Abstract
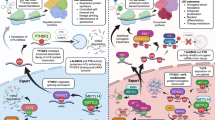
How cancer cells couple metabolic stress sensing to orchestrate specific survival programmes is a key question. Here we show a long non-coding RNA (lncRNA)-guided epitranscriptomic mechanism orchestrating metabolic adaptation by controlling the stability of master stress regulator ATF4. Glucose or glutamine deprivation induces endoplasmic reticulum stress via reactive oxygen species–NRF2-dependent transcription of the lncRNA DAMER. Following its demethylation and nuclear retention by the m6A-eraser ALKBH5, DAMER acts as a scaffold, guiding ALKBH5 to demethylate and stabilize ATF4 mRNA through specific base-pairing. This provides an alternative post-transcriptional pathway for ATF4 upregulation, rewiring asparagine metabolism to promote cancer cell survival under stress. Furthermore, we identified the US FDA-approved drug elbasvir as a potent inhibitor of the DAMER–ALKBH5 interaction. Elbasvir dismantles this adaptive programme, targeting tumour asparagine dependency and exhibiting potent antitumour effects in preclinical models. Our findings reveal a paradigm for lncRNA-guided RNA demethylation that solves a target specificity enigma and offers a strategy targeting metabolic adaptation in cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All RNA sequencing, pull-down sequencing and SHAPE-MAP sequencing data have been deposited in the Gene Expression Omnibus under the accession codes GSE309436, GSE309437, GSE309440 and GSE309724. The human rectum adenocarcinoma data were derived from the TCGA Research Network (http://cancergenome.nih.gov/). The original western blot images have been provided and are publicly available. Source data are provided with this paper. All other data supporting the findings of this study are available.
References
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Martínez-Reyes, I. & Chandel, N. S. Cancer metabolism: looking forward. Nat. Rev. Cancer 21, 669–680 (2021).
Zhang, L. et al. Targets of tumor microenvironment for potential drug development. MedComm Oncol. 3, e68 (2024).
Pavlova, N. N., Zhu, J. & Thompson, C. B. The hallmarks of cancer metabolism: Still emerging. Cell Metab. 34, 355–377 (2022).
Vander Heiden, M. G. & DeBerardinis, R. J. Understanding the intersections between metabolism and cancer biology. Cell 168, 657–669 (2017).
Eagle, H. Nutrition needs of mammalian cells in tissue culture. Science 122, 501–514 (1955).
Jain, M. et al. Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation. Science 336, 1040–1044 (2012).
Hosios, A. M. et al. Amino acids rather than glucose account for the majority of cell mass in proliferating mammalian cells. Dev. Cell 36, 540–549 (2016).
Birsoy, K. et al. Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides. Nature 508, 108–112 (2014).
Yuneva, M. O. et al. The metabolic profile of tumors depends on both the responsible genetic lesion and tissue type. Cell Metab. 15, 157–170 (2012).
Pavlova, N. N. et al. As extracellular glutamine levels decline, asparagine becomes an essential amino acid. Cell Metab. 27, 428–438 (2018).
Krall, A. S. et al. Asparagine couples mitochondrial respiration to ATF4 activity and tumor growth. Cell Metab. 33, 1013–1026 (2021).
Zhao, T., Du, J. & Zeng, H. Interplay between endoplasmic reticulum stress and non-coding RNAs in cancer. J. Hematol. Oncol. 13, 163 (2020).
Zhao, Y. et al. ROS signaling under metabolic stress: cross-talk between AMPK and AKT pathway. Mol. Cancer 16, 79 (2017).
Zielke, S. et al. ATF4 links ER stress with reticulophagy in glioblastoma cells. Autophagy 17, 2432–2448 (2021).
Balsa, E. et al. ER and nutrient stress promote assembly of respiratory chain supercomplexes through the PERK–eIF2α axis. Mol. Cell 74, 877–890 (2019).
Gwinn, D. M. et al. Oncogenic KRAS regulates amino acid homeostasis and asparagine biosynthesis via ATF4 and alters sensitivity to L-asparaginase. Cancer Cell 33, 91–107 (2018).
Magne, L. et al. ATF4 and the integrated stress response are induced by ethanol and cytochrome P450 2E1 in human hepatocytes. J. Hepatol. 54, 729–737 (2011).
Williams, R. T. et al. ZBTB1 regulates asparagine synthesis and leukemia cell response to L-asparaginase. Cell Metab. 31, 852–861 (2020).
Walter, P. & Ron, D. The unfolded protein response: from stress pathway to homeostatic regulation. Science 334, 1081–1086 (2011).
Zhou, Z. et al. Mechanism of RNA modification N6-methyladenosine in human cancer. Mol. Cancer 19, 104 (2020).
Huang, H., Weng, H. & Chen, J. m6A modification in coding and non-coding RNAs: roles and therapeutic implications in cancer. Cancer Cell 37, 270–288 (2020).
