Abstract
Mitochondria export Ca2+ via Na+/Ca2+ exchange machinery (mito-NCX) to regulate intracellular Ca2+ signalling and mitochondrial Ca2+ homeostasis. TMEM65 has recently been implicated as essential for mito-NCX, but its mechanisms and roles remain unclear. Here we show that TMEM65 depletion severely impairs mito-NCX. TMEM65 is highly expressed in the heart and brain but absent in the liver, correlating with mito-NCX activity in these tissues. Biochemical and functional analyses reveal that TMEM65 forms a homodimer, containing plausible ion-coordinating residues critical for function. Heterologous expression of TMEM65 induces Na+/Ca2+ exchange in cells lacking native mito-NCX activity. Moreover, purified, liposome-reconstituted TMEM65 exhibits key mito-NCX features. We further identify the binding site for CGP-37157, a potent, widely used mito-NCX inhibitor. Finally, TMEM65 deletion elevates mitochondrial Ca2+ and primes mitochondria to permeability transition. These findings firmly establish TMEM65 as the protein mediating mito-NCX, offering a new therapeutic target for diseases associated with mitochondrial Ca2+ dysregulation.
This is a preview of subscription content, access via your institution
Access options






Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the main text, Extended Data Figures and Supplementary Information. The raw mass spectrometry data have been deposited in the PRIDE repository under accession number PXD064897. All unique materials generated in this study are available upon request without restriction. Source data files, including unprocessed scans of gels and blots, as well as numerical data giving rise to all bar charts, are provided with this paper. Source data are provided with this paper.
References
Gunter, T. E., Buntinas, L., Sparagna, G., Eliseev, R. & Gunter, K. Mitochondrial calcium transport: mechanisms and functions. Cell Calcium 28, 285–296 (2000).
Bernardi, P. Mitochondrial transport of cations: channels, exchangers, and permeability transition. Physiol. Rev. 79, 1127–1155 (1999).
Garbincius, J. F. & Elrod, J. W. Mitochondrial calcium exchange in physiology and disease. Physiol. Rev. 102, 893–992 (2022).
Giorgi, C., Marchi, S. & Pinton, P. The machineries, regulation and cellular functions of mitochondrial calcium. Nat. Rev. Mol. Cell Biol. 19, 713–730 (2018).
Kamer, K. J. & Mootha, V. K. The molecular era of the mitochondrial calcium uniporter. Nat. Rev. Mol. Cell Biol. 16, 545–553 (2015).
Rizzuto, R., De Stefani, D., Raffaello, A. & Mammucari, C. Mitochondria as sensors and regulators of calcium signalling. Nat. Rev. Mol. Cell Biol. 13, 566–578 (2012).
Konig, T. et al. The m-AAA protease associated with neurodegeneration limits MCU activity in mitochondria. Mol. Cell 64, 148–162 (2016).
Panov, A. V. et al. Early mitochondrial calcium defects in Huntington’s disease are a direct effect of polyglutamines. Nat. Neurosci. 5, 731–736 (2002).
Jadiya, P. et al. Impaired mitochondrial calcium efflux contributes to disease progression in models of Alzheimer’s disease. Nat. Commun. 10, 3885 (2019).
Liu, T., Yang, N., Sidor, A. & O’Rourke, B. MCU overexpression rescues inotropy and reverses heart failure by reducing SR Ca(2+) leak. Circ. Res. 128, 1191–1204 (2021).
Santulli, G., Xie, W., Reiken, S. R. & Marks, A. R. Mitochondrial calcium overload is a key determinant in heart failure. Proc. Natl Acad. Sci. USA 112, 11389–11394 (2015).
Luongo, T. S. et al. The mitochondrial Na+/Ca2+ exchanger is essential for Ca2+ homeostasis and viability. Nature 545, 93–97 (2017).
Marchi, S., Giorgi, C., Galluzzi, L. & Pinton, P. Ca2+ fluxes and cancer. Mol. Cell 78, 1055–1069 (2020).
Chakraborty, P. K. et al. MICU1 drives glycolysis and chemoresistance in ovarian cancer. Nat. Commun. 8, 14634 (2017).
Tosatto, A. et al. The mitochondrial calcium uniporter regulates breast cancer progression via HIF-1α. EMBO Mol. Med. 8, 569–585 (2016).
Luongo, T. S. et al. The mitochondrial calcium uniporter matches energetic supply with cardiac workload during stress and modulates permeability transition. Cell Rep. 12, 23–34 (2015).
