Abstract
The common octopus (Octopus vulgaris) is a highly valued seafood species, whose flavour and palatability are often linked to its geographic origin. However, globalized supply chains have increased the risk of mislabelling. Lipid metabolism in marine organisms is shaped by environmental conditions, then lipidomics offers a promising tool for origin authentication. This study used HILIC-LC-MS&MS/MS to profile the polar lipidome of O. vulgaris arm muscle from three Iberian Atlantic landing sites: Ría de Arousa (Spain), Peniche, and Santa Luzia (Portugal). While total lipid content was consistent across sites, Peniche samples showed lower phospholipid levels, and Santa Luzia samples showed elevated LPE/PE and LPC/PC ratios, possibly reflecting stress or degradation. Ether-linked phospholipids and the sphingolipid CAEP were most abundant in Ría de Arousa, contributing to site discrimination. These findings demonstrate the potential of lipidomic profiling as a tool for seafood traceability, offering insights into environmental and dietary influences on lipid composition.
Similar content being viewed by others
Introduction
The common octopus (Octopus vulgaris Cuvier, 1797) is one of the most significant cephalopod fisheries in the European Union. These fisheries are both economically and ecologically important, particularly in regions such as Southern Europe and Northwest Africa1. Widely distributed across the Northeast Atlantic, Mediterranean, and Eastern Central Atlantic, O. vulgaris is particularly abundant along the Atlantic coasts of Portugal, Spain, Morocco, Mauritania, and Senegal2. In the Iberian Peninsula, it is one of the highest-revenue species, playing a particularly significant role in the fisheries of the Algarve region in Portugal and Galicia in Spain. However, landings have declined in recent years, especially in Galicia, where environmental changes in estuaries (Rias) and anthropogenic factors, such as overexploitation, ineffective control of rules and regulations, or illegal fishing practices, have been identified as key contributors to this trend3.
In the Iberian Peninsula, the common octopus stands out as one of the most significant fishing resources, enrolling thousands of people on its fisheries along the coastal waters of Galicia (Spain) and mainland Portugal (especially on the Algarve)4,5,6. The common octopus’ fishery is not only a cornerstone of the small-scale fishing sector but also a critical driver of regional economies. In Santa Luzia (Algarve, Portugal), octopus accounts for 99% of both the total weight and value of annual landings1. In Galicia, the common octopus fisheries support local economies and small-scale fishing communities, providing substantial economic benefits. Galicia serves as a major hub for octopus exports, supplying both domestic and international demand7. The high market value of octopus has led to increased interest in sustainable fishing practices and mariculture to ensure supply stability8,9. Efforts to improve sustainability within the octopus’ fishery have contributed to price stabilization and expand access to premium markets9.
Octopus vulgaris has a short lifespan of 12–18 months with non-overlapping generations, making its populations highly sensitive to environmental variability10. In Southern European countries, the management of octopus fisheries primarily falls under the jurisdiction of national and/or local governments, likely reflecting the historical significance of exploitation and the economic importance of this resource1. Nevertheless, there is a longstanding acknowledgment that the common octopus has increasingly become a depleted resource, marked by declining catches in the most important supplier countries (Morocco and Mauritania in Northern Africa), largely attributable to overfishing practices10,11, illegal fishing practices, misreporting, and unauthorized sales1.
Octopus vulgaris is a highly versatile feeder and a generalist predator, whose diet is dependent on available prey. The diet of the common octopus encompasses a broad spectrum of species, including bivalves, crustaceans, finfish, and polychaetes12,13,14, while cannibalism and autophagy are also frequently documented in this species13. Factors such as seasonal and geographical differences also shape the feeding habits of the common octopus15. This feeding plasticity enables the common octopus to rapidly adapt to diverse environments, a trait likely mirrored in its lipidome, making it a potential tool for determining geographic origin.
Globalization of seafood markets allowed a better distribution of products around the world, with once local specific products currently being available in a global market16. However, geographic origin is often linked to the quality of specific products in the eyes of consumers, impacting the valuation of these products17,18,19,20. Additionally, a greater consumer awareness on environmental issues has led to an increased demand for more information regarding harvest or production methods (wild vs. farmed), as well as the management of wild stocks21,22. A system providing information at these levels, needs reliable traceability tools to obtain and confirm origin certification of seafood products.
The lipid diversity identified in marine organisms plays a crucial role in several physiological processes, including reproduction, growth, immunological responses, and energy reserves23. Lipidome profiles exhibit species-specific characteristics, making their analysis a valuable tool for detecting mislabelling and species substitution in the supply chain, particularly within marine organisms24,25. Due to the influence of the surrounding environment in the growth of marine species and the feeding resources they use, unique lipid signatures can also be associated with specific geographic origins26,27,28. Fatty acid and lipid analysis have been successfully used to discriminate the geographic origin of several seafood products, such as the green seaweeds Ulva spp.26 and Codium tomentosum28, the brown macroalga Saccharina latissima27, the common cockle Cerastoderma edule29, the common octopus O. vulgaris30, the halophyte plant Salicornia ramosissima31, or the goose neck barnacle Pollicipes pollicipes19.
The present study aimed to evaluate the potential use of lipidomic fingerprints as a biochemical tool to pinpoint the geographic origin of common octopus captured and landed in three locations of the Iberian Atlantic coast: Ria de Arousa (RAr), Peniche (Pe), and Santa Luzia (SL). We hypothesized that the polar lipid composition of O. vulgaris arm muscle will be unique over different landing locations along the Iberian Atlantic coast and that these natural barcodes could be successfully used to verify claims on its geographic origin.
Results
Lipid and phospholipid content
Total lipid content of O. vulgaris revealed that Pe (45.86 ± 7.47 mg g−1 DW) was the landing location that presented specimens yielding the highest lipid content and in decreasing order RAr and SL (41.53 ± 4.29 mg g−1 DW and 40.40 ± 7.53 mg g−1 DW, respectively, Table 1). However, no significant differences were found in the lipid content of common octopus’ samples originating from the different locations being studied. Phospholipid quantification expressed as percentage of total lipids, showed significant differences between Pe and the other sites, with Pe recording the lowest value at 36.98% ± 4.59 (Table 1).
