Abstract
Research on the biology and evolution of sex chromosomes has primarily focused on diploid XX/XY and ZW/ZZ systems. In contrast, the rise, evolution and demise of U/V systems has remained an enigma. Here we analyse genomes of nine brown algal species with different sexual systems to determine the history of their sex determination. U/V sex chromosomes emerged between 450 and 224 million years ago, when a region containing the pivotal male-determinant MIN ceased recombining. Seven ancestral genes within the sex-determining region show remarkable conservation over this vast evolutionary time, although nested inversions caused expansions of the sex locus, independently in each lineage. We evaluate whether these expansions are associated with increased morphological complexity and sexual differentiation, and show that taxonomically restricted genes evolve unexpectedly often in U and V chromosomes. We also investigate two situations in which U/V-linked regions have changed. First, we demonstrate that convergent evolution of two monoicous species occurred by ancestral males acquiring U-specific genes. Second, the Fucus dioecious system involves new sex-determining gene(s), acting upstream of formerly V-specific genes during development. Both situations have led to the demise of U and V chromosomes and erosion of their specific genomic characteristics.
Similar content being viewed by others
Main
The mechanisms controlling the development of male or female identities, or co-sexuality, when individuals express both sex functions, vary widely across different organisms1,2. In species with separate sexes, sex chromosomes may be present, carrying a sex-determining region (SDR)3 that encodes factors directing sex identity and which often does not undergo recombination in the heterogametic sex (XY or ZW)4 of diploid species (dioecious), or in the diploid stage of haploid-dominant (dioicous) species. Sex chromosomes have independently evolved from autosomes multiple times and may be subject to specific evolutionary forces, including differential selection between sexes, asymmetrical expression of deleterious mutations and hemizygosity, meiotic silencing and dosage compensation3.
Research on the biology and evolution of sex chromosomes has primarily focused on diploid XX/XY and ZW/ZZ systems in mammals, birds, fish, Drosophila and diploid plants4,5. U/V haploid sex-determination systems, such as those of bryophytes and algae6,7, have been less explored. In U/V systems, sex is not determined at fertilization but during meiosis, when haploid spores inherit either a U chromosome, and will develop into a female gametophyte, or a V chromosome, controlling male gametophyte formation8. These fundamental inheritance differences between U/V and XX/XY or ZW/ZZ systems have broad evolutionary and genomic implications9,10. However, so far, only the U/V systems of the brown alga Ectocarpus and the U/Vs of four distantly related bryophyte taxa11,12,13,14 have been fully sequenced and assembled into chromosomes. While these studies helped understand the genomic structure of bryophyte U- and V-linked regions, the species involved diverged ~500 Ma (million years ago) and do not share homologous U/V chromosomes15. As a result, we still lack a broad comparative view across multiple homologous U/V systems that would inform a reconstruction of their evolutionary history. Brown algae represent exceptional models for studying sex chromosome evolution because they display diverse reproductive systems, life cycles and sex chromosome systems in a single lineage16. Their ancestral state probably involved separate sexes16, suggesting that their sex chromosomes could share a common origin. Here we study the origin, evolution and demise of U/V sex chromosomes in the brown algae.
Results
The origin of brown algal sex chromosomes
We focused on species covering the phylogenetic, morphological and reproductive diversity of the brown algal clade17 and their closest extant outgroup, Schizocladia ischiensis17,18. We substantially improved the brown algal genome datasets available18,19,20 to reach chromosome or near-chromosome-level genome assemblies (Extended Data Table 1 and Supplementary Table 1). This revealed that brown algae have 27–33 chromosomes and largely conserved macrosynteny (Fig. 1a).
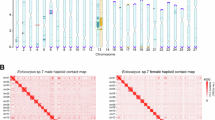
a, Macrosynteny plot comparing genomes of six dioicous (green), two monoicous (red), one dioecious (blue) and one outgroup species (yellow). The chromosomes were originally numbered by their physical size in the Ectocarpus v2 genome141. Note that the dioecious species F. serratus has a fully diploid life cycle (without gametophytes7). Syntenic blocks of the V sex chromosome are highlighted in red, with the emergence of U/V chromosomes shown in the phylogeny. Genome sizes are indicated in brackets. b, Microsynteny plot of V chromosomes in five dioicous species, highlighting the male sex-determining regions (blue) and the PARs (green). The PAR genes whose orthologues are found within the SDR of other species are highlighted in purple. c, Microsynteny plot of U chromosomes in two dioicous species, highlighting the female sex-determining regions (peach) and the PARs (green). The PAR genes whose orthologues are found within the SDR of other species are highlighted in purple. Note that the genome assemblies for C. linearis and S. ischiensis are not chromosome level, leading to a high number of contigs.
We identified the female (U) and male (V) sex-determining regions (SDRs) in the dioicous species using a combination of bioinformatic and experimental approaches (see ‘Discovery of the U/V sex determination regions’ in Methods; Supplementary Figs. 1–5). All U/V species share the same, albeit highly rearranged, ancestral sex chromosome, showing remarkable stability despite the large evolutionary time (Fig. 1b,c and Extended Data Table 1). The recombination suppression event leading to the birth of U/V sex chromosomes occurred after the split of S. ischiensis and Dictyota dichotoma, ~450–224 Ma21 (Fig. 1a). The male-determining gene MIN22 is the only V-specific gene consistently present in all V-SDRs of the dioicous species. We note that one dioecious (Fucus serratus) and two monoicous (haploid, co-sexual Chordaria linearis and Desmarestia dudresnayi) species lack U/V sex chromosomes but still retain MIN on a chromosome homologous to the ancestral U/V (‘U/V-homologue’ hereafter). The outgroup S. ischiensis has low synteny with the brown algae and exhibits putative fusion-with-mixing events23 (Fig. 1a and Extended Data Fig. 1).
We next examined the U/V-SDRs by comparing male and female genome assemblies (Methods). The SDRs contain a small number of genes overall (between 18 and 52), and compared with the pseudoautosomal regions (PARs) (between 229 and 904), with considerable variation in gene content and size across species, the smallest being found in the Ectocarpales (Ectocarpus sp. 7, Ectocarpus crouaniorum, Scytosiphon promiscuus; Fig. 1b,c, Extended Data Fig. 2a and Extended Data Table 1). SDR size differences across species are strongly correlated with the number of genes (R2 = 0.97; Extended Data Fig. 2b) and the repeat content (R2 = 0.99; Extended Data Fig. 2c) inside these regions. Many genes located in the PARs of the Ectocarpales are within the V-SDRs of Undaria pinnatifida, Desmarestia herbacea and D. dichotoma, indicating that the SDR boundaries have changed across species. The boundary differences coincide with extensive structural rearrangements, particularly inversions, even among closely related taxa (Fig. 1b,c and Extended Data Fig. 3). Note that the centromere in the V chromosome of Ectocarpus is found within the SDR19, so we cannot exclude that a centromere-related suppression of recombination may have preceded the inversion events found on the SDR24.
Together, our results indicate that the brown algal U/V sex chromosomes evolved between 450–224 Ma, via suppressed recombination in a genomic region that contained MIN (henceforth male-determining locus). The presence of MIN in distantly related lineages could push the age of the U/V chromosomes further back in time, but more evidence would be required to establish that dioicy existed in these organisms.
The evolution of the SDRs involved boundary expansions and gene gains
The brown algal U- and V-SDRs carry homologous genes (gametologue pairs), indicating descent from a common ancestral region (Supplementary Table 2). Ectocarpus sp. 7 and D. herbacea show similar ratios of gametologues and U- or V-specific genes (16/14 and 11/7 gametologue/sex-specific genes in Ectocarpus and D. herbacea, respectively; Supplementary Table 2). Only ten genes share SDR orthologues between both species, while the rest were mostly acquired independently in the SDR of each species, with one gene that was retained as a gametologue pair in Ectocarpus sp. 7 but lost both copies in D. herbacea (Supplementary Table 2). Five gametologue pairs conserved both copies in the two species, while another three gametologue pairs lost either the male or the female copy in D. herbacea (Supplementary Table 2). In addition, MIN and a U-specific gene are also conserved between species (Supplementary Table 2). Although the total number of U/V-SDR genes differs between Ectocarpus sp. 7 (18 genes) and D. herbacea (30 genes), each species shows an equal number of gametologues and sex-specific genes in its U- and V-SDRs (Supplementary Table 2). This intraspecies symmetry supports the idea that the U and V chromosomes may have undergone parallel evolutionary changes within each lineage10,25,26. The V-SDR of D. herbacea contains 20 additional genes that belong to endogenous viral elements, which are common across brown algal genomes18.
Diploid sex chromosome in animals and plants exhibit evolutionary strata representing different recombination suppression events over time. Strata are identified by analysing synonymous substitutions (Ks) between male/female gametologue pairs27 whose locations in fully X or Z-linked regions are known. However, detecting evolutionary strata in U/V systems is difficult because neither of these fully sex-linked regions recombines and gene movements and chromosome rearrangements disrupt collinearity of both chromosomes between species25,28,29. Moreover, in the absence of a recombining outgroup (which does not exist in brown algae), the ancestral gene order cannot be reliably inferred. In both Ectocarpus sp. 7 and D. herbacea, the V- and U-SDR rearrangements differ by inversions (Fig. 2a,b and Extended Data Fig. 4), consistent with the idea that inversions may lead to suppressed recombination between sex chromosomes. An analysis of gametologue pair divergence revealed saturated levels of Ks values (Fig. 2b,c and Supplementary Table 3), further limiting the inference of evolutionary strata across brown algal SDRs. Nonetheless, the gametologue Ks values are broadly consistent between orthologues in Ectocarpus sp. 7 and D. herbacea, where shared SDR gametologues between species have higher Ks values and probably spent more evolutionary time diverging than the gametologues that are not shared between species (Supplementary Table 3). Furthermore, the location of gametologues with the lowest Ks values in the U-SDR of D. herbacea, relative to the PAR genes in Ectocarpus sp. 7, suggests that inversions involving the entire U-SDR and adjacent PAR segments probably contributed to the expansion of the U/V-SDR boundaries in D. herbacea, in a process we term ‘engulfment’ (Extended Data Fig. 4). The expansion of the SDR boundaries in D. herbacea led to the engulfment of a region containing four genes in the PAR1 of Ectocarpus sp. 7, and a second region with 13 genes located on the PAR2 (Fig. 2d, Supplementary Table 4 and Extended Data Fig. 4). Twelve of these engulfed genes into the SDR of D. herbacea were retained as gametologues. These observations support a scenario where expansions in the SDR boundaries of brown algae occur through nested inversions. Two chromatin-related transcription factors in the Ectocarpus PARs were independently incorporated into the SDRs of four other dioicous species (Supplementary Tables 4 and 5).
a, Microsynteny plot between the U and V chromosomes of Ectocarpus sp. 7 and D. herbacea. b, Synteny between the U and V gametologues within the V-SDRs of both species, coloured by synonymous substitutions per site (Ks). c, Identification of ancestral SDR gametologues (red squares) and independently acquired gametologues (black dots) with respect to their gametologue Ks values and their position in the V-SDR. d, Circos plot linking gametologue pairs in each species with expression levels of all SDR genes (log2(TPM + 1)) across different life stages in Ectocarpus sp.7 and mature gametophytes (matGA) in D. herbacea. Gametologues are highlighted in dark colours, sex-specific genes are highlighted in light colours, viral insertions are marked in grey and asterisks denote conserved SDR genes (also in f). e, The ancestral state reconstruction of V-SDR gene content across brown algae, showing the expected number of genes in the SDR (white circles), gene retention (blue numbers), gene gain through expansion of the SDR boundaries (orange), gene gain through autosomal translocation (purple), gene birth event inside the SDR (yellow) and gene loss (red) along with changes in gamete dimorphism16. f, Schematic of the seven ancestral V-SDR gene orthogroups (OGs), with genomic locations marked: retained in the V-SDR (blue), found in the U/V-homologue of non-dioicous species (green), translocated from the V-SDR to an autosome (yellow), present in a non-scaffolded contig (red), and lost (grey). Bold: MIN. See Supplementary Tables 5 and 6.