Tao, L. et al. Epigenetic regulation in cancer therapy: from mechanisms to clinical advances. MedComm Oncol. 3, e59 (2024).
Chen, X. Y., Zhang, J. & Zhu, J. S. The role of m6A RNA methylation in human cancer. Mol. Cancer 18, 103 (2019).
Liu, H. et al. ALKBH5-mediated m6A demethylation of GLUT4 mRNA promotes glycolysis and resistance to HER2-targeted therapy in breast cancer. Cancer Res. 82, 3974–3986 (2022).
Yu, H. et al. ALKBH5 inhibited cell proliferation and sensitized bladder cancer cells to cisplatin by m6A–CK2α-mediated glycolysis. Mol. Ther. Nucleic Acids 23, 27–41 (2021).
Ahola, S. et al. OMA1-mediated integrated stress response protects against ferroptosis in mitochondrial cardiomyopathy. Cell Metab. 34, 1875–1891 (2022).
Guo, X. et al. Mitochondrial stress is relayed to the cytosol by an OMA1–DELE1–HRI pathway. Nature 579, 427–432 (2020).
Nakamura, A. et al. Inhibition of GCN2 sensitizes ASNS-low cancer cells to asparaginase by disrupting the amino acid response. Proc. Natl Acad. Sci. USA 115, E7776–E7785 (2018).
Sidrauski, C. et al. Pharmacological brake-release of mRNA translation enhances cognitive memory. eLife 2, e00498 (2013).
Smola, M. J., Rice, G. M., Busan, S., Siegfried, N. A. & Weeks, K. M. Selective 2′-hydroxyl acylation analyzed by primer extension and mutational profiling (SHAPE-MaP) for direct, versatile and accurate RNA structure analysis. Nat. Protoc. 10, 1643–1669 (2015).
Siegfried, N. A., Busan, S., Rice, G. M., Nelson, J. A. & Weeks, K. M. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat. Methods 11, 959–965 (2014).
Xia, Z. et al. Epitranscriptomic editing of the RNA N6-methyladenosine modification by dCasRx conjugated methyltransferase and demethylase. Nucleic Acids Res. 49, 7361–7374 (2021).
Nogueira, V. & Hay, N. Molecular pathways: reactive oxygen species homeostasis in cancer cells and implications for cancer therapy. Clin. Cancer Res. 19, 4309–4314 (2013).
Reid, M. A. et al. The B55α subunit of PP2A drives a p53-dependent metabolic adaptation to glutamine deprivation. Mol. Cell 50, 200–211 (2013).
Hast, B. E. et al. Cancer-derived mutations in KEAP1 impair NRF2 degradation but not ubiquitination. Cancer Res. 74, 808–817 (2014).
Berger, A. H. et al. High-throughput phenotyping of lung cancer somatic mutations. Cancer Cell 30, 214–228 (2016).
Jaramillo, M. C. & Zhang, D. D. The emerging role of the Nrf2–Keap1 signaling pathway in cancer. Genes Dev. 27, 2179–2191 (2013).
Romero, R. et al. Keap1 mutation renders lung adenocarcinomas dependent on Slc33a1. Nat. Cancer 1, 589–602 (2020).
Roundtree, I. A. et al. YTHDC1 mediates nuclear export of N6-methyladenosine methylated mRNAs. eLife 6, e31311 (2017).
Komatsu, T. E. et al. Regulatory analysis of effects of hepatitis C virus NS5A polymorphisms on efficacy of elbasvir and grazoprevir. Gastroenterology 152, 586–597 (2017).
Xiao, Z., Dai, Z. & Locasale, J. W. Metabolic landscape of the tumor microenvironment at single cell resolution. Nat. Commun. 10, 3763 (2019).
Lyssiotis, C. A. & Kimmelman, A. C. Metabolic interactions in the tumor microenvironment. Trends Cell Biol. 27, 863–875 (2017).
Elia, I. & Haigis, M. C. Metabolites and the tumour microenvironment: from cellular mechanisms to systemic metabolism. Nat. Metab. 3, 21–32 (2021).
Lin, S. C. & Hardie, D. G. AMPK: sensing glucose as well as cellular energy status. Cell Metab. 27, 299–313 (2018).
González, A., Hall, M. N., Lin, S. C. & Hardie, D. G. AMPK and TOR: the yin and yang of cellular nutrient sensing and growth control. Cell Metab. 31, 472–492 (2020).
Abu-Remaileh, M. et al. Lysosomal metabolomics reveals V-ATPase- and mTOR-dependent regulation of amino acid efflux from lysosomes. Science 358, 807–813 (2017).
Knott, S. R. V. et al. Asparagine bioavailability governs metastasis in a model of breast cancer. Nature 554, 378–381 (2018).