Polar lipidome profiling
The analysis performed using HILIC-LC-MS allowed the identification of 372 lipid species (m/z values) from 13 polar lipid classes in the lipidome of common octopus’ arm muscle, including glycerophospholipids (Supplementary Fig. S1) and sphingolipids (Supplementary Fig. S2). In the glycerophospholipid category were identified phosphatidylcholine—PC (Fig. 1a) whose main lipid species were PC 38:6 (PC 16:0_22:6) and PC 36:5 (PC 16:0_20:5), lyso PC–LPC (Supplementary Fig. S3a) with LPC 22:6 and LPC 20:5 as the most abundant lipid species; phosphatidylethanolamine—PE (Fig. 1b) with the ether species PE O-38:6/P-38:5 (PE O-18:1/20:5 and PE P-18:0/20:5) and PE O-38:5/P-38:4 (PE O-18:1/20:4 and P-18:0/20:4) showing the highest abundance, lyso PE—LPE with the most abundant species corresponding to LPE O-18:1/P-18:0 and LPE 18:0 (Supplementary Fig. S3b); phosphatidylinositol—PI, with PI 38:5 (PI 18:0_20:5) being the most abundant molecular species in specimens sampled from the three locations surveyed, while the second most abundant in RAr and Pe was PI 38:4 (PI 18:0_20:4) and 36:5 (PI 16:0_20:5) in SL (Fig. 2a); phosphatidylserine—PS with the most abundant lipid species corresponding to PS 38:5 (PS 18:0_20:5) and PS O-40:6 (PS O-18:0/22:6) (Fig. 2b); and phosphatidylglycerol—PG with PG 34:1 (PG 16:0_18:1) as the most abundant lipid species (Supplementary Fig. S4a). In the sphingolipid category (Supplementary Fig. S2) were identified sphingomyelin—SM featuring SM 32:1;O2 (SM 16:1;O2/16:0) and SM 38:2;O2 (SM 16:1;O2/22:1) (Supplementary Fig. S5a); ceramide—Cer with Cer 32:1;O2 (Cer 16:1;O2/16:0) and Cer 30:1;O2 (Cer 18:1;O2/12:0) being the most abundant species (Supplementary Fig. S5b); hexosylceramide—HexCer with HexCer 38:2;O2 (HexCer 16:1;O2/22:1) and HexCer d40:1 being the only species identified in this class (Supplementary Fig. S4b); ceramides aminoethylphosphonate—CAEP which presented CAEP 32:1;O2 (CAEP 16:1;O2/16:0) as the most abundant lipid species in all sampling sites, while the second most abundance was CAEP 38:2;O2 (CAEP 16:1;O2/22:1) in RAr and SL, and CAEP 35:3;O2 (CAEP 19:3;O2/16:0) in Pe (Fig. 3a); N-methyl ceramide aminoethylphosphonate—N-methyl-CAEP with the lipid species N-methyl-CAEP 32:1;O2 (N-methyl-CAEP 16:1;O2/16:0) and N-methyl-CAEP 38:2;O2 (N-methyl-CAEP 16:1;O2/22:1) showing the highest abundance (Fig. 3b); and ceramide phosphoethanolamine – PE-Cer showing PE-Cer 35:3;O2 (PE-Cer 19:3;O2/16:0) and PE-Cer 32:1;O2 (PE-Cer 16:1;O2/16:0) as the most abundant lipid species (Fig. 3c)] (Supplementary Tables S1 and S2). The ether-linked phospholipids plasmanyl and plasmenyl are isomers with the same m/z value, and in this study, they were counted as a single molecular lipid species. They were only distinguished when MS/MS data provided clear evidence to identify one of the isomers.
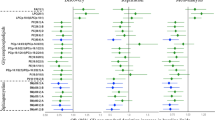
Relative abundance of phospholipid species of a phosphatidylcholine (PC) and b phosphatidylethanolamine (PE) identified in common octopus’ (Octopus vulgaris) arm muscle captured in three locations along the Iberian Atlantic coast (Ria Arousa—RAr, Peniche—Pe, and Santa Luzia—SL). Data represent relative abundance within the lipid class. Only lipid species with relative abundance >1% were represented. Total lipid species and their relative abundance are summarized in Supplementary Table S1. Different letters indicate significant differences between sampling locations (Tukey’s HSD post hoc test, p < 0.05).
Relative abundance of phospholipid species of a phosphatidylinositol (PI) and b phosphatidylserine (PS) identified in common octopus’ (Octopus vulgaris) arm muscle captured in three locations along the Iberian Atlantic coast (Ria Arousa—RAr, Peniche—Pe, and Santa Luzia—SL). Data represent relative abundance within the lipid class. Different letters indicate significant differences between sampling locations (Tukey’s HSD post hoc test, p < 0.05).
Relative abundance of lipid species of a ceramide aminoethylphosphonate (CAEP), b N-methyl ceramide aminoethylphosphonate (N-methyl CAEP), and c ceramide phosphoethanolamine (PE-Cer) identified in common octopus’ (Octopus vulgaris) arm muscle captured in three locations along the Iberian Atlantic coast (Ria Arousa—RAr, Peniche—Pe, and Santa Luzia—SL). Data represent relative abundance within the lipid class. Different letters indicate significant differences between sampling locations (Tukey’s HSD post hoc test, p < 0.05).
Samples from RAr showed the lowest values of LPE/PE and LPC/PC ratios (0.54 ± 0.19 and 0.34 ± 0.14, respectively), while octopus from SL showed the highest ratio (LPE/PE: 0.78 ± 0.21, LPC/PC: 0.83 ± 0.23). The values of LPE/PE and LPC/PC ratios for Peniche samples were 0.72 ± 0.15 and 0.57 ± 0.12, respectively (Fig. 4).
LPE/PE and LPC/PC ratio identified in lipid extracts of common octopus’ (Octopus vulgaris) arm muscle captured in three locations along the Iberian Atlantic coast (Ria Arousa—RAr, Peniche—Pe, and Santa Luzia—SL). Different letters indicate significant differences between sampling locations (Tukey’s HSD post hoc test, p < 0.05).
The relative abundance of essential fatty acids, namely eicosapentaenoic acid (EPA, 20:5 n-3) and docosahexaenoic acid (DHA, 22:6 n-3) in the fatty acyl composition of the main phospholipid classes was determined. PC was the phospholipid class with the highest percentage of lipid species containing DHA in their composition (31% in RAr, 37% in Pe and 36% in SL, Fig. 5a, b, c), while, in PE the lipid species containing EPA in their structure accounted for 36% in RAr, 39% in Pe and 41% in SL (Fig. 5d, e, f). Most of the lipid species in PI presented EPA as an esterified fatty acid in their structure (70% in RAr, 73% in Pe, and 76% in SL, Fig. 6a, b, c). No PI lipid species were identified containing only DHA (Fig. 6a, b, c). In PS, the relative abundance of lipid species containing EPA (18% in RAr, 19% in Pe, 16% in SL) was slightly lower than those containing DHA (22% in RAr, 25% in Pe, 24% in SL, Fig. 6d, e, f).