The observation of greater V-SDR gene content in early diverging lineages (such as D. dichotoma) than in the later-diverging Ectocarpales (Extended Data Table 1) could reflect either gene loss in the V-SDRs of Ectocarpales or independent gene gains in the V-SDRs of each lineage (as predicted in ref. 10), from an ancestral state with low V-SDR gene content that is retained in Ectocarpales. To distinguish between these possibilities, we reconstructed the ancestral SDR gene content (Supplementary Table 5), focusing on the V chromosome, as the genomic data are of better quality (Supplementary Table 1), and assuming parallel U/V-SDR evolution10,25,26 as seen in Ectocarpus sp. 7 and D. herbacea (Fig. 2d and Supplementary Table 2). This analysis revealed that brown algal V-SDR evolution occurred via lineage-specific gene gains rather than gene loss in the Ectocarpales (Fig. 2e,f). Gene gains were caused by a combination of three processes: expansions of the SDR boundaries into the PARs, translocation of autosomal genes into the SDR and lineage-specific gene birth events within the SDR (Fig. 2e and Supplementary Table 5). Consistently, ancestral V-SDR genes were associated with higher gametologue Ks values, while independently acquired gametologues in D. herbacea had lower Ks values (Supplementary Table 2 and Fig. 2c).
The seven genes in the ancestral V-SDR (Fig. 2f and Extended Data Fig. 2d) include the male-determinant MIN22 and six V gametologues of genes that are also carried on the U chromosome (Fig. 2f and Supplementary Table 6). As predicted by early models of U/V-SDR evolution10, all seven genes are probably related to sex determination processes. Gametologue pairs include putative transmembrane proteins that may play a role in gamete recognition30, STE20 serine/threonine kinase gametologues probably involved in pheromone pathways31, and a casein kinase, a MEMO-like domain protein and a GTPase-activating protein which may act in signal transduction (Supplementary Table 6). All of these genes are gametologue pairs in Ectocarpus sp. 7, but in D. herbacea the casein kinase was lost from the U-SDR and the putative transmembrane receptor was lost in both sexes. We noticed that these ancestral V-SDR genes remain in the U/V-homologue of the species that have lost their U/V system (Fig. 2f and Supplementary Table 6), emphasizing their importance for pathways in sex organ development even in the absence of sex chromosomes.
The V-SDR size appears to be associated with the level of sexual dimorphism (Fig. 2e and Extended Data Fig. 2), but the small sample size is insufficient for formal statistical analysis. Species with low sexual dimorphism (anisogamous) retained the ancestral V-SDR genes with very few gene gains, further suggesting that they may represent the V-SDR ancestral state. Although the number of SDR changes is small, oogamous species each independently gained diverse V-SDR genes, and one gene (ATP-dependent RNA helicase) was convergently acquired in all (OG0003211 in Extended Data Fig. 2d). All the detected autosomal translocations into the V-SDRs of Ectocarpus sp. 7 and D. herbacea (Fig. 2e) also involve sex-specific genes (Supplementary Tables 2 and 5), consistent with a model where sexual antagonism in autosomal loci may be solved by gaining sex linkage32. In contrast, we found no correlation between autosomal sex-biased gene (SBG) expression and sexual dimorphism level (false discovery rate (FDR)-corrected P > 0.01; Supplementary Table 7), supporting previous studies33 (Extended Data Fig. 5a). However, we observed an enrichment of male-biased genes on the PARs in all species (chi-square test P < 0.01) except D. dichotoma (Extended Data Fig. 5b).
Most U/V-SDR genes were prominently expressed in fertile haploid gametophytes, consistent with gene preservation via haploid purifying selection (Supplementary Table 8). Gametologues had typically higher expression levels than sex-specific genes (present in only one of the SDRs) (Wilcoxon test, P = 0.00075 in D. herbacea; P = 0.08843 in Ectocarpus sp. 7) (Fig. 2d). A comparative analysis in fertile gametophytes between SDR genes and their autosomal counterparts in other species showed that newly acquired genes on the SDR had similar expression levels to their autosomal counterparts (Extended Data Fig. 6), suggesting either a co-option of autosomal biological activity into male-specific functions in the V-SDR or the general importance of these genes for gametophyte development. Examining expression levels across multiple tissues in Ectocarpus sp. 7 revealed that activity of U/V-SDR genes is not confined to fertile gametophytes (Fig. 2d). Therefore, the SDRs contain not only genes involved in sex determination and gametophyte fertility but also genes playing a broader role in development.
Altogether, our analyses illustrate how brown algal U/V-SDRs undergo structural changes, evolving mainly by lineage-specific gene gains associated with increasing levels of sexual dimorphism. We identified a set of conservatively sex-linked genes in dioicous brown algae, suggesting their role in sex determination and/or differentiation, along with genes potentially involved in other developmental pathways.
Structural features and evolutionary dynamics of brown algal U/V sex chromosomes
We next examined the structural features that differentiate the entire U/V sex chromosomes (V-SDR and PARs) from the rest of the genome (Fig. 3a and Supplementary Fig. 12a). As expected for non-recombining regions25,34, all V sex chromosomes are repeat rich and gene poor (Wilcoxon rank-sum test, FDR-corrected P < 0.01; Fig. 3a, Extended Data Fig. 7 and Supplementary Tables 9–11). V-SDRs have significantly higher repeat density than the PARs or the autosomes (permutation test, FDR-corrected P < 0.001; Extended Data Fig. 8). This low gene density is not influenced by the presence of centromeres within the SDRs, as the coding density in the Ectocarpus sp. 7 V centromere (3.51%) is slightly higher than in the rest of the V-SDR (2.85%), presumably due to the small size of the centromere (153 kbp)19. The PARs were also significantly enriched in repeats when compared with the autosomes, although less so than the V-SDRs (permutation test, FDR-corrected P < 0.001; Extended Data Fig. 8). Among repetitive elements, ‘unclassified’ transposable elements (TEs) were enriched in the PARs and SDRs of the Ectocarpales (permutation test, FDR-corrected P < 0.01), while the V-SDRs of species that underwent genome expansion (for example, U. pinnatifida, D. herbacea, D. dichotoma) predominantly accumulated long terminal repeat (LTR) elements (Supplementary Fig. 6 and Supplementary Table 9).
a, Karyoplots for five dioicous species showing the following features from bottom to top: chromosome compartments (autosomes, PARs and SDR), relative gene ages, interspecies Ks values, and proportion of coding (CDS, red) and repeat (TEs, blue) density. Statistically significant differences for each feature between each autosome and the V chromosome are depicted on top of the track for that autosome (FDR-corrected two-sided Wilcoxon rank-sum test; values indicated with solid colours when P < 0.01 for the tested hypothesis). The precise range of gene age categories and interspecies Ks values for each species can be found in Supplementary Figs. 7–11. b, Violin plots for five dioicous species showing the relative gene age ranks (higher ranks equate to younger ages) of the TRGs across chromosome compartments (autosomes, PARs and SDR). Statistically significant differences in mean values of gene ages (centre line) were assessed using FDR-corrected two-sided permutation tests.
Moreover, sex chromosomes had fewer orthologues conserved between species compared with the autosomes (chi-square test, P < 10−4, Supplementary Table 12), possibly reflecting increased numbers of taxonomically restricted genes (TRGs; that is, genes that are not detectable outside of a defined taxonomic group). Phylostratigraphy analyses35,36 confirmed an enrichment of TRGs in the sex chromosomes of all dioicous species (Wilcoxon rank-sum test, FDR-corrected P < 0.01; Fig. 3b and Supplementary Tables 13 and 14). TRG enrichment was localized in the PARs of the Ectocarpales, but this pattern extended to the entire sex chromosome, including the SDRs, in species with larger V-SDRs (permutation test, FDR-corrected P < 0.001; Fig. 3b). Importantly, sex chromosomes have statistically younger TRGs than the last common ancestor of the five dioicous species (same Order or broader taxonomic groups), indicating that TRG enrichment arose independently in each species (Pearson standardized residuals >2.4; Supplementary Figs. 7–11).
We previously proposed a theoretical model where generation-antagonistic selection may favour the retention of young sporophyte-beneficial loci in the PARs of Ectocarpus sp. 7 U/Vs37. Consistent with this model, sporophyte-biased genes are indeed enriched in the sex chromosomes of Ectocarpus sp. 7 and U. pinnatifida (fold change >2, adjusted P < 0.05), but less so in D. dichotoma and S. promiscuus (Supplementary Table 15). Moreover, we explored additional mechanisms underlying TRG emergence by estimating interspecies Ks values between orthologues in closely related species (Supplementary Table 16) and comparing these values between chromosomes and genomic compartments (V-SDR, PAR, autosomes). If synonymous mutations behave neutrally38,39, then interspecies Ks can be used as a proxy for mutation rates40,41. Consistently, we found higher interspecies Ks values in the V sex chromosomes compared with autosomes across all dioicous species (Wilcoxon rank-sum test, FDR-corrected P < 0.01; Fig. 3a and Supplementary Tables 16 and 17), suggesting higher mutation rates relative to autosomes. Higher interspecies Ks values are also localized in the PARs, mirroring the pattern observed with the TRGs (Supplementary Figs. 7–10). Therefore, the enrichment of TRGs in the U and V is associated with both enrichment of sporophyte-biased genes and higher synonymous substitution rates.
To test the generality of the pattern of TRG enrichment on U and V chromosomes, we applied the same approach in other organisms with haploid sex determination, the plants Ceratodon purpureum, Sphagnum angustifolium, Marchantia polymorpha11,12,42 and the fungus Cryptococcus neoformans43. We observed a clear enrichment of TRGs in the V chromosomes of C. purpureum and S. angustifolium (Wilcoxon rank-sum test, FDR-corrected P < 0.01; Supplementary Figs. 12 and 13 and Supplementary Tables 13 and 14), but not in the U/V chromosomes of M. polymorpha or the mating-type chromosome of C. neoformans (Supplementary Figs. 14 and 15 and Supplementary Tables 13 and 14).
