Abstract
Biodiversity and its associated genetic diversity are being lost at an unprecedented rate. Simultaneously, the distributions of flora, fauna, fungi, microbes and pathogens are rapidly changing. Novel technology can help to capture and record genetic diversity before it is lost and to measure population shifts and pathogen distributions. Here we report the rapid application of shotgun long-read environmental DNA (eDNA) analysis for non-invasive biodiversity, genetic diversity and pathogen assessments from air. We also compared air eDNA with water and soil eDNA. Coupling long-read sequencing with established cloud-based biodiversity pipelines enabled a 2-day turnaround from airborne sample collection to completed analysis by a single investigator. To determine the full utility of airborne eDNA, we also conducted a local bioinformatic analysis and deep short-read shotgun sequencing. From outdoor air eDNA alone, comprehensive genetic analysis was performed, including population genetics (phylogenetic placement) of a charismatic mammal (bobcat, Lynx rufus) and a venomous spider (golden silk orb weaver, Trichonephila clavipes), and haplotyping humans (Homo sapiens) from natural complex community settings, such as subtropical forests and temperate locations. The rich datasets also enabled deeper analysis of specific species and genomic regions of interest, including viral variant calling, human variant analysis and antimicrobial resistance gene surveillance from airborne DNA. Our results highlight the speed, versatility and specificity of pan-biodiversity monitoring via non-invasive eDNA sampling using current benchtop/portable and cloud-based approaches. Furthermore, they reveal the future feasibility of scaling down (equipment and temporally) these approaches for near real-time analysis. Together these approaches can enable rapid simultaneous detection of all life and its genetic diversity from air, water and sediment samples for unbiased non-targeted information-rich genomics-empowered (1) biodiversity monitoring, (2) population genetics, (3) pathogen and disease-vector genomic surveillance, (4) allergen and narcotic surveillance, (5) antimicrobial resistance surveillance and (6) bioprospecting.
Similar content being viewed by others
Main
New approaches are required to capture and record genetic diversity before it is lost and measure shifts in species and pathogen distributions, due to the variety of crises threatening, reducing and redistributing biodiversity on a global scale1,2,3,4,5. Deep-sequencing technologies are rapidly advancing in speed, power and portability. Coupled with improvements in DNA capture from environmental substrates, including airborne DNA, these developments enhance our ability to non-invasively detect species and analyse environmental DNA (eDNA) to obtain unprecedented insights6,7,8,9,10,11.
Airborne DNA is rapidly being developed as a biodiversity monitoring tool, enabling the identification of species presence in a location from air sampling alone7,12,13,14,15,16. While making highly notable advances, the majority of air eDNA effort has focused on targeted approaches such as metabarcoding. Metabarcoding only recovers a small amount of DNA per species/group, but this DNA section (barcode) is informative for the species of origin. The main advantage of this approach is suspected accuracy. The application of whole-genome sequencing approaches to metagenomic (multispecies) samples is known as shotgun sequencing. There is a paucity of studies directly comparing the performance of metabarcoding to shotgun sequencing metagenomic approaches. Additional metabarcoding limitations include requiring a priori genetic sequence knowledge of target species and the inability to accurately identify species from populations with genetic diversity that is not yet documented/investigated and which is unaccounted for in barcoding databases. In instances where metabarcoding is used in populations that are well sequenced, the technology is able to recover informative (barcodes), but limited genetic information per species present (approximately a few hundred base pairs)6,7,12,14,15,17,18,19,20,21,22,23,24. This is also true of metabarcoding studies recovering deposited airborne eDNA by swabbing surfaces, such as foliage or spider webs25,26,27,28.
Deep sequencing is cost effective and efficient enough to enable shotgun approaches to species identification, in which regions from across the entire genome of each species present can be sequenced in parallel, providing information-rich datasets7,8,10,29. By not requiring PCR-based barcode amplification, shotgun sequencing approaches also more accurately represent the original proportions of species DNA in an environmental sample. Long-read sequencing of air eDNA has been applied to study microbes30, humans7,31 and selected multicellular species7,31, but pan-domain-of-life long-read analysis has not been conducted. Deep-sequencing long-read or short-read shotgun approaches are not limited to species identification across preselected taxonomically restricted species assemblages. They can facilitate pan-biodiversity lifeform detection, while simultaneously allowing population genetics, genetic variant analysis, pathogen surveillance, antimicrobial resistance (AMR) gene surveillance and bioprospecting, all from a single sample/assay, providing ultrarich datasets. Bioprospecting and genome mining are being utilized for novel drug discovery from microbes without the need for specimen collection or culture32,33. New drugs are primarily derived from natural products34. Genomic shotgun sequencing-based eDNA-empowered bioprospecting offers ecosystem-scale drug discovery across the entire tree of life, not just microbes. Meanwhile, the study of genetic variation within eDNA samples can be utilized for applications as diverse as population genetics6,7, disease risk-associated allelles7, viral variant surveillance6,11, pesticide resistance markers and AMR surveillance. Long-read sequencing technologies are likely to prove particularly advantageous to eDNA-based bioprospecting, population genetics, genetic variant analysis and pathogen and AMR gene surveillance applications7,35, as they can provide increased specificity and haplotype-phased genetic information on individuals even from within complex pooled environmental samples7.
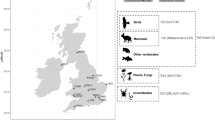
In concert with advances in remote sampling, rapid eDNA library preparation and sequencing, analysis and bioinformatic pipelines are getting ever closer to a point of real-time lifeform, pathogen and genetic diversity detection in the environment, particularly for airborne DNA. Here, we captured airborne DNA from a variety of indoor and outdoor sampling sites (Figs. 1 and 2a and Supplementary Fig. 1a). We show that sampling, shotgun sequencing and cloud-based bioinformatic capabilities have already matured sufficiently to enable lifeform monitoring and genetic surveillance from airborne DNA, even from temperate (Ireland) or subtropical (Florida) complex natural outdoor environments. We compare long-read (Oxford Nanopore Technologies, ONT) and short-read (Illumina) shotgun sequencing technologies. In addition, we demonstrate that field-recovered airborne DNA can be simultaneously leveraged for a broad range of downstream applications (Fig. 1, Table 1 and Extended Data Table 1), including pan-biodiversity assessments of complex prokaryote and eukaryote communities, individual ancestry genetics and variant associations and pathogen and AMR gene monitoring.
A schematic overview of sample types, locations and eDNA analytical approaches utilized in this study and the downstream applications explored. Figure created with BioRender.com.
Results
Air eDNA samples were collected between 2022 and 2024 from a broad range of urban and rural locations (Fig. 2a and Supplementary Fig. 1a). We also collected rainwater, river water, saltwater, soil, scat and sand samples for comparison (Fig. 1 and Supplementary Table 1). This resulted in a total of 78 shotgun sequencing datasets (30 long-read and 48 short-read). Six samples were sequenced with both technologies (Supplementary Table 1). A total of 4 human exome enrichment air eDNA datasets and 14 vertebrate 12S metabarcoding air eDNA datasets were also generated.
a, A schematic of the locations of the Florida, USA and Irish sampling sites. Maps created in ArcGIS online. Satellite imagery from Google Earth; map data ©2021 Google and TerraMetrics. UF, University of Florida; WRI, Wildlife Rehabilitation Ireland; TCD, Trinity College Dublin. b, Human ZNF285 gene species-specific qPCR-based detection from Irish eDNA samples and NFC samples. Each qPCR reaction is a 10-μl reaction containing 1 μl of extracted eDNA template. The absolute quantity is per reaction, derived from a standard curve using human cell whole-genome DNA. Each boxplot is a single sample, and each datapoint (three technical qPCR wells per reaction replicates per sample) is displayed by a grey dot. Tukey whiskers (which extend to data points that are less than 1.5× the interquartile range (IQR) away from the first and third quartile) were utilized for each boxplot. The median for each sample is shown as a horizontal line within each box, and the box edges are the upper and lower quartiles. c, The absolute quantity of 18S rRNA pan-eukaryotic eDNA levels within Irish eDNA samples and NFCs, as detected by qPCR. Each qPCR reaction is a 10-μl reaction containing 1 μl of extracted eDNA template. The absolute quantity is per reaction, derived from a standard curve using human cell whole-genome DNA. Each boxplot is a single sample, and each datapoint (three technical qPCR wells per reaction replicates per sample) is displayed by a grey dot. Tukey whiskers (which extend to data points that are less than 1.5× the IQR away from the first and third quartile) were utilized for each boxplot. The median for each sample is shown as a horizontal line within each box, and the box edges are the upper and lower quartiles. d, Human haplogroup and haplotyping from short-read shotgun sequencing of a forest/low density residential and city centre outdoor air samples and the human mitochondrial sequence relative abundance of total sequenced reads per sample. Capital letters and numbers denote the human mitochondrial haplotypes detected in each air eDNA sample.
Population genetics from airborne DNA, human
We assessed whether human (Homo sapiens) genetic ancestry applications (akin to animal population genetics) were already achievable using shotgun sequencing of airborne eDNA samples from a natural outdoor setting with a complex community of species present. Species-specific human quantitative PCR (qPCR) revealed that air eDNA recovered from Dublin City contains more human DNA than air sampled from rural locations (Fig. 2b). Human eDNA was highest in the city air sample, despite the other air samples having higher overall pan-eukaryotic eDNA recovery (Fig. 2c). The highest level of pan-eukaryotic DNA recovered was from continuous air sampling for 1 month in a rural location (Fig. 2c). The level of human eDNA recovered from city air was of a similar scale to that recovered from a river that we previously showed7,36 to be polluted with high levels of improperly treated human wastewater (Fig. 2b). The short-read shotgun sequencing of airborne DNA revealed a greater number of human ancestry haplotypes in Dublin City air samples than the much lower habitation density forest/mixed residential area sample (Fig. 2d). While the overall proportions were similar (predominantly European–Indian), only 8 distinct phylotypes were identified in the forest compared with 87 (2023) and 84 (2024) distinct phylotypes in the city air (Fig. 2d), with human mitochondrial sequences also more abundant in city air (relative abundances in forest: 0.0001%, city: 0.001–0.002%).
Population genetics from airborne DNA, wildlife
Despite using the lowest output nanopore sequencer and the cloud-based Chan Zuckerberg (CZ) ID37,38 platform only considering 1 million of the generated reads (per sample), long-read sequencing recovered almost the entirety of some metazoan organelle genomes, including mitochondria (Forest air April 2023, Fig. 3a). In this sample, the Carposina sasakki (moth) mitochondrial genome (Fig. 3a) was covered separately four times by single continuous reads, revealing that with enough sequencing depth or enrichment, the entire mitochondrial genomes of individuals can be recovered from the air. This suggests that wildlife population genomic approaches are feasible from shotgun air eDNA sequencing, especially as genomic databases become increasingly populated.
a, The selected species mitochondrial genome and chloroplast genome coverage from long-read shotgun nanopore MinION sequencing of an outdoor air sample (Florida: 1-week forest air, April 2023). b, The maximum-likelihood phylogenetic tree (phylogeny) of the bobcat (L. rufus), with mitochondrial reads from an outdoor Illumina short-read shotgun sequenced air sample (Florida: 1-week forest air, April 2023). For sequences and accession numbers utilized for the tree, see Supplementary Table 3. c, A circular cladogram tree of the golden silk orb weaver (Trichophiles clavipes), showing the placement of the sequences recovered from short-read shotgun sequencing of an outdoor air eDNA sample (Florida: 1-week forest air, April 2023). The cladogram branch lengths are normalized for visual purposes only. The inset photo shows a golden silk orb weaver from the Florida forest study site taken in 2024, approximately 1 year after the air sample used for phylogenetic analysis had been obtained. The photo was taken by D.J.D.
