Abstract
Rice is a staple crop for more than half of the world’s population, and its sustainable production is vital to ensure global food security. However, rice is susceptible to several devastating fungal diseases1, including blast disease caused by Magnaporthe oryzae, sheath blight by Rhizoctonia solani, false smut by Ustilaginoidea virens, brown spot by Bipolaris oryzae, bakanae by Fusarium fujikuroi and head blight by Fusarium graminearum. The mechanisms underlying the susceptibility to these fungal diseases remain unclear. Here we report that the β subunit of SnRK1, SnRK1β1A, confers broad-spectrum susceptibility to these fungal diseases. Our findings show that diverse rice fungal pathogens have convergently evolved an effector-like protein, Gas2, which interacts with SnRK1β1A to prevent its ubiquitination-mediated degradation and promotes its nuclear translocation. SnRK1β1A is markedly induced on fungal infection, promoting susceptibility by inhibiting SnRK1α1, an α subunit of SnRK1 known to positively regulate broad-spectrum resistance in rice2. Notably, rice lines with disrupted SnRK1β1A are resistant to several fungal diseases without compromising growth and yield in the field under normal farming conditions. This study demonstrates that broad-spectrum disease resistance in crops can be achieved by disrupting inducible susceptibility genes whose encoded proteins are targeted by effectors conserved across several pathogens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The RNA-seq dataset has been deposited in the NCBI Sequence Read Archive under accession numbers SRR33600040–SRR33600063. Other sequences can be accessed under the following numbers: SnRK1β1A (LOC_Os05g41220, XP_052157421.1), SnRK1α1 (LOC_Os05g45420, NP_001389545.1), XB24 (LOC_Os01g56470, XP_015612864.1), GAS2 (XP_003719620.1), GAS2L (XP_003709024.1), BoGAS2 (XP_007693280.1), RsGAS2 (CCO30280.1), FfGAS2 (XP_023432757.1), FgGAS2 (XP_011320808.1), UvGAS2 (XP_043001658.1), BoGAS2L (XP_007687900.1), NcMAS1 (XP_960032.1) and NcMAS2 (XP_962708.1). All data are available in this Article and its Supplementary Information. Original gel blots are shown in Supplementary Fig. 1. Source data are provided with this paper.
References
Ou, S. H. Rice Diseases 2nd edn (C.A.B., 1985).
Filipe, O., De Vleesschauwer, D., Haeck, A., Demeestere, K. & Höfte, M. The energy sensor OsSnRK1a confers broad-spectrum disease resistance in rice. Sci. Rep. 8, 3864 (2018).
Savary, S. et al. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 3, 430–439 (2019).
Eckardt, N. A. Plant disease susceptibility genes? Plant Cell 14, 1983–1986 (2002).
Pavan, S., Jacobsen, E., Visser, R. G. F. & Bai, Y. Loss of susceptibility as a novel breeding strategy for durable and broad-spectrum resistance. Mol. Breeding 25, 1–12 (2010).
van Schie, C. C. & Takken, F. L. Susceptibility genes 101: how to be a good host. Annu. Rev. Phytopathol. 52, 551–581 (2014).
Piffanelli, P. et al. A barley cultivation-associated polymorphism conveys resistance to powdery mildew. Nature 430, 887–891 (2004).
Fukuoka, S. et al. Loss of function of a proline-containing protein confers durable disease resistance in rice. Science 325, 998–1001 (2009).
Garcia-Ruiz, H., Szurek, B. & Van den Ackerveken, G. Stop helping pathogens: engineering plant susceptibility genes for durable resistance. Curr. Opin. Biotech. 70, 187–195 (2021).
Zaidi, S. S., Mukhtar, M. S. & Mansoor, S. Genome editing: targeting susceptibility genes for plant disease resistance. Trends Biotechnol. 36, 898–906 (2018).
Li, S. et al. Genome-edited powdery mildew resistance in wheat without growth penalties. Nature 602, 455–460 (2022).
Wang, N. et al. Inactivation of a wheat protein kinase gene confers broad-spectrum resistance to rust fungi. Cell 185, 2961–2974 (2022).
Zhou, Y. et al. Engineering of rice varieties with enhanced resistances to both blast and bacterial blight diseases via CRISPR/Cas9. Plant Biotechnol. J. 20, 876–885 (2022).
