Abstract
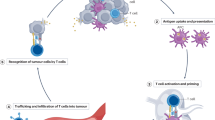
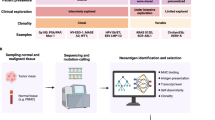
The efficacy of in situ cancer vaccination has been hampered by a poor spatiotemporal orchestration of multiple key steps of the cancer-immunity cycle in most tumours and systemic toxicity related to therapeutic strategies. Here we report a systemic injectable and pyroptosis-enabled nanoadjuvant that evokes the secretion of a vaccine-like pyroptosome in the tumour area for robust antitumour immunity. This systemic injectable and pyroptosis-enabled nanoadjuvant induces vigorous immunogenic pyroptosis, triggering the efficient release of tumour antigen-rich pyroptosomes, damage-associated molecular patterns and proinflammatory cytokines. Photoactivatable release of a TLR7/8 agonist into the pyroptosome generates an in situ cancer vaccine platform that cooperatively activates the cancer-immunity cycle and avoids systemic toxicity. The vaccine boosts both innate and adaptive immune responses, facilitating the eradication of primary tumours and metastatic nodules, thereby providing long-lasting cancer prevention. Overall, the rational design of pyroptosis-inducing nanomedicines for boosting the cancer-immunity cycle reported here might aid in the development of more efficient personalized cancer immunotherapy approaches.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data of this study are available within the Article and its Supplementary Information. The mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium (https://proteomecentral.proteomexchange.org/cgi/GetDataset) via the PRIDE partner repository with the dataset identifier PXD069142. Source data are provided with this paper.
References
Mellman, I., Coukos, G. & Dranoff, G. Cancer immunotherapy comes of age. Nature 480, 480–489 (2011).
Keskin, D. B. et al. Neoantigen vaccine generates intratumoral T cell responses in phase Ib glioblastoma trial. Nature 565, 234–239 (2019).
Lin, M. J. et al. Cancer vaccines: the next immunotherapy frontier. Nat. Cancer 3, 911–926 (2022).
Palmer, C. D. et al. Individualized, heterologous chimpanzee adenovirus and self-amplifying mRNA neoantigen vaccine for advanced metastatic solid tumors: phase 1 trial interim results. Nat. Med. 28, 1619–1629 (2022).
Sahin, U. et al. Personalized RNA mutanome vaccines mobilize poly-specific therapeutic immunity against cancer. Nature 547, 222–226 (2017).
Sahin, U. & Tureci, O. Personalized vaccines for cancer immunotherapy. Science 359, 1355–1360 (2018).
Blass, E. & Ott, P. A. Advances in the development of personalized neoantigen-based therapeutic cancer vaccines. Nat. Rev. Clin. Oncol. 18, 215–229 (2021).
Gong, N. Q. et al. Enhancing in situ cancer vaccines using delivery technologies. Nat. Rev. Drug Discov. 23, 607–625 (2024).
Jhunjhunwala, S., Hammer, C. & Delamarre, L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat. Rev. Cancer 21, 298–312 (2021).
Chen, L. et al. Bacterial cytoplasmic membranes synergistically enhance the antitumor activity of autologous cancer vaccines. Sci. Transl. Med. 13, eabc2816 (2021).
Xu, J. et al. A general strategy towards personalized nanovaccines based on fluoropolymers for post-surgical cancer immunotherapy. Nat. Nanotechnol. 15, 1043–1052 (2020).
Wang, C. et al. Oncolytic mineralized bacteria as potent locally administered immunotherapeutics. Nat. Biomed. Eng. 8, 561–578 (2024).
Zhang, Y. B. et al. Close the cancer-immunity cycle by integrating lipid nanoparticle-mRNA formulations and dendritic cell therapy. Nat. Nanotechnol. 18, 1364–1374 (2023).
Galluzzi, L., Buque, A., Kepp, O., Zitvogel, L. & Kroemer, G. Immunogenic cell death in cancer and infectious disease. Nat. Rev. Immunol. 17, 97–111 (2017).
Aaes, T. L. & Vandenabeele, P. The intrinsic immunogenic properties of cancer cell lines, immunogenic cell death, and how these influence host antitumor immune responses. Cell Death Differ. 28, 843–860 (2021).
Bedoui, S., Herold, M. J. & Strasser, A. Emerging connectivity of programmed cell death pathways and its physiological implications. Nat. Rev. Mol. Cell Biol. 21, 678–695 (2020).