Hinze, L. et al. Exploiting the therapeutic interaction of WNT pathway activation and asparaginase for colorectal cancer therapy. Cancer Discov. 10, 1690–1705 (2020).
Wu, J. et al. Asparagine enhances LCK signalling to potentiate CD8+ T-cell activation and anti-tumour responses. Nat. Cell Biol. 23, 75–86 (2021).
Xiao, S. et al. The RNA N6-methyladenosine modification landscape of human fetal tissues. Nat. Cell Biol. 21, 651–661 (2019).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Chang, G. et al. YTHDF3 induces the translation of m6A-enriched gene transcripts to promote breast cancer brain metastasis. Cancer Cell 38, 857–871 (2020).
Su, R. et al. R-2HG exhibits anti-tumor activity by targeting FTO/m6A/MYC/CEBPA signaling. Cell 172, 90–105 (2018).
He, L. et al. Functions of N6-methyladenosine and its role in cancer. Mol. Cancer 18, 176 (2019).
Deng, L. J. et al. m6A modification: recent advances, anticancer targeted drug discovery and beyond. Mol. Cancer 21, 52 (2022).
Zhou, J. et al. N6-methyladenosine guides mRNA alternative translation during integrated stress response. Mol. Cell 69, 636–647 (2018).
Liu, X. et al. ATOH8 binds SMAD3 to induce cellular senescence and prevent Ras-driven malignant transformation. Proc. Natl Acad. Sci. USA 120, e2208927120 (2023).
Yang, X. et al. m6A-dependent modulation via IGF2BP3/MCM5/Notch axis promotes partial EMT and LUAD metastasis. Adv Sci. 10, e2206744 (2023).
Tian, H. et al. AKT-induced lncRNA VAL promotes EMT-independent metastasis through diminishing Trim16-dependent Vimentin degradation. Nat. Commun. 11, 5127 (2020).
Wu, S. et al. Long non-coding RNA LEISA promotes progression of lung adenocarcinoma via enhancing interaction between STAT3 and IL-6 promoter. Oncogene 40, 3449–3459 (2021).
Busan, S. & Weeks, K. M. Accurate detection of chemical modifications in RNA by mutational profiling (MaP) with ShapeMapper 2. RNA 24, 143–148 (2018).
Pan, X., Fang, Y., Li, X., Yang, Y. & Shen, H. B. RBPsuite: RNA-protein binding sites prediction suite based on deep learning. BMC Genomics 21, 884 (2020).
Cook, K. B., Kazan, H., Zuberi, K., Morris, Q. & Hughes, T. R. RBPDB: a database of RNA-binding specificities. Nucleic Acids Res. 39, D301–D308 (2011).
Agostini, F. et al. catRAPID omics: a web server for large-scale prediction of protein–RNA interactions. Bioinformatics 29, 2928–2930 (2013).
Wang, S. et al. The N6-methyladenosine epitranscriptomic landscape of lung adenocarcinoma. Cancer Discov. 14, 2279–2299 (2024).
Momcilovic, M. et al. The GSK3 signaling axis regulates adaptive glutamine metabolism in lung squamous cell carcinoma. Cancer Cell 33, 905–921 (2018).
Wei, X. et al. METTL3 preferentially enhances non-m6A translation of epigenetic factors and promotes tumourigenesis. Nat. Cell Biol. 24, 1278–1290 (2022).
Acknowledgements
This work was supported by grants from the Noncommunicable Chronic Diseases-National Science and Technology Major Project (grant number 2023ZD0501100 to M.L.), National Natural Science Foundation of China (grant numbers 82488101, 82030088 and 82241231 to M.L.; 82550006 to J.C. and 8240110744 to T.T.), Guangdong Basic and Applied Basic Research Foundation (grant numbers 2023B1111030003 to M.L., 2024B1515040029 and 2023B111020007 to J.C., and 2023A1515010289 and 2022A1515111072 to T.T.), Funding by Science and Technology Projects in Guangzhou (grant number 2024A04J6549 to J.C.), Funding on Guangdong Marine Economy Development Special Project (project number GDNRC [2022]35 to M.L.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. We thank the Facility of Medical Science (Guangzhou Campus), SYSU for its help in equipment use and data analysis and the National Supercomputer Center in Guangzhou for providing high-performance computational resources.
