Abstract
Unravelling the genetic basis of the remarkable phenotypic diversity observed in natural populations remains a central challenge in biology1,2,3,4. Despite major advances5,6,7,8,9,10,11,12,13,14,15,16,17,18,19, no species has yet been characterized with a truly comprehensive atlas of genetic variation. Here we present an extensive genomic and phenotypic resource for the yeast Saccharomyces cerevisiae based on near telomere-to-telomere assemblies of 1,086 natural isolates. Leveraging these high-contiguity assemblies, we generated a highly complete species-wide structural variant atlas, gene-based pangenome and graph pangenome. By incorporating the full spectrum of genetic variation captured across the species, we conducted genome-wide association studies across 8,391 molecular and organismal traits19,20,21,22. The inclusion of structural variants and small insertion–deletion mutations improved heritability estimates by an average of 14.3% compared with analyses based only on single-nucleotide polymorphisms. Structural variants were more frequently associated with traits and exhibited greater pleiotropy than other variant types. Notably, the genetic architecture of molecular and organismal traits differed markedly. Together, this work provides a unique dataset that illuminates how diverse forms of genetic variation shape phenotypic diversity and lays the groundwork for integrative, genome-scale studies in other eukaryotic systems.
Similar content being viewed by others
Main
A comprehensive understanding of the genetic architecture that underlies phenotypic diversity requires looking beyond single-nucleotide polymorphisms (SNPs) to encompass the full spectrum of genetic variation4,5,6,7,8. Although genome-wide association studies (GWASs) have uncovered thousands of loci linked to complex traits, they have historically concentrated on small variants, mainly SNPs, largely owing to constraints in detecting larger, more complex variants at the species level12,13,23. In this context, structural variants (SVs), including insertions, deletions, duplications and rearrangements, remain underexplored despite their potential to exert substantial phenotypic effects in complex traits. Emerging long-read sequencing strategies and pangenome approaches now enable high-resolution detection of SVs at the population level9,14,15,16,17,18,24,25,26, but assembling complete, telomere-to-telomere genomes for large cohorts remains a challenge.
On the phenotypic front, integrating molecular traits such as transcript, protein, and metabolite levels with organismal phenotypes provides a more detailed view of trait architecture20,27,28,29,30,31,32,33. Multilayered phenotypic data across large, natural populations are still uncommon. The budding yeast S. cerevisiae represents a unique opportunity in this context. With over 1,000 natural isolates spanning diverse ecological and geographic origins19, and rich datasets capturing both organismal and molecular phenotypes19,20,21,22, it is ideally suited for dissecting the contribution of complex variants to trait diversity. However, the lack of a population-scale SV catalogue has thus far limited our ability to fully resolve how different variant types shape phenotypic variation.
Here, we assembled near telomere-to-telomere genomes for 1,086 natural S. cerevisiae isolates using long-read sequencing, enabling a comprehensive catalogue of SVs and gene content diversity at the species level. By integrating this genomic resource with 8,391 molecular and organismal traits, we reveal that SVs are more frequently associated with phenotypic variation and exhibit greater pleiotropy than SNPs and small (less than 50 bp) insertions–deletions mutations (indels), particularly for organismal traits. A graph-based pangenome uncovered 2.5 Mb of non-reference sequence, underscoring the extent of uncharted genomic diversity. This study addresses a critical gap in our understanding of how different types of genetic variation contribute to phenotypic diversity.
High-quality assemblies of 1,482 genomes
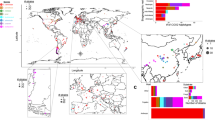
To comprehensively capture species-wide diversity, we sequenced 989 natural S. cerevisiae isolates using Oxford Nanopore technology (ONT)19,34 (Fig. 1a), achieving an average depth of 95× and an N50 of 19.1 kb (Fig. 1b, Supplementary Fig. 1 and Supplementary Table 1). We supplemented this with ONT data from 14 beer isolates35 and 24 Taiwanese isolates36, for a total of 1,027 isolates. A hybrid assembly pipeline was utilized to maximize contiguity and completeness (Supplementary Fig. 2 and Methods), yielding chromosome-scale assemblies for 1,015 isolates. We also included 71 assemblies from the S. cerevisiae reference assembly panel7, resulting in 1,086 isolates overall.
a, Schematics of the pangenome and association analyses. eQTL, expression QTL; pQTL; protein QTL; gCNVs, genomic CNVs. b, Long-read sequencing depth and reads N50 per isolate, for 989 newly sequenced isolates. The middle bar of the box plots corresponds to the median; the upper and lower bounds correspond to the third and first quartiles, respectively. The whiskers correspond to the upper and lower bounds 1.5 times the interquartile range (IQR). c, Haplotype resolution of genome assemblies for 1,086 isolates. d, Assembly contiguity represented as Nx value (length of the shortest contig in the group of the longest contigs that represent x% of the assembly length) for 1,482 assemblies. The white line represents the reference genome assembly, and the blue dashed line corresponds to the mean value for all assemblies.
These isolates vary in ploidy and zygosity, with 75% being diploid, of which 55.2% are heterozygous (Fig. 1c and Supplementary Table 2). Haplotype-resolved assemblies were generated for 396 of the 456 non-polyploid heterozygous isolates, and the remaining 60 were assembled in collapsed form (Fig. 1c and Supplementary Note 1). Altogether, we generated 1,482 high-quality assemblies across the 1,086 isolates (Supplementary Table 2).
Assembly quality was assessed across several metrics. Contiguity matched the reference genome (Fig. 1d), with a median of 1.06 contigs per chromosome and 97.2% of the chromosomes assembled into a single contig (Extended Data Fig. 1a, Supplementary Table 2 and Supplementary Note 2). Assembly sizes ranged from 11.17 Mb to 12.95 Mb (mean = 11.90 ± 0.17 Mb) (Extended Data Fig. 1b). Accuracy, based on Illumina data and Merqury estimates, had an average Merqury quality value of 41.5 (Extended Data Fig. 1c), and completeness averaged 99.1% by BUSCO, closely matching the reference score37 of 99.4% (Extended Data Fig. 1d). These results confirm that, although they do not always encompass the entire telomere-to-telomere sequence (Supplementary Note 2), the assemblies exhibit contiguity and completeness close to those of the reference genome, reaching near telomere-to-telomere status.
Species-wide SV spectrum
This comprehensive set of genome assemblies enabled accurate detection of SVs (SVs larger than 50 bp) in a highly diverse population. Through pairwise alignment of the assemblies with the S288c reference genome (Fig. 1a), we identified a total of 262,629 redundant SVs across 1,086 isolates, corresponding to 6,587 unique events. Systematic validation of 500 SVs based on the mapping of short-read sequencing data confirmed sequence disruption in 95% of the SV calls (Supplementary Note 3). SVs were classified into four categories: presence–absence variations (PAVs, 4,755 events), segmental copy-number variations (CNVs) (1,207), inversions (231) and translocations (394) (Fig. 2a). Together, these SVs span a total of 27.3 Mb of sequences, excluding translocations. Transposable elements (TEs), particularly Ty elements, are major contributors to SVs in S. cerevisiae. Ty elements are found spanning over 50% of the SV sequence in 39% of PAVs (1,834 events), 20% of inversions (46 events) and 9% of CNVs (104 events) (Supplementary Table 3).
a, Number of non-redundant SV events per type and frequency in the population. Frequency categories are rare (MAF < 1%), low-frequency (1% ≥ MAF > 5%) and common (MAF ≥ 5%). INV, inversions; TRA, translocations. b, Rarefaction curves and extrapolation for each type of SV. c, MAF of TE-related and non-TE-related SVs. P value was calculated using a two-sided Mann–Whitney–Wilcoxon test (****P = 5.2 × 10−39). The middle bar of the box plots corresponds to the median; the upper and lower bounds correspond to the third and first quartiles, respectively. The whiskers correspond to the upper and lower bounds 1.5 times the IQR. d, Enrichment of SNPs, indels and SVs in subtelomeric regions. P values were computed using two-sided Fisher’s exact tests with FDR correction (****P = 0 for SNPs-indels, ****P = 3.5 × 10−234 for SNPs-SVs and ****P = 2.9 × 10−92 for indels-SVs,). e, Structural diversity along chromosomes, represented by outer blue rectangles, for each type of SV. Blue points correspond to regions that are outliers in structural diversity. The inner plot represents a map of translocations, coloured according to their MAF. f, Proportion of the SV types in the SV signature of wine, beer, Asian fermentation (AF) and wild isolates. Total represents all the SVs involved in any clade signature. P values were computed using Pearson’s chi-squared test with FDR correction (****P = 7.2 × 10−5). g, Correlation between the number of SNPs and SVs across 970 non-polyploid isolates using a Spearman correlation test (P = 4.4 × 10−211). Larger points correspond to the average value per clade. Coloured points indicate deviation from the correlation using Pearson’s chi-squared test with Bonferroni correction (P = 9.4 × 10−4 (AU wine 2), P = 9.8 × 10−5 (Alpechín), P = 3.2 × 10−4 (Georgian wine), P = 0.024 (Belgium beer 1), P = 0.049 (French dairy), P = 3.8 × 10−16 (Chinese wild)).
The large size of our population enabled precise quantification of SV diversity in S. cerevisiae. A median of 289 SVs was observed between isolates. We calculated a structural diversity of 2.0 × 10−5, defined as the average number of SVs per site between isolate pairs, two orders of magnitude lower than nucleotide diversity in the same population19 (Methods). Extrapolating SV accumulation with increasing sample size, we estimated a total of 7,237 SVs, indicating that our dataset captures more than 90% of all SV events in the species (Fig. 2b). Capture rates varied slightly by SV type, from 92.4% of inversions to 83.7% of translocations (Supplementary Table 4). We also estimated species coverage, the proportion of redundant SVs recovered, which reached 99.5%, suggesting nearly complete representation of shared SVs (Extended Data Fig. 2 and Supplementary Table 4).
In addition, the large sample size also enabled accurate estimation of SV allele frequencies in the population. Similar to SNPs, SVs are skewed towards low frequencies: 69% are rare (minor allele frequency (MAF) < 1%), 20% are low-frequency (1% ≤ MAF < 5%) and only 11% are common (MAF ≥ 5%) (Fig. 2a). Frequency patterns varied by SV type, translocations and inversions were rarer than PAVs and CNVs (Extended Data Fig. 3a). Their site frequency spectra resembled those of nonsense mutations19, suggesting strong deleterious effects (Extended Data Fig. 3b). Additionally, Ty-related SVs were more frequently shared across isolates than non-Ty-related ones (Fig. 2c and Extended Data Fig. 3c).