This same Florida coastal forest/mixed residential air sample (April 2023) was short-read deep sequenced and two example species investigated: a charismatic wild predator (bobcat, Lynx rufus) and a venomous spider (Trichonephila clavipes). The bobcat eDNA was shown to be present in this 1-week air sample, which was taken adjacent to an animal track known to occasionally be used by bobcats (Supplementary Video 1). The bobcat mitochondrial aligning short-reads were used to investigate their source population. From air sampling alone, phylogenetic analysis revealed that the airborne bobcat DNA clustered most closely with that of a wild bobcat and a zoo-housed bobcat in northeast Florida (Fig. 3b). The wild Florida bobcat DNA sequence was from bobcat scat we collected approximately 0.15 km away from the forest air sampling site 11 months after the air sample had been taken (Supplementary Table 1). Next, we found that golden silk orb weaver spider eDNA in the same Florida forest 1-week air sample clustered with other North American isolates, phylogenetically outgrouping from Caribbean or South American sequences (Fig. 3c). Taken together, this demonstrates that current technological capabilities are sufficient for population genetics of a diverse range of wildlife from air samples alone, even when non-targeted shotgun sequencing approaches are utilized.
Pathogen surveillance from airborne DNA
The eDNA genomics demonstrates that pathogen variant/strain analysis and genomic surveillance is highly feasible from outdoor airborne DNA. For example, using long-read shotgun sequencing and cloud-based CZ ID37,38 analysis, 27.6× whole-genome coverage of an avian virus (faeces-associated gemycircularvirus 17) was recovered from a windowpane swab (Mountain May 2023), while 6,483.3× (Estuary July 2023) (Extended Data Fig. 1a, 90.43% identify, fast base calling) and 8.0× (Dublin City 2023) whole-genome coverage of the same virus was recovered from pumped air samples. In total, 63 viruses were detected in the long-read Dublin City air sample. The long-read shotgun sequencing and CZ ID analysis also enabled virus surveillance from aquatic and beach sand eDNA samples. For example, the Florida Gulf Coast nearshore water sample (Keaton Beach June 2023) detected 37 human and animal viruses including frog, cow, fish, human, horse and mollusc viruses and the zoonotic cowpox virus (Extended Data Fig. 1b). Air eDNA long-read shotgun sequencing enabled simultaneous pathogen surveillance for viral, bacterial, archaea and eukaryotic disease-related species. In total, 221 species from across the tree of life with pathogenic potential for humans were detected in the long-read Dublin City air sample (cutoff of minimum 1,000 bp sequences). Agroforestry-relevant pathogens could also be recovered from airborne DNA, such as laminated root rot fungus (Coniferiporia sulphurascens) (Extended Data Fig. 1c), as could tree eDNA (Extended Data Fig. 1d–f) and the plant and human fungal pathogen Alternaria alternata (Extended Data Fig. 1f).
Bioprospecting and AMR gene surveillance from airborne eDNA
The genetic information recovered from airborne DNA can also be utilized for critical applications such as biodiversity bioprospecting or AMR gene surveillance, irrespective of requirements to identify the DNA’s species of origin. We show that AMR genes were detectable in each eDNA sample type analysed, with the number of AMR genes identified in air eDNA samples similar to the number detected in sand eDNA but less than water eDNA (Fig. 4a). Most AMR genes were unique to each sample, but some were shared across samples and substrates (Fig. 4a and Supplementary Fig. 1b).
a, UpSet plot for two sand, air and water samples (short-read shotgun sequenced), with the frequency of unique and shared AMR genes, with total AMR genes detected per sample (identified by CZ ID). Ire, Ireland; FM, Florida Marineland; Dc, Dermochelys coriacea (leatherback sea turtle); Fl, Florida. b, A quantification of H. sapiens single-nucleotide variation (SNV), sequence alteration, insertions and deletion variants recovered from indoor air eDNA human exome enrichment short-read sequencing. On the y axis is the number of variant counts (log6) per variant category. The number of counts (not log transformed) for each sample for each category is provided above each bar.
Human genomic variant detection by exome enrichment from air
The eukaryotic/vertebrate genetic variance information was also obtainable from DNA recovered from the air. The hybridization-based target enrichment can capture genetic variant information from particular species of interest. Human exome hybridization-based enrichment and short-read deep sequencing (whole-exome sequencing) applied to indoor air samples (2 h and 20 min room air, October 2022, and 3 h and 15 min room air, January 2023) recovered genomic variant information, including insertions, deletions and over 217,149 single-nucleotide polymorphisms (SNPs) (Fig. 4b). Together, the AMR and human variant analyses further highlighted the utility of airborne DNA for a wide variety of genetic diversity applications.
Airborne DNA for surveillance, disease vectors and pests
The eDNA from human and animal disease vector species were also readily recoverable from the air by shotgun sequencing, including mosquitoes and biting midges (Culicoides) (Extended Data Fig. 2a,b). This opens new avenues for large-scale disease vector species surveillance, including arthropod population genetics (Fig. 3c) or surveillance for insecticide resistance loci. Due to global warming and human-linked invasive species introductions, approximately 4 billion people are at risk of mosquito-borne viral diseases. Simultaneously, airborne DNA shotgun sequencing detected mammalian vectors of human disease, such as the black rat (Rattus rattus), brown rat (Rattus norvegicus) and house mouse (Mus musculus) (Extended Data Fig. 2c).
Airborne DNA could also detect urban pest species, which have a substantial economic impact. For example, at the mixed residential/hammock forest site (Forest air April 2023), 11 pest species such as cockroach, fire ant and termite were detected (Extended Data Fig. 3a,b). The positive correlation between the abundance of these 11 pest species was remarkably similar whether the sample was sequenced with long-read or short-read technology (linear regression, R2 = 0.994) (Extended Data Fig. 3c).
Airborne DNA for surveillance, host–parasite assessments
Bee and the honeybee mite (Varroa destructor) signals were examined as an example of multisite host–parasite air eDNA data recovery. Bees are vital for agriculture as honey producers and pollinators. The bee and mite eDNA were jointly recovered from a number of air eDNA samples, including the Irish windowpane swab (Extended Data Fig. 3d), highlighting the gravity of potential widespread bee colony infections. The 2023 city centre air sample collected in Trinity College Dublin had a large amount of Bombus genus DNA (Extended Data Fig. 3d), probably due to the university’s campus pollinator plan, which provides bee-friendly planting, management and housing practices. Airborne honeybee mite DNA was most abundant at the Meath estuary site (Extended Data Fig. 3d).
Airborne DNA for surveillance, allergens and narcotics
Overall, our sequencing across different environments revealed genetic information useful for applications beyond biodiversity assessments. For example, indoor air, outdoor air, water and tourist beach sand sampling also readily recovered DNA from common food items, suggesting utility for allergen scanning of enclosed spaces. Peanut (Arachis hypogaea) eDNA was frequently recovered from air samples, including Dublin City air. There was a strong positive correlation (R2 = 0.973) between the intrasample quantity of peanut eDNA recovered from the Irish air samples sequenced with both long-read and short-read technologies. Given the seasonal abundance shifts observed in tree eDNA from air shotgun sequencing, tree pollen was probably predominantly being detected (Extended Data Fig. 1d–f). Opium poppy (Papaver somniferum) DNA was identified in water eDNA from an Irish river (Avoca River RAMP23 C) polluted with human wastewater7,36, indicating a possible food- or narcotics-related origin. P. somniferum eDNA was also detected in Dublin City air in both the 2023 (2.9 nucleotides per million reads sequenced, nt_rpm) and 2024 (7.5 nt_rpm) samples. Of the Irish air sampling locations, Dublin City also had the highest level of Cannabis genus eDNA, with 22.9 nt_rpm and 2.1 nt_rpm in the 2023 and 2024 samples, respectively. Recreational cannabis use is not legal in Ireland, and medical use is under strict control. Psilocybe genus (‘magic mushrooms’) eDNA was also detectable in the 2024 Dublin air sample (17.8 nt_rpm).
Airborne DNA for phyla assessments
Previous studies appear to have underestimated the amount of eukaryotic and metazoan DNA recoverable from eDNA samples. Relatively high proportions of metazoan eDNA were recoverable by shotgun sequencing, and air samples tended to have higher metazoan loads than aquatic samples (Extended Data Fig. 4a). One-week pumped air samples also had higher numbers of eukaryotic than prokaryotic species detected (Fig. 5a). The reverse was true of water and sediment samples, although the proportion of eukaryotic species eDNA recovered from these was still higher than suggested by previous estimates39 (Fig. 5a). The broad taxonomic coverage achieved by eDNA shotgun sequencing enables direct comparisons between DNA within each sample originating from different phyla. Arthropods were the most dominant metazoan phylum recovered from air eDNA (Extended Data Fig. 4b–d). The Irish air samples were collected across a wide geographic distribution (Fig. 2a), yet arthropods, followed by chordates, were the most abundant animal phyla at each location (Extended Data Fig. 4b–d). Of these pumped air samples, Dublin City air had the lowest number of arthropod hits as a percentage of total metazoan hits (Extended Data Fig. 4d). Despite the phyla-level similarities, the metazoan species with the most counts varied between these eDNA samples (Supplementary Fig. 2a,b). For example, rainbow trout (Oncorhynchus mykiss) had the most counts in the Irish river water sample, dust mite (Blomia tropicalis) in the Dublin City centre air sample and midge (Sitodiplosis mosellana) in the Wicklow mountain air sample (Supplementary Fig. 2a,b). Taken together, these results suggest that shotgun sequencing is a feasible approach for eukaryotic and metazoan species, particularly for air eDNA samples.
a, The biodiversity assessments from air, water and sand eDNA conducted using long-read shotgun sequencing. Pan-biodiversity (excluding deuterostomes) OTU assessments of each sample type (minimum of two sequences required for OTU identification). All samples are from 2023, unless otherwise stated in the axis sample descriptor. Irel., Ireland; WL, Whitney Lab; WRI, Wildlife Rehabilitation Ireland; TCD, Trinity College Dublin. b, An investigation of chosen species in a single air eDNA long-read shotgun sequenced sample (1-week forest air, April 2023) taken in a mixed barrier-island hammock forest/residential area, Florida, as measured in nucleotides. The inset photos are fungus and anole images taken at the forest study site in 2024. The photos were taken by D.J.D.
Rapid biodiversity metagenomic assessment, non-deuterostome
Complex species communities in airborne DNA are readily retrievable by long-read shotgun sequencing from urban and non-urban, indoor and outdoor locations, using a range of sampling approaches: active air pumping (vacuum pump), passive air sampling (natural air flow over a filter), swabbing windowpanes or attaching a filter or membrane to a moving vehicle, as well as from water, soil and sand samples (Fig. 5a, Supplementary Fig. 1a and Supplementary Table 1). Rapid, cloud-based37,38 long-read metagenomic analyses revealed biodiversity information from all sample types, including microbial, fungal, plant and protostome-animal species abundances (Fig. 5a). Beach sand samples had among the highest species diversity, attributable to sand acting as a natural filter/accumulator of marine species DNA (Fig. 5a and Supplementary Table 1). However, when only considering CZ ID’s eukaryote database, the highest species diversity was recovered from air samples collected using an air vacuum pump for 7–8 days (929–1,069 operational taxonomic units (OTUs) per sample) and a swab from a windowpane (1,111 OTUs) that had been accumulating deposited airborne eDNA (and window surface-dwelling species) for 1 year (Fig. 5a). The highest pathogen diversity was from a Florida Gulf Coast seawater sample, Keaton Beach, Florida, followed closely by several outdoor pumped air samples and the Irish windowpane swab (Fig. 5a and Supplementary Table 1). The highest bacterial diversity from air eDNA sampling was recovered from the Irish city centre (Fig. 5a, Dublin City 2023). Notably, only minimal biodiversity was recovered from sequenced negative field control (NFC) samples (reagent contaminome) (Supplementary Fig. 3a).