Tao, H. et al. Engineering broad-spectrum disease-resistant rice by editing multiple susceptibility genes. J. Integr. Plant Biol. 63, 1639–1648 (2021).
Oliva, R. et al. Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat. Biotechnol. 37, 1344–1350 (2019).
Sha, G. et al. Genome editing of a rice CDP-DAG synthase confers multipathogen resistance. Nature 618, 1017–1023 (2023).
Li, Y. J. et al. Genome editing of the susceptibility gene ZmNANMT confers multiple disease resistance without agronomic penalty in maize. Plant Biotechnol. J. 21, 1525–1527 (2023).
Thomazella, D. P. D. T. et al. Loss of function of a DMR6 ortholog in tomato confers broad-spectrum disease resistance. Proc. Natl Acad. Sci. USA 118, e2026152118 (2021).
Bi, W. et al. CRISPR/Cas9-guided editing of a novel susceptibility gene in potato improves Phytophthora resistance without growth penalty. Plant Biotechnol. J. 22, 4–6 (2024).
Gao, M. et al. Ca2+ sensor-mediated ROS scavenging suppresses rice immunity and is exploited by a fungal effector. Cell 184, 5391–5404 (2021).
Li, W. et al. A natural allele of a transcription factor in rice confers broad-spectrum blast resistance. Cell 170, 114–126 (2017).
Justesen, A., Somerville, S., Christiansen, S. & Giese, H. Isolation and characterization of two novel genes expressed in germinating conidia of the obligate biotroph Erysiphe graminis f.sp. hordei. Gene 170, 131–135 (1996).
Xue, C. et al. Two novel fungal virulence genes specifically expressed in appressoria of the rice blast fungus. Plant Cell 14, 2107–2119 (2002).
Grell, M. N., Mouritzen, P. & Giese, H. A Blumeria graminis gene family encoding proteins with a C-terminal variable region with homologues in pathogenic fungi. Gene 311, 181–192 (2003).
Martínez-Cruz, J. S. et al. Effectors with chitinase activity (EWCAs), a family of conserved, secreted fungal chitinases that suppress chitin-triggered immunity. Plant Cell 33, 1319–1340 (2021).
Liu, H. et al. Plant immunity suppression by an exo-β-1,3-glucanase and an elongation factor 1α of the rice blast fungus. Nat. Commun. 14, 5491 (2023).
Bouly, J. P., Gissot, L., Lessard, P., Kreis, M. & Thomas, M. Arabidopsis thaliana proteins related to the yeast SIP and SNF4 interact with AKINα1, an SNF1-like protein kinase. Plant J. 18, 541–550 (1999).
Yang, J. et al. SnRK1A-mediated phosphorylation of a cytosolic ATPase positively regulates rice innate immunity and is inhibited by Ustilaginoidea virens effector SCRE1. New Phytol. 236, 1422–1440 (2022).
Wang, W. et al. SnRK1 stimulates the histone H3K27me3 demethylase JMJ705 to regulate a transcriptional switch to control energy homeostasis. Plant Cell 33, 3721–3742 (2021).
Ramon, M. et al. Default activation and nuclear translocation of the plant cellular energy sensor SnRK1 regulate metabolic stress responses and development. Plant Cell 31, 1614–1632 (2019).
Han, X. et al. SnRK1 phosphorylates and destabilizes WRKY3 to enhance barley immunity to powdery mildew. Plant Commun. 1, 100083 (2020).
Dale, S. et al. Similar substrate recognition motifs for mammalian AMP-activated protein kinase, higher plant HMG-CoA reductase kinase-A, yeast SNF1, and mammalian calmodulin-dependent protein kinase I. FEBS Lett. 361, 191–195 (1995).
Swain, S. et al. Arabidopsis thaliana cdd1 mutant uncouples the constitutive activation of salicylic acid signalling from growth defects. Mol. Plant Pathol. 12, 855–865 (2011).
Polge, C. & Thomas, M. SNF1/AMPK/SnRK1 kinases, global regulators at the heart of energy control? Trends Plant Sci. 12, 20–28 (2007).
