Abstract
Alterations in gut microbiota have been linked to chronic kidney disease (CKD), but large-scale studies and mechanistic insights are limited. Here we analysed gut metagenome data from 1,550 older individuals (aged 65–93 years) with comprehensive kidney function measurements. Segatella copri was positively associated with kidney function through microbial ammonia metabolism-related pathways and the asnA gene, which encodes an ammonia-assimilating enzyme. These associations were replicated in two external studies. In mice, ammonia supplementation increased serum levels of creatinine and blood urea nitrogen, accelerating CKD progression. In vitro cultures of S. copri or asnA-overexpressing Escherichia coli reduced ammonia concentrations, which was markedly attenuated in asnA-knockout S. copri. Gavage of either S. copri or asnA-overexpressing E. coli, but not asnA-knockout S. copri, mitigated ammonia-induced CKD progression in mice. These findings highlight the role of gut microbial ammonia metabolism in CKD pathogenesis and underscore the therapeutic potential of microbial-based interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Metagenomic data of the current study have been deposited to the Genome Sequence Archive under the accession codes PRJCA039229 (RLAS cohort) and PRJCA040203 (HS study). All other data supporting the findings of this study are provided as source data files accompanying this paper. The UniProt database is accessible at https://www.uniprot.org/. Source data are provided with this paper.
Code availability
Codes associated with the data analysis and visualization are available at https://github.com/LinscStef/CKDGutMicrobiome (ref. 73).
References
GBD Chronic Kidney Disease Collaboration. Global, regional, and national burden of chronic kidney disease, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet 395, 709–733 (2020).
Kalantar-Zadeh, K., Jafar, T. H., Nitsch, D., Neuen, B. L. & Perkovic, V. Chronic kidney disease. Lancet 398, 786–802 (2021).
Eckardt, K. U. et al. Evolving importance of kidney disease: from subspecialty to global health burden. Lancet 382, 158–169 (2013).
Yang, T., Richards, E. M., Pepine, C. J. & Raizada, M. K. The gut microbiota and the brain–gut–kidney axis in hypertension and chronic kidney disease. Nat. Rev. Nephrol. 14, 442–456 (2018).
Meijers, B., Evenepoel, P. & Anders, H. J. Intestinal microbiome and fitness in kidney disease. Nat. Rev. Nephrol. 15, 531–545 (2019).
Wang, X. et al. Aberrant gut microbiota alters host metabolome and impacts renal failure in humans and rodents. Gut 69, 2131–2142 (2020).
Wu, I. W. et al. Integrative metagenomic and metabolomic analyses reveal severity-specific signatures of gut microbiota in chronic kidney disease. Theranostics 10, 5398–5411 (2020).
Wang, H. et al. Perturbed gut microbiome and fecal and serum metabolomes are associated with chronic kidney disease severity. Microbiome 11, 3 (2023).
Sato, N. et al. Metagenomic profiling of gut microbiome in early chronic kidney disease. Nephrol. Dial. Transplant. 36, 1675–1684 (2021).
Rysz, J. et al. The impact of CKD on uremic toxins and gut microbiota. Toxins 13, 252 (2021).
Krukowski, H. et al. Gut microbiome studies in CKD: opportunities, pitfalls and therapeutic potential. Nat. Rev. Nephrol. 19, 87–101 (2023).
Weiner, I. D., Mitch, W. E. & Sands, J. M. Urea and ammonia metabolism and the control of renal nitrogen excretion. Clin. J. Am. Soc. Nephrol. 10, 1444–1458 (2015).
Davila, A. M. et al. Intestinal luminal nitrogen metabolism: role of the gut microbiota and consequences for the host. Pharmacol. Res. 68, 95–107 (2013).
Leigh, J. A. & Dodsworth, J. A. Nitrogen regulation in bacteria and archaea. Annu. Rev. Microbiol. 61, 349–377 (2007).
Carrero, J. J., Hecking, M., Chesnaye, N. C. & Jager, K. J. Sex and gender disparities in the epidemiology and outcomes of chronic kidney disease. Nat. Rev. Nephrol. 14, 151–164 (2018).