Author information
Authors and Affiliations
Contributions
M.L., J.C., and T.T. conceived and designed the experiments. T.T., X.F., Y. Yang and B. Liu performed the experiments, analysed data and wrote a draft of the paper with the help of J.L., Z.C., S.W., J.Z., X.Y., L.H., Y.Z., Z.G., W.W., J.Z., Y.H., Y. Ye, X.X., J.Y. and R.H. T.T., X.F. and B.L. contributed to the statistical analyses of the data. H.G., J.W., L.X, W.H., B. Li, J.C. and M.L. discussed experiments and edited the manuscript. M.L. and J.C. conceived the ideas, designed and discussed experiments, supervised progress and extensively edited and communicated regarding the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Nutrient deprivation triggers a non-canonical, post-transcriptional ATF4 response dependent on ALKBH5-mediated m6A demethylation.
a, MTT assays of the indicated lung cancer cells under nutrient deficiency. b, A549 xenografts from mice on normal (ND), glucose-deficient (GDD), or glutamine-deficient (QDD) diets, with or without intratumoral ASNase injection (n = 5 mice per group). Left: tumour images; right: tumour weights. Scale bar, 1 cm (left). c, Protein (left) and mRNA (right) levels of ATF4, ASNS, SLCs (SLC3A2, SLC7A1, SLC7A5) under nutrient deficiency. d, ChIP assay showing ATF4 binding to ASNS and SLC promoters under nutrient deficiency. e, Intracellular asparagine (Asn) concentrations in 95D cells with ATF4 overexpression or silencing under nutrient deficiency. f,g, Colony formation assay (f) and Asn concentrations (g) in vector control or ATF4-overexpressing 95D cells with stable ASNS silencing under nutrient deficiency. h, Metabolic map of glycolysis and the TCA cycle for the tracer. i, 13C-labelling patterns from D-Glc-13C6 (4 g/L, 24 h) in 95D, H460, and A549 cells under normal or glutamine-deficient conditions. j, Metabolic flux ratios of Asn(M + 2)/OAA(M + 2) and Cit(M + 4)/OAA(M + 2) in cells from i under normal vs. glutamine-deficient conditions. k, WB analysis of ASNS, CS, and GLUL levels in the indicated cells under normal or glutamine-deficient conditions. l,m, Autophagy analysis in sh-Vector and ATF4-silenced 95D cells under nutrient deficiency, with or without Bafilomycin A1 (Baf-A1, 10 nM). l, WB analysis for ATF4, ASNS, LC3, and p62. m, Autophagic flux assessed by mRFP–GFP-LC3 reporter. Scale bar, 20 μm. n, WB analysis validation of ATF4 and ATG5 shRNA knockdown in 95D cells. o,p, Intracellular Asn concentration (o) and colony formation (p) in indicated 95D cells cultured under control or nutrient deficiency with Baf-A1 or exogenous Asn. Representative images or data summaries are shown for Western blots (c(left),k,l,n), heat map (c(right)), colony formation assays (f,p), xenograft tumours (b(left)), and fluorescence microscopy (m). All experiments were repeated independently three times with similar results. Data in quantitative graphs are presented as mean ± s.d. For the xenograft study (b(right)), n = 5 mice per group; for all other quantitative experiments (a,d,e,g,i,j,o), n = 3 independent biological replicates. P values are calculated by two-tailed unpaired Student’s t-test (b,d(left)) or two-way ANOVA followed by Tukey’s test (a,d(right),e,g,o) or Sidak’s test (i,j). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 2 Nutrient deprivation triggers a non-canonical, post-transcriptional upregulation of ATF4.
a, Luciferase activity from ATF4-Luci or ATF4-uORF2-Luci (uORF2-Luci) reporters in sh-Vector and ALKBH5-silenced 95D cells cultured for 24 h in complete, −Glc, or −Gln DMEM. b, Polysome profile showing the association of ATF4 transcript with polyribosomes versus monoribosomes in sh-Vector and ALKBH5-silenced 95D cells. c, WB analysis of p-GCN2, p-eIF2α, and eIF2α in 95D and H460 cells under nutrient deficiency. d, WB analysis of GCN2, eIF2α and ATF4 levels in 95D sh-Vector, GCN2-silenced or eIF2α-silenced cells under nutrient deficiency. e, WB analysis of GCN2, p-eIF2α, eIF2α and ATF4 levels in 95D sh-Vector and GCN2-silenced cells cultured in glucose- or glutamine-deficient medium for 0, 2, 4, 8, 16, 24, or 48 h. f, WB analysis of GCN2, p-eIF2α, eIF2α, ATF4, ASNS, and SLC3A2 levels in 95D cells treated with Vehicle, GCN2iB, or ISRIB under control, glucose-deficient, glutamine-deficient, or leucine-deficient conditions. g, Asn concentrations in 95D cells treated with GCN2iB, ISRIB, or TUDCA. h, Nuclear run-on assay quantifying nascent ATF4 mRNA transcription rates in 95D cells under nutrient deficiency. i, ATF4 mRNA levels in 95D cells treated with ActD and RNase under nutrient deficiency. j, Asn concentrations in sh-Vector and ALKBH5-silenced 95D cells under nutrient deficiency. Representative images are shown for western blots (c–f). All experiments were repeated independently three times with similar results. Data in quantitative graphs are presented as mean ± s.d. from n = 3 independent biological replicates (a,b,g–j). P values are calculated by two-way ANOVA followed by Tukey’s test (a,b,g–j). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 3 lncRNA DAMER is an essential specificity determinant for ALKBH5-mediated ATF4 demethylation.