Crepin, N. & Rolland, F. SnRK1 activation, signaling, and networking for energy homeostasis. Curr. Opin. Plant Biol. 51, 29–36 (2019).
Hulsmans, S., Rodriguez, M., De Coninck, B. & Rolland, F. The SnRK1 energy sensor in plant biotic interactions. Trends Plant Sci. 21, 648–661 (2016).
Li, X. et al. AKINβ1 is involved in the regulation of nitrogen metabolism and sugar signaling in Arabidopsis. J. Integr. Plant Biol. 51, 513–520 (2009).
Gong, Z. et al. Two Magnaporthe appressoria-specific (MAS) proteins, MoMas3 and MoMas5, are required for suppressing host innate immunity and promoting biotrophic growth in rice cells. Mol. Plant Pathol. 23, 1290–1302 (2022).
Maurus, I. et al. Verticillium dahliae Vta3 promotes ELV1 virulence factor gene expression in xylem sap, but tames Mtf1-mediated late stages of fungus-plant interactions and microsclerotia formation. PLoS Pathog. 19, e1011100 (2023).
Ding, Y. et al. Host-induced gene silencing of a multifunction gene Sscnd1 enhances plant resistance against Sclerotinia sclerotiorum. Front. Microbiol. 12, 693334 (2021).
Yan, X. et al. The transcriptional landscape of plant infection by the rice blast fungus Magnaporthe oryzae reveals distinct families of temporally co-regulated and structurally conserved effectors. Plant Cell 35, 1360–1385 (2023).
Wang, Y. et al. An ERAD-related ubiquitin-conjugating enzyme boosts broad-spectrum disease resistance and yield in rice. Nat. Food 4, 774 (2023).
Himmelbach, A. et al. A set of modular binary vectors for transformation of cereals. Plant Physiol. 145, 1192 (2007).
Jiang, C. et al. An orphan protein of Fusarium graminearum modulates host immunity by mediating proteasomal degradation of TaSnRK1α. Nat. Commun. 11, 4382 (2020).
Zhou, Z., Bi, G. & Zhou, J. Luciferase complementation assay for protein–protein interactions in plants. Curr. Protoc. Plant Biol. 3, 42–50 (2018).
Chen, J. B. et al. An LRR-only protein promotes NLP-triggered cell death and disease susceptibility by facilitating oligomerization of NLP in Arabidopsis. New Phytol. 232, 1808–1822 (2021).
Liu, Y. et al. A designer rice NLR immune receptor confers resistance to the rice blast fungus carrying noncorresponding avirulence effectors. Proc. Natl Acad. Sci. USA 118, e2110751118 (2021).
Chen, H. et al. Cytoplasmic and nuclear Sw-5b NLR act both independently and synergistically to confer full host defense against tospovirus infection. New Phytol. 231, 2262–2281 (2021).
Deng, Y. et al. Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance. Science 355, 962–965 (2017).
Tang, C. et al. Study and application of the classification standard of rice false smut. Plant Prot. 27, 18–21 (2001).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Lin, X. et al. A molecular switch OsWRKY10-OsVQ8 orchestrates rice diterpenoid phytoalexin biosynthesis for broad-spectrum disease resistance. New Phytol. 246, 2243 (2025).
Zhan, C. et al. Selection of a subspecies-specific diterpene gene cluster implicated in rice disease resistance. Nat. Plants 6, 1447 (2020).
Liu, C. et al. eQTLs play critical roles in regulating gene expression and identifying key regulators in rice. Plant Biotechnol. J. 20, 2357–2371 (2022).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (grant no. 32293240 to Y.-L.P.), the Earmarked Fund for China Agricultural Research System (CARS-1-43 to J.Y.), the Pinduoduo-China Agricultural University Research Fund (grant no. PC2023A01005 to Y.-L.P.), the Joint Research Program of State Key Laboratory of Agricultural and Forestry Biosecurity (SKLJRP2508 to Y.-L.P.) and the 2115 Talents project from China Agricultural University (to Y.-L.P.). We are grateful to S. Chen for providing B. oryzae isolates, W. Sun for providing U. virens isolates, D.-X. Zhou for providing snrk1α1 mutant, G. Zhao for assisting in the in vitro kinase assay with radio isotopes, Y. Gao for assisting in bioinformatics analysis, D. Wang for assisting in modelling of the dimeric Gas2 structure, N. J. Talbot for critical review and constructive discussion and J.-R. Xu and S.-Y. He for their suggestions and discussions.
