Abstract
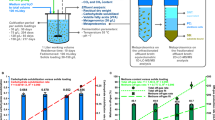
The breakdown of cellulose is one of the most important reactions in nature1,2 and is central to biomass conversion to fuels and chemicals3. However, the microfibrillar organization of cellulose and its complex interactions with other components of the plant cell wall poses a major challenge for enzymatic conversion4. Here, by mining the metagenomic ‘dark matter’ (unclassified DNA with unknown function) of a microbial community specialized in lignocellulose degradation, we discovered a metalloenzyme that oxidatively cleaves cellulose. This metalloenzyme acts on cellulose through an exo-type mechanism with C1 regioselectivity, resulting exclusively in cellobionic acid as a product. The crystal structure reveals a catalytic copper buried in a compact jelly-roll scaffold that features a flattened cellulose binding site. This metalloenzyme exhibits a homodimeric configuration that enables in situ hydrogen peroxide generation by one subunit while the other is productively interacting with cellulose. The secretome of an engineered strain of the fungus Trichoderma reesei expressing this metalloenzyme boosted the glucose release from pretreated lignocellulosic biomass under industrially relevant conditions, demonstrating its biotechnological potential. This discovery modifies the current understanding of bacterial redox enzymatic systems devoted to overcoming biomass recalcitrance5,6,7. Furthermore, it enables the conversion of agro-industrial residues into value-added bioproducts, thereby contributing to the transition to a sustainable and bio-based economy.
Similar content being viewed by others
Main
Cellulose, the most abundant renewable polymer on Earth, poses a challenge for biological depolymerization. Although composed entirely of glucose residues, its crystalline microfibrillar structure, along with its association with lignin and hemicelluloses in plant cell walls, make it highly resistant to degradation. As a result, its breakdown in nature is slow and involves complex multi-component enzymatic systems1,6,8,9,10.
This process can be carried out by a plethora of microorganisms through distinct biochemical routes and the canonical model, based on extensive research on filamentous fungi and bacteria, comprises at least three major hydrolytic activities: endo-β-glucanase, cellobiohydrolase and β-glucosidase. In this model, endo-β-glucanases cleave cellulose chains internally, whereas cellobiohydrolases primarily act on cellulose chain ends, although they can also exhibit some endo-cleavage activity11,12. Both enzymes release cellooligosaccharides, which are then converted into glucose by β-glucosidases1.
This model was subsequently modified with the discovery of redox enzymes known as lytic polysaccharide monooxygenases (LPMOs)5,6,7,13,14,15,16,17,18,19, some of which are capable of acting on crystalline patches of cellulose5. Furthermore, the aforementioned hydrolytic activities can be found in large cell-bound multi-enzymatic complexes known as cellulosomes20,21 or as cell-free multi-modular proteins comprising several catalytic and non-catalytic domains22. This overview underscores the intricate and diverse nature of known microbial systems dedicated to overcoming cellulose recalcitrance. Nevertheless, most microbial life remains unculturable in laboratory conditions, leaving much of its genetic potential obscured.
In this work, we explored the genomic dark matter of microbial communities specialized in plant biomass breakdown using a multidisciplinary approach that included metagenomics, proteomics, carbohydrate enzymology by chromatographic, colorimetric and mass spectrometric methods, fourth-generation synchrotron-based X-ray diffraction, fluorescence and absorption spectroscopies, site-directed mutagenesis, CRISPR–Cas9 fungal genetic engineering and 65-l and 300-l pilot plant bioreactor experiments. We identified a metalloenzyme that enhances cellulose conversion through a previously undescribed mechanism of substrate binding and oxidative cleavage. This discovery establishes a new frontier in redox biochemistry for plant biomass depolymerization, one of the most important bioreactions in nature with far-reaching implications for biotechnology.
A cellulose oxidative cleaving enzyme
To identify previously undescribed biomass-active microorganisms and biocatalysts, we carried out a metagenomic analysis of soil samples covered with sugarcane bagasse that has been maintained over decades in a biorefinery (Quatá, São Paulo, Brazil). We found that microbial diversity in this environment had a sharp decrease (approximately 1,000 operational taxonomic units (OTUs)) compared with the bulk soil from native vegetation in the vicinity of the bagasse pile (approximately 2,200 OTUs) (Fig. 1a,b and Supplementary Fig. 1). Moreover, this decrease in microbial diversity was accompanied by an increase in the number of metagenome-assembled genomes (MAGs) associated with pathways involved in polysaccharide breakdown and metabolism, indicating a microbial specialization towards lignocellulosic bacteria (Extended Data Fig. 1a and Supplementary Fig. 2).
a, Sampling site indicating the area covered with sugarcane bagasse and an adjacent area where a bulk control soil sample was taken. SBS, sugarcane bagasse-covered soil. b, The alpha diversity index shows that the sugarcane bagasse-covered soil has reduced microbial diversity compared with the control soil. c, Phylogenetic tree illustrating the relationships between the 124 recovered MAGs. Previously undescribed genomes in the taxonomy are highlighted with purple stars. d, Predicted metabolic pathways, glycoside hydrolases (GHs) and the newly described CelOCE in ‘Candidatus Telluricellulosum braziliensis’, highlighting its potential role in cellulose conversion. GHs with low sequence identity (<30%) to known GHs in the CAZy database (https://www.cazy.org/), or those with activities matching their predicted family but not subfamily, are shown in red. Predicted enzymatic activities consistent with their GH family or subfamily classification are indicated by Enzyme Commission (EC) numbers in parentheses.
Among the recovered high-quality MAGs (Fig. 1c and Supplementary Tables 1 and 2), one member from a recently proposed23 and uncharacterized uncultured bacterial phylum 4 (UBP4) was further investigated owing to its uncharted potential for plant cell wall breakdown. This potential was evidenced by multiple genes that encode glycoside hydrolases, such as those from the families GH3, GH5, GH9, GH39, GH43, GH44, GH74 and GH148 (Fig. 1d and Extended Data Fig. 1b). Whereas the UBP4 phylum was initially identified in waste water in a large-scale metagenome reconstruction effort23, here we describe a soil-derived MAG that diverges at the family level from existing UBP4 genomes in the GTDB database24. On the basis of its extensive CAZyme repertoire (Fig. 1d and Extended Data Fig. 1c) and Brazilian soil origin, we propose the name ‘Candidatus Telluricellulosum braziliensis’ for this uncultured bacterium (SeqCode, https://seqco.de/)25.
Using hidden Markov models for remote homology detection, we selected eight sequences from this genome that showed at least 10% sequence identity to known carbohydrate-processing proteins but lacked matches in the CAZy database26. These sequences were synthesized, expressed in Escherichia coli and biochemically characterized (Supplementary Tables 3 and 4). One of these proteins showed the capacity to boost the depolymerization of pretreated sugarcane bagasse (approximately 21% improvement) by a cellulolytic enzyme cocktail produced by an industrially competitive T. reesei strain27,28 (Fig. 2a). This fungal cocktail comprises key enzymatic activities for the efficient depolymerization of cellulose and heteroxylans, including cellobiohydrolases (CBH1 and CBH2), endo-β-glucanases (EGL1, EGL2 and EGL5), LPMO (LPMO9A), β-glucosidase (heterologous CEL3A from Talaromyces emersonii), endo-β-xylanases (XYN1, XYN2 and XYN4), β-xylosidase (BXL1) and other accessory enzymes (Supplementary Tables 5 and 6). The fact that a bacterial enzyme can enhance the performance of an optimized fungal CAZyme cocktail is notable and highlights its potential for plant biomass degradation applications.
a, Boosting effect on the saccharification of pretreated sugarcane bagasse, microcrystalline cellulose and amorphous cellulose when CelOCE is combined with a cellulolytic enzyme cocktail. Data are the mean ± s.d. from three independent experiments. Statistical significance was determined by one-way analysis of variance (ANOVA) with Tukey’s post hoc test (**P < 0.01). Percentages indicate the difference between treatments. b, High-performance anion-exchange chromatography with pulsed amperometric detection (HPAEC–PAD) profiles of reactions containing reductant and enzymes CelOCE (dark green line), KdgF (orange line) and BacB (blue line). Control reactions using only sugarcane bagasse (dark grey line), only ASC (grey line), sugarcane bagasse and CelOCE, no ASC (light green line), sugarcane bagasse and ASC (light grey line) and with inactivated enzyme (grey–blue line) are also shown. Standard C1-oxidized and non-oxidized cellooligosaccharides are represented by black lines. DP, degree of polymerization; ox, oxidized. c,d, Amorphous (c) and microcrystalline cellulose (d) binding isotherms comparing the enzyme in the presence or absence of a reductant (ASC). The binding isotherms for PASC were fitted using the Langmuir–Freundlich model. The fit for CelOCE without ASC yielded n = 2.5 (n, Langmuir–Freundlich coefficient) and R2 = 0.99, whereas the fit for CelOCE with ASC resulted in n = 2.1 and R2 = 0.99. Data are the mean ± s.d. from three independent experiments. e, SSN depicting three distinct isofunctional clusters of the reference proteins BacB (Protein Data Bank (PDB) ID: 3H7J), KdgF (PDB ID: 5FPZ) and CelOCE (this study). Connections between nodes indicate at least 30% sequence identity with an alignment e-value cut-off of 1 × 10–5.
The purified protein exhibited no detectable hydrolytic activity on a broad range of substrates, including polysaccharides, oligosaccharides and synthetic substrates (Supplementary Table 7), indicating a non-hydrolytic mode of action, that potentially involves a redox mechanism. Hence, to investigate this hypothesis, we analysed the products released from diverse substrates under redox conditions, specifically in the presence of oxygen and an electron donor (ascorbic acid (ASC)). We found that the enzyme released only one product, identified as cellobionic acid (Supplementary Fig. 3), from lignocellulose (pretreated sugarcane bagasse) (Fig. 2b), microcrystalline cellulose (Avicel) (Supplementary Fig. 4a) and amorphous cellulose (PASC) (Supplementary Fig. 4b). Activity tests on cellooligosaccharides (C2–C6) and other polysaccharides, including chitin, mannan, xylan, xyloglucan, arabinoxylan, mixed-linked β-glucan, laminarin, lichenan, starch and pectin, did not show any degradation products, indicating a clear preference for cellulosic substrates (Supplementary Fig. 4c–l). Particularly, the enzyme showed no binding affinity to cellobiose (Supplementary Fig. 5) and did not lead to the consumption of cellobiose after 16 h in the presence of ASC (Supplementary Fig. 6), ruling out the possibility of cellobionic acid generation from cellobiose. Furthermore, the enzyme enhanced the saccharification efficiency of the same Trichoderma enzyme cocktail, increasing the glucose yields by approximately 8% and 12.5% on Avicel and PASC (Fig. 2a), respectively, supporting its specific activity and role in cellulose conversion.
Distinct techniques, including affinity gel electrophoresis (Supplementary Fig. 7), Fourier-transform infrared spectroscopy (FTIR) (Extended Data Fig. 2a) and X-ray photoelectron spectroscopy (XPS) (Extended Data Fig. 2b) demonstrated qualitatively the capacity of this protein to bind to cellulose. Moreover, only a denaturing agent such as SDS effectively displaced the enzyme from cellulose (Extended Data Fig. 2a,b). Binding isotherms revealed dissociation constants in the low micromolar range (7–9 µM) for PASC (Fig. 2c), whereas saturation was not reached for microcrystalline cellulose (Avicel) (Fig. 2d), similar to what has been observed for cellulose-active LPMOs29. Notably, the redox state of the copper did not affect binding affinity or maximum binding capacity (Bmax) as much as observed with LPMOs, probably owing to the buried nature of the catalytic copper in this enzyme. Competitive binding assays further showed that the enzyme markedly retains its cellulose binding capacity in the presence of bovine serum albumin (BSA), similarly to cellulases30 (Supplementary Fig. 8). Collectively, these cellulose binding assays highlight the specific interaction and high affinity of the discovered enzyme for cellulose.
To document the taxonomic occurrence of this enzyme, we performed sequence similarity network (SSN) and phylogenetic analyses, revealing a number of orthologues (sequence identity >30%) annotated as hypothetical proteins in sequence databases. These orthologues were identified across diverse bacterial phyla associated with biomass breakdown, as well as in archaea (Extended Data Fig. 3). Species including Bacteroides caccae, Draconibacterium halophilum, Rhodocytophaga rosea, Adhaeribacter swui and Lacunisphaera limnophila contain orthologous sequences and distinct CAZymes associated with cellulose breakdown, such as those from families GH5, GH8 and GH9.
The closest characterized homologues, identified by SSN analysis (Fig. 2e), belong to distinct isofunctional clusters (sequence identity <30%) and are involved in antibiotic biosynthesis (BacB)31 or uronate metabolism (KdgF)32. Control experiments with heterologously produced BacB and KdgF did not yield any products from pretreated sugarcane bagasse (Fig. 2b), indicating that the oxidative cleavage of cellulose is specific to the sequence cluster containing the discovered enzyme.
Our results support the discovery of a cellulose oxidative cleaving enzyme (CelOCE) from a previously undescribed phylum associated with plant cell wall breakdown. CelOCE cleaves cellulose by a previously unknown exo-acting mechanism, releasing cellobionic acid as the sole product.
