrocytes also exhibit significant colocalization in the SN, suggesting their potential involvement in PD pathology. This aligns with recent studies hypothesizing that oligodendrocytes may also participate in the pathological expansion of PD10. PD, a disease characterized primarily by pathological changes in the SN, has led to the long-term neglect of oligodendrocytes, mainly present in the midbrain white matter. Only in recent years have studies noticed that genes upregulated in postmortem brain tissue of PD patients are also significantly enriched in oligodendrocytes, with oligodendrocyte fluctuations preceding motor symptoms and pathological changes in the SN28.
Additionally, other studies have found an increase in oligodendrocytes in the MPTP mouse model, accompanied by notable morphological changes closely related to the reduced innervation of striatal DA neurons29. Takagi et al. also confirmed significant damage to oligodendrocytes in the SN of MPTP mice30. Here, using the Cell Chat tool, we characterized the pathways of glial cells in the midbrain of PD mice, identifying FGF signaling as a predominant pathway between oligodendrocytes and astrocytes. Although the overall weight of the FGF signaling pathway was reduced in the PD environment, oligodendrocytes enhanced their communication with astrocytes. This suggests that the interaction between oligodendrocytes and astrocytes via the FGF signaling pathway may also contribute to the development of PD pathology.
As a signaling pathway that plays a crucial role in brain neurodevelopment and neuroinflammation, we investigated the involvement pattern of oligodendrocytes and astrocytes in the FGF signaling pathway within the PD environment. This suggests that the complex intercellular communication within different networks in the PD brain might play a significant role in neuronal inactivation. Specifically, we characterized a significant reduction in the activation of the FGF signaling pathway under PD conditions, but an increased communication between oligodendrocytes and astrocytes. FGF1’s neuroprotective effects in neurodegenerative diseases (such as PD and AD) have been confirmed in both basic research and clinical studies. FGF1 is a critical factor in increasing DA levels in the brain and promoting the recovery of DA neurons from injury31,32,33. Furthermore, recent studies suggest that FGF1 can protect neuroblastoma (SH-SY5Y) cells from p53-dependent apoptosis. Given that p53 is a key gene in ferroptosis, this indicates that FGF1 can inhibit the ferroptosis process34.
Interestingly, the decreased expression levels of FGF9 in different neurological diseases have varying impacts on disease progression. For example, in MS, low levels of FGF9 can promote neuron survival, myelination, and inhibit the expression of pro-inflammatory chemokines35,36. However, in PD and Huntington’s disease, low levels of FGF9 impair neuron survival37,38,39. Importantly, FGF9 can upregulate the expression of heme oxygenase-1 and γ-glutamylcysteine synthetase to protect DA neurons from MPTP toxin-induced oxidative damage. This protective mechanism is not only mediated through the binding to FGFR receptors but also involves the activation of two parallel downstream pathways: ERK1/2 and AKT40. Beyond increasing the activity of CREB and NRF2, ERK1/2, as members of the MAPK family, are also involved in Ca2+ regulation, which can affect mitochondrial oxidative phosphorylation. This potentially indicates a link between FGF9 and ferroptosis41,42. Here, we discovered a significant reduction in the colocalization of FGF1 and its receptors (FGFR1, FGFR2, and FGFR3), as well as FGF9-FGFR2 within the SN in PD. This implies that the impaired communication between oligodendrocytes and astrocytes through the FGF signaling pathway could contribute to neuronal inactivation in PD.
As the biochemical mechanisms of ferroptosis in PD continue to advance, the associated iron accumulation, Ca2+ dysregulation, mitochondrial dysfunction, protein misfolding, and inflammation have provided new insights into the pathogenesis of PD43,44,45. Consequently, increasing research is focusing on the involvement of ferroptosis within the complex mechanisms of PD, aiming to further elucidate the specific mechanisms of neuronal ferroptosis43. However, previous studies have been limited by the causal relationship between iron deposition and neuronal inactivation and the lack of rigorous testing of glial cell activation through single-cell multi-omics. These studies have only described the downstream response processes of neuronal ferroptosis in PD, leaving the mechanisms by which PD predisposes neurons to ferroptosis unresolved10,46.
