Abstract
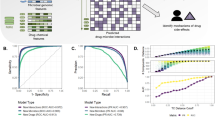
Microbiota-mediated drug metabolism can affect pharmacological efficacy. Here we conducted a systematic comparative metabolomics investigation of drug metabolism modes by evaluating the impacts of human gut commensal bacteria on 127 G-protein-coupled receptor (GPCR)-targeted drugs. For the most extensively metabolized drugs in our screen, we elucidated both conventional and unconventional drug transformations and the corresponding activities of generated metabolites. Comparisons of drug metabolism by a gut microbial community versus individual species revealed both taxon intrinsic and collaborative processes that influenced the activity of the metabolized drugs against target GPCRs. We also observed iloperidone inactivation by generating unconventional metabolites. The human gut commensal bacteria mixture incorporated sulfur in the form of a thiophene motif, whereas Morganella morganii used a cascade reaction to incorporate amino-acid-derived tricyclic systems into the drug metabolites. Our results reveal a broad impact of human gut commensal bacteria on GPCR-targeted drug structures and activities through diverse microbiota-mediated biotransformations.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data generated during this study are included in the Article and its Supplementary Tables 1–25 and Supplementary Figs. 1–309. The information on all GPCR drugs discussed in this Article was sourced from DrugBank Online at https://go.drugbank.com/. Source data are provided with this paper.
References
Rosenbaum, D. M., Rasmussen, S. G. F. & Kobilka, B. K. The structure and function of G-protein-coupled receptors. Nature 459, 356–363 (2009).
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schiöth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Sriram, K. & Insel, P. A. G protein-coupled receptors as targets for approved drugs: how many targets and how many drugs? Mol. Pharmacol. 93, 251–258 (2018).
Yang, D. et al. G protein-coupled receptors: structure- and function-based drug discovery. Signal Transduct. Target. Ther. 6, 1–27 (2021).
Hauser, A. S. et al. Pharmacogenomics of GPCR drug targets. Cell 172, 41–54 (2018).
Salon, J. A., Lodowski, D. T. & Palczewski, K. The significance of G protein-coupled receptor crystallography for drug discovery. Pharmacol. Rev. 63, 901–937 (2011).
Mantas, I., Saarinen, M., Xu, Z.-Q. D. & Svenningsson, P. Update on GPCR-based targets for the development of novel antidepressants. Mol. Psychiatry 27, 534–558 (2022).
Mangoni, A. A. & Jackson, S. H. D. Age-related changes in pharmacokinetics and pharmacodynamics: basic principles and practical applications. Br. J. Clin. Pharmacol. 57, 6–14 (2004).
Ahmed, S., Zhou, Z., Zhou, J. & Chen, S.-Q. Pharmacogenomics of drug metabolizing enzymes and transporters: relevance to precision medicine. Genomics Proteomics Bioinform. 14, 298–313 (2016).
Roden, D. M., Wilke, R. A., Kroemer, H. K. & Stein, C. M. Pharmacogenomics. Circulation 123, 1661–1670 (2011).
Koziolek, M. et al. The mechanisms of pharmacokinetic food–drug interactions—a perspective from the UNGAP group. Eur. J. Pharm. Sci. 134, 31–59 (2019).
Tariq, R. A., Vashisht, R., Sinha, A. & Scherbak, Y. Medication Dispensing Errors and Prevention (StatPearls, 2024).
Li, Y. et al. Current trends in drug metabolism and pharmacokinetics. Acta Pharm. Sin. B 9, 1113–1144 (2019).
Zanger, U. M. & Schwab, M. Cytochrome P450 enzymes in drug metabolism: regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol. Ther. 138, 103–141 (2013).
Zhang, Z. & Tang, W. Drug metabolism in drug discovery and development. Acta Pharm. Sin. B 8, 721–732 (2018).
Lai, Y. et al. Recent advances in the translation of drug metabolism and pharmacokinetics science for drug discovery and development. Acta Pharm. Sin. B 12, 2751–2777 (2022).
Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. & Goodman, A. L. Mapping human microbiome drug metabolism by gut bacteria and their genes. Nature 570, 462–467 (2019).
Javdan, B. et al. Personalized mapping of drug metabolism by the human gut microbiome. Cell 181, 1661–1679.e22 (2020).
Pant, A., Maiti, T. K., Mahajan, D. & Das, B. Human gut microbiota and drug metabolism. Microb. Ecol. 86, 97–111 (2023).
Swanson, H. I. Drug metabolism by the host and gut microbiota: a partnership or rivalry? Drug Metab. Dispos. 43, 1499–1504 (2015).
Zimmermann-Kogadeeva, M. Quantifying host–microbiota interactions. Science 373, 173 (2021).
Li, H., He, J. & Jia, W. The influence of gut microbiota on drug metabolism and toxicity. Expert Opin. Drug Metab. Toxicol. 12, 31–40 (2016).
Verdegaal, A. A. & Goodman, A. L. Integrating the gut microbiome and pharmacology. Sci. Transl. Med. 16, eadg8357 (2024).
Thursby, E. & Juge, N. Introduction to the human gut microbiota. Biochem. J 474, 1823–1836 (2017).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Hou, K. et al. Microbiota in health and diseases. Signal Transduct. Target. Ther. 7, 135 (2022).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Weersma, R. K., Zhernakova, A. & Fu, J. Interaction between drugs and the gut microbiome. Gut 69, 1510–1519 (2020).
Wan, Y. & Zuo, T. Interplays between drugs and the gut microbiome. Gastroenterol. Rep. 10, goac009 (2022).
Spanogiannopoulos, P. et al. Host and gut bacteria share metabolic pathways for anti-cancer drug metabolism. Nat. Microbiol. 7, 1605–1620 (2022).
Haiser, H. J. et al. Predicting and manipulating cardiac drug inactivation by the human gut bacterium Eggerthella lenta. Science 341, 295–298 (2013).
Koppel, N., Maini Rekdal, V. & Balskus, E. P. Chemical transformation of xenobiotics by the human gut microbiota. Science 356, eaag2770 (2017).
Chen, H. et al. A forward chemical genetic screen reveals gut microbiota metabolites that modulate host physiology. Cell 177, 1217–1231 (2019).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Burapan, S., Kim, M. & Han, J. Demethylation of polymethoxyflavones by human gut bacterium, Blautia sp. MRG-PMF1. J. Agric. Food Chem. 65, 1620–1629 (2017).
Schendel, R. R. et al. Structural transformation of 8–5-coupled dehydrodiferulates by human intestinal microbiota. J. Agric. Food Chem. 63, 7975–7985 (2015).
Rich, B. E. et al. Alternative pathway for dopamine production by acetogenic gut bacteria that O‐demethylate 3‐methoxytyramine, a metabolite of catechol O‐methyltransferase. J. Appl. Microbiol. 133, 1697–1708 (2022).
Kumano, T. Specialized metabolites degradation by microorganisms. Biosci. Biotechnol., Biochem. 88, 270–275 (2024).
Bess, E. N. et al. Genetic basis for the cooperative bioactivation of plant lignans by Eggerthella lenta and other human gut bacteria. Nat. Microbiol. 5, 56–66 (2020).
Casey Laizure, S., Herring, V., Hu, Z., Witbrodt, K. & Parker, R. B. The role of human carboxylesterases in drug metabolism: have we overlooked their importance? Pharmacother. J. Hum. Pharmacol. Drug Ther. 33, 210–222 (2013).
Kim, H.-J. et al. Glucuronidation of a sarpogrelate active metabolite is mediated by UDP-glucuronosyltransferases 1A4, 1A9, and 2B4. Drug Metab. Dispos. 41, 1529–1537 (2013).
Malhotra, B., Guan, Z., Wood, N. & Gandelman, K. Pharmacokinetic profile of fesoterodine. Int. J. Clin. Pharmacol. Ther. 46, 556–563 (2008).
