Abstract
Chromosomal linkages formed through crossover recombination are essential for the accurate segregation of homologous chromosomes during meiosis1. The DNA events of recombination are linked to structural components of meiotic chromosomes2. Imperatively, the biased resolution of double Holliday junction (dHJ) intermediates into crossovers3,4 occurs within the synaptonemal complex (SC), the meiosis-specific structure that mediates end-to-end synapsis of homologues during the pachytene stage5,6. However, the role of the SC in crossover-specific dHJ resolution remains unclear. Here we show that key SC components function through dependent and interdependent relationships to protect dHJs from aberrant dissolution into non-crossover products. Conditional ablation experiments reveal that cohesin, the core of SC lateral elements, is required to maintain both synapsis and dHJ-associated crossover recombination complexes (CRCs) during pachytene. The SC central region transverse-filament protein is also required to maintain CRCs. Reciprocally, the stability of the SC central region requires the continuous presence of CRCs effectively coupling synapsis to dHJ formation and desynapsis to resolution. However, dHJ protection and CRC maintenance can occur without end-to-end homologue synapsis mediated by the central element of the SC central region. We conclude that local ensembles of SC components are sufficient to enable crossover-specific dHJ resolution to ensure the linkage and segregation of homologous chromosomes.
Similar content being viewed by others
Main
During meiotic prophase I, cohesin complexes connect sister chromatids and mediate their organization into linear arrays of chromatin loops tethered to a common axis2,5,7,8,9. These cohesin-based axes define interfaces for the pairing and synapsis of homologous chromosomes that culminates in the formation of SCs. An SC is a tripartite structure comprising the two juxtaposed homologue axes, now called lateral elements, connected by a central lattice of transverse filaments5,6. Extension of this lattice to achieve full synapsis requires an additional central element complex5,6,10 (Extended Data Fig. 1a). Meiotic recombination facilitates pairing and synapsis between homologous chromosomes and then connects them through crossing over. These connections are necessary for accurate segregation during the first meiotic division1. To this end, the DNA events of recombination are physically and functionally linked to underlying chromosome structures2. The protein complexes that catalyse DNA double-strand breaks (DSBs) and subsequent strand exchange are tethered to homologue axes. The ensuing joint molecule intermediates and their associated recombination complexes interact with the central region of the SC. A subset of recombination events is assigned a crossover fate with a tightly regulated distribution to ensure that each chromosome pair receives at least one2. At designated sites, nascent joint molecules mature into dHJs that then undergo biased resolution specifically into crossovers3,4. These steps occur in the context of the SC central region and associated CRCs. The post-synapsis roles of SC components in crossing over remain unclear, particularly whether the SC functions after dHJ formation to facilitate crossover-specific resolution.
Cohesin and crossover-biased dHJ resolution
To address the role of SC lateral elements in crossover-specific dHJ resolution, the cohesin core was ablated using the auxin-inducible degron (AID) system11 to conditionally degrade Rec8 (the meiosis-specific kleisin subunit) at the time of dHJ resolution (Fig. 1a). Real-time inactivation of Rec8–cohesin circumvents severe early meiotic defects of cohesin mutants in the formation and processing of DSBs12,13 (Extended Data Fig. 2a–d). In all experiments, cell cultures were synchronized at the resolution transition using an oestradiol-inducible allele of NDT80 (NDT80-IN) that reversibly arrests cells at the pachytene stage, in which chromosomes are fully synapsed and dHJs are poised for resolution14.
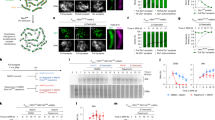
a, Schematic of the experimental strategy. Top, homologue synapsis with chromatin (blue), homologue axes (green) and the SC central region (pink). The X on the far right of the cartoon indicates a crossover formed after Ndt80 is expressed. Middle, cell synchronization at dHJ resolution transition using the inducible NDT80-IN allele and conditional degradation of target proteins using the AID system. Bottom, DNA events of meiotic recombination. Only the two chromosomes engaged in recombination are shown. b, Western blot analysis of Rec8-AID from subcultures (released from arrest) with or without the addition of auxin at 7 h. Arp7 was used as the loading control. c, Quantification of Rec8-AID levels from the experiment in b. d, Quantification of nuclear divisions (MI ± MII, cells with two and four nuclei) from REC8-AID subcultures with or without auxin. e, Representative 1D gel Southern blot analysis of crossover (top) and non-crossover (bottom) formation at the HIS4::LEU2 recombination hotspot from REC8-AID subcultures with or without auxin. The dagger symbol indicates cross-hybridizing background bands. f, Quantification of crossover and non-crossover levels from the experiments in e. g, Quantification of the final levels of crossovers and non-crossovers at 11 h from REC8-AID subcultures with or without auxin. Mean ± s.d. of three independent experiments; ****P < 0.0001, **P = 0.0038, two-tailed unpaired t-test. h, Representative 2D gel Southern blot analysis of joint molecules from REC8-AID subcultures with or without auxin and before and after release from arrest at 7 and 11 h, respectively. The top-left panel highlights joint molecule species: IH-dHJ, inter-homologue double Holliday junction; IS-JMs, inter-sister joint molecules; mc-JMs, 3-chromatid and 4-chromatid joint molecules; SEI, single-end invasion. i, Quantification of total joint molecule levels from experiments in h. Mean ± s.d. of three independent experiments; NS, not significant, P = 0.4561, two-tailed unpaired t-test.
Auxin and oestradiol were simultaneously added to degrade Rec8-AID while releasing cells from pachytene arrest and triggering dHJ resolution (Fig. 1b–d). Without auxin, Rec8-AID levels remained high until cells had completed meiotic divisions (meiosis I (MI) + meiosis II (MII)). By contrast, with auxin, Rec8-AID was completely degraded within 60 min and meiotic divisions were delayed by around 2 h (Fig. 1c,d). Crossing over at the HIS4::LEU2 DSB hotspot15 was reduced by 70% after Rec8-AID degradation and was accompanied by a 77% increase in non-crossover products (Fig. 1e–g; note that our assay reports a subset of non-crossover gene-conversion products and not absolute levels of non-crossovers15). The opposing changes in levels of crossovers and non-crossovers and the comparable kinetics of their formation with and without auxin (Fig. 1e,f) suggest that dHJs are efficiently resolved when Rec8-AID is degraded, but their resolution fate is reversed. This inference was confirmed by 2D gel electrophoresis and Southern blotting (Fig. 1h,i). Thus, Rec8-based cohesin is required after dHJ formation to facilitate crossover-specific resolution. Degradation of a core cohesin subunit, Smc3, confirmed that the cohesin complex, not just Rec8, is required for this process (Extended Data Fig. 2e–h). However, the mitotic kleisin Mcd1 (known as RAD21 in other species) was not required for crossover-specific dHJ resolution, a result that indicates that this function is specific to Rec8–cohesin (Extended Data Fig. 2i,j). The separase Esp1 also did not influence crossover-specific resolution (Extended Data Fig. 2i), a finding consistent with previous observations16 showing that a separase-resistant allele of Rec8 does not affect crossing over.
Distinct functions of cohesin and the Smc5/6 complex
Two classes of meiotic crossovers are distinguished by their dependencies on enzymes that resolve joint molecules. Class I crossovers depend on the crossover-specific dHJ resolvase defined by the endonuclease MutLγ (a complex of Mlh1 and Mlh3)3,4,17,18,19. By contrast, minority class II crossovers require structure-selective endonucleases, primarily Mus81–Mms4 (known as MUS81–EME1 in other species)3,20,21,22. Epistasis analysis revealed that cohesin and MutLγ act in the same crossover resolution pathway (Fig. 2a,b), whereas Mus81–Mms4 acts in a parallel pathway (Fig. 2c,d; in these experiments, Mms4-AID was degraded at the resolution transition in a yen1Δ background, in which the backup resolvase Yen1 is deleted). Crossover levels were indistinguishable after Rec8-AID degradation in the presence or absence of MutLγ (in cells with a mlh3Δ mutation; Fig. 2b). However, in contrast to the REC8-AID single mutant in which non-crossovers were increased (Fig. 1g), non-crossovers in REC8-AID mlh3Δ strains were reduced to control levels (Fig. 2b). This result suggests that MutLγ influences gene conversion of the central polymorphism used to detect non-crossovers15. That is, non-crossover products that contain unrepaired heteroduplex DNA or undergo restoration of the central polymorphism, which are undetectable with our assay, are increased in REC8-AID mlh3Δ cells.
