Abstract
Staphylococcus aureus can contaminate food products and persist in food-related environments, posing significant challenges to food safety and public health. However, the role of plasmids in mediating antimicrobial resistance (AMR) and facilitating host adaptation in S. aureus has been largely underestimated. We conducted a global plasmidome analysis of 1395 isolates (human: 88.2%, animal: 11.8%) spanning 90 years. Plasmids were detected in 66.8% of strains, typically one to two per genome. We identified 35 distinct plasmid-borne antimicrobial resistance genes (ARGs), with a significant temporal increase in their abundance. Over 85.5% of these plasmids were predicted to be mobilizable, indicating a high transmission potential. Notably, clonal complex 5 (CC5) carried the highest plasmid burden, particularly subclade CC5.6, which exhibited high prevalence of conjugative pSK41 plasmids and ARGs. Although this lineage has not been reported elsewhere, its emergence raises concerns about ARG dissemination through conjugative plasmids. These findings emphasize the role of plasmids in the global spread of AMR and have important implications for food safety and resistance control strategies in food production.
Similar content being viewed by others
Introduction
S. aureus is a highly adaptable Gram-positive bacterium responsible for a broad range of infections in both humans and animals. It can contaminate food products and thrive in various food-related environments, such as food waste and food processing settings, making it a major concern for public health and foodborne illness prevention1. In the United States, approximately 241,000 cases of foodborne illness are attributed to S. aureus each year2. The widespread misuse of antimicrobials has led to the emergence and spread of multidrug-resistant (MDR) S. aureus strains, posing a significant challenge to both the treatment of complicated infections and the prevention of their further dissemination3.
Plasmid-mediated dissemination of AMR is a major global public health concern. As key mobile genetic elements carrying ARGs, plasmids facilitate efficient horizontal gene transfer of ARGs across species and genera4. Although the role of plasmids has been extensively characterized in Gram-negative bacteria such as those in the Enterobacterales order, their contribution in S. aureus has remained largely overlooked5,6. Historically, S. aureus was thought to contain only a limited number of plasmids, with horizontal gene transfer believed to occur predominantly via transduction5. However, recent genomic analysis has challenged this assumption, revealing a broader and more diverse plasmid repertoire. Contarin et al. reported that plasmids were one of the primary vectors of ARGs in S. aureus7. Ramsay et al. further found that up to 74% of non-conjugative S. aureus plasmids could be mobilized by conjugative plasmids via a relaxase-in-trans mechanism8. These findings underscore the critical importance of investigating the plasmid biology of S. aureus.
Staphylococcal plasmids exhibit substantial genetic diversity and can be categorized according to their genetic architecture and functional characteristics. Based on conserved regions within their replication sequences, plasmids in S. aureus can be classified into six groups: Rep1, Rep2, Rep3, RepA_N, Rep_trans, and RepL9. With respect to their mobility mechanisms, these plasmids are further divided into conjugative, mobilizable, and non-mobilizable categories. To date, conjugative plasmids in S. aureus have been grouped into three distinct families: pSK41, pWBG749, and pWBG48. Mobilizable plasmids either carry oriT sequence mimics derived from pSK4110 or pWBG74911 or encode a distinct relaxase. Both conjugative and mobilizable plasmids play critical roles in the horizontal transfer of ARGs.
CC5 is recognized as a major lineage of hospital-acquired methicillin-resistant S. aureus (HA-MRSA) and is widely prevalent in healthcare settings globally12. CC5 strains typically exhibit a MDR phenotype, displaying resistance to β-lactams, aminoglycosides, and macrolides13, and carry a range of virulence genes, including fnbB, tst, and sec14. Notably, CC5 has been identified as a predominant genetic background in which S. aureus acquires vancomycin resistance through the uptake of the vanA operon from Enterococcus species15. Accelerating resistance evolution has been observed in CC5 strains, highlighting their adaptive potential and raising public health concerns13,16. In addition, CC5-MRSA demonstrates cross-species adaptability, capable of causing severe infections in humans and livestock17,18.
In this study, we systematically analyzed 1395 S. aureus strains from 51 countries, spanning from 1933 to 2023, with complete genomic sequences. Our analysis focused on (1) the distribution profiles of plasmids among these strains, (2) the plasmid size, replicon type, and mobility potential, (3) a comparative analysis of ARG diversity and abundance between chromosomal and plasmid DNA, (4) the observation that the predominant CC5 exhibited plasmid enrichment, and (5) the identification of subclade CC5.6 exhibiting a particularly high prevalence of conjugative pSK41-family plasmids and ARGs.
Results
Genomic characteristics and plasmid dynamics of a global S. aureus collection
Our assembled dataset included 1395 complete S. aureus genomes (as of July 1, 2023), consisting of 1230 (88.2%) human-derived and 165 (11.8%) animal-derived strains (Fig. 1A). These strains were collected between 1933 and 2023, with the majority obtained during the period from 2010 to 2023 (Supplementary Data 1). The dataset encompassed strains from six continents, with the United States (40.1%), Australia (14.1%), Germany (10.3%), and China (7.2%) contributing the most (Supplementary Data 1). We identified 11 clonal complexes (CCs) comprising 191 distinct sequence types (STs), with CC8 (30.5%) and CC5 (27.5%) being the predominant lineages (Fig. 1B). All 1395 S. aureus genomes exhibited ≥95% completeness and ≤3.5% contamination, ensuring a high-quality dataset for downstream analysis (Supplementary Data 1).