Fate of U/V sex chromosomes following loss of dioicy
We studied the evolutionary trajectory of brown algal genomes after the loss of the U/V system, by exploring two independent transitions to monoicy in C. linearis and D. dudresnayi that undergo sexual reproduction and develop male and female gametangia33. Most genes in the ‘ex’-sex chromosomes (U/V-homologues) of both monoicous species are male derived, indicating that monoicy emerged from a male background (Fig. 4a,b). The U/V-homologue of C. linearis contains several rearrangements spanning the regions that are homologous to the PAR and SDR (SDR-homologue), with 11 V-SDR-derived and 2 U-SDR-derived orthologues located within the SDR-homologue, with an additional V-SDR-derived gene that was translocated elsewhere in the U/V-homologue (Fig. 4a and Supplementary Table 18). Likewise, D. dudresnayi underwent at least two inversion events within the SDR-homologue after splitting from D. herbacea (Fig. 4b), containing 20 V-SDR-derived genes and 4 U-SDR-derived genes (Supplementary Table 19).
a, Comparison of the U/V-homologue in C. linearis against the U and V chromosomes of Ectocarpus sp. 7. b, Comparison of the U/V-homologue in D. dudresnayi against the U and V chromosomes of D. herbacea. The colour code represents the identity of the genes alongside the chromosomes, while the shapes represent the evolutionary fate of each SDR gene in the monoicous genome. The matching shades between the SDRs and the U/V-homologue are either colour coded by their ancestral background or they appear as transparent dotted shades if the gametologue of the other sex was retained. c, Karyoplot of D. dudresnayi showing the following features from bottom to top: chromosome compartment (autosomes and U/V-homologue), relative gene ages, interspecies Ks values, proportion of coding (CDS, red) and repeat (TEs, blue) density. Statistically significant differences for each feature between each autosome and the U/V-homologue are depicted on top of the track for that autosome (FDR-corrected two-sided Wilcoxon rank-sum test; values indicated with solid colours when P < 0.01 for the tested hypothesis). d, Violin plot showing the relative gene age ranks (higher ranks equate to younger ages) of the TRGs between the autosomes and the U/V-homologue of D. dudresnayi. Statistically significant difference in mean values of gene ages (centre line) was assessed using an FDR-corrected two-sided permutation test.
Both monoicous species retained mostly male and a few female copies for most of the U/V-SDR-derived gametologues (91% in C. linearis and 100% in D. dudresnayi), whereas several U- and V-specific orthologues were lost in these species (10 and 17 sex-specific genes in C. linearis and D. dudresnayi, respectively). Of these lost orthologues, 60% and 43% present closely related autosomal paralogues in C. linearis and D. dudresnayi, respectively (Supplementary Tables 18 and 19), although it is unclear whether the expression of these autosomal paralogues is compensating the activity of the lost genes. The only three U-SDR-derived orthologues in C. linearis are flanked by PAR orthologues translocated at the end of the V-SDR-derived region, suggesting that the U/V-homologue (contig 12) of C. linearis acquired its U-SDR-derived genes through two translocations (Extended Data Fig. 9). Three U-SDR-derived genes in D. dudresnayi are dispersed across the V-SDR-derived region of the U/V-homologue, suggesting independent U-SDR translocations into the V-SDR, while the fourth U-SDR-derived gene was translocated to an autosome (Extended Data Fig. 9).
The seven ancestral V-SDR genes are transcriptionally active (log2(TPM + 1) > 2) during reproductive stages of both monoicous species (Supplementary Table 20), emphasizing their role in reproduction despite the absence of a U/V system, particularly MIN22 which is retained in both species. While most U-SDR-derived genes are absent in monoicous species, a single intracellular cholesterol transporter gene was convergently preserved in both monoicous genomes (Supplementary Table 20) and actively expressed during fertility in both Ectocarpus sp. 7 and D. herbacea (Supplementary Table 8).
The U/V-homologue of D. dudresnayi retains some vestiges of its past as a U/V chromosome, such as low coding density, high repeat density and enrichment of TRGs (Wilcoxon rank-sum test, FDR-corrected P < 0.01), although we found non-significant differences in interspecies Ks values across the genome (Fig. 4c, Extended Data Fig. 7, Supplementary Fig. 16 and Supplementary Tables 9–14, 16 and 17).
Finally, we examined the transition from haploid to diploid sex determination, which has remained unstudied in eukaryotes. Although the ancestral state for the brown algae is a U/V sexual system, the Fucales recently transitioned to a diploid life cycle44, with many species, such as F. serratus45, exhibiting diploid separate sexes (dioecy)46. Dioecy probably evolved from monoecy (both sexes in the same diploid individual) in the last common ancestor of the Fucus genus47 (25–5 Ma18), consistent with a young sex chromosome in F. serratus. Our extensive bioinformatic analysis and PCR sex-linkage testing for candidate genes such as MIN (Methods) failed to identify sex-linked sequences in F. serratus (Supplementary Fig. 17), suggesting that the SDR is small and undifferentiated. However, male F. serratus conserves the MIN gene and all the ancestral V-SDR genes in its U/V-homologue (Figs. 2f and 5a). Importantly, although none of the U/V-SDR-derived genes are sex-linked in F. serratus, MIN and four other ancestral V-SDR genes are exclusively expressed in males (fully silenced in females) (fold change (FC) > 2, padj < 0.05; Fig. 5a and Supplementary Table 21). This pattern is consistent in three other Fucales species (Supplementary Table 22). Therefore, the ancestral V-SDR genes probably still play roles in male sex determination or differentiation pathways.
a, Expression of ancestral U/V-SDR genes in the diplontic species F. serratus. Gene expression of mature algae (using 3 males and 3 females, see Methods) is given as log2(TPM + 1) and bars represent standard deviation of the mean. P values shown on the plot are derived from differential expression analysis performed using DESeq2, which applies a two-sided Wald test with Benjamini–Hochberg correction for multiple testing. Bold text represents whether the gene in F. serratus corresponds to an ancestral male or the female gametologue. b, Karyoplot of F. serratus showing the following features from bottom to top: chromosome compartment (autosomes and U/V-homologue), relative gene ages, interspecies Ks values (between 0.00079 and 6.838, with an average value of 0.148), proportion of coding (CDS, red) and repeat (TEs, blue) density. Statistically significant differences for each feature between each autosome and the U/V-homologue are depicted on top of the track for that autosome (FDR-corrected two-sided Wilcoxon rank-sum test; values indicated with solid colours when P < 0.01 for the tested hypothesis). c, Violin plot showing the relative gene age ranks (higher ranks equate to younger ages) of the TRGs between the autosomes and the U/V-homologue of F. serratus. The mean values of gene ages (centre line) are not significantly different (FDR-corrected two-sided permutation test).
Contrary to the observations in D. dudresnayi, the U/V-homologue of F. serratus lacks the TRG enrichment pattern and all the other distinctive features of the U/V chromosomes (Wilcoxon rank-sum test, FDR-corrected P > 0.01). Thus, this ‘ex’-sex chromosome has lost all the evolutionary vestiges of its past as a U/V chromosome (Fig. 5b, Extended Data Fig. 7, Supplementary Fig. 18 and Supplementary Tables 9–14, 16 and 17).
Discussion
Here we characterized the evolutionary trajectory of brown algal sex chromosomes (Fig. 6). Brown algal sex chromosomes date back 450–244 Ma21, at the origin of brown algae. We propose that the male-determining gene MIN underlies the birth of this U/V system22. The ancestral cassette with seven V-SDR genes suggests a very early evolution of these genes into a non-recombining locus during the evolution of the U/V-SDRs. The ancestral V-SDR genes probably contribute to reproduction, but may also be involved in broader developmental functions. Despite their old age, brown algal U/V chromosomes retain large PARs bordering the SDR, unlike haploid systems in non-vascular plants that mostly lack detectable PARs11,12,13,14.
U/V sex chromosomes arose from an ancestral autosome, via suppression of recombination that probably occurred via an inversion. The SDR boundaries expanded into neighbouring PAR via inversions, but also by recruitment of genes from autosomes; expansion occurred in a lineage-specific fashion, concomitant with increased sexual dimorphism of the different species. SDR genes are maintained within the SDR if they have roles in sex, whereas genes with no role in sex are lost. Faster substitution rates, probably a consequence of the heterochromatic context of the sex chromosome, may promote the rise of taxonomically restricted genes, which are selectively maintained on the sex chromosome if they have advantages to the sporophyte generation. In species that switch to a diploid life cycle, the U/V system disappears, but the genes that are in the V-specific region retain roles in sex, although they are no longer masters. Transition from U/V separate sexes to co-sexuality (monoicy) occurred when a male haploid individual acquired female-specific genes via translocations. During the demise of the U/V sex chromosomes, their structural and evolutionary footprints slowly disappear.
Brown algal genomes have a high degree of synteny conservation. The U/V-SDR is however prone to accumulate structural rearrangements, including inversions that may have caused the initial recombination suppression event in proto-sex chromosomes and the later expansion of the U/V-SDRs into the PARs. Similar to other haploid systems48,49, TEs conspicuously accumulated in the SDRs following recombination suppression50, possibly causing further rearrangements through TE-mediated inversions51.
Models of XX/XY and ZW/ZZ sex chromosome evolution suggest that sexually antagonistic selection may lead to SDR expansions, making former PAR genes fully sex linked and gaining new genes52,53. We show that the three brown algal species with greater sexual dimorphism in gametes (oogamy) indeed have enlarged U/V-SDR gene contents, although statistical testing is not possible given the small sample size. Oogamy is proposed to be ancestral in the brown algae16,21, but this trait seems to be highly labile and our analyses suggest multiple independent transitions to oogamy from a less dimorphic ancestor with a small ancestral SDR, accompanied by SDR gene gain. Anisogamous species only experienced few gene gains in their SDRs, as predicted in ref. 10. Similar to M. polymorpha, the relatively simple gene content of the U/V-SDRs in the brown algae could regulate an autosomal effector gene network controlling sexual dimorphism, including differences in somatic development between male and female gametophytes54. Accordingly, we observed substantial sex-biased expression of autosomal genes between mature male and female gametophytes carrying sex organs. This contrasts with C. purpureus, where sex chromosomes carry thousands of genes and few sex-biased autosomal genes11. Consistent with other studies33,55, we found no correlation between levels of sex-biased expression and sexual dimorphism in brown algae. Sexual dimorphism in this lineage may be controlled by a relatively small subset of genes, and most sex-biased genes may influence gametophyte physiology or vegetative development.
In diploid sexual systems, recombination suppression between sex chromosomes explains Y and W degeneration56 (but see refs. 57,58,59). Differences in the evolution of U/V systems are predicted, since sex determination occurs in the haploid stage, in which deleterious mutations are more efficiently removed by selection than in diploid systems29,60 and some of the predicted differences have been documented in bryophytes29. In brown algae, as in diploid systems, the SDRs lost recombination, leading to TE accumulation and consequent reduction in gene density. However, unlike the degeneration documented in some XX/XY and ZW/ZZ systems, our ancestral state reconstruction revealed more gene gains than losses or gene movements out of the V-SDR during evolution of brown algal lineages. Unclassified repeats accumulate in the Ectocarpales sex chromosomes, including the PARs, but not in species with larger genomes, where LTR retroelements become dominant. DNA transposons are overrepresented among unclassified repeats61, and they often insert near the progenitor locus in a process called local hopping62. We propose that the U/V-SDR may thus act as a source of DNA transposons that hop to the PARs, thus increasing their repeat density, whereas increased colonization of LTR elements obscures this pattern in larger genomes.
