Abstract
The mammalian cerebral cortex comprises a complex neuronal network that maintains a precise balance between excitatory pyramidal neurons and inhibitory interneurons. Accumulating evidence indicates that specific interneuron subtypes form stereotyped microcircuits with distinct pyramidal neuron classes1,2,3. Here we show that pyramidal neurons have an active role in this process by promoting the survival and terminal differentiation of their associated interneuron subtypes. In the wild-type cortex, interneuron subtype abundance mirrors the prevalence of their pyramidal neuron partners. In Fezf2 mutants, which lack L5b pyramidal neurons and are expanded in L6 intratelencephalic neurons, corresponding subtype-specific shifts occur through two distinct mechanisms: somatostatin interneurons adjust their programmed cell death, whereas parvalbumin interneurons switch their subtype identity. Silencing neuronal activity or blocking vesicular release in L5b pyramidal neurons revealed that their communication with interneurons does not require voltage-gated synaptic activity but engages both tetanus toxin-sensitive and tetanus toxin-insensitive pathways. Moreover, a targeted bioinformatic screen for ligand–receptor pairs displaying subtype-specific expression and reduced expression of pyramidal neuron-derived ligand in Fezf2 mutants identified candidate secreted factors and adhesion molecules. These findings reveal distinct, pyramidal neuron-driven mechanisms for sculpting interneuron diversity and integrating them into local cortical circuits.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data have been deposited at the GEO database under the accession number GSE272706 and at the Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP2716). The public datasets analysed in this study are available from the following sources: the Allen MERFISH spatial transcriptomics data12 were downloaded from the Allen Brain Cell Atlas (C57BL6J-638850-raw.h5ad; https://alleninstitute.github.io/abc_atlas_access/descriptions/MERFISH-C57BL6J-638850.html); additional snRNA-seq and scRNA-seq datasets were downloaded from the GEO database under the accession numbers GSE164570 (ref. 11), GSE158096 (ref. 25) and GSE153164 (ref. 42); and processed SMART-seq v4 scRNA-seq datasets were obtained from their original publications12,46.
Code availability
Emergene (https://github.com/genecell/Emergene) and PIASO (https://github.com/genecell/PIASO) are available on GitHub.
References
Yoshimura, Y. & Callaway, E. M. Fine-scale specificity of cortical networks depends on inhibitory cell type and connectivity. Nat. Neurosci. 8, 1552–1559 (2005).
Naka, A. et al. Complementary networks of cortical somatostatin interneurons enforce layer specific control. eLife 8, e43696 (2019).
Wu, S. J. et al. Cortical somatostatin interneuron subtypes form cell-type-specific circuits. Neuron 111, 2675–2692.e9 (2023).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Tasic, B. et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018).
Yao, Z. et al. A taxonomy of transcriptomic cell types across the isocortex and hippocampal formation. Cell 184, 3222–3241.e26 (2021).
Krook-Magnuson, E., Varga, C., Lee, S.-H. & Soltesz, I. New dimensions of interneuronal specialization unmasked by principal cell heterogeneity. Trends Neurosci. 35, 175–184 (2012).
Bortone, D. S., Olsen, S. R. & Scanziani, M. Translaminar inhibitory cells recruited by layer 6 corticothalamic neurons suppress visual cortex. Neuron 82, 474–485 (2014).
Muñoz, W., Tremblay, R., Levenstein, D. & Rudy, B. Layer-specific modulation of neocortical dendritic inhibition during active wakefulness. Science 355, 954–959 (2017).
Lu, J. et al. Selective inhibitory control of pyramidal neuron ensembles and cortical subnetworks by chandelier cells. Nat. Neurosci. 20, 1377–1383 (2017).
Nigro, M. J., Hashikawa-Yamasaki, Y. & Rudy, B. Diversity and connectivity of layer 5 somatostatin-expressing interneurons in the mouse barrel cortex. J. Neurosci. 38, 1622–1633 (2018).
Allaway, K. C. et al. Genetic and epigenetic coordination of cortical interneuron development. Nature 597, 693–697 (2021).