Relative abundance (%) of lipid species containing EPA (20:5 n-3), DHA (22:6 n-3), both (EPA & DHA) or other fatty acids (FA) in the fatty acyl composition of phosphatidylcholine (PC) in samples of common octopus’ (Octopus vulgaris) arm muscle from a Ria Arousa (RAr), b Peniche (Pe), c Santa Luzia (SL) and in the composition of phosphatidylethanolamine (PE) in samples from d RAr, e Pe, f SL. Lipid species whose fatty acyl composition was not identified were included in other FA. Different letters under relative abundance indicate significant differences between sampling locations (Tukey’s HSD post hoc test, p < 0.05).
Relative abundance (%) of lipid species containing EPA (20:5 n-3), DHA (22:6 n-3), both (EPA & DHA) or other fatty acids (FA) in the fatty acyl composition of phosphatidylinositol (PI) in samples of common octopus’ (Octopus vulgaris) arm muscle from a Ria Arousa (RAr), b Peniche (Pe), c Santa Luzia (SL) and in the composition of phosphatidylserine (PS) in samples from d RAr, e Pe, f SL. Lipid species whose fatty acyl composition was not identified were included in other FA. Different letters under relative abundance indicate significant differences between sampling locations (Tukey’s HSD post hoc test, p < 0.05).
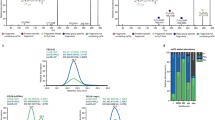
The ANOVA test of normalized extracted ion chromatograms (XIC) areas of polar lipid revealed the existence of significant differences between locations in 268 of the 372 identified lipid species (Supplementary Table S2). The eigenvalues of the two principal components of the principal component analysis (PCA) analysis performed using the normalized XIC areas of polar lipid species explained 62.7% of total variance, with principal component 1 (PC1) and principal component 2 (PC2) axes explaining 49.6% and 13.1%, respectively (Fig. 7a). PC1 allowed the separation of RAr samples from those originating from Pe and SL, while PC2 separated Pe from SL samples. Samples originating from RAr displayed a higher dispersion in the PCA plot when compared with the other two locations (Fig. 7a). These results were supported by the hierarchical clustering dendrogram (Fig. 7b) and heatmap (Fig. 7c) performed using the top 50 lipid species sorted by analysis of variance test displaying the lowest p-values (Supplementary Table S2). The lipid species that most contributed for the discrimination of the samples and their clustering were ether-linked phospholipids, of PC and PE, diacyl forms of PS and PI and the sphingolipid CAEP. All of them were more abundant in samples from RAr, except the lipid species LPC 18:4, PI 38:1, PI39:5, and PE O-38:1/P-38:0, which were more expressed in the samples from Pe and SL (Fig. 7c).
a Principal component analyses (PCA), b dendrogram, and c hierarchical clustering heatmap of the top 50 lipid species sorted by analysis of variance test displaying the lowest p-values of log-transformed normalized extracted-ion chromatogram (XIC) areas of total polar lipid species identified in the arm muscle of Octopus vulgaris samples captured in three locations along the Iberian Atlantic coast (Ria Arousa—RAr, Peniche—Pe, and Santa Luzia—SL).
Discussion
The lipidome of marine organisms is highly influenced by prevailing conditions in their habitat, which are shaped by various factors, such as temperature, diet, feeding behavior, life stage, or physiological state32,33. Total lipid content in the arm muscle of common octopus was not influenced by environmental factors across the studied locations, suggesting that lipid levels in arm muscle remain constant under different growth conditions. The relative abundance of phospholipids in total lipid extracts (~50%) from RAr and SL aligns with previous findings reported by Prato et al.34 for wild-sourced O. vulgaris. In contrast, samples from Pe showed the lowest phospholipid content, resembling levels reported for specimens cultured using a mixed diet of crab, fish, and mussel by Prato et al.34. These results suggest that individuals from Pe likely underwent dietary-driven shifts. Phospholipids, as essential components of cell membranes, play a fundamental role in maintaining membrane integrity and function35. These polar lipids contribute to membrane permeability, facilitating the exchange of nutrients and ions, and regulate metabolism, physiology, and energy production36,37. Environmental changes in marine ecosystems influence polar lipid composition, with shifts in phospholipid profiles serving as biochemical markers of an organism’s physiological state35. Stressors, such as salinity fluctuations38 and dietary changes34, have been linked to variations in phospholipid content.
The polar lipidome of O. vulgaris was recently characterized in detail through a comprehensive study by Gaspar et al.39. The present study revealed consistent findings across the three sampling locations. The lipidome of O. vulgaris presented a high abundance of ether-linked lipids, especially in PC, PE, LPE, and PS. Depending on the groups at sn-1 of the glycerol backbone, the ether-linked phospholipids can be divided into two subclasses: 1-O-alkyl-2-acyl (plasmanyl, identified with O in the nomenclature), and 1-O-(alk-1′-enyl)-2-acyl (plasmenyl or plasmalogen, identified with P in the nomenclature). Specifically, plasmalogens contribute to cellular membrane structure-mediated functions and are key biomolecules in the response to oxidative stress, considered important endogenous anti-oxidant molecules, that provide protection to membrane lipids against oxidation40,41,42. These lipids enhance cellular adaptability by preserving membrane integrity under stress conditions43. Ether-linked phospholipids exhibited the greatest variation among sampling areas, likely reflecting metabolic adjustments in O. vulgaris to different environmental conditions or diets.
In marine organisms, the LPC/PC and LPE/PE ratios highlight the dynamic interplay between lipid metabolism and external factors. These ratios have significant biological implications, as they reflect lipid metabolism, oxidative stress, environmental adaptation, and seafood quality control47. Elevated LPC/PC ratios, as it was observed in SL samples, indicate PC hydrolysis, often due to enzymatic activity (e.g., phospholipase A2) or oxidative stress, affecting membrane stability and fluidity47,48. Additionally, elevated LPE/PE ratios reflect PE breakdown, which compromises membrane structure and function. This is particularly evident under stress conditions like oxidative damage or improper tissue storage47,48. The elevated LPC/PC and LPE/PE ratios in SL samples suggest that octopus from this landing site likely experienced stress conditions, such as food shortage, temperature fluctuations, habitat changes, or even improper storage after death.
