Abstract
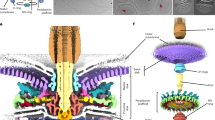
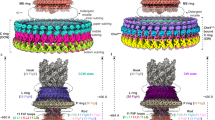
Knowledge of bacterial flagella has largely come from studies of the simpler motors of Escherichia coli and Salmonella enterica. However, many bacteria harbour more complex motors. The function, mechanisms and evolution associated with such auxiliary motor structures are unclear. Here we deploy structural, genetic, biochemical and functional approaches to characterize complex adaptations of the flagellar motor in Campylobacter jejuni. We observed an E ring formed by 17 FlgY homodimers around the MS ring, a cage-like structure made of FcpMNO and PflD, and PflA–PflB interactions in a spoke–rim formation between the E ring and cage. These scaffolds stabilized the 17 torque-generating stator complexes. Phylogenetic analyses suggest an ancient origin and widespread prevalence of the E ring and spokes across diverse flagellated bacteria, and co-option of type IV pilus components in the ancestral motor of phylum Campylobacterota. Collectively, these data provide insight into the assembly, function and evolution of complex flagellar motors.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data are provided in the Article and Supplementary Information. The cryo-ET maps of the motor in WT Campylobacter jejuni and ΔflgY, ΔpflA, ΔpflB, ΔpflC, ΔpflD and ΔfcpMNO mutant cells have been deposited in the Electron Microscopy Data Bank under accession codes EMD-45507, EMD-49254, EMD-49256, EMD-49255, EMD-49325, EMD-49253 and EMD-49252, respectively. The cryo-EM map and atomic coordinates have been deposited in the PDB and the Electron Microscopy Data Bank under accession numbers 9LEQ and EMD-63032, respectively. Further details are provided in Supplementary Tables 7 and 9. Raw reads of RNA-seq studies have been deposited in the Sequence Read Archive (https://www.ncbi.nlm.nih.gov/sra) under BioProject number PRJNA1223640. The transcriptome datasets generated in this study are summarized in Supplementary Table 2. The Campylobacter jejuni 81-176 reference genome (NCBI accession number: GCA_000015525.1) is available for download from NCBI (https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF/000/015/525/GCF_000015525.1_ASM1552v1/). Source data are provided with this paper.
References
Chen, S. et al. Structural diversity of bacterial flagellar motors. EMBO J. 30, 2972–2981 (2011).
Kaplan, M. et al. The presence and absence of periplasmic rings in bacterial flagellar motors correlates with stator type. eLife https://doi.org/10.7554/eLife.43487 (2019).
Zhu, S. et al. In situ structures of polar and lateral flagella revealed by cryo-electron tomography. J. Bacteriol. https://doi.org/10.1128/JB.00117-19 (2019).
Rossmann, F. M. & Beeby, M. Insights into the evolution of bacterial flagellar motors from high-throughput in situ electron cryotomography and subtomogram averaging. Acta Crystallogr. D 74, 585–594 (2018).
Armitage, J. P. & Berry, R. M. Assembly and dynamics of the bacterial flagellum. Annu. Rev. Microbiol. 74, 181–200 (2020).
Hu, H. et al. Structural basis of torque generation in the bi-directional bacterial flagellar motor. Trends Biochem. Sci. 47, 160–172 (2022).
Chang, Y., Xu, H., Motaleb, M. A. & Liu, J. Characterization of the flagellar collar reveals structural plasticity essential for spirochete motility. mBio 12, e0249421 (2021).
Qin, Z., Lin, W. T., Zhu, S., Franco, A. T. & Liu, J. Imaging the motility and chemotaxis machineries in Helicobacter pylori by cryo-electron tomography. J. Bacteriol. 199, e00695-16 (2017).
Lertsethtakarn, P., Ottemann, K. M. & Hendrixson, D. R. Motility and chemotaxis in Campylobacter and Helicobacter. Annu. Rev. Microbiol. 65, 389–410 (2011).
Gao, B., Lara-Tejero, M., Lefebre, M., Goodman, A. L. & Galan, J. E. Novel components of the flagellar system in epsilonproteobacteria. mBio 5, e01349-14 (2014).
