Abstract
DNA damage promotes mutations that fuel cancer, ageing and neurodegenerative diseases1,2,3, but surprisingly, the causes and types of damage remain largely unknown. There are three identified mechanisms that damage DNA during transcription: collision of RNA polymerase (RNAP) with the DNA-replication machinery head-on and co-directionally4,5,6, and R-loop-induced DNA breakage7,8,9,10. Here we identify novel DNA damage reaction intermediates11,12 and uncover a fourth transcription-related source of DNA damage: endogenous DNA damage at sites of terminated transcripts. We engineered proteins to capture single-stranded DNA (ssDNA) ends with 3′ polarity in bacterial and human cells. In Escherichia coli, spontaneous 3′-ssDNA-end foci were unexpectedly frequent, at one or more per cell division, and arose via two identifiable pathways, both of which were dependent on DNA replication. A pathway associated with double-strand breaks was suppressed by overexpression of replicative DNA polymerase (pol) III, suggesting competition between pol III and DNA damage-promoting proteins. Mapping of recurrent 3′-ssDNA-ends identified distinct 3′-ssDNA-end-hotspots, mostly unrelated to double-strand breaks, next to the 5′-CCTTTTTT transcription-terminator-like sequence. These 3′-ssDNA-termini coincide with RNA 3′-termini identified by DirectRNA sequencing13 or simultaneous 5′ and 3′ end RNA sequencing (SEnd-seq)14 and were prevented by a mutant RNAP that reads through terminators. Our findings reveal that transcription termination or pausing can promote DNA damage and subsequent genomic instability.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Sequencing data reported in this paper are available on the Sequence Read Archive under the BioProject ID PRJNA949415. The E. coli genome assembly version used in this study is NC_000913.3. Biological materials generated in this study are available from the Rosenberg laboratory to academic researchers contingent on the Baylor College of Medicine Materials Transfer Agreement. Source data are provided with this paper.
Code availability
The main scripts used for the analysis in this paper are available from GitHub (https://github.com/JLiu0504/ThreeSSeq-processing).
References
Zeman, M. K. & Cimprich, K. A. Causes and consequences of replication stress. Nat. Cell Biol. 16, 2–9 (2014).
Gaillard, H., García-Muse, T. & Aguilera, A. Replication stress and cancer. Nat. Rev. Cancer 15, 276–289 (2015).
Tubbs, A. & Nussenzweig, A. Endogenous DNA damage as a source of genomic instability in cancer. Cell 168, 644–656 (2017).
Srivatsan, A., Tehranchi, A., MacAlpine, D. M. & Wang, J. D. Co-orientation of replication and transcription preserves genome integrity. PLoS Genet. 6, e1000810 (2010).
Tehranchi, A. K. et al. The transcription factor DksA prevents conflicts between DNA replication and transcription machinery. Cell 141, 595–605 (2010).
Dutta, D., Shatalin, K., Epshtein, V., Gottesman, M. E. & Nudler, E. Linking RNA polymerase backtracking to genome instability in E. coli. Cell 146, 533–543 (2011).
Wahba, L., Amon, J. D., Koshland, D. & Vuica-Ross, M. RNase H and multiple RNA biogenesis factors cooperate to prevent RNA: DNA hybrids from generating genome instability. Mol. Cell 44, 978–988 (2011).
Wimberly, H. et al. R-loops and nicks initiate DNA breakage and genome instability in non-growing Escherichia coli. Nat. Commun. 4, 2115 (2013).
Hamperl, S., Bocek, M. J., Saldivar, J. C., Swigut, T. & Cimprich, K. A. Transcription–replication conflict orientation modulates R-loop levels and activates distinct DNA damage responses. Cell 170, 774–786.e719 (2017).
Lang, K. S. et al. Replication–transcription conflicts generate R-loops that orchestrate bacterial stress survival and pathogenesis. Cell 170, 787–799.e718 (2017).
Konforti, B. & Davis, R. DNA substrate requirements for stable joint molecule formation by the RecA and single-stranded DNA-binding proteins of Escherichia coli. J. Biol. Chem. 266, 10112–10121 (1991).
Razavy, H., Szigety, S. K. & Rosenberg, S. M. Evidence for both 3′ and 5′ single-strand DNA ends in intermediates in Chi-stimulated recombination in vivo. Genetics 142, 333–339 (1996).
Adams, P. P. et al. Regulatory roles of Escherichia coli 5′ UTR and ORF-internal RNAs detected by 3′ end mapping. eLife 10, e62438 (2021).
Ju, X., Li, D. & Liu, S. Full-length RNA profiling reveals pervasive bidirectional transcription terminators in bacteria. Nat. Microbiol. 4, 1907–1918 (2019).
Hou, Y., Song, H., Croteau, D. L., Akbari, M. & Bohr, V. A. Genome instability in Alzheimer disease. Mech. Ageing Dev. 161, 83–94 (2017).
Vermeij, W. et al. Restricted diet delays accelerated ageing and genomic stress in DNA-repair-deficient mice. Nature 537, 427–431 (2016).
Xia, J. et al. Bacteria-to-human protein networks reveal origins of endogenous DNA damage. Cell 176, 127–143.e124 (2019).
Seigneur, M., Bidnenko, V., Ehrlich, S. D. & Michel, B. RuvAB acts at arrested replication forks. Cell 95, 419–430 (1998).
