Abstract
Somatic genome editing in mouse models has increased our understanding of the in vivo effects of genetic alterations. However, existing models have a limited ability to create multiple targeted edits, hindering our understanding of complex genetic interactions. Here we generate transgenic mice with Cre-regulated and constitutive expression of enhanced Acidaminococcus sp. Cas12a (enAsCas12a), which robustly generates compound genotypes, including diverse cancers driven by inactivation of trios of tumour suppressor genes or an oncogenic translocation. We integrate these modular CRISPR RNA (crRNA) arrays with clonal barcoding to quantify the size and number of tumours with each array, as well as the impact of varying the guide number and position within a four-guide array. Finally, we generate tumours with inactivation of all combinations of nine tumour suppressor genes and find that the fitness of triple-knockout genotypes is largely explainable by one- and two-gene effects. These Cas12a alleles will enable further rapid creation of disease models and high-throughput investigation of coincident genomic alterations in vivo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All data generated from this study have been deposited at the NCBI Gene Expression Omnibus and are accessible through GEO series accession number GSE286992 (ref. 78). Source data are provided with this paper.
Code availability
Python (v.3.9.12) was used for all analyses of barcode sequencing data and data visualization. Code for the analysis of barcode sequencing data is accessible on GitHub (https://github.com/JasperXuEvolution/Efficient-and-multiplexed-somatic-genome-editing-with-Cas12a-mice)79.
References
Kersten, K., de Visser, K. E., van Miltenburg, M. H. & Jonkers, J. Genetically engineered mouse models in oncology research and cancer medicine. EMBO Mol. Med. 9, 137–153 (2017).
Hebert, J. D., Neal, J. W. & Winslow, M. M. Dissecting metastasis using preclinical models and methods. Nat. Rev. Cancer 23, 391–407 (2023).
Sánchez-Rivera, F. J. & Jacks, T. Applications of the CRISPR–Cas9 system in cancer biology. Nat. Rev. Cancer 15, 387–393 (2015).
Dow, L. E. Modeling disease in vivo with CRISPR/Cas9. Trends Mol. Med. 21, 609–621 (2015).
Murray, C. W. et al. An LKB1-SIK axis suppresses lung tumor growth and controls differentiation. Cancer Discov. 9, 1590–1605 (2019).
Paul, B. & Montoya, G. CRISPR-Cas12a: functional overview and applications. Biomed. J. 43, 8–17 (2020).
Khan, S. & Sallard, E. Current and prospective applications of CRISPR-Cas12a in pluricellular organisms. Mol. Biotechnol. 65, 196–205 (2023).
Zetsche, B. et al. Multiplex gene editing by CRISPR–Cpf1 using a single crRNA array. Nat. Biotechnol. 35, 31–34 (2017).
Campa, C. C., Weisbach, N. R., Santinha, A. J., Incarnato, D. & Platt, R. J. Multiplexed genome engineering by Cas12a and CRISPR arrays encoded on single transcripts. Nat. Methods 16, 887–893 (2019).
Dede, M., McLaughlin, M., Kim, E. & Hart, T. Multiplex enCas12a screens detect functional buffering among paralogs otherwise masked in monogenic Cas9 knockout screens. Genome Biol. 21, 262 (2020).
Dai, X. et al. Massively parallel knock-in engineering of human T cells. Nat. Biotechnol. 41, 1239–1255 (2023).
Magnusson, J. P., Rios, A. R., Wu, L. & Qi, L. S. Enhanced Cas12a multi-gene regulation using a CRISPR array separator. eLife 10, e66406 (2021).
DeWeirdt, P. C. et al. Optimization of AsCas12a for combinatorial genetic screens in human cells. Nat. Biotechnol. 39, 94–104 (2021).
Ma, E. et al. Improved genome editing by an engineered CRISPR-Cas12a. Nucleic Acids Res. 50, 12689–12701 (2022).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing. Nat. Biotechnol. 37, 276–282 (2019).
Gier, R. A. et al. High-performance CRISPR-Cas12a genome editing for combinatorial genetic screening. Nat. Commun. 11, 3455 (2020).