Finally, we used haplotype-resolved assemblies from 396 heterozygous isolates to assess structural heterozygosity in S. cerevisiae. The proportion of heterozygous SVs per isolate ranged from 11% to 94% (Extended Data Fig. 4a,b) and was strongly correlated with SNP heterozygosity (Spearman R = 0.79, P < 2.2 × 10−16; Extended Data Fig. 4c). SVs in subtelomeric regions showed higher heterozygosity (Extended Data Fig. 4d), consistent with the known structural variability of these regions. Inversions and translocations exhibited higher heterozygosity than PAVs and CNVs (Extended Data Fig. 4e). SV length was correlated with heterozygosity (Spearman R = 0.55, P = 2.6 × 10−8; Extended Data Fig. 4f), with SVs greater than 30 kb in size showing a marked shift towards heterozygosity (78% heterozygous) compared with smaller SVs (less than 30 kb, 45% heterozygous). This size effect helps explain the increased heterozygosity observed for typically larger SV classes such as inversions and translocations (Extended Data Fig. 4g).
Genomic distribution of SVs
We analysed the genomic distribution of SVs and found it to be highly uneven, with significant enrichment in subtelomeric regions (two-sided Fisher exact test, P = 1.1 × 10−309). Although SNPs and indels are also enriched in these regions, the enrichment is much stronger for SVs (Fig. 2d). By computing structural diversity along the genome, we identified bursts of diversity that are often specific to a single SV type (Fig. 2e) and defined 46 SV hotspots (Supplementary Table 5), including 21 translocation hotspots, nearly all in subtelomeric regions. One notable exception is a well-described reciprocal translocation between chromosomes 8 and 16, which confers sulfite resistance through the overexpression of the SSU1 gene located near the breakpoint38 (Fig. 2e). Hotspots for PAVs, CNVs and inversions often mapped to TE-rich regions or SV-prone genes, such as FLO39,40, CUP1 (ref. 41), and genes with tandem repeats, such as HPF1, SPA2 and NUM1 (refs. 42,43). An inversion hotspot on chromosome 14 overlaps a 24-kb region flanked by inverted repeats, probably driven by recombination44.
These findings indicate that SV hotspots arise either from genome fragility, linked to Ty elements or repetitive sequences, or from adaptive pressures targeting specific genes. To distinguish these mechanisms, we examined the distribution of hotspot SVs across species clades with known ecological origins45. Out of the 46 hotspots, 23 were evenly distributed across clades and localized to fragile regions (Supplementary Table 5), whereas the remaining 23 were clade-enriched, reflecting either population bottlenecks or local adaptation—for example, SVs associated with sulfite and copper resistance were enriched in wine isolates (Supplementary Table 5).
SV diversity and population structure
To explore the relationship between population structure and SV diversity, we built separate phylogenies using SNP and SV genotypes (Supplementary Fig. 3). Despite minor differences, the overall tree topology was conserved, with clades clustering consistently, indicating distinct SV signatures per clade. Using allelic enrichment of non-singleton SVs, we identified 1,933 SV alleles that were significantly over-represented in at least one clade, resulting in 3,559 clade–SV associations (Supplementary Table 6). The types of SVs involved varied by clade—for example, translocations were enriched in wine clades (Pearson’s chi-squared test, P = 7.2 × 10−5; Fig. 2f and Extended Data Fig. 5).
We also assessed whether SV and SNP diversity scaled similarly across clades. Although SV and SNP counts per isolate were generally correlated (R = 0.69, P < 2.2 × 10−16; Fig. 2g), deviations were observed. Wild clades, particularly the Chinese wild group, the species’ ancestral population19, deviates from this correlation, with fewer SVs than expected based on the number of SNPs, or more SNPs than expected based on the number of SVs (Fig. 2g). A similar trend in the Alpechín clade probably reflects SNP inflation due to introgression. By contrast, domesticated clades (French dairy, beer and wine) showed an excess of SVs, suggesting that SVs contributed to rapid adaptation during domestication.
Overall, our assemblies enabled a near-complete view of SV diversity in S. cerevisiae, revealing a landscape shaped by both population structure and adaptive processes.
A complete gene-based pangenome
The comprehensive analysis of high-quality genome assemblies enabled a complete reconstruction of the S. cerevisiae gene-based pangenome that delineates the exhaustive catalogue of genes that are present in the species. Across the population, we identified 8,541 gene families (hereafter referred to as genes), including 2,199 absent from the reference genome (Methods). Gene counts per isolate ranged from 6,438 to 6,814 (average of 6,651; Supplementary Fig. 4). The pangenome consists of 5,047 core genes shared by all isolates and 3,494 accessory genes with variable presence. Accessory genes were further classified into soft core (1,263 genes present in >90% of isolates), dispensable (2,102 genes in 0.001–90%) and private (129 genes unique to one isolate) categories (Fig. 3a). The high proportion of core and soft core genes (73.9%) indicates moderate gene content variation, consistent with a closed pangenome typical of many eukaryotes14,18,25,46,47,48. The genes captured by our population represent 99.5% of the species estimate (Fig. 3b and Supplementary Table 7), demonstrating the high completeness of our defined gene-based pangenome.
a, Distribution of the frequency of genes in the population. Colours correspond to different frequency categories (core, soft core, dispensable and private), and pie charts represent the number of genes in each category. b, Rarefaction curves of the number of genes for pan, core and accessory genomes. c, Distribution of gene location along chromosomes. Colours represent frequency categories. A large introgression event found in strain CPN produces a private gene signature between 424 and 590 kb on chromosome 7. d, Inferred origin of genes constituting the different frequency categories. e, Distribution of the gene length per origin. Dashed vertical lines represent the median value for each origin. Letters discriminate groups between which a two-sided Mann–Whitney–Wilcoxon test with FDR correction is significant with P < 0.05. P < 2.6 × 10−8 (A versus B), P < 1.8 × 10−31 (A versus C), P < 3.2 × 10−24 (B versus C). f, Number and origin of genes involved in the gene signature of each clade. Stripes indicate candidate origin, whereas the absence of pattern indicates a confident origin (Methods). g, Presence of MEL genes in the population. The gene tree was built from multiple sequence alignment of all genes of the pangenome associated to an alpha-galactosidase activity, in addition to the S. paradoxus and S. mikatea homologous genes. The inner tree represents the 1,086 isolates of our study and was built using a neighbour-joining strategy on SNP markers.
Core and accessory genes display distinct genomic and functional characteristics. Accessory genes are highly enriched in subtelomeric regions (two-sided Fisher’s exact test, odds ratio = 0.03, P < 2.2 × 10−16; Fig. 3c and Supplementary Table 8), further reflecting the genomic variability of these regions. Using previously generated transcriptomic data20, we found that core genes are more highly expressed than accessory genes (Supplementary Fig. 5), consistent with findings in other species14,18,49,50. Functional enrichment analyses confirmed that core genes are involved in essential biological processes (Supplementary Fig. 6 and Supplementary Table 9).
To investigate the origin of non-reference genes, we aligned novel gene sequences to a curated eukaryotic database (Supplementary Table 8 and Methods). Among the 2,199 novel genes, 1,233 (56.1%) showed highest similarity to close Saccharomyces relatives, suggesting introgression, and 358 (16.3%) were most similar to non-Saccharomyces species, which is indicative of horizontal gene transfers (HGTs). Another 516 genes (23.5%) showed low identity to reference homologues but aligned best to S. cerevisiae, suggesting rapid evolution, and were classified as fast-evolving genes. The remaining 92 genes (4.2%) lacked significant similarity and may represent de novo gene birth51. These four categories represent the bulk of the accessory genome (Fig. 3d), all sharing features such as subtelomeric localization and low expression (Extended Data Fig. 6). Gene length varies across categories, with introgressed genes being similar in size to reference genes, while HGTs, fast-evolving, and de novo genes tend to be shorter52 (Fig. 3e).
Gene content variation is structured by population. Clustering isolates by gene presence/absence reveals strong population stratification (Extended Data Fig. 7). Enrichment analyses showed widespread introgression across clades, with notably high levels in Alpechín, Mexican agave, and French Guiana isolates, confirming past hybridization events19,53,54 (Fig. 3f and Supplementary Table 10). HGTs were also identified in wine isolates, consistent with previous reports19,55, and partially shared with the Mixed Origins 1 clade, suggesting post-acquisition intraspecific gene flow.
While overall functional content remains conserved across isolates, some introgressed genes appear to confer novel traits. For example, we identified seven introgressed MEL genes encoding alpha-galactosidase activity, allowing growth on melibiose (Supplementary Table 8). These genes, present in phylogenetically distant clades and closely related to homologues from Saccharomyces paradoxus and Saccharomyces mikatae, are likely to represent parallel acquisitions (Fig. 3g), contributing to convergent functional adaptations.
SVs drive broad trait associations
The genome assemblies of more than 1,000 isolates enabled a comprehensive catalogue of genetic diversity, adding 44,804 SVs to the 1.4 million SNPs and 56,086 indels (<50 bp) identified previously (Methods). This resource complements previous phenotypic data spanning 241 colony growth traits (used here as organismal trait proxies), and 8,150 molecular traits, including transcriptomic and proteomic measurements19,20,21,22 (Fig. 1a). Including SVs and indels alongside SNPs increased trait heritability estimates by 14.3% on average (0.36 versus 0.41; Supplementary Fig. 7 and Supplementary Table 11), in line with earlier reports8,56. More importantly, this dataset enables GWASs at single-variant resolution, allowing for direct analysis of the phenotypic effects of SNPs, indels and SVs.