Air eDNA biodiversity assessments, direct genome alignments
In principle, eDNA shotgun sequencing should provide an unbiased representation of the original proportions of DNA present in the filtrate, not being subject to inter- and intra-barcode amplification biases. Read assembly based metagenome reconstruction approaches such as metaFlye40 have been widely established for prokaryote communities. However, eukaryote footprints from longer and more complex genomes may require additional species identification precautions. Utilizing 6.5 million eDNA long-reads from a vacuum-pumped barrier-island hammock forest air sample (Forest April 2023, Fig. 5a), we assessed the presence of specific deuterostomes by directly mapping to 45 diverse animal reference genomes (Fig. 5b and Supplementary Table 2). While a substantial fraction of sequence fragments mapped to multiple genomes (46.5%), particularly among more closely related taxa, we also identified reads uniquely matching to any of the preselected references (Supplementary Fig. 3b). As expected, the number of genera called by metagenomic analysis reduces with more stringent sequence identity filters (Supplementary Fig. 3c). This probably arises due to a combination of false positives at lower sequence identities and because reference databases do not yet contain all species. Therefore, related species present in the database are currently called.
Simultaneous detection of difficult-to-study species such as cryptic (bobcat), nocturnal (bats and moths), predatory (rattlesnake and alligator), arboreal (squirrels), microscopic, native, invasive or aerial (birds, bats and butterflies) revealed a multilayered ecological fingerprint of the area (Fig. 5b). Moreover, predator and prey species (for example, bats and moths, bobcats and squirrels and osprey and fish) were detected in this same sample, as indeed were entire food webs (Fig. 5b). This notably includes several pathogenic, parasitic and disease vector species (and their hosts) of commercial, agricultural and human health concern (Fig. 5b). Agriculturally and economically beneficial species could also be detected (Fig. 5b). The recovered eDNA abundance for two similarly sized related species of anoles aligned well with known species abundance patterns (https://edis.ifas.ufl.edu/publication/UW486, https://animalia.bio/american-anole): invasive brown anoles (Anolis sagrei) are more common than native green anoles (Anolis carolinensis), and correspondingly, there was a 2.7-fold higher quantity of airborne brown anole DNA recovered (April 2023, Fig. 5b). For the three forest (from April 2024 to May 2024) and two Whitney Lab (July 2024 A and B) short-read shotgun sequenced air samples (Supplementary Table 1), brown anole eDNA was 1.4-fold to 2.2-fold higher than green anole eDNA. eDNA and individual abundance correlations warrant further investigation.
Short-read versus long-read air eDNA signals
Traditional eDNA analyses rely primarily on short-read sequencing with higher data throughput. To enable direct comparison, we sequenced six of the eDNA samples with both long-read and short-read sequencing protocols (Supplementary Table 1). Resulting metagenomic animal species compositions were broadly consistent between the two platforms (Extended Data Fig. 5a). Arthropods were the most abundant animal phylum in the dual-technology-sequenced Forest April 2023 air sample, followed by chordates (Extended Data Fig. 5a). In addition, for the 45 direct genome-aligned species (Supplementary Table 2), there was a strong positive correlation between the sequencing platforms (Extended Data Fig. 5b, log–log regression R2 = 0.64, Forest April 2023; log–log regression R2 = 0.68, Mountain July 2023). Similarly, when considering all genera, the CZ ID metagenomic pipeline detected a high proportion of overlap between long-read and short-read sequencing technologies (Extended Data Fig. 5c).
Shotgun sequencing versus vertebrate metabarcoding
When short-read shotgun sequencing (total extracted DNA) and 12S vertebrate metabarcoding (PCR followed by amplicon short-read sequencing) are conducted on the same air samples, both approaches recover vertebrate eDNA (Extended Data Fig. 6a and Supplementary Table 1). For the 18 vertebrate species compared, metabarcoding was more variable, with an individual vertebrate species range of 0–74% (0–107,851 amplicon sequence variant (ASV) count), while shotgun sequencing had a range of 0–23% (0–257 read counts) (Extended Data Fig. 6a). Of the 18 vertebrates compared, over 74% (sample A) and 19% (sample B) of vertebrate metabarcoding reads were from a single species, the Northern cardinal (Cardinalis cardinalis) (Extended Data Fig. 6a). Conversely, when shotgun sequenced, C. cardinalis only accounted for 0.4% (sample A) and 0.1% (sample B) of the total reads from these 18 vertebrates. Reptile species such as the American alligator (Alligator mississippiensis) were detected by shotgun sequencing but were not detected by the metabarcoding (Extended Data Fig. 6a). Alligator eDNA was not detected in any of the 14 metabarcoded air samples (Extended Data Fig. 6b), probably due to the barcode primer set having lower affinity for reptile sequences. Sample B had 7.5-fold more human eDNA than sample A (shotgun sequencing); however, barcode-based PCR amplification reduced this difference to 1.5-fold (Extended Data Fig. 6c). Metabarcoding boosted vertebrate signals, but this did not occur equally across all species, losing or diminishing the original species eDNA ratios present in the samples, as well as some vertebrates being entirely undetected, probably due to barcode and taxonomic database biases (Extended Data Fig. 6a–c).
Replicability of shotgun sequencing eDNA analysis
The pumped air samples collected together recovered a similar amount of pan-eukaryotic eDNA (Extended Data Fig. 7a), while for surface swabs collected together this level was more variable (Supplementary Fig. 4a). However, both the surface swabs and the air eDNA-based shotgun sequencing analysis showed a high degree of genus call overlap with their respective matching (replicate) samples (Extended Data Figs. 7b,c and 8a and Supplementary Fig. 4b). Even when a large portion of the material of one biological replicate was lost (spillage) before loading on a spin column during DNA extraction and a much lower yield of eukaryotic eDNA was obtained (Supplementary Fig. 5a), there was still a high degree of genus call overlap between the replicates (Supplementary Fig. 5b,c). This further highlights the sensitivity and power of deep-sequencing technologies. The samples taken together had a higher degree of overlap than the samples taken with temporal (at the same site at different times) or spatial (different sites) variation (Extended Data Figs. 7b,c and 8a–d). This was the case even when samples were taken for the same dates and duration from sites that were only 3.2 km apart on the same barrier island (Fig. 2a): the Florida coastal hammock forest and Whitney Lab (ocean–beach–saltmarsh–mangrove interface) (Extended Data Fig. 8d).
We detected overlaps between eDNA shotgun-sequencing-identified species in Florida sand, water and outdoor air collections (Supplementary Fig. 6a). This is unsurprising, given that species and their shed DNA can move between substrates and that the compared samples were taken on or adjacent to barrier-island habitats. However, the majority of species identified were unique to each substrate (Fig. 5a and Supplementary Fig. 6a,b). Beach sand recovered the largest number of species overall from ONT MinION shotgun sequencing (Fig. 5a and Supplementary Fig. 6b), although air eDNA had higher diversity of Protostomia than the other substrates (Fig. 5a and Supplementary Fig. 6a).
Reducing air eDNA extraction time
Finally, we assessed whether air eDNA extraction time could be cut in half, making near real-time applications even more feasible. Short (1–15 min) lysis steps are common in many cell, blood and viral applications, and DNA captured from the environment is suspected to be predominantly cellular, subcellular or cell free, with air being a relatively clean substrate. Therefore, we trialled a reduction from 1 h to 10 min for the 56 °C proteinase K step, having previously shortened this step from overnight (water and sand eDNA) to 1 h for air eDNA. Matching 48-h air samples taken consecutively (from December 2023 to January 2024) in the same location (Supplementary Table 1) showed no loss of pan-eukaryotic eDNA recovery when simultaneously extracted with either the 10 min or 1 h duration, as assessed by qPCR (Supplementary Fig. 7a). Vacuum pumping, swabbing outdoor surfaces or the collection of rainwater all recovered eukaryotic (Fig. 5a and Supplementary Figs. 4a and 7a) and prokaryotic eDNA (Fig. 5a and Supplementary Fig. 7b).
Overall, the accumulated data in this study highlight that the combination of air eDNA and shotgun deep sequencing generates rich datasets and enables a wide range of genomic diversity applications, down to population genetics, from air sampling alone.
Discussion
For extensive discussion and contextualization, please see the Supplementary Information.
Multiple ongoing crises, including biodiversity and habitat loss, increased pollution, reduction in arable land, mass extinction, animal and plant pandemics, invasive species, increase in antimicrobial resistant strains of pathogens, insecticide resistance and a changing climate, are altering the abundance and distribution of life and diseases across the planet1,2,3,4,5,41,42,43,44,45. Effective tools can advance understanding of the impacts of these changes across broad scales of life and record planet Earth’s genetic diversity before it is severely diminished or further portions of it are lost entirely. Emerging eDNA technologies offer a cost-effective scalable approach for non-invasively recording the remaining abundance of genetic diversity on the planet and understanding shifting biodiversity trends6,7,9,12,13,14,46. Shotgun-based lifeform and genetic diversity detection capabilities are applicable to environmental issues and species conservation and also have large-scale utility for economic, agriculture, aquaculture, law enforcement, biosecurity and medical applications7 (Table 1).
Staggering advances have been made in recent air eDNA biodiversity research7,12,13,14,15, but so far, this has primarily focused on metabarcoding, which provides presence information for a selected set of species. Here, PCR amplification bias was visible in air metabarcoding results, with some species becoming overrepresented, including the northern cardinal (C. cardinalis) and eastern grey squirrel (Sciurus carolinensis). Moreover, air eDNA shotgun sequencing recovered species known to be in the sampling vicinity, which metabarcoding failed to detect. Similar to metabarcoding approaches12,14,18, our shotgun sequencing of air samples captured spatial and temporal differences in species DNA abundance. Furthermore, shotgun air eDNA approaches rapidly recovered genetic information across broader groups of life simultaneously, from viruses to vertebrates. Beyond species identification, even in natural complex community settings, airborne DNA was of sufficient quality to enable in-depth genomic study and surveillance of many genetic traits of interest, such as antimicrobial, pesticide or herbicide resistance markers, sex ratios, phenotypic traits, pathogen variants, population genetics or human genome variant calling, including SNP level resolution (Box 1).
New tools for surveying environments for AMR, such as eDNA shotgun sequencing, are particularly required; AMR is recognized by the World Health Organization as one of the most important threats to global health, food security and development47. Simultaneously, shotgun sequencing of eDNA coupled with genome mining32,33 can provide a rapid tool for drug discovery34 bioprospecting, across the entire tree of life, negating the need for specimen collection and/or culture. Such eDNA-empowered pan-biodiversity bioprospecting/pan-genome mining could be termed ‘whole biome mining’.
Utilizing eDNA for population genetic approaches has been a major objective of the eDNA field. However, most of its success so far has come from individual enriched eDNA samples, for example, animal tracks or wakes6,7,8,10,48. We recently conducted human genetic diversity analysis from complex pooled community (species and individual mixed) eDNA samples (river water and indoor air eDNA)7. Here, we extend population genetics to airborne eDNA in outdoor natural community settings, analysing human genetic ancestry, a low abundance wildlife species (bobcat, L. rufus), and a medium abundance invertebrate (spider, T. clavipes). We also recovered single reads covering complete mitochondrial genomes, including for aerial species (for example, moth). Future studies should examine the potential of multilocus population genetics from air eDNA, through long-read sequencing of large somatic chromosome fragments (for example, single reads covering upwards of 50 kb regions7), target enrichment approaches (hybridization probe capture used here or long-read adaptive sequencing) or through single-cell and single-nucleus sequencing approaches.