Zhang, L. et al. AsCas12a ultra nuclease facilitates the rapid generation of therapeutic cell medicines. Nat. Commun. 12, 3908 (2021).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Aaltonen, L. A. et al. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Liu, P. et al. Enhanced Cas12a editing in mammalian cells and zebrafish. Nucleic Acids Res. 47, 4169–4180 (2019).
Yousefi, M. et al. Combinatorial inactivation of tumor suppressors efficiently initiates lung adenocarcinoma with therapeutic vulnerabilities. Cancer Res. 82, 1589–1602 (2022).
Rogers, Z. N. et al. A quantitative and multiplexed approach to uncover the fitness landscape of tumor suppression in vivo. Nat. Methods 14, 737–742 (2017).
Cai, H. et al. A functional taxonomy of tumor suppression in oncogenic KRAS-driven lung cancer. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-20-1325 (2021).
Tang, Y. J. et al. Functional mapping of epigenetic regulators uncovers coordinated tumor suppression by the HBO1 and MLL1 complexes. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/2024.08.19.607671v1 (2024).
Yousefi, M. et al. Fully accessible fitness landscape of oncogene-negative lung adenocarcinoma. Proc. Natl Acad. Sci. USA. 120, e2303224120 (2023).
Schaffer, B. E. et al. Loss of p130 accelerates tumor development in a mouse model for human small-cell lung carcinoma. Cancer Res. 70, 3877–3883 (2010).
Duplaquet, L. et al. KDM6A epigenetically regulates subtype plasticity in small cell lung cancer. Nat. Cell Biol. 25, 1346–1358 (2023).
Iacobuzio-Donahue, C. A., Velculescu, V. E., Wolfgang, C. L. & Hruban, R. H. Genetic basis of pancreas cancer development and progression: insights from whole-exome and whole-genome sequencing. Clin. Cancer Res. 18, 4257–4265 (2012).
Halbrook, C. J., Lyssiotis, C. A., Pasca di Magliano, M. & Maitra, A. Pancreatic cancer: advances and challenges. Cell 186, 1729–1754 (2023).
Izeradjene, K. et al. Kras(G12D) and Smad4/Dpc4 haploinsufficiency cooperate to induce mucinous cystic neoplasms and invasive adenocarcinoma of the pancreas. Cancer Cell 11, 229–243 (2007).
Hingorani, S. R. et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005).
Bardeesy, N. et al. Smad4 is dispensable for normal pancreas development yet critical in progression and tumor biology of pancreas cancer. Genes Dev. 20, 3130–3146 (2006).
Aguirre, A. J. et al. Activated Kras and Ink4a/Arf deficiency cooperate to produce metastatic pancreatic ductal adenocarcinoma. Genes Dev. 17, 3112–3126 (2003).
Hingorani, S. R. et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell 4, 437–450 (2003).
Whittle, M. C. et al. RUNX3 controls a metastatic switch in pancreatic ductal adenocarcinoma. Cell 161, 1345–1360 (2015).
Bardeesy, N. et al. Both p16Ink4a and the p19Arf-p53 pathway constrain progression of pancreatic adenocarcinoma in the mouse. Proc. Natl Acad. Sci. USA 103, 5947–5952 (2006).
Chiou, S.-H. et al. Pancreatic cancer modeling using retrograde viral vector delivery and in vivo CRISPR/Cas9-mediated somatic genome editing. Genes Dev. 29, 1576–1585 (2015).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15, 271–274 (2018).
Hegde, M., Strand, C., Hanna, R. E. & Doench, J. G. Uncoupling of sgRNAs from their associated barcodes during PCR amplification of combinatorial CRISPR screens. PLoS ONE 13, e0197547 (2018).
Glenfield, C. & Innan, H. Gene duplication and gene fusion are important drivers of tumourigenesis during cancer evolution. Genes 12, 1376 (2021).
Schneider, J. L., Lin, J. J. & Shaw, A. T. ALK-positive lung cancer: a moving target. Nat. Cancer 4, 330–343 (2023).
Sabir, S. R., Yeoh, S., Jackson, G. & Bayliss, R. EML4-ALK variants: biological and molecular properties, and the implications for patients. Cancers 9, 118 (2017).
Diaz-Jimenez, A. et al. EML4-ALK variant-specific genetic interactions shape lung tumorigenesis. Preprint at bioRxiv https://doi.org/10.1101/2024.08.26.609730 (2024).