Homodimeric monocopper architecture
To elucidate the mechanism behind its redox activity, we solved the crystal structure of CelOCE in three different states (Extended Data Table 1). CelOCE adopts a compact jelly-roll fold comprising two anti-parallel β-sheets (Extended Data Fig. 4a) and forms a back-to-back homodimer, with the active sites positioned on opposite faces (Fig. 3a and Extended Data Fig. 4b). This dimeric arrangement, stabilized by extensive β-sheet interactions (Extended Data Fig. 4b and Supplementary Table 8), was further validated in solution by analytical size-exclusion chromatography coupled with multi-angle light scattering (Supplementary Fig. 9), supporting the conclusion that it represents the biologically active form.
a, Dimeric arrangement observed in CelOCE crystal structures, highlighting the dimer interface, the location of the active site (blue region encompassing the copper atom) and the cellulose binding site (grey region). b, Octahedral copper coordination sphere in the CelOCE crystal structures, showing the copper-coordinating residues H44, H46, H84 and Q50 as sticks, the copper atom as an orange sphere and water molecules as red spheres. Dashed lines indicate distances in ångström. c, Surface representation of a CelOCE protomer, highlighting the flattened catalytic interface that enables the interaction with the cellulose. The copper-coordinating histidine and proline residues contributing to this unconventional interface are shown. The residue F33, proposed to be involved in disaccharide recognition in the active site pocket, is also shown. d, ITC data for copper binding to CelOCE. The main plot depicts the binding isotherm (green circles) with its theoretical fit (black line). Thermodynamic parameters are shown in the bottom right. Top left inset, thermogram. ΔG, Gibbs free energy change; ΔH, enthalpy change; ΔS, entropy change; T, temperature. e, EPR spectra of CelOCE in the absence (grey line) and presence (green line) of reductant (ASC). The typical EPR spectrum of a Cu2+ centre in the resting state (grey line) is abolished after reduction to Cu+ (green line). a.u., arbitrary units. f, Time-dependent analysis of cellobionic acid production by CelOCE under aerobic and anaerobic conditions and the role of an electron donor (ASC). Anaerobic reactions were conducted with ASC, either in the absence or presence of 100 µM hydrogen peroxide (H2O2). Aerobic reactions were carried out with or without ASC. Data are the mean ± s.d. of three independent experiments.
Each subunit contains a single copper atom in a distorted octahedral coordination sphere, which involves three histidine (H44, H46 and H84), one glutamine (Q50) and two water molecules (Fig. 3b). In the presence of a sugar mimetic, glycerol, one water molecule is replaced by an oxygen atom from glycerol, displacing the remaining water molecule to the equatorial plane alongside H44, H46 and Q50 (Extended Data Fig. 5a). At low pH (approximately pH 3.0), H44 is probably protonated and was observed to flip towards the bulk solvent (Extended Data Fig. 5b), a behaviour recently reported for an LPMO33. The copper atom is buried in the active site, which exhibits a pocket-like topology, in contrast to LPMOs (Fig. 3c and Extended Data Fig. 6a). Furthermore, this active site is nestled in a flattened interface, a structural arrangement well suited for interaction with cellulose (Fig. 3c).
The presence of copper in CelOCE was further confirmed through synchrotron X-ray fluorescence (XRF) analyses (Supplementary Fig. 10), X-ray absorption spectroscopy (XAS) (Supplementary Fig. 11), isothermal titration calorimetry (ITC) (Fig. 3d) and electron paramagnetic resonance (EPR) (Fig. 3e and Supplementary Fig. 12). Long incubation times of CelOCE with other metals, such as nickel after enzyme saturation with copper did not alter the XAS spectrum, indicating a preference for copper (Supplementary Fig. 11). Furthermore, copper binding affinity was measured using ITC, revealing a dissociation constant in the low micromolar range (Kd = 1.14 ± 0.11 µM) (Fig. 3d). The binding process is primarily driven by enthalpic contributions (ΔH = −19.8 ± 0.2 kJ mol−1) and is accompanied by a favourable entropic component (TΔS = −14.1 ± 0.2 kJ mol−1), resulting in an exothermic process with one copper atom binding to each monomer.
The EPR spectrum of CelOCE is characteristic of a copper centre in a +2 oxidation state featuring an axial EPR signature (gz > gy ≈ gx, Az > Ay ≈ Ax) (Fig. 3e and Supplementary Table 9). It contrasts with the predominance of a rhombic EPR signature in LPMOs (gz ≠ gy ≠ gx, Az ≠ Ay ≠ Ax)5,34, indicating that CelOCE has a distinct coordination sphere. Control experiments using EPR in the presence of a reductant (ASC) showed a strong decrease in the signal, indicative of the formation of the EPR-silent Cu(I) form (Fig. 3e). Furthermore, the chelating agent EDTA also silenced the EPR signal (Supplementary Fig. 12) and strongly decreased the release of cellobionic acid (Supplementary Fig. 13), highlighting the importance of copper for catalysis.
Electron donor and co-substrate
Next, we sought to understand the redox requirements for this cellulose oxidative cleaving activity. We first confirmed its strict requirement for an electron donor for activity, as product formation was observed only in the presence of a reductant (ASC) (Fig. 3f, Extended Data Fig. 7a and Supplementary Fig. 14). This reduction step is well documented for redox enzymes and distinct types of small and macromolecular electron donors are capable of driving the activity of, for instance, LPMOs19,35,36.
Regarding the requirement for a co-substrate, the enzyme was inactive in the absence of both oxygen and hydrogen peroxide, even in the presence of a reductant (Fig. 3f, Extended Data Fig. 7b and Supplementary Fig. 15). However, CelOCE displayed activity under anaerobic conditions in the presence of ASC and when supplemented with exogenous hydrogen peroxide, indicative of peroxygenase activity (Fig. 3f and Supplementary Fig. 15). The enzyme showed similar turnover rates of oxidized product formation under aerobic conditions (0.053 ± 0.003 min−1) or under anaerobic conditions in the presence of hydrogen peroxide (0.050 ± 0.003 min−1) (Fig. 3f and Extended Data Fig. 7b). This result was unexpected because LPMOs perform orders of magnitude better in the presence of exogenous hydrogen peroxide than with oxygen37,38. This suggests that the in situ peroxide generation is not a rate-limiting factor for CelOCE. Further support comes from the observation that the turnover rate for peroxide generation (0.23 ± 0.05 min−1) is approximately fourfold higher than that for cellobionic acid production (Supplementary Fig. 16), although the reaction stoichiometry is 1 mol of product generated per 1 mol of hydrogen peroxide consumed (Extended Data Fig. 7c). Notably, the turnover rate observed for CelOCE falls in the range typically seen for LPMOs acting on Avicel under aerobic conditions without the addition of exogenous hydrogen peroxide (Supplementary Table 10). This turnover rate would be expected given its non-processive (pocket-like active site) and exo mode of action, particularly when acting solely on Avicel, for which its efficiency is inherently limited by the low natural abundance of cellulose extremities, which serve as the primary sites for its catalytic activity.
The fact that CelOCE is a homodimer with the active site of each subunit located at the opposite side of the biological assembly might contribute to its capacity to be self-sufficient in generating hydrogen peroxide. In this model, while one active site is protected from the solvent by interacting with the cellulose, the other is probably free and acts as an in situ peroxide supplier (Fig. 4a), to ensure that peroxide is generated near the active site engaged with cellulose and enabling its effective use. Supporting this mechanism, increasing concentrations of cellulose did not inhibit the peroxide production (Supplementary Fig. 17), whereas disrupting the dimer by deleting the four N-terminal residues (Supplementary Figs. 17 and 18) resulted in a reduction in the peroxide generation.
a, Proposed model for the simultaneous interaction of CelOCE with cellulose and in situ hydrogen peroxide generation. While one active site engages the non-reducing end of a cellulose chain, the other active site in the homodimer is probably available to generate hydrogen peroxide, the essential co-substrate for cellulose oxidative cleavage. b, Cellotetraose (represented as spheres and sticks) was docked and equilibrated in the CelOCE structure, demonstrating that two glucosyl residues can be accommodated in the active site pocket. The C1 carbon of the −1 glucosyl moiety is positioned favourably for oxidative attack, leading to the production of cellobionic acid as the sole product, which aligns with the biochemical data.
Together, these results indicate that after reduction of the catalytic copper by an electron donor, the enzyme CelOCE is primed to catalyse a peroxygenase reaction, which is further fuelled by an innovative dimerization strategy for in situ peroxide generation.
Exo-type mechanism on cellulose
To elucidate the molecular basis of how this metalloenzyme recognizes cellulose for oxidative cleavage, we conducted a series of structural and computational analyses. CelOCE exhibits an unconventional active site topology. The catalytic copper is buried approximately 5 Å (Extended Data Fig. 6a) deep in a pocket formed by metal-coordinating residues, along with one tyrosine (Y105), one phenylalanine (F33), one arginine (R102), three leucines (L13, L14 and L41) and one acidic residue (a glutamic acid, E96) (Supplementary Fig. 19). Computational calculations suggest that this pocket is large enough to accommodate a disaccharide (Extended Data Fig. 6b) and is stereochemically compatible with glucosyl moieties (Extended Data Fig. 6c).
In the modelled complex of CelOCE with a cellooligosaccharide, the non-reducing end glucosyl (−2) residue is primarily anchored by the side chains of E96 and Q50, whereas the −1 glucosyl residue stacks against F33, productively positioning the C1 atom near the catalytic copper (Fig. 4a,b and Supplementary Fig. 20). Recognition of the reducing end of cellulose chains seems to be stereochemically unfavourable for the observed C1 regioselectivity; however, the elucidation of the substrate–enzyme complex will be essential to experimentally determine the activity directionality of the enzyme (Extended Data Fig. 8). To validate these in silico predictions, we generated an F33A variant of the enzyme, which showed impaired catalytic activity (Supplementary Fig. 21), supporting the proposed binding mode (Fig. 4a and Extended Data Fig. 6c) and consistent with the release of cellobionic acid as the sole product (Fig. 4b). The exclusive detection of cellobionic acid throughout the reaction time course, including at early stages (Fig. 3f), provides further evidence for its exo mode of action.
Another part of this mechanism relies on the rigidness of a solvent-exposed loop in the vicinity of the copper-binding site, which is stabilized by proline residues. This loop, in conjunction with the C-terminal α-helix and the second N-terminal β-strand, forms a flattened surface (Fig. 3c). Typically, surface loops are prone to be flexible and rarely form flat surfaces; however, owing to the presence of these proline residues and metal coordination, the loop adopts a unique geometry that is stereochemically compatible to interact with cellulose (Fig. 3c). These proline residues are highly conserved in CelOCE orthologues, featuring the motif PXHXHP, which includes two histidine residues (H44 and H46) involved in copper coordination (Supplementary Fig. 22).
These two distinctive features of the catalytic interface, the pocket-like active site and the flattened surface topology, unequivocally demonstrate that this enzyme operates in an exo mode, representing an unprecedented mechanism among carbohydrate-active oxidoreductases.
Cooperative action in converting biomass
To further explore how CelOCE contributes to the conversion of plant biomass, we assessed its cooperative action with key cellulases, including GH539 and GH4540 endoglucanases and a GH7 cellobiohydrolase (Cel7A)41. We observed a remarkable additive effect with endo-acting cellulases (up to around 300% increase), but no positive effect with the exo-acting cellobiohydrolase Cel7A (Fig. 5a). This lack of additive effect with Cel7A is consistent with the exo-acting mode of CelOCE, which does not generate new cellulose ends.
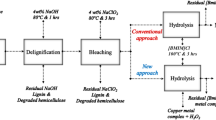
a, Complementary assays showing cooperative action of CelOCE with endo- and exo-acting cellulases on microcrystalline cellulose. b, Genetic engineering approach used to integrate the sequences encoding CelOCE and L. similis AA9A into the T. reesei genome. c, Saccharification efficiency of the enzyme cocktail produced by the engineered strains under industrially relevant conditions, using pretreated sugarcane bagasse as the technical substrate. Data in a and c are the mean ± s.d. of three independent experiments. In a, statistical significance was determined by one-way ANOVA with Tukey’s post hoc test (***P < 0.001).
Next, we evaluated the role of CelOCE in combination with an enzyme cocktail produced by T. reesei Br_TrR03, an engineered strain developed for lignocellulose biorefineries27, mimicking industrial conditions. For this purpose, the sequence encoding CelOCE was integrated into the genome of Br_TrR03 using a customized CRISPR–Cas9 approach27 (Fig. 5b). The secretome produced by this engineered strain (Supplementary Figs. 23 and 24) under industrially relevant conditions considerably increased the glucose release by 21% and 19% for pretreated sugarcane bagasse and eucalyptus materials, respectively (Fig. 5c and Extended Data Fig. 9c). Mass spectrometry confirmed the presence of CelOCE in the secretome (Supplementary Tables 11 and 12) and increased levels of cellobionic acid were observed during biomass saccharification, consistent with its activity and mode of action (Extended Data Fig. 9).