Hendrixson, D. R. & DiRita, V. J. Identification of Campylobacter jejuni genes involved in commensal colonization of the chick gastrointestinal tract. Mol. Microbiol. 52, 471–484 (2004).
Beeby, M. et al. Diverse high-torque bacterial flagellar motors assemble wider stator rings using a conserved protein scaffold. Proc. Natl Acad. Sci. USA 113, E1917–E1926 (2016).
Drobnic, T. et al. In situ structure of a bacterial flagellar motor at subnanometre resolution reveals adaptations for increased torque. Nat. Microbiol. 10, 1723–1740 (2025).
Cohen, E. J. et al. Evolution of a large periplasmic disk in Campylobacterota flagella enables both efficient motility and autoagglutination. Dev. Cell 59, 3306–3321.e5 (2024).
Gao, B. et al. Metabolic and fitness determinants for in vitro growth and intestinal colonization of the bacterial pathogen Campylobacter jejuni. PLoS Biol. 15, e2001390 (2017).
Chen, Y. et al. Tetrameric PilZ protein stabilizes stator ring in complex flagellar motor and is required for motility in Campylobacter jejuni. Proc. Natl Acad. Sci. USA 122, e2412594121 (2025).
Johnson, R. C., Walsh, M. P., Ely, B. & Shapiro, L. Flagellar hook and basal complex of Caulobacter crescentus. J. Bacteriol. 138, 984–989 (1979).
Stallmeyer, M. J., Hahnenberger, K. M., Sosinsky, G. E., Shapiro, L. & DeRosier, D. J. Image reconstruction of the flagellar basal body of Caulobacter crescentus. J. Mol. Biol. 205, 511–518 (1989).
Liu, X. et al. Bacterial flagella hijack type IV pili proteins to control motility. Proc. Natl Acad. Sci. USA 121, e2317452121 (2024).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold3. Nature https://doi.org/10.1038/s41586-024-07487-w (2024).
Hattori, M., Tanaka, Y., Fukai, S., Ishitani, R. & Nureki, O. Crystal structure of the MgtE Mg2+ transporter. Nature 448, 1072–1075 (2007).
Lynch, M. J. et al. Co-folding of a FliF–FliG split domain forms the basis of the MS:C ring interface within the bacterial flagellar motor. Structure 25, 317–328 (2017).
Xue, C. et al. Crystal structure of the FliF–FliG complex from Helicobacter pylori yields insight into the assembly of the motor MS-C ring in the bacterial flagellum. J. Biol. Chem. 293, 2066–2078 (2018).
Baker, M. A. et al. Domain-swap polymerization drives the self-assembly of the bacterial flagellar motor. Nat. Struct. Mol. Biol. 23, 197–203 (2016).
Singh, P. K. et al. CryoEM structures reveal how the bacterial flagellum rotates and switches direction. Nat. Microbiol. https://doi.org/10.1038/s41564-024-01674-1 (2024).
Rocaboy, M. et al. The crystal structure of the cell division amidase AmiC reveals the fold of the AMIN domain, a new peptidoglycan binding domain. Mol. Microbiol. 90, 267–277 (2013).
Guo, S. et al. PilY1 regulates the dynamic architecture of the type IV pilus machine in Pseudomonas aeruginosa. Nat. Commun. 15, 9382 (2024).
Chevance, F. F. & Hughes, K. T. Coordinating assembly of a bacterial macromolecular machine. Nat. Rev. Microbiol. 6, 455–465 (2008).
Zhao, X. et al. Cryoelectron tomography reveals the sequential assembly of bacterial flagella in Borrelia burgdorferi. Proc. Natl Acad. Sci. USA 110, 14390–14395 (2013).
Mastronarde, D. N. & Held, S. R. Automated tilt series alignment and tomographic reconstruction in IMOD. J. Struct. Biol. 197, 102–113 (2017).
Jagannathan, A., Constantinidou, C. & Penn, C. W. Roles of rpoN, fliA, and flgR in expression of flagella in Campylobacter jejuni. J. Bacteriol. 183, 2937–2942 (2001).
Kaplan, M. et al. Novel transient cytoplasmic rings stabilize assembling bacterial flagellar motors. EMBO J. 41, e109523 (2022).
Mo, R. et al. The evolutionary path of chemosensory and flagellar macromolecular machines in Campylobacterota. PLoS Genet. 18, e1010316 (2022).