A Host sources of the 1395 S. aureus strains; B The number of isolates in each clonal complex (CC); C Plasmid-bearing vs. plasmid-free strain counts and the distribution plasmid counts per isolate; D Temporal distribution of isolate collection dates, separated by plasmid-bearing (blue) and plasmid-free (orange) groups; E Global geographic distribution of isolates. Each point represents a sampling location, with the size of the circle proportional to the number of isolates. Plasmid-bearing isolates are shown in blue and plasmid-free in orange.
We conducted a detailed analysis of the plasmid distribution characteristics in these 1395 S. aureus genomes. Our findings revealed that 472 (33.8%) of the strains were plasmid-free, while 923 (66.2%) carried at least one plasmid. Among the plasmid-bearing strains, 578 (62.6%) harbored a single plasmid, followed by 276 (29.9%) that carried two plasmids (Fig. 1C). Temporal analysis revealed a marked increase in plasmid prevalence. Prior to 2012, the proportion of plasmid-bearing strains was comparable to that of plasmid-free strains; however, after 2012, plasmid-carrying strains became predominant (Fig. 1D). Comparative analysis of the geographical distribution of plasmid-bearing and plasmid-free strains revealed distinct patterns. Among the four countries with the highest numbers of isolates, plasmid-bearing strains predominated in the United States and Australia, whereas plasmid-free strains were more common in Germany. In contrast, China exhibited a balanced distribution, with plasmid-bearing and plasmid-free strains occurring at comparable frequencies (Fig. 1E). However, our dataset may not fully capture the temporal increase in plasmid prevalence or the national situation due to uneven sampling. Additionally, it may be influenced by technological advancements and biases in sequencing.
Genetic and functional characterization of the S. aureus plasmidome
Plasmid analysis of 1395 S. aureus strains identified 1371 discrete plasmids (Supplementary Data 2), which were systematically characterized for size distribution, replicon typing, and mobility potential.
Size profiling revealed that 42.4% (582/1371) of the plasmids clustered in the 20–30 kb range, followed by smaller plasmids (≤10 kb), which represented 27.1% (372/1371) of the total (Fig. 2A).
A Size distribution of plasmids; B Frequencies of different plasmid replicon types; C Replicon type and size correlation; D Distribution of plasmid mobility types, the inset pie chart shows the distribution of three conjugative plasmid families; E Comparison of the number of ARGs per chromosome vs. plasmid; F ARG density per 10 kb genomic length carried by chromosome and plasmids; G Temporal trend of average ARGs per chromosome and plasmid over time.
Replicon prediction classified 85.4% (1171/1371) of plasmids into known replication systems, while the remaining plasmids did not match any known replicons. Among the plasmids with predicted known replicons, 948 were single-replicon plasmids, with RepA_N being the most prevalent (558/950, 58.7%), followed by Rep1 (180/950, 18.9%). Double- or multi-replicon configurations accounted for 18.9% (221/1171), predominantly involving RepA_N co-occurring with Rep1 (Fig. 2B).
Furthermore, we selected plasmids in which its replicon presented in more than 50 plasmids for further analysis of the relationship between replicon and plasmid size. The results showed distinct size preferences for different replicons. RepL, Rep_trans, and Rep1 replicons were predominantly associated with plasmids smaller than 5 kb, whereas larger plasmids in the 20–30 kb range were more commonly associated with RepA_N and Rep3 replicons (Fig. 2C).
Mobility prediction classified 85.5% (1173/1371) of the plasmids as mobilizable elements, including 10.1% (139/1371) conjugative plasmids and 75.4% (1034/1371) mobilizable plasmids that rely on conjugative plasmids for transfer. Three distinct conjugative plasmid families were identified, among which the pSK41 family predominated, accounting for 69.1% (96/139) of all conjugative plasmids (Fig. 2D).
Plasmid-borne ARGs show increasing prevalence
The diversity and abundance of ARGs were assessed independently for chromosomal and plasmid elements (Supplementary Data 3 and Supplementary Data 4). A total of 3636 ARGs (42 distinct types) were identified in chromosomal elements and 2,490 (35 types) on plasmids across S. aureus strains. Resistance determinants displayed differential genomic localization: chromosomal elements predominantly harbored mecA, ant(9)-Ia, dfrG, dfrK, erm33, ermA, and tetM, while plasmids carried a distinct set of ARGs. Specifically, we identified a total of 937 plasmids carrying at least one ARG, with a notable prevalence of blaZ, aph(3’)-IIIa, ermC, mphC, msrA, and mupA (Fig. S1). On average, chromosomal elements encoded 2.606 ARGs per genome, while plasmids carried 1.816 ARGs per plasmid (Fig. 2E). Given the differences in sequence length between chromosomes and plasmids, ARG density was normalized to the number of ARGs per 10 kb. The normalized values indicated that plasmids carried an average of 0.253 ARGs per 10 kb, which was significantly higher than the 0.009 observed in chromosomal elements (Fig. 2F). The temporal dynamics of ARGs associated with S. aureus plasmids and chromosomes from 1933 to 2023 revealed a markedly greater increase in plasmid-borne ARGs compared to chromosomal ones (k = 0.26), underscoring the escalating dissemination risk of plasmid-mediated resistance over the past 90-year period. (Fig. 2G).