a, Global m6A dot blot in 95D and H460 cells under nutrient deficiency. b, WB analysis of ALKBH5 levels in 95D and H460 cells under nutrient deficiency. c, DAMER expression in paired lung cancer (T) and adjacent non-tumorous (N) tissues from TCGA (n = 72). d, WB analysis of ATF4, ASNS, and SLCs (SLC3A2, SLC7A1, SLC7A5) levels in sg-ctrl and DAMER-depleted H460 cells under nutrient deficiency. e, Relative DAMER expression in 95D and H460 sg-ctrl, DAMER-depleted cells. f, MTT assay of sg-ctrl and DAMER-depleted 95D and H460 cells under nutrient deficiency. g, WB analysis of ATF4, ASNS, and SLCs, p-GCN2, eIF2α, p-eIF2α, and HIF-1α levels in xenograft tumour periphery (P) and core (C). h, Relative DAMER expression and metabolite (Glc, Gln, Asn) concentrations in xenograft tumour periphery and core. i, Correlation between metabolite and gene expression levels in xenograft tumours periphery and core (related to Fig. 2i). j, ATP levels in the periphery and core of H460 sg-ctrl and DAMER-KO (KO-1) xenograft tumours. k, CLIP assays for FTO interaction with DAMER or ATF4 mRNA in indicated cells under nutrient deficiency. l, IF-FISH showing co-localization of DAMER and ALKBH5 in 95D cells under nutrient deficiency. Scale bar, 10 μm. m, Left: Predicted secondary structures of DAMER-FL and its fragments using RNAfold. Right: RNA pull-down showing ALKBH5 interaction with DAMER fragments. n, Global m6A levels in 95D sg-ctrl or DAMER-depleted cells. o, Global m6A levels in 95D sg-ctrl or DAMER-KO1 cells transfected with indicated ALKBH5 constructs. p, ATF4 luciferase reporter activity in 95D sg-ctrl or DAMER-KO1 cells after ALKBH5 silencing and rescue with indicated ALKBH5 constructs under nutrient deficiency. q, Polysome profile of ATF4 mRNA association with polyribosomes versus monoribosomes in indicated cells. Representative images are shown for dot blots (a,n,o), western blots (b,d,g,m(right)), and IF-FISH (l). All experiments were repeated independently three times with similar results. Data in quantitative graphs are mean ± s.d. from n = 72 patient pairs for TCGA data (c); n = 5 in sg-ctrl group and n = 3 in DAMER-KO group (h,i); n = 3 independent biological replicates for all other quantitative experiments (e,f,k,p,q). P values are calculated by two-tailed paired Student’s t-test (c,j) or two-way ANOVA followed by Tukey’s test (e,f,p,q), or Sidak’s test (h), or two-tailed unpaired Student’s t-test (k), linear regression (i). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 4 DAMER guides ALKBH5 to demethylate ATF4 mRNA through specific base-pairing.
a, SHAPE reactivity for ATF4 mRNA in vitro, in the presence or absence of DAMER. b, SHAPE-directed structural models of ATF4 mRNA regions. c, SHAPE-directed structural models of DAMER (559–569 nt, ATF4-binding Site 2). d,e, SHAPE reactivity for DAMER-M1 (d(left)), DAMER-M2 (d(right)), and DAMER-M1/2 (e) in vitro, in the presence or absence of ATF4 mRNA. f, RNA-FISH showing co-localization of DAMER (Cy3) and ATF4 mRNA (GFP) in 95D cells under nutrient deficiency. Nuclei were stained with DAPI. Scale bar, 10 μm. g, Top-ranked candidate DAMER-binding RNAs identified by RNA–RNA pull-down followed by RNA sequencing. h, Relative fold change of ATF4, RNF157, RAD51B, ATF4P3, and ATF4P4 in 95D cells under nutrient deficiency. i, RIP assays for ALKBH5 interaction with RNF157 or RAD51B mRNA in 95D cells under nutrient deficiency. j, SELECT assays for m6A levels at potential sites in ATF4 mRNA in Vector and ALKBH5-overexpressing 95D cells. k, SELECT assay for m6A levels at A1131, A1227, and A1599 of ATF4 mRNA in 95D sg-ctrl and DAMER-KO1 cells under nutrient deficiency. l,m, Analysis of ATF4 mRNA m6A levels by m6A-RIP assays (l) and ATF4 mRNA half-life (m) in 95D sg-ctrl, DAMER-KO1, and ALKBH5-sh1 cells overexpressing indicated ATF4 constructs. n, ATF4 mRNA half-life in the indicated 95D cells using a dCasRx-ALKBH5 system under nutrient deficiency. Representative images or tables are shown for RNA-FISH (f), heat map (h) and half-life data (m,n); all experiments were repeated independently three times with similar results. Data in quantitative graphs are presented as mean ± s.d. from n = 3 independent biological replicates (i–l). P values are calculated by two-way ANOVA followed by Tukey’s test (i,l) or Sidak’s test (j), or two-tailed unpaired Student’s t-test (k). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 5 The DAMER–ATF4 axis is activated by upstream ER stress via ROS–NRF2 signalling.