Yao, Z. et al. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 624, 317–332 (2023).
Hilscher, M. M., Leão, R. N., Edwards, S. J., Leão, K. E. & Kullander, K. Chrna2-Martinotti cells synchronize layer 5 type A pyramidal cells via rebound excitation. PLoS Biol. 15, e2001392 (2017).
Scala, F. et al. Layer 4 of mouse neocortex differs in cell types and circuit organization between sensory areas. Nat. Commun. 10, 4174 (2019).
Fazzari, P., Mortimer, N., Yabut, O., Vogt, D. & Pla, R. Cortical distribution of GABAergic interneurons is determined by migration time and brain size. Development 147, dev185033 (2020).
Hevner, R. F., Daza, R. A. M., Englund, C., Kohtz, J. & Fink, A. Postnatal shifts of interneuron position in the neocortex of normal and reeler mice: evidence for inward radial migration. Neuroscience 124, 605–618 (2004).
Lodato, S. et al. Excitatory projection neuron subtypes control the distribution of local inhibitory interneurons in the cerebral cortex. Neuron 69, 763–779 (2011).
Ye, Z. et al. Instructing perisomatic inhibition by direct lineage reprogramming of neocortical projection neurons. Neuron 88, 475–483 (2015).
Darbandi, S. F. et al. Neonatal Tbr1 dosage controls cortical layer 6 connectivity. Neuron 100, 831–845.e7 (2018).
Wester, J. C. et al. Neocortical projection neurons instruct inhibitory interneuron circuit development in a lineage-dependent manner. Neuron 102, 960–975.e6 (2019).
Yook, C. et al. A TBR1-K228E mutation induces Tbr1 upregulation, altered cortical distribution of interneurons, increased inhibitory synaptic transmission, and autistic-like behavioral deficits in mice. Front. Mol. Neurosci. 12, 241 (2019).
Molyneaux, B. J., Arlotta, P., Hirata, T., Hibi, M. & Macklis, J. D. Fezl is required for the birth and specification of corticospinal motor neurons. Neuron 47, 817–831 (2005).
Chen, B. et al. The Fezf2–Ctip2 genetic pathway regulates the fate choice of subcortical projection neurons in the developing cerebral cortex. Proc. Natl Acad. Sci. USA 105, 11382–11387 (2008).
Matho, K. S. et al. Genetic dissection of the glutamatergic neuron system in cerebral cortex. Nature 598, 182–187 (2021).
Stogsdill, J. A. et al. Pyramidal neuron subtype diversity governs microglia states in the neocortex. Nature 608, 750–756 (2022).
Tsyporin, J. et al. Transcriptional repression by FEZF2 restricts alternative identities of cortical projection neurons. Cell Rep. 35, 109269 (2021).
Southwell, D. G. et al. Intrinsically determined cell death of developing cortical interneurons. Nature 491, 109–113 (2012).
Priya, R. et al. Activity regulates cell death within cortical interneurons through a calcineurin-dependent mechanism. Cell Rep. 22, 1695–1709 (2018).
Wong, F. K. et al. Pyramidal cell regulation of interneuron survival sculpts cortical networks. Nature 557, 668–673 (2018).
Xu, Q., Tam, M. & Anderson, S. A. Fate mapping Nkx2.1-lineage cells in the mouse telencephalon. J. Comp. Neurol. 506, 16–29 (2008).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Klein, D. et al. Mapping cells through time and space with moscot. Nature 638, 1065–1075 (2025).
Thompson, C. L. et al. A high resolution spatiotemporal atlas of gene expression of the developing mouse brain. Neuron 83, 309–323 (2014).
Lodato, S. et al. Gene co-regulation by Fezf2 selects neurotransmitter identity and connectivity of corticospinal neurons. Nat. Neurosci. 17, 1046–1054 (2014).
Tuncdemir, S. N. et al. Early somatostatin interneuron connectivity mediates the maturation of deep layer cortical circuits. Neuron 89, 521–535 (2016).