Plasmids are enriched in the predominant CC5 clones
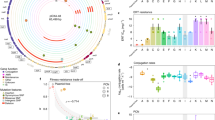
A comprehensive analysis of plasmid distribution across S. aureus clonal complexes revealed that CC5 strains harbored the highest average plasmid burden, with 1.19 plasmids per strain (Fig. 3A). A strong association was observed between the predominant replicon type, RepA_N, and CC5 lineages. Specifically, 44.6% (249/558) of RepA_N-carrying plasmids were identified in CC5 strains (Fig. 3B). Additionally, a substantial proportion of conjugative plasmids (59.0%, 82/139) originated from CC5 strains, a substantial higher representation than in other lineages (Fig. 3C). Four dominant categories of plasmid-borne ARGs were mainly identified in the CC5 lineage, including (i) aminoglycoside resistance genes aac(6’)-aph(2”) (80.6%), aadD (67.0%), ant(6)-Ia (98.8%), aph(2”)-Ia (80.6%), and aph(3’)-III (37.3%); (ii) the β-lactam resistance gene blaZ (40.7%); (iii) macrolide resistance genes msrA (36.9%), and mphC (36.5%); and (iv) the disinfectant resistance gene qacA (38.9%) (Fig. 3D), with percentages indicating the proportion of plasmid-carried ARGs associated with the CC5 lineage.
A Bar plot showing the proportion of isolates carrying 0-7 plasmids across various clonal complexes. The average number of plasmids per isolate is indicated above each bar; B Sankey diagram depicting associations between clonal complexes and plasmid replicon types; C Sankey diagram linking plasmid mobility types (conjugative, mobilizable, non-mobilizable) with clonal complexes; D Sankey diagram connecting plasmid-borne ARGs with clonal complexes.
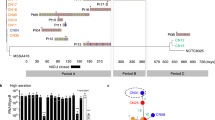
To further investigate conjugative plasmid distribution within the S. aureus CC5 lineage, a core genome-based maximum likelihood phylogenetic tree was constructed using all 384 S. aureus CC5 strains. Six distinct clades were identified within the CC5 phylogeny (Fig. 4). Clade CC5.6, primarily comprising strains from North America (Fig. S2), exhibited a significantly higher proportion of conjugative plasmids (76.4%, 55/72) compared to other subclades (p < 0.01; Fig. S3A). This clade also demonstrated the highest prevalence of plasmid-borne ARGs (Fig. S3B), with an average of 4.3 ARGs per isolate, notably higher than in other clades (p < 0.01). Additionally, a significant positive correlation was observed between genetic distance from the CC5 root and the number of plasmid-borne ARGs (Pearson’s R = 0.546, p < 0.001) in the CC5 core-genome tree (Figs. 4 and S4).
Circular phylogenetic tree of S. aureus CC5 lineage isolates annotated with plasmid features and ARGs. The inner tree branches are colored according to subclades (CC5.1-CC5.6). Four outer rings display: (1) Plasmid replicon types, including RepA, Rep1, Rep2, Rep3, RepL, and Rep_trans. (2) Plasmid mobility, categorized as non-mobilizable (blue), mobilizable (purple), or conjugative (green). (3) Number of plasmid-borne ARGs, ranging from 0 (white) to 16–20 (dark green). (4) Specific plasmid-borne ARGs, colored and listed by name (e.g., blaZ, mecA, aac(6’)-aph(2”), tetK, ermB, vanA, etc.).
The pSK41 conjugative plasmids are enriched in the subclade CC5.6 of CC5 lineage
We further investigated the types of conjugative plasmids present in CC5 strains and found that the pSK41 family was predominant, primarily distributed among CC5.6 isolates. All strains within this lineage were recovered from Homo sapiens hosts in the USA, with isolation dates ranging from 2015 to 2017. Phylogenetic analysis revealed that the pSK41 plasmid was absent in ancestral CC5 strains and was acquired multiple times throughout the evolutionary process, although it did not spread widely. The major acquisition event occurred around 2012 in the USA, followed by the expansion of a clade harboring the pSK41 plasmid (Fig. 5A).
A Time-scaled phylogenetic tree of bacterial isolates showing the presence (teal) or absence (black) of pSK41-family plasmids in subclade CC5.6. Each terminal node represents a single isolate, with color indicating whether a pSK41-family plasmid was identified; B Circular comparison of plasmid sequences against the reference plasmid pSK41. The innermost ring shows the coding sequences (CDSs) of reference pSK41 (NC_005024.1). GC skew is displayed in green (positive) and purple (negative) in the second rings. The outer rings represent alignments of other plasmids (labeled by GenBank accession numbers) with pSK41, with colored regions indicating conserved sequences. Blue gene labels (e.g., rep1, traO, bleO, aadD, mobV) denote the difference between the plasmids in clade CC5.6 and the reference.
The characterization of pSK41-family plasmids within the clade revealed conserved structural features (Fig. 5B). These plasmids varied in size from 41,662 to 52,910 bp and contained two clinically significant ARGs: aadD and mupA. They exhibited more than 99.70% nucleotide identity and over 83% coverage when compared to the reference plasmid pSK41 (NC_005024.1). In comparison to pSK41, these plasmids consistently lacked several key genes, including the replicon rep1, the transfer gene traO, the bleomycin resistance gene bleO, and the relaxase gene mobV.