The main phospholipid classes, namely PC and PE, showed DHA and EPA in their most abundant lipid species. These essential polyunsaturated fatty acids (PUFA) were predominant in both wild and reared octopuses34,44. Octopus vulgaris accumulates DHA and EPA primarily through its diet of crustaceans, molluscs, and fish. However, its predatory nature results in higher DHA content compared to EPA45. The high concentration of these fatty acids in O. vulgaris tissues plays a crucial role in nervous system development and function46.
The lipid species that contributed the most for the discrimination recorded between sampling sites were mainly ether-linked phospholipids (i.e., PC and PE), suggesting an adaptation of cell membranes to the different sampling environments. The high content of ether-linked PC in RAr, when compared to other locations, can be related to the higher availability of bivalves, as these are farmed using aquaculture systems in this area. Marine invertebrates contain a high diversity of ether-linked phospholipids in their lipidomes42,49, which are especially abundant in molluscs50,51. Plasmalogens were estimated to represent 33%, 54% and 63% of total PC, PE, and PS in Mytilus edulis, respectively51. Different capture locations often showcase contrasting oceanographic dynamics and available preys, which can ultimately shape the lipidome of organisms that inhabit them. The Galician coast, located in the northwest part of Spain, is characterized by the Rías (drowned river valleys), which create a unique coastal landscape and provide rich habitats for marine life and favor aquaculture practices targeting the grow-out of bivalves. For decades, the common octopus has been harvested along the Galician coastline, with ports in the Rías Baixas, such as RAr, traditionally experiencing high landings5. These environments are highly influenced by upwelling events, ranking RAr as a highly productive marine ecosystem52,53. It is worth highlighting that RAr is the largest ria in the coast of Galicia and features an impressive aquaculture production of Mediterranean mussel Mytilus galloprovincialis, accounting for nearly 70% of all mussel rafts in Galicia54. Mussel aquaculture significantly influences the diet of common octopus in these locations, as mussel production rafts can attract possible octopus’ prey55, therefore enhancing feeding opportunities. The influence of raft culture in the diet of other invertebrates (e.g., the harbor crab Liocarcinus depurator) has already been reported56. Therefore, samples from RAr presenting a higher abundance of ether-linked PC (plasmanyl and plasmenyl) may be associated with the octopus’ diet in this area.
The CAEP molecular species that contributed to discriminate between landing areas were particularly abundant in samples originating from RAr. This sphingolipid class is not very common in marine organisms, although it has been identified in some mollusc, echinoderm, and cnidarian species57. The most discriminating CAEP lipid species included the sphingoid bases 16:1;O2 and 19:3;O2 in their composition, which are the most frequently found in marine invertebrates58. CAEP are very abundant in different molluscs, such as the squid Uroteuthis chinensis58 and the Mediterranean mussel M. galloprovincialis59. These molecules act as protective cell membrane agents due to their endurance against hydrolytic enzymes, thus enabling cellular adaptation to environmental changes60. CAEP and sterols have been suggested as relevant molecules to form lipid rafts in the cell membranes of marine invertebrates49, contributing to the regulation of their fluidity. In nudibranch molluscs, PI and CAEP have been identified as a conservative part of their polar lipidome, without experiencing any major influence by diet61. As such, the differences observed in the CAEP composition between samples originating from different geographic areas may most likely be related to environmental changes that have promoted pronounced shifts in the cell membranes of O. vulgaris.
In conclusion, the present study revealed the polar lipidome profile of O. vulgaris and its compositional variations across three landing sites along the Iberian Atlantic coast. While total lipid content in O. vulgaris arm muscle remained consistent across locations, phospholipid levels were lower in specimens landed in Pe, most likely due to environmental or dietary differences. Additionally, octopus from SL exhibited the highest LPE/PE and LPC/PC ratios, suggesting an exposure to stressors or inadequate storage conditions. The dominant polar lipid species were rich in essential fatty acids, such as EPA and DHA, highlighting their potential as valuable sources of these PUFA. This study represents a first step in evaluating lipidomics as a reliable tool for the discrimination of the geographic origin of O. vulgaris, as the composition of its lipidome reflects environmental conditions and dietary resources in different fishing grounds. Key membrane lipid classes in marine invertebrates, including PC, PE, CAEP, and ether-linked phospholipids, significantly contributed to the differentiation among sampling locations. Although the findings of this study highlight the potential of lipidomic profiling of O. vulgaris arm muscle to indicate geographic origin, further validation is needed to evaluate its effectiveness, compliance with traceability standards, and feasibility for routine application along the seafood supply chain. Expanding the analysis to include specimens from the Mediterranean and Moroccan/Mauritanian coasts, along with seasonal and long-term sampling, would provide deeper insights into how environmental and trophic factors influence the lipidome and biochemical phenotypes of O. vulgaris.
Methods
Sampling
Samples of O. vulgaris were sourced from fisher’s landing operating on three different areas along the Atlantic coast of Galicia (Spain) [Ria de Arousa (RAr, Porto de Ribeira)] and Portugal [Peniche (Pe) and Santa Luzia (SL)] (Fig. 8), in the summer of 2018. A total of 3 locations and 10 individuals of common octopus (n = 10) per location were sampled (3 × 10 = 30). Sampled specimens were transported in coolers to the laboratory immediately after being landed in the different fishing harbors. The fourth right arm counted from the sagittal plane, front to back, was cut and stored at –80 °C. Samples were subsequently freeze-dried (CoolSafe 55–9 L Pro, Labogene, Lillerød, Denmark) and stored at –80 °C for further analysis.
Sampling areas of Octopus vulgaris specimens along the Atlantic coast of the Iberian Peninsula: Ria de Arousa, Peniche, and Santa Luzia.
All octopuses were collected, slaughtered, and maintained refrigerated by duly licensed professional fishers while still onboard, being already dead when landed in the fishing harbor; as such, ethical issues concerning animal experimentation and welfare do not apply to the present study.