Marques-Smith, A. et al. A transient translaminar GABAergic interneuron circuit connects thalamocortical recipient layers in neonatal somatosensory cortex. Neuron 89, 536–549 (2016).
Anastasiades, P. G. et al. GABAergic interneurons form transient layer-specific circuits in early postnatal neocortex. Nat. Commun. 7, 10584 (2016).
Favuzzi, E. et al. GABA-receptive microglia selectively sculpt developing inhibitory circuits. Cell 184, 4048–4063.e32 (2021).
Lepiemme, F. et al. Oligodendrocyte precursors guide interneuron migration by unidirectional contact repulsion. Science 376, eabn6204 (2022).
Irala, D. et al. Astrocyte-secreted neurocan controls inhibitory synapse formation and function. Neuron 112, 1657–1675.e10 (2024).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seq v2. Nat. Biotechnol. 39, 313–319 (2021).
Hirata, T. et al. Zinc finger gene fez-like functions in the formation of subplate neurons and thalamocortical axons. Dev. Dyn. 230, 546–556 (2004).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Dai, M., Pei, X. & Wang, X.-J. Accurate and fast cell marker gene identification with COSG. Brief. Bioinformatics 23, bbab579 (2022).
Di Bella, D. J. et al. Molecular logic of cellular diversification in the mouse cerebral cortex. Nature 595, 554–559 (2021)
Yao, Z. et al. A transcriptomic and epigenomic cell atlas of the mouse primary motor cortex. Nature 598, 103–110 (2021).
Jin, S. et al. Inference and analysis of cell–cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Moffitt, J. R. et al. High-throughput single-cell gene-expression profiling with multiplexed error-robust fluorescence in situ hybridization. Proc. Natl Acad. Sci. USA 113, 11046–11051 (2016).
Zhang, M. et al. Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598, 137–143 (2021).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Stickels, R. et al. Library generation using Slide-seq v2. protocols.io https://www.protocols.io/view/library-generation-using-slide-seqv2-bxijpkcn (2021).
Cable, D. M. et al. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat. Biotechnol. 40, 517–526 (2022).
Acknowledgements
We thank P. Arlotta and S. Lodato for providing insightful discussions and suggestions on the manuscript; P. Arlotta for providing the Fezf2-KO and Tcerg1lcreER mouse line; P. Keeley for providing the Baxfl mouse line; K. Kullander for providing the Chrna2–cre mouse line; D. Ginty for providing the Rorbcre mouse line; D. A. Stafford for providing the Hpsecre mouse line; R. Raichur for assistance on pre-processing the Slide-seq data; the former co-op and rotational students L. Abbott, M. Kim and A. Shinde for experiments that provided supporting data for this study, and S. Du for additional technical assistance; the Neurobiology Department and the Neurobiology Imaging Facility for consultation and instrument availability that supported this work. This work was supported by grants from the National Institutes of Health (NIH) R01NS081297 and R37MH071679 to G.F., the William Randolph Hearst Fund (FY20) to S.J.W., the F32 fellowship from National Institute of Mental Health (NIMH) F32MH125464 to S.J.W. and the NIMH 1U01MH130962 grant (to P. Arlotta) which supported C.M.
Author information
Authors and Affiliations
Contributions
S.J.W. and G.F. conceptualized the study. The investigation was performed by S.J.W. (snRNA-seq, Slide-seq, MERFISH, viral injection and revision experiments), S.-P.Y. (Slide-seq, RNAscope and immunostaining), Y.Q. (Slide-seq), C.M. (MERFISH), S.H. (snRNA-seq), D.S. (retro-orbital injection) and M.D., J.T. and B.C. (revision experiments). M.D. and S.J.W. conducted formal analysis. V.K., G.J.M., E.Z.M., F.C., S.L.F., Q.X., J.A.S. and D.J.D.B. provided resources. S.J.W., M.D., C.M. and G.F. wrote the original draft. S.L.F., G.F., S.J.W. and M.D. were responsible for writing, reviewing and editing the manuscript. G.F. supervised the project. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
G.F. is a founder of Regel Therapeutics, which has no competing interests with the present article.