Brown algal U/V chromosomes display an excess of TRGs. What mechanisms underlie this pattern? The SDR and the PARs of U/V chromosomes are enriched in heterochromatin63, involved in repressing TEs64, and heterochromatic regions tend to have higher mutation rates due to reduced access of the DNA repair machinery during replication65. Accordingly, we consistently observe higher interspecies Ks values in U/Vs, particularly in the PARs, since these regions recombine between sexes and thus have a higher rate of neutral fixation than the SDR, which experiences the Hill–Robertson effect that reduces fixation probabilities of neutral mutations due to its linkage with other sites under selection66. We speculate that this feature could facilitate the evolution of TRGs. Alternatively, the high density of DNA transposons within the U/V could also promote the co-option of their regulatory motifs and enable de novo transcript birth, as seen in Drosophila67. Note that the pattern could be reinforced through generation-antagonistic selection37, but DNA transposons and higher mutation rates may be sufficient to initiate this pattern in species lacking sporophyte-biased gene expression. Importantly, the TRG enrichment is unique to U/V systems and gradually disappears when these systems are lost. This pattern extends beyond brown algae to other eukaryotes with U/V systems, such as C. purpureus11,68 and S. angustifolium12. However, we could not detect enrichment of TRGs on the small U/V sex chromosomes of M. polymorpha42,69, or on the mating-type chromosome of C. neoformans43. Mosses such as C. purpureus and S. angustifolium display relatively more complex sporophyte body plans than liverworts such as M. polymorpha70, which could underlie stronger generation-antagonistic selection37. Unlike brown algae, most of the sex chromosomes in bryophytes are sex linked, with minimal space for PARs29. In this context, the sex chromosome of C. purpureus expanded its SDR very recently in evolutionary time through fusions of autosomes with earlier established U and V chromosomes71, retaining its evolutionary footprints as PARs, while the sex-linked region in M. polymorpha is much older69, which could also limit the formation of TRGs that are predominantly observed in the PARs of the brown algae. The pattern thus appears to be specific to U/V systems where chromosomal degeneration is mild and linked to haploid–diploid life cycles where the sporophyte stage is sufficiently complex, highlighting a key role for generation selection37. Therefore, our study hints at a unique interplay between complex life cycles, heterochromatic landscape, DNA transposons and higher mutation rates that may lead to TRG enrichment in U/V chromosomes, and this process is pervasive across distant, independently evolved eukaryotic kingdoms.
Monoicous brown algae have transcriptomic profiles resembling ancestral females33. However, our results show that monoicy arose at least twice from a male ancestor that acquired female genes. The male pathway requires MIN22,25,72, which could have facilitated the evolution of monoicy from males, as also seen in the green lineage73,74. We note the presence of U-SDR-derived gene(s) in all monoicous species, particularly a cholesterol transporter gene that is found in all U-SDRs, suggesting that the U-SDR contains femaleness-promoting factor(s), consistent with reports in kelps75. We thus propose that monoicy evolved via translocation events adding essential femaleness-promoting genes to a male genetic background. Since the resulting monoicous individuals were capable of producing both male and female reproductive structures, individuals with U chromosomes were no longer essential for sexual reproduction, ultimately leading to the loss of the U-SDR in monoicous species. Although the combination of key female and male genes is essential for this evolutionary transition, the retention of a sex chromosome is not. For example, in Volvox africanus, monoicy required the retention of female SDR-like regions, while most male SDR genes were lost except for a multicopy array of the male-determining gene MID76.
The evolution of a dioecious system in F. serratus is associated with an irreversible transition to diploidy in Fucales16,47. The U/V to dioecy transition has remained elusive2,47, but our data in brown algae imply that it involved an intermediate monoicous stage, supporting previous predictions from ancestral state reconstruction analyses16. A small, undifferentiated Y-specific region consistent with a young XY system may explain why the sex chromosome in F. serratus was undetectable. Nonetheless, all ancestral V-SDR genes are found in the U/V-homologue of F. serratus, several showing a male-biased gene expression across Fucales species, particularly MIN22. Our findings imply that MIN and possibly other ancestral V-SDR genes are still involved in male differentiation, but shifted downwards in the sex determination cascade. These results thus support and extend the ‘bottom-up’ hypothesis of sex determination, where downstream components of sex differentiation are conserved across taxa, and new master sex regulators can replace older ones77.
Methods
Biological material
Scytosiphon promiscuus, Dictyota dichotoma, Undaria pinnatifida and Desmarestia dudresnayi haploid gametophytes were cultivated under laboratory conditions as in ref. 78. We cultivated the gametophytes at 14 °C with a photoperiod of 12:12 h light:dark at an irradiance of 25 µmol photons m−2 s−1. The media consisted of filtered natural seawater (NSW), which was autoclaved and enriched with half-strength Provasoli nutrient solution (Provasoli-enriched seawater; PES)78. We grew the first biomass in 140 mm Petri dishes and the gametophytes were later transferred to a 1 l flask with gentle aeration. The gametophytes were fragmented once a month and the media were changed every 2 weeks to promote biomass production. Before freezing, gametophytes were treated with antibiotics for 3 days with gentle agitation and under the same culture conditions. The first day, gametophytes were treated with a mix of streptomycin (2 g l−1 of PES), penicillin G (0.5 g l−1 of PES) and chloramphenicol (0.1 g l−1 of PES); the next day with ampicilin (1 g l−1 of PES), and on the last day with kanamycin (1 g l−1 of PES). Between each day of treatment and before freezing, gametophytes were rinsed with 500 ml of NSW to remove traces of antibiotic.
Samples of fucoid algae sexual and vegetative tissue were collected in the intertidal zone during low tides in June 2012 from Viana do Castelo (F. vesiculosus, A. nodosum) and Caminha (Rio Minho; F. ceranoides), northern Portugal. Sexual phenotypes were verified in the field by sectioning and observing receptacles under a field microscope. Tissue samples were flash frozen in liquid nitrogen on the shore and transported to the laboratory in a cryoshipper, after which they were lyophilised and stored dry at room temperature on silica crystals (see Supplementary Table 1 for a list of strains used in this study).
DNA and RNA extraction and sequencing
Genomic DNA was isolated from algal tissue (~100 mg) by grinding into fine powder under liquid nitrogen and subsequent cell lysis in 500 μl of Genomic Lysis Buffer (OMNIPREP for plant kit) for 1 h at 60 °C. The lysate was cleaned up with 200 μl of chloroform and DNA was precipitated in ethanol. The DNA pellet was digested in CF buffer (Macherey–Nagel) for 45 min at 65 °C and purified using NucleoBond AXG20 Mini columns according to the user manual (Macherey–Nagel). Final high molecular weight genomic DNA was quantified (Qubit), analysed for purity (Nanodrop) and checked for size distribution (Femto Pulse System) before preparing the sequencing libraries. We sequenced the libraries using an Oxford Nanopore Technologies (ONT) MinION Mk1B. We prepared the ONT libraries using an SQK-LSK110 library preparation kit for R9.4.1 flow cells and an SQK-LSK114 library preparation kit for R10.4.1 flow cells. Two libraries were sequenced for D. dudresnayi on R9.4.1 flow cells and a third library was sequenced on an R10.4.1 flow cell.
RNA was isolated from mature gametophytes of U. pinnatifida and S. promiscuus following modified procedure of the Qiagen RNAeasy kit, and the TruSeq RNA Library Prep Kit v.2 was used to sequence the transcriptomes in an Illumina NextSeq 2000 platform (150 bp, PE reads). Extraction of total RNA from fucoid algae (F. vesiculosus, A. nodosum and F. ceranoides) was performed following ref. 79 and RNA libraries were sequenced on an Illumina HiSeq 2000 machine (100 bp, PE reads).
Genome assembly and annotation
High-quality, chromosome-level assemblies of brown algal genomes have been notoriously difficult to obtain due to technical challenges in extracting nucleic acids. Whole-genome assemblies and annotations of S. promiscuus male, D. dichotoma male, D. herbacea male and female, E. crouanorium male, C. linearis, S. ischiensis and F. serratus male were obtained from ref. 18. We also downloaded the genome of Ectocarpus sp. 7 (ref. 19) and the male genome of U. pinnatifida20, which were already assembled at a chromosome level. For D. dudresnayi, we performed genome sequencing, de novo genome assembly and ab initio gene annotation. Base calling was done using ONT Guppy80 with the configuration files ‘dna_r9.4.1_450bps_sup.cfg’ and ‘dna_r10.4.1_e8.2_400bps_sup.cfg’ and the options ‘–trim_adapters –trim_primers’, yielding 17.4 Gbp of data in 2,871,152 reads. We merged all the reads and analysed them using Kraken (v.2.1.2)81 and the bacteria database (August 2022) to remove potential contaminant sequences. All data classified as bacterial reads by Kraken were screened using blastN (v.2.13.0+)82 (-evalue 0.001 -num_alignments 20) against the NCBI genbank bacterial database (downloaded November 2023). The blastN output was visualized in MEGAN (v.6.23.4)83, and all reads that were declared as bacterial were extracted and removed from further analyses. We obtained 1,908,772 decontaminated reads with an average length of 5.1 Kbp (9.8 Gbp of data, 20× coverage), which were deposited on the NCBI Sequence Read Archive (Supplementary Table 1).
The decontaminated reads were assembled de novo using flye (v.2.9.1-b1780)84 with the options ‘–nano-raw -g 450 m -t 28 -i 3 –scaffold’. The draft assembly consisted of 1,032 contigs with a total size of 425 Mbp, an N50 of 4.6 Mbp and an L50 of 29 contigs. We used TransposonPSI (http://transposonpsi.sourceforge.net/) to predict the TEs and RepeatScout (v.1.0.6)85 to predict the simple repeats in the genome assembly. Both predictions were combined to soft mask the repetitive content in the genome assembly using bedtools maskfasta (v.2.27.1)86. We mapped the RNA-seq data of D. dudresnayi from the PhaeoExplorer database18 to the soft-masked genome assembly using STAR (v.2.7.1a)87. We used BRAKER v.2.1.6 alongside the RNA-seq data88 to predict the protein-coding genes in the soft-masked genome assembly.
Hi-C library preparation and sequencing for chromosome-level assemblies
We generated Hi-C libraries for three male genomes (S. promiscuus, D. herbacea and D. dichotoma) and two female genomes (Ectocarpus sp. 7 and D. herbacea). Fresh algal tissue was cross-linked for 20 min at room temperature in a solution of 2% formaldehyde with filtered NSW and then transferred into a 400 mM glycine solution with filtered NSW for 5 min to quench the formaldehyde. The samples were then stored at −80 °C until use. The Hi-C libraries were prepared as follows. The samples were de-frosted in 1 ml of 1× DpnII buffer with protease inhibitors (Roche cOmplete), transferred to Precellys VK05 lysis tubes (Bertin) and disrupted using the Precellys apparatus with five grinding cycles of 30 s at 7,800 r.p.m., followed by 20 s pauses. SDS was added to the lysate at 0.5% final concentration and samples were incubated for 10 min at 62 °C, followed by the addition of Triton X-100 to a final concentration of 1% and 10 min of incubation at 37 °C under gentle shaking. We added 500 U of DpnII to 4.6 ml of the digestion mixture and incubated the samples for 2 h at 37 °C under gentle shaking (180 r.p.m. in an inclined rack to prevent sedimentation), followed by the addition of another 500 U of DpnII and an overnight incubation under the same conditions. The digested samples were centrifuged at 4 °C for 20 min at 16,000 × g. The supernatant was discarded and the pellet was incubated for biotinylation at 37 °C for 1 h under a constant shaking (300 r.p.m.) in a 500 ml biotinylation mix with a concentration of 1× ligation buffer, 0.09 mM dATP-dGTP-dTTP, 0.03 mM biotin-14-dCTP and 0.64 U ml−1 Klenow fragments. After biotinylation, the samples were incubated for 3 h at room temperature in a 1.2 ml ligation reaction with a concentration of 1× ligation buffer, 100 mg ml−1 BSA, 1 mM ATP and 0.4 U ml−1 T4 DNA Ligase. The samples were then incubated overnight at 65 °C after adding 20 μl 0.5 M EDTA, 80 μl 10% SDS and 1.6 mg Proteinase K. DNA was extracted with 1 volume of phenol/cholorform/isoamyl alcohol (24:24:1), followed by 30 s of vortexing at top speed and a 5-min centrifugation at top speed. We precipitated the DNA by adding 1/10 volume of 3 M NaAC pH 5 and two volumes of 100% cold ethanol, followed by a 30-min incubation at −80 °C and a 20-min centrifugation at 14,000 × g and 4 °C. The DNA pellet was washed with 1 ml 70% ethanol, then dried at 37 °C for 10 min and resuspended in 100 μl 1× TE buffer with 1 mg ml−1 RNase. DNA was sheared to 250–500 bp fragments using a Covaris S220 ultrasonicator, purified with AMPure beads (0.6×) (Beckman) and eluted in 20 μl 10 mM Tris pH 8.0. Biotinylated but not ligated DNA fragments were first removed by T4 DNA polymerase treatment (final concentration, 300 U per pellet; NEB), and the biotin-labelled fragments were selectively captured by Dynabeads MyOne Streptavidin C1 beads (Invitrogen). The libraries were prepared using the NEB Ultra II library preparation system and sequenced on the NextSeq 2000 Illumina platform (2 × 150 bp) (Supplementary Table 1).