Zeytuni, N. & Zarivach, R. Structural and functional discussion of the tetra-trico-peptide repeat, a protein interaction module. Structure 20, 397–405 (2012).
Eggenhofer, E., Haslbeck, M. & Scharf, B. MotE serves as a new chaperone specific for the periplasmic motility protein, MotC, in Sinorhizobium meliloti. Mol. Microbiol. 52, 701–712 (2004).
Velez-Gonzalez, F. et al. Rotation of the Fla2 flagella of Cereibacter sphaeroides requires the periplasmic proteins MotK and MotE that interact with the flagellar stator protein MotB2. PLoS ONE 19, e0298028 (2024).
Moon, K. H. et al. Spirochetes flagellar collar protein FlbB has astounding effects in orientation of periplasmic flagella, bacterial shape, motility, and assembly of motors in Borrelia burgdorferi. Mol. Microbiol. 102, 336–348 (2016).
Moon, K. H., Zhao, X., Xu, H., Liu, J. & Motaleb, M. A. A tetratricopeptide repeat domain protein has profound effects on assembly of periplasmic flagella, morphology and motility of the lyme disease spirochete Borrelia burgdorferi. Mol. Microbiol. 110, 634–647 (2018).
Tassinari, M., Rudzite, M., Filloux, A. & Low, H. H. Assembly mechanism of a Tad secretion system secretin–pilotin complex. Nat. Commun. 14, 5643 (2023).
Roujeinikova, A. Crystal structure of the cell wall anchor domain of MotB, a stator component of the bacterial flagellar motor: implications for peptidoglycan recognition. Proc. Natl Acad. Sci. USA 105, 10348–10353 (2008).
Tachiyama, S. et al. FlgY, PflA, and PflB form a spoke-ring network in the high-torque flagellar motor of Helicobacter pylori. Proc. Natl Acad. Sci. USA 122, e2421632122 (2025).
Botting, J. M. et al. FlbB forms a distinctive ring essential for periplasmic flagellar assembly and motility in Borrelia burgdorferi. PLoS Pathog. 21, e1012812 (2025).
Pallen, M. J. & Matzke, N. J. From The Origin of Species to the origin of bacterial flagella. Nat. Rev. Microbiol. 4, 784–790 (2006).
Egelman, E. H. Reducing irreducible complexity: divergence of quaternary structure and function in macromolecular assemblies. Curr. Opin. Cell Biol. 22, 68–74 (2010).
Jernigan, K. K. & Bordenstein, S. R. Tandem-repeat protein domains across the tree of life. PeerJ 3, e732 (2015).
Arrias, P. N. et al. Diversity and structural–functional insights of alpha-solenoid proteins. Protein Sci. 33, e5189 (2024).
Ferreira, J. L. et al. The “Jack-of-all-trades” flagellum from Salmonella and E. coli was horizontally acquired from an ancestral beta-proteobacterium. Front. Microbiol. 12, 643180 (2021).
Gould, S. J. & Vrba, E. S. Exaptation—a missing term in the science of form. Paleobiology 8, 4–15 (1982).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Tachiyama, S. et al. The flagellar motor protein FliL forms a scaffold of circumferentially positioned rings required for stator activation. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2118401119 (2022).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Xu, A. & Xu, C. FastTomo: a SerialEM script for collecting electron tomography data. Preprint at bioRxiv https://doi.org/10.1101/2021.03.16.435675 (2021).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Xiong, Q., Morphew, M. K., Schwartz, C. L., Hoenger, A. H. & Mastronarde, D. N. CTF determination and correction for low dose tomographic tilt series. J. Struct. Biol. 168, 378–387 (2009).
Agulleiro, J. I. & Fernandez, J. J. Tomo3D 2.0—exploitation of advanced vector extensions (AVX) for 3D reconstruction. J. Struct. Biol. 189, 147–152 (2015).
Morado, D. R., Hu, B. & Liu, J. Using Tomoauto: a protocol for high-throughput automated cryo-electron tomography. J. Vis. Exp. https://doi.org/10.3791/53608 (2016).
Winkler, H. et al. Tomographic subvolume alignment and subvolume classification applied to myosin V and SIV envelope spikes. J. Struct. Biol. 165, 64–77 (2009).