In contrast, recent studies have actively explored this issue, particularly by discussing how glial cells in PD induce ferroptosis. One study used whole-genome CRISPR screening to identify new regulators of ferroptosis, including the vesicle transport gene SEC24B, suggesting that SEC24B-regulated microglial ferroptosis leads to neurodegenerative changes47. Another study described the crosstalk network based on iron metabolism between PD glial cells and neurons. This includes microglial activation leading to iron accumulation, impaired astrocytic iron transport and reduction capacity, and substantial release of iron and ROS from oligodendrocytes10. Here, using single-nucleus sequencing and spatial transcriptomics, we revealed the involvement pattern of astrocytes and oligodendrocytes in ferroptosis through the FGF signaling pathway in PD. Specifically, the significantly reduced expression of FGF1 and FGF9 within the FGF signaling pathway under PD conditions weakens their protective ability against neuronal oxidative damage. Additionally, Ca2+ involvement and the highly active mitochondrial oxidative phosphorylation processes in astrocytes and oligodendrocytes facilitate the substantial release of ROS. Oligodendrocytes activate the ferroptosis process, and even though their numbers have not decreased, they may preferentially lead to the significant reduction of vulnerable DA neurons.
In summary, this study emphasizes the crucial role of astrocyte-oligodendrocyte crosstalk in the pathological progression of PD, involving the FGF signaling pathway. The reduced expression of FGF1 and FGF9 in the PD environment may promote neuronal ferroptosis, which includes significant increases in Ca2+ and marked attenuation of the NRF2-SLC7A11-GPX4 antioxidant pathway (Fig. 9). Furthermore, our research contributes to a deeper understanding of the involvement patterns of PD glial cells in the PD environment and the specific pathways through which glial crosstalk damages neurons. This could potentially aid in developing more effective anti-inflammatory treatments targeting the inhibition of ferroptosis and neuronal protection in the brains of PD patients, thereby offering promising avenues for advancing PD therapy.
Methods
Study design and Parkinson’s mouse model
The experimental mice used were SPF-grade C57BL/6J mice (male, 6–8 weeks old), with 15 mice in the Ctrl group and 15 mice in the PD group. According to previous research, we employed the MPTP model for PD mice48, with slight adjustments to the protocol: all mice underwent a one-week acclimatization period before the experiment began. All mice were housed in an SPF environment, fed standard mouse chow ad libitum, with the temperature maintained between 20 and 26 °C, relative humidity between 40 and 70%, and a 12-h light/12-h dark cycle. After the acclimatization period, all mice were weighed. The PD group mice received intraperitoneal injections of MPTP at 30 mg/kg/day, while the Ctrl group mice received saline injections for 5 consecutive days. Following the injections, all mice underwent behavioral tests, including the rotarod test and the open field test. Finally, all mice were deeply anesthetized with pentobarbital sodium, and their brains were collected for snRNA-seq, spatial transcriptomics analysis, TH immunohistochemical identification, immunofluorescence, qRT-PCR, and Western blot experiments. All housing and experimental procedures for the animals involved in this study complied with the revised 1986 Animals (Scientific Procedures) Act of the UK and the 2010/63/EU Directive of Europe, with approval from the Experimental Animal Science Ethics Committee of Shandong Sport University (Approval Number: 2023023).
Behavioral analyses
To evaluate PD-related behavioral performance in mice, all 30 mice underwent the rotarod test and open field test after the injection period in the PD group. In the rotarod test, we used a rotarod device to assess the mice’s motor coordination. All mice were trained on the rotarod for 300 s (at 5–10 rpm). After completing more than three rounds of this training, we began the formal experiment, recording the time each mouse took to fall off the rotarod.
In the open field test, we made certain adjustments based on a reported protocol49. The experimental arena was a specialized open field apparatus for mice (50 cm × 50 cm × 50 cm). Before testing, all mice were acclimated to the environment for 5 min. On the second day, the formal experiment was conducted by placing each mouse in the center of the apparatus and recording video for 5 min. The apparatus was cleaned with alcohol and water between each mouse to prevent interference with the results.
TH identification and multiplex IHC (Immunohistochemistry)
Mice were deeply anesthetized with intraperitoneal injection of pentobarbital sodium, followed by cardiac perfusion with PBS to remove blood. The entire brain and midbrain tissue were then harvested, preserved in paraffin, and sectioned into 5 μm slices. For immunohistochemistry, the sections were deparaffinized and rehydrated, followed by antigen retrieval using sodium citrate antigen retrieval solution (Thermo Fisher, 00-4955-58) (pH 6.0, 30 min). Endogenous peroxidase was blocked with 3% H2O2 for 20 min, and non-specific binding sites were minimized using 10% goat serum (Boster, AR1009) for 20 min. Sections were then incubated with TH antibody (58844, 1: 500, CST) overnight at 4 °C, and washed three times for 10 min each in TBST. Subsequently, goat anti-mouse IgG (1:2500) was added and incubated at room temperature for 2 h, with diaminobenzidine (DAB) as the chromogen (Maxim, DAB4033). The sections were counterstained with hematoxylin (Baso, BA4041). All DAB-immunostained sections were mounted, dried, and imaged using a slide scanner (Hamamatsu, NanoZoomer S360). The semiquantitative analysis of TH protein levels was represented by the positive area.