We scaffolded the genomes from ref. 18 into chromosome-level assemblies using the Hi-C data. We filtered the low-quality Hi-C reads using Trimmomatic (v.0.39)89 (ILLUMINACLIP:2:30:10 LEADING:25 TRAILING:25 SLIDINGWINDOW:4:15 MINLEN:75 AVGQUAL:28). We mapped the Hi-C reads against each genome assembly using BWA-mem v.0.7.17-r1188d in the Juicer v.1.6 pipeline90 to generate a contact map, which was then fed to 3D-DNA v190716 (ref. 91) to scaffold the genomes into chromosomes. The obtained scaffolds were manually inspected against the contact maps to solve the limits of each chromosome using Juicebox (v.1.11.08)92. The PhaeoExplorer gene annotations18 were lifted into the new assemblies using Liftoff (v.1.6.1)93, while the annotation of TEs was performed using RepeatModeler2 (ref. 94). We scaffolded the genomes of E. crouaniorum and D. dudresnayi into chromosomes using a reference-guided assembly with RagTag (v.2.0.1)95 against the chromosome-level assemblies of Ectocarpus sp. 7 and D. herbacea, respectively. All genes within the SDRs in the brown algal species studied (see below) were manually curated to exclude any TE-related genes from the annotation.
Discovery of the U/V sex determination regions
Male sex-determining regions (V-SDR) in S. promiscuus, U. pinnatifida, D. herbacea and D. dichotoma, as well as female sex-determining region (U-SDR) in D. herbacea were analysed following two complementary methods: (1) a k-mer-based YGS approach, originally designed to detected Y-linked sequences in heterogametic systems, developed in ref. 96 and (2) genomic coverage analysis, designed to identify sex-linked regions through differences in read depth between male and female individuals97. These methods are well suited for organisms with divergent sex chromosomes, such as brown algae, where U and V haplotypes have diverged over extended evolutionary time.
The YGS method principle is to identify male or female sex-linked scaffolds by comparing k-mer frequencies between reference genome assembly and k-mers generated from DNA-seq reads of the opposite sex. Regions in the male reference genome that contain k-mers that are absent in female reads will indicate candidate male SDR sequences; similarly, female genomic scaffolds with low coverage in male k-mers will denote female SDR region. For each species, 15-base-pair k-mer sequences were generated separately from male and female Illumina reads (see Supplementary Table 1 for data accession numbers) using Jellyfish v.2.3.0 count (-m 15 -s 10 G -C –quality-start=33 –min-quality=20) and converted to fasta format with Jellyfish dump (–lower-count=5)98. Next, non-overlapping 500-kb sliding windows (Desmarestia, Dictyota and Undaria) or 200-kb sliding windows (Scytosiphon) of the reference chromosome genomes (from the sex whose SDR was to be identified) were created using seqkit (v.2.3.1)99 and used as input for the YGS.pl script96 together with the fasta k-mer files produced in the previous step. Each window was then analysed to calculate the proportion of k-mers in the reference window that are not present in the opposite-sex k-mer database. Genomic windows with a minimum of ≥50% of unmatched single-copy k-mers were then retained as candidate male or female SDR sequences. Because the borders of the SDRs cannot be precisely defined at the single-nucleotide level with the available data, we focused on genes within these regions and defined the SDR boundaries on the basis of the flanking genes located at the transition to pseudoautosomal regions (PARs).
Candidate SDR regions identified by YGS were further validated by analysing sex-specific differences in read coverage. In detail, the short Illumina reads coming from males and females of each investigated species were trimmed with Trimmomatic89 (see above) and mapped to the reference genome for which the SDR was to be studied, using HISAT2 (ref. 100) (default settings). Bam files produced by HISAT2 were used as input for Mosdepth101 to calculate coverage in 10-kb windows along the genome sequence (-m -n -b 10000 –fast-mode -Q 30). Read mapping depth in genomic windows was normalized by the genome-wide mean for each sex, and the coverage in genomic intervals was then compared between males and females. Because V-SDR-linked sequences are present only in males, we expect them to have similar read coverage as autosomal regions in males, but little or no coverage in females (and conversely for the U-SDR sequences in D. herbacea). The comparison focused on regions within male reference genomes where the coverage in males fell within the range of 75–125% of the genome average, while the coverage in females remained below 50% of the genome average.
Both coverage and k-mer analysis identified identical genomic regions, providing high-confidence candidate SDRs (Extended Data Table 1 and Supplementary Figs. 1–5). In D. herbacea, where both male and female chromosome-level genome assemblies were available, we directly compared U and V chromosomes to further confirm the SDR borders by analysing the collinearity of pseudoautosomal regions flanking the SDRs. The SDR scaffolds for all studied species were further validated by PCR amplification (Supplementary Table 1) using 4 males and 4 females.
Genetic mapping and search for the sex chromosome in F. serratus
Three different sets of materials were used in this study: (1) 12 male and 12 female field samples, hereafter denoted the 24-individual natural population; (2) 157 sporophyte progeny population derived from a cross between one male sample and one female sample collected from the field and (3) 3 male and 3 female samples collected from the field for whole-genome sequencing. The 157-progeny population and 24-individual natural population were genotyped using the double digest RAD sequencing approach (ddRAD-seq). Briefly, individual genomic DNA was digested with the restricted enzymes PstI and HhaI to obtain fragments that were size selected between 400 and 800 bp before sequencing on in Illumina HiSeq 2500 platform (paired-end 2 × 125 bp). See ref. 102 for detailed protocol of the ddRAD-seq.
We performed whole-genome sequencing on an Illumina HiSeq 2500 system (2 × 150 bp paired-end) for the 3 male and 3 female samples. For ddRAD-seq data, raw reads were cleaned and trimmed with Trimmomatic as above and mapped to the draft genome of F. serratus male. For the progeny population, genotypes were called from the obtained bam files using the Stacks pipeline (v.2.5)103. The obtained vcf files were filtered with VCFtools (v.0.1.16)104 and bcftools105 (max missing per locus: 30%, max missing per sample: 40%, max mean coverage:30, minQG:20).
The filtered vcf file of the progeny population was used to construct a genetic map with Lep-MAP3 (ref. 106). Briefly, the ParentCall2 module was used to call parental genotypes, the SeparateChromosomes2 module was used to split the markers into linkage groups and the OrderMarkers2 module was used to order the markers within each linkage group using 30 iterations per group and finally computing genetic distances. Phased data were converted to informative genotypes with the script map2genotypes.awk.
We used different approaches to identify the SDR in F. serratus
Coverage analysis
We combined whole-genome sequence data from the 3 males and 3 females alongside the ddRAD-seq data of the 24-individual natural population, mapping both datasets to the F. serratus male genome assembly using bwa-mem107. Coverage analyses was done in several ways:
-
Using SATC (sex assignment through coverage)108, a method that uses sequencing depth distribution across scaffolds to jointly identify: (1) male and female individuals and (2) sex-linked scaffolds. This identification was achieved by projecting the scaffold depths into a low-dimensional space using principal component analysis and subsequent Gaussian mixture clustering. Male and female whole-genome sequences were used for this analysis.
-
Using the method SexChrCov described in ref. 109 with the 24-individual natural population.
-
Using the method DifCover110 which identifies regions in a reference genome for which the read coverage of one sample is significantly different from the read coverage of another sample when aligned to a common reference genome. The 24-individual natural population was used for this analysis.
-
Using soap.coverage (v.2.7.9)111 to calculate the coverage (number of times each site was sequenced divided by the total number of sequenced sites) of each scaffold in each sample. For each scaffold, the male to female (M:F) fold change coverage was calculated as log2(average male coverage) – log2(average female coverage). The 24-individual natural population was used for this analysis.
Fixation index (FST) and sex-biased heterozygosity
This approach has been previously used to find sex-linked genomic regions in several studies112,113. Using the 24-individual natural population, FST was calculated using vcftools104. Sex-biased heterozygosity was defined as the log10 of the male heterozygosity:female heterozygosity ratio, where heterozygosity was measured as the fraction of sites that are heterozygous. This ratio is expected to be zero for autosomal scaffolds and elevated on young sex scaffolds due to excess heterozygosity in males.
Identification of eventual female scaffolds that failed to map to the male reference genome
Vcftools and bedtools were used to extract female regions that did not map to the reference genome consistently in the 3 resequenced female samples.
All candidate contigs were tested by PCR in 4 males and 4 females.
Synteny analyses, K s analysis and transitions to co-sexuality
Whole-genome synteny comparisons were performed for each pair of chromosome-level assemblies using MCscan (v.1.2.14)114, both between different species, between sex chromosomes in the same species and between monoicous species and their closest relatives with U/V chromosomes. The putative gametologues between sex chromosomes that were predicted with MCscan were reassessed using OrthoFinder (v.2.5.4)115 and best reciprocal DIAMOND (v.2.1.8.162)116 hits.
We calculated the number of synonymous substitutions per synonymous site (Ks) for each pair of male and female gametologues as a proxy to assess the relative time at which both genes diverged from each other. The amino acid sequences of each pair of gametologues were aligned with MAFFT (v.7.520)117 and subsequently aligned into codons using pal2nal (v.14)118. The gametologue Ks values were calculated using the model in ref. 119 as implemented in KaKs_calculator (v.2.0)120.
We evaluated the male or female identity of the genes in the monoicous species whose orthologues were found within the SDR in their closest dioicous relatives. For this, we compared the results obtained with MCscan114 against the orthogroup prediction performed with OrthoFinder115, with best reciprocal DIAMOND116 hits and by calculating gene trees for each orthogroup using an amino acid alignment with MAFFT117 and gene tree reconstructions using FastTree (v.2.1.11)121.
Ancestral reconstruction of the male SDR
The brown algal phylogeny was obtained from ref. 18. The species tree is based on 32 single-copy nuclear genes whose protein sequences were aligned manually using AliView122, and whose best-fit substitution models were assessed independently using the Akaike information criterion. The tree was generated using a maximum likelihood approach implemented in RAxML bootstraps and the gamma model. Every node in the phylogeny has 99–100% bootstrap support values. Divergence times were subsequently calculated using MCMCtree123 and three calibration points. The MCMC chains were run for 1.5 million generations and the first 200,000 MCMC chains were discarded as burn-in.