In all experiments shown in this figure, cells were released from NDT80-IN arrest. a, Representative 1D gel Southern blot analysis of crossover (top) and non-crossover (bottom) formation in control (REC8-AID no auxin), mlh3Δ, REC8-AID (with auxin) and REC8-AID mlh3Δ (with auxin) strains. b, Final levels of crossovers and non-crossovers at 13 h from the indicated strains. Mean ± s.d. of three independent experiments. Statistical comparisons are with the control unless indicated otherwise. Dunnett’s multiple comparisons test, ****P < 0.0001, NS, P = 0.5281, **P = 0.0034 (REC8-AID versus control), **P = 0.0026 (REC8-AID versus REC8-AID mlh3Δ). c, Representative 1D gel Southern blot analysis of crossover (top) and non-crossover (bottom) formation in control, MMS4-AID yen1Δ (with auxin), SMC3-AID (with auxin) and SMC3-AID MMS4-AID yen1Δ (with auxin) strains. d, Final levels of crossovers and non-crossovers at 11 h from the indicated strains. Mean ± s.d. of n = 4 control, n = 5 MMS4-AID yen1Δ, n = 4 SMC3-AID and n = 3 SMC3-AID MMS4-AID yen1Δ independent experiments. Statistical comparisons are with the control unless indicated otherwise. For crossovers, Tukey’s multiple comparisons test was performed; for non-crossovers, Dunnett’s multiple comparisons test was performed. ****P < 0.0001, ***P = 0.0003, **P = 0.0066, *P = 0.0385 (control versus SMC3-AID), *P = 0.0488 (control versus SMC3-AID MMS4-AID yen1Δ), NS, P = 0.7806. e, Representative 1D gel Southern blot analysis of crossover (top) and non-crossover (bottom) formation in control, NSE4-AID (with auxin), REC8-AID (with auxin) and REC8-AID NSE4-AID (with auxin) strains. f, Final levels of crossovers and non-crossovers at 11 h from experiments in c. Mean ± s.d. of three independent experiments. Statistical comparisons are with the control unless indicated otherwise. Dunnett’s multiple comparisons test, ****P < 0.0001, **P = 0.0049, ***P = 0.0022 (non-crossovers NSE4-AID versus control), ***P = 0.0002 (non-crossovers REC8-AID versus control); NS, P = 0.9955 (non-crossovers REC8-AID NSE4-AID versus control). g, Representative Southern blot 2D gel images at 11 h from no-auxin control, NSE4-AID (with auxin), REC8-AID (with auxin) and REC8-AID NSE4-AID (with auxin). h, Quantification of total joint molecule levels from the experiments in e. Mean ± s.d. of three independent experiments. Tukey’s multiple comparison test, ****P < 0.0001, NS, P = 0.3320.
Mus81–Mms4 works in conjunction with a second SMC complex, Smc5/6 (refs. 21,22,23,24). Consistently, degradation of Nse4-AID (an essential subunit of Smc5/6) during the resolution transition reduced crossing over to the same extent as when Mus81–Mms4 was inactivated by degrading Mms4-AID (Fig. 2c–f). Although non-crossovers were not reduced when Mus81–Mms4 was inactivated, Nse4-AID degradation also reduced non-crossovers by 2.1-fold (Fig. 2e,f). This finding indicates that Smc5/6 controls an additional, Mus81–Mms4-independent pathway of non-crossover formation, most probably through the regulation of a subpopulation of the Sgs1–Top3–Rmi1 complex (Extended Data Figs. 1 and 3).
We next aimed to confirm that Smc5/6 and cohesin act in independent pathways of resolution. Simultaneous degradation of Nse4-AID and Rec8-AID resulted in an additive reduction in crossing over; and non-crossover levels that are the combination of those observed when Nse4-AID and Rec8-AID were individually degraded (Fig. 2e,f). Note that simultaneous degradation of Nse4-AID and Smc3-AID gave analogous results (Extended Data Fig. 4). Further distinguishing these two pathways, a subset of joint molecules remained unresolved when Nse4-AID was degraded alone, whereas resolution remained efficient when cohesin was inactivated through the degradation of either Rec8-AID or Smc3-AID (Figs. 1g,h and 2g,h and Extended Data Fig. 4e,f). These results show that Smc5/6 is essential for the resolution of a subset of joint molecules into both crossovers and non-crossovers. By contrast, Rec8–cohesin promotes crossover-specific dHJ resolution but is not required for resolution per se.
Cohesin, synapsis and crossover complexes
To begin to understand how cohesin facilitates crossover-specific dHJ resolution, Rec8-AID was degraded while pachytene arrest was maintained (no induction of NDT80-IN). Chromosomes were then analysed by immunostaining for markers of homologue axes (Rec8 and Red1), the SC central region (Zip1) and CRCs (Msh5 and Zip3)25 (Fig. 3a). In no-auxin controls, linear Rec8-stained and discontinuous Red1-stained structures colocalized, and synapsed homologues (indicated by lines of Zip1 staining) were decorated with foci of Msh5 and Zip3 (Fig. 3a). One hour after the addition of auxin, Rec8 structures were lost and characteristic features of pachytene chromosomes disappeared. That is, SCs disassembled and Zip1 staining was now confined to a few foci and larger structures resembling polycomplexes that are diagnostic of defective synapsis. Moreover, the numbers of Red1-stained structures were reduced by twofold, a result consistent with the core function of cohesin in organizing homologue axes26 and directly recruiting Red1 (ref. 27). Furthermore, CRCs were dissociated, as indicated by the loss of Msh5 and Zip3 foci (Fig. 3a,b). Thus, cohesin is required to maintain the integrity of pachytene chromosome structures, a finding consistent with analyses of meiotic cohesin in Caenorhabditis elegans28.
a, Representative images of surface-spread meiotic nuclei from REC8-AID cultures in which pachytene arrest was maintained. Cells were sampled 1 h after the addition of auxin or DMSO vehicle at 7 h and immunostained for the indicated markers. Scale bars, 1 μm. b, Quantification of structures immunostained for Red1, Msh5 and Zip3 from experiments in a. Mean ± s.d. Red1 staining, n = 30 and 37 nuclei; Msh5 staining, n = 64 and 57 nuclei; Zip3 staining, n = 37 and 31 nuclei (n indicates the number of nuclei analysed for the auxin and no-auxin experiments, respectively). Unpaired two-tailed t-test. ****P < 0.0001.
Interdependent functions of SC components
To discern which cohesin-dependent features of pachytene chromosomes are important for crossover-specific dHJ resolution, the SC transverse-filament protein Zip1 (refs. 29,30) and the pro-crossover factor MutSγ (a complex of Msh4 and Msh5) were inactivated as cells were released from pachytene arrest (Fig. 4a,b). MutSγ binds and stabilizes nascent joint molecules to promote homologue synapsis and dHJ formation31,32,33. The degradation of either Zip1-AID or Msh4-AID led to phenotypes similar to those resulting from the loss of cohesin. That is, reduced crossovers and increased non-crossovers (Fig. 4c,d), which indicated that these factors have continued roles after dHJs have formed to maintain their crossover resolution fate.
a, Western blot analysis of Zip1-AID from subcultures (released from arrest) with or without the addition of auxin at 7 h. Arp7 was used as the loading control. b, Western blot analysis of Msh4-AID from subcultures (released from arrest) with or without the addition of auxin at 7 h. Arp7 was used as the loading control. c, Representative 1D gel Southern blot analysis of crossover (top) and non-crossover (bottom) formation in control, ZIP1-AID (with auxin) and MSH4-AID (with auxin) strains. d, Final levels of crossovers and non-crossovers at 11 h from the indicated strains. Mean ± s.d. of n = 6 control, n = 3 ZIP1-AID and n = 6 MSH4-AID independent experiments. Statistical comparisons are with the control unless indicated otherwise. Tukey’s multiple comparisons test, ****P < 0.0001, ***P = 0.0003, **P = 0.0087, NS, P = 0.2321 (ZIP1-AID versus MSH4-AID for crossover analysis), P = 0.0524 (ZIP1-AID versus MSH4-AID for non-crossover analysis). e, Representative images of surface-spread meiotic nuclei from pachytene-arrested ZIP1-AID cells sampled 1 h after the addition of auxin or DMSO vehicle at 7 h and immunostained for the indicated markers. f, Quantification of foci immunostained for Msh5 and Zip3 from the experiments in e. Mean ± s.d. Msh4 staining, n = 44 and 62 nuclei; Zip3 staining, n = 40 and 41 nuclei (n indicates the number of nuclei analysed for the auxin and no-auxin experiments, respectively). Unpaired two-tailed t-test, ****P < 0.0001. g, Representative images of surface-spread meiotic nuclei from pachytene-arrested MSH4-AID cells sampled 1 h after the addition of auxin or DMSO vehicle at 7 h and immunostained for the indicated markers. h, Quantification of foci stained for Msh4 and Zip3 from the experiments in g. Mean ± s.d. Msh4 staining, n = 39 and 35 nuclei; Zip3 staining, n = 51 and 49 nuclei (n indicates the number of nuclei analysed for the auxin and no-auxin experiments, respectively). Unpaired two-tailed t-test, ****P < 0.0001. Scale bars, 1 µm (e,g).