For multiplex immunofluorescence, the tissue was fixed with 4% paraformaldehyde for more than 24 h, permeabilized with Triton X-100 (Beyotime, Shanghai, China) for 20 min, and blocked with 5% bovine serum albumin (BSA) (Beyotime, Shanghai, China) at room temperature for 1 h. The sections were incubated with primary antibodies: FGF1 (TA500064, F001, Mouse, 1:100, OriGene Technologies), FGF9 (bs-5906R, F009, Rabbit, 1:100, Bioss), FGFR1 (BS1441, CJ36131, Rabbit, 1:500, Bioworld Technology), FGFR2 (BS74863, CC04241, Rabbit, 1:100, Bioworld Technology), FGFR3 (BS74263, CC19231, Rabbit, 1:100, Bioworld Technology), NRF2 (#BF8017, 0a14106, Mouse, 1:100, Affinity Biosciences), GPX4 (#DF6701, 78p5366, Rabbit, 1:100, Affinity Biosciences). Corresponding HRP-conjugated goat anti-rabbit/goat anti-mouse secondary antibodies (1:2500) were then added and incubated at room temperature for 2 h. Finally, the sections were incubated with DAPI solution at room temperature in the dark for 10 min and imaged for fluorescent signals using a fluorescence microscope (Olympus BX53). The images were analyzed using Case Viewer software, with the SN region (indicated by white squares) identified at 55X magnification. Image J software was used to quantify FGF and its receptors, NRF2, and GPX4 in the SN region.
Generation of single-nucleus suspensions
After euthanizing the mice, 15 ml of PBS solution was injected into the left atrium of the heart for perfusion, followed by extraction of the whole brain. According to the “Franklin and Paxinos Mouse Brain Atlas” (3rd edition), coronal sections were made using bregma as a reference point, covering from 2.92 mm posterior to the anterior fontanelle to 3.88 mm behind (AP: −2.92 to −3.88 mm), dorsal-ventral depth of 1.8–2.5 mm, and 1.2 mm lateral to the midline on both sides, encompassing the SNc and reticulata (SNr). The obtained sections were added to 1 mL of NST buffer, and the tissue was finely minced. The mixture was then filtered with ST Wash buffer, and the suspension was passed through a 40 µm mesh filter before being centrifuged at 4 °C (500 × g for 5 min). After discarding the supernatant, the nuclei were resuspended in 5 ml PBS + 1% BSA for washing and then counted.
Cell loading and 10x library preparation
According to the manufacturer’s protocol (Oebiotech, Shanghai), the 10× Genomics Chromium Next GEM Single Cell 3ʹ Reagent Kits v3 were used to generate single-cell libraries. The previously obtained suspension was resuspended in 1% BSA-PBS and then counted using the Countess® II Automated Cell Counter. The droplets were collected such that mRNA from the cells attached to the Cell Barcodes on the beads, forming Single Cell GEMs (Gel Bead-in-Emulsions). Subsequently, a reverse transcription reaction was run within the droplets to establish cDNA libraries. These libraries were distinguished by the sample index on the library sequences to identify the sample source of the target sequences. Following this, Cell Ranger was used for quality control of the samples, ultimately generating raw data in fastq format (with 46.4 GB for the Ctrl group and 48.4 GB for the PD group).
snRNA-seq merge and QC (Quality Control)
The libraries were quantified using Qubit, and the qualifying libraries were subjected to bridge amplification on a cBot. All libraries prepared for this study were sequenced on the HiSeq4000 system (Illumina) with 150 bp paired-end sequencing, and each library was sequenced across an entire lane. Following initial quality control by Cell Ranger, we further processed the data using the Seurat package (version 4.0.0) for additional quality control. We filtered out low-quality cells based on the distribution of nUMI, nGene, and percent.mito metrics. The specific quality control criteria were as follows: only cells with more than 200 genes, more than 1000 UMI, log10GenesPerUMI >0.7, <5% mitochondrial gene content, and less than 5% hemoglobin gene content were retained as high-quality cells. Additionally, we used DoubletFinder (version 2.0.3) to remove doublets. After quality control, the final number of cells obtained was 6255 for the Ctrl group (6857 before QC) and 8202 for the PD group (9329 before QC), which were included in downstream analyses. The filtered matrix in Seurat was normalized for library size to obtain normalized counts.