Kwong, J. Q. et al. The mitochondrial calcium uniporter selectively matches metabolic output to acute contractile stress in the heart. Cell Rep. 12, 15–22 (2015).
Murphy, E. & Steenbergen, C. Mechanisms underlying acute protection from cardiac ischemia-reperfusion injury. Physiol. Rev. 88, 581–609 (2008).
Tsai, C. W. et al. Evidence supporting the MICU1 occlusion mechanism and against the potentiation model in the mitochondrial calcium uniporter complex. Proc. Natl Acad. Sci. USA 120, e2217665120 (2023).
Fan, M. et al. Structure and mechanism of the mitochondrial Ca2+ uniporter holocomplex. Nature 582, 129–133 (2020).
Wang, Y. et al. Structural mechanism of EMRE-dependent gating of the human mitochondrial calcium uniporter. Cell 177, 1252–1261 e1213 (2019).
Baradaran, R., Wang, C., Siliciano, A. F. & Long, S. B. Cryo-EM structures of fungal and metazoan mitochondrial calcium uniporters. Nature 559, 580–584 (2018).
Tsai, M. F. et al. Dual functions of a small regulatory subunit in the mitochondrial calcium uniporter complex. eLife 5, e15545 (2016).
Csordas, G. et al. MICU1 controls both the threshold and cooperative activation of the mitochondrial Ca2+ uniporter. Cell Metab. 17, 976–987 (2013).
Mallilankaraman, K. et al. MICU1 is an essential gatekeeper for MCU-mediated mitochondrial Ca2+ uptake that regulates cell survival. Cell 151, 630–644 (2012).
De Stefani, D., Raffaello, A., Teardo, E., Szabo, I. & Rizzuto, R. A forty-kilodalton protein of the inner membrane is the mitochondrial calcium uniporter. Nature 476, 336–340 (2011).
Baughman, J. M. et al. Integrative genomics identifies MCU as an essential component of the mitochondrial calcium uniporter. Nature 476, 341–345 (2011).
Perocchi, F. et al. MICU1 encodes a mitochondrial EF hand protein required for Ca2+ uptake. Nature 467, 291–296 (2010).
Sancak, Y. et al. EMRE is an essential component of the mitochondrial calcium uniporter complex. Science 342, 1379–1382 (2013).
Kamer, K. J., Grabarek, Z. & Mootha, V. K. High-affinity cooperative Ca2+ binding by MICU1-MICU2 serves as an on-off switch for the uniporter. EMBO Rep. 18, 1397–1411 (2017).
Patron, M. et al. MICU1 and MICU2 finely tune the mitochondrial Ca2+ uniporter by exerting opposite effects on MCU activity. Mol. Cell 53, 726–737 (2014).
Yoo, J. et al. Cryo-EM structure of a mitochondrial calcium uniporter. Science 361, 506–511 (2018).
Payne, R., Hoff, H., Roskowski, A. & Foskett, J. K. MICU2 restricts spatial crosstalk between InsP3R and MCU channels by regulating threshold and gain of MICU1-mediated inhibition and activation of MCU. Cell Rep. 21, 3141–3154 (2017).
Tsai, C. W. et al. Mechanisms and significance of tissue-specific MICU regulation of the mitochondrial calcium uniporter complex. Mol. Cell 82, 3661–3676 e3668 (2022).
Carafoli, E., Tiozzo, R., Lugli, G., Crovetti, F. & Kratzing, C. The release of calcium from heart mitochondria by sodium. J. Mol. Cell. Cardiol. 6, 361–371 (1974).
Crompton, M., Moser, R., Ludi, H. & Carafoli, E. The interrelations between the transport of sodium and calcium in mitochondria of various mammalian tissues. Eur. J. Biochem. 82, 25–31 (1978).
Crompton, M., Kunzi, M. & Carafoli, E. The calcium-induced and sodium-induced effluxes of calcium from heart mitochondria. Evidence for a sodium-calcium carrier. Eur. J. Biochem. 79, 549–558 (1977).
Fiskum, G. & Lehninger, A. L. Regulated release of Ca2+ from respiring mitochondria by Ca2+/2H+ antiport. J. Biol. Chem. 254, 6236–6239 (1979).