Lipid extraction
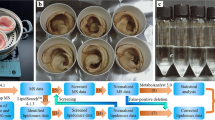
Freeze-dried samples of O. vulgaris arm muscle were individually homogenized using a mortar grinder (RM 200, Retsch, Hann, Germany). Between the homogenization of samples, the mortar grinder was carefully cleaned with silicate followed by alcohol (70%) to avoid cross-contamination. A modified Bligh and Dyer62 method was used to extract total lipids39. An amount of 50 mg of O. vulgaris arm muscle was mixed with 2.5 mL of methanol (MeOH) and 1.25 mL of dichloromethane (CH2Cl2) in a glass centrifuge tube. After homogenization and sonication for 1 min, using an automatic ultrasonic frequency of 35 kHz (Bandelin, Sonorex, RK 100, Berlin, Germany), the sample was incubated on ice on a rocking platform shaker for 30 min. The tube was then centrifuged at 568 × g for 10 min at room temperature (UNIVERSAL 320 R, Hettich, Tuttlingen, Germany). The organic phase containing the lipids was collected in a new tube and the biomass residue was re-extracted. A volume of 1.25 mL of CH2Cl2 and 1.25 mL of Mili Q water was added to the organic phases containing the lipid. After homogenization and centrifugation for 10 min at 568 × g to promote phase separation, the organic phases containing the lipids were collected in a new tube and the aqueous phase of the first extraction was re-extracted with 1.88 mL of CH2Cl2 followed by homogenization and centrifugation for 10 min 568 × g to recover the organic phase. The total lipid extract was dried using a speed vacuum (UNIVAPO-100H coupled with UNIJET II refrigerated aspirator) and transferred to previously dried and weighed dark vials. The lipid extracts were stored at –20 °C. The total lipid content was estimated by gravimetry and used to determine the lipid content per dry weight of biomass (µg mg−1 DW).
Phospholipid quantification
The content of phospholipids in total lipid extracts was performed through the phosphorus assay63. Total lipid extract was resuspended in 300 µL of CH2Cl2 and a volume of 10 µL of lipid extract was transferred to a glass tube and dried under a nitrogen stream. Then, 125 µL of 70% perchloric acid was added followed by incubation at 180 °C for 60 min in a heating block. After cooling at room temperature, 825 µL of MiliQ water, 125 µL of 2.5% aqueous solution of ammonium molybdate ((NH4)6 Mo7O24·4H2O) and 125 µL of 10% ascorbic acid were added to the glass tubes. The standards were prepared using 0.1–2 µg of phosphate (standard solution of NaH2PO4·2H2O, 100 µg phosphorus mL−1) to prepare a calibration curve and followed the same treatment as the samples. Samples and standards were incubated at 100 °C in a water bath (Precisterm, JP Selecta) for 10 min, followed by cooling in cold water. The absorbance of samples and standards was measured at 797 nm using a microplate ultraviolet-visible spectrophotometer (Multiskan GO, Thermo Scientific).
The amount of phospholipids was estimated by multiplying the amount of phosphorus in each sample by 25, the conversion factor between phosphorus and phospholipids64.
Hydrophilic interaction liquid chromatography-mass spectrometry
Lipid extracts were analyzed by hydrophilic interaction liquid chromatography-mass spectrometry (HILIC-LC-MS) on an Ultimate 3000 Dionex ultra-high-performance liquid chromatography (UHPLC) system (Thermo Fisher Scientific) with an autosampler coupled online to a Q-Exactive mass spectrometer with Orbitrap ® technology (Thermo Fisher Scientific). A two-mobile-phase solvent system was used to perform the analysis: a mobile phase A consisting of a mix of water, acetonitrile, and methanol (25/50/25, by volume) and a mobile phase B consisting of acetonitrile and methanol (60/40, by volume), with both phases presenting 5 mM ammonium acetate. To perform the LC-MS analysis an amount of lipid extract corresponding to 5 µg of phospholipids was dissolved in CH2Cl2 (1 µg μL−1) and mix with 75 µL of the starting eluent (95/5, B/A, by volume) and 4 µL of each lipid standard [ceramide (Cer d18:1_17:0), 0.04 µg; 1,2-dimyristoyl-sn-glycero-3-phosphate(dMPA), 0.08 µg; 1,2-dimyristoyl-sn-glycero-3-phospho-(10-rac-)glycerol (dMPG), 0.012 µg; lysophosphatidylcholine (LPC), 0.02 µg; 1,2-dimyristoylsnglycero-3-phosphoethanolamine (dMPE), 0.02 µg) (Avanti Polar Lipids, Inc., Alabaster, AL). The samples were injected into an ACE HILIC-N microbore column (100 × 1.0 × 3 µm) at a flow rate of 50 µL min−1 and a temperature of 35 °C. Initially, 5% of mobile phase A was held isocratically for 2 min, followed by a linear increase to 70% of mobile phase A within 11 min and a new linear increase to 90% within 7 min, maintained for 30 min, and returning to the initial conditions in 5 min, being held for an additional 5 min.
The Orbitrap mass spectrometer was operated using positive/negative switching toggles between positive (electrospray voltage 3.0 kV) and negative (electrospray voltage −2.7 kV) ion modes with a capillary temperature of 250 °C and a sheath gas flow of 15 U. In MS experiments, a high resolution of 70,000 was used, as well as an automatic gain control (AGC) target of 1 × 106. In tandem mass spectrometry (MS/MS), a resolution of 17,500 and AGC target of 1 × 105 were used and cycles consisted of one full-scan mass spectrum and 10 data-dependent MS/MS scans were repeated continuously throughout the analysis with the dynamic exclusion of 60 s and an intensity threshold of 2 × 104. Normalized collision energy TM (CE) ranged between 25, 30, and 35 eV. Data acquisition was performed using the Xcalibur data system (V3.3, Thermo Fisher Scientific, USA). The molecular identification of lipid species was performed by interpretation of the HILIC–ESI–MS/MS spectra, through the typical fragmentation patterns of the polar head group and the fatty acyl chains inherent in the structure of each molecular species39, LC-MS expected retention time, and mass accuracy (Qual Browser) with an error of ≤5 ppm.
Data analysis
Raw data processing was performed using the MZmine 2.53 software65. The MS raw data were pre-processed by filtering and smoothing. Peak assignment and ion identification were based on mass accuracy and performed against in-house polar lipid databases. The mass list was filtered, then peaks were detected and processed. The parameters set for MZmine 2.53 were the following: minimum peak height, above 1 × 104; join alignment; allowable error of retention time, 0.5 min; acceptable error of m/z (m/z tolerance) 5 ppm. Integrated peak area values from lipid species were exported using the comma-separated values (.csv) format. Integrated peak areas of each lipid species were exported and data were normalized by dividing peak areas from extracted ion chromatograms (XIC) for each lipid species by the peak area of selected internal standard for lipid classes as follows: PG, PI with dMPG; CAEP, N-methyl-CAEP, PC, PE, PE-Cer, and PS with dMPE; Cer and HexCer with Cer d18:1_17:0; LPC, LPE, and SM with LPC.