Winkler, H. 3D reconstruction and processing of volumetric data in cryo-electron tomography. J. Struct. Biol. 157, 126–137 (2007).
Santiveri, M. et al. Structure and function of stator units of the bacterial flagellar motor. Cell 183, 244–257.e16 (2020).
Guo, S., Xu, H., Chang, Y., Motaleb, M. A. & Liu, J. FliL ring enhances the function of periplasmic flagella. Proc. Natl Acad. Sci. USA 119, e2117245119 (2022).
Tan, J. et al. Structural basis of assembly and torque transmission of the bacterial flagellar motor. Cell 184, 2665–2679.e19 (2021).
Johnson, S. et al. Structural basis of directional switching by the bacterial flagellum. Nat. Microbiol. https://doi.org/10.1038/s41564-024-01630-z (2024).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Wang, N. et al. Structural basis of human monocarboxylate transporter 1 inhibition by anti-cancer drug candidates. Cell 184, 370–383.e13 (2021).
Xu, K., Wang, Z., Shi, J., Li, H. & Zhang, Q. C. A2-Net: Molecular structure estimation from cryo-EM density volumes. In Proc. AAAI Conference on Artificial Intelligence Vol. 33, 1230–1237 (AAAI Press, 2019).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Battesti, A. & Bouveret, E. The bacterial two-hybrid system based on adenylate cyclase reconstitution in Escherichia coli. Methods 58, 325–334 (2012).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 25, 402–408 (2001).
Gabler, F. et al. Protein sequence analysis using the MPI Bioinformatics Toolkit. Curr. Protoc. Bioinformatics 72, e108 (2020).
Abby, S. S., Denise, R. & Rocha, E. P. C. Identification of protein secretion systems in bacterial genomes using MacSyFinder version 2. Methods Mol. Biol. 2715, 1–25 (2024).
Letunic, I., Khedkar, S. & Bork, P. SMART: recent updates, new developments and status in 2020. Nucleic Acids Res. 49, D458–D460 (2021).
Denise, R., Abby, S. S. & Rocha, E. P. C. Diversification of the type IV filament superfamily into machines for adhesion, protein secretion, DNA uptake, and motility. PLoS Biol. 17, e3000390 (2019).
Na, S. I. et al. UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J. Microbiol. https://doi.org/10.1007/s12275-018-8014-6 (2018).
Galperin, M. Y. et al. COG database update 2024. Nucleic Acids Res. 53, D356–D363 (2025).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 421 (2009).
Chaumeil, P. A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Johnson, L. S., Eddy, S. R. & Portugaly, E. Hidden Markov model speed heuristic and iterative HMM search procedure. BMC Bioinform. 11, 431 (2010).
Xu, H., He, J., Liu, J. & Motaleb, M. A. BB0326 is responsible for the formation of periplasmic flagellar collar and assembly of the stator complex in Borrelia burgdorferi. Mol. Microbiol. 113, 418–429 (2020).
Karpenahalli, M. R., Lupas, A. N. & Soding, J. TPRpred: a tool for prediction of TPR-, PPR- and SEL1-like repeats from protein sequences. BMC Bioinform. 8, 2 (2007).
D’Andrea, L. D. & Regan, L. TPR proteins: the versatile helix. Trends Biochem. Sci. 28, 655–662 (2003).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 40, 1023–1025 (2022).
Moller, S., Croning, M. D. & Apweiler, R. Evaluation of methods for the prediction of membrane spanning regions. Bioinformatics 17, 646–653 (2001).
Raddi, G. et al. Three-dimensional structures of pathogenic and saprophytic Leptospira species revealed by cryo-electron tomography. J. Bacteriol. 194, 1299–1306 (2012).
Liu, J. et al. Intact flagellar motor of Borrelia burgdorferi revealed by cryo-electron tomography: evidence for stator ring curvature and rotor/C-ring assembly flexion. J. Bacteriol. 191, 5026–5036 (2009).
Nothaft, H., Liu, X., McNally, D. J. & Szymanski, C. M. N-linked protein glycosylation in a bacterial system. Methods Mol. Biol. 600, 227–243 (2010).