Remote eDNA sensing by PCR-based detection is being trialled49, and portable sequencers are available. Near real-time detection is particularly feasible for airborne DNA; we streamlined extraction to less than 1 h, and 4G cellular connections from remote field sites were sufficient for eDNA metagenomic analysis. These findings reveal that we are closer than ever to the realization of a near real-time air-based lifeform detection device, as envisaged by Star Trek’s tricorder50,51.
Summary
Together, these results highlight the information-rich genetic data circulating through the open air and how it can be utilized to understand and address medical, commercial and environmental global challenges. Quantifying the abundance of airborne DNA revealed informative differences over space and time, from microscopic to macroscale lifeforms, simultaneously measuring changes in wildlife, human-associated animals, biodiversity, allergens, narcotics, pests, disease vectors (for example, mosquitoes), parasites and pathogens. Applications range from local to global scales, such as pathogen and allergen monitoring and genomic surveillance indoors through to mapping changes in pathogens and biodiversity across large habitats. Airborne eDNA genomics is equally well suited to the study of the growing genetic resistance of vectors, pests, parasites, and pathogens to drugs and chemicals used to control them, including insecticide resistance markers52 and AMR genes. Simultaneously, air eDNA metagenomics obtains information on populations of origin and genetic makeup. These approaches are particularly valuable for inaccessible study sites or dangerous, elusive, nocturnal, cryptic or aerial species and across all life stages. Sampling opportunities and strategies are widespread, with every windowpane in the world already probably acting as an airborne eDNA sampling device. Coupling airborne DNA recovery to rapidly developing remote sensing and sequencing technologies and cloud-based analysis platforms, the realization of a rapid lifeform and biological threat-detecting technology is closer than ever before. Similar to the atmosphere itself, the opportunities afforded by air eDNA genomics are near boundless.
Methods
Permitting
The analysis of human-derived eDNA from field obtained samples and human-related sampling was conducted with University of Florida (UF) Institutional Review Board (IRB-01) ethical approval under project number IRB202201336, with all study participants providing informed consent. One participant provided sand footprint samples, and six participants provided room air samples (pooled room air). Participation was on a voluntary basis, with no compensation received by the participants. Permitting for the collection of our water and sediment samples that were used as outgroups here is as reported in the publications for which those samples were originally collected6,7,8.
Sample collection
In this study, we generated 78 shotgun sequencing datasets (30 ONT and 48 Illumina), 4 human exome-enriched deep-sequencing datasets (Illumina), 14 vertebrate 12S metabarcoding datasets and assessed 62 samples by qPCR (Supplementary Table 1). Of the shotgun sequenced samples, nine were ONT outdoor active pump sampling, and 25 were Illumina outdoor active pump sampling. Sequenced outdoor samples ranged from 7 h to 35 days of pumping, with 1 week being the standard duration employed.
All laboratory procedures from sampling to final analysis (sequencing) were conducted in a way that minimized DNA contamination from investigators or other sources. This included no contact of investigators/samplers with substrates being sampled (water, sand or air), new nitrile gloves for each sample collection and frequent glove changes throughout sample processing, cleaning of equipment and benchtops with bleach before use, and NFCs being treated identically to genuine field samples throughout all processes from extraction through to sequencing or qPCR. All eDNA samples were extracted in laboratories that do not process other biodiversity DNA samples: a sea turtle laboratory in Florida and a chick/mouse developmental biology laboratory in Ireland.
All outdoor air samples and Irish sand samples were collected for this study. The outdoor air samples were collected by one of three methods: 0.22-µm pore Millipore Sterivex-GP Pressure Filter Units (Merk Millipore, cat no. SVGPL10RC) (1) attached to a vacuum pump (Fisherbrand MaximaDry (115 V, 40 l min−1, cat. no. 1388016) or Fisherbrand MaximaDry (230 V, 20 l min−1, cat. no. 1388015)) and filtered for a specific duration (active pump sampling), (2) left in an environment to passively absorb airborne DNA (passive sampling), or (3) taped to the roof of a vehicle with air passing through the filter while the vehicle drove (car mounted, Ford Explorer) (Supplementary Fig. 1a). The Irish air vacuum pump samples were collected for one extra day compared with the Florida 7-day samples, to partially offset the use of a slightly weaker vacuum pump. The car-mounted samples were not actively vacuum pumped but rather air passed through freely as the vehicle drove (Fig. 2a and Supplementary Fig. 1a). The car-mounted sampling route was a round trip (4 h 20 min of driving) from our research institute to our main campus sequencing facility (short amount of barrier-island driving but predominantly inland habitats that included urban towns, agricultural areas and conifer forests). Air NFCs had filters placed in the same environment (adjacent to sampling filters) for the same duration but with luer-lock caps in place to prevent any airflow through the filter.
Non-Irish seawater and beach sand eDNA samples were collected between 2018 and 2023 and sequenced as part of our human, sea turtle and pathogen research6,53. The soil samples were collected (2022) as part of a bobcat eDNA study8. Indoor air samples from our sea turtle hospital were collected (with ethical approval from the UF IRB, 2022–2023) as part of a human eDNA study (2022 and 2023)7, but the indoor air human exome enrichment sequencing was conducted for this current study. The NFC samples of water and sand were also collected and processed as per the study samples. For the NFC water sampling, 1 litre of MilliQ water (Florida) or 1 litre of Qiagen Nuclease-free water (Ireland, cat. no. 129117) was transported from the laboratory to field sampling locations and stored in a cool box with the environmental samples to monitor for potential contamination during sampling, transportation and processing7. The air and water NFCs were filtered and extracted alongside the other collected air and water samples from each sampling trip and subjected to the same next-generation sequencing or qPCR conditions. Between 300 ml and 1 litre of water samples were filtered (Supplementary Table 1); lower volume samples had been filtered until debris clogging prevented larger volumes being filtered. For low debris samples (for example, February 2024 rainwater) a maximum of 1 litre was filtered, despite filters not becoming clogged. For sand/soil eDNA, a 50-ml tube was filled with sand from each sampling event, with 10 ml of this sand/soil used per individual eDNA extraction6. Human-present (sea turtle rehabilitation hospital) air samples were collected from a 280-square-foot room while participants went about their daily work activities (that is, could enter and exit the room throughout sampling period), with a maximum of the same six participants using the room for a portion of the sampling period7. The room was air conditioned (outside air) and had an external door that was opened and closed and occasionally left open for certain work procedures. Indoor air was also collected from an office-style building on a barrier island (24 h vacuum pumped, 24 h office air May 2023), which had been temporarily closed (no occupants) to facilitate old ceiling tile and roof insulation removal. The NFC samples of air were also collected and processed as per the study samples. For outdoor air sampling, pumps were only started once investigators had left the air filter inlet location, and pumping was stopped before the recovery of the filters.
DNA extraction and sequencing
Before filtration and between every sample, laboratory surfaces, filtration equipment and all standard laboratory equipment involved in the washing and filtration process, as well as the filtration pump itself were disinfected with 70% ethanol (general cleaning and biotic disinfection), and sampling equipment (collection bottles) was disinfected by washing thoroughly with 10% bleach (DNA destruction) and rinsed thoroughly with deionized water.
Outdoor air
The outdoor air samples were collected as described above. Immediately after air sampling, 740 µl of Buffer ATL was added into the filter units, and the samples were either immediately extracted or filters stored at −20 °C in Buffer ATL until extraction. Next, 20 µl proteinase K (from 1 h to <24 h samples) or 60 µl proteinase K (samples collected for >24 h) was added to each filter (after filter thawing if it had been stored frozen) and the 56 °C ATL Buffer and proteinase K incubation was conducted for 1 h. For four samples, the 1-h incubation was reduced to a 10-min incubation to assess the feasibility of reduced extraction times for outdoor air samples.
Sand
For sand samples, 1× concentration Tris and EDTA (TE) buffer (IDTE pH 8.0 1× TE solution) (Integrated DNA Technologies, cat. no. 11-05-01-09) was added to each individual sand sample in individual 50-ml Falcon conical centrifuge tubes, at approximately two times the volume of sand (10 ml of sand and 20 ml of 1× TE buffer). The samples were shaken gently by hand, then set on a rocking platform for 1 h at room temperature. The samples were rested until sand had sunk to the bottom of each tube, then the supernatant was immediately pipetted into a 60-ml sterile BD luer-lock syringe (Fisher Scientific, cat. no. 136898). The samples were then hand filtered using 60-ml BD luer-lock syringes through 0.22-µm Sterivex-GP Pressure Filter Units (Millipore, cat. no. SVGPL10RC) and capped with B.Braun luer-lock caps (Medline, cat. no. BMGTMR2000B). A total of 740 µl Buffer ATL and 60 µl proteinase K from a Qiagen DNeasy Blood and Tissue Kit (Qiagen, cat. no. 69504) were added to each sample, and they were placed in 50-ml Falcon conical centrifuge tubes in a rolling incubator overnight at 56 °C.
River/seawater
The water samples were pumped (by hand, Ireland and Florida 2023, or electronically, Florida before 2023) through 0.22-µm Sterivex-GP Pressure Filter Units (Millipore, cat. no. SVGPL10RC) and capped with B.Braun luer-lock caps (Medline, cat. no. BMGTMR2000B). Hand pumping was with sterile 60-ml BD luer-lock syringes. The electronic pumping was carried out using a GeoTech Peristaltic Pump Series II. A total of 740 µl of Buffer ATL and 60 µl of proteinase K from a Qiagen DNeasy Blood and Tissue Kit were added to each sample. and they were placed in 50-ml Falcon conical centrifuge tubes in a rolling incubator overnight at 56 °C.
Rainwater
Rainwater was collected using sterile 1-litre bottles and sterile funnels (Fisherbrand utility 203 mm funnels, cat. no. 1050010) at the Florida hammock forest site (placed on a balcony in an open canopy area with no tree cover). Rainwater was collected in February, August (category 1 Hurricane Debby) and September 2024 (category 4 Hurricane Helene) 2024. The sampling duration ranged from 19 h and 36 min to 20 h and 15 min, and the sampling was stopped once rain ended. The rainwater volumes ranged from 1 litre to 0.45 litre, depending on rainfall volumes. The collected rainwater was then filtered immediately and eDNA extracted as described above for river water, the only exception being that due to the relative clarity of the sample the 56 °C step was conducted only for 2 h and 30 min. For the rainwater NFCs, 1 litre of MilliQ water in a capped sterile bottle was placed beside the rainwater sampling bottles for the duration of sampling and an equal volume processed identically alongside the rainwater samples.
Indoor air
For indoor air eDNA sampling, room air was passed through 0.22-µm pore Millipore Sterivex-GP Pressure Filter Units (Merk Millipore, cat. no. SVGPL10RC), using a Welch vacuum pump (2019LD-4112), a GeoTech Peristaltic Pump Series II or a Fisherbrand MaximaDry vacuum pump (cat. no. 1388016). The eDNA was extracted6,53 as for the water and sand samples, with the exception that only 20 μl of proteinase K was added per filter, and the 56 °C ATL Buffer and proteinase K incubation was conducted for 1 h.