Statistical analysis
One-way ANOVA was performed to assess significant differences in the content of total lipids (mg g−1 DW), phospholipids (% of total lipids), relative abundance of lipid species within the corresponding lipid class, and normalized XIC areas of identified lipid species between landing sites. Shapiro–Wilks and Bartlett’s tests were performed to evaluate ANOVA assumptions of normality and homogeneity of variance, respectively.
Normalized XIC areas of polar lipids were processed using Metaboanalyst 6.066. Missing values were replaced by 1/5 of the minimum positive value of each variable. Data filtering was performed to remove variables that evidenced a low repeatability by relative standard deviation (RSD = SD/mean), followed by log transformation. Principal component analysis (PCA) was performed to visualize the general 2D clustering of O. vulgaris samples from three different locations. A hierarchical clustering dendrogram and a heatmap were performed using Euclidean distances and the Ward clustering algorithm. Heatmap provided a better understanding of the significance of the variables in separating different landing sites. The top 50 molecular species were ranked using p-values from an ANOVA test.
Data Availability
The datasets generated during the current study are available from the corresponding author (felisa.rey@ua.pt) on reasonable request.
Abbreviations
- CAEP:
-
Ceramide aminoethylphosphonate
- Cer:
-
Ceramide
- CH2Cl2 :
-
Dichloromethane
- DHA:
-
Docosahexaenoic acid
- dMPA:
-
1,2-dimyristoyl-sn-glycero-3-phosphate
- dMPE:
-
1,2-dimyristoylsnglycero-3-phosphoethanolamine
- dMPG:
-
1,2-dimyristoyl-sn-glycero-3-phospho-(10-rac-glycerol)
- EPA:
-
Eicosapentaenoic acid
- HexCer:
-
Hexosylceramide
- HILIC:
-
Hydrophilic interaction liquid chromatography
- LPC:
-
Lyso-phosphatidylcholine
- LPE:
-
Lyso-phosphatidylethanolamine
- MeOH:
-
Methanol
- MS:
-
Mass spectrometry
- MS/MS:
-
Tandem mass spectrometry
- N-methyl-CAEP:
-
N-methyl ceramide aminoethylphosphonate
- PC:
-
Phosphatidylcholine PCA Principal component analysis
- PE:
-
Phosphatidylethanolamine
- Pe:
-
Peniche
- PE-Cer:
-
Ceramide phosphoethanolamine
- PG:
-
Phosphatidylglycerol
- PI:
-
Phosphatidylinositol
- PS:
-
Phosphatidylserine
- PUFA:
-
Polyunsaturated fatty acids
- RAr:
-
Ria de Arousa
- SL:
-
Santa Luzia
- SM:
-
Sphingomyelin
- UHPLC:
-
Ultra-high-performance liquid chromatography
- XIC:
-
Extracted ion chromatograms
References
Pita, C. et al. Fisheries for common octopus in Europe: socioeconomic importance and management. Fish. Res. 235, 105820 (2021).
WoRMS Editorial Board. World Register of Marine Species. https://www.marinespecies.org at VLIZ (2024).
Pascual-Fernández, J. J., Florido-del-Corral, D., De La Cruz-Modino, R. & Villasante, S. Small-scale fisheries in Spain: diversity and challenges. In Small-Scale Fisheries in Europe: Status, Resilience and Governance Vol. 23 (eds. Pascual-Fernández, J. J., Pita, C. & Bavinck, M.) 253–281 (Springer International Publishing, Cham, 2020).
Almeida, C., Loubet, P., Laso, J., Nunes, M. L. & Marques, A. Environmental assessment of common octopus (Octopus vulgaris) from a small-scale fishery in Algarve (Portugal). Int J. Life Cycle Assess. 27, 849–867 (2022).
Bañón, R., Otero, J., Campelos-Álvarez, J. M., Garazo, A. & Alonso-Fernández, A. The traditional small-scale octopus trap fishery off the Galician coast (Northeastern Atlantic): historical notes and current fishery dynamics. Fish. Res. 206, 115–128 (2018).
Sonderblohm, C. P. et al. Participatory assessment of management measures for Octopus vulgaris pot and trap fishery from southern Portugal. Mar. Policy 75, 133–142 (2017).
Roumbedakis, K., Pita, C. & Pierce, G. J. Market opportunities for octopus in the Atlantic area: CEPHS & CHEFS—Summary of main achievements. Preprint at, https://doi.org/10.48528/B55A-1D44 (2021).
Certification & Ratings Collaboration. Certification and Ratings Collaboration. https://certificationandratings.org (2025).
MSC. Marine Stewardship Council (MSC). https://www.msc.org/for-business/fisheries (2025).
Vaz-Pires, P., Seixas, P. & Barbosa, A. Aquaculture potential of the common octopus (Octopus vulgaris Cuvier, 1797): a review. Aquaculture 238, 221–238 (2004).
Ospina-Alvarez, A. et al. A network analysis of global cephalopod trade. Sci. Rep. 12, 322 (2022).
Ambrose, R. F. & Nelson, B. V. Predation by Octopus vulgaris in the Mediterranean. Mar. Ecol. 4, 251–261 (1983).
Rosa, R., Marques, A. M., Nunes, M. L., Bandarra, N. & Sousa Reis, C. Spatial-temporal changes in dimethyl acetal (octadecanal) levels of Octopus vulgaris (Mollusca, Cephalopoda): relation to feeding ecology. Sci. Mar. 68, 227–236 (2004).
Smith, C. D. Diet of Octopus vulgaris in False Bay, South Africa. Mar. Biol. 143, 1127–1133 (2003).
Rodríguez, C., Carrasco, J. F., Arronte, J. C. & Rodríguez, M. Common octopus (Octopus vulgaris Cuvier, 1797) juvenile ongrowing in floating cages. Aquaculture 254, 293–300 (2006).
Asche, F., Bellemare, M. F., Roheim, C., Smith, M. D. & Tveteras, S. Fair enough? Food security and the international trade of seafood. World Dev. 67, 151–160 (2015).