Under industrial conditions, this boosting effect surpassed that achieved with the same parental strain expressing a thermostable fungal AA9 LPMO from the fungus Lentinus similis (LsAA9A) (Fig. 5c, Supplementary Figs. 25–27 and Supplementary Tables 13 and 14), highlighting the potential of CelOCE to enhance lignocellulose conversion. This increased additive effect, compared with L. similis AA9A, may be attributed to the unique exo-acting mechanism and in situ hydrogen peroxide generation capacity of CelOCE. Notably, the enzyme cocktail produced by the parental strain is already highly effective for lignocellulose deconstruction, with high cellulase and β-glucosidase activities27, further emphasizing the complementary role of CelOCE to the Trichoderma secretome for biomass saccharification, a well-established workhorse in biotechnology. Furthermore, the CelOCE-containing cocktail was successfully produced in 65-l and 300-l pilot plant bioreactors (Supplementary Fig. 23), demonstrating its industrial relevance.
Collectively, these results indicate that CelOCE can boost the conversion of lignocellulosic biomass both in vitro (exogenously added) and in vivo (co-expressed in Trichoderma) under industrially relevant conditions. This enhancement is primarily attributed to its cooperative action with endocellulases.
Discussion
CelOCE has an innovative redox mechanism of cellulose cleavage. Its unique copper coordination in a pocket-like active site enables an exo mode of action, exclusively releasing cellobionic acid. The flattened catalytic interface mediates the interaction with cellulose, and the homodimeric structure enables in situ hydrogen peroxide generation, fuelling the peroxygenase activity of the enzyme. This combination of features results in a cooperative action with hydrolytic endocellulases, boosting cellulose depolymerization.
Notably, CelOCE comprises only 115 residues, one of the smallest catalytically active proteins found in carbohydrate enzymology. Its compact size and distinct copper coordination, lacking the characteristic N-terminal histidine found in LPMOs, offer advantages for biotechnological applications, including improved diffusion, facilitated protein engineering and even the design of artificial enzymes or new functions. The functional versatility of this jelly-roll scaffold is further evidenced by the diverse enzymatic activities already observed in nature42,43.
The biotechnological potential of this metalloenzyme was demonstrated by its co-expression with cellulases and hemicellulases in Trichoderma, leading to enhanced glucose release from pretreated lignocellulosic biomass under industrially relevant conditions. This boosting effect exceeded that of the same strain expressing a fungal AA9 LPMO. The CelOCE-containing cocktail was produced in both 65-l and 300-l pilot plant bioreactors using a low-cost carbon source, the biomass was pretreated in a pilot plant reactor and saccharification assays were conducted at high solids loading (>15%), closely mimicking real-world biorefinery conditions.
This discovery enables further developments in redox biochemistry for plant polysaccharide depolymerization by revealing that copper-catalysed peroxygenase reactions, until now restricted to LPMOs7,38, have evolved in other biocatalysts. Furthermore, by shedding light on the bacterial redox systems involved in carbohydrate breakdown and metabolism, this study increases our understanding of the mechanisms that underlie the global carbon cycle and provides opportunities for the bioconversion of agro-industrial residues into value-added bioproducts.
Methods
Metagenomic and bioinformatic approaches
Soil sample collection
Soil samples were collected from a sugarcane mill in Quatá, São Paulo, Brazil, where residual sugarcane bagasse had been stored over 20 years (Fig. 1a). After mechanically removing the surface bagasse layer, samples were collected at the soil surface and from a depth of 20 cm. These samples are referred to as sugarcane bagasse-covered soil (SBS). A control sample was collected from nearby soil without bagasse coverage. All samples were immediately frozen in liquid nitrogen and stored at −80 °C until further processing.
Nucleic acid extraction
Microbial DNA was extracted from both SBS and bulk control soils using the FastDNA Spin Kit for Soil (MP Biomedicals). Ten grams of soil samples were pulverized with an oscillating ball mill (TE-350, Tecnal) and used to conduct five extraction batches of 2 g each, resulting in five separate DNA extracts. These extracts were then transferred to Lysing Matrix E Tubes, external contaminants were solubilized with MT buffer and sodium phosphate buffer was added for cell lysis. The samples were homogenized using a FastPrep FP120 instrument (MP Biomedicals). Protein precipitation solution was added to the supernatant to separate nucleic acids from cellular debris, followed by centrifugation at 14,000g for 5 min. The resulting supernatant was then mixed with a binding matrix, incubated for 3 min and transferred to a spin filter. After centrifugation (14,000g for 1 min), the pellet was washed with a salt–ethanol solution (SEWS-M), dried and resuspended in ultrapure water. Further purification was performed using the PowerClean DNA Clean-Up Kit (Mo Bio Laboratories). DNA quality was assessed by 0.8% (w/v) agarose gel electrophoresis.
16S rRNA amplicon sequencing and analysis
The V4 region of the 16S rRNA gene was amplified in triplicate using the primers 515F and 806R. Paired-end sequencing (2 × 300 bp) was performed on an Illumina MiSeq platform (V3 kit, 600 cycles) using the MiSeq reporter software at the high-performance sequencing facility of the Brazilian Biorenewables National Laboratory (LNBR). The ZymoBIOMICS microbial community DNA standard II served as a positive control. For taxonomic analysis, paired-end reads were quality-checked with FastQC v.0.12.0 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and filtered with Trimmomatic44 v.0.36 to remove adapters and low-quality reads. Filtered reads were merged using the fastq_mergepairs function from the Usearch v.10 package45 (minimum overlap of 50 bp and maximum error of 0.5), followed by the removal of primer and singleton sequences. The UPARSE unoise3 function was used for denoising and zOTU (zero radius OTUs) recovery. Taxonomic assignment was performed with the sintax function (cut-off 0.8, RDP database v.16)46. Further analyses were performed using the phyloseq v.1.20 package in R Studio v.1.3.1093 (https://bioconductor.org/packages/release/bioc/html/phyloseq.html). Details on read counts and diversity indices are described in Supplementary Table 15.
Metagenomic short- and long-read sequencing
Metagenomic libraries were constructed using the Nextera library preparation kit (Illumina). Quantification and quality control of the libraries were performed using quantitative PCR and the KAPA library quantification kit (Roche) and the Agilent Bioanalyzer 2100 system (Agilent Technologies). Sequencing was performed on an Illumina HiSeq 2500 device (2 × 250 bp) using the HiSeq 2500 control software. Furthermore, long-read sequencing was conducted on a MinION device (Oxford Nanopore) using the MiniKNOW v.19.12.5 software. For long-read sequencing, 1 µg of high-molecular-mass DNA from the SBS sample was prepared with SQK-LSK109 and Native Barcoding Kits.
Metagenomic de novo assembly and binning
Raw metagenomic sequences underwent quality control and trimming using FastQC v.0.12.0 and Trimmomatic v.0.36, followed by taxonomic classification with Kaiju47 v.1.7.4. Quality-filtered reads were de novo assembled using IDBA_UD v.1.1.1 with pre-correction and k-mer sizes48 of 20–60 (Supplementary Table 16). The resulting assemblies were binned using MetaWRAP v.1.349, generating initial bin sets with MetaBAT2, MaxBin2 and CONCOCT, followed by refinement and reassembly (minimum completion 55%, maximum contamination 15%). Bins were then taxonomically classified and functionally annotated using the modules Classify and Annotate_bins, respectively. Furthermore, long reads from Oxford Nanopore Technologies sequencing were used for scaffolding with SSPACE-long-reads v.1.1 with parameters: -k 5, -a 0.7, -x 1, -m 50, -o 20 and -n 1000. Final MAGs were assessed for completeness and contamination using CheckM250 v.1.0.2 and further classified with GTDB-tk against the GTDB database51 release 214. Detailed information on recovered MAGs is summarized in Supplementary Table 2. Gene prediction and annotation were performed with Prokka52 v.1.11. CAZyme and PUL annotations followed CAZy pipelines based on hidden Markov model profiles and sequence similarity26. To estimate CAZyme gene abundance, metagenomic reads were mapped to MAG gene sets using Kallisto v.0.46.1 with quant function53 and normalized abundance was expressed as transcripts per million (TPM).
Phylogenetic analysis and metabolic reconstruction
The phylogenetic profile of recovered MAGs was reconstructed using UBCG54 v.3.0, involving marker gene identification, multiple sequence alignment refinement and concatenation, and phylogeny reconstruction using Mafft55 v.7.487 and RAxML56 v.8.2.12. The resulting tree was visualized using the iTOL57 web tool v.6.9.1. Metabolic pathways in the recovered MAGs were reconstructed using gapseq58 v.1.1 and KEGG Orthology annotations. Enzyme commission (EC) numbers were assigned using KOFAMscan59 v.1.3.0 (e < 1 × 10−5). Proteins annotated as CAZymes but lacking an EC annotation had their EC transferred from characterized CAZymes of the same family recovered from the CAZy database using DIAMOND60 v.2.0.14.152. Pathway abundance was estimated by aligning metagenomic reads to binned sequences with bowtie2 (ref. 61) v.2.4.5 and calculating bin TPMs (SAMtools62 v.1.15.1). Each pathway predicted in a bin was assigned to its TPM, with total pathway abundance being the sum across relevant bins.
Pipeline for enzyme discovery from microbial dark matter
In silico protein selection approach
Protein sequences from the lignocellulolytic MAG ‘Ca. Telluricellulosum braziliensis’ belonging to the recently discovered and uncharacterized bacterial phylum UBP4 (GTDB database) were initially retrieved. Sequences lacking CAZy annotation were further analysed using HHpred63 v.3.3 and those exhibiting remote homology (10–25% sequence similarity) to proteins involved in carbohydrate breakdown and metabolism were selected for heterologous expression and biochemical assays (Supplementary Table 3).
Gene synthesis, heterologous expression and purification
The eight selected sequences were codon optimized for E. coli expression and synthesized with an N-terminal 6×His-tag. Next, E. coli BL21(DE3) cells were transformed with the target genes in pET-28a(+) expression vectors. Transformants were grown in Luria–Bertani (LB) medium (0.5% (w/v) yeast extract, 1% (w/v) tryptone and 1% (w/v) sodium chloride) at 37 °C to an optical density at 600 nm (OD600 nm) of approximately 0.8. Protein expression was induced with 0.4 mM isopropyl β-d-1-thiogalactopyranoside (IPTG) (Sigma Aldrich) at 18 °C for 16 h. Cells were collected by centrifugation (13,000g, 15 min, 4 °C) and resuspended in lysis buffer (20 mM sodium phosphate, pH 7.4, 300 mM NaCl, 5 mM imidazole, 1 mM phenylmethylsulfonyl fluoride (PMSF), 25 U ml–1 Turbo nuclease, 0.1 mg ml–1 lysozyme, 1.2 mg ml–1 deoxycholic acid). The lysed sample was then centrifuged at 21,000g for 40 min (4 °C). Soluble protein lysates were loaded onto a 5-ml HiTrap Chelating HP column (GE Healthcare) and the 6×His-tag target proteins were eluted using an imidazole gradient (up to 0.5 M). Further purification was achieved through size-exclusion chromatography on a HiLoad 16/600 Superdex 75/200 pg column (Cytiva) equilibrated with 20 mM sodium phosphate (pH 7.4) and 150 mM NaCl. Protein purity was assessed by SDS–PAGE and protein concentration were determined by measuring absorbance at 280 nm using the calculated extinction coefficient (ε280 nm) for each sequence.
Activity screening assays
Recombinant proteins (Supplementary Table 3) were screened for activity against a broad panel of substrates, including polysaccharides, oligosaccharides and synthetic p-nitrophenol derivatives (Supplementary Table 7). Assays were conducted by incubating the purified protein with the substrate (0.5% (w/v) for polysaccharides or 1 mM for oligosaccharides and synthetic substrates) in 50 mM sodium acetate buffer (pH 5.0) at 40 °C for up to 24 h. Enzymatic activity on polysaccharides was assessed by quantifying reducing sugar release using the 3,5-dinitrosalicylic acid (DNS) method64. For synthetic substrates, activity was determined by monitoring the release of p-nitrophenol at 405 nm. Enzyme assays consist of at least three independent experiments.
Saccharification boosting screening
To assess the potential of purified proteins to enhance the saccharification efficiency of the T. reesei Br_TrR03 cellulase cocktail27,28, we conducted saccharification experiments using steam-exploded sugarcane bagasse as described in the ‘Lignocellulosic biomass pretreatment’ section. In the screening phase, reactions were performed with 5% (w/v) total solids and an enzyme cocktail dosage of 1 mg g–1 of pretreated bagasse. Reactions were incubated for 24 h at 40 °C in 50 mM sodium acetate buffer (pH 5.0) using a combi-D24 hybridization incubator (FinePCR). The boosting effect was assessed by determining the total reducing sugar released by the DNS method64. Saccharification assays consist of at least three independent experiments.