Discussion
S. aureus is a significant zoonotic pathogen with the potential for cross-transmission among animals, food, and humans19. Although plasmids are recognized for their crucial role in the dissemination of ARGs in Gram-negative bacteria, their distribution, characteristics, and contribution to ARG transfer and bacterial adaptability, especially in S. aureus, remain relatively underexplored. In this study, we conducted a large-scale, global analysis of 1395 publicly available complete S. aureus genomes spanning 90 years. To the best of our knowledge, this is the first comprehensive characterization of the S. aureus plasmidome and the evolutionary dynamics of plasmid-borne resistome, with important implications for understanding the spread of AMR.
To minimize the impact of incomplete assemblies, only closed genomes were included in this study. However, as with all analysis based on publicly available data, our results are subject to certain biases. Owing to the growing accessibility of long-read sequencing technologies, the majority of available S. aureus genomes have been generated since 2000, leading to an underrepresentation of earlier isolates. As S. aureus is primarily human-associated, the dataset is dominated by human-derived strains, with only 165 isolates obtained from animals and none clearly documented as food-related (Fig. 1A). Geographic unevenness also exists, largely reflecting socioeconomic differences and research priorities across regions. Moreover, we observed an overrepresentation of CC5 and CC8 isolates (Fig. 1B), which can be attributed to their clinical relevance: CC5 is the predominant hospital-associated MRSA (HA-MRSA) lineage12, while CC8 is the primary community-acquired MRSA (CA-MRSA) clone20. These sampling biases, particularly in clonal complex composition, host source, and post-2000 temporal coverage, should be considered when interpreting our results. Consequently, our conclusions reflect patterns observed in the currently available public genomic data.
Among all the involved strains with complete genomes, our findings reveal that 66.8% of the strains harbor plasmids, with the vast majority (92.5%) carrying one to two plasmids (Fig. 1C). These results differ from previous reports, such as those from Malaysia and the Czech Republic, which documented plasmid carriage rates of 90% and 89%, respectively21,22. The discrepancies may arise from the broader geographic and temporal scope of our dataset (1933–2023), which likely introduced greater diversity and variability in plasmid dynamics. Regional variation in antimicrobial usage, host adaptation, and ecological niches may also contribute to differences in plasmid prevalence and maintenance across S. aureus populations23. Additionally, the observed increase in plasmid-carrying strains after 2012, as well as the regional differences, may reflect not only genuine biological phenomena but also the effects of advances in sequencing technology and uneven sampling. Therefore, the results presented here should be interpreted within the context of the available dataset.
Regarding the plasmid number per strain, compared to Enterobacteriaceae, which often carry multiple plasmids per strain (averaging 2–4, and sometimes more than 5 in MDR isolates24), S. aureus exhibits a more restricted plasmid carriage profile. This contrast likely reflects fundamental differences in plasmid ecology: the extensive plasmid repertoire in Enterobacteriaceae may result from higher rates of horizontal gene transfer and broader environmental adaptability, whereas the limited plasmid load in S. aureus may represent an evolutionary trade-off between adaptive benefits and the metabolic costs of plasmid maintenance25. In addition, compared with Enterobacteriaceae, S. aureus relies more on bacteriophage-mediated transduction for horizontal gene transfer5, which may partly explain its lower plasmid carriage.
Regarding plasmid size and architecture, large-scale genomic analysis has shown that Enterobacteriaceae strains, such as E. coli and K. pneumoniae, typically harbor large plasmids (40–150 kb), often associated with AMR and conjugative transfer functions24. In contrast, our study highlighted a distinct plasmid size distribution in S. aureus, with the majority of plasmids falling within the 20–30 kb (42.4%) and <10 kb (27.1%) ranges, while plasmids larger than 60 kb were notably rare (Fig. 2A). This observed size profile aligns with previous findings in S. aureus21,26, where small to medium-sized plasmids predominate. This distribution likely reflects intrinsic differences in plasmid architecture and replication strategies between S. aureus and Enterobacteriaceae species. The predominance of small to medium-sized plasmids in S. aureus may represent an adaptive balance between maintaining beneficial ARGs and minimizing the fitness cost of plasmid carriage23. Plasmid size correlated with ARGs load in S. aureus, as smaller plasmids mainly carried single genes like ermC and tetK, whereas larger ones often contained multiple ARGs such as blaZ, mphC, and msrA (Figs. S5 and S6).
Regarding the plasmid replicon type, plasmids in S. aureus are commonly classified based on the staphylococcal rep gene family system27. In our dataset, 69% of plasmids carried a single replicon type, while 16% were multi-replicon, exhibiting lower diversity in replication types compared with plasmids of other bacterial groups. Notably, a strong correlation was observed between replicon type and plasmid size. The RepA_N replicon was the most prevalent, predominantly associated with mid-sized plasmids (20–30 kb), whereas Rep1 was enriched in small plasmids (<5 kb) (Fig. 2C). This size-dependent distribution aligns with established replication strategies in Staphylococcus species: plasmids smaller than 10 kb typically utilize rolling-circle replication (RCR), which favors compact genomes, whereas larger plasmids predominantly adopt theta-mode replication, enabling the stable maintenance of extensive genetic content5. Similar patterns are observed in other Gram-positive bacteria. For example, RepA_N dominates mid-sized resistance plasmids in Enterococcus spp.28, while Rep1-like systems are common in small cryptic plasmids of Bacillus spp.29. Although previous studies suggested lineage-specific distribution of replicon types in S. aureus30, we found no significant overall correlation. One notable exception was RepA_N, which was highly enriched in CC5 and CC8 strains (89%). The absence of broader associations may reflect sampling bias or limited representation of certain clonal complexes in public genomic databases.