Ishizuka, T. et al. Human carboxymethylenebutenolidase as a bioactivating hydrolase of olmesartan medoxomil in liver and intestine. J. Biol. Chem. 285, 11892–11902 (2010).
Kuze, Y., Kogame, A., Jinno, F., Kondo, T. & Asahi, S. Development, validation and application of the liquid chromatography tandem mass spectrometry method for simultaneous quantification of azilsartan medoxomil (TAK-491), azilsartan (TAK-536), and its 2 metabolites in human plasma. J. Chromatogr. B 1001, 174–181 (2015).
Asaki, T. et al. Selexipag: an oral and selective IP prostacyclin receptor agonist for the treatment of pulmonary arterial hypertension. J. Med. Chem. 58, 7128–7137 (2015).
Helf, M. J., Fox, B. W., Artyukhin, A. B., Zhang, Y. K. & Schroeder, F. C. Comparative metabolomics with Metaboseek reveals functions of a conserved fat metabolism pathway in C. elegans. Nat. Commun. 13, 782 (2022).
Vizcaino, M. I., Engel, P., Trautman, E. & Crawford, J. M. Comparative metabolomics and structural characterizations illuminate colibactin pathway-dependent small molecules. J. Am. Chem. Soc. 136, 9244–9247 (2014).
Böcker, S., Letzel, M. C., Lipták, Z. & Pervukhin, A. SIRIUS: decomposing isotope patterns for metabolite identification. Bioinformatics 25, 218–224 (2009).
Dührkop, K. et al. SIRIUS 4: a rapid tool for turning tandem mass spectra into metabolite structure information. Nat. Methods 16, 299–302 (2019).
Zheng, R., Wu, Y.-H., Jiang, D.-X. & Zhang, D. Determination of metabolite of nicergoline in human plasma by high-performance liquid chromatography and its application in pharmacokinetic studies. J. Pharm. Anal. 2, 62–66 (2012).
Zhu, H.-J. et al. Carboxylesterase 1 as a determinant of clopidogrel metabolism and activation. J. Pharmacol. Exp. Ther. 344, 665–672 (2013).
Kaye, B., Cussans, N., Faulkner, J., Stopher, D. & Reid, J. The metabolism and kinetics of doxazosin in man, mouse, rat and dog. Br. J. Clin. Pharmacol. 21, 19S–25S (1986).
Jia, M. et al. Simultaneous determination of iloperidone and its two active metabolites in human plasma by liquid chromatography–tandem mass spectrometry: application to a pharmacokinetic study. J. Chromatogr. B 928, 52–57 (2013).
Subramanian, N. & Kalkman, H. O. Receptor profile of P88-8991 and P95-12113, metabolites of the novel antipsychotic iloperidone. Prog. Neuropsychopharmacol. Biol. Psychiatry 26, 553–560 (2002).
Reher, R. et al. A convolutional neural network-based approach for the rapid annotation of molecularly diverse natural products. J. Am. Chem. Soc. 142, 4114–4120 (2020).
Dunbar, K. L., Scharf, D. H., Litomska, A. & Hertweck, C. Enzymatic carbon–sulfur bond formation in natural product biosynthesis. Chem. Rev. 117, 5521–5577 (2017).
Zhang, X. et al. Biosynthesis of chuangxinmycin featuring a deubiquitinase-like sulfurtransferase. Angew. Chem. Int. Ed. 60, 24418–24423 (2021).
Wang, Y. et al. Identifying the minimal enzymes for unusual carbon–sulfur bond formation in thienodolin biosynthesis. ChemBioChem 17, 799–803 (2016).
Ouyang, H. et al. Mining the metabolic capacity of Clostridium sporogenes aided by machine learning. Angew. Chem. Int. Ed. 63, e202319925 (2024).
Bittremieux, W. et al. Comparison of cosine, modified cosine, and neutral loss based spectrum alignment for discovery of structurally related molecules. J. Am. Soc. Mass. Spectrom. 33, 1733–1744 (2022).