Stengel, B. Chronic kidney disease and cancer: a troubling connection. J. Nephrol. 23, 253–262 (2010).
Fabrizi, F. et al. Decreased serum aminotransferase activity in patients with chronic renal failure: impact on the detection of viral hepatitis. Am. J. Kidney Dis. 38, 1009–1015 (2001).
Louis, P. & Flint, H. J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 19, 29–41 (2017).
Li, W. et al. Ecological and network analyses identify four microbial species with potential significance for the diagnosis/treatment of ulcerative colitis (UC). BMC Microbiol. 21, 138 (2021).
Nath, K. A., Hostetter, M. K. & Hostetter, T. H. Pathophysiology of chronic tubulo-interstitial disease in rats. Interactions of dietary acid load, ammonia, and complement component C3. J. Clin. Invest. 76, 667–675 (1985).
Tolins, J. P., Hostetter, M. K. & Hostetter, T. H. Hypokalemic nephropathy in the rat. Role of ammonia in chronic tubular injury. J. Clin. Invest. 79, 1447–1458 (1987).
Vince, A. J. & Burridge, S. M. Ammonia production by intestinal bacteria: the effects of lactose, lactulose and glucose. J. Med. Microbiol. 13, 177–191 (1980).
Richardson, A. J., McKain, N. & Wallace, R. J. Ammonia production by human faecal bacteria, and the enumeration, isolation and characterization of bacteria capable of growth on peptides and amino acids. BMC Microbiol. 13, 6 (2013).
Chen, Z. et al. The role of intestinal bacteria and gut–brain axis in hepatic encephalopathy. Front. Cell. Infect. Microbiol. 10, 595759 (2020).
Lau, W. L., Kalantar-Zadeh, K. & Vaziri, N. D. The gut as a source of inflammation in chronic kidney disease. Nephron 130, 92–98 (2015).
Bartochowski, P. et al. Gut–kidney axis investigations in animal models of chronic kidney disease. Toxins 14, 626 (2022).
Diwan, V., Brown, L. & Gobe, G. C. Adenine-induced chronic kidney disease in rats. Nephrology 23, 5–11 (2018).
Lun, H. et al. Altered gut microbiota and microbial biomarkers associated with chronic kidney disease. MicrobiologyOpen 8, e00678 (2019).
Yang, R. et al. Coprococcus eutactus, a potent probiotic, alleviates colitis via acetate-mediated IgA response and microbiota restoration. J. Agric. Food Chem. https://doi.org/10.1021/acs.jafc.2c06697 (2023).
Fan, Y. et al. The gut microbiota contributes to the pathogenesis of anorexia nervosa in humans and mice. Nat. Microbiol. 8, 787–802 (2023).
Chaves, L. D. et al. Chronic kidney disease, uremic milieu, and its effects on gut bacterial microbiota dysbiosis. Am. J. Physiol. Renal Physiol. 315, F487–F502 (2018).
Andrianova, N. V. et al. Microbiome–metabolome signature of acute kidney injury. Metabolites 10, 142 (2020).
Verbrugghe, P. et al. Evaluation of hypoglycemic effect, safety and immunomodulation of Prevotella copri in mice. Sci. Rep. 11, 21279 (2021).
Abdelsalam, N. A., Hegazy, S. M. & Aziz, R. K. The curious case of Prevotella copri. Gut Microbes 15, 2249152 (2023).
Pedersen, H. K. et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 535, 376–381 (2016).
Scher, J. U. et al. Expansion of intestinal Prevotella copri correlates with enhanced susceptibility to arthritis. eLife 2, e01202 (2013).
Wen, C. et al. Quantitative metagenomics reveals unique gut microbiome biomarkers in ankylosing spondylitis. Genome Biol. 18, 142 (2017).
De Vadder, F. et al. Microbiota-produced succinate improves glucose homeostasis via intestinal gluconeogenesis. Cell Metab. 24, 151–157 (2016).
Metwaly, A. & Haller, D. Strain-level diversity in the gut: the P. copri case. Cell Host Microbe 25, 349–350 (2019).