Meier, P. et al. Immunogenic cell death in cancer: targeting necroptosis to induce antitumour immunity. Nat. Rev. Cancer 24, 299–315 (2024).
Chen, B. et al. A pyroptosis nanotuner for cancer therapy. Nat. Nanotechnol. 17, 788–798 (2022).
Chen, B., Zhang, Q., Gao, J. & Wang, Y. Control cell death by endosomal trafficking of proton transistor nanophotosensitizers. Life Med. 1, 264–266 (2022).
Nel, A. E. et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat. Mater. 8, 543–557 (2009).
Zhao, R. et al. A pH-responsive nanoparticle library with precise pH tunability by co-polymerization with non-ionizable monomers. Angew. Chem. Int. Ed. 61, e202200152 (2022).
Wang, Y. et al. Digitization of endocytic pH by hybrid ultra-pH-sensitive nanoprobes at single-organelle resolution. Adv. Mater. 29, 1603794 (2017).
Wang, Z. et al. pH-amplified CRET nanoparticles for in vivo imaging of tumor metastatic lymph nodes. Angew. Chem. Int. Ed. 60, 14512–14520 (2021).
Tang, M. et al. pH-gated nanoparticles selectively regulate lysosomal function of tumour-associated macrophages for cancer immunotherapy. Nat. Commun. 14, 5888 (2023).
Akishiba, M. et al. Cytosolic antibody delivery by lipid-sensitive endosomolytic peptide. Nat. Chem. 9, 751–761 (2017).
Teo, W. et al. Nile Red fluorescence spectroscopy reports early physicochemical changes in myelin with high sensitivity. Proc. Natl Acad. Sci. USA 118, e2016897118 (2021).
Fuchs, Y. & Steller, H. Live to die another way: modes of programmed cell death and the signals emanating from dying cells. Nat. Rev. Mol. Cell Biol. 16, 329–344 (2015).
Han, X. et al. Adjuvant lipidoid-substituted lipid nanoparticles augment the immunogenicity of SARS-CoV-2 mRNA vaccines. Nat. Nanotechnol. 18, 1105–1114 (2023).
Adamik, J. & Butterfield, L. H. What’s next for cancer vaccines. Sci. Transl. Med. 14, eabo4632 (2022).
Guo, Y. et al. In situ generation of micrometer-sized tumor cell-derived vesicles as autologous cancer vaccines for boosting systemic immune responses. Nat. Commun. 13, 6534 (2022).
Chao, Y. et al. Combined local immunostimulatory radioisotope therapy and systemic immune checkpoint blockade imparts potent antitumour responses. Nat. Biomed. Eng. 2, 611–621 (2018).
Li, W. et al. Targeting photodynamic and photothermal therapy to the endoplasmic reticulum enhances immunogenic cancer cell death. Nat. Commun. 10, 3349 (2019).
Shekarian, T. et al. Repurposing rotavirus vaccines for intratumoral immunotherapy can overcome resistance to immune checkpoint blockade. Sci. Transl. Med. 11, eaat5025 (2019).
Xia, H. et al. A pH-/enzyme-responsive nanoparticle selectively targets endosomal toll-like receptors to potentiate robust cancer vaccination. Nano Lett. 22, 2978–2987 (2022).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (NSFC) grants (82225044 and 82530110 to Y.W.; 82522084, 82373804 and 82450112 to B.C.), Fundamental and Interdisciplinary Disciplines Breakthrough Plan of the Ministry of Education of China (JYB2025XDXM609 to Y.W.), National Key Research and Development Program of China (2024YFB3814600 to B.C.), Beijing Natural Science Foundation (Z240028 to Y.W.) and Beijing Nova Program (20250484828 to B.C.). We would like to thank the State Key Laboratory of Natural and Biomimetic Drugs, Peking University Biological Imaging and Flow Cytometry Core Facilities, for flow cytometry, confocal, animal and tissue imaging services.
Author information
Authors and Affiliations
Contributions
B.C. and Y.W. are responsible for all phases of the research. F.W. and L.W. helped with the fabrication and characterization of the EE-stressed nanophotosensitizer. X.P., J.L. and Y.Y. helped with the synthesis and characterization of the copolymers. H.X., Y.W. and M.T. participated in the evaluation of the immune response. Y.Y. and M.Q. helped with the in vivo antitumour studies. J.R. assisted in the western blot. B.Z. helped with the in vivo imaging. W.C. performed the high-performance liquid chromatography analysis. L.Z. conducted the shotgun proteomics analysis. C.H. and M.D. helped with the autoimmune response analysis. Q.Z. provided conceptual suggestions. Y.W. and B.C. provided constructive guidance for the overall design of the project. B.C. wrote the manuscript. Y.W. and B.C. revised the paper. All authors discussed the results and assisted in the preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Materials thanks Victor Bosteels and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Nanophotosensitizers containing EPA monomers exhibit superior membrane affinity and early endosome stress-mediated cytotoxicity.