Non-filter based collection
For the car-mounted (Ford Explorer, Supplementary Fig. 1a) air eDNA collection syringe-only sample (4.3 h car-mounted syringe, May 2023), the car-mounted sterile syringe (BD luer-lock syringe, Fisher Scientific, cat. no. 136898) was capped (leur lock cap) and the walls of the syringe washed (pipetting and hand rotation) with 2,960 µl of ATL Buffer. The syringe was then uncapped, and a sterile plunger was used to pass the liquid into a sterile 15-ml tube. A total of 80 μl of proteinase K was then added to the ATL Buffer, and the sample was incubated for 1 h at 56 °C. The Irish windowpane swab (Wicklow Mountain, May 2023) was collected from a windowpane (outdoor facing side, exposed to the elements) at the air sampling site (Supplementary Fig. 1a). The windowpane had been washed 1 year prior (with water and detergent), so it had been accumulating DNA for 1 year. A sterile swab was dipped in ATL Buffer and used to swab a small section of the windowpane. For the Florida swabs, a variety of surfaces at the hammock forest air sampling site were similarly swabbed within 10 min of each other: windowpane, varnished wooden plank, metal fence post and a Palmetto palm (Sabal palmetto) leaf. For all the swab samples, an area of approximately 3 cm2 was swabbed for 20 s. The swabs were immediately placed in a sterile 50-ml Falcon conical centrifuge tubes containing 500 µl of ATL Buffer and stored at −20 °C. For the swab NFCs, a swab was placed directly into a sterile 50-ml tube containing ATL, without being used to swab any surface. After thawing, 60 μl of proteinase K was added to the ATL Buffer, and the sample was incubated for 1 h at 56 °C.
For all three filtered sample types (sand, water and air), after the 56 °C incubation, the solutions of Buffer ATL (post water or air filtration or post sand washing and filtration), proteinase K and eDNA were transferred from the Sterivex-GP Pressure Filter Units to 2-ml microcentrifuge tubes using 10 ml of BD Slip Tip Sterile Syringes (Fisher Scientific, cat. no. 14823434). The DNA was then isolated using a modified Qiagen DNeasy Blood and Tissue Kit protocol6,53,54. Following incubation, equal volume 800 µl (sand and water) AL Buffer and 800 µl (sand and water) ice-cold ethanol were added to each sample, and they were vortexed vigorously and microcentrifuged after each addition. For air samples, 500 µl of each solution was used (as a lower volume of ATL solution is recovered from the initially dry Sterivex filters). For swab samples, 500 µl of each solution was used (directly into the 50-ml tube). Each sample was extracted according to the remainder of the kit protocol, with the only alteration being that DNA was eluted with 70 µl of AE Buffer, incubated at 70 °C before addition to spin column and incubated on the column at room temperature for 7 min before final centrifugation. The DNA concentration was measured on a ThermoScientific Nanodrop 2000 Spectrophotometer (Fisher Scientific), and the samples were stored at −20 °C until shotgun sequencing or qPCR.
Library preparation and Florida Illumina shotgun sequencing was conducted at the UF’s Interdisciplinary Center for Biotechnology Research (ICBR) Core Facilities on an Illumina NovaSeq 6000 (from 2022 to July 2023, 2× 150-bp PE sequencing on an S4 flow cell) or NovaSeq Series X Plus (from October 2023 to 2024, 2× 150-bp PE sequencing on an 25B flow cell) (see Supplementary Table 1 for the sequencing machine utilized for each sample). Indoor air human Illumina exome capture libraries were also constructed and sequenced at the UF ICBR Gene Expression and NextGen Sequencing Cores. Four indoor air eDNA samples (two air eDNA samples and two NFCs) were used for Illumina DNA Prep with Enrichment, Exome Panel (cat. no. 20020183, captures 45 Mb of exonic content) exome capture analysis, according to the manufacturer’s instructions. Exome sequencing was conducted on an Illumina NovaSeq 6000 S4 flow cell for 2× 150-bp cycles (with NFCs expected to return fewer reads due to a lack of human eDNA) (Supplementary Table 1). The Irish samples were shotgun sequenced commercially at BMKGENE, with Illumina libraries sequenced on a NovaSeq 6000 (2023 sequencing, 2× 150-bp PE, eight PCR amplification cycles during library preparation) or a NovaSeq Series X Plus (2024 sequencing, 2× 150-bp PE sequencing on an 25B flow cell) and ONT libraries sequenced on a PromethION 48. All Florida Oxford nanopore samples were sequenced on a MinION in the Duffy lab at the UF’s Whitney Laboratory for Marine Bioscience. MinION libraries were prepared according to manufacturer’s instructions, using the following kits: ONT Ligation Sequencing Kit (SQK-LSK110, cat. no. 76487-106 or SQK-LSK114), NEBNext Companion Module for ONT Ligation Sequencing Kit (cat. no. E7180S), ONT Flow Cell Wash Kit (EXP-WSH004, cat. no. 76487-116) and sequenced on ONT Minion Flow Cells (cat. no. 76487-106). The samples sequenced up to May 2023 were with v10 chemistry (Ligation Sequencing Kit, SQK-LSK110) and R9.4.1 flow cells, while samples sequenced after May 2023 were sequenced with v14 chemistry (SQK-LSK114) and R10.4.1 flow cells (FLO-MIN114.8) (Supplementary Table 1). The MinION samples sequenced after December 2023 used the new ONT light shield (Supplementary Table 1 and see Supplementary Fig. 7c for indicative read length distributions). Throughout the project, the latest sequencing approaches (chemistries/machines) were applied as they became available, to represent the most current technology and to avail of the associated cost savings.
The vertebrate metabarcoding was conducted according to Lynggaard et al.55 using the 12SV05 vertebrate mitochondrial metabarcoding primer set 12SV05 forward: TTAGATACCCCACTATGC and 12SV05 reverse: TAGAACAGGCTCCTCTAG14,55,56. Input DNA (air eDNA or NFC) was amplified on a Bio-Rad C1000 Touch Thermal Cycler with AmpliTaq Gold DNA Polymerase (ThermoFisher, cat. no. N8080241) for the following cycles: 1 cycle of 95 °C for 10 min, then 45 cycles of 94 °C for 30 s, 51 °C for 30 s and 72 °C for 1 min. The PCR products were then purified using a QIAquick PCR Purification Kit (Qiagen, cat. no. 28104), according to the manufacturer’s instructions. The PCR products were used to construct Illumina sequencing libraries at the UF ICBR Gene Expression core facility, and the libraries were paired-end sequenced (2× 150 bp) at the UF ICBR Next-Generation Sequencing core facility, on an Illumina NovaSeq Series X Plus (10B flow cell). Equal pooling ratios utilized 4.42% of a lane for these three 14 eDNA samples and 65 other metabarcoding samples (not for this study). Each of the 14 samples had 0.03–0.09% of the final lane output (or 466,890–1,285,665 reads).
Bioinformatic analysis
All bioinformatic tools were utilized using default parameters, unless otherwise stated.
Non-deuterostome metagenomics
Nanopore fast-based calling with Dorado was utilized. CZ ID37,38, a free, cloud-based metagenomics platform (https://czid.org/) was utilized for biodiversity bioinformatic analysis (ONT and Illumina). Approximately half of the ONT samples analysed were uploaded to the CZ ID bioinformatics pipeline through a mobile/cellular internet connection (Huawei B525s-23a domestic cellular connection router with a Three Ireland mobile network SIM card, over an older 4G network) from our Irish Croghan Mountain field site. The CZ ID platform queries genetic information from all species, excluding deuterostomes. The CZ ID nanopore metagenomic pipeline was utilized for ONT samples. The CZ ID Illumina metagenomic pipeline was utilized for short-read samples. As the CZ ID platform was primarily developed for microbial metagenomics, it does not query deuterostome sequences, as these are identified as host species, and it actively subtracts host reads such as human reads. Moreover, for samples with over 1 million reads, the pipeline actively subsamples only 1 million reads per analysis. OTUs from the CZ ID data were identified twice: those identified with a single sequence and those identified by at least two. The heat maps were generated using the CZ ID heatmap tool. The CZ ID platform38 was also used to determine the presence of AMR genes (CZ ID AMR Pipeline v1.3.1) in Illumina shotgun sequenced samples (Supplementary Table 1).
Metazoan metagenomics (including deuterostomes)
To conduct metazoan long-read and short-read metagenomics, including deuterostomes, the CZ ID nanopore and CZ ID Illumina metagenomic pipelines were recreated, respectively, in a high-performance computing (HPC) environment using Bash and Python, without host filtering, without subsampling to 1 million reads and aligning to all metazoan species (NCBI Blast). To classify metagenomic reads (both long and short) based on taxonomy, DIAMOND v2.1.7 (ref. 57) was utilized for its inherent high-speed alignment, sensitivity and the translation of nucleotide reads for protein alignments capabilities. First, metagenomic reads from eDNA samples were preprocessed, for adaptor trimming and quality control using Trim galore v0.6.7 (Trim Galore! (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/)) for Illumina or Porechop v0.2.4 (ref. 58) for ONT reads. The two-step DIAMOND process involved database preparation and read alignment. A reference database was constructed using the non-redundant protein sequences from NCBI. Taxonomic information was integrated using DIAMOND’s ‘makedb’ option with the following input files: a protein accession to taxonomy ID mapping file prot.accession2taxid (https://ftp.ncbi.nih.gov/pub/taxonomy/accession2taxid/), a taxonomy nodes file nodes.dmp (https://ftp.ncbi.nih.gov/pub/taxonomy/new_taxdump/) and a taxonomy names file names.dmp (https://ftp.ncbi.nih.gov/pub/taxonomy/new_taxdump/). For both long-read and short-read samples, the processed reads were aligned to the prepared database using DIAMOND’s blastx mode with ‘very-sensitive’ option, ensuring comprehensive taxonomic classification as per the most stringent options of the package. This step translated nucleotide sequences into protein sequences, aligning them against the database. The alignment output format was specified so as to include detailed taxonomic information for each read. Only the best hit for each read was retained using the option ‘max-target-seqs 1’. The results were parsed with in-house python script to aggregate reads on the basis of the NCBI taxonomy structure for different phyla of protostomes and deuterostomes. This reduced version of the CZ ID pipeline was used to overcome the read subset limitations and the exclusive identification of eukaryotes. Therefore, the contig generation step of the pipeline was not implemented, as it is not feasible to reconstruct species-specific contigs for eukaryotes due to the large genome sizes, the pooled nature and the unequal representation of species in eDNA NGS data. The metazoan metagenomics bioinformatics pipeline (MetaBioTax v1.0) has been deposited in GitHub (https://github.com/nousiaso/MetaBioTax, https://gitfront.io/r/nousiaso/QZwYQLPm9dJQ/MetaBioTax/).
Direct reference genome and mitogenome alignments
The Galaxy platform58,59 (https://usegalaxy.eu/) was utilized for bioinformatic analysis with selected reference genomes (Supplementary Table 2), with NanoGalaxy also utilized for nanopore sequenced data60. All samples were checked for quality (FastQC, v0.73 (ref. 61)), adaptors and low-quality reads trimmed (<20 quality score) (Trim Galore! (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/) v0.6.7 for Illumina data; Porechop v0.2.4 (ref. 62) for ONT sequence data), and high-quality reads aligned to the human reference genome (T2T-CHM13v1.1 (ref. 63)) using Bowtie2, v2.4.2 (ref. 64) for Illumina or minimap2, v2.24 (ref. 65) for ONT. Paired-read alignments were conducted for Illumina data and single read alignments for ONT data. The trimmed air eDNA nanopore data were also aligned (minimap2, v2.24 (ref. 65)) to reference genomes (Supplementary Table 2). Human mitochondrial haplogroups were classified from Illumina data using Haplogrep (V3.2.1)66 with the Phylotree 17.2 (ref. 67) and the Kulczynski distance with only sequences passing quality control (>90%) retained. Human mitochondrial haplogroup charts were produced in RStudio68, using the webr package v0.1.5 (https://github.com/cardiomoon/webr). Moth (C. sasakki) and Loblolly pine (Pinus taeda) mitochondrial and chloroplast sequences (ONT data) were concatenated in Geneious, and the genome coverage plots were created using the ggplot package v3.4.0 (ref. 69). Illumina mitochondrial sequences of the bobcat (L. rufus) and the golden silk orb weaver spider (T. clavipes) were aligned to Genbank references (Supplementary Table 3) using the Geneious Mapper with maximum sensitivity and slow mapping options selected (GeneiousPrime 2023.0, www.geneious.com/prime). Jukes–Cantor maximum-likelihood trees were similarly created in GeneiousPrime with automatic sequence determination, global alignments with free end gaps and Identity (1.0/0.0) Cost Matrix options selected, using cougar (Puma concolor) and Joro spider (Trichonephila clavata) as outgroups for the bobcat and spider trees, respectively. UpSet plots and Venn diagrams were created using the ‘UpSetR’ (V. 1.4.0) and venneuler packages (X.x). Area-proportional Venn diagrams of genus lists, with 20 reads per million total reads and eukaryotic genera-only cutoffs, were generated using BioVenn70.