These results suggest that cohesin may facilitate crossover-specific dHJ resolution by stabilizing the SC central region, which in turn stabilizes CRCs. This interpretation was tested by immunostaining the chromosomes of pachytene-arrested cells after the degradation of Zip1-AID (Fig. 4e,f) or Msh4-AID (Fig. 4g,h). In budding yeast, full synapsis of the 16-homologue pairs is indicated by around 16 Zip1-stained lines and discontinuous linear Red1-stained structures (Fig. 4e,g, left). When Zip1-AID was degraded, desynapsis ensued, whereby Zip1 lines were lost and the numbers of linear Red1-stained structures approximately doubled, which is indicative of unsynapsed but intact homologue axes (Fig. 4e). As predicted, CRCs marked by Msh5 and Zip3 foci were diminished (Fig. 4e,f). As Zip3 and Msh4 protein levels remained high in these cells (Extended Data Fig. 5), we inferred that CRCs are disassembled when Zip1-AID is degraded. Notably, desynapsis also occurred when Msh4-AID was degraded, which revealed that the maintenance of SCs during pachytene requires the continued presence of CRCs (Fig. 4g). Zip3 foci were also lost when Msh4-AID was degraded. This result highlights the interdependence between Holliday junction binding (MutSγ) and the regulatory (Zip3) components of CRCs (Fig. 4g,h).
dHJs are protected from aberrant dissolution
dHJs normally remain stable in pachytene-arrested NDT80-IN cells because the expression of the polo-like kinase Plk1 (also known as Cdc5), which activates dHJ resolution, requires the transcription factor Ndt80 (refs. 14,34). However, dHJ levels decreased by around threefold when Rec8-AID was degraded but pachytene arrest was maintained (Fig. 5b,c). This result implied that pachytene chromosome structures protect dHJs against aberrant resolution by a Plk1-independent resolution activity. A good candidate for this activity is the Sgs1–Top3–Rmi1 (STR) complex, the budding yeast orthologue of the human BLM complex BLM–TOPIIIα–RMI1/2, and a robust decatenase enzyme that can dissolve dHJs specifically into non-crossover products35,36,37,38,39. Confirming this prediction, dHJs were stabilized when Rec8-AID and Top3-AID were simultaneously degraded (Fig. 5a–c). Note that the degradation of both Smc3-AID and Top3-AID gave similar results (Extended Data Fig. 6). Moreover, dHJ resolution after Rec8-AID degradation alone produced non-crossover products that did not form when Top3-AID was also degraded (Fig. 5d,e).
a, Western blot analysis of Top3-AID and Rec8-AID from subcultures with or without the addition of auxin at 7 h and pachytene arrest was maintained. Arp7 was used as the loading control. The degradation-resistant fraction of Top3-AID is mitochondrial. b, Representative 2D gel Southern blot analysis of joint molecules from REC8-AID and REC8-AID TOP3-AID strains with and without the addition of auxin. c, Quantification of total joint molecule levels from the indicated strains. d, Western blot analysis of Top3-AID and Rec8-AID from subcultures with or without the addition of auxin at 7 h with release from pachytene arrest. e, Representative 1D gel Southern analysis of crossover (top) and non-crossover (bottom) formation in control, TOP3-AID (with auxin), REC8-AID (with auxin) and REC8-AID TOP3-AID (with auxin) strains. f, Final levels of crossovers and non-crossovers at 11 h from experiments in g. Mean ± s.d. of three independent experiments. Statistical comparisons are with the control unless indicated otherwise. Tukey’s multiple comparisons test, ****P < 0.0001, **P = 0.0018 (REC8-AID versus REC8-AID TOP3-AID for crossover comparison),*P = 0.0489 (control versus REC8-AID TOP3-AID for crossover comparison), NS, P = 0.9219 (control versus TOP3-AID for crossover comparison), **P = 0.0021 (control versus REC8-AID for non-crossover comparison), **P = 0.0034 (REC8-AID versus REC8-AID TOP3-AID for non-crossover comparison), NS, P = 0.4753 (control versus TOP3-AID for non-crossover comparison), P = 0.9721 (control versus REC8-AID TOP3-AID for non-crossover comparison).
The stability of dHJs in pachytene-arrested cells when Rec8-AID and Top3-AID were degraded together indicated that resolution was again dependent on Plk1 and suggests that the crossover defect resulting from Rec8-AID degradation may be rescued. Indeed, expression of NDT80-IN while simultaneously degrading Rec8-AID and Top3-AID resulted in a 2.6-fold increase in crossovers relative to the degradation of Rec8-AID alone, whereas non-crossovers decreased by 1.6-fold (Fig. 5f–h). Note that the degradation of both Rec8-AID and Sgs1-AID gave similar results (Extended Data Fig. 7). Moreover, stabilization of dHJs and partial rescue of the crossover defects resulting from Zip1-AID or Msh4-AID degradation were also seen when Top3-AID was simultaneously degraded (Extended Data Figs. 8 and 9). Partial restoration of crossover levels is probably explained by our observation that when STR function was ablated, essentially all resolution was mediated by the Smc5/6–Mus81–Mms4 pathway (Extended Data Fig. 3).
We wondered whether stabilizing dHJs also rescues the cytological defects that result from the degradation of Rec8-AID, Zip1-AID or Msh4-AID. To this end, Top3-AID was degraded together with Rec8-AID, Zip1-AID or Msh4-AID in pachytene-arrested cells, and chromosomes were immunostained for markers of synapsis (Zip1) and CRCs (Msh4 and Zip3) (Extended Data Fig. 10). In each case, cytological phenotypes were largely indistinguishable from those observed when Rec8-AID, Zip1-AID or Msh4-AID were degraded alone. SCs and CRCs disassembled after the degradation of both Rec8-AID and Top3-AID (Extended Data Fig. 10a,b), and CRCs were dissociated when Zip1-AID and Top3-AID were degraded together (Extended Data Fig. 10c,d). Moreover, SCs disassembled after the degradation of both Msh4-AID and Top3-AID (Extended Data Fig. 10e,f). Thus, the maintenance of inter-homologue DNA connections through the stabilization of dHJs does not bypass the interdependencies among cohesin, SCs and CRCs.
We conclude that the key components of pachytene chromosomes—cohesin-based homologue axes, SC transverse filaments and CRCs—protect crossover-designated dHJs from being aberrantly dissolved into non-crossovers by the STR complex (Extended Data Fig. 1b).
Full synapsis is dispensable for crossovers
High levels of crossing over can occur without end-to-end homologue synapsis in cells that lack the SC central element, which is required to mature and extend the transverse filament lattice to achieve full synapsis5,6,10,40. To determine whether the maintenance of full synapsis is required for crossover-specific dHJ resolution, the central element protein Ecm11 (ref. 10) was ablated as cells were released from pachytene arrest (Fig. 6). Crossover and non-crossover levels were indistinguishable from no-auxin controls when Ecm11-AID was degraded (Fig. 6a,b). This result was in marked contrast to the phenotypes seen after the loss of Rec8, Zip1 or Msh4 (Figs. 1 and 4). Moreover, in arrested NDT80-IN cells, dHJs remained stable after Ecm11-AID degradation (Fig. 6c,d).
a, Representative 1D gel Southern blot analysis of crossover (top) and non-crossover (bottom) formation from ECM11-AID subcultures (released from arrest) with or without auxin. b, Final levels of crossovers and non-crossovers at 11 h from the experiments in a. Mean ± s.d. of three independent experiments. NS, P > 0.9999, two-tailed, Mann–Whitney U-test. c, Representative 2D gel Southern blot analysis of joint molecules from ECM11-AID subcultures with and without the addition of auxin and pachytene arrest was maintained. d, Quantification of total joint molecule levels from the experiment in c. e,f, Representative images of surface-spread meiotic nuclei from pachytene-arrested ECM11-AID cells sampled 1 h after the addition of auxin or DMSO vehicle at 7 h and immunostained for the indicated markers. Scale bars, 1 μm. g, Quantification of foci stained for Zip3 and Msh5 from the ECM11-AID experiments in e and f, and comparison with corresponding data from ZIP1-AID (Fig. 4f). Mean ± s.d. Zip3 staining, n = 61, 51 and 41 nuclei; Msh5 staining, n = 45, 45 and 62 nuclei (n indicates the number of nuclei analysed for the auxin, no-auxin and ZIP1-AID experiments, respectively). Unpaired two-tailed t-test, ****P < 0.0001. h,i, Quantification of colocalization (mean ± s.d.) between foci immunostained for Zip1 and Zip3 (h, n = 40 nuclei), and Zip1 and Msh5 (i, n = 31 nuclei) from auxin-treated ECM11-AID nuclei in e and f.