For many years, the role of plasmids in AMR dynamics in S. aureus remained underappreciated and poorly understood. Although several important studies7,21,31 have demonstrated that plasmids play a key role in carrying ARGs in S. aureus, these investigations were largely confined to specific geographic regions. In contrast, our study conducted a comprehensive analysis of plasmid-associated ARGs in 1395 S. aureus genomes collected globally over a 90-year period, encompassing broad geographic and temporal diversity. We identified 35 distinct plasmid-borne ARGs conferring resistance to most major antimicrobial classes, including those classified as critically important antimicrobials (CIAs) for both human and veterinary medicine32,33. Yuan et al.34 reported a temporal increase in S. aureus resistance based on large-scale genomic analysis. Our findings build on this by revealing that this expansion is largely plasmid-driven. Specifically, our longitudinal analysis (1933–2023) showed a significantly faster accumulation of plasmid-borne ARGs compared to chromosomal ARGs, underscoring the central role of plasmids in the long-term evolution of resistance in S. aureus (Fig. 2G). Although this trend may be influenced by sampling bias, particularly the overrepresentation of CC5 and CC8 isolates in our dataset and the limitation for pre-2000 isolates. Nevertheless, several recent studies35,36 have also demonstrated a similar global increase in the abundance and diversity of ARGs carried by plasmids over time, supporting the biological plausibility of the observed pattern in our dataset.
Moreover, the faster accumulation of plasmid-borne ARGs is likely driven by the high mobility and transfer efficiency of plasmids. Unlike chromosomal ARGs, plasmid-mediated resistance genes can rapidly disseminate across bacterial populations via horizontal gene transfer mechanisms, such as conjugation. Our data showed that over 85.5% of plasmids were predicted to be mobilizable, either by encoding their own conjugative machinery or by relying on co-resident conjugative plasmids for mobilization. These mobile genetic elements serve as effective vectors for the acquisition and propagation of ARGs under selective pressure, thus playing a central role in the emergence and persistence of resistance over time37. Further research is necessary to elucidate the molecular mechanisms underlying plasmid transfer, stability, and host range in different environmental settings to better control plasmid-mediated resistance.
In this study, we highlight a typical clonal complex CC5 harboring the highest mean plasmid burden (Fig. 3A), suggesting this clonal complex may possess unique capabilities in acquiring, maintaining, and disseminating plasmids15. A significant positive correlation was observed between genetic distance from the CC5 root and the number of plasmid-borne ARGs (Figs. 4 and S3, Pearson’s r = 0.5546, p < 0.001), indicating a stepwise enrichment of resistance determinants during strain evolution that may enhance the fitness of CC5 strains in antimicrobial-rich environments, such as hospital settings38. Notably, clade CC5.6 stands out due to its significantly higher prevalence of conjugative pSK41 family plasmids. Our data indicate that a major acquisition event of pSK41-like plasmids occurred around 2012 in the United States (Figs. 4 and 5A), coinciding with the emergence of MDR healthcare-associated MRSA and the implementation of the CDC’s Core Elements of Hospital Antibiotic Stewardship Programs39,40. Given that pSK41 conjugative plasmids can not only self-transfer but also mobilize non-conjugative plasmids carrying additional resistance genes, the expansion of CC5.6 strains harboring these plasmids likely accelerated the cooperative dissemination of resistance within this lineage23,41. Supporting this, CC5.6 showed a significant increase in both the quantity and diversity of plasmid-borne ARGs (Fig. S3), which may promote its ecological persistence and potential dominance in both clinical and environmental settings.
Within the One Health framework, food is a crucial interface linking animals, humans, and the environment, facilitating the transmission of S. aureus and AMR42. Food-producing animals can disseminate S. aureus or ARGs to humans through the food chain43, while humans may contaminate food-processing and retail environments, promoting human–food–human or reverse zoonotic transmission44. Although CC5 is primarily recognized as a hospital-associated MRSA lineage, it has increasingly been detected in livestock and food-associated environments, particularly in poultry45 and farm-settings18. In our dataset, 384 CC5 isolates were identified, including 376 from human sources and 4 from food animals. The limited number of food-animal isolates may reflect the historical focus of CC5 research on clinical infections, suggesting that its role in food and animal reservoirs requires further investigation. Importantly, CC5 strains in this study exhibited a high plasmid carriage rate with enrichment of ARGs. Notably, the CC5.6 subclade harboring conjugative plasmids could enhance horizontal ARG transfer and accelerate the dissemination of resistance within the food–animal–human continuum. Collectively, these findings underscore the potential of CC5, particularly CC5.6, to contribute to resistance transmission and persistence across sectors, posing emerging challenges to food safety and public health46.