We searched for orthologue genes within the V-SDR of five species (Ectocarpus sp. 7, S. promiscuus, U. pinnatifida, D. herbacea and D. dichotoma) in our OrthoFinder results. For each V-SDR gene, we coded its orthologue in the other species as ‘present’ (1) if it is also sex linked in the V-SDR, whereas it was coded as ‘absent’ (0) if the orthologue resides in the PARs, in an autosome or if there is no detectable orthologue in that species. Once we generated this presence/absence matrix with the evolutionary relationship of the genes within the V-SDR (Supplementary Table 5), we used it as the input file for the software Count (v.10.04)124 to estimate the ancestral content of the V-SDR throughout a phylogeny and determine the most likely scenario of V-SDR evolution in the brown algae. We employed posterior probabilities under a phylogenetic birth-and-death model with independent gain and loss rates across each branch in the phylogeny. We modelled the independent gain and loss rates through 10 gamma categories and performed 1,000 optimization rounds with a convergence threshold on the likelihood >0.1 to find the best fitting model for the data. The branch lengths in the tree that were used for the ancestral state reconstruction were retrieved from the molecular clock analysis performed in ref. 16. We distinguished between conserved V-SDR genes that are ancestral and parallel acquisitions of the same gene in the V-SDR by analysing gene trees between male and female genomes, in addition to female transcriptome assemblies of D. dichotoma and U. pinnatifida. Sequence alignments were done using MAFFT117 with default settings and uploaded to the https://www.phylogeny.fr/ platform. Alignments were further curated using Gblocks (v.0.91b)125 (min. seq. for flank pos.: 85%, max. contig. nonconserved pos.: 8, min. block length: 10). Trees were produced using PhyML (v.3.1l)126 with the default model and visualized in TreeDyn (v.198.3)127. The approximate likelihood-ratio test was chosen as the statistical test for branch support. We inferred the function of the ancestral V-SDR genes through the annotation of genes in Ectocarpus sp. 7 belonging to that orthogroup. The most likely acquisition mechanism of each SDR gene in each species was assessed on the basis of the position of each orthologue in the other species (pseudoautosomic, autosomic or missing; Supplementary Table 5).
Genomic content across chromosomes
We used closely related genome assemblies available in the PhaeoExplorer database18 to assess the depletion of orthologues in the sex chromosome. We predicted one-to-one orthologues using OrthoFinder115 between the following species pairs: Ectocarpus sp. 7 with Ectocarpus siliculosus, S. promiscuus with C. linearis, U. pinnatifida with Saccharina japonica, F. serratus with Fucus distichus, D. herbacea with D. dudresnayi, and D. dichotoma with Halopteris paniculata (Supplementary Table 16). We calculated the expected number of detectable orthologues for each chromosome and compared it against the observed number of detected orthologues using chi-square tests. We performed Benjamini–Hochberg corrections to the P values of the chi-square tests to control the FDR in the analysis128.
GenEra36 was used by running DIAMOND in ultra-sensitive mode116 against the NCBI NR database and all the PhaeoExplorer proteins18 to perform a phylostratigraphic analysis (e-value threshold of 10−5) and calculate the relative ages of each gene in each genome (Supplementary Table 13). Phylostratigraphy is a genetic statistical method developed to date the putative origin of all the genes contained in the genome of a target species by detecting homologues across species at different evolutionary distances (all the way from species within the same genus to species from different domains of life). Finding the most distant homologues of each gene can link them to their founder events (that is, the first instance where a gene homologue is found in the history of that lineage), allowing us to then determine their relative ages, coded as the taxonomic group where that gene is detected35,36,129. The gene age categories outside of the brown algae and S. ischiensis were based on the taxonomic classification of each species within the NCBI Taxonomy database130, while the gene ages within the brown algae were manually assessed to reflect the evolutionary relationships obtained in the PhaeoExplorer maximum likelihood tree18. We performed Wilcoxon rank-sum tests in R (v.4.3.1)131 to assess non-random differences in gene age distributions between pairs of chromosomes (Supplementary Table 14). We performed Benjamini–Hochberg corrections to the P values of the Wilcoxon rank-sum tests to control the FDR in the analysis128. The gene ages responsible for these differences were found by evaluating the standardized residuals using mosaic plots.
We used the interspecies Ks values between pairs of species as a proxy for neutral mutation rates across six of the seven chromosome-level assemblies by using the most closely related genome assemblies available in the PhaeoExplorer database18. We used the same set of one-to-one orthologues detected between species pairs as for the orthologue-depletion test (Supplementary Table 16). However, the evolutionary distance between D. dichotoma and H. paniculata prevented us from calculating reliable interspecies Ks values for this species since synonymous substitutions reached the point of saturation. The amino acid sequences of each pair of orthologues were aligned with MAFFT117 and subsequently aligned into codons using pal2nal118. The interspecies Ks values were calculated using the model in ref. 119 as implemented in KaKs_calculator (v.2.0)120. We also evaluated the difference in interspecies Ks values between the autosomes and the sex chromosomes through FDR-corrected Wilcoxon rank-sum tests (Supplementary Table 17). We calculated the protein-coding density, the density of TEs and the taxonomic identity of these TEs within 100-kb non-overlaping windows across each chromosome using bedtools86 (Supplementary Table 9). The differences in protein-coding space and repeat content between the autosomes and the sex chromosomes were also tested using FDR-corrected Wilcoxon rank-sum tests (Supplementary Tables 10 and 11). The differences in repeat density, percentage of unclassified repeats and gene ages across genomic compartments (SDR, PARs and autosomes) were tested using FDR-corrected permutation tests with 10,000 permutations. All genomic features were plotted using karyoploteR (v.1.20.3)132.
Gene expression analysis
We used kallisto (v.0.44.0)133 to calculate gene expression levels using 31-base-pair-long k-mers and 1,000 bootstraps. Transcript abundances were then summed within genes using the tximport v.3.19 package134 to obtain the expression level for each gene in transcripts per million (TPM). Differential expression analysis was done in the DESeq2 v.3.19 package135 in R v.4.3.1, applying FC ≥ 2 and padj < 0.05 cut-offs. Sex-biased gene expression analysis in Ectocarpus sp. 7, S. promiscuus, U. pinnatifida, D. herbacea and D. dichotoma was performed between mature male and female gametophytes (gametophytes bearing reproductive structures). To discover genes with sporophyte-biased expression in Ectocarpus sp.7, S. promiscuus, U.pinnatifida and D. dichotoma, we first calculated the differential expression between male gametophytes and sporophytes, as well as female gametophytes and sporophytes. Genes that showed significant sporophyte-biased expression (FC ≥ 2, padj < 0.05) in both comparisons were considered sporophyte biased.
A total of 314.2 M RNA-seq reads from F. vesiculosus male, female and vegetative tissue were assembled de novo with rnaSPAdes136 using k-mer values of 33 and 49. Assembly quality was assessed by (pseudo)mapping reads back onto the resulting assembly and retaining ‘good’ contigs as defined using TransRate (v.1.0.3)137 with default settings. The resulting 159,108 contigs were aligned with BLASTx82 against a database of Stramenopile proteins, and those with top hits against brown algae (Phaeophyceae) were retained as the final curated reference transcriptome (36,394 contigs, N50 = 1,770 bp). Transcript expression levels were determined by mapping the reads from all samples against the reference transcriptome using Bowtie2 (ref. 138) and the RSEM-EBSeq (v.1.3.3)139 pipeline, and relative expression values were recorded as TPM. All samples used in the gene expression analysis can be found in Supplementary Table 1.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The accession numbers and download links for all the chromosome-level genome assemblies and annotations that were generated and used in this study are available in Supplementary Table 1 and in the Edmond Repository140 at https://doi.org/10.17617/3.OOWB2Y. The raw sequence reads for the Oxford Nanopore data, Hi-C libraries and RNA-seq libraries are available in the Sequence Read Archive under BioProject accession number PRJNA1059008. All genome assemblies and annotations are also accessible through the Phaeoexplorer database (https://phaeoexplorer.sb-roscoff.fr/) for comparative genomics analyses. This paper does not report original data. Further information and requests for resources and reagents should be directed to and will be fulfilled by S.M.C. (susana.coelho@tuebingen.mpg.de).
Code availability
This paper does not report original code.
References
Bachtrog, D. et al. Sex determination: why so many ways of doing it? PLoS Biol. 12, e1001899 (2014).
Beukeboom, L. W. & Perrin, N. The Evolution of Sex Determination (Oxford Univ. Press, 2014).
Vicoso, B. Molecular and evolutionary dynamics of animal sex-chromosome turnover. Nat. Ecol. Evol. 3, 1632–1641 (2019).
Mignerot, L. & Coelho, S. M. The origin and evolution of the sexes: novel insights from a distant eukaryotic linage. C. R. Biol. 339, 252–257 (2016).
Ma, W. & Rovatsos, M. Sex chromosome evolution: the remarkable diversity in the evolutionary rates and mechanisms. J. Evol. Biol. 35, 1581–1588 (2022).
Umen, J. & Coelho, S. Algal sex determination and the evolution of anisogamy. Annu. Rev. Microbiol. 73, 267–291 (2019).
Coelho, S. M. & Umen, J. Switching it up: algal insights into sexual transitions. Plant Reprod. 34, 287–296 (2021).
Avia, K. et al. Genetic diversity in the UV sex chromosomes of the brown alga Ectocarpus. Genes 9, 286 (2018).
Coelho, S. M., Mignerot, L. & Cock, J. M. Origin and evolution of sex‐determination systems in the brown algae. New Phytol. 222, 1751–1756 (2019).
Bull, J. J. Sex chromosomes in haploid dioecy: a unique contrast to Muller’s theory for diploid dioecy. Am. Nat. 112, 245–250 (1978).
Carey, S. B. et al. Gene-rich UV sex chromosomes harbor conserved regulators of sexual development. Sci. Adv. 7, eabn2488 (2021).
Healey, A. L. et al. Newly identified sex chromosomes in the Sphagnum (peat moss) genome alter carbon sequestration and ecosystem dynamics. Nat. Plants 9, 238–254 (2023).
Bowman, J. L. et al. Insights into land plant evolution garnered from the Marchantia polymorpha genome. Cell 171, 287–304.e15 (2017).
Silva, A. T. et al. To dry perchance to live: insights from the genome of the desiccation‐tolerant biocrust moss Syntrichia caninervis. Plant J. 105, 1339–1356 (2021).
Bechteler, J. et al. Comprehensive phylogenomic time tree of bryophytes reveals deep relationships and uncovers gene incongruences in the last 500 million years of diversification. Am. J. Bot. 110, e16249 (2023).
Heesch, S. et al. Evolution of life cycles and reproductive traits: insights from the brown algae. J. Evol. Biol. 34, 992–1009 (2021).
Bringloe, T. T. et al. Phylogeny and evolution of the brown algae. Crit. Rev. Plant Sci. 39, 281–321 (2020).
Denoeud, F. et al. Evolutionary genomics of the emergence of brown algae as key components of coastal ecosystems. Cell 187, 6943–6965.e39 (2024).
Liu, P. et al. 3D chromatin maps of a brown alga reveal U/V sex chromosome spatial organization. Nat. Commun. 15, 9590 (2024).
Shan, T. et al. First genome of the brown alga Undaria pinnatifida: chromosome-level assembly using PacBio and Hi-C technologies. Front. Genet. 11, 140 (2020).
Choi, S.-W. et al. Ordovician origin and subsequent diversification of the brown algae. Curr. Biol. https://doi.org/10.1016/j.cub.2023.12.069 (2024).
Luthringer, R. et al. Repeated co-option of HMG-box genes for sex determination in brown algae and animals. Science 383, eadk5466 (2024).
Simakov, O. et al. Deeply conserved synteny and the evolution of metazoan chromosomes. Sci. Adv. 8, eabi5884 (2022).
Sun, Y., Svedberg, J., Hiltunen, M., Corcoran, P. & Johannesson, H. Large-scale suppression of recombination predates genomic rearrangements in Neurospora tetrasperma. Nat. Commun. 8, 1140 (2017).
Ahmed, S. et al. A haploid system of sex determination in the brown alga Ectocarpus sp. Curr. Biol. 24, 1945–1957 (2014).