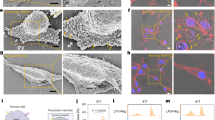
(a) Cell membrane binding capacity of nanophotosensitizers with various EPA monomers on A549 cells at pH 6.0. Fluorescence signal was measured by flow cytometry and normalized to EPA-0 group. (b) Oxidative stress-induced LDH release of nanophotosensitizers with various EPA monomers on A549 cells after treatment of membrane binding and 660 nm irradiation (6 J cm−2). (c) Quantification of erythrocytes hemolysis induced by the nanophotosensitizer-mediated oxidative stress when binding with membrane at pH 7.4 or pH 6.0. (d) Confocal images of cellular transport and endocytic organelle colocalization analysis of PEPA and PDPA at 0.5 h and 2 h after internalization. Scale bar = 5 μm. (e, f) The membrane-binding of PEPA and PDPA (3 μg mL−1 PPa, equivalent to 40 μg mL−1 polymer with negligible physical membrane disruption upon protonation and in the absence of irradiation) evokes erythrocytes hemolysis under 660 nm irradiation. (g) Interaction of Nile red-labelled copolymers with artificial lipid membrane at pH 6.0 measured by fluorimeter. (h) Photocytotoxicity of PEPAEE and PDPAEE nanophotosensitizer on A549 cells. (i) Quantification of EE stress-evoked lipid peroxidation detected with oxidation-sensitive sensor C11-BODIPY581/591 probe. (j) Cytosolic calcium release elicited by the EE stress of PEPA or PDPA. All data are presented as mean ± s.d. (n = 3 biologically independent experiments). (k) Intracellular caspase-3 activity of A549 cells measured by flow cytometry with Fluorometric Assay Kit. (l) The cell death morphology and PI staining of A549-GFP cells treated with PEPA or PDPA-mediated EE stress. The elevated red signal of nuclear PI and loss of cellular GFP indicate cell death accompanied by plasma membrane rupture. Evident cell swelling and giant bubbles from the plasma membrane along with quickly enhanced permeability indicated by PI staining are typical characteristics of pyroptosis. Scale bar = 10 μm. Statistical significance was analyzed by two-sided Student’s t-test.
Extended Data Fig. 2 Early endosome stress of PEPA nanophotosensitizer evokes GSDME-mediated pyroptosis in murine tumor cells.
(a) Confocal images of intracellular singlet oxygen generation measured by SOSG nanoprobe (nSOSG, green) and colocalization with PEPA and endocytic organelles at 0.5 h (Rab5a, early endosome marker, blue) and 2 h (Lamp1, lysosome marker, blue) post-internalization. The green signal of nSOSG indicates singlet oxygen generated by PEPA nanophotosensitizer after 660 nm irradiation. Scale bar = 10 μm. (b) Representative phase-contrast images of cell death morphology induced by PEPAEE or PEPALy on CT26 and CT26-GSDME−/− cells. Scale bar = 20 μm. (c) MTT assay and LDH release assay of PEPAEE stress on CT26 and CT26-GSDME−/− cells. GSDME knockout rescued EE stress-induced pyroptosis. Data are presented as mean ± s.d. (n = 3 biologically independent experiments). Statistical significance was analyzed by two-sided Student’s t-test. (d) Immunoblots of endogenous GSDME expression in different murine cancer cell lines. (e) Heatmap of GSDME expression quantified from panel d. (f) Photocytotoxicity of PEPAEE or PEPALy on different murine cancer cell lines. Data are presented as mean ± s.d. (n = 3 biologically independent experiments). (g) Phase-contrast imaging of cell death morphology on different murine cancer cells treated with PEPA6.5-mediated EE stress. Scale bar = 20 μm.
Extended Data Fig. 3 EE stress-evoked pyroptosis enhances the ICD effect of cancer cells in vitro and in vivo.