In conclusion, this study provides a comprehensive global analysis of 1395 S. aureus genomes spanning 90 years, revealing the pivotal role of plasmids in shaping the AMR landscape of this zoonotic pathogen. Plasmids were identified in 66.8% of strains, with most (92.5%) carrying one to two plasmids, primarily of small to medium size (20–30 kb: 42.4%; <10 kb: 27.1%), while large plasmids (>60 kb) were rare. A strong association was observed between replicon type and plasmid size, with RepA_N predominant in mid-sized plasmids (20–30 kb) and Rep1 enriched in small plasmids (<5 kb). Thirty-five distinct plasmid-borne ARGs were identified. Longitudinal analysis revealed that plasmid-borne ARGs accumulated significantly faster than chromosomal ARGs, underscoring their central role in the long-term evolution of resistance, likely driven by the high mobility of plasmids, with 85.5% predicted to be mobilizable. Clonal complex analysis showed that CC5 strains harbored the highest mean plasmid burden, with subclade CC5.6 standing out for its high prevalence of conjugative pSK41-family plasmids and elevated diversity and abundance of plasmid-borne ARGs. A major pSK41 acquisition event occurred around 2012 in the U.S., followed by the expansion of a resistant clade, raising concerns about the potential for these conjugative plasmids to accelerate ARG dissemination. These findings underscore the central role of plasmids in the spread of AMR in S. aureus, stressing the need for targeted surveillance of plasmid-mediated resistance in high-risk strains in both food production/processing environments and clinical settings, and the development of strategies to hinder plasmid transfer or persistence to combat the global threat of AMR.
Methods
Dataset assembly of complete S. aureus genomes
We retrieved complete genome data for 1395 S. aureus strains (1230 human-derived and 165 animal-derived), spanning 1933 to 2023, from the GenBank database. Associated metadata, including collection dates, geographic locations, and host information, were also compiled (Supplementary Data 1).
Genome quality control was performed using CheckM v1.2.147 with default parameters. Only assemblies with completeness ≥95% and contamination ≤5% were retained. Geospatial visualization was performed in R 4.5.148 by mapping strain coordinates derived from isolation metadata. Gene annotation was performed using Prokka v1.14.649 with default parameters. Multilocus sequence typing (MLST) profiles and clonal complex (CC) assignments were predicted through the PubMLST database50 using a BLASTn-based approach51.
Plasmid sequence identification, replicon typing, and mobility prediction
SeqKit52 was used to extract plasmid sequences from complete S. aureus genomes and calculate their sizes in base pairs (bp). Replicon typing was performed using a locally installed version of PlasmidFinder v1.353, with the accompanying database downloaded from the Center for Genomic Epidemiology (CGE). Replicon typing was performed using BLAST+ v2.6.051, with thresholds of ≥80% sequence identity and ≥60% query coverage.
Relaxase genes were identified by querying plasmid sequences against the MOBfamDB database using HMMER3 v3.3.2, with a domain coverage cutoff of ≥60% and an E-value threshold of ≤0.0154. OriT-mimic sequences were retrieved from known conjugative staphylococcal plasmids (pWBG749 and pSK41) using a local BLASTn search51. Conjugative genes were identified by comparing plasmid sequences with known conjugative staphylococcal plasmids (pWBG749: smpA-smpW; pSK41: traA-traM; pWBG4: detA-detV) using local BLASTN alignment51. Plasmids containing these conjugative transfer gene clusters were classified as conjugative plasmids. Plasmids harboring either a conjugative plasmid oriT mimic or a relaxase were categorized as mobilizable. Those lacking both characteristics were designated as non-mobilizable.
Identification of ARGs
The ARGs located on chromosomes and plasmids were predicated separately using ResFinder55, with the thresholds set at >80% sequence identity and >60% query coverage.
Phylogenetic analysis of CC5 strains
A phylogenetic tree of 384 CC5 strains was constructed using KSNP456 based on core genome single nucleotide polymorphisms (cgSNPs) and was rooted with a S. aureus CC1 genome (CGA_024741775) as the outgroup. The maximum-likelihood tree was generated from 25,567 high-quality cgSNPs. Branch support was evaluated using 1000 bootstrap replicates, bootstrap values ≥ 80% are shown at nodes. The Interactive Tree of Life (ITOL)57 was utilized to visualize and annotate the phylogenetic tree.
To estimate the evolutionary timescale of clade CC5.6 within the CC5 lineage, Bayesian evolutionary analysis was performed using BEAST v1.8.258. Jmodeltest59 was employed to determine the best-fitting substitution model. Three molecular clock models-strict, relaxed lognormal, and relaxed exponential-were evaluated. Each model combination involved chains of 20 million generations, sampled every 2000 generations. The analysis indicated that the relaxed lognormal clock paired with the GTR + G + I substitution model was the best-supported molecular clock model. The maximum clade credibility tree was then constructed using TreeAnnotator v2.6.758 and visualized with Figtree v1.4.460.
Data availability
All data analyzed in this study were obtained from NCBI and are included in this published article and its supplementary information files.
References
Costa, F. G., Mills, K. B., Crosby, H. A. & Horswill, A. R. The Staphylococcus aureus regulatory program in a human skin-like environment. mBio. 15, e00453–24 (2024).
Wu, S. et al. Prevalence and characterization of Staphylococcus aureus isolated from retail vegetables in China. Front. Microbiol. 9, 1263 (2018).
Wang, D. et al. Understanding bacterial ecology to combat antibiotic resistance dissemination. Trends Biotechnol. 43, 1566–1582 (2025).
Garcillán-Barcia, M. P., Redondo-Salvo, S. & de la Cruz, F. Plasmid classifications. Plasmid 126, 102684 (2023).