Charlesworth, D. Plant contributions to our understanding of sex chromosome evolution. New Phytol. 208, 52–65 (2015).
Lahn, B. T. & Page, D. C. Four evolutionary strata on the human X chromosome. Science 286, 964–967 (1999).
Lipinska, A. P. et al. Multiple gene movements into and out of haploid sex chromosomes. Genome Biol. 18, 104 (2017).
Charlesworth, D. The mysterious sex chromosomes of haploid plants. Heredity 129, 17–21 (2022).
Bolwell, G. P., Callow, J. A., Callow, M. E. & Evans, L. V. Fertilization in brown algae: II. evidence for lectin-sensitive complementary receptors involved in gamete recognition in Fucus serratus. J. Cell Sci. 36, 19–30 (1979).
Zhao, Z.-S., Leung, T., Manser, E. & Lim, L. Pheromone signalling in Saccharomyces cerevisiae requires the small GTP-binding protein Cdc42p and its activator CDC24. Mol. Cell. Biol. 15, 5246–5257 (1995).
Ponnikas, S., Sigeman, H., Abbott, J. K. & Hansson, B. Why do sex chromosomes stop recombining? Trends Genet. 34, 492–503 (2018).
Cossard, G. G. et al. Selection drives convergent gene expression changes during transitions to co-sexuality in haploid sexual systems. Nat. Ecol. Evol. 6, 579–589 (2022).
Bachtrog, D. Accumulation of Spock and Worf, two novel non-LTR retrotransposons, on the neo-Y chromosome of Drosophila miranda. Mol. Biol. Evol. 20, 173–181 (2003).
Domazet-Lošo, T., Brajković, J. & Tautz, D. A phylostratigraphy approach to uncover the genomic history of major adaptations in metazoan lineages. Trends Genet. 23, 533–539 (2007).
Barrera-Redondo, J., Lotharukpong, J. S., Drost, H.-G. & Coelho, S. M. Uncovering gene-family founder events during major evolutionary transitions in animals, plants and fungi using GenEra. Genome Biol. 24, 54 (2023).
Luthringer, R. et al. The pseudoautosomal regions of the U/V sex chromosomes of the brown alga Ectocarpus exhibit unusual features. Mol. Biol. Evol. 32, 2973–2985 (2015).
Kimura, M. Preponderance of synonymous changes as evidence for the neutral theory of molecular evolution. Nature 267, 275–276 (1977).
King, J. L. & Jukes, T. H. Non-Darwinian evolution. Science 164, 788–798 (1969).
Eyre-Walker, A. & Keightley, P. D. High genomic deleterious mutation rates in hominids. Nature 397, 344–347 (1999).
Keightley, P. D. & Eyre-Walker, A. Deleterious mutations and the evolution of sex. Science 290, 331–333 (2000).
Montgomery, S. A. et al. Chromatin organization in early land plants reveals an ancestral association between H3K27me3, transposons, and constitutive heterochromatin. Curr. Biol. 30, 573–588.e7 (2020).
Loftus, B. J. et al. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science 307, 1321–1324 (2005).
Clayton, M. N. Isogamy and a fucalean type of life history in the Antarctic brown alga Ascoseira mirabilis (Ascoseirales, Phaeophyta). Bot. Mar. 30, 447–454 (1987).
Coyer, J. A., Peters, A. F., Hoarau, G., Stam, W. T. & Olsen, J. L. Hybridization of the marine seaweeds, Fucus serratus and Fucus evanescens (Heterokontophyta: Phaeophyceae) in a 100-year-old zone of secondary contact. Proc. R. Soc. B 269, 1829–1834 (2002).
Hatchett, W. J. et al. Evolutionary dynamics of sex‐biased gene expression in a young XY system: insights from the brown alga genus Fucus. New Phytol. 238, 422–437 (2023).
Cánovas, F. G., Mota, C. F., Serrão, E. A. & Pearson, G. A. Driving south: a multi-gene phylogeny of the brown algal family Fucaceae reveals relationships and recent drivers of a marine radiation. BMC Evol. Biol. 11, 371 (2011).
Lee, S. C., Ni, M., Li, W., Shertz, C. & Heitman, J. The evolution of sex: a perspective from the fungal kingdom. Microbiol. Mol. Biol. Rev. 74, 298–340 (2010).
Duhamel, M., Hood, M. E., Rodríguez de la Vega, R. C. & Giraud, T. Dynamics of transposable element accumulation in the non-recombining regions of mating-type chromosomes in anther-smut fungi. Nat. Commun. 14, 5692 (2023).
Dolgin, E. S. & Charlesworth, B. The effects of recombination rate on the distribution and abundance of transposable elements. Genetics 178, 2169–2177 (2008).
Gray, Y. H. M. It takes two transposons to tango: transposable-element-mediated chromosomal rearrangements. Trends Genet. 16, 461–468 (2000).
Charlesworth, D. Evolution of recombination rates between sex chromosomes. Phil. Trans. R. Soc. B 372, 20160456 (2017).
Jordan, C. Y. & Charlesworth, D. The potential for sexually antagonistic polymorphism in different genome regions. Evolution 66, 505–516 (2012).
Luthringer, R. et al. Sexual dimorphism in the brown algae. Perspect. Phycol. 1, 11–25 (2014).
Scharmann, M., Rebelo, A. G. & Pannell, J. R. High rates of evolution preceded shifts to sex-biased gene expression in Leucadendron, the most sexually dimorphic angiosperms. Elife 10, e67485 (2021).
Charlesworth, B. Model for evolution of Y chromosomes and dosage compensation. Proc. Natl Acad. Sci. USA 75, 5618–5622 (1978).
Lenormand, T., Fyon, F., Sun, E. & Roze, D. Sex chromosome degeneration by regulatory evolution. Curr. Biol. 30, 3001–3006.e5 (2020).
Jeffries, D. L., Gerchen, J. F., Scharmann, M. & Pannell, J. R. A neutral model for the loss of recombination on sex chromosomes. Phil. Trans. R. Soc. B 376, 20200096 (2021).
Jay, P., Tezenas, E., Véber, A. & Giraud, T. Sheltering of deleterious mutations explains the stepwise extension of recombination suppression on sex chromosomes and other supergenes. PLoS Biol. 20, e3001698 (2022).
Coelho, S. M., Gueno, J., Lipinska, A. P., Cock, J. M. & Umen, J. G. UV chromosomes and haploid sexual systems. Trends Plant Sci. 23, 794–807 (2018).
Peona, V. et al. Teaching transposon classification as a means to crowd source the curation of repeat annotation—a tardigrade perspective. Mob. DNA 15, 10 (2024).
Ivics, Z. & Izsvák, Z. The expanding universe of transposon technologies for gene and cell engineering. Mob. DNA 1, 25 (2010).
Gueno, J. et al. Chromatin landscape associated with sexual differentiation in a UV sex determination system. Nucleic Acids Res. 50, 3307–3322 (2022).
Bourdareau, S. et al. Histone modifications during the life cycle of the brown alga Ectocarpus. Genome Biol. 22, 12 (2021).
Makova, K. D. & Hardison, R. C. The effects of chromatin organization on variation in mutation rates in the genome. Nat. Rev. Genet. 16, 213–223 (2015).
Charlesworth, B. The effects of deleterious mutations on evolution at linked sites. Genetics 190, 5–22 (2012).
Lebherz, M. K., Fouks, B., Schmidt, J., Bornberg-Bauer, E. & Grandchamp, A. DNA transposons favor de novo transcript emergence through enrichment of transcription factor binding motifs. Genome Biol. Evol. https://doi.org/10.1093/gbe/evae134 (2024).
McDaniel, S. F. Divergent outcomes of genetic conflict on the UV sex chromosomes of Marchantia polymorpha and Ceratodon purpureus. Curr. Opin. Genet. Dev. 83, 102129 (2023).
Iwasaki, M. et al. Identification of the sex-determining factor in the liverwort Marchantia polymorpha reveals unique evolution of sex chromosomes in a haploid system. Curr. Biol. 31, 5522–5532.e7 (2021).
Ligrone, R., Duckett, J. G. & Renzaglia, K. S. The origin of the sporophyte shoot in land plants: a bryological perspective. Ann. Bot. 110, 935–941 (2012).
McDaniel, S. F., Neubig, K. M., Payton, A. C., Quatrano, R. S. & Cove, D. J. Recent gene-capture on the UV sex chromosomes of the moss Ceratodon purpureus. Evolution https://doi.org/10.1111/evo.12165 (2013).
Vigneau, J. et al. Interactions between U and V sex chromosomes during the life cycle of Ectocarpus. Development 151, dev202677 (2024).
Singh, S., Davies, K. M., Chagné, D. & Bowman, J. L. The fate of sex chromosomes during the evolution of monoicy from dioicy in liverworts. Curr. Biol. 33, 3597–3609.e3 (2023).
Takahashi, K. et al. Reorganization of the ancestral sex-determining regions during the evolution of trioecy in Pleodorina starrii. Commun. Biol. 6, 590 (2023).
Liesner, D. et al. Developmental pathways underlying sexual differentiation in the U/V sex chromosome system of giant kelp. Dev. Cell https://doi.org/10.1016/j.devcel.2024.12.022 (2025).
Yamamoto, K. et al. Three genomes in the algal genus Volvox reveal the fate of a haploid sex-determining region after a transition to homothallism. Proc. Natl Acad. Sci. USA 118, e2100712118 (2021).
Herpin, A. & Schartl, M. Plasticity of gene‐regulatory networks controlling sex determination: of masters, slaves, usual suspects, newcomers, and usurpators. EMBO Rep. 16, 1260–1274 (2015).
Starr, R. C. & Zeikus, J. A. UTEX—the culture collection of algae at the University of Texas at Austin 1993 list of cultures. J. Phycol. 29, 1–106 (1993).
Pearson, G., Lago-Leston, A., Valente, M. & Serrão, E. Simple and rapid RNA extraction from freeze-dried tissue of brown algae and seagrasses. Eur. J. Phycol. 41, 97–104 (2006).
Wick, R. R., Judd, L. M. & Holt, K. E. Performance of neural network basecalling tools for Oxford Nanopore sequencing. Genome Biol. 20, 129 (2019).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20, 257 (2019).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Huson, D. H., Auch, A. F., Qi, J. & Schuster, S. C. MEGAN analysis of metagenomic data. Genome Res. 17, 377–386 (2007).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, i351–i358 (2005).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Hoff, K. J., Lange, S., Lomsadze, A., Borodovsky, M. & Stanke, M. BRAKER1: unsupervised RNA-seq-based genome annotation with GeneMark-ET and AUGUSTUS. Bioinformatics 32, 767–769 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Shumate, A. & Salzberg, S. L. Liftoff: accurate mapping of gene annotations. Bioinformatics 37, 1639–1643 (2021).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl Acad. Sci. USA 117, 9451–9457 (2020).
Alonge, M. et al. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol. 23, 258 (2022).
Carvalho, A. B. & Clark, A. G. Efficient identification of Y chromosome sequences in the human and Drosophila genomes. Genome Res. 23, 1894–1907 (2013).
Vicoso, B., Emerson, J. J., Zektser, Y., Mahajan, S. & Bachtrog, D. Comparative sex chromosome genomics in snakes: differentiation, evolutionary strata, and lack of global dosage compensation. PLoS Biol. 11, e1001643 (2013).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962 (2016).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pedersen, B. S. & Quinlan, A. R. Mosdepth: quick coverage calculation for genomes and exomes. Bioinformatics 34, 867–868 (2018).