De Filippis, F. et al. Distinct genetic and functional traits of human intestinal Prevotella copri strains are associated with different habitual diets. Cell Host Microbe 25, 444–453.e3 (2019).
Tett, A. et al. The Prevotella copri complex comprises four distinct clades underrepresented in westernized populations. Cell Host Microbe 26, 666–679.e7 (2019).
Ramezani, A. et al. Role of the gut microbiome in uremia: a potential therapeutic target. Am. J. Kidney Dis. 67, 483–498 (2016).
Hinchman, S. K., Henikoff, S. & Schuster, S. M. A relationship between asparagine synthetase A and aspartyl tRNA synthetase. J. Biol. Chem. 267, 144–149 (1992).
Chan, M. J. et al. Breath ammonia is a useful biomarker predicting kidney function in chronic kidney disease patients. Biomedicines 8, 468 (2020).
Vallet, M. et al. Urinary ammonia and long-term outcomes in chronic kidney disease. Kidney Int. 88, 137–145 (2015).
Andrade, S. L. & Einsle, O. The Amt/Mep/Rh family of ammonium transport proteins. Mol. Membr. Biol. 24, 357–365 (2007).
van Heeswijk, W. C., Westerhoff, H. V. & Boogerd, F. C. Nitrogen assimilation in Escherichia coli: putting molecular data into a systems perspective. Microbiol. Mol. Biol. Rev. 77, 628–695 (2013).
Joshi, S., McMacken, M. & Kalantar-Zadeh, K. Plant-based diets for kidney disease: a guide for clinicians. Am. J. Kidney Dis. 77, 287–296 (2021).
Kalantar-Zadeh, K. et al. Plant-dominant low-protein diet for conservative management of chronic kidney disease. Nutrients 12, 1931 (2020).
Precup, G. & Vodnar, D. C. Gut Prevotella as a possible biomarker of diet and its eubiotic versus dysbiotic roles: a comprehensive literature review. Br. J. Nutr. 122, 131–140 (2019).
Goldberg, M. R. et al. Microbial signature in IgE-mediated food allergies. Genome Med. 12, 92 (2020).
Trefflich, I., Dietrich, S., Braune, A., Abraham, K. & Weikert, C. Short- and branched-chain fatty acids as fecal markers for microbiota activity in vegans and omnivores. Nutrients 13, 1808 (2021).
Su, G. et al. Fiber intake and health in people with chronic kidney disease. Clin. Kidney J. 15, 213–225 (2022).
Chou, Y. H. & Chen, Y. M. Aging and renal disease: old questions for new challenges. Aging Dis. 12, 515–528 (2021).
Nishijima, S. et al. Fecal microbial load is a major determinant of gut microbiome variation and a confounder for disease associations. Cell 188, 222–236.e15 (2025).
Liu, Z. et al. Cohort profile: the Rugao Longevity and Ageing Study (RuLAS). Int. J. Epidemiol. 45, 1064–1073 (2016).
Guan, H. et al. Comparison of fecal collection methods on variation in gut metagenomics and untargeted metabolomics. mSphere 6, e0063621 (2021).
Inker, L. A. et al. Estimating glomerular filtration rate from serum creatinine and cystatin C. N. Engl. J. Med. 367, 20–29 (2012).
van Anken, H. C. & Schiphorst, M. E. A kinetic determination of ammonia in plasma. Clin. Chim. Acta 56, 151–157 (1974).
Wang, Y. et al. Sample collection, DNA extraction, and library construction protocols of the human microbiome studies in the International Human Phenome Project. Phenomics 3, 300–308 (2023).
Pu, Y. et al. Impact of DNA extraction methods on gut microbiome profiles: a comparative metagenomic study. Phenomics 5, 76–90 (2025).
Sun, Z. et al. Gut microbiome alterations and gut barrier dysfunction are associated with host immune homeostasis in COVID-19 patients. BMC Med. 20, 24 (2022).
Blanco-Miguez, A. et al. Extending and improving metagenomic taxonomic profiling with uncharacterized species using MetaPhlAn 4. Nat. Biotechnol. 41, 1633–1644 (2023).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with bioBakery 3. eLife 10, e65088 (2021).