Firth, N. et al. Staphylococcal plasmids, transposable and integrative elements. Microbiol. Spectr. 6, GPP3–0030-2018 (2018).
Acman, M., van Dorp, L., Santini, J. M. & Balloux, F. Large-scale network analysis captures biological features of bacterial plasmids. Nat. Commun. 11, 2452 (2020).
Contarin, R. et al. The interplay between mobilome and resistome in Staphylococcus aureus. mBio. 15, e02428-24 (2024).
Ramsay, J. P. et al. An updated view of plasmid conjugation and mobilization in Staphylococcus. Mob. Genet. Elem. 6, e1208317 (2016).
Lozano, C. et al. Expansion of a plasmid classification system for Gram-positive bacteria and determination of the diversity of plasmids in Staphylococcus aureus strains of human, animal, and food origins. Appl. Environ. Microbiol. 78, 5948–5955 (2012).
Pollet, R. M. et al. Processing of nonconjugative resistance plasmids by conjugation nicking enzyme of Staphylococci. J. Bacteriol. 198, 888–897 (2016).
O’Brien, F. G. et al. Origin-of-transfer sequences facilitate mobilisation of non-conjugative antimicrobial-resistance plasmids in Staphylococcus aureus. Nucleic Acids Res. 43, 7971–7983 (2015).
Chen, F. et al. Global genetic diversity and Asian clades evolution: a phylogeographic study of Staphylococcus aureus sequence type 5. Antimicrob. Agents Chemother. 68, e01175-23 (2024).
Challagundla, L. et al. Phylogenomic classification and the evolution of clonal complex 5 methicillin-resistant Staphylococcus aureus in the Western Hemisphere. Front. Microbiol. 9, 1901 (2018).
Fan, Y. X. et al. Sequence Type 5 (ST5) as a possible predictor of bacterial persistence in adult patients with methicillin-resistant Staphylococcus aureus pneumonia treated with vancomycin. Microbiol. Spectr. 10, e01348–22 (2022).
Kos, V. N. et al. Comparative genomics of vancomycin-resistant Staphylococcus aureus strains and their positions within the clade most commonly associated with methicillin-resistant S. aureus hospital-acquired infection in the United States. mBio. 3, e00112-12 (2012).
Strommenger, B. et al. Evolution of methicillin-resistant Staphylococcus aureus towards increasing resistance. J. Antimicrob. Chemother. 69, 616–622 (2014).
Lowder, B. V. et al. Recent human-to-poultry host jump, adaptation, and pandemic spread of Staphylococcus aureus. Proc. Natl. Acad. Sci. USA 106, 19545–19550 (2009).
Kawanishi, M. et al. Prevalence and genetic characterization of methicillin-resistant Staphylococcus aureus isolated from pigs in Japan. Antibiotics 13, 155 (2024).
Silva, V. et al. Staphylococcus aureus and MRSA in livestock. Microorganisms 11, 124 (2023).
Tabatabaie Poya, F. S. et al. Unveiling the genetic landscape of Staphylococcus aureus isolated from hospital wastewaters: emergence of hypervirulent CC8 strains in Tehran, Iran. Int. J. Microbiol. 2025, 5458315 (2025).
Al-Trad, E. I. et al. The plasmidomic landscape of clinical methicillin-resistant Staphylococcus aureus isolates from Malaysia. Antibiotics 12, 733 (2023).
Kuntová, L. et al. Characteristics and distribution of plasmids in a clonally diverse set of methicillin-resistant Staphylococcus aureus strains. Arch. Microbiol. 194, 607–614 (2012).
Liu, Z., Zhao, Q., Xu, C. & Song, H. Compensatory evolution of chromosomes and plasmids counteracts the plasmid fitness cost. Ecol. Evol. 14, e70121 (2024).
Galata, V., Fehlmann, T., Backes, C. & Keller, A. PLSDB: a resource of complete bacterial plasmids. Nucleic Acids Res. 47, D195–D202 (2019).
Hall, J. P. et al. Plasmid fitness costs are caused by specific genetic conflicts enabling resolution by compensatory mutation. PLoS Biol. 19, e3001225 (2021).
Neyaz, L., Rajagopal, N., Wells, H. & Fakhr, M. K. Molecular characterization of Staphylococcus aureus plasmids associated with strains isolated from various retail meats. Front. Microbiol. 11, 223 (2020).
Jensen, L. B. et al. A classification system for plasmids from enterococci and other Gram-positive bacteria. J. Microbiol. Methods 80, 25–43 (2010).
Weaver, K. E., Kwong, S. M., Firth, N. & Francia, M. V. The RepA_N replicons of Gram-positive bacteria: a family of broadly distributed but narrow host range plasmids. Plasmid 61, 94–109 (2009).
Reyes-Ramírez, A. & Ibarra, J. E. Plasmid patterns of Bacillus thuringiensis type strains. Appl. Environ. Microbiol. 74, 125–129 (2008).
McCarthy, A. J. & Lindsay, J. A. The distribution of plasmids that carry virulence and resistance genes in Staphylococcus aureus is lineage associated. BMC Microbiol. 12, 104 (2012).
Alhejaili, A. Y. et al. Methicillin-resistant Staphylococcus aureus in Saudi Arabia: genomic evidence of recent clonal expansion and plasmid-driven resistance dissemination. Front. Microbiol. 16, 1602985 (2025).
World Health Organization. WHO’s list of medically important antimicrobials (World Health Organization, 2024).