Trichoderma enzyme cocktail production for screening assays
The T. reesei Br_TrR03 strain was cultivated in a BioFlo/CelliGen 115 system (Eppendorf) to produce the enzyme cocktail used in the saccharification boosting screening28. The cultivation medium contained 20 g l–1 (NH4)2SO4, 1.0 ml l–1 J647 antifoaming agent (Struktol), 20 g l–1 whole yeast cells (dry mass) and 50 g l–1 total reducing sugars (TRS) from sugarcane molasses. Bioreactors were initialized with 1.0 l of medium, including a 10% (v/v) inoculum prepared from fungal spores. The cultivation process was controlled as follows: pH was maintained at 4.5 ± 0.5 using 2 M phosphoric acid and 10% (w/v) ammonium hydroxide; temperature was kept at 28.0 °C; aeration was provided at 0.7 standard litres per minute (slpm) compressed air; and an agitation cascade (400–1,000 rpm) was used to ensure that the dissolved oxygen remained above 20%. From 25 h onwards, a sugarcane molasses solution (approximately 350 g kg–1 TRS) with 1.0 ml l–1 antifoaming agent was fed at 1.3 g TRS kg–1 h–1 (based on instant bioreactor mass) until 1 h before collection, creating a non-linear feeding profile. Samples were collected every 24 h, centrifuged and the supernatants stored at −20 °C. The final fermentation broth was used for saccharification experiments, protein quantification (Lowry method, BSA standard)65 and enzymatic activity analysis.
Lignocellulosic biomass pretreatment
Sugarcane bagasse and eucalyptus residues were pretreated at the LNBR pilot plant using a SuPR 2G reactor (AdvanceBio Systems). Sugarcane bagasse underwent steam explosion using 0.5% (v/v) sulfuric acid at 140 °C for 15 min, followed by centrifugation (1,610g, 20 min) to separate the C5 stream. Eucalyptus residue was first subjected to alkaline deacetylation with 0.4% (w/w) NaOH at 70 °C for 60 min, then steam exploded with 0.25% (v/v) sulfuric acid at 190 °C for 3 min and the C5 stream was separated. The resulting pretreated sugarcane bagasse contained 53.4% cellulose, 33.3% lignin and 5.8% hemicellulose, whereas the pretreated eucalyptus residue consisted of 61.5% cellulose, 33.8% lignin and 3.3% hemicellulose.
Production and purification of CelOCE, variants and other enzymes
Site-directed mutagenesis
CelOCE variants (CelOCE(Δ2–5) and CelOCE(F33A)) were generated using inverse PCR. Primers were designed with complementary sequences longer than 15 nucleotides and a Tm of 50 °C (see Supplementary Table 17 for primer sequences). PCR amplicons were circularized using Gibson Assembly66. The resulting plasmids were transformed into E. coli BL21(DE3) cells, and variant proteins were expressed and purified following the same protocol as for the wild-type enzyme. All mutations were confirmed by Sanger sequencing.
Preparative expression and purification of CelOCE and variants
CelOCE and its variants were overexpressed in E. coli BL21(DE3) using the pET-28a(+) expression vector without a 6×His-tag. Following the same expression protocol as described in the ‘Gene synthesis, heterologous expression and purification’ section, soluble protein lysates were subjected to ion exchange chromatography on a 5-ml HiTrap Q-FF column (Cytiva). CelOCE was eluted with a saline gradient up to 0.5 M, followed by size-exclusion chromatography using a HiLoad 16/600 Superdex 75 pg column (Cytiva) equilibrated with 20 mM sodium phosphate buffer (pH 7.4), containing 150 mM NaCl. To remove excess salt and prevent precipitation, proteins were buffer-exchanged using a HiTrap Desalting 5 ml column (Cytiva). CelOCE was then doped with sub-equimolar CuSO4 to enhance stability, followed by removal of excess copper using a VIVASPIN TURBO concentrator (10 kDa molecular weight cut-off (MWCO), Sartorius). To rule out interference by contaminations, three independent enzyme preparations were included, each starting from fresh transformations and including previously unused purification columns (Supplementary Fig. 28). In all three preparations, the activity on Avicel PH-101 (Sigma Aldrich) was validated by detecting cellobionic acid (Supplementary Fig. 29).
Size-exclusion chromatography with multi-angle light scattering
Size-exclusion chromatography with multi-angle light scattering (SEC–MALS) experiments were performed to determine the molecular mass and oligomeric state of CelOCE and its variants. In brief, 100 µl of purified protein samples were injected into a Superdex 200 (10/300) analytical size-exclusion column (Cytiva) connected to a high-performance liquid chromatography (HPLC) 1260 Infinity II system (Agilent). The column was equilibrated with 20 mM HEPES buffer (pH 7.4) containing 150 mM NaCl. Elution was monitored using a DAWN8 eight-angle static light scattering detector and an Optilab refractive index monitor (Wyatt Technology). Data acquisition and molecular mass calculations for CelOCE and its variants were performed using ASTRA v.8.1.2 software (Wyatt Technology).
BacB production and purification
The Bacillus subtilis bacilysin biosynthesis protein (BacB, PDB: 3H7J) sequence was codon optimized for E. coli expression, synthesized with an N-terminal 6×His-tag and subcloned into the pET-28a(+) vector. Next, E. coli BL21(DE3) cells containing the plasmid were grown in LB medium at 37 °C to an OD600 of approximately 0.8, then induced with 0.4 mM IPTG at 18 °C for 16 h. Cells were collected by centrifugation (13,000g, 15 min, 4 °C), lysed and centrifuged as described in the ‘Gene synthesis, heterologous expression and purification’ section. Recombinant BacB was purified using nickel-affinity and size-exclusion chromatography, using the same protocols and conditions outlined in the ‘Gene synthesis, heterologous expression and purification’ section.
KdgF production and purification
The Yersinia enterocolitica subsp. enterocolitica 8081 uronate metabolism protein (KdgF, PDB: 5FPX) sequence was codon optimized for E. coli expression and synthesized with an N-terminal 6×His-tag. The gene was subcloned into the pET-28a(+) expression vector. Expression and purification of KdgF followed the same protocol outlined in the ‘Gene synthesis, heterologous expression and purification’ section.
Cel5A production and purification
The B. subtilis endo-β-1,4-glucanase (Cel5A, GH5_2) was produced as described previously39. In brief, Cel5A was expressed in BL21(DE3)slyD– cells in LB medium at 37 °C for 4 h after induction with 0.5 mM IPTG. Collected cells were resuspended in lysis buffer (50 mM sodium phosphate, pH 7.4, 100 mM NaCl, 1 mM PMSF, 5 mM benzamidine), then lysed with lysozyme (80 μg ml–1, 30 min, on ice) and sonication. The lysate was centrifuged (10,000g, 30 min) and the supernatant was loaded onto a 5 ml HiTrap Chelating column (GE Healthcare) at 1 ml min–1. Proteins were eluted with a 0–500 mM imidazole gradient. Further purification was achieved using a 5 ml HiTrap SP HP column (Cytiva) with a 0–1 M NaCl gradient at 1 ml min–1. Size-exclusion chromatography was performed on a Superdex 75 16/60 column (Cytiva) equilibrated with 50 mM sodium phosphate, pH 7.4, 150 mM NaCl.
Cel45A production and purification
The Thermothielavioides terrestris endo-β-1,4-glucanase (Cel45A, GH45_1)40 was synthesized with an N-terminal 6×His-tag and subcloned into the pET-28a(+) vector. E. coli BL21(DE3) SHuffle cells containing the plasmid were grown in LB medium at 37 °C to an OD600 of 0.8. Cel45A expression was then induced with 0.4 mM IPTG at 18 °C and 180 rpm for 16 h. Collected cells were lysed by resuspension in lysis buffer containing sodium deoxycholate (60 mg l–1 of culture), lysozyme (20 mg l–1 of culture), 1 mM PMSF and DNase (20 µg ml–1) in buffer (20 mM sodium phosphate, pH 7.4, 300 mM NaCl, 5 mM imidazole). After incubation on ice for 1 h with gentle agitation, the lysate was centrifuged (21,000g, 45 min). The supernatant was subjected to nickel-affinity chromatography on a 5 ml HiTrap Chelating column (Cytiva), eluting the 6×His-tag protein with a 0–500 mM imidazole gradient. Final purification was achieved by size-exclusion chromatography on a HiLoad 16/600 Superdex 200 column (Cytiva) equilibrated with 20 mM sodium phosphate (pH 7.4) and 150 mM NaCl.
Cel7A production and purification
The cellobiohydrolase from T. reesei (Cel7A, GH7) was purified from the fungus secretome as described previously41. The T. reesei Br_TrR03 secretome was obtained as outlined in the ‘Trichoderma enzyme cocktail production for screening assays’ section. The secretome solution was vacuum-filtered through Miracloth (EMD Biosciences) using a 0.45-μm PES membrane and concentrated by tangential ultrafiltration (10 kDa MWCO). Buffer exchange was performed with 20 mM Bis-Tris (pH 6.5) to remove low-molecular-mass contaminants, followed by another filtration step. The filtrate was adjusted to 1.5 M (NH4)2SO4 and loaded onto a 26/10 Phenyl Sepharose Fast Flow column. Unbound material was washed off with 80% of 20 mM Bis-Tris pH 6.5 containing 2 M (NH4)2SO4, followed by elution with a descending gradient (80% to 0%) over eight column volumes. Active fractions were identified using a pNP-lactose activity assay (2 mM pNPL, 50 mM acetate pH 5.0, 30 min, 45 °C). These fractions were pooled, concentrated and desalted into 20 mM Bis-Tris (pH 6.5) using Superdex 25 HiPrep columns. The desalted protein was then loaded onto a Source 15Q 10/100 anion-exchange column and eluted with a 0–50% gradient of 20 mM Bis-Tris pH 6.5 containing 1 M NaCl over 30 column volumes. Active fractions were identified by pNP-lactose activity. The final purification step involved size-exclusion chromatography on a Superdex 75 16/60 column using 20 mM acetate buffer (pH 5.0) with 100 mM NaCl.
LPMO production and purification
The LPMOs TtAA9J from Thermothelomyces thermophilus, LsAA9A from L. similis, PaAA9E from Podospora anserina and NcAA9C from Neurospora crassa were expressed in Komagataella phaffii X-33 using the pPICZα vector and their native signal peptides. Cells were cultured in YPD medium (1% (w/v) yeast extract, 2% (w/v) peptone, 2% (w/v) glucose) at 30 °C and 200 rpm until glucose depletion. Protein expression was induced by adding 1% (v/v) methanol every 24 h for 72 h. The supernatants, obtained after centrifugation (13,000g, 15 min), were concentrated and buffer-exchanged into 20 mM Tris-HCl (pH 7.0) using a 10 kDa MWCO hollow fibre cartridge coupled to a tangential flow filtration system (GE Healthcare). The concentrates were applied to a DEAE-Sepharose XK 16/100 column (Cytiva) and eluted with a 0–1 M NaCl gradient, except for NcAA9C, which was applied to a CM-Sepharose XK 16/100 column (Cytiva) and eluted with a 0–1 M NaCl gradient. LPMO-containing fractions were pooled, concentrated and incubated with CuSO4 (3:1 molar ratio) on ice for 1 h. After centrifugation (20,000g, 10 min), the samples were further purified on a HiLoad 16/600 Superdex 75 pg column (Cytiva) equilibrated with 20 mM Tris-HCl (pH 7.0) and 150 mM NaCl. The final LPMO fractions were pooled, concentrated and stored at 4 °C.
Sample purity and quantification
Protein purity was assessed by SDS–PAGE (10%, w/v), followed by staining with Imperial Protein Stain (Thermo Fisher Scientific). Molecular mass under denaturing conditions was estimated using a PageRuler Prestained Protein Ladder (Thermo Fisher Scientific). Protein concentrations were determined either by the Bradford assay (Bio-Rad) or by measuring absorbance at 280 nm and using the calculated extinction coefficient (ε280) for each protein sequence.
Cellulose interaction assays
Qualitative cellulose binding assay
The cellulose binding capacity of CelOCE was evaluated qualitatively as described previously67. In brief, 80 µg of CelOCE was incubated with 5% (w/v) Avicel in 50 mM sodium acetate buffer (pH 5.2) for 24 h at 4 °C with gentle agitation. The final reaction volume was 200 µl, and incubations were performed in the presence or absence of 1 mM ASC. After incubation, insoluble cellulose was pelleted by centrifugation at 13,000g for 2 min. The supernatant, containing unbound proteins, was carefully removed. The Avicel pellet was washed twice by resuspension in buffer and centrifugation. The washed pellet was then resuspended in 200 µl of SDS-loading buffer (without dye) and heated at 95 °C for 10 min. Both the soluble (supernatant) and insoluble (pellet) fractions were analysed by SDS–PAGE using a 4–20% gradient gel.
Quantitative cellulose binding assay
The binding of CelOCE, with or without reductant (ASC), to microcrystalline (Avicel) and amorphous (PASC) cellulose was quantified as described previously29. In brief, reactions containing 5% (w/v) Avicel or 0.2% (w/v) PASC in 50 mM sodium acetate buffer (pH 5.0) were incubated with 0.1–1.5 mg ml–1 CelOCE in a final volume of 500 µl. BSA served as a negative control in the same concentration range. Reactions were incubated for 16 h at 4 °C with constant agitation using a Revolver rotator (Labnet International). Insoluble substrate with bound CelOCE was removed by centrifugation at 20,000g for 3 min. The concentration of free enzyme (biological unit) in the supernatant was determined spectrophotometrically at 280 nm. For reactions containing 1 mM ASC, 240 mM phosphoric acid was also added, and absorbance measurements were taken at 290 nm (ref. 29). Standard curves for CelOCE and BSA in the presence of 1 mM ASC were obtained at 290 nm. The experimental data were fitted using the Langmuir–Freundlich model68. Data are the mean ± s.d. of three independent experiments.