D’Amico, P., Armani, A., Gianfaldoni, D. & Guidi, A. New provisions for the labelling of fishery and aquaculture products: difficulties in the implementation of Regulation (EU) n. 1379/2013. Mar. Policy 71, 147–156 (2016).
El Sheikha, A. F. & Montet, D. How to determine the geographical origin of seafood? Crit. Rev. Food Sci. Nutr. 56, 306–317 (2016).
Mamede, R. et al. Fatty acids profiling of goose barnacle (Pollicipes pollicipes) tissues to evaluate nutritional quality and confirm harvesting location. J. Food Comp. Anal. 127, 105930 (2024).
Ricardo, F. et al. Assessing the elemental fingerprints of cockle shells (Cerastoderma edule) to confirm their geographic origin from regional to international spatial scales. Sci. Total Environ. 814, 152304 (2022).
Leal, M. C., Pimentel, T., Ricardo, F., Rosa, R. & Calado, R. Seafood traceability: current needs, available tools, and biotechnological challenges for origin certification. Trends Biotechnol. 33, 331–336 (2015).
Turchini, G. M., Quinn, G. P., Jones, P. L., Palmeri, G. & Gooley, G. Traceability and discrimination among differently farmed fish: a case study on Australian murray cod. J. Agric. Food Chem. 57, 274–281 (2009).
Parrish, C. C. Lipids in marine ecosystems. ISRN Oceanogr. 2013, 1–16 (2013).
Haridas, P. C. et al. Authentication of the species identity of squid rings using UHPLC-Q-Orbitrap MS/MS-based lipidome fingerprinting and chemoinformatics. Food Chem. 442, 138525 (2024).
Lopes, D. et al. Insights of species-specific polar lipidome signatures of seaweeds fostering their valorization in the blue bioeconomy. Algal Res. 55, 102242 (2021).
da Costa, E. et al. Site-specific lipidomic signatures of sea lettuce (Ulva spp., chlorophyta) hold the potential to trace their geographic origin. Biomolecules 10, 489 (2020).
Monteiro, J. P. et al. The unique lipidomic signatures of Saccharina latissima can be used to pinpoint their geographic origin. Biomolecules 10, 107 (2020).
Rey, F. et al. Domesticated populations of Codium tomentosum display lipid extracts with lower seasonal shifts than conspecifics from the wild — Relevance for biotechnological applications of this green seaweed. Mar. Drugs 18, 188–188 (2020).
Ricardo, F., Maciel, E., Domingues, M. R. & Calado, R. Spatio-temporal variability in the fatty acid profile of the adductor muscle of the common cockle Cerastoderma edule and its relevance for tracing geographic origin. Food Control 81, 173–180 (2017).
Arechavala-Lopez, P. et al. Fatty acids and elemental composition as biomarkers of Octopus vulgaris populations: does origin matter? Mar. Pollut. Bull. 139, 299–310 (2019).
Ricardo, F., Veríssimo, A., Maciel, E., Domingues, M. & Calado, R. Fatty acid profiling as a tool for fostering the traceability of the halophyte plant Salicornia ramosissima and contributing to its nutritional valorization. Plants 13, 545 (2024).
Gladyshev, M. I., Sushchik, N. N., Tolomeev, A. P. & Dgebuadze, Y. Y. Meta-analysis of factors associated with omega-3 fatty acid contents of wild fish. Rev. Fish. Biol. Fish. 28, 277–299 (2018).
Parrish, C. C. Production, transport, fate and effects of lipids in the marine environment. Mar. Drugs 23, 52 (2025).
Prato, E., Portacci, G. & Biandolino, F. Effect of diet on growth performance, feed efficiency and nutritional composition of Octopus vulgaris. Aquaculture 309, 203–211 (2010).
Fokina, N. N., Ruokolainen, T. R. & Nemova, N. N. Lipid composition modifications in the blue mussels (Mytilus edulis L.) from the White Sea. In Organismal and Molecular Malacology (ed. Ray, S.) (InTech, 2017).
Tocher, D. R., Bendiksen, E. Å, Campbell, P. J. & Bell, J. G. The role of phospholipids in nutrition and metabolism of teleost fish. Aquaculture 280, 21–34 (2008).
Navarro, J. C. & Villanueva, R. Lipid and fatty acid composition of early stages of cephalopods: an approach to their lipid requirements. Aquaculture 183, 161–177 (2000).
Nemova, N. N., Fokina, N. N., Nefedova, Z. A., Ruokolainen, T. R. & Bakhmet, I. N. Modifications of gill lipid composition in littoral and cultured blue mussels Mytilus edulis L. under the influence of ambient salinity. Polar Rec. 49, 272–277 (2013).
Gaspar, L. et al. Lipidomics of common octopus’ (Octopus vulgaris) arm muscle using untargeted high-resolution liquid chromatography-mass spectrometry. J. Food Comp. Anal. 115, 104871 (2023).
Braverman, N. E. & Moser, A. B. Functions of plasmalogen lipids in health and disease. Biochim. et Biophys. Acta Mol. Basis Dis. 1822, 1442–1452 (2012).
Broniec, A. et al. Interactions of plasmalogens and their diacyl analogs with singlet oxygen in selected model systems. Free Radic. Biol. Med. 50, 892–898 (2011).
Yamashita, S., Miyazawa, T., Higuchi, O., Kinoshita, M. & Miyazawa, T. Marine plasmalogens: a gift from the sea with benefits for age-associated diseases. Molecules 28, 6328 (2023).
Rangholia, N., Leisner, T. M. & Holly, S. P. Bioactive ether lipids: primordial modulators of cellular signaling. Metabolites 11, 41 (2021).
Ben-Youssef, S., Selmi, S., Ezzeddine-Najai, S. & Sadok, S. Total lipids and fatty acids composition of the coastal and the deep-sea common octopus (Octopus vulgaris) Populations: a comparative study. Nutr. Health 19, 195–201 (2008).
Lourenço, S., Roura, Á, Fernández-Reiriz, M.-J., Narciso, L. & González, ÁF. Feeding Relationship between Octopus vulgaris (Cuvier, 1797) early life-cycle stages and their prey in the western Iberian upwelling system: correlation of reciprocal lipid and fatty acid contents. Front. Physiol. 8, 467 (2017).