Using a linear mixed model57, we identified 7,768 significant associations linking 3,717 traits to 4,564 QTL (Fig. 4a and Supplementary Table 12), with 3,471 SNP-QTL, 230 indel-QTL and 863 SV-QTL. This corresponds to 6.5%, 10.5%, and 19.8% of tested SNPs, indels and SVs, respectively (Fig. 4b), revealing a strong enrichment of SV-QTL (two-sided Fisher’s exact tests with false discovery rate (FDR) correction, P = 6.6 × 10−161 and 4.9 × 10−20 versus SNPs and indels). SV-QTL also show greater pleiotropy, affecting 2.82 traits on average compared with 1.45 for SNP-QTL and 1.34 for indel-QTL (Fig. 4c; two-sided Wilcoxon tests with FDR correction, P < 10−14). Pleiotropic QTL, which are associated with more than one trait, account for 48.6% (419) of SV-QTL, whereas only 20.3% (2,766) and 21.3% (49) of SNP-QTL and indel-QTL (two-sided Fisher’s exact test, P < 10−14).
a, Distribution of 4,564 QTL detected along the genome. The type of the leading variant involved is colour-coded. b, Proportion of each type of variant among QTL (inner circle) and the total set of common variants (outer circle). The bar plot indicates the odds ratio of the QTL in reference to the total set of variants. Letters discriminate groups between which a two-sided Fisher’s exact test with FDR correction is significant with P < 0.05. P = 1.9 × 10−12 (A versus B), P = 6.6 × 10−161 (A versus C), P = 4.9 × 10−20 (B versus C). c, Distribution of the number of traits associated per QTL, coloured by variant type. The dashed vertical line indicates the average number of traits. d, Number of traits associated depending on the position of QTL along the genome. QTL hotspots (associated with 20 traits or more) are highlighted. Colour and orientation correspond to the type of QTL.
SV-QTL are enriched in subtelomeric regions (Fig. 4a; two-sided Fisher’s exact test, P = 0 and 6.2 × 10−50 versus SNP-QTL and indel-QTL), in line with their known genomic location (Fig. 2d). Additionally, SVs contribute disproportionately to QTL hotspots: 15 SVs are each associated with at least 20 traits, compared with just 3 SNPs and no indels (Fig. 4d). One major SV-QTL hotspot involves a recombination-driven fusion of the ALD2 and ALD3 genes, associated with 66 expression and 30 growth traits. This SV arose independently multiple times, producing five alternate coding sequences (Extended Data Fig. 8) and is strongly enriched in Beer and French dairy isolates (two-sided Fisher’s exact tests with FDR correction, P = 7.06 × 10−30 and 7.45 × 10−18), underlining a possible positive selection in these specific environments.
Effect size estimates show that indel-QTL have the largest average effect (6.0 × 10−2), followed by SNP-QTL (3.9 × 10−2) and SV-QTL (3.4 × 10−2) (Supplementary Fig. 8). Indels exhibit significantly stronger effects than SNPs (1.5×, two-sided Mann–Whitney–Wilcoxon test, P = 3.4 × 10−12) and SVs (1.8×, two-sided Mann–Whitney–Wilcoxon test, P = 3.9 × 10−17), despite their lower pleiotropy. For molecular traits, QTL were classified as local or distant relative to the affected gene (Extended Data Fig. 9). We identified 2,131 local and 5,208 distant associations. Local QTL exhibit significantly higher effect sizes than distant QTL (6.2 × 10−2 versus 3.0 × 10−2; two-sided Mann–Whitney–Wilcoxon test, P = 3.4 × 10−12; Extended Data Fig. 10a), a pattern that holds across all variant types (Extended Data Fig. 10b). Notably, indels are strongly enriched for local associations, with 54.5% of indel-QTL classified as local, compared to 28.8% for SNPs and 26.2% for SVs (Extended Data Fig. 10c; two-sided Fisher’s exact tests with FDR correction, P < 10−18). This higher proportion of local QTL among indels likely contributes to their overall greater effect size relative to SNPs and SVs.
Distinct phenotypic effects of SV types
The precise characterization of SVs has enabled further investigation into the phenotypic effects of the different types of SVs. We identified 615 CNV-QTL, 192 deletion-QTL, 54 insertion-QTL and 2 translocation-QTL. Of the three common inversions present in our dataset, none was associated with a phenotypic variation. The limited number of associated translocations prevents any comparison of their phenotypic effect with other types of SVs. Associated SVs constitute 20.9% of the total deletions, 19.2% of CNVs and 13.5% of insertions. This finding indicates an enrichment of QTL in deletions and CNVs in comparison to insertions (two-sided Fisher’s exact test, P = 9.4 × 10−3 and 0.026, respectively). Deletion-QTL exhibit an average effect size of 4.1 × 10−2, which is 1.2-fold that of CNV-QTL (3.3 × 10−2; two-sided Mann–Whitney–Wilcoxon test, P = 3.5 × 10−4) and 2.2-fold that of insertion-QTL (1.9 × 10−2; two-sided Mann–Whitney–Wilcoxon test, P = 3.3 × 10−9) (Supplementary Fig. 9). In addition, associated deletions have a local effect in 25.0% of the cases, which is analogous to the 25.4% of local associations for CNVs but lower than the 47.7% of local associations for insertions (two-sided Fisher’s exact test, P value = 9.4 × 10−4). Overall, these results reveal the limited phenotypic effect of insertions in comparisons to other SVs, as evidenced by a reduced fraction of QTL and a diminished effect size. Insertions appear to be constrained to their local effect and are less frequently acting in trans.
We further aimed to assess the difference in phenotypic effect of SVs related or non-related to TE sequences. Among common SVs, 13.1% of the TE-related SVs were found to be associated with the variation of at least one trait, which is similar to the 13.6% of non-TE-related associated SVs (two-sided Fisher’s exact test, P value = 0.91). Unlike SV-QTL, TE-related SV-QTL are never located within subtelomeric regions, which is expected given the scarcity of TEs in these regions (95.6% of all TE-related SVs are located outside subtelomeric regions). QTL involving TE and non-TE-related SVs exhibit an average effect size of 2.41 × 10−2 and 2.43 × 10−2, respectively, which represents minimal variation (two-sided Mann–Whitney–Wilcoxon test, P value = 0.026). Overall, TE-related SVs exhibit a similar phenotypic effect than other SVs.
Complexity differs across trait types
A key strength of this dataset is the inclusion of both molecular and organismal phenotypes within the same population, allowing direct comparison of their genetic architectures. We identified 4,444 QTL for molecular traits and 168 for organismal traits, averaging 0.9 and 1.7 QTL per trait, respectively (Fig. 5a). This suggests that organismal traits are probably genetically more complex, involving a larger number of contributing loci (two-sided Mann–Whitney–Wilcoxon test, P = 1.3 × 10−8). By contrast, QTL for molecular traits showed significantly higher effect sizes on average (3.9 × 10−2 versus 2.7 × 10−2; two-sided Mann–Whitney-Wilcoxon test, P = 8.1 × 10−12) (Fig. 5b).
a, Distribution of the number of QTL identified per trait. The dashed lines represent the average number of QTL associated per trait. The type of trait (growth or molecular) is colour-coded. b, Effect size of QTL associated with variation of molecular and growth traits. The P value was computed using a two-sided Mann–Whitney–Wilcoxon test (****P = 8.1 × 10−12). The middle bar of the box plots corresponds to the median; the upper and lower bounds correspond to the third and first quartiles, respectively. The whiskers correspond to the upper and lower bounds 1.5 times the IQR. n denotes the number of associations. c, Proportion of the different types of variants found within all variants, molecular and growth QTL. d, Graphical representation of the Minigraph pangenome. The path of the linear reference genome is indicated in orange. Segments are coloured according to their nucleotide sequence identity with the reference genome to highlight non-reference sequences.
The type of associated variants also differs. SV-QTL make up 18.6% and 41.1% of the total QTL for molecular and organismal traits, respectively, both enriched relative to the 7.4% frequency of SVs among common variants (two-sided Fisher’s exact test, P = 2.4 × 10−118 and 2.6 × 10−33). However, the enrichment is stronger for organismal traits (5.6-fold) than for molecular traits (2.5-fold), suggesting that SVs have a more prominent role in shaping complex organismal phenotypes (Fig. 5c and Supplementary Fig. 10).
These findings highlight distinct genetic architectures: organismal traits tend to involve more, weaker-effect variants that are likely to be spread across regulatory layers, whereas molecular traits are influenced by fewer but stronger-effect variants. The pronounced enrichment of SV-QTL in organismal traits supports the idea that large variants may have more persistent phenotypic effects across multiple regulatory layers.
Diversity with a graph pangenome
To facilitate SV genotyping in other S. cerevisiae collections and capture the full spectrum of structural diversity, we constructed a reference graph pangenome using 500 haplotypes, including the linear reference genome. All 6,587 SVs in our catalogue were represented by at least one haplotype. A first graph, built from whole-genome alignments using Minigraph58, included only SVs and reached a length of 48.6 Mb, 4 times the size of the linear reference (Fig. 5d and Supplementary Table 13). This expansion is mainly due to sequence redundancy, with 88% of bases aligning back to the reference (Supplementary Note 4), and the over-representation of Ty elements, which make up 52.2% of the graph length compared to 1.6% of the average genome (Supplementary Table 14). After removing redundancy, the sequence content reached 11.9 Mb, with 2.5 Mb (21%) absent from the linear reference, matching estimates from the gene-based pangenome. A subset of these non-reference regions (267 kb) is likely to correspond to introgressions from other Saccharomyces species.
We then constructed a more detailed graph using Minigraph-Cactus59, integrating both small variants and SVs for comprehensive genotyping. This graph spans 57.7 Mb and encodes variation in 2,081,695 snarls, subgraphs representing alternative alleles, genotyped across 2,874 isolates45. Of these, 4,400 snarls represent SVs larger than 50 bp, and 13.2% are multiallelic. Multiallelic snarls are more prevalent in subtelomeric regions (two-sided Fisher’s exact test, odds ratio = 3.1, P < 2.2 × 10−16) and are enriched in SVs (two-sided Fisher’s exact test, odds ratio = 13.6, P < 2.2 × 10−16). The graph captures 97.5% of low-frequency and 98.8% of common SNPs (Supplementary Table 15), and improves genotyping accuracy compared to a linear reference. Notably, it yields a 10% average increase in heritability estimates across 8,153 traits (Supplementary Note 4).
Together, these results highlight the power of a graph-based approach to capture a broad spectrum of genetic variation in S. cerevisiae, enabling accurate genotyping of both small and SVs at scale. This reference graph pangenome serves as a robust resource for future studies of population genomics, trait mapping and SV-driven adaptation across diverse yeast lineages. However, it is important to note the limitations of current graph-building algorithms in providing a truly exhaustive representation of genomic variants within a population. Particularly, a construction based on the alignments of homologous chromosomes prevents the detection of reciprocal translocations. This underscores the relevance of an assembly-based pangenome approach.