a, Representative transmission electron microscopy images of endoplasmic reticulum morphology in 95D cells treated with or without TUDCA under nutrient deficiency. Scale bar, 1 μm. b, Luciferase assay of NRF2-binding sites in DAMER promoter in 95D and H460 cells under nutrient deficiency. c, MTT cell growth assays of NRF2-overexpressing sg-ctrl and DAMER-KO1 95D cells under nutrient deficiency. d, m6A-RIP analysis of m6A levels on ATF4 mRNA in NRF2-silenced 95D cells. e,f, ATF4 mRNA half-life assessed after ActD treatment (e) and WB analysis of NRF2, ATF4, ASNS, and SLC3A2 levels (f) in NRF2-silenced 95D cells under nutrient deficiency. g, MTT assays of NRF2-silenced 95D cells under nutrient deficiency. h,i, Relative DAMER expression (h) and ATF4 mRNA half-life (i) in Keap1- and NRF2-silenced sg-ctrl and DAMER-KO1 95D cells under nutrient deficiency. j, m6A-RIP analysis of m6A levels on ATF4 mRNA in the indicated 95D cells. k, MTT assays of the indicated 95D cells under nutrient deficiency. l, Flow cytometry analysis of ROS levels in 95D and H460 cells under hypoxia (hypo). m, m6A-RIP analysis of m6A levels on ATF4 mRNA in sg-ctrl or DAMER-KO1 cells treated with NAC under hypoxia. n, Relative fold change of DAMER (left) and ATF4 mRNA (right) in the indicated cells treated with NAC under hypoxia. o, WB analysis of ATF4 levels in 95D sg-ctrl and DAMER-KO1 cells treated with NAC under hypoxia. p, Asn concentrations in 95D sg-ctrl and DAMER-KO1 cells treated with NAC under hypoxia. Representative images or tables are shown for transmission electron microscopy (a), western blots (f,o), half-life data (i) and flow cytometry (l). All experiments were repeated independently three times with similar results. Data in quantitative graphs are presented as mean ± s.d. from n = 3 independent biological replicates (b–e,g,h,j,k,m,n,p). P values are calculated by two-way ANOVA followed by Tukey’s test (b,c,e,g,h,k,m,n,p) or two-tailed unpaired Student’s t-test (d,j). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 6 ALKBH5-mediated demethylation controls DAMER’s nuclear localization and function.
a, DAMER distribution in nuclear and cytoplasmic fractions from 95D sh-Vector, ALKBH5-sh1, and FTO-sh1 cells under nutrient deficiency. b,c, SELECT assay for m6A levels at sites in DAMER in 95D and H460 cells under nutrient deficiency. d, SELECT assay for m6A levels at DAMER sites A755 and A774 in sh-Vector and ALKBH5-silenced 95D cells under nutrient deficiency. e, SELECT assay for m6A levels at DAMER sites A755 and A774 in the indicated cells under nutrient deficiency. f, Nuclear-to-total DAMER fluorescence ratio in 95D sh-Vector and ALKBH5-sh1 cells in control, −Glc, or −Gln medium for 6, 12, and 24 h (quantification of Fig. 5d). g,h, CLIP (g) and RNA pull-down (h) assays of DAMER–ALKBH5 interaction in indicated cells under nutrient deficiency. i, m6A-RIP assays of DAMER (left) and ATF4 mRNA (right) m6A levels in 95D sg-ctrl and DAMER-KO cells overexpressing indicated DAMER constructs. j, ATF4 mRNA half-life in 95D sg-ctrl and DAMER-KO cells overexpressing indicated DAMER constructs under nutrient deficiency, after ActD treatment. k, Asn concentrations in the indicated cells under nutrient deficiency. l,m, MTT (l) and colony formation (m) assays of 95D sg-ctrl and DAMER-KO cells overexpressing indicated DAMER constructs under nutrient deficiency. n, SELECT assay for m6A levels at DAMER sites A755 and A774 in 95D cells with dCasRx-METTL3 system under control or dCasRx-ALKBH5 system under glutamine deficiency. Representative images or tables are shown for RNA pull-down assays (h), half-life data (j) and colony formation assays (m). All other experiments were repeated independently three times with similar results. Data in quantitative graphs are presented as mean ± s.d. from n = 3 independent biological replicates (a–f,i,k,l,n). P values are calculated by two-way ANOVA followed by Tukey’s test (a–g,i,k,l) or two-tailed unpaired Student’s t-test (n). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 7 The DAMER–ALKBH5-ATF4 axis is a linchpin for metabolic adaptation.