Author information
Authors and Affiliations
Contributions
Y.-L.P. conceived and designed the project. X.L. and G.Y. designed experiments and analysed data. G.Y. performed most of the experiments, including the generation of diverse fungal and rice mutants, rice transgenic lines and RNA-seq data, effector protein assays, Y2H, co-immunoprecipitation, BiFC, LCI, pull-down assays of protein–protein interactions, plant infection assays, subcellular localization, protein degradation assay, ubiquitination assay, kinase assays, ROS detection, MAPK detection, RT-qPCR/RT-PCR, western blots and phylogenetic analysis. X.W. contributed to the generation of fungal mutants, screening Gas2-interacting proteins by Y2H, plant infection assays and agronomic trait investigation. M.L. and Z.H. performed expression and purification of Gas2 and SnRK1β1A, structural analyses of the Gas2 and SnRK1β1A interaction, ROS detection in infected rice leaf sheath cells and SnRK1α1 kinase isotope assays. Z.L. analysed the RNA-seq data. F.Z. performed field disease surveys. S.W. and X.Z. predicted and analysed the structures of MAS proteins. X.W., J.Y., H.G., V.B., W.-S.Z., W. Z., M.Y. and J.-M.Z. analysed the data and provided critical feedback. Y.-L.P., G.Y., X.L. and V.B. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare a granted Chinese patent pertaining to SnRK1β1A (no. ZL 202310112719.3), filed by China Agricultural University with inventors Y.-L.P., G.Y., X.L. and X.W. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature thanks Yong Liang, Amir Sharon, Bing Yang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Gas2 is conserved in rice fungal pathogens.
a, Phylogenetic analysis and secondary structures of Gas1 and Gas2-homologous MAS proteins from rice fungal pathogens and Neurospora crassa. The phylogenetic tree was constructed in MEGA X using the Maximum Likelihood method. The secondary structures of MAS proteins were predicted with PSIPRED 4.0. b, Comparisons of the AlphaFold2 predicated 3D-structure of Gas2 from M. oryzae with its orthologues from Bipolaris oryzae (XP_007693280.1/BoGas2 and XP_007687900.1/BoGas2L), Rhizoctonia solani (CCO30280.1/RsGas2 and CCO30304.1/RsGas2L), F. fujikuroi (XP_023432757.1/FfGas2 and XP_023437241.1/FfGas2L), Fusarium graminearum (XP_011320808.1/FgGas2 and XP_011315585.1/FgGas2L) and Ustilaginoidea virens (XP_043001658.1/UvGas2), which cause brown spot, sheath blight, bakanae, head blight and false smut in rice, respectively. Different colour dots indicate MAS proteins from different rice fungal pathogens. c–g, Schematic diagrams of knockout strategies for generating Δgas2 (c), ΔBogas2 (d), ΔUvgas2 (e), ΔFggas2 (f) and ΔFfgas2 (g) and their verification. Δgas2 by Southern blot analysis; genomic DNA isolated from wild type (WT) P131 and the two Δgas2 mutants was digested with SalI. The blot analysis was performed by the digoxin kit instructions. HindIII-digested λ-DNA was used as a marker in c. Genomic DNA isolated from the other fungal mutants and its corresponding WT strains were amplified by PCR using specific primers (d–g). h, Virulence of F. graminearum ΔFggas2 mutant on Nipponbare (NPB) could be rescued by GAS2-orthologues from rice fungal pathogens but not by unrelated MAS from saprophytic N. crassa. Relative fungal biomass was measured by qPCR at 5 dpi. i, The WT virulence of Δgas2 mutant could not be rescued by introducing GAS2 without the C-terminus (Δgas2/GAS2ΔC). Rice leaves were sprayed with M. oryzae P131, Δgas2 and Δgas2/GAS2ΔC strains. Relative fungal biomass was measured by qPCR at 5 dpi. Data represent mean ± s.d.; n = biologically independent samples in the graphs. One-way ANOVA followed by post-hoc Tukey tests were used in (h,i); letters denote significantly different groups. Experiments were repeated three times (h,i) with similar results.