Chiesi, M., Schwaller, R. & Eichenberger, K. Structural dependency of the inhibitory action of benzodiazepines and related compounds on the mitochondrial Na+-Ca2+ exchanger. Biochem. Pharmacol. 37, 4399–4403 (1988).
Vaghy, P. L., Johnson, J. D., Matlib, M. A., Wang, T. & Schwartz, A. Selective inhibition of Na+-induced Ca2+ release from heart mitochondria by diltiazem and certain other Ca2+ antagonist drugs. J. Biol. Chem. 257, 6000–6002 (1982).
Cai, X. & Lytton, J. The cation/Ca2+ exchanger superfamily: phylogenetic analysis and structural implications. Mol. Biol. Evol. 21, 1692–1703 (2004).
Palty, R. et al. NCLX is an essential component of mitochondrial Na+/Ca2+ exchange. Proc. Natl Acad. Sci. USA 107, 436–441 (2010).
Xue, J. et al. Structural mechanisms of the human cardiac sodium-calcium exchanger NCX1. Nat. Commun. 14, 6181 (2023).
Marinelli, F. et al. Sodium recognition by the Na+/Ca2+ exchanger in the outward-facing conformation. Proc. Natl Acad. Sci. USA 111, E5354–E5362 (2014).
Stavsky, A. et al. Aberrant activity of mitochondrial NCLX is linked to impaired synaptic transmission and is associated with mental retardation. Commun. Biol. 4, 666 (2021).
Assali, E. A. et al. NCLX prevents cell death during adrenergic activation of the brown adipose tissue. Nat. Commun. 11, 3347 (2020).
Taha, M. et al. NCLX controls hepatic mitochondrial Ca2+ extrusion and couples hormone-mediated mitochondrial Ca2+ oscillations with gluconeogenesis. Mol. Metab. 87, 101982 (2024).
Emrich, S. M. et al. The mitochondrial sodium/calcium exchanger NCLX (Slc8b1) in B lymphocytes. Cell Calcium 108, 102667 (2022).
Pathak, T. et al. Dichotomous role of the human mitochondrial Na+/Ca2+/Li+ exchanger NCLX in colorectal cancer growth and metastasis. eLife 9, e59686 (2020).
Flicker, D., Sancak, Y., Mick, E., Goldberger, O. & Mootha, V. K. Exploring the in vivo role of the mitochondrial calcium uniporter in brown fat bioenergetics. Cell Rep. 27, 1364–1375 e1365 (2019).
Pan, X. et al. The physiological role of mitochondrial calcium revealed by mice lacking the mitochondrial calcium uniporter. Nat. Cell Biol. 15, 1464–1472 (2013).
Zhang, Y. et al. Loss of TMEM65 causes mitochondrial disease mediated by mitochondrial calcium. Preprint at bioRxiv https://doi.org/10.1101/2022.08.02.502535 (2022).
Vetralla, M. et al. TMEM65-dependent Ca2+ extrusion safeguards mitochondrial homeostasis. Preprint at bioRxiv https://doi.org/10.1101/2023.10.10.561661 (2023).
Garbincius, J. F. et al. TMEM65 regulates NCLX-dependent mitochondrial calcium efflux. Preprint at bioRxiv https://doi.org/10.1101/2023.10.06.561062 (2023).
Almagro Armenteros, J. J. et al. Detecting sequence signals in targeting peptides using deep learning. Life Sci. Alliance 2, e201900429 (2019).
Nishimura, N., Gotoh, T., Oike, Y. & Yano, M. TMEM65 is a mitochondrial inner-membrane protein. PeerJ 2, e349 (2014).
Schnaitman, C. & Greenawalt, J. W. Enzymatic properties of the inner and outer membranes of rat liver mitochondria. J. Cell Biol. 38, 158–175 (1968).
Yernool, D., Boudker, O., Folta-Stogniew, E. & Gouaux, E. Trimeric subunit stoichiometry of the glutamate transporters from Bacillus caldotenax and Bacillus stearothermophilus. Biochemistry 42, 12981–12988 (2003).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Wang, C., Baradaran, R. & Long, S. B. Structure and reconstitution of an MCU-EMRE mitochondrial Ca2+ uniporter complex. J. Mol. Biol. 432, 5632–5648 (2020).
Hockerman, G. H., Peterson, B. Z., Johnson, B. D. & Catterall, W. A. Molecular determinants of drug binding and action on L-type calcium channels. Annu. Rev. Pharmacol. Toxicol. 37, 361–396 (1997).