Wen, W.-H. et al. Reductive inactivation of the hemiaminal pharmacophore for resistance against tetrahydroisoquinoline antibiotics. Nat. Commun. 12, 7085 (2021).
Lichman, B. R. The scaffold-forming steps of plant alkaloid biosynthesis. Nat. Prod. Rep. 38, 103–129 (2021).
Mancinotti, D., Michiko Frick, K. & Geu-Flores, F. Biosynthesis of quinolizidine alkaloids in lupins: mechanistic considerations and prospects for pathway elucidation. Nat. Prod. Rep. 39, 1423–1437 (2022).
Cao, Y. et al. Commensal microbiota from patients with inflammatory bowel disease produce genotoxic metabolites. Science 378, eabm3233 (2022).
Wu, Q. et al. Metabolomics and genomics enable the discovery of a new class of nonribosomal peptidic metallophores from a marine micromonospora. J. Am. Chem. Soc. 145, 58–69 (2023).
Rampino, A. et al. Antipsychotic drug responsiveness and dopamine receptor signaling; old players and new prospects. Front. Psychiatry 9, 702 (2019).
Wang, S. et al. Structure of the D2 dopamine receptor bound to the atypical antipsychotic drug risperidone. Nature 555, 269–273 (2018).
Rabin, C. et al. In vitro and in vivo demonstration of risperidone implants in mice. Schizophr. Res. 98, 66–78 (2008).
Spina, E. & De Leon, J. Metabolic drug interactions with newer antipsychotics: a comparative review. Basic Clin. Pharmacol. Toxicol. 100, 4–22 (2007).
Wilson, I. D. & Nicholson, J. K. Gut microbiome interactions with drug metabolism, efficacy, and toxicity. Transl. Res. 179, 204–222 (2017).
Vich Vila, A. et al. Impact of commonly used drugs on the composition and metabolic function of the gut microbiota. Nat. Commun. 11, 362 (2020).
Zhao, Q., Chen, Y., Huang, W., Zhou, H. & Zhang, W. Drug–microbiota interactions: an emerging priority for precision medicine. Signal Transduct. Target. Ther. 8, 386 (2023).
Park, H. B. et al. Bacterial autoimmune drug metabolism transforms an immunomodulator into structurally and functionally divergent antibiotics. Angew. Chem. Int. Ed. 59, 7871–7880 (2020).
Bustion, A. E., Nayak, R. R., Agrawal, A., Turnbaugh, P. J. & Pollard, K. S. SIMMER employs similarity algorithms to accurately identify human gut microbiome species and enzymes capable of known chemical transformations. eLife 12, e82401 (2023).
Heinken, A. et al. Genome-scale metabolic reconstruction of 7,302 human microorganisms for personalized medicine. Nat. Biotechnol. 41, 1320–1331 (2023).
Zimmermann, M., Zimmermann-Kogadeeva, M., Wegmann, R. & Goodman, A. L. Separating host and microbiome contributions to drug pharmacokinetics and toxicity. Science 363, eaat9931 (2019).
Loureiro, A. I. et al. Absorption, metabolism and excretion of opicapone in human healthy volunteers. Br. J. Clin. Pharmacol. 88, 4540–4551 (2022).
Surapaneni, S. et al. Absorption, metabolism, and excretion, in vitro pharmacology, and clinical pharmacokinetics of ozanimod, a novel sphingosine 1-phosphate receptor modulator. Drug Metab. Dispos. 49, 405–419 (2021).
Overton, H. A., Fyfe, M. C. T. & Reynet, C. GPR119, a novel G protein-coupled receptor target for the treatment of type 2 diabetes and obesity. Br. J. Pharmacol. 153, S76–S81 (2008).
Flock, G., Holland, D., Seino, Y. & Drucker, D. J. GPR119 regulates murine glucose homeostasis through incretin receptor-dependent and independent mechanisms. Endocrinology 152, 374–383 (2011).