Discussion
The development of high-quality, long-read sequencing and telomere-to-telomere (T2T) assemblies has transformed the analysis of genome variation in eukaryotes. Genomics is undergoing a major paradigm shift, with pangenomes and graph-based models now at the forefront of population-scale analyses14,17,46,60,61,62,63. These frameworks enable high-resolution detection of gene content diversity and SVs, offering transformative insights into genomic variation5,9,10,50,64,65,66.
Constructing truly representative variant catalogues demands not only comprehensive variant detection, but also genome sampling that captures the full breadth of population diversity. Many current eukaryotic datasets remain undersampled and fall short of achieving saturation. This Article presents an extensive exploration of sequence and structural variation across an entire eukaryotic species, leveraging high-quality, near telomere-to-telomere assemblies from 1,086 diverse natural isolates of S. cerevisiae. This exhaustive dataset captures the full spectrum of genetic diversity, ranging from SNPs and indels to complex SVs, and reveals how distinct types of genetic variants contribute to phenotypic variation at both molecular and organismal levels.
Compared with previous preliminary studies in yeast7,67,68, our high-contiguity, population-wide assemblies enabled the construction of a comprehensive species-wide map of SVs, revealing widespread structural heterogeneity across populations. This improved resolution allowed accurate estimation of SV allele frequencies, identification of numerous SV hotspots and detection of lineage-specific SVs, including causal variants linked to adaptive traits. These results refine and extend previous work, highlighting the importance of broad sampling for uncovering the evolutionary and functional effects of SVs in any eukaryotic population, such as humans.
Leveraging this variant catalogue, we also dissected the genetic basis of a large number of traits, 8,391 molecular (transcript and protein abundances) and organismal traits (growth traits)19,20,21,22. SVs were more frequently associated with traits and exhibited greater pleiotropy than SNPs and indels, often underlying QTL hotspots, particularly for complex organismal traits. Our results highlight the distinct phenotypic effects of different variant types and underscore the importance of an exhaustive SV atlas for fully resolving trait architecture.
Our findings reinforce a broader principle in genomics: SVs often harbour causal genetic variation and contribute disproportionately to complex traits, especially at QTL hotspots and in pleiotropic contexts. This observation supports the notion that SVs are a major source of missing heritability8,56. Therefore, genome-wide approaches integrating comprehensive SVs and gene content variation are not only warranted but are also essential to fully resolve genotype–phenotype relationships across species. The framework demonstrated here in S. cerevisiae, combining population-wide telomere-to-telomere assemblies, graph-based genotyping and unified multilayer phenotyping, provides a framework for unpacking trait architecture across eukaryotic genomes.
Methods
Strain culture and DNA extraction
We used a collection of S. cerevisiae isolates that were previously sequenced using short-read sequencing19. For each isolate, we obtained single colonies from frozen stock on solid YPD (1% yeast extract, 2% peptone, and 2% glucose) and cultured one colony per strain in 25 ml of liquid YPD at 30 °C under shaking (120 rpm). After the culture reached saturation (approximately 1.5 days), the nuclear DNA was extracted from the cells using either a previously described protocol69 or a Monarch HMW DNA Extraction Kit (New England Biolabs). The cells from the saturated culture were treated for 2 h with zymolyase (1,000 U ml−1) in 1 M sorbitol to produce spheroplasts. The spheroplasts were then processed with the Monarch HMW DNA Extraction Kit. Samples with a DNA concentration higher than 30 ng µl−1 were retained for DNA sequencing.
Sequencing data
Long reads sequencing data were obtained using Oxford Nanopore sequencing technology. The library was prepared according to the following protocol, using the Oxford Nanopore SQK-LSK109 and SQK-LSK114 kits. Genomic DNA fragments were repaired and 3′-adenylated with the NEBNext FFPE DNA Repair Mix and the NEBNext Ultra II End Repair/dA-Tailing Module (New England Biolabs). Sequencing adapters provided by Oxford Nanopore Technologies (Oxford Nanopore Technologies) were then ligated using the NEBNext Quick Ligation Module (NEB). After purification with AMPure XP beads (Beckmann Coulter), the library was mixed with the sequencing buffer (ONT) and the loading bead (ONT) and loaded on PromethION R9.4.1 and R10.4.1 flowcells. Basecalling was performed with guppy 5.0.16 (https://nanoporetech.com). To confirm the correspondence of novel long-read sequences with previously generated short-read sequences, we compared SNPs inferred from both types of data. Long and short reads were mapped independently on the reference genome using minimap2 v.2.24 (ref. 70) and bwa-mem2 v.2.2.1 (ref. 71), respectively, and SNPs were inferred with longshot v.0.4.5 (ref. 72) and gatk v.4.5.0.0 (ref. 73). The reference genome version R64-3-1 was downloaded as a fasta file from the Saccharomyces genome database74 website (https://www.yeastgenome.org). We computed the pairwise distance between all samples based on short reads and long reads SNPs using plink v.1.9 (ref. 75). Cases with unclear correspondence between short and long reads were discarded.
Reads phasing
Sequencing data from non-polyploid heterozygous samples with coverage higher than 20x were phased to obtain one read set for each haplotype. Long reads were mapped on the reference genome using minimap2 v.2.24 (ref. 70) with the option -ax map-ont. SNPs were called with longshot v.0.4.5 (ref. 72) --no_haps --min_cov 7 --min_alt_count 7 --min_alt_frac 0.2. Regions of loss of heterozygosity, defined as 50-kb windows containing fewer than 10 SNPs, were detected and removed from the phasing process. SNPs were phased using whatshap phase v.1.4 (ref. 76), and each sequencing read was tagged HP1, HP2 or unassigned. Unassigned reads were downsampled at 50% coverage with filtlong v.0.2.1 (https://github.com/rrwick/Filtlong) --min_length 1000 --length_weight 10 --keep_percent 50 --min_mean_q 9 to maintain similar coverage to the phased reads. Read set for each haplotype was finally obtained by combining phased reads and unassigned reads.
Genome assembly
The genome assembly pipeline (Supplementary Fig. 2) was run with raw sequencing data of 1,027 samples with more than 10x sequencing coverage, in addition to phased sequencing data for 433 non-polyploid heterozygous samples. Sequencing data were systematically downsampled to 30x using filtlong --min_length 1000 --length_weight 10 --target_bases 360000000 --min_mean_q 9 and additionally to 40x when raw coverage was higher than 40x with --target_bases 480000000. Raw and downsampled sequencing reads were assembled with 3 genome assemblers: (1) Necat v.0.0.1_update20200803 (ref. 77); (2) Flye v.2.9 (ref. 78); and (3) SMARTdenovo -c 1 (ref. 79) using reads cleaned with Necat. Redundancy within each genome assembly was removed by discarding contigs covered on more than 95% by other contigs of the draft assembly. Nuclear contigs were then selected by sequence similarity with a database of S. cerevisiae nuclear chromosomes built from 142 genome assemblies7, discarding chromosomes containing mitochondrial insertions. For each sample and each phased haplotype, the best genome assembly was selected with seven criteria chosen to favour completeness and contiguity: (1) each genome assembly must cover the reference genome over 95% of its length; (2) cover at least 80% of each reference chromosome (except for chromosome 1 for which the threshold was lowered to 75% because of a more variable size); (3) does not cover more than 50% of the mitochondrial genome; (4) does not contains fused chromosomes, identified as contigs containing multiple centromeres; (5) favours the lowest number of contigs required to cover 95% of the reference genome; (6) favours the lowest number of contigs; and finally (7) favours the largest total length. The second-best genome assembly, obtained with a different assembler, was also kept for further utilization in the SV detection pipeline. Genome assemblies were then polished with both long reads using medaka consensus -m r941_prom_sup_g507 1.8.0 (https://github.com/nanoporetech/medaka) and Illumina short reads using HapoG 1.3.3 (ref. 80). For phased haplotypes, contigs of each haplotype were concatenated to perform the short reads polishing. Finally, scaffolding against the reference genome was performed using ragout --solid-scaffolds 2.3.1 (ref. 81). For cases for which the scaffolding generated fused chromosomes, the non-scaffolded genome assembly was retained. Assembly contigs were named and ordered according to their sequence similarity to reference chromosomes. For strain XTRA_FHL, whose sequencing data was contaminated by a Kluveromyces marxianus isolate, K. marxianus contigs were manually removed.
Quality assessment and genome annotation
Correctness of genome assemblies was evaluated with Merqury82, and completeness was assessed with miniBusco37. Gene prediction and detection of TEs, centromeres and subtelomeric elements were performed through the LRSDAY pipeline v.1.7.0 (ref. 83). Telomeric sequences were identified across all assemblies using Telofinder7.
SV detection
SVs were detected by individually comparing the generated assemblies with the reference genome (SGD R64 genome assembly of strain S288c, GenBank ID: GCA_000146045.2) using MUM&Co v.3.8 (ref. 84) with the -g 12000000 option. SV calling was run on 1,482 genome assemblies from 1,086 isolates (including 396 isolates with phased genome assemblies). The pipeline uses whole-genome alignments obtained via the MUMmer4 software85 to detect insertions, deletions, duplications, contractions, inversions, and reciprocal translocations exhibiting a size larger than 50 bp. To be validated, SVs had to be detected in at least two independent assemblies. To avoid removing singleton SVs—that is, those present in a single haplotype—we considered an additional set of 1,329 ‘second-best assemblies’ for 959 isolates of our collection, obtained from an alternative assembler and that met the completeness quality threshold defined. A total of 2,811 single sample VCF files were obtained and merged into a single multisample VCF file. First, insertions, deletions, duplications, contractions, and inversions were merged using Jasmine v.1.1.5 (ref. 86), which is based on an SV proximity graph that consider SV breakpoint position and length. Given that Jasmine’s algorithm does not consider two breakpoints for a single SV, a custom merging strategy was developed for translocations, as these involve two distinct breakpoints in the genome. This strategy is based on the construction of a translocation graph, linking pairs of translocations with both breakpoints within a 10 kb region. Each connected component of this graph was treated as a single SV and appended to the Jasmine’s output.
To minimize false positives, we retained only SVs detected in at least two genome assemblies, coming from different isolates, haplotypes, or genome assemblers. We then discarded the second-best assemblies from the VCF file. Finally, phased haplotypes from the same isolate were merged into phased heterozygous genotypes, resulting in the final SV VCF file containing 1,086 samples. For further analyses, we classified insertions and deletions as presence–absence variants (PAVs), and duplications and contractions as CNVs, to avoid reference-biased terminology. Therefore, PAVs and indels are only distinguished by their sizes—higher or lower than 50 bp, respectively.