Holmes, I., Harris, K. & Quince, C. Dirichlet multinomial mixtures: generative models for microbial metagenomics. PLoS ONE 7, e30126 (2012).
Ali, B. H. et al. New model for adenine-induced chronic renal failure in mice, and the effect of gum acacia treatment thereon: comparison with rats. J. Pharmacol. Toxicol. Methods 68, 384–393 (2013).
Li, J. et al. A versatile genetic toolbox for Prevotella copri enables studying polysaccharide utilization systems. EMBO J. 40, e108287 (2021).
Wang, P. et al. Development of an efficient conjugation-based genetic manipulation system for Pseudoalteromonas. Microb. Cell Fact. 14, 11 (2015).
Chen, B. et al. Gut bacteria alleviate smoking-related NASH by degrading gut nicotine. Nature 610, 562–568 (2022).
Li, X. et al. Ketogenic diet-induced bile acids protect against obesity through reduced calorie absorption. Nat. Metab. 6, 1397–1414 (2024).
Wu, L. L. et al. Transforming growth factor beta 1 and renal injury following subtotal nephrectomy in the rat: role of the renin-angiotensin system. Kidney Int. 51, 1553–1567 (1997).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442 (2021).
Lin, S. CKDGutMicrobiome. GitHub https://github.com/LinscStef/CKDGutMicrobiome (2025).
Acknowledgements
We thank the participants for their participation in this research and the research staff in the Rugao Longitudinal Ageing Study. This work was jointly supported by the National Key R&D Program of China (2021YFA1301000, 2020YFC2005000 and 2021YFC2500202 to Y.Z. and J.C.) and the Shanghai Municipal Science and Technology Major Project (Grant No. 2023SHZDZX02 to Y.Z.). The computations in this research were performed using the CFFF platform and the Human Phenome Data Center at Fudan University, and we thank the centre staff for their support.
Author information
Authors and Affiliations
Contributions
Y.Z., X.L., and S.L. conceived of and designed the study. Y.Z., J.N., X.W. and J.C. were involved in the funding acquisition. X.W., Y.Z. and X.S. designed the RLAS cohort. S.L., Z.S., H.Z. and X.W. recruited the study participants and collected faecal and blood samples. S.L., M.W., Q.Z., J.Q. and J.C. recruited the HS study participants and collected faecal and blood samples. S.L., Z.S., M.K. and X. Zhou prepared the samples for sequencing and analysed the data. X. Zhu, X. Zhang and X.L. performed the in vitro and in vivo experiments. S.L., Z.S. and X.L. drafted the paper. Z.M., Y.P., P.G., J.L., L.M., F.Z. and S.H. contributed to the revision of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Andrew Holmes, Brandilyn Peters-Samuelson, Till Strowig and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The variation in kidney function explained by host characteristics.
The bars represented the proportion of variation in eGFR (a) and impaired kidney function (b) explained by host characteristics and gut microbiome. The R2 was calculated using a univariate linear regression, with each explanatory variable analyzed separately in individual models. P values were not adjusted for multiple comparisons. All statistical tests were two-sided. The eGFR was calculated by using the 2012 Chronic Kidney Disease Epidemiology Collaboration equation based on creatinine and cystatin C. Impaired kidney function was defined as eGFR less than 60 mL/min/1.73 m2. Participants with impaired kidney function, n = 328; Participants with normal kidney function, n = 1,222. eGFR, estimated glomerular filtration rate. BMI, body mass index. Medication Use, whether people have had at least five types of drugs. TG, triglyceride. HDL-c, high-density lipoprotein cholesterol. LDL-c, low-density lipoprotein cholesterol. PCo, principal coordinates.
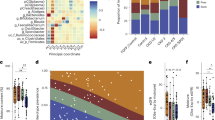
Extended Data Fig. 2 Associations of host characteristics with the gut microbial composition.