Qian, Y. et al. Activation and signaling mechanism revealed by GPR119-Gs complex structures. Nat. Commun. 13, 7033 (2022).
Zhao, J., Zhao, Y., Hu, Y. & Peng, J. Targeting the GPR119/incretin axis: a promising new therapy for metabolic-associated fatty liver disease. Cell. Mol. Biol. Lett. 26, 32 (2021).
Goodman, A. L. et al. Identifying genetic determinants needed to establish a human gut symbiont in its habitat. Cell Host Microbe 6, 279–289 (2009).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Smith, C. A., Want, E. J., O’Maille, G., Abagyan, R. & Siuzdak, G. XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 78, 779–787 (2006).
Marzullo, P. et al. Ammonium formate-Pd/C as a new reducing system for 1,2,4-oxadiazoles. synthesis of guanidine derivatives and reductive rearrangement to quinazolin-4-ones with potential anti-diabetic activity. Int. J. Mol. Sci. 22, 12301 (2021).
Hanwell, M. D. et al. Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J. Cheminformatics 4, 17 (2012).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Acknowledgements
We thank members of the Crawford lab for helpful discussions. We also thank T. Wu and J. Kim at the Yale West Campus Analytical Core for their assistance with LC–MS. This work was primarily supported by the National Institute of General Medical Sciences (1RM1GM141649 to J.M.C. and N.W.P.). The mouse model experiment (DP2DK125119 to N.W.P.) and preparation of the microbial community (R01AT010014 to A.L.G.) were supported by the NIH. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. T.T. was in part supported by the NIH Chemistry-Biology Interface Training Program (T32GM067543) and the NSF Predoctoral Fellowship Program (2020293597). A.A.V. was in part supported by the NIH Individual Predoctoral Fellowship program (F31DK132941). B.D.-L. is a Cancer Research Institute Irvington Fellow supported by the Cancer Research Institute (CRI4515). Y.Z. was in part supported by a Peter Moore fellowship in the Department of Chemistry.
Author information
Authors and Affiliations
Contributions
Q.W., D.S., A.L.G., N.W.P. and J.M.C. conceived the project. Q.W., D.S., Y.Z., A.A.V. and T.T. conducted the experiments. B.D.-L. performed molecular docking. Q.W. and D.S. carried out data analyses. Q.W. prepared graphical illustrations and wrote the first draft of the manuscript. All authors reviewed, edited and contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
N.W.P. is a co-founder of Artizan Biosciences and has received research funding for unrelated studies from Artizan Biosciences and F. Hoffmann-La Roche. A.L.G. serves on the scientific advisory boards for Seres Therapeutics, Piton Therapeutics, Nuanced Health and Taconic Biosciences. The others authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks Aaron Wright and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Mapping of GPCR drug activities.
(a) Number of GPCRs and their class that interacted with the 127 selected drugs. The drug target information was obtained from the DrugBank database. (b) Mapping specific interactions between drugs and GPCRs. Agonist interactions were labeled in red and antagonist interactions were labeled in grey. Aligned with the prevalence of antagonistic GPCR drugs in the DrugBank database, our drug panel was composed primarily of antagonists (74.8 %, 95/127), but they were complemented with some agonists (20.5 %, 26/127) and dual-function drugs (4.7 %, 6/127).
Extended Data Fig. 2 High-throughput identification of 12 GPCR drugs heavily consumed through drug metabolism screening by the human gut bacterial collection.