Cell type sub-clustering and annotation
We used the FindVariableGenes function in the Seurat package (mean.function = FastExpMean, dispersion.function = FastLogVMR) to select the top 2000 highly variable genes (HVGs). We then performed PCA (principal component analysis) for dimensionality reduction using the expression profiles of these highly variable genes. The results were visualized in a two-dimensional space using UMAP (non-linear dimensionality reduction).
Marker gene identification
We used the FindAllMarkers function in the Seurat package to identify marker genes and visualized these marker genes using the VlnPlot and FeaturePlot functions. Based on the results obtained from the SingleR package (version 1.4.1), we cross-referenced the top 20 marker genes of each cluster with the Cell Marker database (http://xteam.xbio.top/CellMarker/index.jsp), the Clinical Bioinformatics Database (https://www.aclbi.com/static/index.html#/), and relevant literature. The reference criteria were set to “mouse” and “brain.” We calculated the correlation between the cells to be identified and the reference datasets, which, to some extent, eliminated the interference of subjective human factors.
Cell-cell communication analysis with CellChat
We selected the CellChat package (version 1.1.3) to analyze potential intercellular communication in the cell data obtained from snRNA-seq50. We preprocessed the data using the default parameters and the functions identifyOverExpressedGenes, identifyOverExpressedInteraction, and ProjectData. Then, we used the functions computeCommunProb, filterCommunication, and computeCommunProbPathway to determine any possible ligand-receptor interactions. Finally, we aggregated the cell communication network using the aggregateNet function. In this study, we identified a total of 57 significant ligand-receptor interaction pairs.
Differential gene analysis and enrichment analysis
We used the FindMarkers function (test.use = presto) in the Seurat package to screen for differentially expressed genes. Genes with p < 0.05 and a fold change greater than 1.5 were considered significantly different. The significantly different genes were then subjected to GO and KEGG enrichment analysis using the hypergeometric distribution test.
Spatial transcriptomics sequencing
According to the “Franklin and Paxinos Mouse Brain Atlas” (3rd edition), coronal sections were made in reference to the bregma, from 2.92 mm anterior to 3.88 mm posterior (AP: −2.92 to −3.88 mm), with a dorsoventral depth of 1.8–2.5 mm, and a mediolateral range of ±1.2 mm from the midline. These sections included the SNc and SNr regions. Subsequent spatial transcriptomic analysis was performed on the samples. In the quantification and quality control results from Space Ranger, the average number of UMIs per spot was 28,400 for the Ctrl group and 54,978 for the PD group. The average number of genes per spot was 6276 for the Ctrl group and 8009 for the PD group. Post quality control, the data was used with the FindAllMarkers function (test.use = bimod) in the Seurat package to identify marker genes. The identified marker genes were visualized using the VlnPlot and FeaturePlot functions.
For mapping spatial cell types, we annotated the cell types defined in snRNA-seq to the spatial sequencing results by running the RCTD algorithm, following the study by Dylan M. Cable et al.51. During execution, the create.RCTD function used the default parameters, but with at least one cell per cell type and at least one UMI (unique molecular identifier) per spot. Additionally, the doublet_mode parameter for the run.RCTD function was set to FALSE to infer the cell composition at each spot.
Cell calcium ion content detection
Following the method of Qian Wang et al.52, we first rinsed fresh mouse SN tissue with XF-DMEM (containing 25 mM HEPES) and cut it into small pieces (~3 mg). After extensive washing, the tissue pieces were incubated with Liberase-TM (2.5 μg/ml, Roche) and DNase I (20 U/ml, Roche) at 37 °C with gentle agitation for 1 h. A single-cell suspension was obtained by passing the mixture through a 100 μm cell strainer (Thermo Fisher Scientific). We isolated astrocytes using magnetic beads labeled with Atp13a4 (BioLegend, eBioscience, R&D Systems, Miltenyi Biotec).
Following the protocol of the calcium content colorimetric assay kit (#S1063S, Beyotime, China), cells were collected by centrifugation at 600 rpm for 5 min at 4 °C. Cell lysis buffer was added to the cells in a ratio of 150 µl per well in a 6-well plate. After thorough lysis, the mixture was centrifuged at 10,000–14,000 rpm for 3–5 min at 4 °C, and the supernatant was collected. We added 50 µl of standard and 50 µl of sample to each well, followed by 150 µl of assay working solution per well. After mixing and incubating at room temperature in the dark for 5 min, the absorbance at 575 nm was measured using a microplate reader. The Ca2+ content of oligodendrocytes and astrocytes was determined based on the standard curve prepared from calcium standards.