Peer review
Peer review information
Nature thanks Trygve Bakken and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization and regional transcriptomic analysis of PVALB and SST interneuron subtypes.
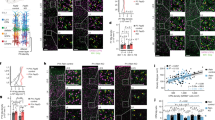
a, Dot plot showing expression of marker genes for each PVALB and SST interneuron subtype. b, Correspondence between subtypes identified in this study and previously defined supertypes6. c, Regional transcriptomic differences of PVALB and SST interneurons between MOs and VISp, determined using Emergene (see Methods). Emergene calculates a P value for region-specific gene signatures in individual nuclei via permutation tests. d, Heatmap showing the percentage of cells per subtype with significant regional differences. Subtypes with ≥50% significant cells are annotated with the median P value (calculated by Emergene). n.s., not significant; P ≥ 0.05 *P < 0.05; N/A, not analyzed (cluster size <10). Exact P values are provided in Supplementary Table 3. e, Proportions of all PVALB and SST subtypes in MOs and VISp, complete version of Fig. 1b. f, Intersectional genetic strategy preferentially targeting PVALB–Rorb interneurons, showing their axons concentrated in L4 (n = 4 mice). g, Intersectional genetic strategy preferentially targeting PVALB–Fzd6 interneurons, showing that these interneurons reside in L5b and extend their axons laterally within L5b. Sparse labeling achieved by low-dose tamoxifen administration allowed reconstruction of individual PVALB–Fzd6 morphologies, with one example shown (right), representative of n = 8 mice tested with various Tamoxifen dosages. For f-g, note that both strategies have off-target labeling of L5 PT neurons specifically in SSp regions. Scale bars, 100 µm.
Extended Data Fig. 2 Regional differences in the proportion of different PVALB and SST interneuron subtypes.
a, Proportion of all PN, PVALB, and SST interneuron subtypes in MOs, SSp, and VISp cortices based on MERFISH data. b, Violin plots showing the laminar distribution of selected PVALB and SST subtypes across three cortical regions. c, Bar plots showing the layer-by-layer composition of PVALB and SST subtypes in each region. d, Boxplots showing the ratio of interneurons (INs) to PNs and the contribution of PVALB and SST subtypes to the total interneuron pool across regions (Center line, median; box limits, 25th and 75th percentiles; whiskers, 1.5xIQR). Same dataset as in Fig. 1d, n = 9 (MOs), 13 (SSp), 9 (VISp) ROIs. Two-sided Wilcoxon rank-sum test, n.s. not significant, P ≥ 0.05; * P < 0.05; ** P < 0.01; *** P < 0.001. Exact P values are provided in Supplementary Table 3. e–h, Validation of regional interneuron densities by immunolabeling and genetic targeting. e, PVALB interneurons were labeled by immunostaining (n = 3 mice; P26-53). Note: This antibody-based method was chosen as it Is largely specific to PVALB interneurons, whereas Pvalb in situ hybridization (see Extended Data Fig. 7a) or Pvalb driver based genetic labeling, which also label L5 PT neurons in SSp. f, SST interneurons were visualized using genetic labeling. Comparable distributions were confirmed by in situ hybridization for Sst mRNA (n = 5 mice total across both methods; P26-61). g, Labeling of L4-targeting SST–Hpse interneurons with some off-target SST-Calb2 interneurons (n = 3 mice; P30-36); h, Labeling of L5b SST–Chrna2 interneurons (n = 5 mice; P34-P40). For e-h, Scale bars, 500 µm (main) and 100 µm (higher magnification of outlined regions on the right). Exact sample genotypes are provided in Supplementary Table 2. i, UMAP of P28 snRNA-seq data based on only the genes included in the MERFISH probe set, demonstrating that this gene set is sufficient to resolve the defined subtypes.
Extended Data Fig. 3 PVALB and SST interneuron subtypes show stereotyped laminar distributions.
a, Violin plots showing the laminar distribution of all PVALB and SST subtypes across MOs, SSp, and VISp. b-c, Gallery of MERFISH spatial maps from additional coronal sections showing the consistent distribution of all identified PVALB and SST interneurons.