Neelsen, K. J. & Lopes, M. Replication fork reversal in eukaryotes: from dead end to dynamic response. Nat. Rev. Mol. Cell Biol. 16, 207–220 (2015).
Rupp, W. D. & Howard-Flanders, P. Discontinuities in the DNA synthesized in an excision-defective strain of Escherichia coli following ultraviolet irradiation. J. Mol. Biol. 31, 291–304 (1968).
Sogo, J. M., Lopes, M. & Foiani, M. Fork reversal and ssDNA accumulation at stalled replication forks owing to checkpoint defects. Science 297, 599–602 (2002).
Yeeles, J. T. & Marians, K. J. The Escherichia coli replisome is inherently DNA damage tolerant. Science 334, 235–238 (2011).
Xia, J. et al. Holliday junction trap shows how cells use recombination and a junction-guardian role of RecQ helicase. Sci. Adv. 2, e1601605 (2016).
Pennington, J. M. & Rosenberg, S. M. Spontaneous DNA breakage in single living Escherichia coli cells. Nat. Genet. 39, 797–802 (2007).
Paull, T. T. et al. A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr. Biol. 10, 886–895 (2000).
Britton, S., Coates, J. & Jackson, S. P. A new method for high-resolution imaging of Ku foci to decipher mechanisms of DNA double-strand break repair. J. Cell Biol. 202, 579–595 (2013).
Shee, C. et al. Engineered proteins detect spontaneous DNA breakage in human and bacterial cells. eLife 2, e01222 (2013).
Canela, A. et al. DNA breaks and end resection measured genome-wide by end sequencing. Mol. Cell 63, 898–911 (2016).
Lensing, S. V. et al. DSBCapture: in situ capture and sequencing of DNA breaks. Nat. Methods 13, 855–857 (2016).
Mei, Q. et al. Two mechanisms of chromosome fragility at replication-termination sites in bacteria. Sci. Adv. 7, eabe2846 (2021).
Caldecott, K. W. Single-strand break repair and genetic disease. Nat. Rev. Genet. 9, 619–631 (2008).
Sassanfar, M. & Roberts, J. W. Nature of the SOS-inducing signal in Escherichia coli: the involvement of DNA replication. J. Mol. Biol. 212, 79–96 (1990).
Blackford, A. N. & Jackson, S. P. ATM, ATR, and DNA-PK: the trinity at the heart of the DNA damage response. Mol. Cell 66, 801–817 (2017).
Sivaramakrishnan, P. et al. The transcription fidelity factor GreA impedes DNA break repair. Nature 550, 214–218 (2017).
Sollier, J. et al. Transcription-coupled nucleotide excision repair factors promote R-loop-induced genome instability. Mol. Cell 56, 777–785 (2014).
Kuzminov, A. & Stahl, F. W. Double-strand end repair via the RecBC pathway in Escherichia coli primes DNA replication. Genes Dev. 13, 345–356 (1999).
Pomerantz, R. T. & O’donnell, M. The replisome uses mRNA as a primer after colliding with RNA polymerase. Nature 456, 762 (2008).
Kushner, S. R., Nagaishi, H., Templin, A. & Clark, A. J. Genetic recombination in Escherichia coli: the role of exonuclease I. Proc. Natl Acad. Sci. USA 68, 824–827 (1971).
Thoms, B., Borchers, I. & Wackernagel, W. Effects of single-strand DNases ExoI, RecJ, ExoVII, and SbcCD on homologous recombination of recBCD+ strains of Escherichia coli and roles of SbcB15 and XonA2 ExoI mutant enzymes. J. Bacteriol. 190, 179–192 (2008).
Lloyd, R. G. & Buckman, C. Identification and genetic analysis of sbcC mutations in commonly used recBC sbcB strains of Escherichia coli K-12. J. Bacteriol. 164, 836–844 (1985).
Dillingham, M. S. & Kowalczykowski, S. C. RecBCD enzyme and the repair of double-stranded DNA breaks. Microbiol. Mol. Biol. Rev. 72, 642–671 (2008).
Gumbiner-Russo, L. M., Lombardo, M.-J., Ponder, R. G. & Rosenberg, S. M. The TGV transgenic vectors for single-copy gene expression from the Escherichia coli chromosome. Gene 273, 97–104 (2001).
Kim, J. J., Kumbhar, R., Gong, F. & Miller, K. M. In time and space: laser microirradiation and the DNA damage response. Methods Mol. Biol. 1999, 61–74 (2019).
Joshi, M. C. et al. Regulation of sister chromosome cohesion by the replication fork tracking protein SeqA. PLoS Genet. 9, e1003673 (2013).
Fehér, T., Cseh, B., Umenhoffer, K., Karcagi, I. & Pósfai, G. Characterization of cycA mutants of Escherichia coli: an assay for measuring in vivo mutation rates. Mutat. Res. 595, 184–190 (2006).
Hastings, P. et al. Competition of Escherichia coli DNA polymerases I, II and III with DNA Pol IV in stressed cells. PLoS ONE 5, e10862 (2010).
Kath, J. E. et al. Exchange between Escherichia coli polymerases II and III on a processivity clamp. Nucleic Acids Res. 44, 1681–1690 (2016).
Maki, H., Horiuchi, T. & Kornberg, A. The polymerase subunit of DNA polymerase III of Escherichia coli. I. Amplification of the dnaE gene product and polymerase activity of the alpha subunit. J. Biol. Chem. 260, 12982–12986 (1985).