(a) The bar graph illustrates that out of the screened 127 drugs, 12 were heavily consumed (drug consumed ≥ 80%). n = 3 biological replicates. Data represent mean ± s.d. (b) The EICs of parent drug ions for both Bac+ and Bac- (after 24 h of drug incubation) offer a relative quantification of their concentration in the supernatants. The significant differences in drug concentration observed between the presence and absence of bacteria strongly suggest that drug consumption is dependent on the presence of bacteria. The EICs (with a ± 20 ppm window) of drug ions were determined using the following ions: [M + H]+, m/z 559.2300 for olmesartan medoxomil; [M + H]+, m/z 569.1667 for azilsartan medoxomil; [M + H]+, m/z 389.2071 for trimethobenzamide; [M + H]+, m/z 412.2846 for fesoterodine; [M + H]+, m/z 497.2217 for selexipag; [M + H]+, m/z 427.2028 for iloperidone; [M + H]+, m/z 405.1921 for ozaminod; [M + H]+, m/z 461.1296 for ponesimod; [M + H]+, m/z 430.2224 for sarpogrelate; [M + H]+, m/z 501.1360 for SB756050; [M + H]+, m/z 388.2118 for Trimebutine; [M + H]+, m/z 457.1904 for GSK1292263. The experiment was conducted in triplicates for both Bac+ and Bac- conditions. Data are means ± SEM. n.d., not detected. Statistical significance was determined using two-tailed t-test.
Extended Data Fig. 3 Impact of gut bacterial drug metabolism on GPCR activity.
(a) Sample preparation procedure for GPCR activity tests. (b) GPCR activity of 12 drugs were tested and compared under two conditions: drugs were administered without undergoing metabolism (at 0 h, blue), and drugs undergoing complete metabolization (at 24 h, red). Drug activities were quantified by EA50 values, Log10[Conc, M] refers to the starter drug concentration of 10 μM in the complex medium extracts. n = 3 biological replicates. Data are means ± SEM.
Extended Data Fig. 4 Comparative metabolomics-based drug metabolite discovery and characterization.
Volcano plot illustrates the FC (molecular features with Bac+ versus molecular features with Bac-) and FDR values in LC-QTOF-MS positive ion mode. (a) The raw LC-MS files of Bac+ and Bac- were processed using XCMS modules to transform them into molecular feature information through feature detection, grouping, and filtering ( < 3,000 molecular features were prioritized based on a strict intensity cutoff to focus on robust drug transformations in this study). (b) Molecular feature selection involved volcano plot analysis with a p-value and fold change cutoff, followed by a manual inspection to create a conclusive validated hit list. (c) Structural elucidation of molecular features of interest was carried out through MS/MS and NMR techniques, as well as chemical standard comparisons.
Extended Data Fig. 5 Drug metabolite profiles for six drug metabolomes.
(a) Information on the metabolites of six drugs. The bar graphs on the left and the EIC traces on the right depict a comparison between Bac- and Bac+ for ponesimod (b), SB756050 (c), trimethobenzamide (d), GSK1292263 (e), ozanimod (f), and trimebutine (g). The metabolism of five other drugs, including sarpogrelate, olmesartan medoxomil, fesoterodine, and azilsartan medoxomil, was characterized through standard comparison (Supplementary Fig. S2). The metabolome of iloperidone is detailed in Fig. 4. n = 3 biological replicates. Data are means ± SEM. Statistical significance was determined using two-tailed t-test.
Extended Data Fig. 6 Mapping drug metabolism by individual bacterial species.
(a) Selected 12 isolates from bacterial collection represent their corresponding clades for the examination of drug metabolism. Proteobacteria, red; Actinobacteria, purple; Bacteroidetes, green; Firmicutes, blue. Mono bacterial drug metabolism for azilsartan medoxomil (b), olmesartan medoxomil (c), sarpogrelate (d), fesoterodine (e), selexipag (f), ozanimod (g), GSK1292263 (h), ponesimod (i), and iloperidone (j). No significant O-demethylation metabolism of the three drugs (trimebutine, trimethobenzamide, and SB756050) was detected in the 12 selected bacterial isolates. Further details are provided in Supplementary Fig. S81. n = 3 biological replicates. Data are means ± SEM.
Extended Data Fig. 7 Proposed iloperidone drug metabolism pathways.