Immunostaining revealed that SCs disassembled after Ecm11-AID degradation in arrested NDT80-IN cells, although numerous foci of Zip1 persisted (Fig. 6e). Thus, the central element is required to both establish and maintain full synapsis. Msh5 and Zip3 foci were maintained at about 50% of no-auxin control levels (Fig. 6f,g), which was in contrast to the almost complete loss seen after the degradation of Rec8-AID, Zip1-AID or Msh4-AID. Moreover, the remaining Msh5 and Zip3 foci showed high degrees of colocalization with residual Zip1 structures (Fig. 6h,i). These observations suggest that although mature SCs may stabilize a subset of CRCs, local ensembles of Zip1 and CRCs are sufficient to protect dHJs and support crossover-specific resolution.
Discussion
The SC is an ancient structure that evolved with the emergence of sexual reproduction. Several functions are attributed to the SC. The assembly of homologue axes and pairwise synapsis of homologues establishes topological order in the nucleus. Once assembled, the SC also prevents the formation of DSBs and facilitates their repair to attenuate DNA-damage checkpoint signalling. At designated crossover sites, SC components facilitate dHJ formation5. This study, together with a companion paper41, reveals that key SC components, but not necessarily end-to-end synapsis per se, also enable crossover-specific resolution of dHJs by protecting them from unscheduled dissolution into non-crossovers by the conserved STR/BLM complex (Extended Data Fig. 1b,c). Failure of this crucial final step of meiotic recombination can result in unlinked univalent chromosomes that are prone to missegregation, which results in aneuploid gametes that are associated with infertility, miscarriage and congenital disease in humans.
Our data indicate that dHJs formed at crossover-designated recombination sites do not possess an intrinsic structure or topology that constrains their resolution fate to ensure crossing over. We previously proposed a model of crossover-specific resolution whereby asymmetric loading of PCNA during DNA synthesis associated with dHJ formation subsequently directs strand-specific nicking by the endonuclease MutLγ on both sides of the two Holliday junctions but distal to the exchange points4. This pattern of incisions always specifies a crossover outcome when the nicked dHJs are disassembled by the STR/BLM complex. Our two-step resolution model reconciles the counterintuitive pro-crossover role of STR/BLM during meiosis, which is in contrast to its well-characterized anticrossover roles in unwinding D-loops and dissolving unnicked dHJs into non-crossover products (Extended Data Figs. 1b and 3b,c). In this framework, we propose that mature dHJs are unnicked and therefore vulnerable to STR/BLM-catalysed dissolution until MutLγ is activated through the Ndt80-dependent expression of Plk1. By preventing STR/BLM from acting on dHJs until they have been nicked by MutLγ, SC components impose the sequential steps required for crossover-specific resolution (Extended Data Fig. 1b).
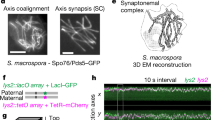
We suggest that local ensembles of cohesin and SC transverse filaments can be sufficient to protect dHJs by stabilizing CRCs, components of which can directly bind Holliday junctions, including MutSγ, MutLγ, Zip2–Spo16 (known as SHOC1–SPO16 in mammals) and Mer3 (known as HFM1 in mammals)25,42, and may directly compete with STR/BLM for binding dHJs and/or may constrain its activity. This proposal is consistent with electron microscopy visualization of late recombination nodules associated with short patches of SC in Sordaria macrospora43 and with the inference that in C. elegans, the cohesin local to designated crossover sites is distinctly regulated28. Moreover, components of the SC central region help recruit and stabilize CRCs32,44,45,46,47. Also in C. elegans, SC central region proteins seem to assemble transient crossover-specific compartments or ‘bubbles’ that may protect CRCs until crossover-specific resolution is executed45.
Our study, together with a companion paper41, also found that CRCs are required to maintain synapsis during pachytene. This observation is consistent with studies indicating that the SC central region is initially dynamic and labile but becomes more stable, contingent on the development of CRCs48,49,50,51. Notably, new subunits are continually incorporated into the SC central region after synapsis48,51. Moreover, in budding yeast, incorporation of new Zip1 molecules occurs predominantly at CRC sites48. Given that Zip1 seems to directly recruit CRC components46,52, this suggests a possible mechanism for the mutual stabilization of SCs and CRCs. This relationship may result in SCs with non-uniform structure and stability48, which could influence resolution fates and explain why sites of crossing over are the last to desynapse during diplotene53. The interdependence and dynamic nature of SCs and CRCs will render chromosomal interactions readily reversible until dHJs are resolved into crossovers. These attributes could help adjust and proofread homologous synapsis, minimize and resolve synaptic interlocks54 and preserve genome stability by destabilizing interactions between non-allelic and diverged sequences.
Importantly, dependency between synapsis and CRCs probably helps impose the ordered sequence of events required to ensure crossing over between homologues. At designated crossover sites, CRCs promote and maintain homologue synapsis, facilitate the formation of dHJs and protect them from aberrant dissolution. In turn, synapsis globally attenuates DSB formation55, thereby diminishing the DNA-damage kinase signalling that inhibits Ndt80 activity. Ndt80-dependent expression of Plk1 then activates crossover-specific dHJ resolution, CRCs disassemble and desynapsis ensues (Fig. 1c,d).
Our results also revealed that two SMC complexes mediate essentially all Plk1-dependent joint-molecule resolution during meiosis (Extended Data Fig. 1c). Rec8–cohesin is required to maintain synapsis and CRCs and thereby protects dHJs to facilitate their crossover-specific resolution. Whether local or global functions of Rec8–cohesin are important for crossover-specific dHJ resolution, and the roles of cohesive versus chromatin-loop-organizing populations of Rec8–cohesin, remain unclear. A distinct Smc5/6-dependent pathway is essential for resolution through Mus81–Mms4 and a subpopulation of STR/BLM, producing a mixture of crossovers and non-crossovers. Despite these distinctions, Rec8–cohesin and Smc5/6 could have common functions to constrain favourable joint-molecule topology56 and control access by resolving enzymes.
Methods
Data reporting
No statistical methods were used to predetermine sample sizes. The experiments were not randomized, and the investigators were not blinded to allocation during experiments. Blinding was used during outcome assessment for the cytology experiments.
Yeast strains
For full genotypes, see Supplementary Table 1. The AID system11 was optimized for meiosis by replacing the promoter of the pADH1-OsTIR1 cassette with the CUP1 promoter36. Carboxy-terminal fusion of a minimal AID to targeted proteins was constructed using the plasmid pHyg-AID*−9Myc as the template for PCR-mediated allele replacement57. To construct an internal degron allele of ZIP1, AID sequences were inserted into plasmid pMPY-3×HA and integrated after codon 700 through PCR epitope tagging58. The primers used to construct AID alleles are listed in Supplementary Table 2. The oestrogen-inducible NDT80-IN GAL4-ER system has been previously described59,60,61.
Meiotic time courses and DNA physical assays
Detailed protocols for meiotic time courses and DNA physical assays at the HIS4::LEU2 locus have been previously described62. At 6.5 h after the induction of meiosis, CuSO4 (100 mM stock in dH2O) was added to obtain a final concentration of 50 μM to induce the expression of pCUP1-OsTIR1 (which encodes the E3 ligase Tir1) and cell cultures were split. For experiments in which NDT80-IN was induced, oestradiol (5 mM stock, Sigma E2758 (β-estradiol) in ethanol) was added at 7 h to obtain a final concentration of 1 μM to both subcultures. Simultaneously, auxin (3-indoleacetic acid, Sigma, 13750, 2 M stock in DMSO) was added to one subculture to obtain a final concentration of 2 mM; an equivalent volume of DMSO was added to the no-auxin control subculture. At 7.5 h, auxin was added again at 1 mM. To analyse the timing and efficiency of meiotic divisions and sporulation, cells were fixed in 40% ethanol, 0.1 M sorbitol, stained with DAPI, and around 200 cells were categorized for each time point. For imaging, DAPI-stained cells were mounted in antifade (Vectashield, Vector Laboratories) and digital images were captured using a Zeiss Axioplan II microscope, Hamamatsu ORCA-ER CCD camera and Volocity software.