Liu, J. et al. Cellular architecture of Treponema pallidum: novel flagellum, periplasmic cone, and cell envelope as revealed by cryo electron tomography. J. Mol. Biol. 403, 546–561 (2010).
Chaban, B., Coleman, I. & Beeby, M. Evolution of higher torque in Campylobacter-type bacterial flagellar motors. Sci. Rep. 8, 97 (2018).
Rossmann, F. M., Hug, I., Sangermani, M., Jenal, U. & Beeby, M. In situ structure of the Caulobacter crescentus flagellar motor and visualization of binding of a CheY-homolog. Mol. Microbiol. 114, 443–453 (2020).
Carroll, B. L. et al. The flagellar motor of Vibrio alginolyticus undergoes major structural remodeling during rotational switching. eLife https://doi.org/10.7554/eLife.61446 (2020).
Ferreira, J. L. et al. γ-proteobacteria eject their polar flagella under nutrient depletion, retaining flagellar motor relic structures. PLoS Biol. 17, e3000165 (2019).
Zhu, S. et al. Molecular architecture of the sheathed polar flagellum in Vibrio alginolyticus. Proc. Natl Acad. Sci. USA 114, 10966–10971 (2017).
Acknowledgements
We thank B. Crane and M. Lynch of Cornell University for discussions regarding the ARM-like domain; the High-Performance Computing Division of the South China Sea Institute of Oceanology for data analysis; J. Aronson (Yale University) for editing and valuable comments on the paper; R. Kumar for suggestions on model building; the Yale Center for Research Computing for guidance and use of the research computing infrastructure; and X. Li, J. Zhu and Z. Li of Shandong University Core Facilities for Life and Environmental Sciences for their help with the cryo-EM and MST experiments. This study was supported by the National Natural Science Foundation of China (32470031 and 32370189), the National Key Research and Development Program of China (2022YFC3102003), the Science and Technology Planning Project of Guangdong Province of China (2021B1212050023) and the Innovation Academy of South China Sea Ecology and Environmental Engineering, Chinese Academy of Sciences (numbers ISEE2021ZD03 and ISEE2021PY05). J.H. and X.G. were supported by the Shandong Provincial Natural Science Foundation (ZR2024ZD47), SKLMT Frontiers and Challenges Project (SKLMTFCP-2023-01) and Young Talent Development Program of SKLMT (M2025YA01). S.T., J.M.B. and J.L. were supported by grants R01AI087946 and R01AI132818 from the National Institute of Allergy and Infectious Diseases; cryo-ET data were collected at Yale CryoEM Resource, which was funded in part by National Institutes of Health grant 1S10OD023603-01A1. H.Z. was supported by funds from the State Key Laboratory of Crop Stress Adaptation and Improvement of Henan University.
Author information
Authors and Affiliations
Contributions
B.G. and J.L. conceived and designed the study. Genetic, protein interaction, RNA-seq and partial biochemical experiments were performed by X.F. with assistance from Y.L., Y.C. and M.A.B.B. Cryo-ET experiments and related structural analyses were performed by S.T. with assistance from H.Z. and C.H., and modelling was performed by J.M.B. and S.T. Single-particle cryo-EM experiments of PflAB and biochemical analyses of FlgY were performed by J.H. under the supervision of X.G. Phylogenetic analysis and homologue searches were performed by S.Z. with assistance from Y.L. Partial mutants were provided by C.H. and M.L.-T., and LC–MS/MS experiments were performed by M.L.-T. The paper was written by B.G., J.L., S.T. and X.F. All authors edited the paper and support the conclusions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Alejandro Buschiazzo, Daniel Kearns and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Sequence and structural analyses of FlgY and FcpMNO.
a, SMART motif analysis of FlgY (WP_002855458.1) suggested a single peptide (SP) at its N-terminus and a MgtE_N domain at C-terminus. b, Sequence alignment of FcpM (WP_002868795.1), FcpN (WP_002868796.1), and FcpO (WP_002859017.1) to their homologs in H. pylori (WP_000911905.1, WP_001212813.1, WP_001212813.1). c, Comparison of AlphaFold3-predicted structures of FcpM, FcpN, and FcpO from Campylobacter jejuni and H. pylori, with the structures of PilM, PilN, and PilO from Myxococcus xanthus (PDB:3JC8). Sequences for structural prediction are the same as in b.