Competitive binding assay
For the BSA blocking assay, reactions containing 5% (w/v) Avicel in 50 mM sodium acetate buffer (pH 5.0) were incubated with 0.2 mg ml–1 CelOCE and/or 0.2 mg ml–1 BSA. CelOCE was tested in the absence or presence of reductant (1 mM ASC) forms. The final reaction volume was 500 µl. Incubations were performed at 4 °C with constant agitation using a Revolver rotator (Labnet International) for 2, 8, 16 and 24 h. The insoluble substrate with bound proteins was removed by centrifugation at 20,000g for 3 min. The concentration of free enzymes in the supernatant was measured using the Bradford method69. Data are the mean ± s.d. from three independent experiments.
XPS and FTIR spectroscopy
To further investigate the interaction between CelOCE and microcrystalline cellulose or bleached cellulose fibres (extracted from sugarcane bagasse), cellulose dispersions (0.4% w/v) were prepared in 20 mM sodium phosphate buffer (pH 5.4) containing 150 mM NaCl and soaked overnight. Enzyme and reductant (ASC) were then added to achieve final concentrations of 1 mM and 480 mg g–1 cellulose, respectively. The reaction mixtures were incubated at 40 °C for 72 h with orbital shaking. Two distinct washing procedures were evaluated. In the first, cellulose pellets were collected by centrifugation at 13,000g for 15 min and then subjected to multiple cycles of washing and centrifugation with deionized water until the conductivity of the wash solution approached that of pure water. The second procedure involved washing the cellulose pellets (200 mg) ten times with 30–40 ml of a 1 mM aqueous SDS solution at 40 °C, followed by rinsing with water to remove residual SDS. The surface elemental composition of the cellulose samples was acquired using a K-Alpha instrument (Thermo Fisher Scientific) with energy resolution of approximately 1 eV and the Avantage v.5.9931 software. Chemical characterization of the samples was performed using a FT-IR Spectrometer (PerkinElmer) and the Spectrum v.10 control software. Spectra were collected at a resolution of 4 cm−1, in the range of 4,000 to 700 cm−1, with a total of 128 scans. Data were analysed using the OriginPro v.2023b software (OriginLab).
Enzyme and complementary assays
Analysis of oxidized and non-oxidized oligosaccharides
Monosaccharides, cellooligosaccharides (degree of polymerization (DP) from 2 to 6), polysaccharides and their corresponding aldonic acid forms resulting from the cleavage of Avicel, PASC, sugarcane bagasse, eucalyptus, chitin, mannan, xylan, xyloglucan, arabinoxylan, β-glucan, laminarin, lichenan, starch and pectin were analysed by high-performance HPAEC–PAD using a Dionex ICS6000 system (Thermo Fisher Scientific) and the Chromeleon v.7.3 software. Reactions containing 0.1% (w/v) of the distinct polysaccharides, 50 mM sodium acetate (pH 5.0), 1 µM of the tested enzyme (CelOCE, KdgF or BacB) and 1 mM reductant were incubated for 16 h at 37 °C. All samples were homogenized by vortexing, filtered through a Millex 0.22-µm syringe filter. Then, 5 µl of each reaction were analysed on an HPAEC–PAD system equipped with a 2 × 50 mm CarboPac PA1 guard column and a 2 × 250 mm CarboPac PA1 analytical column (Thermo Fisher Scientific) maintained at 30 °C. The analysis was performed at a flow rate of 0.1 ml min–1. The column was equilibrated with 0.1 M NaOH (eluent A) and bound oligosaccharides were eluted using a gradient of 1 M sodium acetate (eluent B) as follows: 0–10% B (linear) over 10 min, 10–30% B (linear) over 25 min, 30–100% B (exponential) over 5 min, 100–0% B (linear) over 1 min and 0% B for 9 min for re-equilibration. Electrochemical detection was carried out using a gold working electrode and an Ag/AgCl pH reference electrode. Soluble cellooligosaccharides (DP 2–6, Megazyme) were used as standards. Corresponding C1-oxidized standards (DP 2–6) were produced by treating non-oxidized cellooligosaccharides with the cellobiose dehydrogenase from T. thermophilus (TtCDH, XM003664495)70. To enable quantification of cellobionic acid from all reactions, a calibration curve was generated using varying concentrations of cellobionic acid standard. Each assay consisted of at least three independent experiments. Data were analysed using the OriginPro v.2023b software (OriginLab).
Cellobionic acid detection by liquid chromatography coupled to mass spectrometry
Cellobionic acid generated by CelOCE activity on Avicel in the presence of ASC was identified using liquid chromatography coupled to mass spectrometry (LC–MS). An ACQUITY Premier Ultra Performance Liquid Chromatograph (UPLC) coupled to a Synapt XS mass spectrometer (Waters) was used for the analysis. Enzyme reactions were diluted 20-fold with a 1:1 (v/v) mixture of deionized water and acetonitrile. Then, 1 µl of the diluted sample was injected onto a Z-HILIC column (1.7 µm, 95 Å pore, 2.1 mm × 150 mm, Waters). The mobile phases consisted of 30% (v/v) acetonitrile (A) and 95% (v/v) acetonitrile (B), both containing 0.1% ammonium hydroxide. The elution gradient was as follows: initial, 85% B for 5 min; linear gradient, 85% B to 45% B; isocratic, 45% B for 5 min; return to initial, 45% B to 85% B and re-equilibration, 85% B for 5 min. The flow rate was maintained at 0.2 ml min–1 throughout the analysis. The mass spectrometer was operated in negative ion mode with the following settings: capillary voltage: 2 kV, cone voltage: 25 V, m/z range: 100–1,500, scan cycle: 0.5 s and collision energy: 4 V. Lock mass correction was applied every 30 s using the peptide standard leucine-enkephalin at 100 pg µl–1 (Waters) with a mass window of 0.5 Da.
Anaerobic experiments
As previously described for LPMOs38, a 1 g l–1 suspension of Avicel PH-101 (Sigma Aldrich) in 50 mM sodium acetate buffer (pH 5.0) was prepared in a reaction glass vial and deoxygenated by flushing with nitrogen gas for 5 min under magnetic stirring. Solutions of 50 mM ASC, 10 mM hydrogen peroxide and 50 µM CelOCE, along with a water control, were deoxygenated using a Schlenk line (three cycles of 10 min vacuum and 2 min N2). All solutions were then placed in a Whitley DG250 anaerobic workstation for 16 h to ensure complete removal of oxygen. To initiate the reactions, 1 µM CelOCE was added to both anaerobic and aerobic Avicel suspensions. After 20 min of incubation, 100 µM hydrogen peroxide was added to half of the anaerobic reactions, whereas the remaining anaerobic reactions and all aerobic reactions received an equivalent volume of water. CelOCE activity was then triggered in all reaction mixtures by the addition of 1 mM ASC (final reaction volume: 600 µl in 2 ml tubes). Time-dependent reactions were conducted at 0, 1, 2, 3, 4, 8 and 16 h. Aerobic reactions served as positive controls to verify that the treatment of the stock solutions did not compromise reactant integrity. Reactions were terminated by boiling, followed by centrifugation at 21,000g for 15 min. The resulting supernatants were analysed by HPAEC–PAD. Each assay consisted of at least three independent experiments.
Hydrogen peroxide detection and quantification
Hydrogen peroxide production was evaluated using an assay adapted from a previous study71 originally designed for LPMOs. To measure the hydrogen peroxide production rate in the absence of a polysaccharide substrate, 1 µM CelOCE was mixed with 50 µM Amplex Red reagent (Invitrogen) and 5 U ml–1 horseradish peroxidase type II (HRP, Sigma Aldrich), in 50 mM sodium phosphate buffer pH 6.0, in a 96-well microplate and incubated at 37 °C in a spectrophotometer. After 2 min, 1 mM ASC (Sigma Aldrich) was added to initiate the reaction. Absorbance at 563 nm was measured every 5 min for a total of 30 min. To measure the change in hydrogen peroxide production in the presence of increasing concentrations of polysaccharide, 1 µM enzyme (wild-type CelOCE, CelOCE(Δ2–5) or TtAA9J) was mixed with 50 mM sodium phosphate buffer pH 6.0 and 0–100 g l–1 Avicel. Then, 1 mM ASC was added to initiate the reaction (final volume of 200 µl in 2 ml tubes). Incubation was performed in a ThermoMixer C (Eppendorf) at 37 °C and 850 rpm for 16 h (wild-type and mutant CelOCE enzymes) or 2 h (TtAA9J). After incubation, the reaction tubes were centrifuged at 20,000g for 10 min to separate the substrate. Then, 50 µl of the supernatant was added to 50 µl of a mix containing Amplex Red, HRP and sodium phosphate buffer pH 6.0, and measured as in the previous assay. Hydrogen peroxide produced during the reactions was quantified using a standard curve. Each assay consisted of at least three independent experiments.
Peroxidase activity and thermal stability of AA9 enzymes
Peroxidase activity was measured following a protocol described previously72. In brief, 1 mM of 2,6-dimethoxyphenol was mixed with 100 µM of hydrogen peroxide and 2–4 µM of AA9 in 50 mM sodium citrate buffer pH 5.0. The reactions were incubated in a ThermoMixer C at 50 °C and 850 rpm for 10, 20 and 30 min to ensure the linear phase of the activity. Immediately, the samples were transferred to a 96-well plate and the absorbance was measured at 469 nm in a spectrophotometer (Infinite M200 Pro, Tecan) using the i-Control v.1.10.4.0 software. Only the linear phase was considered to calculate the specific activity of each AA9. The quantification was done using the extinction coefficient of coerulignone (ɛ469 nm = 53,200 M−1 cm−1). For the thermal stability measurements, the stock solutions of 40 µM of each AA9 enzyme were incubated in a ThermoMixer C at 50 °C in 50 mM sodium citrate pH 5.0 for 6, 12, 24, 48 and 72 h. The peroxidase activity was measured at 30 °C in 50 mM Tris-HCl pH 7.5 for 15 min incubated in a spectrophotometer (Infinite M200 Pro, Tecan).
Complementary assays
Saccharification experiments were conducted to evaluate the ability of CelOCE to enhance the efficiency of the T. reesei Br_TrR03 enzyme cocktail27,28. Avicel, PASC and steam-exploded sugarcane bagasse were used as substrates. Reactions were performed in 2 ml microtubes with a final volume of 1 ml, using total solid contents ranging from 0.1 to 5% (w/v). Each reaction contained 50 mM sodium acetate buffer (pH 5.0), 2 µg enzyme cocktail and 50 µg CelOCE. Whereas ASC (1 mM) was included for Avicel and PASC reactions, it was excluded from those using sugarcane bagasse. Reactions were incubated at 40 °C for 24–72 h in a combi-D24 hybridization incubator (FinePCR), following the same saccharification assay protocol used for the enzyme cocktail. The cooperative effect of CelOCE with individual purified endo- and exo-enzymes was assessed using Avicel and PASC as substrates. Control reactions lacking enzyme, reductant (ASC) or both were also included. These reactions were incubated overnight at 37 °C and 850 rpm in a Thermomixer Comfort (Eppendorf). All reactions were terminated by boiling at 95 °C for 10 min, followed by centrifugation at 20,000g for 15 min. The supernatants were analysed by HPAEC–PAD and the boosting effect of CelOCE was determined by measuring the TRS released using the DNS method64. Complementary assays consisted of at least three independent experiments.
Turnover rate calculation
For a standard reaction, 0.1% (w/v) Avicel was incubated with 1 µM CelOCE and 1 mM ASC in 50 mM sodium acetate buffer (pH 5.0) at 37 °C for 0, 1, 2, 3, 4, 8 and 16 h. The apparent turnover rate was calculated on the basis of the linear correlation between cellobionic acid concentration (µM) and time (h) in the 1- to 4-h time frame, as described previously37.
Biophysical approaches for copper characterization
XRF and XAS
CelOCE in 20 mM HEPES buffer pH 7.4 (control) was incubated with 1 mM CuSO4 or with 1 mM NiSO4 (4 h) followed by the addition of 1 mM CuSO4 overnight. A final concentration of 1.4 mM CelOCE was loaded into a MicroRT Capillary (MiTeGen) for measurements. The XRF and XAS experiments were performed at the Extreme condition Methods of Analysis (EMA) beamline from the Brazilian Synchrotron Light Laboratory (LNLS/CNPEM) using a Vortex-ME4 detector (Hitachi). Data collection was performed using EMA control software and included energy steps of 0.5 keV and an acquisition time of 1 s per scan. The energy of the beamline was calibrated using the \({k}_{{a}_{1}}\) emission line and the first inflection point of a copper foil, while an ionization chamber monitored the incident beam intensity. The foil was positioned upstream of the sample and measured under the same conditions as CelOCE, with the first inflection point set to 9.074 keV. The spectra of all datasets were calibrated, normalized and merged using Athena73 software v.0.9.26.