Measurement of tissue iron content
According to the protocol of the Tissue Iron Assay Kit (#A039-2-1, Nanjing Jiancheng Bioengineering Institute, China), take 20 mg of mouse SN tissue and add 180 µl of normal saline. Homogenize the tissue mechanically in an ice-water bath, then centrifuge at 2500 rpm for 10 min. Take 50 µl of the supernatant as the sample to be tested, add 150 µl of iron chromogenic reagent, and mix well. Boil the mixture in a water bath for 5 min, then cool it under running water. Centrifuge at 2500 rpm for 10 min and take 100 µl of the supernatant. Measure the absorbance at 520 nm with double-distilled water as a blank. Quantify the iron content of each sample based on the absorbance values and the standard curve prepared from the iron standard solution.
Measurement of tissue GSH content
According to the GSH assay kit operation manual (A006-1-1, Nanjing Jiancheng Bio, China), prepare a 10% homogenate from the mouse SN tissue, centrifuge at 2500 rpm for 10 min, then take the supernatant and mix it with reagent one application solution and centrifuge again. Use the supernatant for the color reaction. After standing at room temperature for 5 min, detect the absorbance of each well at 420 nm using a microplate reader and calculate the GSH content in the SN tissue according to the formula.
Measurement of total SOD activity in tissue
According to the total superoxide dismutase (T-SOD) assay manual (#A001-1, Nanjing Jiancheng Bio, China), prepare a 10% homogenate from the mouse SN tissue, centrifuge at 2500 rpm for 10 min, and identify the supernatant. Mix the supernatant with the working solution, leave it at room temperature for 10 min, and detect the absorbance of each well at 550 nm using a microplate reader. Calculate the SOD activity in the SN tissue according to the formula.
Tissue malondialdehyde (MDA) content determination
According to the protocol of the Malondialdehyde (MDA) Assay Kit (#A003-1, Nanjing Jiancheng Bioengineering Institute, China), take 20 mg of mouse SN tissue and add 180 µl of normal saline to prepare a 90% tissue homogenate. Add 200 µl of reagent one, 3000 µl of reagent two, and 1000 µl of reagent three, as per the manual instructions. Cover the centrifuge tube and make a small hole in the lid with a needle. Mix thoroughly using a vortex mixer, followed by incubation in a 95 °C water bath for 40 min. After boiling, cool the mixture under running water, then centrifuge at 3500 rpm for 10 min. Take 200 µl of the supernatant and measure the absorbance at 532 nm. The MDA content in the tissue is then determined based on the absorbance values obtained.
qRT-PCR
Cut 20 mg of SN tissue from the sample and lyse the tissue using Trizol. After lysis, centrifuge the sample at 12000 rpm for 15 min at 4 °C. Transfer the uppermost supernatant to an EP tube containing 500 µl of isopropanol, mix well, and let it stand for 10 min. Centrifuge again at 12,000 rpm for 10 min at 4 °C. Discard the upper isopropanol layer, add 1 ml of 70% ethanol, and centrifuge at 7500 rpm for 5 min at 4 °C. Check the RNA pellet, discard the supernatant, and obtain the RNA solution. Dissolve the RNA in 20 µl of DEPC water, based on the RNA quantity.
According to the cDNA synthesis kit instructions: First, take 15 µl of Total RNA and add it to a new PCR tube. Add 4 µl of 5× TransScript and 1 µl of gDNA Remover (total volume 20 µl). Mix gently, incubate at 50 °C for 5 min, heat at 85 °C for 2 min for inactivation, and finally set the cycle at 4 °C. Following the qPCR SuperMix kit instructions, add 1 µl of cDNA, 0.5 µl of forward primer, and 0.5 µl of reverse primer (NRF2: F-ATTGCCTGTAAGTCCTGGTCA, R-GCTCTTTGGACATCATTTCG; SLC7A11: F-GGTGGTGTGTTTGCTGTC, R-GCTGGTAGAGGAGTGTGC; GPX4: F-CCGAAGTAAACTACACTCAGCTCG, R-TTGATCTCTTCGTTACTCCCTGGC; β-actin: F-ACTGCCGCATCCTCTTCCT, R-TCAACGTCACACTTCATGATGGA; TH: F-TCAGAGGAGCCCGAGGTC, R-GGGCGCTGGATACGAGAG), 10 µl of 2×SuperMix, and 8 µl of water. Use a fluorescent quantitative PCR machine and select the three-step method to set the qPCR reaction system: 94 °C for 30 s, 94 °C for 5 s, 55 °C for 15 s, and 72 °C for 10 s. Cycle 40 times, and finally obtain the curve.