Maddalo, D. et al. In vivo engineering of oncogenic chromosomal rearrangements with the CRISPR/Cas9 system. Nature 516, 423–427 (2014).
Chen, Y. et al. Club cells employ regeneration mechanisms during lung tumorigenesis. Nat. Commun. 13, 4557 (2022).
Parrish, P. C. R. et al. Discovery of synthetic lethal and tumor suppressor paralog pairs in the human genome. Cell Rep. 36, 109597 (2021).
Nguyen, B. et al. Genomic characterization of metastatic patterns from prospective clinical sequencing of 25,000 patients. Cell 185, 563–575 (2022).
Tang, K. et al. Cas12a-knock-in mice for multiplexed genome editing, disease modelling and immune-cell engineering. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-025-01371-2 (2025).
Jin, W. et al. Advancing the genetic engineering toolbox by combining AsCas12a knock-in mice with ultra-compact screening. Nat. Commun. 16, 974 (2025).
Li, P. et al. Cas12a mediates efficient and precise endogenous gene tagging via MITI: microhomology-dependent targeted integrations. Cell. Mol. Life Sci. 77, 3875–3884 (2020).
Li, A. et al. AAV-CRISPR gene editing is negated by pre-existing immunity to Cas9. Mol. Ther. 28, 1432–1441 (2020).
Hughes, N. W. et al. Machine-learning-optimized Cas12a barcoding enables the recovery of single-cell lineages and transcriptional profiles. Mol. Cell 82, 3103–3118 (2022).
Yang, D. et al. Lineage tracing reveals the phylodynamics, plasticity, and paths of tumor evolution. Cell 185, 1905–1923 (2022).
Li, L. et al. A mouse model with high clonal barcode diversity for joint lineage, transcriptomic, and epigenomic profiling in single cells. Cell 186, 5183–5199 (2023).
Bowling, S. et al. An engineered CRISPR-Cas9 mouse line for simultaneous readout of lineage histories and gene expression profiles in single cells. Cell 181, 1410–1422 (2020).
Chan, M. M. et al. Molecular recording of mammalian embryogenesis. Nature 570, 77–82 (2019).
Santinha, A. J. et al. Transcriptional linkage analysis with in vivo AAV-Perturb-seq. Nature 622, 367–375 (2023).
Jin, X. et al. In vivo Perturb-seq reveals neuronal and glial abnormalities associated with autism risk genes. Science 370, eaaz6063 (2020).
Zheng, X. et al. Massively parallel in vivo Perturb-seq reveals cell type-specific transcriptional networks in cortical development. Cell 187, 3236–3248 (2024).
Lin, R. et al. Directed evolution of adeno-associated virus for efficient gene delivery to microglia. Nat. Methods 19, 976–985 (2022).
Li, B. E. et al. In vivo CRISPR/Cas9 screening identifies Pbrm1 as a regulator of myeloid leukemia development in mice. Blood Adv. 7, 5281–5293 (2023).
Romero, R. et al. Keap1 loss promotes Kras-driven lung cancer and results in dependence on glutaminolysis. Nat. Med. 23, 1362–1368 (2017).
Ng, S. R. et al. CRISPR-mediated modeling and functional validation of candidate tumor suppressor genes in small cell lung cancer. Proc. Natl Acad. Sci. USA 117, 513–521 (2020).
Lee, M. C. et al. A multiplexed in vivo approach to identify driver genes in small cell lung cancer. Cell Rep. 42, 111990 (2023).
Belk, J. A. et al. Genome-wide CRISPR screens of T cell exhaustion identify chromatin remodeling factors that limit T cell persistence. Cancer Cell 40, 768–786 (2022).
Song, X. et al. Genome editing with AAV-BR1-CRISPR in postnatal mouse brain endothelial cells. Int. J. Biol. Sci. 18, 652–660 (2022).
Thürkauf, M. et al. Fast, multiplexable and efficient somatic gene deletions in adult mouse skeletal muscle fibers using AAV-CRISPR/Cas9. Nat. Commun. 14, 6116 (2023).
Kim, H. K. et al. Deep learning improves prediction of CRISPR-Cpf1 guide RNA activity. Nat. Biotechnol. 36, 239–241 (2018).