Miko, I. Epistasis: gene interaction and phenotype effects. Nat. Educ. 1, 197 (2008).
Yanofsky, C. & Horn, V. Rifampin resistance mutations that alter the efficiency of transcription termination at the tryptophan operon attenuator. J. Bacteriol. 145, 1334–1341 (1981).
McDowell, J. C., Roberts, J. W., Jin, D. J. & Gross, C. Determination of intrinsic transcription termination efficiency by RNA polymerase elongation rate. Science 266, 822–825 (1994).
Kogoma, T. Escherichia coli RNA polymerase mutants that enhance or diminish the SOS response constitutively expressed in the absence of RNase HI activity. J. Bacteriol. 176, 1521–1523 (1994).
Gusarov, I. & Nudler, E. The mechanism of intrinsic transcription termination. Mol. Cell 3, 495–504 (1999).
Kotlajich, M. V. et al. Bridged filaments of histone-like nucleoid structuring protein pause RNA polymerase and aid termination in bacteria. eLife 4, e04970 (2015).
Zenkin, N., Yuzenkova, Y. & Severinov, K. Transcript-assisted transcriptional proofreading. Science 313, 518–520 (2006).
Dey, S. et al. Structural insights into RNA-mediated transcription regulation in bacteria. Mol. Cell 82, 3885–3900.e10 (2022).
Mirkin, E. V., Castro Roa, D., Nudler, E. & Mirkin, S. M. Transcription regulatory elements are punctuation marks for DNA replication. Proc. Natl Acad. Sci. USA 103, 7276–7281 (2006).
Borukhov, S., Sagitov, V. & Goldfarb, A. Transcript cleavage factors from E. coli. Cell 72, 459–466 (1993).
Phillips, G. J., Prasher, D. & Kushner, S. R. Physical and biochemical characterization of cloned sbcB and xonA mutations from Escherichia coli K-12. J. Bacteriol. 170, 2089–2094 (1988).
Liu, J., Mei, Q., Nimer, S., Fitzgerald, D. M. & Rosenberg, S. M. in Methods in Enzymology, Vol. 661 (ed. Eichman, B. F.) 155–181 (Elsevier, 2021).
Cohen, A. & Clark, A. J. Synthesis of linear plasmid multimers in Escherichia coli K-12. J. Bacteriol. 167, 327–335 (1986).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Bonura, T. & Smith, K. C. Sensitization of Escherichia coli C to gamma-radiation by 5-bromouracil incorporation. Int. J. Radiat. Biol. Relat. Stud. Phys. Chem. Med. 32, 457–464 (1977).
Gong, F., Clouaire, T., Aguirrebengoa, M., Legube, G. & Miller, K. M. Histone demethylase KDM5A regulates the ZMYND8–NuRD chromatin remodeler to promote DNA repair. J. Cell Biol. 216, 1959–1974 (2017).
Henricksen, L. A., Umbricht, C. B. & Wold, M. S. Recombinant replication protein A: expression, complex formation, and functional characterization. J. Biol. Chem. 269, 11121–11132 (1994).
Ducret, A., Quardokus, E. M. & Brun, Y. V. MicrobeJ, a tool for high throughput bacterial cell detection and quantitative analysis. Nat. Microbiol. 1, 16077 (2016).
Bolte, S. & Cordelieres, F. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 224, 213–232 (2006).
Kim, D. R., Pritchard, A. E. & McHenry, C. S. Localization of the active site of the alpha subunit of the Escherichia coli DNA polymerase III holoenzyme. J. Bacteriol. 179, 6721–6728 (1997).
Zheng, Q. rSalvador: an R package for the fluctuation experiment. G3: Genes Genomes Genet. 7, 3849–3856 (2017).
Bonocora, R. P. & Wade, J. T. in Bacterial Transcriptional Control (eds Artsimovitch, I. & Santangelo, T. J.) 327–340 (Springer, 2015).
Thomason, L. C., Sawitzke, J. A., Li, X., Costantino, N. & Court, D. L. Recombineering: genetic engineering in bacteria using homologous recombination. Curr. Protoc. Mol. Biol. 106, 1.16.11–11.16. 39 (2014).
Reisch, C. R. & Prather, K. L. The no-SCAR (Scarless Cas9 Assisted Recombineering) system for genome editing in Escherichia coli. Sci. Rep. 5, 15096 (2015).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Wold, M. S., Weinberg, D. H., Virshup, D. M., Li, J. J. & Kelly, T. J. Identification of cellular proteins required for simian virus 40 DNA replication. J. Biol. Chem. 264, 2801–2809 (1989).
Razavy, H. Single-Strand DNA Ends in Recombination In Vivo. MSc thesis, Univ. of Alberta (1997).
Rinken, R., Thomas, B. & Wackernagel, W. Evidence that recBC-dependent degradation of duplex DNA in Escherichia coli recD mutants involves DNA unwinding. J. Bacteriol. 174, 5424–5429 (1992).
Connelly, J. C., de Leau, E. S. & Leach, D. R. DNA cleavage and degradation by the SbcCD protein complex from Escherichia coli. Nucleic Acids Res. 27, 1039–1046 (1999).
Chase, J. W. & Richardson, C. C. Exonuclease VII of Escherichia coli: mechanism of action. J. Biol. Chem. 249, 4553–4561 (1974).