(a–d) Species-level and (e–h) pathway-level alpha diversity across participants with different kidney function status. P values for group differences were determined using multivariate linear regression, with adjustments for age, sex, BMI, diabetes, hypertension, cancer, medication use, HDL-c, and LDL-c. The boxes represented the median and interquartile range (IQR) of the distributions, and top and bottom whiskers marked the point at 1.5×IQR. Participants of impaired kidney function, n = 328; Participants of normal kidney function, n = 1,222; Participants of eGFR ≥ 90 mL/min/1.73 m2, n = 187; Participants of eGFR ≥ 60 & < 90 mL/min/1.73 m2, n = 1035; Participants of eGFR < 60 mL/min/1.73 m2, n = 328. (i) The microbial composition among participants with impaired kidney function or normal kidney function at the pathway-level. Group difference was calculated by permutational multivariate analysis of variance (PERMANOVA) using Bray-Curtis dissimilarity. Participants with impaired kidney function, n = 328; Participants with normal kidney function, n = 1,222. (j–l) Proportion of the variance in microbiome taxonomic composition explained by each measured kidney function parameters and other host characteristics. (m–p) Proportion of the variance in microbiome functional composition explained by each measured kidney function parameters and other host characteristics. The explained variance (R2) for (j–p) was calculated by PERMANOVA using Bray-Curtis dissimilarity, and all variables were jointly fitted in one model for each panel. ns, not significant (P > 0.05). eGFR, estimated glomerular filtration rate. BMI, body mass index. Medication Use, whether people have had at least five types of drugs. HDL-c, high-density lipoprotein cholesterol. LDL-c, low-density lipoprotein cholesterol. P values were not adjusted for multiple comparisons. All statistical tests were two-sided.
Extended Data Fig. 3 Association of Phocaeicola vulgatus with kidney function.
The data represented the AST-transformed relative abundance of P. vulgatus across participants with different kidney disease severity in the RLAS cohort, HS study, and published dataset. P values for the group difference were tested by MaAsLin2 (two-sided test) with the adjustment of age, sex, body mass index, diabetes, hypertension, high-density lipoprotein cholesterol, and low-density lipoprotein cholesterol in the HS study, and further adjusted for cancer and medication use in the RLAS cohort. The Benjamini-Hochberg method was used to calculate FDR-adjusted P values only in the RLAS cohort. The boxes represented the median and interquartile range (IQR) of the distributions, and top and bottom whiskers marked the point at 1.5×IQR. Participants of eGFR ≥ 90 mL/min/1.73 m2, n = 187; Participants of eGFR ≥ 60 & < 90 mL/min/1.73 m2, n = 1,035; Participants of eGFR < 60 mL/min/1.73 m2, n = 328. Participants in HS study: control, n = 15; CKD, n = 47; KF, n = 52. Participants in the published dataset: control, n = 69; KF, n = 223. ns, not significant (P > 0.05). eGFR, estimated glomerular filtration rate; CKD, chronic kidney disease; KF, kidney failure.
Extended Data Fig. 4 Associations of plasma ammonia with eGFR (a) and the relative abundance of S. copri (b) in HS study.
The associations were estimated by multivariable linear regression (a) and MaAsLin2 (b), with adjustment for age, sex, body mass index, diabetes, hypertension, high-density lipoprotein cholesterol, low-density lipoprotein cholesterol, and abnormal liver function. All statistical tests were two-sided. Abnormal liver function was defined as any of the following: alanine aminotransferase > 40 U/L, aspartate aminotransferase > 35 U/L, or total bilirubin > 21 μmol/l, or direct Bilirubin > 8 μmol/l, or alkaline phosphatase > 100 U/L, or gamma-glutamyl transferase > 45 U/L. Participants with plasma ammonia data: control, n = 15; CKD, n = 39; KF = 50. eGFR, estimated glomerular filtration rate; CKD, chronic kidney disease; KF, kidney failure.
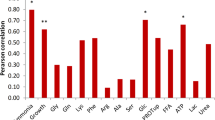
Extended Data Fig. 5 Correlation between microbial species and functional features.
Data represented the correlations of 7 pathways and 6 enzymes related to ammonia production (x-axis) with 16 microbial species related to kidney function and the enzyme/gene related to ammonia assimilation (y-axis) in the RLAS cohort. We included the 15 replicated species associated with eGFR, and the species that showed the strongest inverse association with eGFR, that is, P. vulgatus. Color and numbers indicated the Spearman’s correlation coefficient. *P < 0.05, **P < 0.01, ***P < 0.001. P values were not adjusted for multiple comparisons. All statistical tests were two-sided.