Detection of Ty-related SVs
The sequence of PAVs, CNVs and inversions were aligned to a Ty retrotransposon database using blast v.2.12.0 (ref. 87) with the -dust no -perc_identity 95 options. The database was constructed from the sequences of the 48 Ty elements present in the reference genome in addition to the 4 solo LTRs sequences. SVs with more than 50% of their length covered were defined as Ty-related SVs.
Structural diversity
To quantify structural diversity, we adapted the classical formula for pairwise nucleotide diversity88 to account for SVs. We defined the structural diversity \({\pi }_{{\rm{SV}}}\) as the average number of structural differences per site between two sequences within a population:
where \({x}_{i}\) and \({x}_{j}\) are the frequencies of haplotypes \(i\) and \(j\), \({\pi }_{{ij}}\) is the number of SV differences between the haplotypes i and j and \(n\) is the total number of haplotypes. Each SV, including PAVs, CNVs, inversions and translocations, was treated as a discrete event, regardless of its size. An SV was considered present in a window if it overlapped the window by at least 1 bp for PAVs, CNVs, and inversions, or if a translocation breakpoint fell within the window.
We computed \({\pi }_{{\rm{SV}}}\) for each type of SV individually on 10-kb sliding windows (1 kb step). Outlier regions were defined as regions with \({\pi }_{{\rm{SV}}}\) greater than the third quartile plus 5 times the IQR for PAVs and translocations, plus 10 times the IQR for inversions, and plus 20 times the IQR for CNVs, to account for baseline variability. To detect regions associated with specific clades, we tested for the over-representation of SVs located in the region of interest in each clade using a two-sided fisher’s exact test with FDR correction.
SNP and indels detection
SNPs and indels were detected for the 1,086 isolates based on the alignment of paired-end Illumina reads7,19,36 to the reference genome. The reads were mapped to the reference genome using bwa-mem2 mem v.2.2.1 (ref. 71) with default parameters and samtools sort v.1.15.1 (ref. 89). The HaplotypeCaller command from gatk v.4.2.3.0 (ref. 73) was used with option --emit-ref-confidence GVCF to generate single sample GVCF files. These files were then gathered into a single multisample vcf file using commands GenomicsDBImport and GenotypeGVCFs --include-non-variant-sites, following gatk’s germline short variant discovery workflow (https://gatk.broadinstitute.org/hc/en-us/articles/360035535932-Germline-short-variant-discovery-SNPs-Indels). Low-quality genotypes (DP < 10 and GQ < 20) were set to missing using bcftools v.1.18.1 (ref. 89) with the +set-gt command. Sites with fewer than 99% informed genotypes and sites exhibiting excess of heterozygosity (ExcHet > 0.99) were removed. Finally, SNPs and indels were separated into two vcf files using bcftools, and complex loci spanning both SNPs and indels were discarded.
Neighbour-joining trees
Neighbour-joining trees were constructed independently from SNPs and SV matrices (1,474,884 and 6,587 markers, respectively, for 1,086 isolates) using the R packages ape90 and SNPRelate91.
Site frequency spectrum
Annotations of SNPs and indels were obtained using SnpEff v.5.1 (ref. 92) with the -no-downstream -no-upstream options.
Comparison of the number of SVs and SNPs per isolate
We used a simple linear regression to model the relationship between the number of SNPs and SVs in each isolate. We further used the R package chisq.posthoc.test93 to test for clades and super clades that deviated from the linear relationship between the number of SNPs and SVs. The chisq.posthoc.test function was used with a matrix containing the mean number of SNPs and SVs for each clade, with the method = ‘bonferroni’ option.
Gene-based pangenome
The gene-based pangenome was built on the de novo annotated coding sequence (CDS) of genomes with a Merqury quality value (QV) superior to 40, as a lower QV is associated with an increased number of singleton gene families (present in a single isolate), potentially being false positive CDS (Supplementary Fig. 11a). After the Merqury QV filtering, we considered 762 genomes corresponding to 651 isolates (Supplementary Fig. 12) for the construction of the gene-based pangenome (Supplementary Fig. 13).
First, we transferred the annotation of the reference CDS on the CDS identified de novo in the assemblies with a nucleotide sequence similarity search, using blastn v.2.12.0 (ref. 87) with options -dust no -prec_identity 95 -strand plus. The reference annotations version R64-4-1 were downloaded from the Saccharomyces genome database74 website (https://www.yeastgenome.org) in gff3 format. For each pair of de novo and reference CDS, the annotation was transferred when one of these cases was true: (1) the de novo CDS is covered by the reference CDS on more than 90% of its length; (2) the reference CDS is covered by the de novo one on more than 50% of its length; (3) the reference CDS is covered by the de novo one on more than 30% of its length and both start and end of the reference CDS are covered (that is, the alignment spans the first and last 10 bp of the CDS). De novo CDS covering less than 80% of the reference sequence have been annotated as truncated. Additionally, de novo CDS were annotated as Truncated when their reference homologue was covered on less than 80% of their length and alignment did not cover the start and end of the CDS. However, this additional information is purely informative and was not considered for the pangenome construction. We then filtered out all the genes with a length inferior to 100 bp.
Second, we identified gene families with a graph-based strategy. We ran a nucleotide sequences similarity search on the 6,673 reference CDS (larger than 100 bp) and 77,322 de novo CDS for which no annotation was transferred in the previous step, using blastn with the same options mentioned before. A graph was then built using CDS as nodes and sequence homology as edges, using the python package NetworkX94. Homology was considered when an alignment between two CDS covered both at 50% of their length or either at 90%, or when a reference annotation was transferred. Connected components with a density lower than 0.4 were further split into Louvain’s communities, and each component or community was then considered as a gene family. For each gene family, a representative sequence was chosen as the reference CDS with the highest degree in the family, or the de novo CDS with the highest degree when no reference CDS was present. Finally, CDS with less than 100 bp of unique sequence in the pangenome (that is, fragments of sequences strictly identical between multiple CDS) were iteratively removed. Identical matches within the pangenome were identified using blastn with options -dust no -strand plus -wordsize 100 -penalty -10000 -ungapped.
Third, to estimate gene presence/absence within the 1,086 isolates, we used a sequencing depth-based approach. Illumina reads were mapped to the representative CDS of each gene family using bwa-mem2 mem v.2.2.1 (ref. 71) with options -U 0 -L 0,0 -O 4,4 -T 20, which remove penalty for unpaired reads, reduce penalty for reads clipping and gap opening and lower the minimum score required. These relaxed parameters were chosen to prevent mapping issues due to diversity within gene families. Read depth over each CDS was calculated using samtools depth v.1.16.1 (ref. 89) with option -aa. For each CDS in each isolate, a normalized depth was computed as the ratio of the median depth of the CDS (discarding non-unique fragment identified in the previous step) over the median depth of all sequences. Because a given normalized depth could have different meaning according to the ploidy of the isolate, we adjusted the normalized depth with the ploidy and a correcting factor x:
To identify the optimal value of x, we built a gold standard gene presence/absence matrix considering 553 isolates and 5,792 genes for which both de novo annotated genome assemblies and transcriptomic data20 were available. A gene was considered present in an isolate when it was: (1) annotated in the genome assembly; and (2) expressed at ≥2 transcripts per million (TPM) in the corresponding RNA-seq data. A gene was considered absent when it was: (1) not annotated; and (2) had expression <2 TPM. All other cases were excluded to avoid ambiguity. This gold standard was solely used to evaluate the accuracy of gene presence/absence calls from sequencing depth, by generating a precision-recall curve for various values of x (Supplementary Fig. 11b). The optimal performance (highest area under the precision-recall curve; Supplementary Fig. 11c) was achieved with x = 0.15. We further computed the precision and recall values using different thresholds (Supplementary Fig. 11d) and chose a threshold of 0.3 as the recall declined sharply beyond this point. Using this threshold, we estimated the presence of all the gene families of the pangenome across the 1,086 isolates.
Importantly, expression data was used solely to calibrate the sequencing depth-based model, and was not used to call gene presence or absence in any genome. Final gene presence/absence calls were made entirely from DNA sequence data using mapped short-read depth.
Pangenome annotation
We sought to annotate the origin and function of novel genes with protein sequences similarity search against a curated database. We built a blast database by coupling the RefSeq protein database with a custom database containing Fungi protein sequences from Shen et al.95 and S. paradoxus protein sequences from Yue et al.96. The sequence similarity search was run using blastp87 with default parameters, and the obtained results were further filtered with a minimum protein identity of 30% and a minimum query coverage of 50%. We categorized the origin of each novel gene as: (1) fast-evolving gene when the best hit was a S. cerevisiae protein; (2) introgression when the best hit was a Saccharomyces protein other than S. cerevisiae; (3) HGT when the protein came out of the Saccharomyces genus; and (4) unknown when no sequence similarity was found. We also transferred the gene ontology (GO) terms associated with the best protein hit in RefSeq to each novel gene (using the same identity and coverage filters as above) and inferred the GO terms of the whole pangenome based on sequence using InterProScan v.4.65-97.0 (ref. 97).
Transcriptomics
Reads were mapped on the CDS of the pangenome using STAR v.2.7.9.a98 with default parameters. Number of reads mapped to each gene family was retrieved using samtools idxstats v.1.16.1 (ref. 89) and TPM were computed with a custom python script.
Rarefaction curves
For both SVs and gene families, rarefactions curves were obtained using the R package iNEXT v.3.0.0 (ref. 99). The iNEXT function was used with a presence–absence matrix of SVs or genes and the options datatype = ‘incidence_raw’ and k = 400. We interpreted the species richness as the total number of SV or gene families in the species and the sample coverage estimate as the species coverage. For the core genome, we first used a matrix of missing genes (filled with 1 when the gene was absent and 0 when the gene was present) as input of the iNEXT function, with the same parameters as before. We then subtracted the rarefaction obtained from the missing genes to the species estimate of the pangenome (8,583 genes), to obtain the rarefaction of the core genome. Finally, we obtained the rarefaction of the accessory genome by subtracting the core genome rarefaction to the pangenome rarefaction.