Schaaper, R. M. Base selection, proofreading, and mismatch repair during DNA replication in Escherichia coli. J. Biol. Chem. 268, 23762–23765 (1993).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Skordalakes, E., Brogan, A. P., Park, B. S., Kohn, H. & Berger, J. M. Structural mechanism of inhibition of the Rho transcription termination factor by the antibiotic bicyclomycin. Structure 13, 99–109 (2005).
Zhou, Y. & Martin, C. T. Observed instability of T7 RNA polymerase elongation complexes can be dominated by collision-induced “bumping”. J. Biol. Chem. 281, 24441–24448 (2006).
Acknowledgements
The authors thank G. Ira, A Nussenzweig and C. Zong for comments; K. Sheng, W. Cao, A. McMullin and P. Chen for technical help; J. T. Wade for DirectRNA-seq data analysis; R. Reyes-Lamothe for sharing strains; and H. Razavy whose MS thesis supplied the data for Extended Data Fig. 1g. The study was supported by the W. M. Keck Foundation (S.M.R. and K.M.M.), National Institutes of Health (NIH) Director’s Pioneer Awards DP1-AG072751 (S.M.R.) and DP1-AI152073 (C.H.), NIH grants R01-CA250905 (S.M.R. and K.M.M.), R35-CA241801 (P.M.S.), R01-GM067153 (I.A.), R01-GM106373 (P.J.H.), R01-GM135368 (D.B.) and R00ES033259 (J.X.); National Aeronautics and Space Administration, Translational Health Research Institute grant NNX16AO69A (S.M.R. and K.M.M.), American Cancer Society Postdoctoral Fellowships PF-18-035-01 (D.M.F.) and PF-22-034-01-DMC (C.M.R.); the Dan L Duncan Comprehensive Cancer Center NIH P30-CA125123, and its support of the Baylor College of Medicine (BCM) Integrated Microscopy Core and the Center for Advanced Microscopy and Image Informatics (also CAMII, NIH DK56338 and ES030285) and BCM Cytometry and Cell Sorting core (also NIH P30-AI36211 and S10-RR024574; Cancer Prevention and Research Institute of Texas RP150578 and RP170719), and the John S. Dunn Gulf Coast Consortium for Chemical Genomics.
Author information
Authors and Affiliations
Contributions
S.M.R. conceived the study. J.L., C.M.R., J.A.H., D.B., L.L., I.A., C.H., K.M.M., P.M.S. and S.M.R. designed or supervised experiments. J.L., D.M.F., Q.M., J.A.H., J.O.P., C.M.R., M.B.C., S.N., R.B.N., M.C., A.X.W., S.G.K., J.X., Y.Z., L.L. and I.A. performed experiments. Bioinformatics analysis was performed by J.L., D.M.F. and Q.M. J.L. analysed sequencing data. J.L., C.H., P.J.H. and S.M.R. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Sabari Sankar Thirupathy and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Construction and validation of E. coli DExI fusion proteins.
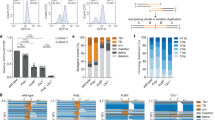
a, Diagram of the chromosomal inducible non-tagged DExI, DExI-GFP, DExI-mCherry and DExI-FLAG fusion constructs used in this study, strains SMR22601, SMR22593/SMR24774, SMR24800 and SMR24776, respectively. The doxycycline- and temperature-inducible expression cassettes were engineered at a non-genic site in the E. coli chromosome (Methods), in cells that carry the wild-type gene in its native locus. Because DExI is dominant to the wild-type gene12, DExI phenotypes are observed here, as previously12. The doxcycline-inducible expression system consists of the phage PN25 constitutive promoter-driven tetR repressor gene and the inducible P N25tetO promoter-driven DExI fusions (left and middle). The temperature-inducible expression system consists of the phage lambda cIts857 temperature-sensitive repressor gene and lambda PR promoter-driven DExI fusion (right). b, DExI-mCherry forms fewer spontaneous foci than DExI-GFP. DExI focus appearance frequencies with indicated fusions in log-phase cells. Mean ± range, n = 2 experiments. c, Coomassie-stained image of recombinant DExI-GFP-6xHis (herein referred to as DExI-GFP). Representative image of two, with similar results. For gel source data, see Supplementary Fig. 1. d-f, DExI-GFP binds only 3’-ssDNA-end-bearing substrates detectably. d, DExI-GFP binds ssDNA, but not dsDNA or ssRNA. Left, the polynucleotide binding properties of DExI-GFP were examined by electrophoretic mobility shift assay (EMSA). 1 nM 32P-radiolabeled ss/dsDNA and ssRNA probes were incubated with indicated concentrations of DExI-GFP. Shift of polynucleotide relative to no-protein control indicates protein binding. Right, quantification from three experiments: Whereas the binding to ssDNA with a 3′ end has a dissociation constant (Kd) of 0.6 ± 0.4 nM (KDs averaged from n = 2 experiments, mean ± range) the binding to blunt dsDNA; ssRNA; and 5’-ssDNA overhanging tail are undetectable (ND) even with 50 µM DExI-GFP protein, indicating that their binding by DExI-GFP is weaker than 50 µM, at least 680 times weaker than to 3’-ssDNA ends. This was also the case for binding to e, circular ss or dsDNA, mean ± S.E.M for n = 3 experiments for ssDNA. f, binding to closed circular dsDNA, nicked circular dsDNA and dsDNA nicked at the consensus sequence. EMSA analysis of 350 ng circular ss or dsDNA substrates was performed with indicated concentrations of DExI-GFP or human