Human exome-based variant analysis
The Illumina human exome variant analysis was performed using the nf-core Sarek pipeline71, leveraging Nextflow for workflow management and Docker for consistent execution environments. The preparation for base quality score recalibration involved generating the necessary files required for the recalibration process using the GATK.GRCh38 reference genome. This step ensured that the recalibration could be accurately performed, correcting any systematic biases introduced during sequencing. The base quality scores of the sequencing reads were recalibrated to adjust for any biases introduced during the sequencing process. This recalibration, performed using the GATK.GRCh38 reference genome, improved the accuracy of the quality scores, leading to more reliable variant calling. Variant calling was performed using the HaplotypeCaller tool from GATK, with the GATK.GRCh38 reference genome. This step involved identifying potential variants, including SNPs and insertions/deletions (indels), from the recalibrated sequencing data. The coordinates from the phase 1 high confidence SNPs, and known indels at https://genome.ucsc.edu/cgi-bin/hgTrackUi?db=hg19&g=tgpPhase1 were used to filter the variant calls. The identified variants were then outputted in VCF format for further analysis. The final step involved annotating the identified variants using the Variant Effect Predictor with the GATK.GRCh38 reference genome. This annotation process provided functional information about the variants, such as their potential impact on genes and proteins, which aids in the interpretation of their biological importance.
Assessment of reference genome specificity and read clustering of ONT eDNA reads
Nanopore eDNA reads were prefiltered via the CZ ID ONT mNGS pipeline v0.7. We then used minimap v2.26 (ref. 65) to align 6.5 million reads (Forest air April 2023) and 1.4 million reads (Mountain air July 2023) (‘sample.humanfiltered.fastq’) against a set of 45 publically available reference genomes from NCBI RefSeq (Supplementary Table 2). To provide a more stringent categorization and further minimize human sequence contamination risks, we also aligned reads against the human telomere-to-telomere (T2T-CHM13v2.0) assembly63. All resulting files were indexed, binarized and sorted using samtools v1.17 (ref. 72). We extracted the IDs of all nanopore sequences mapping to any of the 45 species but not T2T-CHM13v2.0 (378,856 reads, 5.8% of the original set) by use of samtools v1.17, before generating a binary presence/absence matrix of reads versus reference genomes in R. Given the large size of this matrix, we undertook two hierarchical clustering experiments on down-sampled representations of nanopore reads: (1) group pruning: for a pruned representation of the sequences mapping to different individual species or shared subsets of reference genomes, we reduced each fraction to a maximum of ten reads, thereby reducing the dimension to a total of 1,100 reads, and (2) random sampling: to provide a quantitative perspective of the different set sizes, we uniformly sampled 1,100 reads from the original matrix. The matrix entries were then hierarchically clustered and displayed using the pheatmap function in R.
Metabarcoding analysis
The metabarcoding 12S rRNA gene sequence reads were initially examined for quality scores with FastQC61 and primers were trimmed using Cutadapt73. All sequences were subsequently processed using the DADA2 pipeline74. Briefly, reads were filtered out (maxN = 0, maxEE = c(2, 2), truncQ = 2, minLen = 50), and remaining unpaired sequences were then denoised and aligned. The resulting ASV table was then cross-referenced against the 12S Vertebrate North American Reference Set75 (https://doi.org/10.5281/zenodo.7888133), using the ‘blast_nucleotide_to_nucleotide’ function in the ‘metablastr’ R-package76, with a >99% identity cut off.
qPCR
Florida
A pan-eukaryotic 18S ribosomal RNA (rRNA) gene (Applied Biosystem, 4352930E) prevalidated Taqman Gene Expression assay, which also has both primers and probe within a single exon (that is, can be used to detect genomic DNA or complementary DNA), was used to quantify the total level of pan-eukaryotic gDNA in samples6,7,8,53. The qPCR reaction mixtures were performed on 384-well plates in a total volume of 10 μl per well: 5 μl TaqMan Fast Advanced Master Mix (Fisher Scientific, cat. no. 4444557); 3.5 μl Nuclease-free water (Fisher Scientific); 0.5 μl of the 18 s rRNA assay (at manufacturer-supplied concentration); 1 μl DNA template (or, for no-template controls, an additional 1 μl of nuclease-free water per well). Each air sample or NFC was run in three technical replicates. No-template controls were run in triplicate on every qPCR plate. The qPCR reactions were performed on an Applied Biosystems QuantStudio 6 Pro with the following cycling parameter: 95 °C for 20 s for one cycle, followed by 30 cycles of 95 °C for 1 s and 60°C for 20 s. Serial dilution of total DNA that had previously been extracted from green sea turtle (Chelonia mydas) blood was used as the template for the standard curve. The serial dilution range was 621 ng μl−1 to 0.00621 ng μl−1, with a 1-μl template used per well.
Ireland
The Applied Biosystem prevalidated Taqman Gene Expression qPCR assay directed against the human ZNF285 gene (assay ID Hs00603276_s1) was used as a species-specific human assay, based on having no cross-reactivity with over 29 other species from mice to plants (https://www.thermofisher.com/order/genome-database/ and refs. 7,31) and having both primers and probe within a single exon (that is, detect DNA). A pan-eukaryotic 18S rRNA gene (Applied Biosystem, 4319413E) prevalidated Taqman Gene Expression assay, which also has both primers and probe within a single exon (that is, detects DNA), was used to quantify the total level of pan-eukaryotic DNA in each of the Irish samples7. The qPCR reaction mixtures were performed on 384-well plates in a total volume of 10 μl per well: 5 μl TaqMan Gene Expression Master Mix (Fisher Scientific, cat. no. 4369016), 3.5 μl nuclease-free water (Fisher Scientific), 0.5 μl of the respective assay (primer/probe mix, manufacturer-supplied concentration) and 1 μl DNA template (or, for no-template controls, an additional 1 μl of nuclease-free water per well). Each biological sample, NFC and standard curve point was run in three technical replicates. No-template controls were run in triplicate on every qPCR plate. The qPCR reactions were performed on an Applied Biosystems QuantStudio 7 Flex, at the Conway Institute’s Genomics Core, University College Dublin, with the following cycling parameter: 50 °C for 2 min and 95 °C for 10 min for 1 cycle, followed by 45 cycles of 95 °C for 15 s and 60 °C for 1 min. For absolute quantity qPCRs, a standard curve was generated using six DNA one-in-ten serial dilutions, ranging from 111.3 ng μl−1 to 0.001113 ng μl−1. A 1-μl template was used per standard curve reaction. The standard curve template used for both pan-eukaryotic and human-specific qPCRs was DNA extracted from cultured human IMR32 neuroblastoma cells in May 2013 for the Duffy et al. 2018 study77. For each standard curve concentration and each eDNA sample, technical triplicate wells/reactions were performed.
qPCR results were plotted with BoxPlotR78 (http://shiny.chemgrid.org/boxplotr/) with every datapoint displayed. Tukey whiskers (extend to data points that are less than 1.5× interquartile range away from the first and third quartile) were utilized for every boxplot. One box is graphed per single sample, consisting of all qPCR technical replicate wells for that sample.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequenced samples including raw reads are available via NCBI (https://www.ncbi.nlm.nih.gov/) under BioProject IDs PRJNA956956 (shotgun, https://www.ncbi.nlm.nih.gov/bioproject/PRJNA956956) and PRJNA1175642 (metabarcoding, https://www.ncbi.nlm.nih.gov/sra/PRJNA1175642). See Supplementary Table 1 for individual NCBI SRR accession numbers for each sequenced sample.
Code availability
The metazoan metagenomics bioinformatics pipeline (MetaBioTax v1.0) is available via GitHub at https://github.com/nousiaso/MetaBioTax and https://gitfront.io/r/nousiaso/QZwYQLPm9dJQ/MetaBioTax/.
References
Razgour, O. et al. An integrated framework to identify wildlife populations under threat from climate change. Mol. Ecol. Resour. 18, 18–31 (2018).
Ceballos, G., Ehrlich, P. R. & Raven, P. H. Vertebrates on the brink as indicators of biological annihilation and the sixth mass extinction. Proc. Natl Acad. Sci. USA 117, 13596–13602 (2020).
Hogg, C. J. Translating genomic advances into biodiversity conservation. Nat. Rev. Genet. 25, 362–373 (2023).
Xu, H. et al. Ensuring effective implementation of the post-2020 global biodiversity targets. Nat. Ecol. Evol. 5, 411–418 (2021).
Driscoll, D. A. et al. A biodiversity-crisis hierarchy to evaluate and refine conservation indicators. Nat. Ecol. Evol. 2, 775–781 (2018).
Farrell, J. A. et al. Detection and population genomics of sea turtle species via non-invasive environmental DNA analysis of nesting beach sand tracks and oceanic water. Mol. Ecol. Resour. 22, 2471–2493 (2022).
Whitmore, L. et al. Inadvertent human genomic bycatch and intentional capture raise beneficial applications and ethical concerns with environmental DNA. Nat. Ecol. Evol. 7, 873–888 (2023).
Koda, S. A. et al. A novel eDNA approach for rare species monitoring: application of long-read shotgun sequencing to Lynx rufus soil pawprints. Biol. Conserv. 287, 110315 (2023).
Giolai, M. et al. Air-seq: measuring air metagenomic diversity in an agricultural ecosystem. Curr. Biol. 34, 3778–3791.e4 (2024).
Urban, L. et al. Non-invasive real-time genomic monitoring of the critically endangered kākāpō. eLife 12, RP84553 (2023).
Crits-Christoph, A. et al. Genetic tracing of market wildlife and viruses at the epicenter of the COVID-19 pandemic. Cell 187, 5468–5482.e11 (2024).
Littlefair, J. E. et al. Air-quality networks collect environmental DNA with the potential to measure biodiversity at continental scales. Curr. Biol. 33, R426–R428 (2023).
Bohmann, K. & Lynggaard, C. Transforming terrestrial biodiversity surveys using airborne eDNA. Trends Ecol. Evol. 38, 119–121 (2023).
Lynggaard, C. et al. Airborne environmental DNA for terrestrial vertebrate community monitoring. Curr. Biol. 32, 701–707.e5 (2022).
Serrao, N. R., Weckworth, J. K., McKelvey, K. S., Dysthe, J. C. & Schwartz, M. K. Molecular genetic analysis of air, water, and soil to detect big brown bats in North America. Biol. Conserv. 261, 109252 (2021).
Rowney, F. M. et al. Environmental DNA reveals links between abundance and composition of airborne grass pollen and respiratory health. Curr. Biol. 31, 1995–2003.e4 (2021).
Clare, E. L. et al. Measuring biodiversity from DNA in the air. Curr. Biol. 32, 693–700. e5 (2022).
Garrett, N. R. et al. Airborne eDNA documents a diverse and ecologically complex tropical bat and other mammal community. Environ. DNA 5, 350–362 (2023).