Chromosome spreading and immunofluorescence microscopy
Cell samples were collected and processed for chromosome spreading and immunostaining essentially as previously described63. The following primary antibodies were provided by A. Shinohara: chicken anti-Red1 (1:500 dilution); rabbit anti-Msh5 (1:750); and rabbit anti-Zip3 (1:500). Guinea pig anti-Zip1 (1:400) was a gift from S. Keeney. Monoclonal anti-cMyc (1:1,000, Roche 11667149001) was used to detect AID–9myc fused proteins. All primary antibodies were incubated overnight at room temperature in 100 μl TBS–BSA buffer (10 mM Tris pH 7.5, 150 mM NaCl and 1% BSA). The secondary antibodies anti-rabbit 568 (A11036, Molecular Probes, 1:1,000), anti-mouse 488 (A11029, Molecular Probes, 1:1,000), anti-rabbit 647 (A21245, Invitrogen) and anti-guinea pig 555 (A21435, Life Technologies) were incubated for 1 h at 37 °C. Coverslips were mounted with Prolong Gold antifade reagent (Invitrogen, P36930). Digital images were captured using a Zeiss Axioplan II microscope, Hamamatsu ORCA-ER CCD camera and analysed using Volocity software. Scatterplots were generated using the GraphPad program in Prism.
Western blot analysis
Whole-cell extracts were prepared using a TCA extraction method essentially as previously described64. Samples were analysed by standard SDS–PAGE and western blotting using the following primary antibodies: monoclonal anti-cMyc (1:1,000, Roche, 11667149001); monoclonal anti-HA (1:1,000, Sigma, 11583816001); goat polyclonal Arp7 (1:10,000, Santa Cruz, SC-8960); goat polyclonal Zip1 (1:500, Santa Cruz, y-N16); rabbit anti-Msh4 (1:500); rabbit anti-Msh5 (1:500); and rabbit anti-Zip3 (1:500; Msh4, Msh5 and Zip3 antibodies were a gift from A. Shinohara). The secondary antibodies (1:5,000) were IRDye 800CW donkey anti-mouse IgG (LI-COR, 925-32212), IRDye 680LT donkey anti-goat IgG (LI-COR, 925-68024), IRDye 680LT donkey anti-rabbit IgG (LI-COR, 925-68023) and IRDye 800CW donkey anti-rabbit IgG (LI-COR, 925-32213). Western blots were imaged on an Odyssey Infrared Imager (LI-COR), and quantification of protein bands was performed using Image Studio Lite (v.5.0.021) software.
Statistical analysis and reproducibility
Statistical analyses were performed using Prism 8 (GraphPad Software). For bar graphs and scatter plots comparing two samples (aggregated from three or more replicate experiments), unpaired t-tests were performed. For multiple comparisons, one-way analysis of variance was performed (Tukey’s or Dunnett’s tests were used depending on the specific comparisons being made). For scatter plots and bar graphs, error bars show the mean value with standard deviations. Western blots are representative of at least three repeats (Figs. 1b, 4ab and 5a,d and Extended Data Figs. 2a, 3a,d,e, 4a,b, 6a,d, 8a,d and 9a,d). Representative immunofluorescence images were chosen from more than 30 samples analysed from at least three biological replicates (Fig. 3a, 4e,g and 6e,g and Extended Data Fig. 10a,c,e).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Relevant data generated or analysed during this study are included in this Article and its Supplementary Information files. Biological materials are available from the corresponding author.
References
Hunter, N. Meiotic recombination: the essence of heredity. Cold Spring Harb. Perspect. Biol. 7, a016618 (2015).
Zickler, D. & Kleckner, N. Meiosis: dances between homologs. Annu. Rev. Genet. 57, 1–63 (2023).
Zakharyevich, K., Tang, S., Ma, Y. & Hunter, N. Delineation of joint molecule resolution pathways in meiosis identifies a crossover-specific resolvase. Cell 149, 334–347 (2012).
Kulkarni, D. S. et al. PCNA activates the MutLγ endonuclease to promote meiotic crossing over. Nature 586, 623–627 (2020).
Zickler, D. & Kleckner, N. Recombination, pairing, and synapsis of homologs during meiosis. Cold Spring Harb. Perspect. Biol. 7, a016626 (2015).
Adams, I. R. & Davies, O. R. Meiotic chromosome structure, the synaptonemal complex, and infertility. Annu. Rev. Genomics Hum. Genet. 24, 35–61 (2023).
Ito, M. & Shinohara, A. Chromosome architecture and homologous recombination in meiosis. Front. Cell Dev. Biol. 10, 1097446 (2022).
Schalbetter, S. A., Fudenberg, G., Baxter, J., Pollard, K. S. & Neale, M. J. Principles of meiotic chromosome assembly revealed in S. cerevisiae. Nat. Commun. 10, 4795 (2019).
Muller, H. et al. Characterizing meiotic chromosomes’ structure and pairing using a designer sequence optimized for Hi-C. Mol. Syst. Biol. 14, e8293 (2018).
Humphryes, N. et al. The Ecm11–Gmc2 complex promotes synaptonemal complex formation through assembly of transverse filaments in budding yeast. PLoS Genet. 9, e1003194 (2013).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Kugou, K. et al. Rec8 guides canonical Spo11 distribution along yeast meiotic chromosomes. Mol. Biol. Cell 20, 3064–3076 (2009).
Kim, K. P. et al. Sister cohesion and structural axis components mediate homolog bias of meiotic recombination. Cell 143, 924–937 (2010).
Sourirajan, A. & Lichten, M. Polo-like kinase Cdc5 drives exit from pachytene during budding yeast meiosis. Genes Dev. 22, 2627–2632 (2008).
Owens, S., Tang, S. & Hunter, N. Monitoring recombination during meiosis in budding yeast. Methods Enzymol. 601, 275–307 (2018).
Yoon, S. W. et al. Meiotic prophase roles of Rec8 in crossover recombination and chromosome structure. Nucleic Acids Res. 44, 9296–9314 (2016).
Cannavo, E. et al. Regulation of the MLH1–MLH3 endonuclease in meiosis. Nature 586, 618–622 (2020).
Manhart, C. M. et al. The mismatch repair and meiotic recombination endonuclease Mlh1–Mlh3 is activated by polymer formation and can cleave DNA substrates in trans. PLoS Biol. 15, e2001164 (2017).
Claeys Bouuaert, C. & Keeney, S. Distinct DNA-binding surfaces in the ATPase and linker domains of MutLγ determine its substrate specificities and exert separable functions in meiotic recombination and mismatch repair. PLoS Genet. 13, e1006722 (2017).
De Los Santos, T. et al. The mus81/mms4 endonuclease acts independently of double-Holliday junction resolution to promote a distinct subset of crossovers during meiosis in budding yeast. Genetics 164, 81–94 (2003).
Copsey, A. et al. Smc5/6 coordinates formation and resolution of joint molecules with chromosome morphology to ensure meiotic divisions. PLoS Genet. 9, e1004071 (2013).
Xaver, M., Huang, L., Chen, D. & Klein, F. Smc5/6-mms21 prevents and eliminates inappropriate recombination intermediates in meiosis. PLoS Genet. 9, e1004067 (2013).
Wehrkamp-Richter, S., Hyppa, R. W., Prudden, J., Smith, G. R. & Boddy, M. N. Meiotic DNA joint molecule resolution depends on Nse5–Nse6 of the Smc5–Smc6 holocomplex. Nucleic Acids Res. 40, 9633–9646 (2012).
Peng, X. P. & Zhao, X. The multi-functional Smc5/6 complex in genome protection and disease. Nat. Struct. Mol. Biol. 30, 724–734 (2023).
Borner, G. V., Hochwagen, A. & MacQueen, A. J. Meiosis in budding yeast. Genetics 225, iyad125 (2023).
Sakuno, T. & Hiraoka, Y. Rec8 cohesin: a structural platform for shaping the meiotic chromosomes. Genes 13, 200 (2022).
Sun, X. et al. Transcription dynamically patterns the meiotic chromosome–axis interface. eLife 4, e07424 (2015).
Castellano-Pozo, M. et al. Surveillance of cohesin-supported chromosome structure controls meiotic progression. Nat. Commun. 11, 4345 (2020).