a, MTT cell growth assays of ALKBH5-silenced cells transfected with ATF4 or DAMER under nutrient deficiency. b, Asn concentrations in ALKBH5-silenced cells transfected with ATF4 or DAMER under nutrient deficiency. c, WB analysis of ALKBH5, ATF4, and ASNS levels in the indicated cells. d, MTT cell growth assays of 95D sg-ctrl and DAMER-KO1 cells overexpressing ALKBH5-WT, ALKBH5-H204A, or ALKBH5-Y141A under nutrient deficiency. e, Schematic of N-terminally Flag-tagged full-length ALKBH5 (FL) and its deletion mutants. f, Exogenous RNA pull-down assays validating the interaction between ALKBH5 deletion mutants and DAMER or ATF4 mRNA. g, SELECT assay showing m6A levels at sites 755 and 774 of DAMER in the indicated cells under nutrient deficiency. h, WB analysis of ATF4 levels in 95D sg-ctrl and DAMER-KO1 cells transfected with ATF4-WT or ATF4-A1599G under nutrient deficiency. i, m6A-RIP assay of ATF4 mRNA m6A levels in 95D sg-ctrl cells using the dCasRx-METTL3 system under nutrient deficiency. j, Asn concentrations in 95D sg-ctrl cells using the dCasRx-METTL3 system under nutrient deficiency. k–n, Indicated cells were subcutaneously inoculated in nude mice. k,l, Representative images of xenograft tumours at the end point. Scale bar, 1 cm. m,n, Glc, Gln, and Asn concentrations in the periphery and core tumour tissues. Representative images or tables are shown for Western blots (c,h), RNA pull-down assays (f), and xenograft tumours (k,l). All other experiments were repeated independently three times with similar results. Data in quantitative graphs are presented as mean ± s.d. from n = 3 independent biological replicates (a,b,d,g,i,j,m,n). P values are calculated by two-way ANOVA followed by Tukey’s test (a,b,d,g,j,m,n) or two-tailed unpaired Student’s t-test (i). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 8 Repurposed anti-HCV drug elbasvir targets the DAMER–ALKBH5 interaction and reverses metabolic adaptation.
a, RNA pull-down showing the DAMER–ALKBH5 interaction in 95D sh-Vector, ALKBH5-sh1 and DAMER-KO1 cells treated with elbasvir. b, RNA-FISH showing DAMER localization in ALKBH5-silenced 95D cells treated with elbasvir under nutrient deficiency. Nuclei were stained with DAPI and DAMER was detected by Cy3-conjugated probes. Scale bar, 10 μm. c, m6A-RIP analysis of DAMER m6A levels in 95D sh-Vector cells treated with elbasvir under nutrient deficiency. d,e, ATF4 mRNA half-life (d) and WB analysis of ATF4, ASNS, and SLC3A2 expression (e) in ALKBH5-, ATF4-, ASNS-silenced, or DAMER-depleted cells treated with elbasvir under nutrient deficiency. f, Asn concentrations (f) in the indicated 95D cells under nutrient deficiency and elbasvir treatment. g, Colony formation of the indicated 95D cells treated with elbasvir under nutrient deficiency. h, ASNS-silenced H460 xenografts in mice treated with elbasvir or a vehicle control (n = 5 per group). Representative images of tumours are shown. i,j, Colony formation (i) and MTT (j) assays of non-cancerous cell lines treated with elbasvir under nutrient deficiency. k, Kaplan–Meier survival analysis of nude mice (n = 5 per group) with H460 xenografts and treated with elbasvir or a vehicle control (end point: 63 days). l, H&E staining of heart, liver, spleen, kidney, brain, and lung tissue samples from elbasvir-treated mice. Scale bar, 10 μm. Representative images or tables are shown for RNA pull-down (a), RNA-FISH (b), half-life data (d), western blots (e), colony formation assays (g,i), xenograft tumours (h), and H&E staining (l). All other experiments were repeated independently three times with similar results. Data in quantitative graphs are presented as mean ± s.d. from n = 3 independent biological replicates (c,f,j) or n = 5 mice per group (k). P values are calculated by two-tailed unpaired Student’s t-test (c), two-way ANOVA followed by Tukey’s test (f,j), or the log-rank test (k). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 9 The DAMER–ATF4 axis is clinically relevant and prognostic in human lung cancer.