Western blot
This experiment is divided into two parts. The first part involves extracting total protein from oligodendrocytes and astrocytes. The extraction of a single-cell suspension of oligodendrocytes is the same as the previously described method for obtaining an astrocyte suspension. Magnetic beads labeled with Plpl (Oligodendrocyte) are used to obtain the oligodendrocyte suspension. Pre-cooled RIPA lysis buffer and protease inhibitor are mixed to a working solution in a ratio of 100:1. The second part involves extracting total protein from mouse SN. Take 20 mg of SN tissue and add 200 µl of working solution. Then, use a high-throughput tissue grinder to homogenize the tissue (50 Hz, 10 s/time, with a 5 s interval each time), repeating 6 times. After centrifugation, collect the protein supernatant.
First, we determine the protein concentration using the BCA method. Then, for each well, set the loading volume of the protein and loading buffer mixture to 10 µl. The electrophoresis conditions are an initial voltage of 80 V for 30 min, followed by a constant voltage of 120 V until the protein completely migrates out of the gel. Remove the gel for membrane transfer under conditions of 200 mA for 50 min. After transfer, block the membrane with 5% skim milk at room temperature on a shaker for 1 h. Cut the membrane according to the Marker and protein molecular weights and incubate the cut membrane with primary antibodies overnight at 4 °C. The information and dilution ratio for each antibody are as follows: Slc25a4 (#DF6674, 35d7800, Rabbit, 1:1000, Affinity Biosciences, US), Ndufa4 (#AF5277, 8j9620, Rabbit, 1:1000, Affinity Biosciences, US), Atp6v0c (K113651P, 20231106, Rabbit, 1:1500, Solarbio, China), Mt1 (#DF13877, 37h7757, Rabbit, 1:1000, Affinity Biosciences, US), TH (#BS1432, CA36131, Mouse, 1:500, Bioworld Technology, China), NRF2 (#BS1258, AA12152, Mouse, 1:1000, Bioworld Technology, China), SLC7A11 (#BS67354, CN16212, Rabbit, 1:1000, Bioworld Technology, China), GPX4(#DF6701, 78p5366, Rabbit, 1:100, Affinity Biosciences), β-actin (#GB11001, 20231213, Rabbit, 1:1000, Servicebio, China).
After washing the membrane with TBST buffer, incubate the membrane with secondary antibodies at room temperature on a shaker for 1.5 h. The secondary antibody information is as follows: HRP Goat anti-mouse IgG Antibody (#Poly4053, 1:2500, Biolegend, US), Goat anti-Rabbit IgG Antibody (#S0001, 1:5000, Affinity Biosciences, US). Finally, use ECL substrate (Beyotime, China) for membrane development and analyze the protein grayscale values using the Image J software in the Image Club system.
Data availability
Sequence data that support the findings of this study have been deposited in the GSA database of the China National Center for Bioinformation under GSA number CRA017010 and CRA015600.
References
Cacabelos, R. Parkinson’s disease: from pathogenesis to pharmacogenomics. Int. J. Mol. Sci. 18, 551 (2017).
Dauer, W. & Przedborski, S. Parkinson’s disease: mechanisms and models. Neuron 39, 889–909 (2003).
Inamdar, N. N., Arulmozhi, D. K., Tandon, A. & Bodhankar, S. L. Parkinson’s disease: genetics and beyond. Curr. Neuropharmacol. 5, 99–113 (2007).
LeWitt, P. A. Levodopa therapy for Parkinson’s disease: pharmacokinetics and pharmacodynamics. Mov. Disord. 30, 64–72 (2015).
Giridharan, V. V., Barichello De Quevedo, C. E. & Petronilho, F. Microbiota-gut-brain axis in the Alzheimer’s disease pathology - an overview. Neurosci. Res. 181, 17–21 (2022).
Depierreux, F. et al. Parkinson’s disease multimodal imaging: F-DOPA PET, neuromelanin-sensitive and quantitative iron-sensitive MRI. NPJ Parkinsons Dis. 7, 57 (2021).
Biondetti, E. et al. The spatiotemporal changes in dopamine, neuromelanin and iron characterizing Parkinson’s disease. Brain 144, 3114–3125 (2021).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Zhang, S. et al. Ferroptosis increases obesity: crosstalk between adipocytes and the neuroimmune system. Front. Immunol. 13, 1049936 (2022).