Clade-specific variants
Clade-specific SVs and genes were obtained using a simple-over-representation analysis based on hypergeometric tests. We used the fora function from the fgsea R package v.1.27.0 (ref. 100) using the list of clades and super clades as ‘pathways’, the isolates having the SV or gene as ‘gene’ and the total list of isolates as ‘universe’. The function was run for each SV or gene present in at least two isolates, except SV or genes present in all isolates. Results were further concatenated, and P value were adjusted using the FDR method.
GO analysis
We used the GO annotations of the reference genes available from SGD (https://current.geneontology.org/annotations/sgd.gaf.gz), in addition to the transferred GO terms for the novel genes (see ‘Pangenome annotation’) and the GO terms inferred with InterProScan97 for the entire pangenome. We discarded terms with a size larger than 500, as well as terms with a size smaller than 2 (except for terms with a reliable evidence code—that is, inferred from mutant phenotype (IMP), inferred from direct assay (IDA) and inferred from genetic interaction (IGI)). For some analyses, we performed a GO term semantic similarity reduction using the calculateSimMatrix and reduceSimMatrix(threshold = 0.7) functions of the rrvgo R package v.1.10.0 (ref. 101). GO term enrichments were performed using the fora function from the fgsea v.1.27.0 R package100. Clade-specific GO terms were detected similarly to clade-specific genes.
Construction of an exhaustive genotype matrix
To build the most comprehensive genotype matrix possible, we combined SNPs and indels with SVs called from both genome assembly comparisons and gene-based pangenome construction. We transformed the normalized depth computed for each CDS–isolate pair (see ‘Gene-based pangenome’) into biallelic variants by setting multiple depth thresholds (starting from 0.25 and increasing by steps of 0.5, as we would expect for a diploid isolate). We discriminated isolates having a normalized depth below or above each threshold for each CDS. In that way, we capture both the presence–absence of each CDS in the population, in addition to the variation in copy number. The complete loss of a gene was considered as deletion.
Although combining multiple strategies for SV calling ensures the comprehensiveness of our variant catalogue, it results in redundancy from the segmental SV detections (that is, assembly comparisons) and the gene-based CNV and deletion detection. For example, a deletion spanning three consecutive genes would result in one segmental deletion and the deletion of each individual gene. The four SV records would exhibit high linkage disequilibrium as they all correspond to a single event. Additionally, aneuploidies are captured by CNVs of all genes present on the aneuploid chromosome. Although it corresponds to a single event, it results in many gene–CNV records, that exhibit high linkage disequilibrium with each other. This redundancy can be easily removed with linkage pruning to prevent duplicated genetic associations in further analyses.
Heritability estimates
Heritability estimates were computed for complex traits, defined as traits having no association with a P value lower than 1 × 10−20 by GWAS, as high effect predictors may bias the estimations. LDAK v.4.2 (ref. 102) was used for the computation. Phenotypes were normalized using a rank-based inverse normal transformation. Plink matrices of SNPs, indels and SVs were first used to generate independent kinship matrices using the LDAK thin model. Weights for each variant were first computed using LDAK with arguments --thin --window-prune .98 --window-kb 20, and kinship were generated using arguments --calc-kins-direct --weights --power -.25. Trait heritability was then estimated using all three kinship matrices together (--mgrm option), using ploidy as covariate and the option --constrain YES to ensure positive values of heritability.
GWAS
We ran GWAS using a linear mixed model implemented in FaST-LMM v.0.4.6 (ref. 57). Phenotypes were normalized in the same way as for heritability estimates. SNPs, indels and SVs were filtered for MAF at 5%. This MAF filtering retained 89,906 SNPs (6.4% of all SNPs), 2,415 indels (4.3%) and 7,708 SVs (10.7%). These allele frequency differences were not accounted for. Genotypes were pruned for linkage disequilibrium using plink v.1.9 (ref. 75) with option --indep-pairwise 50 kb 1 0.8, yielding 54,544 SNPs, 2,203 indels and 4,540 SVs. Variants were further combined in a single plink matrix, which was used as both kinship and test set for GWAS. To preclude the effect of ploidy and aneuploidies on the genetic associations, both were added as covariates. Ploidy was encoded as a numerical covariate and aneuploidies were encoded for each chromosome as −1, 0 or 1, representing loss, expected copy number or gain, respectively. To correct for the large number of variants tested, a trait-specific P value threshold was defined using a permutation test with 100 permutations and alpha = 0.05. In brief, for each trait, associations were run 100 times on permutated phenotypes retaining the lowest P value for each run. The P value threshold corresponds to the 5% quantile (that is, the fifth-lowest P value across permutations), corresponding to an FDR correction of 5%.
Local variants were defined as located in a 25-kb region around the gene of interest or linked to a pruned variant located in this region. For translocations, both breakpoints were considered for the definition of local variants. To further account for linkage disequilibrium between associated variants, groups of linkage were identified, defined as connected components of variants associated with a same trait and in linkage disequilibrium (based on a 0.5 r2 threshold and a maximal physical distance of 50 kb). For each group, only the variant with the lowest P value was retained, leading to the final number of 4,564 QTL.
Graph construction and novel sequence detection
We build a graph pangenome using the Minigraph-Cactus pipeline v.2.6.4 (ref. 59) with 500 haplotypes, including the reference genome and 499 genomes selected to represent a maximum number of SVs. We used the first graph generated by Minigraph58, that uniquely contains SVs, in order to identify repetitive reference segments in the graph and novel sequences. Only segments larger than 100 bp were considered for these analyses. The segments were mapped to the reference genome using minimap2 -ax asm5 v.2.24 (ref. 70) and the coverage depth along the genome was retrieved using samtools depth v.1.16.1 (ref. 89). To determine the fraction of the graph corresponding to TEs, the segments were aligned on a Ty sequence database constructed from the previous analyses of 1,011 genomes103, using blastn v.2.12.0 (ref. 87), with option -perc_identity 70. The same alignment was performed on each of the genome assemblies used for the graph pangenome construction. Additionally, we sought for sequence redundancy in the graph using blastn -perc_identity 95 and applied a 50% coverage threshold. We built a sequence similarity graph and selected a single representative segment for each connected component. Components containing no reference segments were considered as novel sequences. Novel segments were considered as introgression when they show sequence similarity higher than 95% on a database composed of Saccharomyces genomes (GCF_000292725.1, GCF_001298625.1, GCF_002079055.1, GCF_947241705.1, GCF_947243775.1, GCA_002079085.1, GCA_002079115.1, GCA_002079145.1, GCA_002079175.1) using blastn -perc_identity 95.
Variant calling using the graph
The pangenome graph (gfa format) was converted to gbz using the vg toolkit v.1.54.0 (ref. 104) with the vg autoindex command, and snarls were detected using vg snarls. Illumina reads of 3,039 isolates with publicly available sequencing data were mapped on the graph using vg giraffe105 with options --fragment-mean 350 --fragment-stdev 100 -b fast. Gam files were converted using vg pack with option -Q 5 to remove reads with low mapping quality. We performed variant calling using vg call106 with options --genotype-snarls --all-snarls --snarls --ref-sample S288c to obtain single sample vcf files. Calling from the graph pangenome worked for 2,874 out of 3,039 isolates, the remaining ones being discarded because of an aberrantly long computing time. Single sample vcf files were merged into a single multisample vcf using bcftools merge v.1.16.1 (ref. 89). Variants supported by less than two reads were set to missing using bcftools +setGT -- -t q -n. -e ‘FMT/DP > = 2’. The resulting vcf was further trimmed for non-present alternate alleles with bcftools view --trim-alt-alleles and variants were atomized into multiple ones with bcftools norm --atomize --atom-overlap “.” --multiallelics +any. The difference of length between alternative and reference alleles were used to classify alleles as SNPs, indels of SVs. SNPs show no length difference, indels have length differences inferior to 50 bp and SVs show length differences larger or equal to 50 bp.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequencing reads associated with this work are available at the European Nucleotide Archive under the accessions PRJEB77686 and PRJEB81147. Genomes and annotations, gene-based pangenome, graph-based pangenomes, SV matrix and phenotypes generated are available on Zenodo (https://doi.org/10.5281/zenodo.15698884 (ref. 107)).
Code availability
Scripts used for this work are available on Zenodo (https://doi.org/10.5281/zenodo.15698884 (ref. 107)) and on GitHub (https://github.com/HaploTeam/1086YeastGenomes).
References
Mackay, T. F. C., Stone, E. A. & Ayroles, J. F. The genetics of quantitative traits: challenges and prospects. Nat. Rev. Genet. 10, 565–577 (2009).
Manolio, T. A. et al. Finding the missing heritability of complex diseases. Nature 461, 747–753 (2009).
Fournier, T. & Schacherer, J. Genetic backgrounds and hidden trait complexity in natural populations. Curr. Opin. Genet. Dev. 47, 48–53 (2017).
Harris, L. et al. Genome-wide association testing beyond SNPs. Nat. Rev. Genet. 26, 156–170 (2025).
Liao, W.-W. et al. A draft human pangenome reference. Nature 617, 312–324 (2023).
Lian, Q. et al. A pan-genome of 69 Arabidopsis thaliana accessions reveals a conserved genome structure throughout the global species range. Nat. Genet. 56, 982–991 (2024).
O’Donnell, S. et al. Telomere-to-telomere assemblies of 142 strains characterize the genome structural landscape in Saccharomyces cerevisiae. Nat. Genet. 55, 1390–1399 (2023).
Zhou, Y. et al. Graph pangenome captures missing heritability and empowers tomato breeding. Nature 606, 527–534 (2022).
Gao, Y. et al. A pangenome reference of 36 Chinese populations. Nature 619, 112–121 (2023).
Schloissnig, S. et al. Structural variation in 1,019 diverse humans based on long-read sequencing. Nature 644, 442–452 (2025).
Igolkina, A. A. et al. A comparison of 27 Arabidopsis thaliana genomes and the path toward an unbiased characterization of genetic polymorphism. Nat. Genet. 57, 2289–2301 (2025).
Alonso-Blanco, C. et al. 1,135 genomes reveal the global pattern of polymorphism in Arabidopsis thaliana. Cell 166, 481–491 (2016).
Backman, J. D. et al. Exome sequencing and analysis of 454,787 UK Biobank participants. Nature 599, 628–634 (2021).
Chen, J. et al. Pangenome analysis reveals genomic variations associated with domestication traits in broomcorn millet. Nat. Genet. 55, 2243–2254 (2023).