(a, b) PEPA-mediated EE stress facilitates intracellular protein release on CT26 cells as evaluated by BCA assay (a) and SDS-PAGE (b). n = 3 biologically independent experiments. (c) Confocal images of annexin V-FITC/PI and HMGB1 staining of CT26 cells at 24 h post-treatment with PEPAEE or PEPALy. Scale bar = 20 μm. (d) ATP release level, and (e, f) HSP70/CRT exposure of CT26 cells at 24 h post-treatment with PEPAEE or PEPALy. n = 3 biologically independent experiments. (g-j) BMDC activation after co-culture with PEPAEE or PEPALy treated CT26-OVA cells. n = 3 biologically independent experiments. Representative contour plots of CD80+CD86+ cells among CD11c+ BMDC (g). Percentage of CD80+CD86+ BMDC (h). Percentage of BMDCs with presentation of OVA antigen (i). Secretion of IL-12 and TNF-α measured by ELISA assay (j). (k) Representative results of specific cell killing study in CT26-OVA tumour-bearing mice after different treatments. (l) Average tumour growth curves and (m) individual tumour growth kinetics of CT26-OVA tumour-bearing mice treated with PEPAEE or PEPALy. n = 5 biologically independent mice. All data are shown as mean ± s.d. Statistical significance was analyzed by one-way ANOVA followed by Dunnett’s multiple comparisons test.
Extended Data Fig. 4 SPEN exhibits safe and effective TLR activation.
(a) Chemical structure of the pH and light-gated polymer-agonist conjugate and schematic of light-triggered TLR activation. (b) 660 nm laser-induced release profiles of SPEN and SPIN quantified by HPLC (n = 3 biologically independent experiments). (c) TLR agonistic activity of various IMDQ treatments with or without 660 nm irradiation via a TLR reporter cell assay (n = 6 biologically independent experiments). (d-g) BMDC activation after treatment of free IMDQ, SPEN, or SPIN with or without 660 nm irradiation. n = 3 biologically independent experiments. Representative contour plots of CD80+CD86+ cells among CD11c+ BMDC (d). Percentage of CD80+CD86+ BMDC (e). Secretion of IL-12 (f) and TNF-α (g) measured by ELISA assay. (h-m) In vivo biosafety and systemic immunotoxicity of SPEN (n = 5 biologically independent mice). Mice were administrated with free IMDQ or SPEN via tail vein injection, and the body weight, cytokines in blood, and spleens were evaluated (h). Mice body weight change after different treatments (i). Photographs (j) and weight (k) of spleens excised at day 8. The profiles of IL-12 (l) and IP-10 (m) level in blood. All data are shown as mean ± s.d. Statistical significance was analyzed by one-way ANOVA followed by Dunnett’s multiple comparisons test.
Extended Data Fig. 5 In vitro and in vivo cancer vaccination capacity of EE stress-derived pyroptosomes.
(a) Morphology of EE stress-derived pyroptosome characterized by TEM. Scale bar = 500 nm. (b) Morphology of Au nanoparticle-encapsulated SPEN characterized by TEM. Scale bar = 20 nm. (c) Quantification of IMDQ in the cell membrane or pyroptosomes derived from CT26-OVA cells (n = 3 biologically independent experiments). (d) Enrichment of GSDME-N, calreticulin (CRT), and ovalbumin (OVA) in the cell membrane or pyroptosomes derived from CT26-OVA (normalized to Na-K-ATPase, n = 3 biologically independent experiments). (e-i) BMDC activation after treatment with pyroptosomes elicited by EE stress of various nanoadjuvants (n = 3 biologically independent experiments). Quantitative expression of CD86 (e) and CD80 (f) on BMDCs. Secretion of TNF-α (g) and IL-12 (h) measured by ELISA assay. Presentation of OVA antigen (i). (j-l) In vivo lymph node (LN) draining of DiR-labelled pyroptosomes (n = 6 biologically independent experiments). Quantitative DiR signal profiles in popliteal LNs (j). Ex vivo fluorescence imaging (k) and quantification (l) of excised LNs. (m-p) Activation of LNs in mice immunized with pyroptosomes (n = 5 biologically independent mice). Photographs of excised LNs (m). Presentation of OVA antigen on DCs in LNs (n). The cell counts of total lymphocytes and T cells in LNs (o). The cell counts of CD8+ T cells and CD4+ T cells in LNs (p). (q) Tumour growth profiles of CT26 tumour-bearing mice immunized with pyroptosomes from 0, 2, 4, 10 or 20 × 104 pyroptotic cells, n = 3 biologically independent mice. (r) Photographs and (s) numbers of spontaneous tumours on MMTV-PyMT/FVB mice immunized with personalized pyroptotic cancer vaccine (n = 10 biologically independent mice). All data are shown as mean ± s.d. For c,l,s, statistical significance was analyzed by two-sided Student’s t-test with Welch’s correction. For d-I and n-p, statistical significance was analyzed by one-way ANOVA followed by Dunnett’s multiple comparisons test.