Extended Data Fig. 6 Ammonia accumulation exacerbated kidney disease severity.
(a) Experimental design. ACD-fed SPF mice were treated with NaCl or NH4Cl water for 4 weeks. The diagram was created using BioRender. (b, c) Daily water intake (b) and food intake (c) per mouse basis in NaCl and NH4Cl water treated groups (n = 6 replicates/treatment). (d) Body weight of ACD-fed mice after 4 weeks of NaCl or NH4Cl water treatment. (e–g) Plasma ammonia (e), serum creatinine (f), and BUN (g) levels in CKD mice after 4 weeks of NaCl or NH4Cl water treatment. (h–l) Representative images (h) and quantification (i–l) of kidney sections from mice in (a), from top to bottom: PAS staining (400×, red arrow indicates glomerular atrophy and black arrow indicates tubule dilation) and quantification of the glomerulosclerotic index in renal glomeruli (i); Masson’s trichrome staining (400×) and quantification of the proportion of the fibrotic area in renal cortex (j); Immunohistochemistry of TGF-β1 (200×) and quantification of the anti-inflammatory cytokine TGF-β1 positive area in renal cortex (k); Immunohistochemistry for 8-OHdG (200×) and quantification of the DNA damage marker 8-OHdG positive area in renal cortex (l). n = 6 mice per group. Data are presented as the mean ± s.e.m. ns, not significant (P > 0.05). *P < 0.05, **P < 0.01, ***P < 0.001 determined by two-tailed Student’s t-test (b-g, and i-l). ACD, adenine-containing diet; SPF, specific-pathogen-free; CKD, chronic kidney disease; BUN, blood urea nitrogen; PAS, Periodic Acid-Schiff; TGF-β1, transforming growth factor-β1; 8-OHdG, 8-hydroxy-2′-deoxyguanosine.
Extended Data Fig. 7 S. copri transplantation delayed kidney damage.
(a–c) The relative abundance of S. copri (a), asnA gene (b), and P. vulgatus (c) in fecal samples from mice gavaged with S. copri, P. vulgatus, or vehicle, determined by qPCR. (d, e) Daily water intake (d) and food intake (e) per mouse basis in vehicle or bacteria strain gavaged groups (n = 6 replicates/treatment). (f) Body weight of CKD mice after 4 weeks gavage-feeding vehicle or bacteria strain. (g, h) Representative immunohistochemistry for TGF-β1 (200×, g) and 8-OHdG (200×, h) staining of kidney sections from S. copri, P. vulgatus, or vehicle gavaged CKD mice. (i) Quantification of the anti-inflammatory cytokine TGF-β1 positive area in renal cortex based on TGF-β1 immunohistochemical sections. (j) Quantification of the DNA damage marker 8-OHdG positive area in renal cortex based on 8-OHdG immunohistochemical sections. n = 6 mice per group. Data are presented as the mean ± s.e.m. ns, not significant (P > 0.05). *P < 0.05, ***P < 0.001 determined by one-way ANOVA with Tukey’s post-hoc test (a-f and i-j). CKD, chronic kidney disease; TGF-β1, transforming growth factor-β1; 8-OHdG, 8-hydroxy-2′-deoxyguanosine.
Extended Data Fig. 8 Knocking out of asnA in S. copri abolished its protection against ammonia-induced kidney damage.