Extended Data Fig. 2 Gas2 is expressed during the appressorial penetration and suppresses chitin-triggered rice immunity.
a, Gas2 lacks chitinase activity. 2HA–Gas2 expressed in E. coli was purified and used to detect chitinase activity. Chitinase from Oryzae sativa was used as a positive control, and the boiled chitinase was used as a negative control. b, GAS2 is specifically expressed during the appressorial penetration. Relative expression of GAS2 normalized to MoActin was calculated using RT-qPCR with RNA extracted from mycelia hyphae (HY) and rice leaves inoculated with M. oryzae strain P131 at the indicated times. c, Gas2 expressed in the appressorium. Microscopic observation of Gas2–GFP driven by the native promoter was performed using inoculated rice leaf sheaths at the indicated time points. Scale bar, 20 μm. d, The signal peptide from Gas2 could lead to the secretion of SUC2 to allow the growth of yeast transformants on SD-Trp medium and YPRAA medium (top) and secretion of invertase into liquid medium (bottom). e, Gas2–mCherry–NLS could be observed in the nucleus of barely epidermal cells infected by an M. oryzae transformant expressing Gas2–mCherry–NLS at 30 h post-inoculation (hpi). Gas2–mCherry signals clearly overlapped with DAPI stained nucleus. Scale bar, 20 μm. f, Transgenic rice lines of NPB overexpressing 3HA-GAS2 (GAS2-OE) verified at transcriptional (top) and protein (bottom) levels. RNA and protein were extracted from the indicated rice leaves for RT-PCR assays and western blot analysis with anti-HA antibody, respectively. Actin was used as an internal control. g, Rice leaf sheath cells infected with Δgas2 showed stronger ROS production than cells infected with P131 or the complementary strain Δgas2/GAS2. At 36 hpi, rice leaf sheath cells were stained with 3,3′-diaminobenzidine (DAB), and infected sites with cells stained with DAB were counted. Scale bars, 25 μm. h, Rice leaves infected with Δgas2 showed stronger induction of PR genes than leaves infected with P131 or the complementary strain Δgas2/GAS2. RNA samples were collected from rice leaf sheaths inoculated with the indicated strains at 36 hpi. OsActin was used as an internal control. i,j, ROS production in rice leaf sheath cells infected by Δgas2 could be inhibited by diphenyleneiodonium (DPI), an inhibitor of NADPH oxidase. Scale bars, 25 μm. k, The DPI treatment significantly improved the invasive growth of Δgas2 in rice leaf sheath cells. l–n, GAS2-OE lines showed weaker ROS burst (l), weaker induction of PR genes (m), and weaker MAPK activation (n) upon chitin treatment compared with their WT NPBs. Rice tissues were immersed in 0.5 μM chitin for the detection of ROS using the Luminol assay, to measure the expression of PR genes by RT-qPCR, and to perform immunoblotting with an anti-phospho-p44/42 antibody, respectively. OsActin was used as an internal control. o, The leaf sheath cells of GAS2-OE lines infected with M. oryzae strain P131 showed weaker ROS accumulation as compared with WT NPB. At 36 hpi, rice leaf sheath cells were stained with DAB, and infection sites with cells by DAB were counted. Scale bars, 20 μm. Data represent mean ± s.d.; n = biologically independent samples in the graphs. Two-tailed Student’s t tests were employed in (g–k,m,o), *P < 0.05, **P < 0.01. For exact P values, see Source Data. Experiments were repeated at least twice (c–e,n) or three times (a,b,g–m,o) with similar results.
Extended Data Fig. 3 Gas2 from multiple rice fungal pathogens interacts with SnRK1β1A.