Wang, Z. L., Yuan, L., Li, W. & Li, J. Y. Ferroptosis in Parkinson’s disease: glia-neuron crosstalk. Trends Mol. Med. 28, 258–269 (2022).
Festa, L. K., Grinspan, J. B. & Jordan-Sciutto, K. L. White matter injury across neurodegenerative disease. Trends Neurosci. 47, 47–57 (2024).
Friedrich, I. et al. Cell specific quantitative iron mapping on brain slices by immuno-microPIXE in healthy elderly and Parkinson’s disease. Acta Neuropathol. Commun. 9, 47 (2021).
Jhelum, P. et al. Ferroptosis mediates cuprizone-induced loss of oligodendrocytes and demyelination. J. Neurosci. 40, 9327–9341 (2020).
McTigue, D. M. & Tripathi, R. B. The life, death, and replacement of oligodendrocytes in the adult CNS. J. Neurochem. 107, 1–19 (2008).
Yang, K. et al. White matter changes in Parkinson’s disease. NPJ Parkinsons Dis. 9, 150 (2023).
Zhang, S. et al. Investigating the mechanisms of inflammation and immune alterations in Parkinson’s disease using spatial transcriptomics techniques. Brain Res. Bull. 217, 111076 (2024).
Zhang, Q., Chen, Z., Zhang, K., Zhu, J. & Jin, T. FGF/FGFR system in the central nervous system demyelinating disease: recent progress and implications for multiple sclerosis. CNS Neurosci. Ther. 29, 1497–1511 (2023).
Huang, J. Y. & Chuang, J. I. Fibroblast growth factor 9 upregulates heme oxygenase-1 and gamma-glutamylcysteine synthetase expression to protect neurons from 1-methyl-4-phenylpyridinium toxicity. Free Radic. Biol. Med. 49, 1099–1108 (2010).
Yuan, Y. et al. Lyssavirus M protein degrades neuronal microtubules by reprogramming mitochondrial metabolism. mBio 15, e0288023 (2024).
Rossi, A., Pizzo, P. & Filadi, R. Calcium, mitochondria and cell metabolism: a functional triangle in bioenergetics. Biochim. Biophys. Acta Mol. Cell Res. 1866, 1068–1078 (2019).
Kadenbach, B. Regulation of mammalian 13-subunit cytochrome c oxidase and binding of other proteins: role of NDUFA4. Trends Endocrinol. Metab. 28, 761–770 (2017).
Boiko, D. I. et al. Melatonergic receptors (Mt1/Mt2) as a potential additional target of novel drugs for depression. Neurochem. Res. 47, 2909–2924 (2022).
Lv, Q. K. et al. Melatonin MT1 receptors regulate the Sirt1/Nrf2/Ho-1/Gpx4 pathway to prevent alpha-synuclein-induced ferroptosis in Parkinson’s disease. J. Pineal Res. 76, e12948 (2024).
de Lau, L. M. & Breteler, M. M. Epidemiology of Parkinson’s disease. Lancet Neurol. 5, 525–535 (2006).
Tremblay, M. E., Cookson, M. R. & Civiero, L. Glial phagocytic clearance in Parkinson’s disease. Mol. Neurodegener. 14, 16 (2019).
Reemst, K., Noctor, S. C., Lucassen, P. J. & Hol, E. M. The indispensable roles of microglia and astrocytes during brain development. Front. Hum. Neurosci. 10, 566 (2016).
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Annese, V. et al. Evidence of oligodendrogliosis in 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP)-induced Parkinsonism. Neuropathol. Appl. Neurobiol. 39, 132–143 (2013).
Takagi, S., Hayakawa, N., Kimoto, H., Kato, H. & Araki, T. Damage to oligodendrocytes in the striatum after MPTP neurotoxicity in mice. J. Neural Transm.114, 1553–1557 (2007).
Liu, Y., Deng, J., Liu, Y., Li, W. & Nie, X. FGF, Mechanism of action, role in parkinson’s disease, and therapeutics. Front. Pharm. 12, 675725 (2021).
Thorns, V. & Masliah, E. Evidence for neuroprotective effects of acidic fibroblast growth factor in Alzheimer disease. J. Neuropathol. Exp. Neurol. 58, 296–306 (1999).
McLay, R. N., Freeman, S. M. & Zadina, J. E. Administration of FGF-1 through transfected cells alleviates MPTP toxicity in mice. Neurotox. Res. 3, 249–253 (2001).
Pirou, C. et al. FGF1 protects neuroblastoma SH-SY5Y cells from p53-dependent apoptosis through an intracrine pathway regulated by FGF1 phosphorylation. Cell Death Dis. 8, e3023 (2017).