Reis, D. B. et al. An insight on Octopus vulgaris paralarvae lipid requirements under rearing conditions. Aquac. Nutr. 21, 797–806 (2015).
Losito, I. et al. Tracing the thermal history of seafood products through lysophospholipid analysis by hydrophilic interaction liquid chromatography–electrospray ionization fourier transform mass spectrometry. Molecules 23, 2212 (2018).
Krautbauer, S. et al. Application of lipid class ratios for sample stability monitoring—evaluation of murine tissue homogenates and SDS as a stabilizer. Metabolites 11, 277 (2021).
Imbs, A. B., Ermolenko, E. V., Grigorchuk, V. P., Sikorskaya, T. V. & Velansky, P. V. Current progress in lipidomics of marine invertebrates. Mar. Drugs 19, 660 (2021).
Facchini, L., Losito, I., Cataldi, T. R. I. & Palmisano, F. Seasonal variations in the profile of main phospholipids in Mytilus galloprovincialis mussels: a study by hydrophilic interaction liquid chromatography-electrospray ionization Fourier transform mass spectrometry. J. Mass Spectrom. 53, 1–20 (2018).
Wang, J. et al. Quantitative and comparative study of plasmalogen molecular species in six edible shellfishes by hydrophilic interaction chromatography mass spectrometry. Food Chem. 334, 127558 (2021).
Outeiro, L., Byron, C. & Angelini, R. Ecosystem maturity as a proxy of mussel aquaculture carrying capacity in Ria de Arousa (NW Spain): a food web modeling perspective. Aquaculture 496, 270–284 (2018).
Tenore, K. R. et al. Coastal upwelling in the Rías Bajas, NW Spain: contrasting the benthic regimes of Rías de Arosa and de Muros. J. Mar. Res. 40, 701–772 (1982).
Mexillon de Galicia. https://www.mexillondegalicia.org/?page_id=22 (2024).
Romero, P., González-Gurriaránn, E. & Penas, E. Influence of mussel rafts on spatial and seasonal abundance of crabs in the Ría de Arousa, North-West Spain. Mar. Biol. 72, 201–210 (1982).
Freire, J. Feeding ecology of Liocarcinus depurator (Decapoda: Portunidae) in the Ria de Arousa (Galicia, north-west Spain): effects of habitat, season and life history. Mar. Biol. 126, 297–311 (1996).
Rey, F. et al. Applications of lipidomics in marine organisms: progress, challenges and future perspectives. Mol. Omics 357–386 https://doi.org/10.1039/D2MO00012A (2022).
Wang, R. et al. Identification of ceramide 2-aminoethylphosphonate molecular species from different aquatic products by NPLC/Q-Exactive-MS. Food Chem. 304, 125425 (2020).
Kariotoglou, D. M. & Mastronicolis, S. K. Phosphonolipids in the mussel Mytilus galloprovincialis. Z. für Naturforsch. C. 53, 888–896 (1998).
Facchini, L., Losito, I., Cataldi, T. R. I. & Palmisano, F. Ceramide lipids in alive and thermally stressed mussels: an investigation by hydrophilic interaction liquid chromatography-electrospray ionization Fourier transform mass spectrometry: Ceramide lipids in mussels: a HILIC-ESI-FTMS study. J. Mass Spectrom. 51, 768–781 (2016).
Imbs, A. B. & Grigorchuk, V. P. Lipidomic study of the influence of dietary fatty acids on structural lipids of cold-water nudibranch molluscs. Sci. Rep. 9, 20013 (2019).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Bartlett, E. M. & Lewis, D. H. Spectrophotometric determination of phosphate esters in the presence and absence of orthophosphate. Anal. Biochem. 36, 159–167 (1970).
Chapman, G. W. A conversion factor to determine phospholipid content in soybean and sunflower crude oils. J. Am. Oil Chem. Soc. 57, 299–302 (1980).
Pluskal, T., Castillo, S., Villar-Briones, A. & Orešič, M. MZmine 2: modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform. 11, 395 (2010).
Chong, J. et al. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 46, W486–W494 (2018).
Acknowledgements
The authors acknowledge the University of Aveiro, Fundação para a Ciência e a Tecnologia I.P. (FCT, Portugal), and Ministério da Ciência e Tecnologia (MCT). This work was funded by national funds through FCT, under the project/grant UID/50006 + LA/P/0094/2020 (doi.org/10.54499/LA/P/0094/2020) —Centro de Estudos do Ambiente e Mar (CESAM), and UID/50006—Laboratório Associado para a Química Verde—Tecnologias e Processos Limpos. This work was also funded by national funds through FCT in the scope of the Individual Call to Scientific Employment Stimulus 2017 with a Junior Researcher contract to Felisa Rey (reference CEECIND/00580/2017, (https://doi.org/10.54499/CEECIND/00580/2017/CP1459/CT0005) and Cristina Pita (reference 2020.02510.CEECIND/CP1589/CT0018, https://doi.org/10.54499/2020.02510.CEECIND/CP1589/CT0018). This study was supported by research project CITAQUA “Desenvolvimento do Projeto de Reforço do Polo de Aveiro (H4)”, framed within Measure 10 of Investment TC-C10-i01—Hub Azul—Rede de Infraestruturas para a Economia Azul, financed by the Recovery and Resilience Plan (PRR) and supported by Fundo Azul of the Portuguese Government” and the Cephs and Chefs Project (https://www.cephsandchefs.com/) funded by the European Regional Development Fund through the Interreg Atlantic Area Programme grant number EAPA_282/2016. We also thank Renato Mamede and Seila Díaz for their technical support. This is a contribution of the Marine Lipidomics Laboratory at the University of Aveiro.
Author information
Authors and Affiliations
Contributions
Conceptualization: F. Rey, F. Ricardo, M.R. Domingues, R. Calado; Sampling: F. Ricardo; Experimental work: L. Gaspar, F. Rey; Data curation: L. Gaspar, F. Rey, M.R. Domingues; Formal analysis: L. Gaspar, F. Rey; Statistical analysis: F. Rey; Writing— original draft: L. Gaspar, F. Rey; Funding acquisition: M.R. Domingues, R. Calado, C. Pita; Writing—review and editing: F. Rey. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rey, F., Gaspar, L., Ricardo, F. et al. Lipidomic signatures in Octopus vulgaris arm muscle reveal geographic variation along the Iberian Atlantic Coast. npj Sci Food 9, 173 (2025). https://doi.org/10.1038/s41538-025-00520-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-025-00520-w