Extended Data Fig. 2 Stator occupancy and motility of Campylobacter jejuni wild type and mutants.
a, Cross-section of motor structure in stator ring region from wild type and ΔfcpMNO mutant to compare stator occupancy with cage (wild type) and without cage (ΔfcpMNO mutant). b-d, The averaged structure of the motor in ΔpflC mutant before 3D classification (ΔpflCPre). The central section (b), cross section (c) of the spoke-rim, and cross section of stator ring in the cytoplasmic side (d). e-g, The averaged structure of the motor in ΔpflC mutant after 3D classification of (ΔpflCPost). The central section (e), cross section (f) of the spoke-rim, and cross section of stator ring in the cytoplasmic side (g).
Extended Data Fig. 3 Swimming velocity of Campylobacter jejuni wild type and ΔflgY mutant in media with different viscosities.
Swimming velocity of wild type or ΔflgY mutant populations after growth for 24 h in BHI broth alone or BHI broth with increasing concentrations of methylcellulose 4000 (a–c), methylcellulose 400 (d–f), or Ficoll 400 (g–i). Velocities of >100 individual cells per condition were tracked, and data from three independent assays were combined. Exact sample numbers for each panel are as follows: panel a, n = 112–222; panel b, n = 148–226; panel d, n = 112–241; panel e, n = 147–244; panel g, n = 101–183; panel h, n = 100–166. a–b, d–e, g–h show distributions of swimming velocities with mean values indicated. Differences for each treatment relative to the control (BHI) were calculated by Kruskal–Wallis test with Dunn’s post hoc multiple comparisons. c, f, i, Violin plots summarizing the data from panels a and b (c), d and e (f), and g and h (i). Red bars indicate the median; grey dash bars indicate the 25th and 75th percentiles. Comparisons between wild type (grey) and ΔflgY mutant (white) at each viscosity were also calculated by Kruskal–Wallis test with Dunn’s post hoc multiple comparisons.
Extended Data Fig. 4 Structural analyses of FlgY.
a, Structural overlap of FlgY C-terminal globular domain (predicted by AlphaFold3) with MgtE_N domain of MgtE (PDB: 2YVX) or three ARM-like domains of FliG (PDB: 3HJL). b, Structural overlap of FlgY C-terminal globular domain with canonical ARM repeat of β-catenin (PDB: 3BCT). c, Size-exclusion chromatography (SEC) profile of purified FlgY15-172 and different truncation variants. d, Crosslinking of purified FlgY15-172 and FlgY90-172 by adding Ethylene glycol bis(succinimidyl succinate) (EGS). Since FlgY90-172 is mainly composed of the ARM-like domain and forms a dimer that fits the cryo-ET map of the E-ring, we use FlgY90-172 to represent the dimeric FlgYARM. e, AlphaFold3-predicted structure of FlgY dimer. f, MST binding curves of FlgY (red) or FlgYARM (pink) with PflA. Data are presented as the mean ± SD from three independent experiments. g, Complex structure of FlgY dimer and PflA predicted by AlphaFold3.
Extended Data Fig. 5 The location of PflD in motor and its interaction with FcpNO.
a, The central cross section of the complete motor structure (top) and side view of the cage region (bottom) from wild type and three mutants. The position of PflD is indicated with a green arrow. b, Left: AlphaFold3-predicted structure of PflD with transmembrane motif highlighted in red. Right: The C-terminal domain and middle loop region of PflD show structural similarity to the N1 domain of PilQ and the loop region of PilP, respectively. c, Comparison of complex structure of PflD-FcpMNO predicted by AlphaFold3 and PilMNOP of T4P (PDB:3JC8). d, BTH analysis for the interaction of PflD and FcpNO or other proteins. Left: the interaction of PflD and FcpNO with their transmembrane motif and the T25 or T18 tags were cloned at the N-terminal end next to transmembrane motif; Right: the interaction of PflD and other proteins with periplasmic region only.