Tung, K. S. & Roeder, G. S. Meiotic chromosome morphology and behavior in zip1 mutants of Saccharomyces cerevisiae. Genetics 149, 817–832 (1998).
Sym, M., Engebrecht, J. A. & Roeder, G. S. ZIP1 is a synaptonemal complex protein required for meiotic chromosome synapsis. Cell 72, 365–378 (1993).
Snowden, T., Acharya, S., Butz, C., Berardini, M. & Fishel, R. hMSH4–hMSH5 recognizes Holliday junctions and forms a meiosis-specific sliding clamp that embraces homologous chromosomes. Mol. Cell 15, 437–451 (2004).
He, W. et al. Regulated proteolysis of MutSγ controls meiotic crossing over. Mol. Cell 78, 168–183 (2020).
Borner, G. V., Kleckner, N. & Hunter, N. Crossover/noncrossover differentiation, synaptonemal complex formation, and regulatory surveillance at the leptotene/zygotene transition of meiosis. Cell 117, 29–45 (2004).
Allers, T. & Lichten, M. Differential timing and control of noncrossover and crossover recombination during meiosis. Cell 106, 47–57 (2001).
Kaur, H., De Muyt, A. & Lichten, M. Top3–Rmi1 DNA single-strand decatenase is integral to the formation and resolution of meiotic recombination intermediates. Mol. Cell 57, 583–594 (2015).
Tang, S., Wu, M. K. Y., Zhang, R. & Hunter, N. Pervasive and essential roles of the Top3–Rmi1 decatenase orchestrate recombination and facilitate chromosome segregation in meiosis. Mol. Cell 57, 607–621 (2015).
Fasching, C. L., Cejka, P., Kowalczykowski, S. C. & Heyer, W. D. Top3–Rmi1 dissolve Rad51-mediated D loops by a topoisomerase-based mechanism. Mol. Cell 57, 595–606 (2015).
Cejka, P., Plank, J. L., Bachrati, C. Z., Hickson, I. D. & Kowalczykowski, S. C. Rmi1 stimulates decatenation of double Holliday junctions during dissolution by Sgs1–Top3. Nat. Struct. Mol. Biol. 17, 1377–1382 (2010).
Bythell-Douglas, R. & Deans, A. J. A structural guide to the Bloom syndrome complex. Structure 29, 99–113 (2021).
Voelkel-Meiman, K., Cheng, S. Y., Morehouse, S. J. & MacQueen, A. J. Synaptonemal complex proteins of budding yeast define reciprocal roles in MutSγ-mediated crossover formation. Genetics 203, 1091–1103 (2016).
Henggeler, A., Orlić, L., Velikov, D. & Matos, J. Holliday junction–ZMM protein feedback enables meiotic crossover assurance. Nature https://doi.org/10.1038/s41586-025-09559-x (2025).
Pyatnitskaya, A., Borde, V. & De Muyt, A. Crossing and zipping: molecular duties of the ZMM proteins in meiosis. Chromosoma 128, 181–198 (2019).
Zhang, L., Espagne, E., de Muyt, A., Zickler, D. & Kleckner, N. E. Interference-mediated synaptonemal complex formation with embedded crossover designation. Proc. Natl Acad. Sci. USA 111, E5059–E5068 (2014).
Shinohara, M., Oh, S. D., Hunter, N. & Shinohara, A. Crossover assurance and crossover interference are distinctly regulated by the ZMM proteins during yeast meiosis. Nat. Genet. 40, 299–309 (2008).
Woglar, A. & Villeneuve, A. M. Dynamic architecture of DNA repair complexes and the synaptonemal complex at sites of meiotic recombination. Cell 173, 1678–1691 (2018).
Voelkel-Meiman, K. et al. Crossover recombination and synapsis are linked by adjacent regions within the N terminus of the Zip1 synaptonemal complex protein. PLoS Genet. 15, e1008201 (2019).
Cahoon, C. K., Helm, J. M. & Libuda, D. E. Synaptonemal complex central region proteins promote localization of pro-crossover factors to recombination events during Caenorhabditis elegans meiosis. Genetics 213, 395–409 (2019).
Voelkel-Meiman, K., Moustafa, S. S., Lefrancois, P., Villeneuve, A. M. & MacQueen, A. J. Full-length synaptonemal complex grows continuously during meiotic prophase in budding yeast. PLoS Genet. 8, e1002993 (2012).
Rog, O., Kohler, S. & Dernburg, A. F. The synaptonemal complex has liquid crystalline properties and spatially regulates meiotic recombination factors. eLife 6, e07424 (2017).
Nadarajan, S. et al. Polo-like kinase-dependent phosphorylation of the synaptonemal complex protein SYP-4 regulates double-strand break formation through a negative feedback loop. eLife 6, e23437 (2017).
Pattabiraman, D., Roelens, B., Woglar, A. & Villeneuve, A. M. Meiotic recombination modulates the structure and dynamics of the synaptonemal complex during C. elegans meiosis. PLoS Genet. 13, e1006670 (2017).
Agarwal, S. & Roeder, G. S. Zip3 provides a link between recombination enzymes and synaptonemal complex proteins. Cell 102, 245–255 (2000).
Qiao, H. et al. Interplay between synaptonemal complex, homologous recombination, and centromeres during mammalian meiosis. PLoS Genet. 8, e1002790 (2012).
Olaya, I., Burgess, S. M. & Rog, O. Formation and resolution of meiotic chromosome entanglements and interlocks. J. Cell Sci. 137, jcs262004 (2024).
Mu, X., Murakami, H., Mohibullah, N. & Keeney, S. Chromosome-autonomous feedback down-regulates meiotic DNA break competence upon synaptonemal complex formation. Genes Dev. 34, 1605–1618 (2020).
Roy, S., Adhikary, H. & D’Amours, D. The SMC5/6 complex: folding chromosomes back into shape when genomes take a break. Nucleic Acids Res. 52, 2112–2129 (2024).
Morawska, M. & Ulrich, H. D. An expanded tool kit for the auxin-inducible degron system in budding yeast. Yeast 30, 341–351 (2013).
Schneider, B. L., Seufert, W., Steiner, B., Yang, Q. H. & Futcher, A. B. Use of polymerase chain reaction epitope tagging for protein tagging in Saccharomyces cerevisiae. Yeast 11, 1265–1274 (1995).
Picard, D. Regulation of protein function through expression of chimaeric proteins. Curr. Opin. Biotechnol. 5, 511–515 (1994).
Benjamin, K. R., Zhang, C., Shokat, K. M. & Herskowitz, I. Control of landmark events in meiosis by the CDK Cdc28 and the meiosis-specific kinase Ime2. Genes Dev. 17, 1524–1539 (2003).
Carlile, T. M. & Amon, A. Meiosis I is established through division-specific translational control of a cyclin. Cell 133, 280–291 (2008).
Oh, S. D. et al. Stabilization and electrophoretic analysis of meiotic recombination intermediates in Saccharomyces cerevisiae. Methods Mol. Biol. 557, 209–234 (2009).
Grubb, J., Brown, M. S. & Bishop, D. K. Surface spreading and immunostaining of yeast chromosomes. J. Vis. Exp. 102, e53081 (2015).
Johnson, E. S. & Blobel, G. Cell cycle-regulated attachment of the ubiquitin-related protein SUMO to the yeast septins. J. Cell Biol. 147, 981–994 (1999).
Hollingsworth, N. M. & Gaglione, R. The meiotic-specific Mek1 kinase in budding yeast regulates interhomolog recombination and coordinates meiotic progression with double-strand break repair. Curr. Genet. 65, 631–641 (2019).
Schmekel, K. & Daneholt, B. Evidence for close contact between recombination nodules and the central element of the synaptonemal complex. Chromosome Res. 6, 155–159 (1998).
Acknowledgements
We thank A. Shinohara and S. Keeney for reagents; J. Matos for communicating unpublished data; and members of the Hunter Laboratory for support and discussions. S.H. was supported by the predoctoral Training Program in Molecular and Cellular Biology at UC Davis supported by NIH T32 training grant GM007377. This work was supported by NIH NIGMS grant GM074223 to N.H. N.H. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
S.T. and N.H. conceived the study and designed the experiments. S.H., R.B., J.E.M., J.K., M.P., E.N., N.L., C.M., H.L. and M.L. assisted S.T. with experiments and data analyses. S.T. and N.H. wrote the manuscript with input and edits from all the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Summary and Model.