Avia, K. et al. High-density genetic map and identification of QTLs for responses to temperature and salinity stresses in the model brown alga Ectocarpus. Sci. Rep. 7, 43241 (2017).
Rochette, N. C., Rivera‐Colón, A. G. & Catchen, J. M. Stacks 2: analytical methods for paired‐end sequencing improve RADseq‐based population genomics. Mol. Ecol. 28, 4737–4754 (2019).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Rastas, P. Lep-MAP3: robust linkage mapping even for low-coverage whole genome sequencing data. Bioinformatics 33, 3726–3732 (2017).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://doi.org/10.48550/arXiv.1303.3997 (2013).
Nursyifa, C., Brüniche‐Olsen, A., Garcia‐Erill, G., Heller, R. & Albrechtsen, A. Joint identification of sex and sex‐linked scaffolds in non‐model organisms using low depth sequencing data. Mol. Ecol. Resour. 22, 458–467 (2022).
Malde, K., Skern, R. & Glover, K. A. Using sequencing coverage statistics to identify sex chromosomes in minke whales. Preprint at https://doi.org/10.48550/arXiv.1902.06654 (2019).
Smith, J. J. et al. The sea lamprey germline genome provides insights into programmed genome rearrangement and vertebrate evolution. Nat. Genet. 50, 270–277 (2018).
Darolti, I. et al. Extreme heterogeneity in sex chromosome differentiation and dosage compensation in livebearers. Proc. Natl Acad. Sci. USA 116, 19031–19036 (2019).
Brelsford, A., Lavanchy, G., Sermier, R., Rausch, A. & Perrin, N. Identifying homomorphic sex chromosomes from wild‐caught adults with limited genomic resources. Mol. Ecol. Resour. 17, 752–759 (2017).
Toups, M. A., Rodrigues, N., Perrin, N. & Kirkpatrick, M. A reciprocal translocation radically reshapes sex‐linked inheritance in the common frog. Mol. Ecol. 28, 1877–1889 (2019).
Tang, H. et al. Synteny and collinearity in plant genomes. Science 320, 486–488 (2008).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 34, 609–612 (2006).
Yang, Z. & Nielsen, R. Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 17, 32–43 (2000).
Wang, D., Zhang, Y., Zhang, Z., Zhu, J. & Yu, J. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genomics Proteomics Bioinformatics 8, 77–80 (2010).
Price, M. N., Dehal, P. S. & Arkin, A. P. Fasttree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 26, 1641–1650 (2009).
Larsson, A. AliView: a fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 30, 3276–3278 (2014).
Yang, Z. PAML 4: Phylogenetic Analysis by Maximum Likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Csűös, M. Count: evolutionary analysis of phylogenetic profiles with parsimony and likelihood. Bioinformatics 26, 1910–1912 (2010).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Chevenet, F., Brun, C., Bañuls, A.-L., Jacq, B. & Christen, R. TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics 7, 439 (2006).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (1995).
Lotharukpong, J. S. et al. A transcriptomic hourglass in brown algae. Nature 635, 129–135 (2024).
Schoch, C. L. et al. NCBI Taxonomy: a comprehensive update on curation, resources and tools. Database 2020, baaa062 (2020).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2021); https://www.r-project.org/
Gel, B. & Serra, E. karyoploteR: an R/Bioconductor package to plot customizable genomes displaying arbitrary data. Bioinformatics 33, 3088–3090 (2017).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res. 4, 1521 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bushmanova, E., Antipov, D., Lapidus, A. & Prjibelski, A. D. rnaSPAdes: a de novo transcriptome assembler and its application to RNA-seq data. Gigascience 8, giz100 (2019).
Smith-Unna, R., Boursnell, C., Patro, R., Hibberd, J. M. & Kelly, S. TransRate: reference-free quality assessment of de novo transcriptome assemblies. Genome Res. 26, 1134–1144 (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Leng, N. et al. EBSeq: an empirical Bayes hierarchical model for inference in RNA-seq experiments. Bioinformatics 29, 1035–1043 (2013).
Haas, F. B. Several brown algal chromosome level assemblies, V1. Edmond https://doi.org/10.17617/3.OOWB2Y (2025).
Cormier, A. et al. Re‐annotation, improved large‐scale assembly and establishment of a catalogue of noncoding loci for the genome of the model brown alga Ectocarpus. New Phytol. 214, 219–232 (2017).
Acknowledgements
This work was supported by the MPG, the CNRS, Sorbonne University, the ERC (grant no. 864038 and 638240 to S.M.C.), the France Génomique National infrastructure project Phaeoexplorer (ANR-10-INBS-09), the JSPS Overseas Research Fellowships (to M.H.), the BMBF-funded de.NBI Cloud within the German Network for Bioinformatics Infrastructure (de.NBI) (031A532B, 031A533A, 031A533B, 031A534A, 031A535A, 031A537A, 031A537B, 031A537C, 031A537D, 031A538A), the Investissements d’Avenir project Idealg (ANR-10-BTBR-04-01), the European BG-01 Blue Growth H2020 project Genialg (727892) and the ANR project Epicycle (ANR-19-CE20-0028-01). S.M.C. was supported by the Moore Foundation (GBMF11489) and the Bettencourt-Schueller Foundation. J.B.-R. was supported by a Humboldt Research Fellowship for postdoctoral researchers from the Alexander von Humboldt Foundation. We thank the members of the Phaeoexplorer consortium, in particular C. Jolivet, L. Mest and D. Scornet for assistance with algae cultures, C. Cruaud for help with sequencing libraries preparation, E. Corre and A. Le Bars for support with the Phaeoexplorer database and A. Couloux for the genome assemblies and annotations. We also thank the Roscoff Bioinformatics platform ABiMS (http://abims.sb-roscoff.fr), part of the Institut Français de Bioinformatique (ANR-11-INBS-0013) and Biogenouest network, for providing computing and storage resources.
Funding
Open access funding provided by Max Planck Society.
Author information
Authors and Affiliations
Contributions
J.B.-R. and A.P.L. conducted investigation (equal) and formal analysis (equal), designed the methodology (equal), performed visualization (equal), wrote the original draft (equal), and reviewed and edited the manuscript (equal). P.L. conducted investigation (supporting) and formal analysis (supporting). E.D., G.C., O.G., K.B., M.H., R.J.C., K.A., G.L., E.A., D.L., R.L., O.G., S.H., Z.N., L.B.-G. and A.F.P. conducted investigation (supporting). G.H., J.-M.A,. G.P., P.W., F.D. and J.M.C. performed data curation (supporting) and data acquisition (supporting). F.B.H. conducted investigation (supporting), designed the methodology (equal), and performed data curation (equal) and formal analysis (supporting). S.M.C. conceptualized the project (lead), acquired funding (lead), administered and supervised the roject (lead), designed the methodology (equal), performed visualization (supporting), wrote the original draft (equal), and reviewed and edited the manuscript (lead).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Macrosynteny plot between Schizocladia ischiensis and Dictyota dichotoma using 1,828 orthologs.
We highlight two fusion-with-mixing events (red squares) between chromosomes 4 and 9, and between chromosomes 23 and 24 in D. dichotoma.
Extended Data Fig. 2 SDR size differences and detection of independently-acquired V-SDR genes across species.
(a) Differences in the size of the male SDR between brown algal species based on the total sequence length, the relative size of the SDR compared to the length of the V chromosome and the number of protein-coding genes retained within the SDR. The bars are colored accord-ing to the level of gamete dimorphism in each species (based on 16). (b) Correlation between the V (blue) and U (pink) SDR sizes and the SDR gene content across species. (c) Correlation between the V (blue) and U (pink) SDR sizes and the SDR repeat content across species. (d) Gene trees showing the independent acquisition of SDR gametologues across species that were previously interpreted as part of the ancestral male SDR genes.
Extended Data Fig. 3 Male SDR synteny between Ectocarpus sp. 7 and Ectocarpus crouaniorum.
One of the species underwent a recent inversion event within the SDR. The arrows in the boxes represent the orientation of each gene within the chromosome.
Extended Data Fig. 4 Synteny analysis plot illustrating the expansion of the Desmarestia herbacea U- and V-sex-determining regions (SDRs) into the surrounding pseudoautosomal region (PAR).
The Ectocarpus sp. 7 V-chromosome is shown in the middle as a reference, with the SDR regions outlined by grey boxes. Green lines trace syntenic relationships between Ectocarpus PAR genes and the recently-acquired Desmarestia SDR genes, with each gene pair represented in a distinct shade of green. This demonstrates that nearly all PAR genes from Ectocarpus, which have entered the expanded SDR in Desmarestia, are retained as gametologues. Orange lines highlight the PAR boundary genes in Desmarestia, which remain within the PAR of Ectocarpus.
Extended Data Fig. 5 Sex-biased gene expression per dioicous species.
a) Proportion of sex biased genes in each of the five dioicous species. MBG: male-biased genes; FBG: female biased genes. (b) Number of sex-biased genes in the pseudoautosomal regions of sex chromosomes (U-V-SDRs excluded), male-biased genes are shown in blue and female-biased genes in red. Exact p-values above the bars mark significant enrichment of the sex-biased genes on the PAR (Chi-square test).
Extended Data Fig. 6 Expression of genes (log2(TPM + 1)) that entered the SDR independently in different species.
Expression is measured in mature male and female gametophytes, hashing marks missing orthologues, stars inside the cells indicate that the gene is inside the male non-recombining region (V-SDR). Orthogroups containing orthologues in less than three species or with multicopy genes were excluded from this analysis. M: male; F: female.
Extended Data Fig. 7 Structural features across the sex chromosomes and U/V-homologues of brown algae.
(a) V and U chromosomes of Ectocarpus sp. 7, Scytosiphon promiscuus, Undaria pinnatifida, Desmarestia herbacea and Dictyota dichotoma. (b) U/V-homologues of Desmarestia dudresnayi and Fucus serratus. Features displayed from bottom to top: chromosome compartments (PARs, SDR, U/V-homologue); relative gene ages, inter-species Ks values, and proportion of coding (CDS, red) and repeat (TEs, blue) density.
Extended Data Fig. 8 Accumulation of repetitive elements in the V-SDRs and PARs of five dioicous species.
Statistically significant differences in median values of repeat density (center line) were assessed using FDR-corrected permutation tests.
Extended Data Fig. 9 Proposed scenarios for the transition from dioicy to monoicy in Chordaria linearis and Desmarestia dudresnayi.
(a) The ancestor of Chordaria linearis likely underwent an initial translocation event from the U chromosome to the V chromosome, inserting part of the U-SDR and a piece of the 3′ PAR towards the 5′ end of the V-SDR potentially through an ectopic recombination event. A subsequent inversion within this translocation spread the 3′PAR genes to both sides of the U-SDR insertion. Finally, a second translocation led to the insertion of an additional piece of the U-SDR within the 3′ PAR translocation. (b) The ancestor of Desmarestia dudresnayi underwent three translocations of U-SDR genes into the V-SDR. Additionally, a fourth translocation event happened between the U-SDR and an autosome (chr_04).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–18.
Supplementary Tables 1–22 (download XLSX )
Supplementary Tables 1–22.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barrera-Redondo, J., Lipinska, A.P., Liu, P. et al. Origin and evolutionary trajectories of brown algal sex chromosomes. Nat Ecol Evol 9, 2127–2144 (2025). https://doi.org/10.1038/s41559-025-02838-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41559-025-02838-w
This article is cited by
-
Revisiting giant virus-host dynamics in brown algae: old stories and new perspectives
The EMBO Journal (2026)
-
Structural and evolutionary features of red algal UV sex chromosomes
Genome Biology (2025)