Extended Data Fig. 4 Characterization of pyramidal neuron and interneuron changes in Fezf2 mutants.
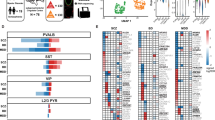
a–d, Analysis of PN subtypes in the P14 Fezf2 KO SSp cortex26. b, Dot plot showing marker gene expression. c, Heatmap showing transcriptomic correspondence between control and KO subtypes. d, Quantification of deep-layer PN subtype proportions. e–h, Analysis of interneuron subtypes in the Fezf2 KO. e, UMAP (as in Fig. 2a) highlighting all subtypes with altered proportions or transcriptomes. f, Bar plot (as in Fig. 2b), highlighting additional subtypes that showed changes in Fezf2 mutants. g, Transcriptomic differences between PVALB and SST interneurons in control and Fezf2 KO conditions, determined using Emergene as in Extended Data Fig. 1c. h, Heatmap showing the percentage of cells per subtype with significant differences across genotypes. Subtypes with ≥50% significant cells are annotated with the median P-value as calculated by Emergene. N/A, not analyzed (cluster size <10). n.s., not significant, P ≥ 0.05; * P < 0.05; *** P < 0.001). Exact P values are provided in Supplementary Table 3. i, Genetic labeling of SST–Chrna2 interneurons in different cortical regions of control and Fezf2 mutant mice (representative of n = 3 control and n = 5 KO mice; age P28-P29). j, RNAscope in situ hybridization (ISH) for Nmbr and Crhr2 mRNA (markers for SST-Nmbr-1/2 subtypes) in control and Fezf2 KO mice. The image shown (P28) is representative of n = 4 control and n = 4 KO mice (age range P27-P31). k, Viral genetic labeling of SST–Nmbr-1/2 interneurons in control and Fezf2 mutant SSp cortices (n = 3 control, n = 3 KO; age P46-62). For i-k, Scale bar: 100 µm. CC, corpus callosum; HP, hippocampus. Exact sample genotypes are provided in Supplementary Table 2.
Extended Data Fig. 5 Spatial Transcriptomic analysis of cortical interneuron changes in Fezf2 mutants.
a, Representative Slide-seq data on a single coronal brain sections of the SSp cortex from WT (left) and Fezf2 KO (right) mouse at P37. b-c, Aggregate Slide-seq data from multiple brain sections, showing the overall distribution of PNs in control (b) and Fezf2 KO (c) cortices. Sample sizes are presented as n = mice (from X ROIs). WT control: MO, n = 3 (4); SSp, n = 5 (8); VISp: n = 4 (4). Age: P28-37. Fezf2 KO: MO, n = 3 (3); SSp, n = 5 (6); VISp, n = 2 (4). Age: 4–6 weeks. d-e, Identification of deep-layer PVALB and SST interneuron subtypes within the aggregate control (d) and Fezf2 KO (e) datasets. (same dataset as in b-c). f, Ridge plots showing the distribution of selected PVALB and SST subtypes in the deep layers of control and Fezf2 KO brains, based on MERFISH data as in Fig. 2c.
Extended Data Fig. 6 Fezf2 deletion causes a spatial redistribution of mid-layer interneuron subtypes.