Extended Data Fig. 6 Tumour immune microenvironment priming of SPEN-mediated EE stress.
(a) Representative flow cytometry plots of IFN-γ+ cytotoxic T cells among CD8+ T cells in immunized mice (n = 5 biologically independent mice). (b-f) The percentage of NK cells (b), CD8+ T cells (c), M1-type macrophages (d), M2-type macrophages (e) and Foxp3+ T cells (f) in tumour tissues quantified from multiplex immunohistochemistry staining of tumour slides. The box represents the interquartile range, with the median represented by the centre line. The whiskers indicate the range of data, extending to the maximum and minimum values (n = 28 regions for Saline group, n = 26 regions for SPEN group, n = 27 regions for PEPAEE, n = 17 regions for SPENEE group, biologically independent). (g) Representative flow cytometry plots of IFN-γ+ NK cells and granzyme B+ NK cells among NKp46+ cells in immunized mice. (h) The percentage of IFN-γ+ cytotoxic NK cells among NKp46+ cells in immunized mice (n = 5 biologically independent mice). (i) Representative confocal images of DQ-ovalbumin (green) degradation assays in BMDMs pretreated with various pyroptosomes. Scale bar = 20 μm. (j) Presentation of OVA antigen on cell membrane of BMDM after treatment with various pyroptosomes (n = 3 biologically independent experiment). (k) Representative flow cytometry plots of NK activation by various pyroptosomes as indicated by membrane expression of NKp46. (l) Quantification of CD107a+ cells among NKp46+ NK cells (n = 3 biologically independent experiment). All data are shown as mean ± s.d. Statistical significance was analyzed by one-way ANOVA followed by Dunnett’s multiple comparisons test.
Extended Data Fig. 7 Combination therapy of SPEN-mediated EE stress and αPD-1.
(a-f) Combination immunotherapy of SPEN with αPD-1 on the immunosuppressive PANC02 tumour-bearing mice (n = 11 mice for saline and αPD-1 groups; n = 12 mice for SPENEE and SPENEE + αPD-1 groups). Schematic illustration of experiment schedule (a). Growth profiles of the primary PANC02 tumour with various treatments (b; two-way ANOVA followed by Bonferroni’s multiple comparisons test). Individual tumour growth curves (c), representative photographs of tumour-bearing mice at day 60 (d), animal survival curves (e; log-rank test; P = 0.0004 for αPD-1 versus SPENEE, P = 0.0477 for SPENEE versus SPENEE + αPD-1), and tumour relapse curves after rechallenge (f, log-rank test; P = 0.0131 for αPD-1 versus SPENEE, P = 0.0003 for SPENEE versus SPENEE + αPD-1). (g-j) Abscopal therapeutic efficacy of combination therapy in an established metastatic CT26-luc tumour model with multiple peritoneal tumour nodules. Schematic illustration of experiment schedule (g). The photographs of bioluminescence imaging (h), metastatic tumour nodules (i), and bioluminescence quantification (j) of excised gastrointestinal tissues. Two-sided Student’s t-test with Welch’s correction. n = 6 biologically independent mice. (k-n) Abscopal therapeutic efficacy of combination therapy in an established lung metastatic 4T1-luc tumour model. Lung metastatic tumour growth profiles quantified by in vivo bioluminescence imaging (k). The bioluminescent photographs (l), quantification (m, n = 5 mice for Saline group and n = 6 mice for SPENEE + αPD-1 group, biologically independent), and H&E staining (n) of excised lungs. Two-sided Student’s t-test with Welch’s correction. All data are shown as mean ± s.d.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–29, Tables 1 and 2, methods, discussion and references.
Supplementary Video 1 (download MOV )
Rapid membrane ballooning and pyroptosis triggered by PEPA-mediated EE stress on CT26-GFP cells. CT26-GFP cells were treated with PEPA (1 μg ml−1 of PPa) for 0.5 h, and immediately irradiated with a 660-nm laser (6 J cm−2). Representative field recorded immediately after irradiation is shown (time duration, h:min:s:ms).
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data blots (download JPG )
Unprocessed western blots.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, B., Wan, F., Xia, H. et al. In situ-generated vaccine-like pyroptosome for personalized cancer immunotherapy. Nat. Mater. (2026). https://doi.org/10.1038/s41563-026-02506-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41563-026-02506-9