De Coster, W., Weissensteiner, M. H. & Sedlazeck, F. J. Towards population-scale long-read sequencing. Nat. Rev. Genet. 22, 572–587 (2021).
Eizenga, J. M. et al. Pangenome graphs. Annu. Rev. Genomics Hum. Genet. 21, 139–162 (2020).
He, Q. et al. A graph-based genome and pan-genome variation of the model plant Setaria. Nat. Genet. 55, 1232–1242 (2023).
Kang, M. et al. The pan-genome and local adaptation of Arabidopsis thaliana. Nat. Commun. 14, 6259 (2023).
Peter, J. et al. Genome evolution across 1,011 Saccharomyces cerevisiae isolates. Nature 556, 339–344 (2018).
Caudal, É. et al. Pan-transcriptome reveals a large accessory genome contribution to gene expression variation in yeast. Nat. Genet. 56, 1278–1287 (2024).
Muenzner, J. et al. Natural proteome diversity links aneuploidy tolerance to protein turnover. Nature 630, 149–157 (2024).
Teyssonnière, E. M. et al. Species-wide quantitative transcriptomes and proteomes reveal distinct genetic control of gene expression variation in yeast. Proc. Natl Acad. Sci. USA 121, e2319211121 (2024).
Yengo, L. et al. A saturated map of common genetic variants associated with human height. Nature 610, 704–712 (2022).
Li, H. & Durbin, R. Genome assembly in the telomere-to-telomere era. Nat. Rev. Genet. 25, 658–670 (2024).
Tong, X. et al. High-resolution silkworm pan-genome provides genetic insights into artificial selection and ecological adaptation. Nat. Commun. 13, 5619 (2022).
Wu, Z. et al. Human pangenome analysis of sequences missing from the reference genome reveals their widespread evolutionary, phenotypic, and functional roles. Nucleic Acids Res. 52, 2212–2230 (2024).
Aguet, F. et al. Molecular quantitative trait loci. Nat. Rev. Methods Primers 3, 4 (2023).
Akiyama, M. Multi-omics study for interpretation of genome-wide association study. J. Hum. Genet. 66, 3–10 (2021).
Ferkingstad, E. et al. Large-scale integration of the plasma proteome with genetics and disease. Nat. Genet. 53, 1712–1721 (2021).
Gallois, A. et al. A comprehensive study of metabolite genetics reveals strong pleiotropy and heterogeneity across time and context. Nat. Commun. 10, 4788 (2019).
Orozco, L. D. et al. Integration of eQTL and a single-cell atlas in the human eye identifies causal genes for age-related macular degeneration. Cell Rep. 30, 1246–1259.e6 (2020).
Suhre, K. et al. Connecting genetic risk to disease end points through the human blood plasma proteome. Nat. Commun. 8, 14357 (2017).
The GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Legras, J.-L. et al. Adaptation of S. cerevisiae to fermented food environments reveals remarkable genome plasticity and the footprints of domestication. Mol. Biol. Evol. 35, 1712–1727 (2018).
Saada, O. A. et al. Phased polyploid genomes provide deeper insight into the multiple origins of domesticated Saccharomyces cerevisiae beer yeasts. Curr. Biol. 32, 1350–1361.e3 (2022).
Lee, T. J. et al. Extensive sampling of Saccharomyces cerevisiae in Taiwan reveals ecology and evolution of predomesticated lineages. Genome Res. 32, 864–877 (2022).
Huang, N. & Li, H. miniBUSCO: a faster and more accurate reimplementation of BUSCO. Preprint at bioRxiv https://doi.org/10.1101/2023.06.03.543588 (2023).
Pérez-Ortín, J. E., Querol, A., Puig, S. & Barrio, E. Molecular characterization of a chromosomal rearrangement involved in the adaptive evolution of yeast strains. Genome Res. 12, 1533–1539 (2002).
Smukalla, S. et al. FLO1 is a variable green beard gene that drives biofilm-like cooperation in budding yeast. Cell 135, 726–737 (2008).
Tofalo, R. et al. Genetic diversity of FLO1 and FLO5 genes in wine flocculent Saccharomyces cerevisiae strains. Int. J. Food Microbiol. 191, 45–52 (2014).
Fogel, S. & Welch, J. W. Tandem gene amplification mediates copper resistance in yeast. Proc. Natl Acad. Sci. USA 79, 5342–5346 (1982).
Barré, B. P. et al. Intragenic repeat expansion in the cell wall protein gene HPF1 controls yeast chronological aging. Genome Res. 30, 697–710 (2020).
Verstrepen, K. J., Jansen, A., Lewitter, F. & Fink, G. R. Intragenic tandem repeats generate functional variability. Nat. Genet. 37, 986–990 (2005).
Salzberg, L. I. et al. A widespread inversion polymorphism conserved among Saccharomyces species is caused by recurrent homogenization of a sporulation gene family. PLoS Genet. 18, e1010525 (2022).
Loegler, V., Friedrich, A. & Schacherer, J. Overview of the Saccharomyces cerevisiae population structure through the lens of 3,034 genomes. G3 14, jkae245 (2024).
Lan, D. et al. Pangenome and multi-tissue gene atlas provide new insights into the domestication and highland adaptation of yaks. J. Anim. Sci. Biotechnol. 15, 64 (2024).
Zhang, F. et al. Long-read sequencing of 111 rice genomes reveals significantly larger pan-genomes. Genome Res. 32, 853–863 (2022).
Li, H. et al. Graph-based pan-genome reveals structural and sequence variations related to agronomic traits and domestication in cucumber. Nat. Commun. 13, 682 (2022).
Chen, S. et al. Gene mining and genomics-assisted breeding empowered by the pangenome of tea plant Camellia sinensis. Nat. Plants 9, 1986–1999 (2023).
Hufford, M. B. et al. De novo assembly, annotation, and comparative analysis of 26 diverse maize genomes. Science 373, 655–662 (2021).
Tautz, D. & Domazet-Lošo, T. The evolutionary origin of orphan genes. Nat. Rev. Genet. 12, 692–702 (2011).
Blevins, W. R. et al. Uncovering de novo gene birth in yeast using deep transcriptomics. Nat. Commun. 12, 604 (2021).
D’Angiolo, M. et al. A yeast living ancestor reveals the origin of genomic introgressions. Nature 587, 420–425 (2020).
Tellini, N. et al. Ancient and recent origins of shared polymorphisms in yeast. Nat. Ecol. Evol. 8, 761–776 (2024).
Novo, M. et al. Eukaryote-to-eukaryote gene transfer events revealed by the genome sequence of the wine yeast Saccharomyces cerevisiae EC1118. Proc. Natl Acad. Sci. USA 106, 16333–16338 (2009).
Gui, S. et al. A pan-Zea genome map for enhancing maize improvement. Genome Biol. 23, 178 (2022).
Lippert, C. et al. FaST linear mixed models for genome-wide association studies. Nat. Methods 8, 833–835 (2011).
Li, H., Feng, X. & Chu, C. The design and construction of reference pangenome graphs with minigraph. Genome Biol. 21, 265 (2020).
Hickey, G. et al. Pangenome graph construction from genome alignments with Minigraph-Cactus. Nat. Biotechnol. 42, 663–673 (2024).
Guo, D. et al. A pangenome reference of wild and cultivated rice. Nature 642, 662–671 (2025).
Lynch, R. C. et al. Domesticated cannabinoid synthases amid a wild mosaic cannabis pangenome. Nature 643, 1001–1010 (2025).
Sun, H. et al. The phased pan-genome of tetraploid European potato. Nature 642, 389–397 (2025).
Cheng, L. et al. Leveraging a phased pangenome for haplotype design of hybrid potato. Nature 640, 408–417 (2025).
Jayakodi, M. et al. Structural variation in the pangenome of wild and domesticated barley. Nature 636, 654–662 (2024).
Tang, D. et al. Genome evolution and diversity of wild and cultivated potatoes. Nature 606, 535–541 (2022).
Ebert, P. et al. Haplotype-resolved diverse human genomes and integrated analysis of structural variation. Science 372, eabf7117 (2021).
Istace, B. et al. De novo assembly and population genomic survey of natural yeast isolates with the Oxford Nanopore MinION sequencer. GigaScience 6, giw018 (2017).
Weller, C. A. et al. Highly complete long-read genomes reveal pangenomic variation underlying yeast phenotypic diversity. Genome Res. 33, 729–740 (2023).
Tsouris, A., Brach, G., Friedrich, A., Hou, J. & Schacherer, J. Diallel panel reveals a significant impact of low-frequency genetic variants on gene expression variation in yeast. Mol. Syst. Biol. 20, 362–373 (2024).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Vasimuddin, Md., Misra, S., Li, H. & Aluru, S. Efficient architecture-aware acceleration of BWA-MEM for multicore systems. In 2019 IEEE International Parallel and Distributed Processing Symposium 314–324 (2019).
Edge, P. & Bansal, V. Longshot enables accurate variant calling in diploid genomes from single-molecule long read sequencing. Nat. Commun. 10, 4660 (2019).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. Preprint at bioRxiv https://doi.org/10.1101/201178 (2018).
Engel, S. R. et al. Saccharomyces Genome Database: advances in genome annotation, expanded biochemical pathways, and other key enhancements. Genetics 229, iyae185 (2025).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J. Hum. Genet. 81, 559–575 (2007).
Martin, M. et al. WhatsHap: fast and accurate read-based phasing. Preprint at bioRxiv https://doi.org/10.1101/085050 (2016).
Chen, Y. et al. Efficient assembly of nanopore reads via highly accurate and intact error correction. Nat. Commun. 12, 60 (2021).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Liu, H. et al. SMARTdenovo: a de novo assembler using long noisy reads. Gigabyte 2021, 15 (2021).
Aury, J.-M. & Istace, B. Hapo-G, haplotype-aware polishing of genome assemblies with accurate reads. NAR Genomics Bioinformatics 3, lqab034 (2021).
Kolmogorov, M., Raney, B., Paten, B. & Pham, S. Ragout—a reference-assisted assembly tool for bacterial genomes. Bioinformatics 30, i302–i309 (2014).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245 (2020).
Yue, J.-X. & Liti, G. Long-read sequencing data analysis for yeasts. Nat. Protoc. 13, 1213–1231 (2018).
O’Donnell, S. & Fischer, G. MUM&Co: accurate detection of all SV types through whole-genome alignment. Bioinformatics 36, 3242–3243 (2020).