a, b, RNAscope ISH confirms an upward laminar shift of the total PVALB (a) and SST (b) interneuron populations in the SSp of P28 Fezf2 KO mice. Scale bar, 100 µm. Bar plots (right) quantify cell proportion per layer (mean ± SD). Sample sizes are n = mice (ROIs). control: n = 5 (10), age P26-28, n = 3932 PVALB, n = 2280 SST. Fezf2 KO: n = 3 (5), age P27-28 n = 2377 PVALB, n = 1677 SST. Statistical significance was determined using a LMM with animal ID as a random effect. n.s., not significant, P ≥ 0.05; * P < 0.05; ** P < 0.01; *** P < 0.001. c, Bar plots showing the composition of each PVALB and SST interneuron subtypes in different cortical layers of control and Fezf2 KO brains in the SSp region, based on MERFISH dataset. control: n = 2 (3), Fezf2 KO: n = 3 (3), age: P14-30. d, Representative RNAscope ISH images of SST–Calb2 interneurons in control and Fezf2 KO SSp cortex, which show no obvious laminar shift. Signal is visualized by a Calb2–Vip subtraction. Note: this strategy also labels L6 SST–Chodl interneurons (n = 3 control, n = 3 KO; P28). e, RNAscope ISH images of Hpse mRNA in SSp of control and Fezf2 KO brains, showing a denser and broader accumulation of SST-Hpse interneurons near the L4/L5 boundary in the Fezf2 KO (arrow). Note: L5 PNs express low levels of Hpse (n = 4 control, n = 5 KO; P27-P31). f, Representative images of viral genetically labeled SST–Hpse interneurons in control and Fezf2 mutants at P47 (n = 1 KO; cohort size was ethically limited due to high post-operative mortality in the HpseCre;Fezf2lacZ/lacZ background). For d-f, Scale bar: 100 µm. Exact sample genotypes, statistical model details, and P values are provided in Supplementary Tables 2 and 3.
Extended Data Fig. 7 Conditional removal of Bax in PVALB and SST interneurons increases their number.
a, PVALB immunolabeling on Nkx2.1-Cre labeling, showing incomplete coverage of superficial interneurons (n = 4, P26-28). b, Representative Pvalb and Gad1 ISH in SSp, showing Pvalb mRNA in Gad1-negative L5 PT neurons (n = 1; finding confirmed in n = 4 mice with Pvalb/Lhx6 co-labeling). c, RNAscope ISH for Pvalb and Sst mRNA in SSp of P28-33 mice from four genotypes. DAPI (left) provides laminar reference. d, Quantification of Nkx2.1-Cre labeled interneurons in control and Bax cKO conditions. n=mice (ROIs); control: n = 4 (4), 2316 interneurons; Bax cKO: n = 4 (4), 3548 interneurons. e, Laminar distribution of genetically labeled interneurons from d. f, Quantification of PVALB and SST interneurons in control and Bax cKO SSp cortices (L5b Pvalb+/Lhx6− neurons excluded). Numbers normalized to pial length (mm). control: n = 5 (10), age P26-28, 3932 PVALB, 2280 SST, 337 Pvalb +/Sst+ interneurons. Bax cKO: n = 4 (8), age P26-28, 3470 PVALB, 2150 SST, 221 total Pvalb +/Sst+ interneurons. g, Quantification of PVALB and SST interneurons in Fezf2 KO and Fezf2 KO_Bax cKO SSp cortices. Fezf2 KO: n = 4 (6), age P28-33, 1668 PVALB, 1257 SST, 138 Pvalb +/Sst+ interneurons. Fezf2 KO_Bax cKO: n = 4 (6), age P28-33, 2161 PVALB, 1768 SST, 175 Pvalb +/Sst+ interneurons. h, Laminar distribution of interneurons from f. i, UMAP plots of P14 snRNA-seq data on interneurons from four genotypes, showing Bax cKO-specific clusters. j, UMAPs plots showing normalized expression level of marker genes for Bax cKO-specific clusters. k, Stacked bar plots showing PVALB and SST subtype proportions across biological replicates of Bax cKO and Fezf2 KO_Bax cKO snRNA-seq datasets. Scale bars (a-c), 100 µm. For (e, h), Error bars, s.d. d, two-sided Wilcoxon rank-sum test. For e-h, LMM with animal ID as a random effect. n.s. not significant, P ≥ 0.05; * P < 0.05. Exact sample genotypes, statistical model details, and P values are provided in Supplementary Tables 2 and 3.