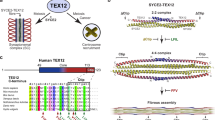
a, Electron micrograph of SC from Blaps cribrosa and schematic highlighting the key features of pachytene-stage chromosomes. Ch, chromatin; LE, lateral element; CE, central element, RN, recombination nodule (the site of a crossover recombination complex, CRC). b, Rec8-cohesin, Zip1, and CRCs constitute local dHJ maturation and protection ensembles that facilitates dHJ formation and protects unnicked dHJs from being dissolved into non-crossovers by the STR/BLM complex. dHJ protection must be maintained until Ndt80-dependent expression of polo-like kinase Plk1Cdc5 triggers crossover-specific resolution via a two-step mechanism: (i) strand-specific nicking of dHJs by the MutLγ endonuclease and associated factors4; (ii) resolution of nicked dHJs by the BLM/STR complex. Placement of the nicks, on both sides of the two Holliday junctions but distal to the exchange points, specifies a crossover outcome. c, Pathways of joint-molecule resolution during meiotic recombination. Resected DSB ends undergo homology search and DNA strand exchange to form nascent displacement loops (D-loops) as common precursors to all pathways. Non-crossover fated D-loops are unwound by BLM/STR to effect synthesis-dependent strand annealing (SDSA), independently of Plk1Cdc5. D-loops that are designated a crossover fate mature into dHJs. In this class-I pathway, Plk1Cdc5 triggers two-step crossover-specific resolution as described in b. All other joint-molecule structures are resolved via a Smc5/6-dependent pathway that is also dependent on Plk1Cdc5 and yields class-II crossovers and non-crossovers. d, Dependencies that help ensure crossing over between homologs. At designated crossover sites, CRC assembly promotes and maintains dHJs and synapsis. In turn, synapsis globally attenuates DSB formation55, thereby diminishing the DNA-damage kinase signaling that inhibits Ndt80 activity65. Ndt80-dependent expression of Plk1Cdc5 then activates crossover-specific dHJ resolution, CRCs disassemble, and desynapsis ensues. The image in a was adapted from ref. 66 (Springer Nature Limited).
Extended Data Fig. 2 Early functions of Rec8 and roles of Smc3, Kleisin Mdc1Rad21, and Separase Esp1 in crossover-specific dHJ resolution.
a, Western analysis of early Rec8-AID degradation following addition of auxin at 2.5 h. b, 1D-gel Southern analysis of DSBs and crossovers following early degradation of Rec8-AID. c, Quantification of the DSB and crossover level from the Southern blot shown in C. d, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation from SMC3-AID subcultures with or without auxin. e, Quantification of final levels of crossovers and non-crossovers at 13 h from SMC3-AID subcultures with or without auxin (mean ± SD, n = 4 independent experiments, ****P < 0.0001, NS, not significant P = 0.0561, two-tailed unpaired t-test). f, Representative 2D gel Southern analysis of joint molecules from SMC3-AID subcultures with or without auxin, before and after release from arrest at 7 and 11 h, respectively. g, Quantification of total joint molecule levels from experiments represented in panel f (mean ± SD, n = 3 independent experiments). NS, not significant, P = 0.3167 for levels at 7hrs; NS, not significant, P > 0.9999 for levels at 11hrs; two-tailed unpaired t-test. h, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation in control, ESP1-AID (with auxin) or MCD1-AID (with auxin) strains. i, Final levels of crossovers and non-crossovers at 11 h from the indicated strains (mean ± SD, n = 3 independent experiments). *P = 0.0146; NS, not significant, P = 0.9610 (control vs. ESP1-AID, crossovers), P = 0.1793 (control vs. MCD1-AID crossovers), P = 0.2073 (control vs. ESP1-AID, non-crossovers), Dunnett’s multiple comparisons test, control.
Extended Data Fig. 3 Sgs1-Top3-Rmi1 and Smc5/6 mediate essentially all joint-molecule resolution.
a, Western blot analysis of Top3-AID and Nse4-AID co-degradation in cells released from pachytene-arrest and addition of auxin at 7 h. b, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation in control, TOP3-AID (with auxin), NSE4-AID (with auxin), TOP3-AID NSE4-AID (with auxin) and TOP3-AID NSE4-AID yen1Δ (with auxin) following release from pachytene arrest. c, Final levels of crossovers and non-crossovers at 11 h from experiments represented in b. (mean ± SD, n = 3 independent experiments). Statistical comparisons with control: ****P < 0.0001, ***P = 0.0002 (NSE4-AID vs. control, crossovers), **P = 0.0251 (TOP3-AID vs. control, non-crossovers), NS not significant P = 0.4157 (TOP3-AID vs. control, crossovers), Tukey’s multiple comparisons test. d, Western blot analysis of RMI1-AID and Nse4-AID co-degradation in cells released from pachytene-arrest and addition of auxin at 7 h. e, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation in control, RMI1-AID (with auxin), NSE4-AID (with auxin) and RMI1-AID NSE4-AID (with auxin) following release from pachytene arrest. f, Final levels of crossovers and non-crossovers at 11 h from experiments represented in b. (mean ± SD, n = 3 independent experiments for control, RMI1-AID, and RMI1-AID NSE4-AID; n = 4 for NSE4-AID). Statistical comparisons with control: ****P < 0.0001, ***P = 0.0003 (NSE4-AID vs. RMI1-AID NSE4-AID, crossovers), **P = 0.0012 (NSE4-AID vs. control, crossovers),**P = 0.0086 (RMI1-AID NSE4-AID vs. control, non-crossovers), *P = 0.0474 (NSE4-AID vs. control, non-crossovers), NS, not significant P = 0.9395 (RMI1-AID vs. control, crossovers), P = 0.7225 (RMI1-AID vs. control, non-crossovers,), Tukey’s multiple comparisons test. g, Representative 2D gel Southern analysis of joint molecules from control, RMI1-AID, NSE4-AID and RMI1-AID NSE4-AID subcultures with auxin after release from arrest at 11 h. h, Quantification of total joint molecule levels from experiments represented in panel g (mean ± SD, n = 3 independent experiments for control, RMI1-AID, and NSE4-AID; n = 4 for RMI1-AID NSE4-AID). NS, not significant P = 0.8596, ***P = 0.0002, ****P < 0.0001, Dunnett’s multiple comparisons test).
Extended Data Fig. 4 Cohesin and the Smc5/6 complex act in parallel pathways of joint-molecule resolution.
a, Western blot images of Smc3-AID Nse4-AID co-degradation following auxin addition at 7 h (corresponds to experiments in Fig. 2e–g). Arp7 is a loading control. b, Western blot images of Rec8-AID Nse4-AID co-degradation following auxin addition at 7 h. c, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation in control, NSE4-AID (with auxin), SMC3-AID (with auxin), and SMC3-AID NSE4-AID (with auxin) strains. d, Final levels of crossovers and non-crossovers at 11 h from the indicated strains (mean ± SD, n = 3 independent experiments for NSE4-AID, SMC3-AID, and SMC3-AID NSE4-AID; n = 4 for control). Statistical comparisons with control unless indicated. For crossovers, Tukey’s multiple comparisons test was performed; for non-crossovers, Dunnett’s multiple comparisons test was performed. ****P < 0.0001; *P = 0.0178 (control vs. NSE4-AID crossovers); *P = 0.0277(control vs. NSE4-AID non-crossovers), *P = 0.0360 (control vs. SMC3-AID non-crossover). NS, not significant, P = 0.7563 (control vs. SMC3-AID NSE4-AID non-crossovers). e, Representative 2D gel Southern analysis of joint molecules from control, NSE4-AID (with auxin), SMC3-AID (with auxin) and SMC3-AID NSE4-AID (with auxin) strains. f, Quantification of total joint-molecule levels from the indicated strains. (mean ± SD, n = 3 independent experiments). Tukey’s multiple comparisons test: *P = 0.0269 (control vs. NSE4-AID), *P = 0.0162(control vs. SMC3-AID NSE4-AID), *P = 0.0207 (SMC3-AID vs. SMC3-AID NSE4-AID), NS, not significant P = 0.9976 (control vs. SMC3-AID).
Extended Data Fig. 5 Targeted degradation does not cause off-target effects.
a, Representative Western blot analysis of Zip1 and Zip3 with or without Rec8-AID degradation following auxin addition at 7 h. b, Quantification of Zip1 and Zip3 level from the experiments shown in a. c, Representative Western blot analysis of Zip3 and Msh4 with or without Zip1-AID degradation following auxin addition at 7 h. d, Quantification of Zip1-AID, Zip3, and Msh4 levels from the experiments shown in c. e, Representative Western blot analysis of Red1, Zip3, and Zip1 with or without Msh4-AID degradation following auxin addition at 7 h. f, Quantification of Msh4-AID, Red1, Zip3, and Zip1 level from the experiments shown in e. Arp7 is the loading control in all experiments.