Zhao, Y. et al. Molecular basis for ligand modulation of a mammalian voltage-gated Ca2+ channel. Cell 177, 1495–1506 e1412 (2019).
Ashrafi, G., de Juan-Sanz, J., Farrell, R. J. & Ryan, T. A. Molecular tuning of the axonal mitochondrial Ca2+ uniporter ensures metabolic flexibility of neurotransmission. Neuron 105, 678–687 e675 (2020).
Duchen, M. R. Mitochondria and calcium: from cell signalling to cell death. J. Physiol. 529, 57–68 (2000).
Andersen, C. G., Bavnhoj, L. & Pedersen, B. P. May the proton motive force be with you: a plant transporter review. Curr. Opin. Struct. Biol. 79, 102535 (2023).
Waight, A. B. et al. Structural basis for alternating access of a eukaryotic calcium/proton exchanger. Nature 499, 107–110 (2013).
Nishizawa, T. et al. Structural basis for the counter-transport mechanism of a H+/Ca2+ exchanger. Science 341, 168–172 (2013).
MacKenzie, D., Arendt, A., Hargrave, P., McDowell, J. H. & Molday, R. S. Localization of binding sites for carboxyl terminal specific anti-rhodopsin monoclonal antibodies using synthetic peptides. Biochemistry 23, 6544–6549 (1984).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Lamboley, C. R. et al. New method for determining total calcium content in tissue applied to skeletal muscle with and without calsequestrin. J. Gen. Physiol. 145, 127–153 (2015).
Acknowledgements
This study was supported by the NIH under award numbers R01-GM144485 (M.-F.T.), R35-GM151958 (M.-F.T.) and R35-GM153424 (L.F.). J.L.Z. is supported by the National Science Foundation Graduate Research Fellowship. Mass spectrometry work was supported by the NIH under Award Number S10OD030441 (to S.-L.X.) and by the Carnegie endowment fund for the Carnegie Mass Spectrometry Facility.
Author information
Authors and Affiliations
Contributions
M.-F.T. and L.F. conceptualized and supervised the overall project. J.L.Z., Y.-C.C., P.-H.L., H.-I.Y., C.-W.T., Y.-L.H., T.-Y.L., I.-C.L., N.F., Y.X., B.R., H.-M.S. and Y.-C.T. performed experiments. J.-L.Z., Y.-C.C., P.-H.L., H.-I.Y., C.-W.T., Y.-L.H. and T.-Y.L. analysed data and prepared figures. M.-F.T. and L.F. wrote the paper with assistance from J.L.Z. and Y.-C.C. and input from all authors. A.V.R. and S.-L.X. carried out mass spectrometry.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks Fabiana Perrochi, Mohamed Trebak, Weiwei Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 IMM localization of TMEM65.
a, Alkaline carbonate extraction assay. Mitochondria isolated from WT HEK cells were subjected to carbonate treatment under alkaline (pH 10.5) or control (pH 7.5) conditions for 30 min at 4 °C. After ultracentrifugation, the membrane pellet (M), which contains integral membrane proteins from the IMM and the outer mitochondrial membrane, and the supernatant (S), which contains soluble and peripheral membrane proteins, were analysed by Western blot. MCU (a known integral membrane protein), TMEM65, and cytochrome C (Cyt C, a peripheral membrane protein) were detected using specific antibodies against these proteins. At pH 10.5, TMEM65, like MCU, remained in the membrane fraction, indicating that TMEM65 is an integral membrane protein. In contrast, cytochrome C shifted from the membrane fraction at pH 7.5 to the supernatant following alkaline extraction, consistent with its known property as a peripheral membrane protein. b, Digitonin extraction assay. Isolated mitochondria from WT HEK cells were incubated with or without 0.1% digitonin for 30 min at 4 °C, a treatment that dissolves the outer mitochondrial membrane while leaving the IMM intact. This process generates submitochondrial vesicles, known as mitoplasts, which retain matrix and IMM-associated proteins. Mitoplasts were pelleted and analysed by Western blot using antibodies against VDAC (an outer membrane protein), MCU (an IMM protein), and TMEM65. The bar chart depicts the fraction of protein not extracted by digitonin, calculated as the ratio of Western blot signals in digitonin-treated samples to those in untreated controls. VDAC is efficiently extracted by digitonin, consistent with its localization in the outer membrane. TMEM65, like MCU, resists digitonin extraction, indicating that it is either an IMM or matrix protein. These results, together with the findings from (a) that TMEM65 is an integral membrane protein, lead to the conclusion that TMEM65 is an IMM protein. Data in the bar chart are presented as mean ± SEM. Statistical analysis was performed using unpaired, two-tailed t-test. All experiments in this figure were performed with four independent biological repeats. Molecular weight marker unit: kDa.