Schwenk, F., Baron, U. & Rajewsky, K. A cre-transgenic mouse strain for the ubiquitous deletion of loxP-flanked gene segments including deletion in germ cells. Nucleic Acids Res. 23, 5080–5081 (1995).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Bae, S., Park, J. & Kim, J.-S. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475 (2014).
Domingo, J., Diss, G. & Lehner, B. Pairwise and higher-order genetic interactions during the evolution of a tRNA. Nature 558, 117–121 (2018).
Poelwijk, F. J., Krishna, V. & Ranganathan, R. The context-dependence of mutations: a linkage of formalisms. PLoS Comput. Biol. 12, e1004771 (2016).
Shuldiner, E. G. et al. Aging represses lung tumorigenesis and alters tumor suppression. Preprint at BioRxiv https://doi.org/10.1101/2024.05.28.596319 (2024).
Hebert, J. D. et al. Data for ‘Efficient and multiplexed somatic genome editing with Cas12a mice’. GEO www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE286992 (2025).
Hebert, J. D. et al. Source code for ‘Efficient and multiplexed somatic genome editing with Cas12a mice’. GitHub github.com/JasperXuEvolution/Efficient-and-multiplexed-somatic-genome-editing-with-Cas12a-mice (2025).
Acknowledgements
We thank the Stanford Veterinary Animal Care staff for expert animal care; G. Charville for advice on histological characterization of tumours; the staff at Cyagen Biosciences for transgenic mouse generation; the staff at the Massachusetts General Hospital Center for Computational and Integrative Biology DNA Core for CRISPR off-target sequencing; and the members of the Winslow laboratory for comments. J.D.H. was supported by an American Cancer Society Fellowship (PF-21-112-01-MM) and a TRDRP Postdoctoral fellowship (T31FT1619); H.X. by a TRDRP Postdoctoral fellowship (T34FT8013); Y.J.T. was partly supported by the Canadian Institute of Health Research (CIHR) postdoctoral fellowship (MFE-176568); P.A.R. by the National Science Foundation Graduate Research Fellowship (DGE-2146755) and the Lucille P. Markey Stanford Graduate Fellowship; S.K. by the National Cancer Institute Predoctoral to Postdoctoral Fellow Transition Award (K00-CA234962). This work was supported by NIH R01-CA230025 (to M.M.W.), NIH R01-CA231253 (to M.M.W. and D.A.P.), NIH R01-CA234349 (to M.M.W. and D.A.P.), NIH R35-CA231997 (to J.S.), NIH R35-HG011316 (to L.C.), R01-GM141627 (to L.C.), NIH P01-CA244114 and in part by the Stanford Cancer Institute support grant (NIH P30-CA124435).
Author information
Authors and Affiliations
Contributions
J.D.H. and H.X. contributed equally to this study. H.X. conducted all data analyses. Y.J.T. developed Tuba-seqUltra and associated methods. J.D.H., P.A.R., C.R.D., J.W., O.D., V.P.S., L.A., I.A. and R.T. performed experiments. J.D.H., H.X., N.W.H., S.K., R.S., J.S., L.C., D.A.P. and M.M.W. designed the study. J.D.H., N.W.H. and M.M.W. designed the enAsCas12a targeting construct and used it to create the H11LSL-Cas12a and H11Cas12a mice. J.D.H., H.X. and M.M.W. wrote the paper with input from all of the authors.
Corresponding author
Ethics declarations
Competing interests
D.A.P. and M.M.W. are founders and hold equity in Guide Oncology. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Comparison of salient features of Cas9 and Cas12a, and broad nuclear expression of Cas12a in H11Cas12a mice.
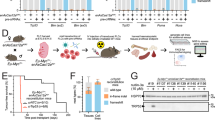
a. Summary of features of Cas9 and Cas12a relevant to multiplexed genome editing. Note that the potential impact of off-target effects is increased when targeting more genes. b. PCR genotyping of mice of the indicated H11LSL-Cas12a and H11Cas12a genotypes. c. Western blots on tissue lysates from H11Cas12a or H11LSL-Cas12a/WT mice. α-Tubulin shows loading. Data representative of 3 independent experiments. d. Immunohistochemical staining for the Cas12a HA tag in lung, liver and kidney sections from the indicated genotypes of mice. Scale bars, 50 µm. Higher magnification images of these liver sections are shown in Fig. 1f. e. Outline of the one-step cloning to generate barcoded Cas12a pre-crRNA vector pools. Guides, Cas12a spacers; Repeats, Cas12a direct repeat sequences (diverse); BsmBI, type IIS restriction enzyme sites; bU6, bovine RNA Pol III promoter; BC, barcode; PGK, phosphoglycerate kinase promoter; LTR, long terminal repeat.