Johnson, M. D., Cox, R. D. & Barnes, M. A. The detection of a non-anemophilous plant species using airborne eDNA. PLoS One 14, e0225262 (2019).
Johnson, M. D., Fokar, M., Cox, R. D. & Barnes, M. A. Airborne environmental DNA metabarcoding detects more diversity, with less sampling effort, than a traditional plant community survey. BMC Ecol. Evol. 21, 218 (2021).
Johnson, M. D., Barnes, M. A., Garrett, N. R. & Clare, E. L. Answers blowing in the wind: detection of birds, mammals, and amphibians with airborne environmental DNA in a natural environment over a yearlong survey. Environ. DNA 5, 375–387 (2023).
Aalismail, N. A., Díaz-Rúa, R., Geraldi, N., Cusack, M. & Duarte, C. M. Diversity and sources of airborne eukaryotic communities (AEC) in the global dust belt over the Red Sea. Earth Syst. Environ. 5, 459–471 (2021).
Klepke, M. J., Sigsgaard, E. E., Jensen, M. R., Olsen, K. & Thomsen, P. F. Accumulation and diversity of airborne, eukaryotic environmental DNA. Environ. DNA 4, 1323–1339 (2022).
Abrego, N. et al. Airborne DNA reveals predictable spatial and seasonal dynamics of fungi. Nature 631, 835–842 (2024).
Lynggaard, C. et al. Vertebrate environmental DNA from leaf swabs. Curr. Biol. 33, R853–R854 (2023).
Aucone, E. et al. Drone-assisted collection of environmental DNA from tree branches for biodiversity monitoring. Sci. Robot. 8, eadd5762 (2023).
Gregorič, M. et al. Spider webs as eDNA samplers: biodiversity assessment across the tree of life. Mol. Ecol. Resour. 22, 2534–2545 (2022).
Newton, J. P., Nevill, P., Bateman, P. W., Campbell, M. A. & Allentoft, M. E. Spider webs capture environmental DNA from terrestrial vertebrates. iScience 27, 108904 (2024).
Andres, K. J., Lodge, D. M., Sethi, S. A. & Andrés, J. Detecting and analysing intraspecific genetic variation with eDNA: from population genetics to species abundance. Mol. Ecol. 32, 4118–4132 (2023).
Reska, T. et al. Air monitoring by nanopore sequencing. ISME Commun. 4, ycae099 (2024).
McCauley, M., Koda, S. A., Loesgen, S. & Duffy, D. J. Multicellular species environmental DNA (eDNA) research constrained by overfocus on mitochondrial DNA. Sci. Total Environ. 912, 169550 (2024).
Atanasov, A. G. et al. Natural products in drug discovery: advances and opportunities. Nat. Rev. Drug Discov. 20, 200–216 (2021).
Chen, J. et al. Global marine microbial diversity and its potential in bioprospecting. Nature 633, 371–379 (2024).
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs over the last 25 years. J. Nat. Prod. 70, 461–477 (2007).
Marx, V. Method of the year: long-read sequencing. Nat. Methods 20, 6–11 (2023).
Nousias, O., Duffy, F. G., Duffy, I. J., Whilde, J. & Duffy, D. J. Long-read nanopore shotgun eDNA sequencing for river biodiversity, pollution and environmental health monitoring. Preprint at bioRxiv https://doi.org/10.1101/2024.11.01.621618 (2024).
Kalantar, K. L. et al. IDseq—an open source cloud-based pipeline and analysis service for metagenomic pathogen detection and monitoring. GigaScience 9, giaa111 (2020).
Simmonds, S. E. et al. CZ ID: a cloud-based, no-code platform enabling advanced long read metagenomic analysis. Preprint at bioRxiv https://doi.org/10.1101/2024.02.29.579666 (2024).
Stat, M. et al. Ecosystem biomonitoring with eDNA: metabarcoding across the tree of life in a tropical marine environment. Sci. Rep. 7, 12240 (2017).
Kolmogorov, M. et al. metaFlye: scalable long-read metagenome assembly using repeat graphs. Nat. Methods 17, 1103–1110 (2020).
UN Environment Programme and International Livestock Research Institute. Preventing the Next Pandemic—Zoonotic Diseases and How to Break the Chain of Transmission (United Nations, 2020).
Mora, C. et al. Over half of known human pathogenic diseases can be aggravated by climate change. Nat. Clim. Change 12, 869–875 (2022).
White, R. J. & Razgour, O. Emerging zoonotic diseases originating in mammals: a systematic review of effects of anthropogenic land-use change. Mammal. Rev. 50, 336–352 (2020).
Cavallero, S., Gabrielli, S., Gazzonis, A. L., Pombi, M. & Šnábel, V. Editorial: zoonotic parasitic diseases in a changing world. Front. Vet. Sci. 8, 715112 (2021).
Whilde, J., Martindale, M. Q. & Duffy, D. J. Precision wildlife medicine: applications of the human-centred precision medicine revolution to species conservation. Glob. Change Biol. 23, 1792–1805 (2017).
Métris, K. L. & Métris, J. Aircraft surveys for air eDNA: probing biodiversity in the sky. PeerJ 11, e15171 (2023).
O’Neill, J. Review on antimicrobial resistance: tackling drug-resistant infections globally: final report and recommendations. (Wellcome Trust, 2016).
Dugal, L. et al. Individual haplotyping of whale sharks from seawater environmental DNA. Mol. Ecol. Resour. 22, 56–65 (2022).
Doi, H. et al. On-site environmental DNA detection of species using ultrarapid mobile PCR. Mol. Ecol. Resour. 21, 2364–2368 (2021).
Chan, J., Raju, S. C. & Topol, E. Towards a tricorder for diagnosing paediatric conditions. Lancet 394, 907 (2019).
Siegel, E. Treknology: The Science of Star Trek from Tricorders to Warp Drive (Voyageur, 2017).
Estep, A. S. et al. Quantification of permethrin resistance and kdr alleles in Florida strains of Aedes aegypti (L.) and Aedes albopictus (Skuse). PLoS Negl. Trop. Dis. 12, e0006544 (2018).
Farrell, J. A. et al. Environmental DNA monitoring of oncogenic viral shedding and genomic profiling of sea turtle fibropapillomatosis reveals unusual viral dynamics. Commun. Biol. 4, 565 (2021).
Seymour, M. et al. Acidity promotes degradation of multi-species environmental DNA in lotic mesocosms. Commun. Biol. 1, 4 (2018).
Lynggaard, C., Frøslev, T. G., Johnson, M. S., Olsen, M. T. & Bohmann, K. Airborne environmental DNA captures terrestrial vertebrate diversity in nature. Mol. Ecol. Resour. 24, e13840 (2024).
Riaz, T. et al. ecoPrimers: inference of new DNA barcode markers from whole genome sequence analysis. Nucleic Acids Res. 39, e145 (2011).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Jalili, V. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2020 update. Nucleic Acids Res. 48, W395–W402 (2020).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
de Koning, W. et al. NanoGalaxy: nanopore long-read sequencing data analysis in Galaxy. GigaScience 9, giaa105 (2020).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Wick, R. Porechop. GitHub https://github.com/rrwick (2017).
Nurk, S. et al. The complete sequence of a human genome. Science 376, 44–53 (2022).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Schönherr, S., Weissensteiner, H., Kronenberg, F. & Forer, L. Haplogrep 3—an interactive haplogroup classification and analysis platform. Nucleic Acids Res. 51, W263–W268 (2023).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, E386–E394 (2009).
Team, R. RStudio: integrated development for R. RStudio http://www.rstudio.com/ (2020).
Wickham, H., Chang, W. & Wickham, M. H. Package ‘ggplot2’. Create elegant data visualisations using the grammar of graphics. Version 2, 1–189 (2016).
Hulsen, T., de Vlieg, J. & Alkema, W. BioVenn—a web application for the comparison and visualization of biological lists using area-proportional Venn diagrams. BMC Genomics 9, 488 (2008).
Garcia, M. et al. Sarek: a portable workflow for whole-genome sequencing analysis of germline and somatic variants. F1000Res. 9, 16665.2 (2020).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Porter, T. M. terrimporter/12SvertebrateClassifier: 12S Vertebrate North American Classifier v3.0.0-ref. Zenodo https://doi.org/10.5281/zenodo.7888133 (2024).
Benoit, M. & Drost, H.-G. A predictive approach to infer the activity and natural variation of retrotransposon families in plants. In Plant Transposable Elements: Methods and Protocols Vol. 2250 (ed. Cho, J.) 1–14 (Springer, 2021).
Duffy, D. J. et al. Identification of a MYCN and Wnt-related VANGL2-ITLN1 fusion gene in neuroblastoma. Gene Rep. 12, 187–200 (2018).
Spitzer, M., Wildenhain, J., Rappsilber, J. & Tyers, M. BoxPlotR: a web tool for generation of box plots. Nat. Methods 11, 121–122 (2014).
Ram, N. The ethics of human sequences in environmental samples. Nat. Ecol. Evol. 7, 796–797 (2023).
Doi, H. & Kelly, R. P. Ethical considerations for human sequences in environmental DNA. Nat. Ecol. Evol. 7, 1334–1335 (2023).
Brown, E. A. Your DNA can now be pulled from thin air. Privacy experts are worried. The New York Times (16 May 2023).
Kleinman, Z. & McMahon, L. UK and US refuse to sign international AI declaration. BBC News https://www.bbc.com/news/articles/c8edn0n58gwo (2025).
Dass, M. A. et al. Assessing eDNA capture method from aquatic environment to optimise recovery of human mt-eDNA. Forensic Sci. Int. 361, 112085 (2024).
Fantinato, C., Gill, P. & Fonneløp, A. E. Investigative use of human environmental DNA in forensic genetics. Forensic Sci. Int. Genet. 70, 103021 (2024).
Novroski, N. M. M. Environmental DNA: forensic friend or foe? Forensic Genomics 3, 35–37 (2023).
Fantinato, C., Gill, P. & Fonneløp, A. E. Detection of human DNA in the air. Forensic Sci. Int. Genet. Suppl. Ser. 8, 282–284 (2022).
Dass, M. A. et al. Assessing the use of environmental DNA (eDNA) as a tool in the detection of human DNA in water. J. Forensic Sci. 67, 2299–2307 (2022).
Trájer, A. J. Which mosquitoes (Diptera: Culicidae) are candidates for DNA extraction in forensic practice? J. Forensic Leg. Med. 58, 183–191 (2018).
Stammnitz, M. R., Hartman Scholz, A. & Duffy, D. J. Environmental DNA without borders. EMBO Rep. 25, 4095–4099 (2024).
Acknowledgements
This study was funded by D.J.D’s UF start-up funds. Maximilian Stammnitz was supported by an EMBO long-term fellowship (grant no. ALTF 544-2021 to M.R.S.). We thank M. Q. Martindale (UF), N. Condron (UF), C. Schnitzler (UF), A. Pacetti (UF), J. Hines (UF), S. Eastman (Florida DEP), P. Thompson (UF), D. Cummings (UF), C. Mavian (UF), R.-D. Xue (Anastasia Mosquito Control District), E. Higgs (Wildlife Rehabilitation Ireland) and L. Jones (Wildlife Rehabilitation Ireland). Sincerest thanks to P. Murphy and R. Rolfe for facilitating eDNA extraction of Irish samples in their lab in the Zoology Department, Trinity College Dublin, and to A. Krstic and W. Kolch for facilitating quality tests and qPCR of Irish samples at Systems Biology Ireland at University College Dublin. We also thank K. Foote, A. Rich, NPS Fort Matanzas National Monument, T. Wolfgang, A. Nelsen, T. Fenn, Jacksonville Zoo and Gardens, D. Feliciano, D. G. Hernández and the Chelonia Team Puerto Rico and C. Manes, R. Herren and T. Osborne, UF for valuable assistance with originally obtaining the sediment and water eDNA outgroup samples. Thanks are also due to the anonymous study participants who permitted us to collect sand from their footprints and room air for human eDNA analysis, with full ethical approval and informed consent. We also thank the staff of the UF ICBR core facility and BMKGENE for sequencing services. Some maps in this research study were created using ArcGIS software by Esri. ArcGIS and ArcMap are the intellectual property of Esri and are used herein under license. Copyright Esri. All rights reserved. For more information about Esri software, please visit www.esri.com. Any use of trade, firm or product name is for descriptive purposes only and does not imply endorsement by the US Government.