replication protein A (RPA). The latter was used as a positive control for DNA binding75. Representative image of n = 6, n = 6, n = 3 experiments for ssDNA, intact and nicked dsDNA, respectively. For gel source data, see Supplementary Fig. 1. g, DExI production mimics the phenotype of cells lacking Exo I and Exo VII, supporting its role blocking Exo VII degradation of DNA. Data from76. Assays of “RecF-pathway” homologous recombination (HR) of linear phage lambda DNAs show that the sbcB15 (or DExI) mutation mimics the phenotype of cells that carry both the deletion of the Exo I (∆xonA) and of Exo VII (∆xseA) simultaneously, and so lack all known single-strand-dependent 3′ exonuclease activity in E. coli. Removal of Exo VII 3’- and 5’-single-strand-specific exonuclease (∆xseA) simultaneously with Exo I (∆xonA) restores HR proficiency to recBC sbcC cells as well as the SbcB15-mutant (DExI) protein does. [xthA encoding Exo III, an apurinic-apyrimidinic (AP) endonuclease, made no detectable contribution.] Diagram (above): two phage lambda genotypes co-infected (“crossed”) in the strains indicated below (single lines represent double-stranded DNA molecules). The lambda lack their own red HR system (bio1 substitution or ∆b1453) and HR-related genes encoded in the nin region (∆nin5) and so recombine via E. coli proteins. The lambda replicate via rolling circle and so do not require any HR for formation of lambda genome multimers, required for lambda genome packaging (and so for progeny phage, per12). These data imply that the inability of ∆xonA to restore HR to recBC sbcC cells results from Exo VII action, and that blocking Exo VII with SbcB15-mutant (DExI) protein prevents Exo VII-inhibition of HR. HR is quantified as the titer of lambda E + S+ recombinant particles among Ets3 parental phage from crosses with limiting Ets parent, 50-times fewer of than of the other lambda genotype employed, using methods per12,76. These are means ± s.e.m. of ratios, normalized to the value from the recBC sbcC sbcB15 strain done in parallel for each genotype with the numbers of independent experiments for each bar as follows, from left to right: n = 6, n = 5, n = 6, n = 3, n = 3. ***p ≤ 0.001, one-way ANOVA with multiple comparison test.; n.s., not significant. h, How DExI promotes and Exo I, Exo VII, and SbcCD hairpin endo/exonuclease inhibit “RecF-pathway” recombination at DSB ends. Cells that lack the RecBCD double-strand endo/exonuclease require both an sbcCD null mutation and DExI (SbcB15 protein) to restore UV-resistance and HR proficiency40 (and Fig. 1b). Above: in ∆recB DExI-producing cells: DExI (SbcB15 protein, blue) prevents Exo I and Exo VII ssDNA-dependent exonucleases (red notched circle) from degrading 3’-ssDNA ends12,77 (and g, above). The SbcCD hairpin endo/exonuclease78 (orange notched circle) degrades DNA with hairpin secondary structures independently of Exo I and Exo VII action. Exo I degrades 3’-ssDNA ends and Exo VII degrades ssDNA ends of 3′ or 5′ polarity79. Below: in ∆recB ∆sbcC DExI cells, blocking degradation of 3’-ssDNA ends with DExI protein and preventing hairpin endo/exo activity by sbcC deletion is proposed to allow persistence of the single-stranded DNA with 3’-ends, required for RecA loading and homology-directed repair (HDR). i, Efficiency of DExI-GFP focus formation after gamma irradiation. The efficiency of DExI focus formation depends on whether each DSB contains one or two resected DSB ends. Thus, it would be 0.5 foci per DSB if these breaks were mostly one-ended, or 0.25 foci per DSB if they were mostly two-ended. In E. coli cells, the directional distribution of nuclease-impeding Chi sites36 leads to more rapid directional nucleolytic destruction of one of the two chromosome arms or DSB ends than the other34, and the ratio of one-to-two-ended species is not known. j, Representative example (above) and quantification of western blots for Exo I-GFP under the native xonA promoter or DExI-GFP driven by the chromosomal inducible PN25tetO promoter (means ± s.e.m., n = 6 experiments for all except the rightmost value, for which n = 3). From these data we can approximate DExI concentrations. On induction, about 94-times more DExI is produced than native Exo I, also present; and uninduced, about 1.7 times more DExI than native Exo I. For gel source data, see Supplementary Fig. 1. E. coli strains used: b: SMR24774, SMR24800; g: SMR121, SMR1700, SMR1701, SMR1775, SMR3108; i: SMR24774; j, SMR27431, SMR27378, SMR24774.
Extended Data Fig. 2 E. coli DExI rapidly accumulates at laser-induced damage in human cancer cells.
a, Diagram of EmGFP-DExI induction and live-cell imaging after laser micro-irradiation in U2OS cells. b, EmGFP-DExI accumulates at sites of laser-induced DNA damage. Scale bar, 10 μm. c, Quantification of b. EmGFP fluorescence intensity at damage sites was calculated by comparing the EmGFP-DExI intensity at damaged versus undamaged sites within the same cell as a function of time. Mean ± s.e.m. n = 12 cells.