Extended Data Fig. 2 Cas12a-mediated somatic genome editing induces indels at target genetic loci, while incorporation of Cas12a-mediated genome editing with Tuba-seqUltra enables quantification of the effects of each crRNA array.
a-c. Indel frequencies for genomic regions targeted by each crRNA within bulk tumour-bearing lung tissue (a), microdissected lung tumours (b) or bulk tumour-bearing pancreas tissue (c) from H11LSL-Cas12a mice. Tumours were initiated with the indicated lentiviral vector pool. Pancreas tissue samples (c) had variable tumour burden, likely explaining observed variability in indel frequencies among samples. Percentages shown indicate the presence of Cas12a-mediated indels, rather than the absolute measurement of Cas12a or individual crRNA efficiency. For a detailed assessment of the efficiency of Cas12a-mediated editing in vivo, see Fig. 6 and Extended Data Figs. 6–8. Symbol shapes and colours identify values from the same sample. Bars represent means. d. Detailed schematic of tumour barcoding with high-throughput BC-crRNA sequencing to determine the size of each Cas12a-induced clonal tumour. e-f. Number of tumours (e) and mean tumour size assuming a log-normal distribution (f) for each crRNA array (27 combinations in total) relative to the median value for all arrays in H11LSL-Cas12a mice. Medians and 95% confidence intervals are shown based on bootstrap resampling with 1,000 iterations.
Extended Data Fig. 3 Inactivation of Trp53, Cdkn2a, and Smad4 greatly increases tumour size and leads to the development of PDAC with differentiated areas with CK19+ cancer cells and desmoplastic stroma as well as more poorly differentiated areas.
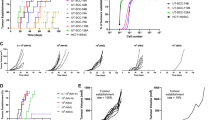
a. Immunohistochemistry images of pancreas sections from the indicated genotypes, showing Trichrome (upper) and Cytokeratin-19 (CK19; lower) staining. KT;H11LSL-Cas12a panels show representative poorly differentiated (left) and more differentiated (right) regions. Scale bars, 100 μm. b. Immunohistochemical staining for the Cas12a HA tag on pancreas tissue from representative KT;H11LSL-Cas12a, KT (lacking Cas12a) and H11LSL-Cas12a mice transduced with Lenti-U6BC-crTrp53/Cdkn2a/Smad4-Cre. Scale bars, 100 μm. c. Total number of neoplastic cells (Total tumour burden) in each mouse normalized to viral titre. Each dot represents a mouse and the bar is the median. Note that two H11LSL-Cas12a mice did not have detectable tumour burden and are not plotted. P value calculated with two-sided Wilcoxon rank-sum test (KT;H11LSL-Cas12a, n = 4 mice; KT, n = 4; H11LSL-Cas12a, n = 2). d. Probability distribution of the size of tumours in the indicated genotypes of mice. e. Mean tumour size assuming a log-normal distribution for each crRNA (guide) relative to the median value for each gene. Medians and 95% confidence intervals are shown based on bootstrap resampling with 1,000 iterations.
Extended Data Fig. 4 Generation of Cas12a Lenti-U6BC-crRNA-Cre vector libraries is associated with minimal shuffling of crRNA sequences.
a,c,e. Fraction of all U6-integrated barcode reads that are associated with only a single crRNA array within each vector pool in the indicated mouse genotypes: Lenti-U6BC-crNf1/Rasa1/Pten-Cre (a), Lenti-U6BC-crRb1/Trp53/Rbl2-Cre (c), and Lenti-U6BC-crTrp53/Cdkn2a/Smad4-Cre (e). Each dot represents a mouse. Bars represent medians. b,d,f. Fraction of total neoplastic cells (total tumour burden) remaining after filtering out spurious “tumour” reads arising during library preparation and sequencing of each pool in the indicated mouse genotypes: Lenti-U6BC-crNf1/Rasa1/Pten-Cre (b), Lenti-U6BC-crRb1/Trp53/Rbl2-Cre (d), and Lenti-U6BC-crTrp53/Cdkn2a/Smad4-Cre (f). Each dot represents a mouse. Bars represent medians.