Extended Data Fig. 2 Mitochondrial Ca2+ uptake in TMEM65-expressing cells.
a-b, No effect of TMEM65 depletion on mitochondrial Ca2+ uptake. Mitochondrial Ca2+ uptake was measured in digitonin-permeabilized WT, TMEM65-KD1, and TMEM65-KO1 HEK293 cells (the KO line was described in Fig. 5). Representative Ca2+ flux traces (a) and quantified uptake rates (b) demonstrate that TMEM65 depletion does not alter mitochondrial Ca2+ uptake. These results confirm that TMEM65 contributes specifically to mito-NCX without affecting other mitochondrial Ca2+ transport pathways. Uptake rates (AU/s) were normalized to the amplitude of fluorescence increase upon Ca2+ addition, yielding normalized Ca2+ uptake rates (s−1) shown in panel b. Independent biological replicates: WT = 9; TMEM65-KD = 6; TMEM65-KO = 5. c, No effect of TMEM65 expression on mitochondrial Ca2+ uptake in Sf9 cells. Mitochondrial Ca2+ uptake was measured in permeabilized Sf9 cells with or without human TMEM65 expression. Representative Ca2+ flux traces (left) and quantified uptake rates (right) demonstrate that TMEM65 expression does not alter mitochondrial Ca2+ uptake. In contrast, hME expression enhances Ca2+ uptake, consistent with the uniporter serving as the primary pathway for Ca2+ entry into mitochondria. Independent biological replicates: No TMEM65 = 5; TMEM65 = 6; hME+TMEM65 = 6. d, No contribution of TMEM65 to Ca2+ influx. Digitonin-permeabilized Sf9 cells were treated with Ru360 to inhibit the uniporter. Following the addition of Ca2+, no mitochondrial Ca2+ uptake was observed regardless of TMEM65 expression. These results indicate that TMEM65 does not mediate Ca2+ influx into mitochondria. Each experiment, repeated 3 times independently, led to similar results. Data in this figure are presented as mean ± SEM. Statistical analysis was performed using unpaired, two-tailed t-test.
Extended Data Fig. 3 Mito-NCX activity across mouse tissues.
a, Expression of TMEM65 relative to MCU. Western blot analysis of MCU levels in various mouse tissues is shown on the left. Bar charts show quantification of MCU levels normalized to Tim23 (middle) and the ratio of TMEM65 (Western blot data presented in Fig. 1e) to MCU (right), both normalized to Tim23. Molecular weight marker unit: kDa. b, Mito-NCX and mitochondrial Ca2+ uptake in mouse tissues. Calcium green-5N fluorescence was used to measure external Ca2+ in isolated mitochondria. The addition of 20 mM NaCl caused a sharp signal drop (reason unclear), followed by Ca2+ efflux mediated by mito-NCX. Hepatic mitochondria exhibited no mito-NCX activity, as indicated by the lack of response to CGP-37157. Bar chart (right) compares the ratio of mito-NCX rate to Ca2+ uptake rate across tissues, correlating with the TMEM65 to MCU expression ratio (panel a, right). Data are presented as mean ± SEM. Statistical analysis was performed using unpaired, two-tailed t-test. All experiments in this figure were performed with 5 independent biological replicates.
Extended Data Fig. 4 Lack of evidence for TMEM65-NCLX interactions or regulation.