Extended Data Fig. 3 Spontaneous 3’-ssDNA-end foci are frequent.
a, Representative images of cells grown in microfluidic chamber, images that record complete rounds of cell division. DExI-GFP-producing cells revealed short- or long-lived spontaneous foci, whereas no foci were observed in GFP-producing cells. b, DExI inhibits cell division. Scatter plot of division times in indicated strains. “no foci” indicates induced cells that are producing DExI-GFP or GFP, but had no visible focus at the time photographed. Cumulative data from n = 3 experiments, one-way ANOVA with multiple comparison test. c, Dose response of increased DExI-GFP induction with increasing inhibition of growth rate. n = 3 experiments, two-sided Student’s t test, asterisks from left to right, **p = 0.0010, **p = 0.0013, ***p = 0.0002, ***p = 0.0001. d, Reduction of chromosome number in Exo I-producing cells/flow cytometry histogram of DNA content (DAPI fluorescence intensity) of Exo I-producing cells after replication run-off. These controls for the replication run-off assays in Fig. 3g produce wild-type Exo I and showed fewer chromosome copies per cell than the wild-type (Fig. 3g), even though they did not display the failure to complete replication seen in DExI-producing cells (Fig. 3g). These data support the interpretation that the spontaneous 3’-ssDNA-ends detected by DExI (Figs. 1 and 3) are generated normally by replication, not as a result of the mutant DExI protein, and so are rapidly degraded in Exo I-producing cells. That is, the spontaneous 3’-ssDNA-ends detected by DExI (Figs. 1 and 3) are not an artifact of producing DExI, and its persistent binding to DNA. e, Although the frequency of mutations is increased by DExI, DExI induced little change in the sequences of cycA gene mutations compared with spontaneous mutations. Summary of mutation sequences from 49 independent clones each, isolated from each of spontaneous and DExI-induced mutants. The two blue dots indicate mutations that occurred together in one mutant isolate, and the two green dots in a different mutant isolate. Each of the rest occurred singly in each sequenced isolate. f, Additional controls for Fig. 3i. No significant reduction of spontaneous mutation rate by rpoB2 or ∆greA ∆greB in cells without DExI. Mean ± s.e.m. n = 4 experiments; one-way ANOVA, ns not significant. g, Increased (not reduced) mutagenesis from increased production of pol IIIα suggests that suppressing the pol IIIα-suppressible DSB pathway may have increased the contribution to mutagenesis of the RNAP-associated pathway, which promotes mutations at cycA. Given that the RNAP-associated pathway requires replication (Fig. 3a), the mutagenesis increase might result from the proposed STABD hypothesis (Fig. 5g), which requires two events to form the RNAP-promoted 3’-ssDNA ends trapped by DExI: (i) RNAP cleavage of the DNA, and (ii) replisome displacement of the cleaved strand (Fig. 5g center picture). Perhaps increased processivity of the replisome with pol IIIα overproduction increases the probability of the second event (displacement), which would increase the frequency of DExI trapping of the 3′ end created by RNAP, and therefore mutagenesis. Mean ± s.e.m. n = 4 experiments; one-way ANOVA with multiple comparison test. Asterisks from left to right,**p = 0.007, *p = 0.038, *p = 0.021. E. coli strains used: a: SMR22427, SMR24774; b: SMR22427, SMR24774; c: SMR26532, SMR26551; d: SMR23317; f: SMR3070, SMR26532, SMR26553, SMR27374, g: SMR26532, SMR26551, SMR27366.
Extended Data Fig. 4 Production of the DNA pol IIIα subunit reduces spontaneous 3’-ssDNA-ends by reducing DSBs.
a, Western-blot analyses confirm that doxycycline (dox) induces pol IIIα production, and show its leaky expression with no dox inducer. Representative image of two with similar results. b, Pol IIIα levels have little effect on cell growth rate. The growth curve (optical density, OD = 600 nm) of DExI-GFP- and pol IIIα-producing cells with indicated dox concentrations. Saturated overnight cultures diluted in M9 glucose medium with different concentrations of doxycycline were grown at 30°C for 1.5 h, then shifted to 37 °C for DExI-GFP induction, mean ± range, n = 2 experiments. c, The corresponding DExI-GFP foci, examined after 4 h. n = 3 experiments (0 dox), n = 2 experiments (1 ng/ml), n = 2 experiments (10 ng/ml), n = 3 experiments (100 ng/ml), mean ± SEM, *p = 0.017, two-sided Student’s t test. d, Excessive, but not normal levels of DNA pol III replication errors promote 3’-ssDNA-end foci. We manipulated levels of the ε subunit of pol III, encoded by the dnaQ gene, which performs replication “proofreading” (removal) of mis-incorporated bases in replication80. Deletion of dnaQ increased DExI foci, indicating that excessive persistence of misincorporated bases can induce DNA damage containing 3’-ssDNA-ends. Mean ± s.e.m. n = 3 experiments, **p = 0.0012, two-sided Student’s t test. e, However, DnaQ overproduction had little effect, implying that, normally, proofreading by DnaQ/ε does not cause most spontaneous 3’-ssDNA-ends; proofreading activity is sufficient to prevent those ends. n = 2 experiments. f, Trans-lesion synthesis (TLS) DNA polymerases are not required for spontaneous 3’-ssDNA-end focus formation. Frequencies of DExI-GFP foci in strains that lack one or more TLS DNA polymerase. n = 2 experiments, mean ± range. g, Western blot analysis of the pol IIIα subunit and Gam amounts in indicated