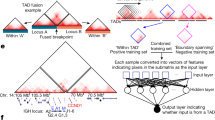
Extended Data Fig. 5 Cas12a-induced oncogenic translocations enable generation of tumours driven by Eml4-Alk fusion.
a. The Lenti-U6BC-crEml4V3/Alk/Trp53-Cre pool has guides targeting intron 6 of Eml4 and intron 19 of Alk to generate the Eml4-Alk Variant 3 (V3) fusion, along with a guide targeting Trp53. Eml4 and Alk are targeted by three crRNAs each in all 9 combinations. b. Intratracheal delivery of Lenti-U6BC-crEml4V3/Alk/Trp53-Cre to H11Cas12a, H11LSL-Cas12a and Wild-type (WT) mice. Viral titre (infectious units, ifu) and mouse numbers are indicated. c. Lung weights from mice 12 weeks after transduction with Lenti-U6BC-crEml4V3/Alk/Trp53-Cre. Each dot represents a mouse. Mean +/- standard deviation is indicated. Data from H11LSL-Cas12a and H11Cas12a mice are merged. d. Light images (upper) and hematoxylin & eosin (H&E) staining of sections (lower) of lungs from the indicated genotypes. Scale bars, 5 mm (upper), 100 μm (lower). e. PCR amplification across all expected Eml4 and Alk junctions from bulk tumour-bearing lung tissue, using primers specific to cut sites created by the indicated crRNAs. Note that the cut sites for crAlk#1 and crAlk#2 are detected by the same primer due to their proximity to each other. Data representative of three independent experiments.
Extended Data Fig. 6 Using increasing numbers of guides modestly but variably increases target gene inactivation.
a-b. Mean tumour size given a log-normal distribution for tumours with arrays containing 1-4 guides targeting the tumour suppressor genes Rbm10 (a) or Setd2 (b) in KT;H11LSL-Cas12a mice, relative to control tumours with inert arrays (median of all arrays containing only Safe or NT guides). c-d. Number of tumours with arrays containing 1 to 4 guides targeting the essential genes Pcna (c) or Rpa3 (d), relative to control tumours with inert arrays. e-g. Mean tumour size given a log-normal distribution for tumours with arrays containing 1 to 4 guides targeting the essential genes Rps9 (e), Pcna (f) or Rpa3 (g), relative to control tumours with inert arrays. h. Mean tumour size given a log-normal distribution for tumours with arrays containing 1 to 4 guides targeting the tumour suppressor gene Rb1 in KT mice (lacking Cas12a), relative to control tumours with inert arrays. All comparisons are not significant. i. Number of tumours with arrays containing 1 to 4 guides targeting the essential gene Rps9 in KT mice, relative to control tumours with inert arrays. All comparisons are not significant. Additional data for arrays in KT mice are shown in Supplementary Table 4. All data (a-i) are medians with 95% confidence intervals based on bootstrap resampling with 1,000 iterations. Empirical P values are calculated by comparing bootstrapped metrics between groups, using the proportion of cases where one group’s values exceed the other’s, with two-sided P values derived by doubling the one-sided values.
Extended Data Fig. 7 Targeting synthetic lethal gene pairs with increasing numbers of guides only modestly increases efficiency, and inactivation of the predicted synthetic lethal paralog gene pair Gsk3a and Gsk3b synergistically promotes tumour growth.