Extended Data Fig. 8 Spatial transcriptomics analysis of Bax removal in PVALB and SST interneurons.
a, Additional MERFISH data at P14 from different genotypes, complementing Fig. 3c. b, Quantification of selected interneuron subtype numbers in the SSp region (MERFISH data), normalized to deep-layer PN count. Values are not directly comparable to Slide-seq due to the higher spatial resolution and cell-calling recovery rate of MERFISH. No statistical test was performed due to limited sample size. n=mice (ROIs); Control: n = 2(3), Bax cKO: n = 1(2), Fezf2 KO: n = 3(3), Fezf2 KO_Bax cKO: n = 1(2). c, Proportion of selected PVALB and SST interneuron subtypes within total L5/6 PVALB and SST interneurons, based on Slide-seq data, used to justify pooling of heterozygous genotypes in control group. A detailed statistical justification for pooling is provided in Supplementary Note. Fezf2 WT: n = 5 (8); Fezf2 HET_Bax cHET: n = 2 (3); Fezf2 KO: n = 5 (6); Fezf2 KO_Bax cHET: n = 2 (4). These data are included in Figs. 2d and 3d. d, Aggregate Slide-seq maps of coronal sections from Bax cKO and Fezf2 KO_Bax cKO mice, highlighting selected PVALB and SST interneuron subtypes identified in L5/6. Bax cKO: n = 3 (5); Fezf2 KO_Bax cKO: n = 3 (5). Age range: P28-33. e, Quantification of selected PVALB and SST subtypes (Slide-seq data), normalized to L5/6 PN counts, comparing control (n = 7 mice, 11 ROIs) and Bax cKO (n = 3 mice, 5 ROIs). f, Quantification of selected PVALB and SST subtypes (Slide-seq data), normalized to L5/6 PN counts, comparing between Fezf2 KO (n = 7 mice, 10 ROIs) and Fezf2 KO_Bax cKO (n = 3 mice, 4 ROIs). Boxplots: center line, median; box limits, 25th–75th percentiles; whiskers, 1.5xIQR. For c,e,f, LMM with animal ID as random effect. n.s., not significant, P ≥ 0.05; * P < 0.05; ** P < 0.01. Exact sample genotypes, sample ages, statistical model details and P values are provided in Supplementary Tables 2 and 3.
Extended Data Fig. 9 Interneuron subtype changes in Fezf2 mutants occur after P2.
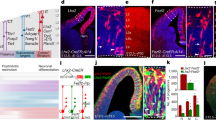
a, UMAP visualization of snRNA-seq data of cortical interneurons collected from control and Fezf2 KO mouse brains at P7, highlighting key deep-layer subtypes. b, Stacked bar plots showing the proportion of deep-layer PVALB and SST subtypes in control and Fezf2 KO brains based on snRNA-seq data. c, UMAP visualization of published snRNA-seq data46 of excitatory neurons from P1 Fezf2 HET and Fezf2 KO mouse cortices. d, River plot showing the transcriptomic correspondence between control and KO subtypes. e, Stacked bar plots showing PN subtype composition in P1 Fezf2 HET and KO brains, based on snRNA-seq data. f-g, Feature genes identified in P7 snRNA-seq dataset as selective for PVALB–Fzd6 (f) and SST–Chrna2 (g) are already selectively expressed at P2. (Left) UMAP plots showing feature gene expression. (Right) Violin plots comparing the expression level of feature genes in putative PVALB-Fzd6 and SST-Chrna2 clusters versus all other nuclei; dots and error bars represent mean ± s.d. Two-sided Wilcoxon rank-sum test. Sample size: (f) PVALB–Fzd6 (n = 717) vs. Others (n = 7263); (g) SST–Chrna2 (n = 538) vs. Others (n = 7442). h, UMAP visualization of snRNA-seq data of cortical interneurons collected from control and Fezf2 KO mouse brains at P2, with PVALB–Fzd6 and SST–Chrna2 interneuron subtypes highlighted. i, MERFISH spatial map of coronal brain sections from the SSp region of control and Fezf2 KO mice at P2, showing PN and selected interneuron subtypes. CR, Cajal-Retzius cells; i.p., intermediate progenitor; m.n., migrating neurons.