Author information
Authors and Affiliations
Contributions
D.J.D. designed and supervised the project. D.J.D., J.W., F.G.D. and I.J.D. conducted sampling. D.J.D., J.A.F., S.A.K., C.B.E. and V.S. generated the data. O.N., M.McC., M.R.S. and D.J.D. conducted bioinformatics analysis. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Zachary Gold, Lara Urban and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Airborne DNA for surveillance of pathogen and host species.
a) Gemycircularvirus genome coverage from shotgun long-read sequencing of an outdoor air sample (8-day Meath estuary July 2023). b) Animal-infecting DNA virus genus-level eDNA quantification in nucleotides per million bases sequenced (nt_bpm) of long-read shotgun sequenced sea water (Keaton Beach, Florida, June 2023). Host animal species name is provided for each bar. c) Coniferiporia (plant root rot fungal pathogen) genus-level eDNA quantification in nucleotides per million reads sequenced (nt_rpm) from short-read shotgun sequenced 1-week-long outdoor air samples from the Florida forest (FOR), and Whitney Lab (WL) sites, and their NFCs. d) Pine tree genus-level eDNA quantification in nucleotides per million reads sequenced (nt_rpm) from Illumina short-read sequenced 1-week-long outdoor air samples from the Florida forest (FOR), and Whitney Lab (WL) sites, and their NFCs. e) Oak tree genus-level eDNA quantification in nucleotides per million reads sequenced (nt_rpm) from short-read shotgun sequenced 1-week-long outdoor air samples from the Florida forest (FOR), and Whitney Lab (WL) sites, and their NFCs. c-e) all samples shown are Illumina NovaSeq Series X Plus shotgun sequenced. f) Changes in the nucleotides per million bases sequenced (nt_bpm) for Oak (Quercus genus), pathogenic fungus (Alternaria alternata) and vine (Vitis genus) identified by CZ ID analysis of shotgun long-read MinION sequencing.
Extended Data Fig. 2 Airborne DNA for surveillance of disease vectors & pests.
a) Mosquito genus-level detection from short-read shotgun sequenced 1-week-long outdoor air samples from the Florida forest (FOR) and Whitney Lab (WL) sites. X-axis is taxon nucleotides per million reads sequenced (nt_rpm). Left: Anopheles genus. Right: the other five mosquito genera detected in at least one air eDNA sample. b) Culicoides (biting midge) genus-level eDNA quantification (nt_rpm) from Illumina short-read sequenced 1-week-long outdoor air samples from the Florida forest (FOR) and Whitney Lab (WL) sites. c) Aligned read counts of short-read shotgun sequencing reads for human-associated mammals, including rats and mice, across the Dublin City July 2023, Meath estuary July 2023, Wicklow mountain July 2023, Wicklow Avoca River (water), and Florida forest April 2023 sampling locations. The y-axis lists both the common and Latin names for each species.
Extended Data Fig. 3 Additional human urban pest species and host-parasite analysis from air eDNA shotgun sequencing.
a) Counts of long-read shotgun sequencing (MinION) reads hitting species considered to be common Florida urban pests in the 1-week forest air April 2023 sample. This air eDNA sample was collected in a mixed hammock coastal forest and low density human residential area. The y-axis lists both the common and Latin names for each species. b) Counts of short-read shotgun sequencing (NovaSeq 6000) reads hitting species considered to be common Florida urban pests in the 1-week forest air April 2023 sample. The y-axis lists both the common and Latin names for each species. c) Urban pest species, linear regression, long-read versus short-read shotgun sequencing based detection from the same 1-week forest air April 2023 sample. The linear regression is for the same pest species displayed in a) and b). d) Nucleotides per million bases sequenced (nt_bpm) of long-read shotgun sequencing reads hitting bee genera and bee mite, across Florida forest and Irish locations.
Extended Data Fig. 4 Airborne DNA for broadscale phyla assessments.
a) Ratio of metazoan mapped reads to total mapped reads from air and water eDNA shotgun sequencing. b) Cumulative hits for arthropod species, for reads from short-read shotgun sequenced Irish 8-day air and the Florida 1-week forest air April 2023 samples. c) Cumulative hits for species in each metazoan phylum (excluding arthropods), for reads from the same short-read shotgun sequenced samples as in b), allowing for a direct comparison of sequencing output across the phyla. d) Percentage of hits for each metazoan phylum as a proportion of total metazoan hits for that sample, for reads from the same short-read shotgun sequenced samples as in b). b-d) Samples shown are: negative field control (NFC 8-day Dublin air capped July 2023), Florida forest air eDNA (1-week forest air April 2023), Ireland water eDNA (Avoca River RAMP23C July 2023), Ireland air eDNA (8-day Wicklow mountain air July 2023, 8-day Dublin city air July 2023 and 8-day Meath estuary air July 2023).
Extended Data Fig. 5 Comparison of air eDNA long-read versus short-read shotgun sequencing.
a) Cumulative hits for species in each metazoan phylum, for long- and short-reads generated from the same air eDNA sample (1-week forest air April 2023), allowing for a direct comparison of sequencing output across the phyla. The y-axis lists the phyla, and the x-axis shows the total matches per phylum. b) Proportion of long- and short-reads stemming from the same air eDNA filtrate which map to any of 45 Florida or Ireland diverse eukaryote reference genomes with high quality (MAPQ ≥ 30). Log-log scatter plot of the fraction of species-uniquely mapped short-reads (Illumina, X-axis) or long-reads (ONT, Y-axis). Lines represent log-log linear regression, gray shading 95% confidence interval. R2, coefficient of determination. Left: correlations for the 1-week Florida forest air April 2023 sample. Right: correlations for the 8-day Wicklow mountain air July 2023 sample. Left: relative species abundance of forest air eDNA, as measured by short-read (n = 44,319 out of 6 Million subsampled reads) and long-read (n = 73,772 out of 6,509,032 reads) sequencing. Right: relative species abundance of mountain air eDNA, as measured by short-read (n = 92,421 out of 6 Million subsampled reads) and long-read (n = 160,104 out of 1,424,249 reads) sequencing. Overlaid species silhouette images from https://www.phylopic.org. c) Genus call overlaps between dual long- and short-read sequencing of the same air eDNA samples: 8-day Dublin city air July 2023 (left) and 3-week Florida forest air November 2023 (right). Includes all domain-of-life genera, except deuterostomes. CZ ID cloud-based metagenomics analysis.
Extended Data Fig. 6 Shotgun sequencing versus vertebrate metabarcoding.
a) Comparison of 12S vertebrate metabarcoding with shotgun sequencing of the same samples (1-week Whitney Lab beach air July 2024, A and B) for detection of eighteen vertebrates. X-axis denotes the percentage of the total number of read counts (shotgun sequencing) or all ASV counts (metabarcoding) for the eighteen vertebrate animals, that relate to each individual species. Percentages enable the visualization of both metabarcoding and shotgun sequencing data at the same scales. Whitney Lab is abbreviated to WL in the sample names in figure panels a-c. b) American alligator (Alligator mississippiensis) detection across all air vertebrate 12 s air eDNA metabarcoding samples. c) Comparison of 12S vertebrate metabarcoding with shotgun sequencing of the same samples (1-week Whitney Lab beach air July 2024, A and B) for human (Homo sapiens) detection by either approach. X-axis denotes the total number of read counts (shotgun sequencing) or ASV counts (metabarcoding).
Extended Data Fig. 7 Replicability of co-sampled air eDNA.
a) Absolute quantity of 18 s rRNA pan-eukaryotic eDNA levels across 1-week air eDNA samples and NFCs, as detected by qPCR. Pumped air samples were taken in duplicate at the same time at the same location (Florida, Whitney Lab), using identical vacuum pumps (Supplementary Table 1); each duplicate is denoted by an A or B. Each qPCR reaction is a 10 μl reaction containing 1 μl of extracted eDNA template. Absolute quantity is per reaction, derived from a standard curve using sea turtle whole genome DNA. Each boxplot is a single sample, and each datapoint (3 technical qPCR well/reaction replicates per sample) is displayed by a grey dot. Tukey whiskers (extend to data points that are less than 1.5 × interquartile range (IQR) away from 1st/3rd quartile) were utilized for each boxplot. The median for each sample is shown as a horizontal line within each box, and box edges are the upper and lower quartiles. Sample name abbreviation: WL = Whitney Lab, Jul/Aug = sampling week ran from end of July into August, Aug = sampling week fell fully in August, Sept = sampling week fell fully in September. b) Area-proportional Venn diagrams of non-deuterostome eukaryotic genus hit overlaps between co-sampled 1 week Whitney Lab air eDNA; each duplicate is denoted by an A or B. Short-read shotgun sequencing, only genera with over 20 supporting reads are shown [NT r]. Top: overlap between two co-sampled samples (and an NFC). Bottom: overlap between two samples from the same location, sampled for the same duration, but collected 10 days apart (Supplementary Table 1) and an NFC. c) All domains of life (excl. deuterostomes) top genera heatmap of the two sets of co-sampled 1-week Whitney Lab air eDNA replicates and their NFCs. Short-read shotgun sequencing, heatmap displays the 50 highest read genera from each sample.
Extended Data Fig. 8 Spatial and temporal variation of shotgun sequenced air and swab eDNA.
a) Area-proportional Venn diagrams of non-deuterostome eukaryotic genus hit overlaps between co-sampled (same time at the same site, Supplementary Table 1) Florida forest swab January 2024 eDNA samples. Short-read shotgun sequencing, only genera with over 20 supporting reads are shown [NT r]. Left: wood plank, palm leaf and metal post swabs. Right: Windowpane, palm leaf and metal post swabs. b) Temporal comparison, area-proportional Venn diagram of non-deuterostome eukaryotic genus hit overlaps between air samples collected at the same site, at different times, or different durations, Florida forest air eDNA, i) 1 week Nov. 2023, ii) 3 weeks Nov. 2023 or iii) 1 week April 2024 (Supplementary Table 1). Sample name abbreviation: FOR = forest, 1wk = 1 week, 3wk = 3 weeks. Short-read shotgun sequencing, only genera with over 20 supporting reads are shown [NT r]. c) All domains of life (excl. deuterostomes) top genera heatmap of the same three air eDNA samples shown in b) above. Short-read shotgun sequencing, heatmap displays the 30 highest read genera from each sample. d) Spatial comparison, area-proportional Venn diagram of non-deuterostome eukaryotic genus hit overlaps between 1 week air samples taken for the same time and duration, two sites 3.2 km apart (Florida forest [FOR] & Whitney Lab [WL], Supplementary Table 1).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–7 and discussion.
Supplementary Table 1 (download XLSX )
Supplementary Tables 1–3.
Supplementary Video 1 (download MP4 )
Supplementary Video 1. Bobcat walking through the coastal hammock forest study site, March 2022. Credit to J.W.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nousias, O., McCauley, M., Stammnitz, M.R. et al. Shotgun sequencing of airborne eDNA achieves rapid assessment of whole biomes, population genetics and genomic variation. Nat Ecol Evol 9, 1043–1060 (2025). https://doi.org/10.1038/s41559-025-02711-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41559-025-02711-w