a. Mean tumour size given a log-normal distribution for tumours with arrays containing 0 to 2 guides each targeting the synthetic lethal genes Ccnl1 and Ccnl2, relative to control tumours with inert arrays (median of all arrays containing only Safe or NT guides). b. Number of tumours with arrays containing 0 to 2 guides each targeting the synthetic lethal genes Ccnl1 and Ccnl2 in KT mice (lacking Cas12a), relative to control tumours with inert arrays. All comparisons are not significant. Additional data for arrays in KT mice are shown in Supplementary Table 4. c-d. Number of tumours (c) and mean tumour size given a log-normal distribution (d) for tumours with arrays containing 0-2 guides each targeting the synthetic lethal genes G3bp1 and G3bp2, relative to control tumours with inert arrays. e-f. Number of tumours (e) and mean tumour size given a log-normal distribution (f) for tumours with arrays containing 0 to 2 guides each targeting the predicted synthetic lethal genes Gsk3a and Gsk3b, relative to control tumours with inert arrays. All data (a-f) are medians with 95% confidence intervals based on bootstrap resampling with 1,000 iterations. Empirical P values are calculated by comparing bootstrapped metrics between groups, using the proportion of cases where one group’s values exceed the other’s, with two-sided P values derived by doubling the one-sided values.
Extended Data Fig. 8 Later guide positions within a Cas12a crRNA array have slightly decreased efficiency when targeting essential genes.
a-c. Mean tumour size given a log-normal distribution for tumours with arrays containing 1 guide targeting the tumour suppressor genes Rb1 (a), Rbm10 (b) or Setd2 (c) in positions 1 through 4 in KT (lacking Cas12a; a) or KT;H11LSL-Cas12a mice (b-c), relative to control tumours with inert arrays (median of all arrays containing only Safe or NT guides). All comparisons for KT mice (a) are not significant. Additional data for arrays in KT mice are shown in Supplementary Table 4. d. Mean tumour size given a log-normal distribution for tumours with arrays containing 1 guide targeting the essential gene Rps9 in positions 1 through 4, relative to control tumours with inert arrays. e. Number of tumours with arrays containing 1 guide targeting the essential gene Rps9 in positions 1 through 4 in KT mice (lacking Cas12a), relative to control tumours with inert arrays. All comparisons are not significant. f-g. Number of tumours (f) and mean tumour size given a log-normal distribution (g) for tumours with arrays containing 1 guide targeting the essential gene Pcna in positions 1 through 4, relative to control tumours with inert arrays. h-i. Number of tumours (h) and mean tumour size given a log-normal distribution (i) for tumours with arrays containing 1 guide targeting the essential gene Rpa3 in positions 1 through 4, relative to control tumours with inert arrays. All data (a-i) are medians with 95% confidence intervals based on bootstrap resampling with 1,000 iterations. Empirical P values are calculated by comparing bootstrapped metrics between groups, using the proportion of cases where one group’s values exceed the other’s, with two-sided P values derived by doubling the one-sided values.
Extended Data Fig. 9 Calculation of fitness effects uncovers positive and negative epistasis.
a. Lung weights from mice of the indicated genotypes 7 weeks after transduction with Lenti-U6BC-crTSGTriple-Cre. Each dot represents a mouse. Means +/- standard deviations are indicated. Data from KT;H11LSL-Cas12a and KT;H11Cas12a/WT mice are merged (for all analyses). b. Definitions of key fitness and epistasis metrics involving single and double knockout (KO) genotypes. c. Fitness values for all tumours containing arrays designed to create single, double or triple KOs, as measured in KT mice (lacking Cas12a and therefore with no actual KOs), shown as violin and box plots (median, interquartile range, and whiskers showing values within 1.5 x interquartile range, based on bootstrap resampling with 1,000 iterations). A two-sample rank-sum test was used to compare the fitness of tumours with different array categories. d-e. Observed fitness values for single and double KO tumour genotypes involving the gene pairs Rb1-Trp53 (d) and Arid1a-Mga (e), compared to what would be expected if the double KO fitness were merely the addition of the constituent single KO fitness values. Empirical P values are calculated by comparing bootstrapped tumour metrics between groups, using the proportion of cases where one group’s values exceed the other’s, with two-sided P values derived by doubling the one-sided values. Medians with 95% confidence intervals are shown based on bootstrap resampling with 1,000 iterations. Complete fitness and epistasis data for all arrays are shown in Supplementary Table 5.