Marçais, G. et al. MUMmer4: a fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Kirsche, M. et al. Jasmine and Iris: population-scale structural variant comparison and analysis. Nat. Methods 20, 408–417 (2023).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Nei, M. & Li, W. H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl Acad. Sci. USA 76, 5269–5273 (1979).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly 6, 80–92 (2012).
Beasley, T. M. & Schumacker, R. E. Multiple regression approach to analyzing contingency tables: post hoc and planned comparison procedures. J. Exp. Educ. 64, 79–93 (1995).
Hagberg, A., Swart, P. J. & Schult, D. A. Exploring Network Structure, Dynamics, and Function Using NetworkX. Office of Scientific and Technical Information https://www.osti.gov/biblio/960616 (U.S. Department of Energy, 2008).
Shen, X.-X. et al. Tempo and mode of genome evolution in the budding yeast subphylum. Cell 175, 1533–1545.e20 (2018).
Yue, J.-X. et al. Contrasting evolutionary genome dynamics between domesticated and wild yeasts. Nat. Genet. 49, 913–924 (2017).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Hsieh, T. C., Ma, K. H. & Chao, A. iNEXT: an R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol. Evol. 7, 1451–1456 (2016).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Sayols, S. rrvgo: a Bioconductor package for interpreting lists of Gene Ontology terms. microPublication Biol. https://doi.org/10.17912/micropub.biology.000811 (2023).
Speed, D., Hemani, G., Johnson, M. R. & Balding, D. J. Improved heritability estimation from genome-wide SNPs. Am. J. Hum. Genet. 91, 1011–1021 (2012).
Bleykasten-Grosshans, C., Fabrizio, R., Friedrich, A. & Schacherer, J. Species-wide transposable element repertoires retrace the evolutionary history of the Saccharomyces cerevisiae host. Mol. Biol. Evol. 38, 4334–4345 (2021).
Garrison, E. et al. Variation graph toolkit improves read mapping by representing genetic variation in the reference. Nat. Biotechnol. 36, 875–879 (2018).
Sirén, J. et al. Pangenomics enables genotyping of known structural variants in 5202 diverse genomes. Science 374, abg8871 (2021).
Hickey, G. et al. Genotyping structural variants in pangenome graphs using the vg toolkit. Genome Biol. 21, 35 (2020).
Loegler, V. et al. Code and data for the analysis of 1,086 near telomere-to-telomere yeast genomes. Zenodo https://doi.org/10.5281/zenodo.15698884 (2025).
Acknowledgements
We thank J. Bloom and M. Ralser for insightful discussions and comments on the paper. This work was supported by a National Institutes of Health grant R01 (GM147040-01) and a European Research Council Advanced grant (UnMiss) to J.S. In addition, this work was supported by a French National Research Agency grant (ANR-24-CE12-0998-01, PolyPhase). This work is also part of Interdisciplinary Thematic Institutes (ITI) Integrative Molecular and Cellular Biology (IMCBio), as part of the ITI 2021-to-2028 program of the University of Strasbourg, CNRS, and Inserm, supported by IdEx Unistra (ANR-10-IDEX-0002). J.S. is a Fellow of the University of Strasbourg Institute for Advanced Study and a member of the Institut Universitaire de France.
Author information
Authors and Affiliations
Contributions
J.S. conceived and supervised the project. P.T., E.T., A.T. and G.B. performed the experiments. C.C., E.P., S.E. and G.B. sequenced the genomes and produced the data. V.L. assembled, annotated the genomes, and constructed gene-based and graph pangenomes. V.L. also performed the SV and GWAS analysis. V.L., J.H., A.F. and J.S. analysed the data. A.F. and J.S supervised the analysis. M.J.D. and J.S. contributed tools and materials. V.L. and J.S. wrote the paper with the input of A.F.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Statistics for 1,482 genome assemblies.
Each statistic is given according to the zygosity of the isolate and the phasing status of the assembly (homozygous isolates (n = 559), haplotype-resolved assemblies of heterozygous isolates (n = 792) and collapsed assemblies of heterozygous isolates (n = 131)). a. Distribution of the number of contigs per assembly. The red line corresponds to the number of chromosomes in the reference genome. b. Distribution of the total assembly length. The red line corresponds the reference genome length. c. Distribution of the Merqury quality value, which correspond to a Phred quality score. d. Distribution of the BUSCO score per isolate.
Extended Data Fig. 2 Rarefaction curves for the species coverage per SV type.
Species coverage represent the fraction of species non-redundant SVs captured by a population.
Extended Data Fig. 3 SV site frequency spectra (SFS).
a. SFS per SV type. b. Flattened SFS for different types of SV and SNPs. Letters on the right discriminate groups between which a two-sided Fisher’s exact test with FDR correction is significant with P < 0.05. c. SFS for Ty and non-Ty-related SVs.
Extended Data Fig. 4 Heterozygosity in structural variation.
All figures in this panel are based on 396 heterozygous isolates for which phased assemblies were constructed. a. Number of SV per isolate, colored by zygosity. b. Distribution of the fraction of heterozygous SVs per isolate. c. Spearman correlation between the fraction of heterozygous SVs and the SNP heterozygosity per isolate, computed as the number of heterozygous loci over the total number of callable positions. d. Heterozygosity level (defined as the frequency at which an SV is found heterozygous) between SVs located in subtelomeric and non-subtelomeric regions. The middle bar of the box plots corresponds to the median; the upper and lower bounds correspond to the third and first quartiles, respectively. The whiskers correspond to the upper and lower bounds 1.5 times the interquartile range (IQR). P value was calculated using two-sided Mann-Whitney-Wilcoxon test (**** indicates P < 2.2 × 10−16). e. Heterozygosity level per type of SV. The middle bar of the box plots corresponds to the median; the upper and lower bounds correspond to the third and first quartiles, respectively. The whiskers correspond to the upper and lower bounds 1.5 times the interquartile range (IQR). Letters discriminate groups between which a two-sided Mann-Whitney-Wilcoxon test with FDR correction is significant with P < 0.05. f. Average heterozygosity level according to SVs length (1 kb bins). The upper and lower bounds indicate the mean plus or minus the standard deviation, respectively. Spearman correlation between the length and heterozygosity level of SVs was computed. g. SV length according to the type of SV. The middle bar of the box plots corresponds to the median; the upper and lower bounds correspond to the third and first quartiles, respectively. The whiskers correspond to the upper and lower bounds 1.5 times the interquartile range (IQR). Letters discriminate groups between which a two-sided Mann-Whitney-Wilcoxon test with FDR correction is significant with P < 0.05. The bimodal distribution of PAVs length reflects the presence of SVs associated with Ty elements ( ~ 6 kb) and solo long terminal repeats (LTR, ~300 bp).
Extended Data Fig. 5 Proportion of the SV types in the clades SV signature.
The number of SVs on the right represent unique SV events overrepresented in each clade (i.e., clade SV signature). The “Total” group represents all the SVs involved in any clade signature. The number of isolates per group is indicated between brackets on the y axis. P values were computed using Pearson’s chi-squared tests with FDR correction in comparison to the Total group. P value significance is indicated with ** (P < 0.01), *** (P < 0.001) and **** (P < 0.0001).
Extended Data Fig. 6 Gene features depending on gene origin.
a. Distribution of gene location along chromosomes. b. Transcript level for genes per origin. Transcriptomes from 969 isolates were considered (Caudal et al. 20). Letters discriminate groups between which a two-sided Mann-Whitney-Wilcoxon test with FDR correction is significant with P < 0.05. The middle bar of the box plots corresponds to the median; the upper and lower bounds correspond to the third and first quartiles, respectively. The whiskers correspond to the upper and lower bounds 1.5 times the interquartile range (IQR).
Extended Data Fig. 7 Heatmap of gene presence/absence in 1,086 Saccharomyces cerevisiae isolates.
Rows correspond to isolates, colored by ploidy, clade, and super clade, which regroup clades of analog ecological origins. Columns correspond to 3,365 non-private accessory genes, colored by origin (introgression, HGT, fast evolving, de novo, or gene present in the reference genome).
Extended Data Fig. 8 Deletion at the ALD2/ALD3 locus.
a. Schematic representation of the deletion carried by strain AAB, compared to the reference strain. Shaded regions indicate sequence identity higher than 95%. b. Frequency of the deletion detected in isolates from wine, beer, Asian fermentation, and wild ecological origins. P values were computed using two-sided Fisher’s exact tests with FDR correction between groups. **** indicates significance of the tests between Beer and any other group with P < 10−17 c. Projection of the isolates in which the deletion was detected on the neighbor joining tree of the 1,086 isolates constructed based on SNP markers. Colors represent the clades previously defined45. d. Schematic of the reconstruction of the ALD2/3 haplotype for isolates in which the deletion was detected. Each line corresponds to an isolate, and its clade is color coded according to c. Each column represents one discriminating site between ALD2 and ALD3 gene sequences (n = 125). Color indicates the correspondence with ALD2 or ALD3 gene sequence. The schematic representations below show the five reconstructed ALD2/3 haplotypes from the above heatmap.
Extended Data Fig. 9 Location of QTL along the genome.
Position of the QTL (x-axis) compared to the gene (y-axis) affected at transcript level (a) or protein abundance (b), according to the type of variant considered. Colors indicate if the QTL is considered as local or distant.
Extended Data Fig. 10 Effect size of QTL according to their distance from the affected gene (local or distant).
a. All types of variants were combined. b. Effect size per type of variant. P values were calculated using a two-sided Mann-Whitney-Wilcoxon test (**** indicates P < 3 × 10−9). The middle bar of the box plots corresponds to the median; the upper and lower bounds correspond to the third and first quartiles, respectively. The whiskers correspond to the upper and lower bounds 1.5 times the interquartile range (IQR). c. Proportion of local and distant association involving SNP-, InDel- and SV-QTL. Letters indicate significance of a two-sided Fisher’s exact test.
Supplementary information
Supplementary Information (download DOCX )
These files contain Supplementary Figs. 1–21, Supplementary Notes 1–4 and supplementary references.
Supplementary Tables (download XLSX )
Supplementary Tables 1–15.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Loegler, V., Thiele, P., Teyssonnière, E. et al. From genotype to phenotype with 1,086 near telomere-to-telomere yeast genomes. Nature 648, 649–658 (2025). https://doi.org/10.1038/s41586-025-09637-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09637-0