(a) Schematic representation of the workflow for knocking out the asnA gene in S. copri by allelic exchange via a suicide plasmid (left). The workflow includes the construction of donor E. coli carrying the suicide plasmid, the conjugation between donor E. coli and recipient S. copri, homologous recombination, and screening and identification of asnA-knockout S. copri. The diagram was created using BioRender. The picture shows the screening of erythromycin-sensitive colonies using blood plates, and the electropherogram shows the confirmation of the asnA gene deletion by PCR validation (right). (b, c) The relative abundance of S. copri (b) and asnA gene (c) in fecal samples from mice gavaged with vehicle, WT S. copri, or asnA-KO S. copri, determined by qPCR. (d, e) Daily water intake (d) and food intake (e) per mouse basis in vehicle, WT S. copri, or asnA-KO S. copri gavaged groups (n = 6 replicates/treatment). (f) Body weight of CKD mice after four weeks gavage-feeding vehicle, WT S. copri, or asnA-KO S. copri. (g, h) Representative immunohistochemistry for TGF-β1 (200×, g) and 8-OHdG (200×, h) staining of kidney sections from vehicle, WT S. copri, or asnA-KO S. copri gavaged CKD mice. (i) Quantification of the anti-inflammatory cytokine TGF-β1 positive area in renal cortex based on TGF-β1 immunohistochemical sections. (j) Quantification of the DNA damage marker 8-OHdG positive area in renal cortex based on 8-OHdG immunohistochemical sections. n = 6 mice per group. Data are presented as the mean ± s.e.m. ns, not significant (P > 0.05). *P < 0.05, **P < 0.01, ***P < 0.001 determined by one-way ANOVA with Tukey’s post-hoc test (b-f and i-j). asnA-KO S. copri, asnA-knockout S. copri; WT S. copri, wild type S. copri; CKD, chronic kidney disease; TGF-β1, transforming growth factor-β1; 8-OHdG, 8-hydroxy-2′-deoxyguanosine.
Extended Data Fig. 9 The transplantation of asnA-overexpressing E. coli delayed kidney damage.
(a) Schematic diagram illustrating the experimental workflow for the introduction of the asnA gene into E. coli, and the transfection of the control E. coli with an unloaded plasmid (left). The diagram was created using BioRender. Western blot analysis depicting the expression of His-tagged asnA protein in the engineered E. coli strains (right), indicating successful protein expression at specific molecular weights. (b) The relative abundance of the asnA gene in fecal samples from mice gavaged with vehicle or engineered E. coli, determined by qPCR. (c, d) Daily water intake (c) and food intake (d) per mouse basis in vehicle or engineered E. coli gavaged groups (n = 6 replicates/treatment). (e) Body weight of CKD mice after 4 weeks gavage-feeding vehicle or engineered E. coli. (f–g) Representative immunohistochemistry for TGF-β1 (200×, f) and 8-OHdG (200×, g) staining of kidney sections from vehicle, asnA-OE E. coli, or control E. coli gavaged CKD mice. (h) Quantification of the anti-inflammatory cytokine TGF-β1 positive area in renal cortex based on TGF-β1 immunohistochemical sections. (i) Quantification of the DNA damage marker 8-OHdG positive area in renal cortex based on 8-OHdG immunohistochemical sections. n = 6 mice per group. Data are presented as the mean ± s.e.m. ns, not significant (P > 0.05). *P < 0.05, **P < 0.01, ***P < 0.001 determined by one-way ANOVA with Tukey’s post-hoc test (b-e and h-i). asnA-OE E. coli, asnA-overexpressing E. coli; CKD, chronic kidney disease; TGF-β1, transforming growth factor-β1; 8-OHdG, 8-hydroxy-2′-deoxyguanosine.
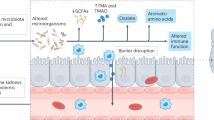
Extended Data Fig. 10 Graphical abstract of the study workflow and findings.
Workflow was created with Biorender.com.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1 and 2.
Supplementary Tables 1–9 (download XLSX )
Supplementary Tables 1–9.
Supplementary Data (download XLSX )
Source data for Supplementary Fig. 1.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data of Extended Data Fig. 8b–j.
Source Data Extended Data Figs. 8 and 9 (download PDF )
Unprocessed gels of Extended Data Fig. 8a and unprocessed western blots of Extended Data Fig. 9a.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data of Extended Data Fig. 9b–i.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, S., Sun, Z., Zhu, X. et al. Segatella copri and gut microbial ammonia metabolism contribute to chronic kidney disease pathogenesis. Nat Microbiol 10, 1684–1697 (2025). https://doi.org/10.1038/s41564-025-02039-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02039-y