Extended Data Fig. 10 Certain advantageous multi-KO tumour genotypes have limited accessibility of fitness trajectories.
a. Definitions of key fitness and epistasis metrics involving triple KO genotypes. b-c. Observed versus expected fitness values of all tumours containing arrays designed to create single, double or triple KOs, as measured in KT mice (lacking Cas12a and therefore with no actual KOs), where the expected values are calculated based on single KO effects only (b) or single KO and two-way epistasis effects (c). Medians with 95% confidence intervals are shown based on bootstrap resampling with 1,000 iterations. Dashed lines are identity lines. d-e. Fitness trajectories for all possible single, double and triple KO tumour genotypes involving Trp53, Keap1 and Mga (d), and Arid1a, Rb1 and Mga (e). Line colours indicate the successive fitness impact of inactivation of the indicated gene. 95% confidence intervals are shown for each genotype. Medians with 95% confidence intervals are shown based on bootstrap resampling with 1,000 iterations. Complete fitness and epistasis data for all arrays are shown in Supplementary Table 5.
Supplementary information
Supplementary Tables 1–5 (download XLSX )
Supplementary Table 1: guide and full pre-crRNA array sequences. Arrays for each experiment were synthesized as pools containing every array listed. Each array contains BsmBI restriction enzyme sites on either end for cloning into the vector backbone and three or four guides (spacers) flanked by direct repeat sequences. Arrays are shown for targeting Nf1, Rasa1 and Pten to generate oncogene-negative lung adenocarcinoma (a); Rb1, Trp53 and Rbl2 to generate SCLC (b); Trp53, Cdkn2a and Smad4 to generate PDAC (c); Eml4, Alk and Trp53 (d); a variety of TSGs, essential genes and synthetic lethal gene pairs to assess Cas12a efficiency (e); and single-, double- and triple-knockout combinations of Arid1a, Keap1, Lkb1, Mga, Rb1, Rbm10, Tet2, Trp53 and Tsc2 (f). Supplementary Table 2: primer sequences for indel analysis of target sites for Cas12a-mediated cleavage. Forward and reverse primers are listed for each site for each of the three targeted positions for each gene. Supplementary Table 3: analysis of potential Cas12a-mediated cutting at top predicted off-target sites. a, The percentage of sequencing reads for each off-target site with insertions or deletions at the predicted cut site. b, The top predicted off-target sites for each guide. c, Primers used to amplify each predicted off-target site for sequencing. Supplementary Table 4: metrics for tumours with arrays to measure Cas12a efficiency. All calculated metrics for tumours with each array, including the mean tumour size assuming a log-normal distribution, 95th percentile tumour size and the total tumour number (all relative to tumours with inert arrays), as well as associated P values. a, All arrays with the same guides combined (regardless of guide order/position within the array) to assess the effects of guide number. b, Arrays listed individually for each order of guides to assess the effects of guide position. c, Arrays with 1–4 guides targeting safe-harbour intergenic regions (safe) or non-targeting (NT) guides. Supplementary Table 5: metrics for tumours with arrays to measure TSG epistasis. a, Fitness metrics for tumours with each array. b, Fitness and epistasis metrics for tumours with arrays targeting all two-gene combinations of TSGs. c, Fitness and epistasis metrics for all three-gene combinations of TSGs.
Source Data Supplementary Figs. 1 and 5 (download PDF )
Uncropped scans of western blots shown in Supplementary Fig. 1c (left panel) and Supplementary Fig. 1c (right panel) and uncropped scan of gel shown in Supplementary Fig. 5e.
Source data
Source Data Fig. 1 (download PDF )
Uncropped scans of western blots shown in Fig. 1c, Fig. 1e (left panel) and Fig. 1e (right panel).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hebert, J.D., Xu, H., Tang, Y.J. et al. Efficient and multiplexed somatic genome editing with Cas12a mice. Nat. Biomed. Eng 9, 1982–1997 (2025). https://doi.org/10.1038/s41551-025-01407-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41551-025-01407-7
This article is cited by
-
Overcoming barriers in CAR-NK immunotherapy: CRISPR-Driven advances in checkpoint editing and allogeneic design
Functional & Integrative Genomics (2026)
-
Methods and applications of in vivo CRISPR screening
Nature Reviews Genetics (2025)
-
Cas12a mice enable the rapid generation of highly complex tumour genotypes
Nature Reviews Cancer (2025)
-
SWITCHER, a CRISPR-inducible floxed wild-type Cre regulating CRISPR activity
Communications Biology (2025)