Vestergaard, M., Frees, D. & Ingmer, H. Antibiotic resistance and the MRSA problem. Microbiol. Spectr. 7, GPP3–0057-2018 (2019).
Yuan, H. et al. The global antimicrobial resistance trends of Staphylococcus aureus and influencing factors. Microbiol. Res. 16, 118 (2025).
Wang, Y. et al. Global increase of antibiotic resistance genes in conjugative plasmids. Microbiol. Spectr. 11, e04478–22 (2023).
Cazares, A. et al. Pre- and postantibiotic epoch: the historical spread of antimicrobial resistance. Science 387, eadr1522 (2025).
Lerminiaux, N. A. & Cameron, A. D. S. Horizontal transfer of antibiotic resistance genes in clinical environments. Can. J. Microbiol. 65, 34–44 (2019).
Benz, F. & Hall, A. R. Host-specific plasmid evolution explains the variable spread of clinical antibiotic-resistance plasmids. Proc. Natl. Acad. Sci. USA 120, e2212147120 (2023).
King, L. M., Bartoces, M., Fleming-Dutra, K. E., Roberts, R. M. & Hicks, L. A. Changes in U.S. outpatient antibiotic prescriptions from 2011–2016. Clin. Infect. Dis. 70, 370–377 (2020).
Cosgrove, S. E. & Srinivasan, A. Antibiotic stewardship: a decade of progress. Infect. Dis. Clin. North Am. 37, 659–667 (2023).
Millan, S. A. & MacLean, R. C. Fitness costs of plasmids: a limit to plasmid transmission. Microbiol. Spectr. 5, MTBP–0016-2017 (2017).
Pan, Y. et al. One health genomic insights into the host-specific evolution and cross-host transmission of Staphylococcus aureus in animal farm environments, food of animal origin, and humans. Int. J. Antimicrob. Agents 62, 106932 (2023).
Qian, J., Wu, Z., Zhu, Y. & Liu, C. One Health: a holistic approach for food safety in livestock. Sci. One Health 1, 100015 (2023).
Al Noman, Z. et al. A systematic review on reverse zoonosis: global impact and changes in transmission patterns. J. Adv. Vet. Anim. Res. 11, 601–617 (2024).
Paula, E. et al. High prevalence of avian-adapted CC5 Staphylococcus aureus isolates in poultry meat in Spain: food chain as vehicle of MRSA and MSSA CC398-t1451. Int. J. Food Sci. Technol. 59, 9180–9188 (2024).
Lv, G. et al. Molecular characteristics of Staphylococcus aureus from food samples and food poisoning outbreaks in Shijiazhuang, China. Front. Microbiol. 12, 652276 (2021).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Team, R.C. R: a language and environment for statistical computing. R Foundation for Statistical Computing. Available at: https://www.R-project.org/ (2020).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Jolley, K. A., Bray, J. E. & Maiden, M. C. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 3, 124 (2018).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 15, 421 (2009).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962 (2016).
Carattoli, A. & Hasman, H. PlasmidFinder and in silico pMLST: identification and typing of plasmid replicons in whole-genome sequencing (WGS). Methods Mol. Biol. 2075, 285–294 (2020).
Garcillán-Barcia, M. P., Redondo-Salvo, S., Vielva, L. & de la Cruz, F. MOBscan: automated annotation of MOB relaxases. Methods Mol. Biol. 2075, 295–308 (2019).
Bortolaia, V. et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 75, 3491–3500 (2020).
Hall, B. G. & Nisbet, J. Building phylogenetic trees from genome sequences with kSNP4. Mol. Biol. Evol. 40, msad235 (2023).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Drummond, A. J., Suchard, M. A., Xie, D. & Rambaut, A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol. Biol. Evol. 29, 1969–1973 (2012).
Posada, D. jModelTest: phylogenetic model averaging. Mol. Biol. Evol. 25, 1253–1256 (2008).
Rambaut, A. FigTree, A graphical viewer of phylogenetic trees. Available at: http://tree.bio.ed.ac.uk/software/figtree/ (2009).
Acknowledgements
This study was supported by the National Key R&D program of China (No. 2024YFE0199000), the National Natural Science Foundation of China (No. 32472458) and the Natural Science Foundation of Shanghai (No. 24ZR1436200). L.L. is additionally supported by the UK Medical Research Council (No. MR/Y015169/1) and the Biotechnology and UK Biological Sciences Research Council (No. BB/P013740/1).
Author information
Authors and Affiliations
Contributions
T.X. contributed to the original draft writing, methodology, investigation, formal analysis, visualization, and data curation. Z. Z. contributed to the original draft writing, methodology, and review & editing of the manuscript. H.W. performed formal analysis and data curation. L.J. contributed to methodology and manuscript review & editing. K.S., M.Z., and Z.N. were involved in manuscript review & editing. L.L. contributed to methodology and manuscript review & editing. S.C. contributed to review & editing, funding acquisition, visualization, validation, supervision, methodology, resources, and conceptualization. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tian, X., Zhang, Z., Hou, W. et al. Plasmidomic landscape of Staphylococcus aureus and the emergence of a CC5 subclade harboring the conjugative plasmid pSK41: implications for food safety and antimicrobial resistance. npj Sci Food 10, 78 (2026). https://doi.org/10.1038/s41538-026-00733-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41538-026-00733-7