EPR
CelOCE samples (0.5 mM) were prepared in 20 mM HEPES buffer (pH 6.5). EPR spectra were recorded on a Bruker Elexsys E580 spectrometer (Bruker) operating at X-band (9.14 GHz) at 100 K (BVT 3000 digital temperature controller) with the following acquisition parameters: modulation frequency, 100 kHz; modulation amplitude, 5 G; conversion time, 90 ms; sweep time, 92.1 s; and microwave power, 20 mW. Data acquisition was performed using the Xepr v.2.6b.119 software. EPR spin-Hamiltonian parameters were determined using a set of computational tools in two steps as follows. Initial parameter estimation: g and A tensors were estimated using laboratory-developed scripts in Python (SciPy/NumPy)74. The g‖ and A‖ values were inferred by analysing the singularities near the low-field edge of the spectrum (260–310 mT). Near the high-field edge (310–330 mT), the hyperfine splitting A⟂ was not resolved, so the average distance between the most intense peaks and shoulders was used to estimate the hyperfine couplings. The g⟂ value was determined from the central position between these peaks (Supplementary Table 9). Simulation and optimization: using these initial guesses, simulations were conducted with the Pepper module of the EasySpin 6.0.0 toolbox75, running in MATLAB software76 (MathWorks, v.9.13.0, R2022b). Diagonal components of the g and A tensors were allowed to vary independently in specified bounds during the fit. A global optimization was first performed using a genetic algorithm (25 generations), followed by local optimization with the Nelder–Mead downhill simplex algorithm. The final Hamiltonian parameters were obtained by second-order perturbation and exact diagonalization for the final simulations.
ITC
ITC experiments were conducted using a MicroCal PEAQ-ITC Automated system (Malvern Panalytical) and the PEAQ-ITC control v.1.50 software. CelOCE samples (100 µM) in 20 mM MES buffer (pH 6.5), treated with Chelex (Sigma-Aldrich) to remove trace metals, were placed in the reaction cell (200 µl volume). Either 1.0 mM CuCl2 or 1 mM cellobiose was loaded into the ITC syringe. Titrations were performed by injecting 2 µl aliquots into the reaction cell at 150 s intervals with a stirring speed of 500 rpm, for a total of 19 injections. ITC data were collected automatically using the MicroCal PEAQ-ITC automated control software and corrected for the heat of dilution by subtracting the heat generated by titrating the ligand into buffer alone. A single-site binding model was fitted to the data using the non-linear least-squares algorithm provided by the MicroCal PEAQ-ITC Automated analysis software. The fitting yielded the stoichiometry (n), dissociation constant (Kd) and enthalpy change (ΔH) of the reaction. Errors in ΔH, Kd and Gibbs free energy change (ΔG) were calculated as the s.d. for at least three independent experiments. Error in entropy change (ΔS) was determined through error propagation.
Structural and computational biology approaches
Crystallization, X-ray diffraction and structure determination and refinement
Crystals of CelOCE were obtained by vapour diffusion in solutions containing: 20% (w/v) polyethylene glycol (PEG) 8000, 3% (v/v) 2-methyl-2,4-pentanediol and 0.1 M imidazole, pH 6.5 (structure 1); 1.4 M trisodium citrate and 0.1 M HEPES buffer pH 7.5 (structure 2); or 20% PEG 6000, 0.1 M citric acid pH 3.0 and 5% glycerol (structure 3). The second crystal (structure 2) was cryoprotected in a solution containing the mother liquor added by 25% (v/v) glycerol for data collection. All crystals were collected and then flash cooled in liquid nitrogen. X-ray diffraction data were collected at 100 K on the MANACA beamline (LNLS-Sirius/CNPEM) using a Pilatus 2 M detector (Dectris) and the fine ϕ-slicing approach77. Diffraction data were obtained using the MxCuBE78 v.2 program. Three datasets, each comprising 3,600 images with a 0.1° rotation and 0.1 s exposure time, were collected. Data processing was performed using XDS79 (version of 30 June 2023, build 20230630). The structures were solved by molecular replacement with Phaser80 v.2.7.0, using a RoseTTAFold (v.1)-generated model81 as a search model. The initial model was refined using Phenix.Refine82 v.1.8.3 and manually adjusted in Coot83 v.0.8.9. The final model and metal coordination were verified using MolProbity84,85 v.4.5 and the CheckMyMetal86 v.2.1, respectively. Dimeric stability and interface analysis were assessed using PDBePISA87 v.1.52.
Molecular docking and computational simulations
The KVFinder88 v.1.1.1 software was used to identify the cavity corresponding to the CelOCE active site. A cellotetraose (C4) molecule, obtained from PDB entry 3WDY, was docked into the CelOCE structure using Autodock Vina72 v.1.1.2. Docking trials were performed in a 10 × 10 × 10 Å3 box centred on the copper region of the CelOCE monomer. To refine the ligand position in the active site, the docked structure was further relaxed, considering the functional dimer of CelOCE. In this set-up, one protomer was docked with cellotetraose, whereas the other remained ligand-free. Residue protonation states were assigned on the basis of a pH of 5.5, with H5 and H85 protonated at Nε, and H44, H46, H83 and H84 protonated at Nδ in both protomers. The system was solvated with TIP3P water molecules, neutralized with two sodium ions and minimized to eliminate steric clashes. Subsequently, the system was heated in four 1-ns steps to 300 K under the NVT ensemble, followed by a 10-ns equilibration under the NpT ensemble (T = 300 K, p = 1 bar). Position restraints were applied to protein, ligand and copper atoms during the initial minimization and heating steps. Distance restraints, maintaining the octahedral copper coordination with the N atoms of H44, H46 and H84, the O atom of Q50, and the O2 and O3 atoms of the second (−1) and third (+1) non-reducing end units of the ligand, respectively, were used throughout the remaining simulation steps. Simulations were performed using the Amber20 package, and structural and trajectory analyses were conducted with visual molecular dynamics (VMD)73. Protein structure images were generated with PyMOL v.2.3 (The PyMOL Molecular Graphics System).
Genetic engineering and characterization of T. reesei
Integration of CelOCE and LsAA9A into T. reesei Br_TrR03 strain
The T. reesei strain Br_TrR03 was engineered to express CelOCE and LsAA9A (an AA9 LPMO from L. similis) using a customized CRISPR–Cas9 approach as described in our previous work27. The genes encoding CelOCE and LsAA9A were optimized for codon usage in Trichoderma and integrated into the xyn4 and xyn5 loci, respectively. The 20-nucleotide protospacers designed to specifically target xyn4 (GCCAAACATACAGACTGAGT) and xyn5 (GCCTGCTCTCTGTCTACGGC) in T. reesei flanked by (5′-end) hammerhead (HH) and (3′-end) hepatitis delta virus (HDV) ribozyme sequences were inserted into the Bsp1407I-digested pTrCas9gRNA1 plasmid, which was further used in protoplast transformation. A markerless donor cassette was assembled in vivo by Saccharomyces cerevisiae, containing 1-kb flanking sequences homologous to regions upstream and 1-kb flanking sequences homologous to downstream targeted genome regions. The cassette was PCR amplified from the plasmid to generate linear DNA fragments, which were then used for fungal transformation. Obtained PCR products were purified and concentrated in a SpeedVac concentrator before use in transformation assays. The oligonucleotides used are listed in Supplementary Table 18. Then, T. reesei was transformed through protoplast-mediated transformation27,89. In each transformation event, 5 μg of the appropriate CRISPR–Cas9 vector and 5 μg of a linear DNA fragment for genomic integration were used. The genetic modifications to the genome of T. reesei strain Br_TrR03 were confirmed by PCR using different combinations of primers that anneal (upstream or downstream) of the targeted genome regions or internally to each integration cassette (Supplementary Table 18).
Shake-flask cultivation
Engineered T. reesei strains were grown in 250-ml Erlenmeyer flasks containing 50 ml of medium that comprised 20.0 g l–1 (NH4)2SO4, 20.0 g l–1 whole yeast cells (dry mass), 50 g l–1 of TRS from sugarcane molasses and pH adjusted to 4.8. The whole yeast cells were generated as described previously27. Sugarcane molasses (Mellaço de Cana) was diluted with water to a TRS concentration of approximately 350 g kg–1 and autoclaved separately before addition to the sterilized base medium. Each flask was inoculated with 0.3 ml of 107 spores per ml and cultivations were carried out in shaker incubators at 28 °C with 200 rpm for 5 days. After this period, samples were collected, centrifuged at 14,000g for 10 min at 4 °C and the supernatants were stored at −20 °C until analysis.
Bench-scale bioreactor cultivation
Bioreactor experiments were conducted using a BioFlo/CelliGen 115 system (Eppendorf) and water-jacketed 3.0 l vessels with a working volume of 1.0–1.7 l (ref. 28). The medium composition was identical to that used in shake-flask experiments, with the addition of 1.0 ml l–1 of J647 antifoaming agent (Struktol). The initial volume in the bioreactors was 1.0 l, including the 10% (v/v) inoculum. The inoculum was obtained by growing fungal spores in flasks, which were shaken as described above for four days at 28 °C and 200 rpm. The pH was maintained at 4.5 ± 0.5 using a solution of 2 M phosphoric acid and 10% (v/v) ammonium hydroxide. The temperature and aeration were maintained at 28.0 °C and 0.7 slpm compressed air, respectively. An agitation cascade (400–1,000 rpm) was used to ensure that the dissolved oxygen remained above 20%. A sugarcane molasses solution (approximately 350 g kg–1 of TRS) containing 1.0 ml l–1 of antifoaming agent was fed from 25 h of cultivation until 1 h before the end of the experiments at a feeding rate of 1.3 gTRS kg–1 h–1 in relation to the instant mass in the bioreactor, generating a non-linear feeding profile. During cultivation, samples were collected at regular intervals (24 h), centrifuged at 14,000g for 10 min at 4 °C and the supernatants stored at −20 °C for subsequent analysis. Final fermentation samples were obtained for hydrolysis experiments, protein quantification and enzymatic activity analysis.
Pilot plant scale bioreactor cultivation
The Trichoderma strain expressing CelOCE was cultivated in both 65-l (Bioflo 610, Eppendorf) and 300-l (Bioflo Pro 300 L, Eppendorf) pilot plant bioreactors. The composition of the molasses-based medium, as well as the cultivation and feeding conditions, were identical to those used in the bench-scale experiments. For the 65-l bioreactors, aeration was set to 13–18.5 slpm, dissolved oxygen 20%, pressure 5–10 psi and agitation 150–427 rpm. In the 300-l bioreactors, aeration was 84–120 slpm, dissolved oxygen 20%, pressure 5–10 psi and agitation 100–450 rpm. In both bioreactors, the inoculum volume was 10% of the total working volume.
Secretome analysis by mass spectrometry
Proteins (200 µg) from the secretome of engineered T. reesei strains were precipitated with acetone and resuspended in 500 µl of 25 mM ammonium bicarbonate buffer (pH 7.8) containing 50 mM dithiothreitol and 5% (w/v) sodium deoxycholate. After incubation at 60 °C for 30 min with agitation, samples were transferred to a 30 kDa MWCO Amicon filter (Merck Millipore) and centrifuged (14,000g, 20 min, 20 °C). The flowthrough was discarded and 450 µl of buffer containing 8 M urea was added, repeating this step twice. After the final centrifugation, 450 µl of 20 mM iodoacetamide in buffer was added for alkylation (45 min, in the dark at room temperature), followed by centrifugation and five desalting washes with buffer. Trypsin (1:30 w/w ratio) was added for overnight digestion at 37 °C. After centrifugation, peptides were collected in the flowthrough, washed twice with deionized water and dried. An internal standard (digested yeast alcohol dehydrogenase, P00330) was added, and 25 fmol of the peptide mixture was injected for LC–MS/MS analysis on a Synapt XS (Waters) coupled to an ACQUITY Premier UPLC. Data acquisitions were performed using the MassLynx v.4.2 program. Separation was performed on a peptide CSH column (1 mm × 100 mm, 1.7 µm, 130 Å) using a 4–55% gradient of over 103 min at 25 µl min–1. Data acquisition was in high-definition data-independent acquisition mode (50–2,000 m/z, 0.5 s scan cycle), with low collision energy at 6 eV and a ramp from 15 to 40 eV for elevated collision energy. Lock mass correction with the peptide standard leucine-enkephalin at 100 pg µl–1 was applied every 30 s with a mass window of 0.5 Da. Data processing was carried out in Progenesis 4.2 with a 1% false discovery rate, 20 ppm MS1 error tolerance and automatic MS2 error adjustment. Carbamidomethylation of cysteine was set as a fixed modification, methionine oxidation as variable and one missed trypsin cleavage was allowed. The T. reesei Br_TrR03 genome (NCBI: PRJNA1031947), including the CelOCE and LsAA9A sequences, served as the reference. Protein abundance was quantified using the top three peptides approach90.