Extended Data Fig. 6 Cohesin protects dHJs from dissolution by the STR/BLM complex.
a, Western blot analysis of Smc3-AID and Top3-AID co-degradation in pachytene-arrested cells following the addition of auxin at 7 h. Arp7 is a loading control. b, Representative 2D gel Southern analysis of joint molecules from subcultures of pachytene arrested SMC3-AID and SMC3-AID TOP3-AID strains, with or without the addition of auxin. c, Quantification of total joint molecules from experiments shown in b. d, Western blot analysis of Smc3-AID and Top3-AID co-degradation in cells released from pachytene-arrest and addition of auxin at 7 h. Arp7 is used as loading control. e, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation in control, TOP3-AID (with auxin), SMC3-AID (with auxin) and SMC3-AID TOP3-AID (with auxin) following release from pachytene arrest. f, Final levels of crossovers and non-crossovers at 11 h from experiments represented in e (mean ± SD, n = 3 independent experiments for TOP3-AID and SMC3-AID TOP3-AID; n = 4 for control and SMC3-AID). Statistical comparisons with control: ****P < 0.0001, ** P = 0.0038, NS, not significant P = 0.9876 (TOP3-AID vs. control crossovers), *P = 0.0423. NS, not significant P = 0.8549 (TOP3-AID vs. control non-crossovers), NS, not significant P > 0.9999 (TOP3-AID SMC3-AID vs. control non-crossovers), Dunnett’s multiple comparisons test.
Extended Data Fig. 7 Sgs1-AID degradation partially suppresses the crossover defect resulting from degradation of Rec8-AID.
a, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation following release from pachytene arrest in control, SGS1-AID (with auxin), REC8-AID (with auxin), REC8-AID SGS1-AID (with auxin). b, Final levels of crossovers and non-crossovers at 11 h from the indicated strains (mean ± SD, n = 3 independent experiments). Data for TOP3-AID and REC8-AID TOP3-AID from Fig. 5e are shown for comparison. ****P < 0.0001, **P = 0.0037 (REC8-AID SGS1-AID vs. control crossovers), **P = 0.0027(REC8-AID TOP3-AID vs. REC8-AID crossovers), **P = 0.0016 (REC8-AID vs. control non-crossovers), *P = 0.0356 (REC8-AID SGS1-AID vs. REC8-AID crossovers). NS, not significant: P = 0.9712 (TOP3-AID vs. control crossovers), P = 0.2430 (SGS1-AID vs. control crossovers), P = 0.0563 (REC8-AID TOP3-AID vs. control crossovers), P = 0.5846 (TOP3-AID vs. control non-crossovers), P > 0.9999 (SGS1-AID vs. control non-crossovers), P = 0.9949 (REC8-AID TOP3-AID vs. control non-crossovers), P = 0.9998 (REC8-AID SGS1-AID vs. control non-crossovers), Dunnett’s multiple comparisons tests were used to compare auxin treated cells with control. Tukey’s multiple comparisons tests were used to compare individual treatment cells.
Extended Data Fig. 8 Zip1 protects double Holliday junctions from aberrant resolution mediated by the STR/BLM complex.
a, Western blot analysis of Zip1-AID and Top3-AID co-degradation in pachytene-arrested cells following the addition of auxin at 7 h. Arp7 is a loading control. b, Representative 2D gel Southern analysis of joint molecules from subcultures of pachytene arrested ZIP1-AID and ZIP1-AID TOP3-AID strains, with or without the addition of auxin. c, Quantification of total joint molecules from experiments shown in b. d, Western blot analysis of Zip1-AID and Top3-AID co-degradation in cells released from pachytene-arrest and addition of auxin at 7 h. e, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation in control, TOP3-AID (with auxin), ZIP1-AID (with auxin) and ZIP1-AID TOP3-AID (with auxin) following release from pachytene arrest. f, Final levels of crossovers and non-crossovers at 11 h from experiments represented in e. (mean ± SD for 3 independent experiments). Statistical comparisons with control unless stated: ****P < 0.0001, ***P = 0.0005, **P = 0.0071(ZIP1-AID TOP3-AID vs. ZIP1-AID crossovers), **P = 0.0065(ZIP1-AID vs. control non-crossovers), **P = 0.0072 (ZIP1-AID TOP3-AID vs. ZIP1-AID non-crossovers), NS not significant P = 0.1642 (TOP3-AID vs. control crossovers), P = 0.2717 (TOP3-AID vs. control non-crossovers), P = 0.9998 (ZIP1-AID TOP3-AID vs. control non-crossovers), Tukey’s multiple comparisons tests.
Extended Data Fig. 9 MutSγ protects double Holliday junctions from aberrant resolution mediated by the STR/BLM complex.
a, Western blot analysis of Msh4-AID and Top3-AID co-degradation in pachytene-arrested cells following the addition of auxin at 7 h. Arp7 is a loading control. b, Representative 2D gel Southern analysis of joint molecules from subcultures of pachytene arrested MSH4-AID and MSH4-AID TOP3-AID strains, with or without the addition of auxin. c, Quantification of total joint molecules from experiments shown in b. d, Western blot analysis of Msh4-AID and Top3-AID co-degradation in cells released from pachytene-arrest and addition of auxin at 7 h. e, Representative 1D-gel Southern analysis of crossover (upper panel) and non-crossover (lower panel) formation in control, TOP3-AID (with auxin), MSH4-AID (with auxin) and MSH4-AID TOP3-AID (with auxin) following release from pachytene arrest. f, Final levels of crossovers and non-crossovers at 11 hrs from experiments represented in e. (mean ± SD, n = 3 for TOP3-AID and MSH4-AID TOP3-AID; n = 6 for control and MSH4-AID). Statistical comparisons with control unless stated: ****P < 0.0001, ***P = 0.0005 (TOP3-AID MSH4-AID vs. control crossovers), ***P = 0.0001 (TOP3-AID MSH4-AID vs MSH4-AID crossovers), NS not significant P > 0.9999 (TOP3-AID vs. control) crossovers, P = 0.1250 (TOP3-AID vs. control non-crossovers), P = 0.0988 (TOP3-AID MSH4-AID vs. control non-crossovers), Tukey’s multiple comparisons test.
Extended Data Fig. 10 Stabilizing dHJs in pachytene does not bypass interdependencies between cohesin, synapsis, and crossover recombination complexes.
a, Representative images of surface-spread meiotic nuclei from pachytene-arrested REC8-AID TOP3-AID cells sampled 1 hr after the addition of auxin or DMSO vehicle at 7 h, and immunostained for the indicated markers. b, Quantification of Red1 and Zip3 immunostaining structures from the REC8-AID TOP3-AID experiments represented in panel a, and comparison with corresponding data from REC8-AID (from Fig. 3b). Means ± SD for Red1 n = 32, 57, and 30 nuclei; and Zip3 n = 33, 57, and 42 nuclei. c, Representative images of surface-spread meiotic nuclei from pachytene-arrested ZIP1-AID TOP3-AID cells sampled 1 hr after the addition of auxin or DMSO vehicle at 7 h, and immunostained for the indicated markers. d, Quantification of Msh5 and Zip3 immunostaining foci from the experiments represented in panel c, and comparison with corresponding data from ZIP1-AID (from Fig. 4f). Means ± SD for Msh5 n = 44, 47, and 67 nuclei; and Zip3 n = 51, 44, and 41 nuclei. e, Representative images of surface-spread meiotic nuclei from pachytene-arrested MSH4-AID TOP3-AID cells sampled 1 hr after the addition of auxin or DMSO vehicle at 7 h, and immunostained for the indicated markers. f, Quantification of Msh4 and Zip3 immunostaining foci from the experiments represented in panel e, and comparison with corresponding data from MSH4-AID (from Fig. 4h). Means ± SD for Msh4 n = 42, 35, and 57 nuclei; and Zip3 n = 57, 63, and 49 nuclei. Scale bars = 1 μm. Unpaired two-tailed t test, ****P < 0.0001.
Supplementary information
Supplementary Fig. 1 (download XLSX )
Source gel images with cropping indicated by red or white boxes.
Supplementary Tables (download PDF )
Supplementary Table 1. Saccharomyces cerevisiae strains used in this study. Supplementary Table 2. Oligonucleotides used in this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tang, S., Hariri, S., Bohn, R. et al. Protecting double Holliday junctions ensures crossing over during meiosis. Nature 647, 776–785 (2025). https://doi.org/10.1038/s41586-025-09555-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09555-1