a, BN-PAGE analysis of NCLX. C-terminally 1D4-tagged NCLX was expressed in TMEM65-KD1 HEK cells with or without TMEM65 (Flag-tagged) co-expression and analysed by BN-PAGE followed by Western blot using anti-1D4 antibody (top). C-terminally 1D4-tagged MCU, expressed in MICU1/MCU/EMRE-KO cells, served as a control. The migration pattern of NCLX remained unchanged in the presence of TMEM65, indicating no stable complex formation between these two proteins. The NCLX band migrated between the 148- and 480-kDa molecular weight markers, suggesting that NCLX, like MCU, forms oligomers. The three Western blot images (bottom) confirmed expression of MCU, NCLX, and TMEM65, detected using anti-1D4, anti-1D4, and anti-TMEM65 antibodies, respectively. Three independent biological replicates were performed, leading to similar results. b, No effect of tagging on NCLX migration. BN-PAGE was performed using untagged NCLX expressed with or without untagged TMEM65 in TMEM65-KO cells (top). Western blot with anti-NCLX antibody showed no change in NCLX migration, suggesting that the absence of NCLX-TMEM65 interactions observed in panel a is not due to interference from 1D4 or FLAG tags. The two Western blot images (bottom) confirmed expression of NCLX and TMEM65, detected with anti-NCLX and anti-TMEM65 antibodies, respectively. Three independent biological replicates were performed, leading to similar results. c, FSEC analysis of TMEM65. GFP-tagged TMEM65 was expressed in HEK293 cells with or without NCLX co-expression. Detergent solubilized whole-cell lysate (left) and purified TMEM65 protein (right) were analysed by FSEC. The elution profile of TMEM65-GFP remained unchanged in the presence of NCLX, suggesting no interactions between these two proteins. d, Lack of NCLX enhancement of TMEM65 function. Overexpression (OE) of C-terminally 1D4-tagged NCLX in WT or TMEM65 (also 1D4-tagged)-OE HEK cells did not enhance TMEM65-induced mito-NCX activity, suggesting that NCLX does not act as a TMEM65 activator. Western blots using anti-1D4 antibody confirmed expression of TMEM65 and NCLX, both 1D4-tagged. Independent biological replicates: WT(Control) = 4; WT(NCLX OE) = 4; TMEM65 OE(Control) = 5; TMEM65 OE(NCLX OE) = 5. e, Functional independence of TMEM65 and NCLX in Sf9 cells. Human TMEM65 was expressed in Sf9 cells with or without co-expression of 1D4-tagged human NCLX. Mitochondrial Ca2+ flux measurements showed that TMEM65-mediated mito-NCX was unaffected by NCLX expression, again supporting the lack of functional regulation between these two proteins. Western blot confirmed the expression of NCLX and TMEM65, detected using anti-1D4 and anti-TMEM65 antibodies, respectively. Independent biological replicates: TMEM65 = 4; TMEM65 + NCLX = 4. Data are presented as mean ± SEM. Statistical analysis was performed using unpaired, two-tailed t-test. Molecular weight marker unit: kDa.
Extended Data Fig. 5 Structural predictions of TMEM65.
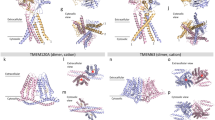
Structures of TMEM65 oligomers predicted by AlphaFold 3. The models for TMEM65 monomer, dimer, trimer and tetramer are shown in ribbon diagrams and coloured by the predicted local distance difference test (pLDDT) score. Higher pLDDT scores indicate higher confidence levels. The Predicted Aligned Error plots show expected positional error between residue pairs. The predicted boundaries of the IMM are highlighted in grey.
Extended Data Fig. 6 Multiple sequence alignment of TMEM65.
Putative ion binding site residues (D132, N163, S166, D167) and CGP-37157 binding site residues (F125, F128, L171, A174) are shaded in red and blue, respectively. MTS: mitochondrial targeting sequence. TMH: transmembrane helix.
Extended Data Fig. 7 Impact of TMEM65 mutations on mito-NCX function.
WT or mutant TMEM65 constructs were expressed to comparable levels in TMEM65-KD HEK cells as shown in Western blot images. Control: untransfected cells. Data are presented as mean ± SEM. Statistical analysis was performed using unpaired, two-tailed t-test. Actin: loading control. Molecular weight marker unit: kDa. All conditions (WT, Control, E144A, S154A, and T155A) were analysed with 3 independent biological replicates.
Extended Data Fig. 8 TMEM65 fluorescence measurements.