Lindner, M. et al. Fibroblast growth factor signalling in multiple sclerosis: inhibition of myelination and induction of pro-inflammatory environment by FGF9. Brain 138, 1875–1893 (2015).
Thummler, K. et al. Fibroblast growth factor 9 (FGF9)-mediated neurodegeneration: Implications for progressive multiple sclerosis?. Neuropathol. Appl. Neurobiol. 49, e12935 (2023).
Huang, J. Y., Hong, Y. T. & Chuang, J. I. Fibroblast growth factor 9 prevents MPP+-induced death of dopaminergic neurons and is involved in melatonin neuroprotection in vivo and in vitro. J. Neurochem. 109, 1400–1412 (2009).
Kanda, T. et al. Self-secretion of fibroblast growth factor-9 supports basal forebrain cholinergic neurons in an autocrine/paracrine manner. Brain Res. 876, 22–30 (2000).
Yusuf, I. O. et al. Fibroblast growth factor 9 stimulates neuronal length through NF-kB signaling in striatal cell Huntington’s disease models. Mol. Neurobiol. 58, 2396–2406 (2021).
Chuang, J. I. et al. FGF9-induced changes in cellular redox status and HO-1 upregulation are FGFR-dependent and proceed through both ERK and AKT to induce CREB and Nrf2 activation. Free Radic. Biol. Med. 89, 274–286 (2015).
Veeranna, K. T. et al. Calpain mediates calcium-induced activation of the erk1,2 MAPK pathway and cytoskeletal phosphorylation in neurons: relevance to Alzheimer’s disease. Am. J. Pathol. 165, 795–805 (2004).
Huntington, T. E. & Srinivasan, R. Astrocytic mitochondria in adult mouse brain slices show spontaneous calcium influx events with unique properties. Cell Calcium 96, 102383 (2021).
Ding, X. S. et al. Ferroptosis in Parkinson’s disease: molecular mechanisms and therapeutic potential. Ageing Res. Rev. 91, 102077 (2023).
Dionisio, P. A., Amaral, J. D. & Rodrigues, C. M. P. Oxidative stress and regulated cell death in Parkinson’s disease. Ageing Res. Rev. 67, 101263 (2021).
Angelova, P. R. et al. Alpha synuclein aggregation drives ferroptosis: an interplay of iron, calcium and lipid peroxidation. Cell Death Differ. 27, 2781–2796 (2020).
Stockwell, B. R. et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell 171, 273–285 (2017).
Ryan, S. K. et al. Microglia ferroptosis is regulated by SEC24B and contributes to neurodegeneration. Nat. Neurosci. 26, 12–26 (2023).
Lee, E. et al. MPTP-driven NLRP3 inflammasome activation in microglia plays a central role in dopaminergic neurodegeneration. Cell Death Differ. 26, 213–228 (2019).
Heng, Y. et al. Ginsenoside Rg1 attenuates motor impairment and neuroinflammation in the MPTP-probenecid-induced parkinsonism mouse model by targeting alpha-synuclein abnormalities in the substantia nigra. Toxicol. Lett. 243, 7–21 (2016).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Wang, Q. et al. IL-27 signalling promotes adipocyte thermogenesis and energy expenditure. Nature 600, 314–318 (2021).
Acknowledgements
This study was supported by grants from the central government, guiding local science and technology development funds (YDZX2022091) and the Shandong Province medical health science and technology project (202303030782). We give great gratitude to OE Biotech Limited for their technique support.
Author information
Authors and Affiliations
Contributions
S.Z.: Writing—original draft, Data curation, M.Y.: Conceptualization, Data curation; X.J.: Data curation; Y.L.: Methodology; W.M.: Software; L.D. and Z.L.: Investigation; Y.L. and X.T.: Resources, Funding acquisition; Q.W.: Supervision, Writing—review and editing, Validation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, S., Yan, M., Jiang, X. et al. Oligodendrocyte-astrocyte crosstalk in Parkinson’s disease mediates neuronal ferroptosis via the FGF signaling pathway. npj Parkinsons Dis. 11, 136 (2025). https://doi.org/10.1038/s41531-025-00995-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41531-025-00995-0
This article is cited by
-
Biomimetic nanodelivery system with simultaneous blood–brain barrier-crossing and neuroprotective abilities for anti-parkinsonian therapy
Chinese Medicine (2025)
-
Mitochondrial Calcium Channels and MAM Interaction in Calcium Homeostasis Dysregulation in Parkinson’s Disease
Neurochemical Research (2025)