Extended Data Fig. 10 Transcriptomic plasticity of PVALB and SST interneurons in Fezf2 mutants.
a, RNA velocity analysis of snRNA-seq data on SST interneurons from control and Fezf2 KO cortices at P7, showing gene expression dynamics of three SST subtypes. b, Optimal transport analysis mapping the developmental trajectories of SST subtypes from P7 to P14 in control and Fezf2 KO conditions. Descendent analysis (left) shows predictive assignment from P7 to P14 subtypes; ancestor analysis (right) shows predicted P7 origins of P14 subtypes. c, Expression scores for PVALB–Fzd6 (P7), PVALB–Slc39a8 (P7), and immature PVALB–Fzd6 (P2) gene sets (see Methods) in PVALB–Fzd6 interneurons from Fezf2 mutants, benchmarked against control references. Sample sizes: Fzd6@P2 (n = 471), Fzd6@P7 (n = 394), Fzd6@P7_Fezf2 KO (n = 274), Slc39a8@P7 (n = 365). d, Gene set analysis, as in (c), for SST–Chrna2 interneurons in Fezf2 mutants at P7, benchmarked against control references. Sample sizes: Chrna2@P2 (n = 355), Chrna2@P7 (n = 314), Chrna2@P7_Fezf2 KO (n = 214), Nmbr1/2@P7 (n = 324). e, Triangular Affinity Map of PVALB–Fzd6 interneurons in mutant condition at P7 (see Methods), showing relative transcriptomic similarity to the three reference groups from (c). f, Triangular Affinity Map, as in (e), for SST–Chrna2 interneurons in P7 mutant, showing similarity to the reference groups from (d). For c, d, two-sided Wilcoxon rank-sum test.
Extended Data Fig. 11 Additional Spatial data and transcriptomic validation of PN manipulation experiments.
a,c-e, Additional MERFISH spatial maps of control, Kir2.1, and TeNT conditions, supplementing Fig. 5a and contributing to quantification in Fig. 5b. b, RNAscope ISH confirming induced Kcnj2 (encoding Kir2.1) and mCherry expression in Fezf2+ L5b PNs (n = 3 control, n = 3 Kir2.1 group). f, UMAP visualization of integrated snRNA-seq datasets from control, Fezf2 KO (dataset as in Fig. 2a), and TeNT conditions. The analysis compares the transcriptomic state of PVALB-Fzd6 interneurons in the TeNT condition (left) and the Fezf2 KO condition (right) relative to pooled controls, revealing a more pronounced transcriptomic shift in the remaining cells of the TeNT condition. Exact sample genotypes are provided in Supplementary Table 2.
Extended Data Fig. 12 Additional candidate ligand–receptor interactions and Wnt5a expression.
a, Lollipop diagram, as in Fig. 5c, showing the 51st–100th ranked candidate ligand/receptor pairs from the bioinformatic screen. See Supplementary Tables 4 and 5 for a complete list of the top 100 ligand-receptor pairs.b, Representative in situ hybridization image for Wnt5a at P4, showing specific expression in L5b. Image from Allen developing mouse brain atlas: in situ hybridization (ISH) data34 (https://developingmouse.brain-map.org/experiment/show/100101325).
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods: Additional methodology description for FACS procedure, immunostaining, stereotaxic AAV injection, RO AAV injection, Tamoxifen administration, RNAscope in situ hybridization, Image Acquisition. Supplementary Notes: Statistical Validation for Pooling Heterozygous Genotypes.
Supplementary Table 1 (download XLSX )
List of MERFISH probe set.
Supplementary Table 2 (download XLSX )
Sample genotypes.
Supplementary Table 3 (download XLSX )
Statistical test and P values.
Supplementary Table 4 (download XLSX )
Top 100 ligand-receptor pairs for PVALB-Fzd6.
Supplementary Table 5 (download XLSX )
Top 100 ligand-receptor pairs for SST-Chrna2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, S.J., Dai, M., Yang, SP. et al. Pyramidal neurons proportionately alter cortical interneuron subtypes. Nature 651, 421–428 (2026). https://doi.org/10.1038/s41586-025-09996-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-09996-8