a, Size-exclusion chromatography of purified TMEM65. TMEM65 eluted at 11.9 mL on a Superdex 200 column. Image: SDS–PAGE and Coomassie staining of the peak fraction. b-d, Fluorescence changes of WT (red) and D132A (black) TMEM65 upon addition of cations. Panels b and d display Trp fluorescence changes with 20 mM Na+ and 20 µM Ca2+, respectively, while panel c shows Tyr fluorescence changes with 20 mM Na+. Notably, Na+ and Ca2+ induce opposite changes in Trp fluorescence, suggesting that these cations drive the conformational distribution of TMEM65 towards two distinct conformations, possibly matrix- versus IMS-facing conformations. We used 30% more D132A TMEM65 (13 µg per experiment) than WT (10 µg) to create a more stringent test of our hypothesis that D132A would diminish TMEM65 fluorescence changes. Despite this, WT TMEM65 still exhibits significantly larger fluorescence changes upon cation addition. Each data point represents results from an independent protein preparation. Statistical analysis was performed using unpaired, two-tailed t-test. We note that a titration experiment to determine Na+ or Ca2+ affinity of TMEM65—while impractical in the current study due to the modest fluorescence changes observed even at high Na+ or Ca2+ concentrations—may be feasible in future studies in which Trp residues are engineered to positions expected to undergo more substantial changes in their chemical environments upon ion binding. All data in this figure are presented as mean ± SEM. Statistical analysis was performed using unpaired, two-tailed t-test. All experiments were performed with 3 independent biological replicates.
Extended Data Fig. 9 Purification of TMEM65 for liposome reconstitution.
a, Size-exclusion chromatography trace of GFP-TMEM65 purified in DM for reconstitution. b, Analysis of TMEM65 incorporation into proteoliposomes. Liposomes containing GFP-tagged TMEM65 were isolated by ultracentrifugation and analysed by SDS–PAGE under reducing conditions. The protein was visualized by Coomassie stain (left) and GFP fluorescence imaging (right). A single protein band was detected by Coomassie staining, which was confirmed to be TMEM65-GFP based on its GFP fluorescence. No protein was detected at 50 kDa, the expected molecular weight of NCLX. These results demonstrate that TMEM65 was successfully incorporated as the predominant protein species in our reconstituted liposomes. c, Western blot analysis of TMEM65 proteoliposomes. Purified NCLX, purified TMEM65-GFP, and TMEM65-GFP proteoliposomes were analysed by Western blot under reducing conditions using antibodies against TMEM65 and NCLX. Anti-TMEM65 antibody detected TMEM65-GFP in both the purified protein sample and reconstituted proteoliposomes, while anti-NCLX antibody detected NCLX only in the purified NCLX control sample. These results confirm that NCLX does not co-purify with TMEM65 and that the reconstituted proteoliposomes are free of NCLX contamination. All experiments in this figure were performed with three independent biological replicates, leading to similar results. Molecular weight marker unit: kDa.
Extended Data Fig. 10 Mutations at the putative CGP-37157 binding site in TMEM65.
a, Western blot analysis of TMEM65 mutant expression. Actin serves as a loading control. b, Dose-response relationship for CGP-37157 inhibition of mito-NCX induced by WT TMEM65. Experiments were performed using TMEM65-KD cells transiently expressing WT TMEM65. Data were fitted to a saturating function I([CGP]) = [CGP]/(IC50 + [CGP]), where I represents the percentage of inhibition, [CGP] is the CGP-37157 concentration, and IC50 is the half maximal inhibitory concentration. Independent biological replicates: 1 µM = 4; 2.5 µM = 3; 5 µM = 7; 10 µM = 3. c, Rates of Mito-NCX induced by WT TMEM65 and TMEM65 mutants. Independent biological replicates: WT, F125A, and L171W = 4; all other mutants = 3. Data are presented as mean ± SEM.
Supplementary information
Supplementary Information (download PDF )
Mass spectrometry experiments described in Supplementary Text, Fig. 1 and Table 1.
Source data
Source Data Figs. 1–3 and 5 and Extended Data Figs. 1, 3, 4 and 7–10 (download PDF )
Uncropped scans of blot or gel images for Figs. 1–3 and 5 and Extended Data Figs. 1, 3, 4 and 7–10.
Source Data Figs. 1–6 and Extended Data Figs. 1–4, 7, 8 and 10 (download XLSX )
Source data for Figs. 1–6 and Extended Data Figs. 1–4, 7, 8 and 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, J.L., Chang, YC., Lai, PH. et al. TMEM65 functions as the mitochondrial Na+/Ca2+ exchanger. Nat Cell Biol 27, 1301–1310 (2025). https://doi.org/10.1038/s41556-025-01721-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-025-01721-x
This article is cited by
-
Tmbim5 and Slc8b1 cooperate in tissue-specific mitochondrial calcium regulation in zebrafish
Communications Biology (2026)
-
TMEM65-dependent Ca2+ extrusion safeguards mitochondrial homeostasis
Nature Communications (2025)