Extended Data Fig. 6 Motility assay of PflD-loop truncation and PilP-loop exchange.
a-b, Soft agar motility assay of wild type, ΔpflD, and ΔpflD complemented with PflD or various truncations of the PflD loop region. The domain organization of full-length PflD and its various truncation constructs made for soft agar motility analyses were indicated on the top. Bars show mean ± SD; dots represent individual halos (n = 6 from three experiments). Statistical significance was calculated by two-tailed unpaired Student’s t tests between wild type and each mutants.
Extended Data Fig. 7 Visualization of the peptidoglycan (PG) layer in Campylobacter jejuni.
a-b, Cryo-ET images of the bacterial tip regions in wild type and ∆pflA mutant captured the PG layer in the periplasmic region. c-d, The subtomogram averaged structure of Campylobacter jejuni flagellar motors in wild type (c) and ∆pflA mutant (d) present layer features around the basal and medial disks region.
Extended Data Fig. 8 Dynamic changes and C-ring symmetry of Campylobacter jejuni motor.
a, c, Central section of motor structures from wild type Campylobacter jejuni with two distinctive heights from LP-ring to MS-ring. b, d, Inset of a and c with measured height from LP-ring to MS-ring, respectively. e-f, 10 classified structures of wild type motor. The relationships between the numbers of the particles of each class averaged structure and the distance between the LP- and MS-rings are shown by column and line, respectively (f). g, Focused classification on the C-ring reveals its 40 subunits.
Extended Data Fig. 10 Summary of motor structures imaged by cryo-ET and mapped to bacterial species tree.
The phylogenetic tree in the center is derived from Fig. 6c. The motor structures investigated by cryo-ET are displayed on the periphery, with one representative species from each genus and taxon group labeled at the top of the image. Species names highlighted in blue indicate that their stator complexes are dynamic and invisible, while those marked in red represent that their stator complexes are visible. Potential E-ring is indicated by pink arrow and question marks in the image of Acetonema longum and Arcobacter butzleri mean that the position of E-ring is uncertain. Information of motor structure for species (follow a clockwise order) were taken from references: Acetonema longum1, Leptospira interrogans89, Borrelia burgdorferi49,90, Treponema pallidum91,92, Arcobacter butzleri93, Campylobacter jejuni (this study), Wolinella succinogenes93, Helicobacter pylori50, Bdellovibrio bacteriovorus93, Hyphomonas neptunium1, Caulobacter crescentus94, Bordetella bronchiseptica47, Hylemonella gracilis1, Legionella pneumophila2, Pseudomonas aeruginosa2, Shewanella oneidensis2, Vibrio alginolyticus95, Plesiomonas shigelloides96, Salmonella enterica3, Escherichia coli97.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–10.
Supplementary Tables (download XLSX )
Supplementary Tables 1–10.
Supplementary Data 1 (download XLSX )
Raw co-immunoprecipitation data for FcpO/N/M.
Supplementary Data 2 (download XLSX )
Normalized and processed qPCR log2(fold change) data.
Supplementary Video 1 (download MP4 )
Video showing conformational fluctuations of the auxiliary periplasmic scaffolds and the surrounding cell envelope, which do not rotate with the rotor.
Supplementary Video 2 (download MP4 )
Video showing that the C ring consists of 40 subunits and visualizing rotation at the molecular scale.
Supplementary Video 3 (download MP4 )
Video showing the early incorporation of stator complexes and surrounding scaffolds before rod formation, as well as a transient periplasmic ring located above the E ring and MS ring.
Supplementary Video 4 (download MP4 )
Video showing multiple flagellar components with structural similarity to type IV pilus proteins, including regions corresponding to PilQ, PilP, PilMNO and PilF.
Source data
Source Data Extended Data Fig. 3 (download XLSX )
Raw swimming-velocity data for Campylobacter jejuni wild type and ΔflgY mutant under media of different viscosities.
Source Data Extended Data Fig. 4d (download JPG )
Raw, uncropped SDS–PAGE gel.
Source Data Extended Data Fig. 4f (download XLSX )
Original MST measurement data.
Source Data Extended Data Fig. 6b (download XLSX )
Original measurements of soft-agar motility halo diameters.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feng, X., Tachiyama, S., He, J. et al. Structural insights into the assembly and evolution of a complex bacterial flagellar motor. Nat Microbiol 11, 770–785 (2026). https://doi.org/10.1038/s41564-025-02248-5
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41564-025-02248-5