Saccharification assays and secretome activity profiling
Enzymatic activities (β-glucosidase, β-xylosidase, endo-β-1,4-xylanase and CMCase) were determined as described previously27 and filter paper activity (FPase) was measured as described previously91. Saccharification reactions were conducted in 50 ml Nalgene Oak Ridge Centrifuge tubes using a combi-D24 hybridization incubator (FinePCR) at 50 °C with maximum rotation (level 9). Biomass pH was adjusted to 5.0 with 5 M NaOH. Each reaction contained 20.0 g total mass with 15% (w/w) final solids loaded in 100 mM acetate buffer (pH 5.0). Two stainless steel spheres (3.5 mm diameter, 1 g each) were added to ensure homogeneity. Distilled water replaced the enzyme in the blank reaction. Sampling and analysis followed an established protocol27. Three independent experiments were performed, and samples were collected every 24 h. Released sugars were quantified by HPLC (Agilent 1260 Infinity) with a refractive index detector. One-way ANOVA with post hoc Tukey’s honest significant difference tests were used to compare mean values of enzymatic activities and protein and sugar concentrations.
Inclusion and ethics statement
All researchers that fulfilled the authorship criteria by Nature Portfolio journals have been included in the author list. Their contributions were essential to the design, execution and interpretation of the study. The roles and responsibilities of each collaborator were clearly defined and mutually agreed ahead of the research. This research faced no severe restrictions or prohibitions in the setting of the researchers and was conducted in a manner that avoids causing stigmatization, incrimination, discrimination or personal risk to any parties involved. We support inclusive, diverse and equitable conduct of research.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Crystallographic data were deposited at the Protein Data Bank under accession codes 9BWF, 9BWH and 9BWI. Metagenomic data were deposited at the National Center for Biotechnology Information (NCBI) database under BioProject number PRJNA1103821. The remaining data are available in the main paper, Supplementary Information and source data. Any additional information is available upon request. Source data are provided with this paper.
Code availability
The code used for EPR spin-Hamiltonian parameter calculations is available at GitHub (https://github.com/colombarifm/celoce_epr)74.
Change history
21 March 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41586-025-08872-9
References
Cragg, S. M. et al. Lignocellulose degradation mechanisms across the tree of life. Curr. Opin. Chem. Biol. 29, 108–119 (2015).
Bomble, Y. J. et al. Lignocellulose deconstruction in the biosphere. Curr. Opin. Chem. Biol. 41, 61–70 (2017).
Lynd, L. R. et al. How biotech can transform biofuels. Nat. Biotechnol. 26, 169–172 (2008).
Chundawat, S. P. S., Beckham, G. T., Himmel, M. E. & Dale, B. E. Deconstruction of lignocellulosic biomass to fuels and chemicals. Annu. Rev. Chem. Biomol. Eng. 2, 121–145 (2011).
Quinlan, R. J. et al. Insights into the oxidative degradation of cellulose by a copper metalloenzyme that exploits biomass components. Proc. Natl Acad. Sci. USA 108, 15079–15084 (2011).
Vaaje-Kolstad, G. et al. An oxidative enzyme boosting the enzymatic conversion of recalcitrant polysaccharides. Science 330, 219–222 (2010).
Munzone, A., Eijsink, V. G. H., Berrin, J.-G. & Bissaro, B. Expanding the catalytic landscape of metalloenzymes with lytic polysaccharide monooxygenases. Nat. Rev. Chem. 8, 106–119 (2024).
Vuong, T. V. & Wilson, D. B. Glycoside hydrolases: catalytic base/nucleophile diversity. Biotechnol. Bioeng. 107, 195–205 (2010).
Rye, C. S. & Withers, S. G. Glycosidase mechanisms. Curr. Opin. Chem. Biol. 4, 573–580 (2000).
Bornscheuer, U., Buchholz, K. & Seibel, J. Enzymatic degradation of (ligno)cellulose. Angew. Chem. Int. Ed. 53, 10876–10893 (2014).
Sandgren, M. et al. The structure of a bacterial cellobiohydrolase: the catalytic core of the Thermobifida fusca family GH6 cellobiohydrolase Cel6B. J. Mol. Biol. 425, 622–635 (2013).
Kurašin, M. & Väljamäe, P. Processivity of cellobiohydrolases is limited by the substrate. J. Biol. Chem. 286, 169–177 (2011).
Vu, V. V., Beeson, W. T., Span, E. A., Farquhar, E. R. & Marletta, M. A. A family of starch-active polysaccharide monooxygenases. Proc. Natl Acad. Sci. USA 111, 13822–13827 (2014).
Couturier, M. et al. Lytic xylan oxidases from wood-decay fungi unlock biomass degradation. Nat. Chem. Biol. 14, 306–310 (2018).
Sabbadin, F. et al. An ancient family of lytic polysaccharide monooxygenases with roles in arthropod development and biomass digestion. Nat. Commun. 9, 756 (2018).
Filiatrault-Chastel, C. et al. AA16, a new lytic polysaccharide monooxygenase family identified in fungal secretomes. Biotechnol. Biofuels 12, 55 (2019).
Sabbadin, F. et al. Secreted pectin monooxygenases drive plant infection by pathogenic oomycetes. Science 373, 774–779 (2021).
Hemsworth, G. R., Henrissat, B., Davies, G. J. & Walton, P. H. Discovery and characterization of a new family of lytic polysaccharide monooxygenases. Nat. Chem. Biol. 10, 122–126 (2014).
Phillips, C. M., Beeson, W. T. IV, Cate, J. H. & Marletta, M. A. Cellobiose dehydrogenase and a copper-dependent polysaccharide monooxygenase potentiate cellulose degradation by Neurospora crassa. ACS Chem. Biol. 6, 1399–1406 (2011).
Bayer, E. A., Chanzy, H., Lamed, R. & Shoham, Y. Cellulose, cellulases and cellulosomes. Curr. Opin. Struct. Biol. 8, 548–557 (1998).
Artzi, L., Bayer, E. A. & Moraïs, S. Cellulosomes: bacterial nanomachines for dismantling plant polysaccharides. Nat. Rev. Microbiol. 15, 83–95 (2017).
Brunecky, R. et al. Revealing nature’s cellulase diversity: the digestion mechanism of Caldicellulosiruptor bescii CelA. Science 342, 1513–1516 (2013).
Parks, D. H. et al. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat. Microbiol. 2, 1533–1542 (2017).
Parks, D. H. et al. GTDB: an ongoing census of bacterial and archaeal diversity through a phylogenetically consistent, rank normalized and complete genome-based taxonomy. Nucleic Acids Res. 50, D785–D794 (2022).
Hedlund, B. P. et al. SeqCode: a nomenclatural code for prokaryotes described from sequence data. Nat. Microbiol. 7, 1702–1708 (2022).
Drula, E. et al. The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res. 50, D571–D577 (2022).
Fonseca, L. M., Parreiras, L. S. & Murakami, M. T. Rational engineering of the Trichoderma reesei RUT-C30 strain into an industrially relevant platform for cellulase production. Biotechnol. Biofuels 13, 93 (2020).
de Lima, E. A. et al. Development of an economically competitive Trichoderma-based platform for enzyme production: bioprocess optimization, pilot plant scale-up, techno-economic analysis and life cycle assessment. Bioresour. Technol. 364, 128019 (2022).
Kracher, D., Andlar, M., Furtmüller, P. G. & Ludwig, R. Active-site copper reduction promotes substrate binding of fungal lytic polysaccharide monooxygenase and reduces stability. J. Biol. Chem. 293, 1676–1687 (2018).
Yang, B. & Wyman, C. E. BSA treatment to enhance enzymatic hydrolysis of cellulose in lignin containing substrates. Biotechnol. Bioeng. 94, 611–617 (2006).
Rajavel, M., Mitra, A. & Gopal, B. Role of Bacillus subtilis BacB in the synthesis of bacilysin. J. Biol. Chem. 284, 31882–31892 (2009).
Hobbs, J. K. et al. KdgF, the missing link in the microbial metabolism of uronate sugars from pectin and alginate. Proc. Natl Acad. Sci. USA 113, 6188–6193 (2016).
Isaksen, I., Jana, S., Payne, C. M., Bissaro, B. & Røhr, Å. K. The rotamer of the second-sphere histidine in AA9 lytic polysaccharide monooxygenase is pH dependent. Biophys. J. 123, 1139–1151 (2024).
Gómez-Piñeiro, R. J. et al. Decoding the ambiguous electron paramagnetic resonance signals in the lytic polysaccharide monooxygenase from Photorhabdus luminescens. Inorg. Chem. 61, 8022–8035 (2022).
Haddad Momeni, M. et al. Discovery of fungal oligosaccharide-oxidising flavo-enzymes with previously unknown substrates, redox-activity profiles and interplay with LPMOs. Nat. Commun. 12, 2132 (2021).
Hemsworth, G. R. Revisiting the role of electron donors in lytic polysaccharide monooxygenase biochemistry. Essays Biochem. 67, 585–595 (2023).
Müller, G., Chylenski, P., Bissaro, B., Eijsink, V. G. H. & Horn, S. J. The impact of hydrogen peroxide supply on LPMO activity and overall saccharification efficiency of a commercial cellulase cocktail. Biotechnol. Biofuels 11, 209 (2018).
Bissaro, B. et al. Oxidative cleavage of polysaccharides by monocopper enzymes depends on H2O2. Nat. Chem. Biol. 13, 1123–1128 (2017).
Santos, C. R. et al. Dissecting structure–function–stability relationships of a thermostable GH5-CBM3 cellulase from Bacillus subtilis 168. Biochem. J. 441, 95–104 (2012).
Gao, J. et al. Characterization and crystal structure of a thermostable glycoside hydrolase family 45 1,4-β-endoglucanase from Thielavia terrestris. Enzyme Microb. Technol. 99, 32–37 (2017).
Linger, J. G. et al. A constitutive expression system for glycosyl hydrolase family 7 cellobiohydrolases in Hypocrea jecorina. Biotechnol. Biofuels 8, 45 (2015).
Dunwell, J. M., Purvis, A. & Khuri, S. Cupins: the most functionally diverse protein superfamily? Phytochemistry 65, 7–17 (2004).
Dunwell, J. M., Culham, A., Carter, C. E., Sosa-Aguirre, C. R. & Goodenough, P. W. Evolution of functional diversity in the cupin superfamily. Trends Biochem. Sci. 26, 740–746 (2001).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Cole, J. R. et al. Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42, D633–D642 (2014).
Menzel, P., Ng, K. L. & Krogh, A. Fast and sensitive taxonomic classification for metagenomics with Kaiju. Nat. Commun. 7, 11257 (2016).
Peng, Y., Leung, H. C. M., Yiu, S. M. & Chin, F. Y. L. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. MetaWRAP—a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Chklovski, A., Parks, D. H., Woodcroft, B. J. & Tyson, G. W. CheckM2: a rapid, scalable and accurate tool for assessing microbial genome quality using machine learning. Nat. Methods 20, 1203–1212 (2023).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk v2: memory friendly classification with the genome taxonomy database. Bioinformatics 38, 5315–5316 (2022).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Na, S.-I. et al. UBCG: Up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J. Microbiol. 56, 280–285 (2018).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
Zimmermann, J., Kaleta, C. & Waschina, S. gapseq: informed prediction of bacterial metabolic pathways and reconstruction of accurate metabolic models. Genome Biol. 22, 81 (2021).
Aramaki, T. et al. KofamKOALA: KEGG ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 36, 2251–2252 (2020).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Hildebrand, A., Remmert, M., Biegert, A. & Söding, J. Fast and accurate automatic structure prediction with HHpred. Proteins 77, 128–132 (2009).
Miller, G. L. Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 31, 426–428 (1959).
Lowry, O. H., Rosebrough, N. J., Farr, A. L. & Randall, R. J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193, 265–275 (1951).
Gibson, D. G. Enzymatic assembly of overlapping DNA fragments. Methods Enzymol. 498, 349–361 (2011).
Crouch, L. I., Labourel, A., Walton, P. H., Davies, G. J. & Gilbert, H. J. The contribution of non-catalytic carbohydrate binding modules to the activity of lytic polysaccharide monooxygenases. J. Biol. Chem. 291, 7439–7449 (2016).
Ayawei, N., Ebelegi, A. N. & Wankasi, D. Modelling and interpretation of adsorption isotherms. J. Chem. 2017, 3039817 (2017).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Oliva, B. et al. Recombinant cellobiose dehydrogenase from Thermothelomyces thermophilus: its functional characterization and applicability in cellobionic acid production. Bioresour. Technol. 402, 130763 (2024).
Kittl, R., Kracher, D., Burgstaller, D., Haltrich, D. & Ludwig, R. Production of four Neurospora crassa lytic polysaccharide monooxygenases in Pichia pastoris monitored by a fluorimetric assay. Biotechnol. Biofuels 5, 79 (2012).
Breslmayr, E. et al. A fast and sensitive activity assay for lytic polysaccharide monooxygenase. Biotechnol. Biofuels 11, 79 (2018).
Ravel, B. & Newville, M. ATHENA, ARTEMIS, HEPHAESTUS: data analysis for X-ray absorption spectroscopy using IFEFFIT. J. Synchrotron Radiat. 12, 537–541 (2005).
Colombari, F. M. colombarifm/celoce_epr: v1.0. Zenodo https://doi.org/10.5281/zenodo.14245344 (2024).