strains. Note that the apparent decrease in the pol IIIα subunit in rpoB2 cells would be expected to increase foci whereas spontaneous foci were decreased in rpoB2 cells (Fig. 4a,f). Representative blots. h, Models for generation of pol IIIα-suppressible, DSB-dependent spontaneous 3’-ssDNA-ends. We hypothesize that (i) high levels of the highly processive DNA pol III reduce replication stalling or stabilize forks until blocks are removed; (ii) conversely, low pol III levels, e.g., by stochastic variation between cells, might allow fork reversal which liberates nascent 3’-ends of the leading strand at a DSB end; (iii) reversed forks can be cleaved by Holliday junction-specific endonucleases18, creating a DSB end, (iv) then DSB-end resection produces 3’-ssDNA-ends. (v) DSB ends might also arise from single-strand interruptions. Fork reversal has been associated with transcription termination57. However, the pausing/termination-promoted, rpoB2-suppressible component of 3’-ssDNA-end foci was mostly non-overlapping with the DSB (Gam-suppressed) component (Fig. 4a), implying that most RNAP pausing/termination-related 3’-ssDNA-ends are not likely to be at reversed forks, which possess DSB ends. By any of many models, including those above, pol IIIα might block or out-compete clamp interactions with other proteins (repair or synthesis) that contribute to spontaneous DSBs that acquire 3’-ssDNA-ends by resection. i, Single deletions of ΔgreA or ΔgreB show wild-type levels of DExI-GFP foci. n = 4 experiments, mean ± s.e.m. For gel source data, see Supplementary Fig. 1. E. coli strains used: a: SMR26382; b: SMR26382; c: SMR23252, SMR26382; d: SMR24774, SMR26631; e: SMR26649, SMR26651, SMR26653, SMR26655; f: SMR24774, SMR26371, SMR26373, SMR26374; g: SMR23252, SMR26382, SMR26483, SMR26485, SMR26487, SMR26488, SMR26385, SMR26486; i: SMR24774, SMR25015, SMR25016.
Extended Data Fig. 5 ThreeSSeq 3’-ssDNA-end genomic profiling with DExI detects dispersed, reproducible, hotspots.
a, Integrated genome viewer (IGV)81 views of signals from ThreeSSeq with DExI in the indicated strains reveal hotspots throughout the genome. Signals were normalized by Reads Per Kilobase per Million mapped reads (RPKM). For calling of peaks and all quantification, the DExI signal (negative control, background) is subtracted from the signal with DExI-FLAG to obtain the DExI-FLAG-specific reads. b, Two DExI-FLAG experiments are more correlated with each other than with the DExI control. Heatmap of Pearson correlations between the experiments. c, Venn diagrams of peak overlap between two replicates in DExI-FLAG-producing, otherwise wild-type cells. d, Heatmaps of ThreeSSeq signals within ± 300 nt regions around peak summits of 113 TOP-strand and 111 BTM-strand 3’-ssDNA-end hotspot sites. Sites were centered along peak summits and sorted by signal intensity. Signals were normalized by RPKM. e, The distribution of 175 sequence-specific 3’-ssDNA-end sites (shown as black dots) throughout the genome. The leading- and lagging-strand in each replichore are indicated. f, Sequence-specific 3’-ssDNA-end hotspots are distributed randomly throughout the genome. To test the randomness of the 3’-ssDNA-end-hotspot distribution, we sampled 175, 92 and 83 genomic sites, corresponding to the number of all DExI-FLAG-immunoprecipitated sites throughout the genome, the number of TOP-strand sites and the number of BTM-strand sites, randomly 1000 times, and calculated the average distance of neighboring sites. The barplots display the distribution of the sample means, and the arrows indicate the average distances of tested data, including all 175 sites (left, p = 0.10), 92 TOP-strand sites (center, p = 0.48), and 83 BTM-strand sites (right, p = 0.16). The distribution of actual 3’-ssDNA-end hotspots was not different from the 1000 random samples. Strains used: a: SMR24776, SMR24777; b: SMR24776, SMR24777; d: SMR24776.
Extended Data Fig. 6 The rpoB2 RNA-polymerase read-through mutation reduces spontaneous ThreeSSeq 3’-ssDNA-end-hotspot signals at the terminator-like consensus sequence.
a, Heatmap of ThreeSSeq signal in rpoB2, phage Mu Gam-producing, and dnaE pol IIIα subunit-producing cells across 175 spontaneous sequence-specific 3’-ssDNA-end hotspot sites. Signals are normalized by RPKM and ordered according to signal intensity in the wild-type cells. Signals are displayed along ± 300-nt region around, and centered along, peak summits. Whereas each rpoB2 data set differs significantly from the WT done in parallel with it (b, below), the data from Gam production and pol IIIα production do not differ from the WT. b, Violin plot showing ThreeSSeq signals of 175 3’-ssDNA-end sites in indicated strains. Read counts were normalized by RPKM using the function dba.count in R package DiffBind. One-way ANOVA with multiple comparison test. n = the total cumulative dots from 2 experiments. c, Integrative genomic viewer (IGV) views of ThreeSSeq signals in indicated strains at two 3’-ssDNA-end hotspot sites. d, CRISPR-ablation of consensus sites (C-sites) that flank cycA reduce DExI-induced mutagenesis at cycA. Mean ± s.e.m. n = 4 experiments. One-way ANOVA with multiple comparison test, asterisks from left to right, p < 0.0001 (GraphPad software does not provide exact p value), p = 0.0005, p = 0.001. E. coli strains used: a: SMR24776, SMR26369, SMR26525, SMR26366; SMR24776, SMR26369; d: SMR27380, SMR27381, SMR26532, SMR26551, SMR27384, SMR27385.