(a) Iloperidone metabolism pathways by human gut commensal bacteria described in this study. (b) Reported biosynthetic pathway for indolethiophene skeleton in Streptomyces sp. (c) Reported quinolizidine alkaloids in the biosynthesis of lupins.
Extended Data Fig. 8 Verification of the activity of metabolites for two drugs, GSK1292263 and iloperidone.
(a) Chemical transformation route for generating a GSK1292263 metabolite, GSK-459. (b) Structural elucidation of synthetic GSK-459 by NMR analyses. (c) LC-MS-based retention time comparison between synthetic standard (trace A in blue), drug metabolite from bacterial transformation (trace B in black) and co-injection (trace C in red). The EIC (with a ± 20 ppm window) for GSK-459 ion was determined using [M + H]+, m/z 459.2061. (d) MS2 similarity comparison with standard obtained from chemical reaction (top spectrum in blue) and drug metabolite from bacterial transformation (bottom spectrum in red). The predicted fragments from MS2 are indicated in panel b. (e) GPCR activity test (GPR119 agonist) on both GSK1292263 and its inactive metabolite GSK-459. n = 3 biological replicates. Data represents mean ± s.d. (f) Chemical structures of iloperidone and its metabolites. All metabolites were isolated from bacterial transformation culture broth, with their structures determined using NMR, excluding metabolite ilo-477, which was inferred by comparative tandem MS and isotope labelling studies. (g) Key 2D NMR (COSY and HMBC) correlations for the structural assignments of ilo-430. (h) GPCR activity test (DRD2 antagonist) on iloperidone and its metabolites. n = 3 biological replicates. Data represents mean ± s.d.
Extended Data Fig. 9 Summary of the metabolites of 17 drugs analyzed in our screen.
The metabolism of Iloperidone is summarized in Extended Data Fig. 8f. The chemical structures of azilsartan medoxomil (a), olmesartan medoxomil (b), ponesimod (c), clopidogrel (d), selexipag (e), risperidone (f), candesartan cilexetil (g), sarpogrelate (h), trimebutine (i), nicergoline (j), trimethobenzamide (k), MK-0557 (l), ozanimod (m), doxazosin (n), fesoterodine (o), SB756050 (p), GSK1292263 (q) and their metabolites are listed. The chemical structures of SB756050 metabolites, including SB-487, SB-473, and SB-459, were determined through 2D NMR analysis of materials isolated from the culture broth. A chemical synthesis approach and/or standard comparison was used to determine the structures of the metabolites of GSK1292263 (GSK-459), olmesartan medoxomil (olmesartan), azilsartan medoxomil (azilsartan), selexipag (ACT-333679), sarpogrelate (sarpogrelate-330), and fesoterodine (fesoterodine-342). MS/MS analysis was used to predict the chemical structures of ponesimod-463a/463b, trimebutine-374, trimethobenzamide-361, trimethobenzamide-347, ozanimod-407, ozanimod-408, doxazosin-438, doxazosin-424, SB-445, and GSK-460. The chemical structures of the remaining metabolites were proposed based on MS analysis. Ten microbial transformations, including those of azilsartan medoxomil, olmesartan medoxomil, clopidogrel, selexipag, candesartan cilexetil, sarpogrelate, nicergoline, ozanimod, doxazosin, and fesoterodine, were conserved with the transformations observed in human metabolism.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–309 and Tables 1–25.
Supplementary Data (download ZIP )
NMR data of nine identified compounds.
Supplementary Data (download XLSX )
The properties of all GPCR drugs studied in this Article.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, Q., Song, D., Zhao, Y. et al. Activity of GPCR-targeted drugs influenced by human gut microbiota metabolism. Nat. Chem. 17, 808–821 (2025). https://doi.org/10.1038/s41557-025-01789-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41557-025-01789-w
This article is cited by
-
Gut microbiome-mediated transformation of dietary phytonutrients is associated with health outcomes
Nature Microbiology (2025)
-
Gut microbiota-mediated GPCR drug metabolism affects downstream pharmacological activity
Nature Chemistry (2025)