Stoll, S. & Schweiger, A. EasySpin, a comprehensive software package for spectral simulation and analysis in EPR. J. Magn. Reson. 178, 42–55 (2006).
MATLAB v.9.13.0 (R2022b) (The MathWorks, Inc., 2022); https://www.mathworks.com
Mueller, M., Wang, M. & Schulze-Briese, C. Optimal fine φ-slicing for single-photon-counting pixel detectors. Acta Crystallogr. D 68, 42–56 (2012).
Gabadinho, J. et al. MxCuBE: a synchrotron beamline control environment customized for macromolecular crystallography experiments. J. Synchrotron Radiat. 17, 700–707 (2010).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Williams, C. J. et al. MolProbity: more and better reference data for improved all-atom structure validation. Protein Sci. 27, 293–315 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Gucwa, M. et al. CMM—an enhanced platform for interactive validation of metal binding sites. Protein Sci. 32, e4525 (2023).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Guerra, J. V. S., Ribeiro-Filho, H. V., Pereira, J. G. C. & Lopes-de-Oliveira, P. S. KVFinder-web: a web-based application for detecting and characterizing biomolecular cavities. Nucleic Acids Res. 51, W289–W297 (2023).
Penttilä, M., Nevalainen, H., Rättö, M., Salminen, E. & Knowles, J. A versatile transformation system for the cellulolytic filamentous fungus Trichoderma reesei. Gene 61, 155–164 (1987).
Silva, J. C., Gorenstein, M. V., Li, G.-Z., Vissers, J. P. C. & Geromanos, S. J. Absolute quantification of proteins by LCMSE: a virtue of parallel MS acquisition. Mol. Cell. Proteomics 5, 144–156 (2006).
Ghose, T. K. Measurement of cellulase activities. Pure Appl. Chem. 59, 257–268 (1987).
Acknowledgements
We thank the Brazilian Center for Research in Energy and Materials (CNPEM), specifically the Brazilian Synchrotron Light Laboratory (LNLS) for the use of MANACÁ (proposal id: 20231934) and EMA (proposal id: 20221693 and 20222047) beamlines, the Biosciences National Laboratory (LNBio) for the automated crystallization of macromolecules (ROBOLAB), the Brazilian Biorenewables National Laboratory (LNBR) for the use of the high-performance sequencing, metabolomics, biophysics of macromolecules and development and scaling of bioprocess facilities, the National Laboratory for Scientific Computing (LNCC/MCTI) for the computing resources granted in the Santos Dumont (SDumont) supercomputer, E. E. D. Silva for his support with SEC–MALS and ITC measurements, R. Senen, E. S. Ribeiro, E. G. Silva, E. L. A. Silva, F. R. C. Souza, L. O. Romão, C. L. A. Menezes and E. L. Bernardi for their support in pilot plant cultivations, A. F. Lima and M. S. Costa for routine high-performance liquid chromatography analyses, M. P. Martins for her support in soil sample collection, C. C. C. Tonoli and G. L. Garrido for their support with crystallization experiments. Part of the work described was performed using services provided by the 3PE platform, a member of IBISBA-FR (https://doi.org/10.15454/08BX-VJ91; www.ibisba.fr), the French node of the European research infrastructure, EU-IBISBA (www.ibisba.eu). This work was supported in part by São Paulo Research Foundation (FAPESP) grants 21/04891-3 (M.T.M.) and 22/03059-5 (G.F.P.) and National Council for Scientific and Technological Development (CNPq) grant 305013/2020-3 (M.T.M.).
Author information
Authors and Affiliations
Contributions
C.A.S., R.Y.M., P.M.R.H., M.L.M., L.G.M., L.D.W. and S.G. performed and analysed biochemical data. C.A.S., M.A.B.M., E.A.A., C.R.S. and M.T.M. performed and analysed synchrotron data. C.R.F.T., P.T.R. and F.M. performed the T. reesei engineering experiments. E.A.L., N.R.B. and J.A.D. developed the bench-top fungal bioprocess and pilot plant bioreactors. D.A.A.P., J.M.J., R.S.A.S., C.B.C.S., V.L., N.T., B.H. and G.F.P. performed and analysed bioinformatics data. C.A.S., F.M.C., A.J.C.F. and M.T.M. performed and analysed EPR data. D.B.S. and J.S.B. performed and analysed XPS and FTIR data. C.A.S. and F.J.F. performed and analysed mass spectrometric data. C.A.S. and M.L.M. performed and analysed ITC data. M.A.B.M. and F.M.C. performed and analysed molecular docking and simulations. C.A.S., M.A.B.M., R.Y.M., P.M.R.H., B.H., B.B., J.-G.B., G.F.P. and M.T.M. wrote the manuscript. M.T.M. conceived the study, provided supervision and acquired funding. All authors discussed the results, reviewed the draft manuscript and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
C.A.S., F.M., E.A.L., G.F.P. and M.T.M. are named inventors on patent application number BR10202401483 filed by the Brazilian Center for Research in Energy and Materials, covering the use of the enzyme discovered in this study for biomass conversion and related biotechnological applications. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Kiyohiko Igarashi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Glycoside hydrolase (GH) annotation.
(a) Top-most abundant GH families identified in each metagenome sample. The normalized abundance of each GH family is compared between the data from Sugarcane Bagasse-covered Soil collected 20 cm below ground (SBS) and control soil sample collected from an adjacent not covered with sugarcane bagasse (Bulk). (b) Top-most abundant GH families identified in the uncultured ‘Candidatus Telluricellulosum braziliensis’ MAG. This panel provides a focused view of the GH repertoire potentially involved in plant cell wall degradation by this bacterium. (c) GH distribution in the recovered MAGs. The top 30 MAGs with the highest number of predicted GHs are shown. The heatmap color intensity indicates the relative abundance of each GH family within a given MAG, facilitating visual comparison of GH profiles across the different MAGs.
Extended Data Fig. 2 CelOCE interaction with microcrystalline cellulose.
(a) Fourier-transform infrared spectroscopy (FTIR) spectra of Avicel untreated or treated with CelOCE. The peaks at 1639 and 1520 cm−1 correspond to the –C = O and –N–H functional groups of CelOCE, respectively. (b) X-ray photoelectron spectroscopy (XPS) patterns for control (untreated) Avicel and Avicel treated with CelOCE, both before and after washing with anionic surfactant (SDS). The survey spectrum of CelOCE-treated cellulose fibers reveals a nitrogen peak (N1s), indicating the presence of N-O, C = N, and C-N groups introduced by the adsorption of CelOCE. The plotted spectra are representative curves from three independent experiments.
Extended Data Fig. 3 Distribution of CelOCE orthologs across bacterial and archaeal phyla.
Phylogenetic tree illustrates the distribution and potential evolutionary history of CelOCE-like proteins across diverse microbial taxa, encompassing 406 microbial species from 37 bacterial and archaeal phyla. The tree was constructed based on 402 complete genomes from the NCBI RefSeq database and 4 MAGs generated in this study (in red). The ‘Candidatus Telluricellulosum braziliensis’ MAG harboring the celOCE gene is denoted as SBS.bin.55 and indicated with a red asterisk. Phyla with fewer than two species (Thermosulfidibacterota, Thermodesulfobacteriota, Nanoarchaeota, Nitrospirota, and Calditrichota) are unlabeled, except when closely related to the CelOCE-containing MAG.
Extended Data Fig. 4 CelOCE structural properties.
(a) CelOCE adopts a compact jelly-roll fold, which consists of two anti-parallel β-sheets, one containing 6 strands (β-sheet A, including the following strands: β2-β4, β6, β9 and β11) and the other containing 4 strands (β-sheet B, including the following strands: β5, β7, β8 and β10). (b) Cartoon representation of the two CelOCE protomers, indicating the homodimeric arrangement in a back-to-back configuration where their active sites face opposite directions. Key interactions stabilizing the dimer interface are indicated, including M1-C77, A3-S75, K4-D74, and E26-E49. These dimeric interactions primarily involve the N-terminal β-strands (β1) and distances are shown in ångström.
Extended Data Fig. 5 Copper coordination sphere in CelOCE.
(a) When glycerol, a sugar mimetic, is present (structure 2), one water molecule in the copper coordination sphere is replaced by an oxygen atom from glycerol. This causes the remaining water molecule to shift to the equatorial plane, alongside H44, H46, and Q50. (b) Conformational change in copper coordinating histidine at acidic pH. Comparison of two CelOCE crystal structures highlights a conformational change in H44 that occurs under acidic pH conditions. The structure obtained under acidic crystallization condition is shown in light orange (structure 3), while the CelOCE structure with non-flipped H44 is shown in white (structure 2) for comparison. The copper-coordinating residues are represented as sticks, the copper atom as an orange sphere, water molecules as red spheres and glycerol molecules as spheres/sticks following the protein color scheme. Distance measurements (in ångström) pertain to the structure obtained under acidic conditions, specifically showing the altered position of H44 relative to the copper.
Extended Data Fig. 6 Active site of CelOCE.
(a) Surface representation of CelOCE, emphasizing the buried nature of the catalytic copper, located approximately 5 Å from the protein surface. The copper is shown as an orange sphere. (b) Cartoon representation of CelOCE with the docked cellotetraose, demonstrating room for the accommodation of a disaccharide within the active-site pocket (−2 and −1 subsites). (c) Detailed view of the CelOCE active site, demonstrating its stereochemical compatibility with glucosyl moieties. The docked cellooligosaccharide shows the non-reducing end glucosyl (−2) residue anchored primarily by interactions with E96 and Q50, while the −1 glucosyl residue stacks against F33, productively positioning the C1 atom for oxidative attack by the catalytic copper.
Extended Data Fig. 7 The role of electron donor and co-substrate for catalysis.
(a) The role of an electron donor (ASC) in product release under aerobic conditions for CelOCE activity. (b) Cellobionic acid production by CelOCE under aerobic and anaerobic conditions in the presence of ASC as a reductant. (c) Stoichiometric ratio for co-substrate and product formation. The reactions containing 50 mM sodium acetate pH 5.0, 0.1% (w/v) Avicel, 1 mM ASC, 5 or 10 µM H2O2, and 1 µM CelOCE were incubated at 37 °C for 16 h under anaerobic conditions. For a, b and c, results are expressed as mean ± standard deviation from three independent experiments. Statistical significance was determined by one-way ANOVA with Tukey’s post hoc test (***p < 0.001).
Extended Data Fig. 8 Putative modes of substrate recognition and regioselectivity.
Cellobionic acid was the only product detected, supporting C1-carbon regioselectivity. Although molecular docking studies indicate that the enzyme likely recognizes the non-reducing end of cellulose, we cannot rule out the possibility of reducing end recognition since subsequent catalytic cycles on the same cellulose chain could also generate cellobionic acid. NR, non-reducing end; R, reducing end; Ox, oxidized.
Extended Data Fig. 9 Cellobionic acid production and saccharification efficiency by the secretome of engineered strains.
The secretome of the parental (Br_TrR03) and engineered (CelOCE::Br_TrR03) T. reesei strains were assayed on technical substrates. (a, b) Cellobionic acid production using pretreated sugarcane bagasse. Saccharification efficiency on pretreated eucalyptus chips including the comparison with Cellic® CTec2 and the (LsAA9A::Br_TrR03) strain secretome (c). Cellobionic acid production using pretreated eucalyptus chips (d, e). Increased release of cellobionic acid was observed in the saccharification assays performed with the secretome of the strain co-expressing CelOCE in both technical substrates. Cellobionic acid production was monitored at 24, 48 and 72 h and the HPAEC-PAD product profiles are shown. Results from panels a, c and d are expressed as mean ± standard deviation from three independent experiments. In a and d, statistical significance was determined by one-way ANOVA with Tukey’s post hoc test (*** p < 0.001).
Supplementary information
Supplementary Figs. 1–30 (download PDF )
Figures include supporting data for HPAEC–PAD, LC–MS, ITC, XRF, XAS and EPR analyses, uncropped gels and additional structural and bioinformatic analyses, including metagenomics and phylogenetics.
Supplementary Tables 1–18 (download XLSX )
Tables encompass oligonucleotides used in this study, proteomics and EPR data and additional bioinformatic analyses.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Santos, C.A., Morais, M.A.B., Mandelli, F. et al. A metagenomic ‘dark matter’ enzyme catalyses oxidative cellulose conversion. Nature 639, 1076–1083 (2025). https://doi.org/10.1038/s41586-024-08553-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-024-08553-z
This article is cited by
-
Green bioconversion of insoluble chitin: chitinase development pathways via multi-strategy synergy
Bioresources and Bioprocessing (2026)
-
Determination of optimal conditions for the cellulase production ability of selected soil-borne fungal isolates under submerged fermentation
Biotechnology Letters (2026)
-
A disulfide redox switch mechanism regulates glycoside hydrolase function
Nature Communications (2026)
-
Enhancing low-temperature fermentation quality and modulating bacterial community of whole-plant maize silage using a novel cold-tolerant Lacticaseibacillus paracasei
BMC Plant Biology (2025)
-
N deposition affects litter decomposition in evergreen broad-leaf forests by reducing soil enzyme activities and altering the structure of microbial communities
Plant and Soil (2025)