a,b, Concentrations of Glc, Gln, and Asn (a) and expression of DAMER, ATF4, ASNS, SLCs (SLC3A2, SLC7A1, and SLC7A5) (b) in 30 paired normal (N) and lung cancer (T) tissues. c, Correlation analysis of Glc, Gln, and Asn concentrations, and DAMER, ATF4, ASNS, and SLCs expression from data in Fig. 8a. d, Expression of DAMER, ATF4, ASNS, SLCs, and HIF-1α in TCGA lung cancer datasets. e, ATP levels in 10 paired normal (N) and lung cancer (T) tissues. f, Relative expression of ALKBH5, ATF4, ASNS, SLCs, HIF-1α, p-GCN2, eIF2α, and p-eIF2α in 10 paired normal and lung cancer tissues. g, Relative DAMER expression in lung cancer tissues versus paired normal tissues from 10 patients. h, Correlation of relative DAMER expression with ATF4 or ASNS expression (data from f,g). i, Correlation of relative HIF-1α with ATF4 expression in TCGA lung cancer datasets. j, Correlation matrix for ATP concentration and expression of DAMER, ATF4, HIF-1α, p-GCN2, eIF2α, and p-eIF2α (data from e,f). k, Representative images (left) and correlation (right) of DAMER (RNA-FISH) and G3BP1 (IF) in lung cancer tissue stratified by low Glc/Gln (Glc/Glnl) or high Glc/Gln (Glc/Glnh). Scale bar, 20 μm (left). l, Correlation of nuclear DAMER (nDAMER), ATF4, ASNS, or SLC3A2 in lung cancer tissue with (G3BP1+) or without (G3BP1−) stress granules (SGs). m, Representative images and correlation of DAMER (RNA-FISH) with ATF4, ASNS, and SLC3A2 (IHC) in lung cancer tissue with AsnhGlc/Glnl or AsnlGlc/Glnh. Scale bar, 20 μm (left). Data are presented as correlation heat map summaries (c,j), western blots (f), or representative images (k(left),m(left)). The heat maps summarize data from n = 30 (c) and n = 10 (j) patient pairs. The western blot in f is representative of the 10 patient pairs analysed. The images are representative of observations from 20 independent patient samples (k(left),m(left)). Data in quantitative graphs are presented as mean ± s.d. Sample sizes are as follows: n = 30 patient pairs (a,b); n = 10 patient pairs (e,g); for the TCGA cohort (d), n = 980 tumour samples and n = 109 normal tissue samples; n = 10 patient samples for the linear regression analysis (h); n = 20 patient samples for the association analyses (k–m). P values are calculated by two-tailed paired Student’s t-test (a,b,e), linear regression (c,h–j), two-tailed unpaired Student’s t-test (d), two-way ANOVA followed by Sidak’s test (g), or two-tailed Fisher’s exact test (k,m), or two-tailed Chi-square test (l). Source numerical data, precise P values and unprocessed blots are provided.
Extended Data Fig. 10 Validation and controls for CLIP and RNA-FISH experiments.
a, Validation of the CLIP assay. Image of biotinylated RNA demonstrating the specific, high-molecular-weight ALKBH5-RNA complex under optimized RNase conditions (L, lane 4). L: low RNase I, 1:500; H: high RNase I, 1:50. The red box indicates the membrane region excised for subsequent qRT-PCR analysis. b–j, Representative RNA-FISH images showing no-probe (negative control, NC) and 18S (positive control for cytoplasmic RNA) staining in 95D cells. These controls correspond to experiments shown in the following figures: b, Fig. 5a; c, Fig. 5b; d, Fig. 5d; e, Fig. 5g; f, Fig. 5i; g, Fig. 5m; h, Fig. 6f; i, Fig. 7d; j, Extended Data Fig. 8b. k–m, Additional no-probe negative controls for RNA-FISH experiments performed on lung cancer tissue sections shown in Fig. 8d (k), Fig. 8e and Extended Data Fig. 9m (l), Extended Data Fig. 9k (m). The CLIP validation gel (a) is from a representative experiment. The FISH control experiments shown in (b–m) are representative and were performed in parallel with their corresponding main experiments. All main experiments were repeated independently three times with similar results. Scale bars, 20 μm. Unprocessed blots are provided.
Supplementary information
Supplementary Table 1 (download XLSX )
Oligonucleotides used for gene expression knockdown.
Supplementary Table 2 (download XLSX )
Sense and antisense primers used for quantitative reverse transcriptase PCR.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Fig. 4 (download PDF )
Unprocessed western blots.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 8 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 9 (download PDF )
Unprocessed western blots.
Source Data Extended Data Fig. 10 (download PDF )
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tao, T., Feng, X., Yang, Y. et al. The lncRNA DAMER selectively guides m6A-dependent regulation of ATF4 and asparagine metabolism under nutrient stress in cancer. Nat Cell Biol 28, 797–811 (2026). https://doi.org/10.1038/s41556-026-01905-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41556-026-01905-z