Extended Data Fig. 7 Endogenous 3’-ssDNA-ends at consensus sites are associated with termination of unannotated transcripts.
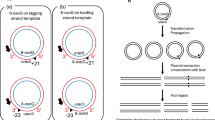
a, Classification, and distribution of recurrent 3’-ssDNA-end sites relative to annotated genes. 3’-ssDNA-ends are defined as: non-template (blue; located on the sense strand within a gene), intergenic (yellow), and template (mageneta; located on the antisense strand within a gene). Arrows represent open reading frames. b, Subclassification and distribution of intergenic 3’-ssDNA-end sites. They are defined as: 3’ UTR (located on the extended sense strand of an upstream gene); 5’ UTR (located on the extended sense strand of a downstream gene); both and neither. c, Bicyclomycin treatment has little effect on DirectRNA-seq signal13 at the 3’-ssDNA-end sites, indicating Rho-independent transcript terminations82. Average profile of DirectRNA-seq signal in bicyclomycin- (BCM)-treated cells versus untreated cells.
Extended Data Fig. 8 DNA endonuclease activity of RNAP at the ThreeSSeq consensus sequence, the rpoB2-sensitive component.
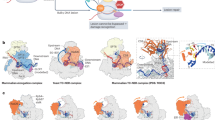
a, Transcription complexes were assembled with E. coli core RNAP on nucleic acid scaffolds composed of template DNA (T DNA, gray), non-template DNA (NT DNA, blue), and RNA (red). The RNA, which anneals to the T DNA strand, defines the geometry of the RNAP-DNA complex. In the absence of RNA, RNAP binds in either of two orientations: binding either the blue or gray DNA strand as template. In panel b, only one scaffold orientation is shown: blue strand on top. But in all DNA substrates with neither protein nor RNA bound, RNAP binds either top or bottom strand as template. b, Indicated DNA strands were labeled at the 5’ end with γ32P-ATP and T4 PNK (label position shown by a yellow star). Assembled complexes were incubated at 37 °C for 15, 30, and 60 min; E. coli RfaH, which specifically binds to the GCGGTAGC element in the NT DNA, was present where shown. After quenching, DNA products were analysed on a 10% urea-acrylamide gel and visualized by phosphorimaging. A dashed line indicates samples from different gels. DNA ladder was made by mixing DNA oligonucleotides of indicated lengths and labeling the mixture with γ32P-ATP and T4 PNK; the 30-mer was present in 3x molar excess. The red X indicates the position of two apparent cleavage products in the non-template DNA; the resulting products migrate as 26- and 27-mers. Each experiment was performed three times with similar results. Primary data are provided in the Source Data File. Two negative controls indicate consensus-sequence specificity of the cleavage: (i) the 5’- labeled T-DNA, which would show cleavage in the bottom gray strand if cleavage were non-specific and could occur next to the 5’ACCCGCGT sequence there; and (ii) the 5’-labeled NT AA DNA, which carries the 5’AATTTTTT sequence in place of the 5’CCTTTTTT consensus, at the same position, with same strand labeled. Neither of these substrates elicits cleavage 5′ of those sequences as is seen with the active consensus sequence in labeled strand (leftmost, 5’-labeled NT-CC DNA). For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 9 Consensus-sequence sites with little DNA damage display transcript termination at nearby upstream sites.
a, DirectRNA-seq reveals two termination peaks (RNA 3’-ends) near sites of the consensus sequence (CS) at which ThreeSSeq did not detect DNA damage. Average profile of DirectRNA-seq14 and ThreeSSeq signals around the CSs of 2375 sites on the TOP (left) and 2277 sites on the BTM (right) strand. DirectRNA-seq signals that mapped to TOP (green) and BTM (yellow) strands, along −300-nt to +100-nt regions, around the CSs, are superimposed on ThreeSSeq signals (TOP: orange; BTM: purple). b, Examples of integrative genomic viewer (IGV) views of DirectRNA-seq signals (green) and ThreeSSeq signals (orange) at two sites that have the CS but are not detected by ThreeSSeq and, c, at two sites at which 3’-ssDNA-ends are detected. d, Model for DNA-damage prevention at the “cold” sites of the CS, following the idea from Zhou and Martin83 on the effects of multiple RNAPs, or the lack thereof, on RNAP dynamics. The model can explain our data that a closely upstream RNAP termination site may prevent the ThreeSSeq-detectable DNA damage. An upstream RNAP might occlude the CS downstream, or displace an RNAP at the downstream CS, so that the DNA cannot be cleaved when the transcript ends. Alternatively (not shown), more termination at the upstream terminator leaves fewer RNAPs and transcripts to end at the downstream site.
Supplementary information
Supplementary Information (download DOCX )
This file contains the Supplementary Discussion, Supplementary References, Supplementary Fig. 1 and Supplementary Tables 1–3.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, J., Perren, J.O., Rogers, C.M. et al. Endogenous DNA damage at sites of terminated transcripts. Nature 640, 240–248 (2025). https://doi.org/10.1038/s41586-024-08578-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-024-08578-4
This article is cited by
-
R-loop editing by DNA cytosine deaminase APOBEC3B modulates the activity of oestrogen receptor enhancers
Nature Communications (2026)