a, The alignment of Arabidopsis AKINβ1, rice SnRK1β1A, SnRK1β1B, SnRK1β1C and SnRK1β3. Lines indicate the CBM domain and βCTD domain, and asterisks indicate the conserved lysine residues in rice SnRK1 β subunits that may serve as potential ubiquitination sites. b, Y2H assays showing interaction of Gas2 with the CTD domain of SnRK1β1A. The curve in the N-terminal indicates potential myristoylated glycine (G), the blue and green boxes indicate the CBM and CTD domains, respectively. c, Y2H assays showing interaction of SnRK1β1A with DUF3129 domain of Gas2. The blue box and the orange box indicate the DUF3129 domain and the β-strand of Gas2, respectively. d, LCI assays showing the interaction of SnRK1β1A with Gas2 from M. oryzae. The relative luciferase activity was measured, and protein expression was confirmed by immunoblot analysis with anti-HA and anti-cluc antibodies. e, Y2H assays showing interaction of SnRK1β1A with Gas2 from M. oryzae, but neither with Gas2-like proteins from M. oryzae and B. oryzae nor with unrelated MAS proteins from N. crassa. f, LCI assays showing the interaction of SnRK1β1A with Gas2 from B. oryzae, U. virens, R. solani, F. fujikuroi and F. graminearum, but not with the Gas2-like protein from B. oryzae nor with unrelated MAS proteins from N. crassa. The relative luciferase activity was measured, and protein expression was confirmed by immunoblot analysis with anti-HA and anti-cluc antibodies. Data represent mean ± s.d.; n = biologically independent samples in the graphs. Experiments were repeated at least three times (b–f) with similar results. RLU, relative luminescence units.
Extended Data Fig. 4 SnRK1β1A is up-regulated upon infection by rice major fungal pathogens and negatively regulates rice immunity.
a–f, Expression levels of SnRK1β1A in NPB after infection by M. oryzae (a), F. graminearum (b), F. fujikuroi (c), B. oryzae (d) R. solani (e) and U. virens (f) at the indicated time points. H2O treatment served as a control. g, Schematic representation of the SnRK1β1A gene structure and gene editing sites. Bold letters indicate PAM, “-” indicates nucleotide deletion, and red letter indicates nucleotide insertion. h, Infection assay showing that snrk1β1a mutants are resistant to diverse isolates of M. oryzae, including SZ-4 and SZ-2. Fungal biomass was measured by qPCR at 5 dpi. i, The leaf sheath cells of snrk1β1a lines showed a higher frequency of ROS production than those of WT ZH11 after infection by M. oryzae strain SZ-5. At 36 hpi, rice leaf sheath cells were stained with DAB, and the percentage of infected sites with DAB-stained cells was calculated. Scale bars, 20 μm. j, Higher levels of ROS production were induced in snrk1β1a lines than in WT ZH11 upon chitin treatment. Rice leaf discs were immersed in 0.5 μM chitin to detect ROS using the Luminol assay. k,l, snrk1β1a lines showed stronger MAPK activation and PR gene induction upon chitin treatment than WT ZH11. One-week-old rice seedlings were treated with 0.5 μM chitin for 0–60 min and collected for phosphorylated MAPK detection using an anti-phospho-p44/42 antibody, with anti-Actin as a loading control, and for measuring the expression of PR genes by RT-qPCR after 6 h. Data represent mean ±s.d.; n = biologically independent samples in the graphs. Two-tailed Student’s t tests were employed in (h,l), **P < 0.01. For exact P values, see Source Data. All experiments were repeated at least three times with similar results. RLU, relative luminescence units.
Extended Data Fig. 5 snrk1β1a mutants show enhanced resistance in rice fields.
a, Identification of transgene-free snrk1β1a mutants. Primers specific to the Cas9, HPT and Actin genes, respectively, were used in genotyping. The Actin gene was used as the DNA quality control. b, snrk1β1a lines showed significantly reduced leaf blast disease indices than their WT ZH11 in rice fields of Shangzhuang (SZ), Panjin (PJ), Donggang (DG) and Enshi (ES). c–e, snrk1β1a lines were resistant to panicle blast, sheath blight and false smut disease in the DG nursery. The white triangle in (d) indicates the symptoms of sheath blight disease and the black triangle in (e) indicates the symptoms of false smut ball. (b–e) were from natural infection with multiple duplicates. Data represent mean ± s.d.; n = biologically independent samples in the graphs. Two-tailed Student’s t tests were employed in (b–e), **P < 0.01. For exact P values, see Source Data. Experiments were repeated three times in (b–e) with similar results.
Extended Data Fig. 6 snrk1β1a are similar to its WT in major agronomic traits except early heading and hypersensitivity to high-salinity stress.
The plant morphology (a) and panicle morphology (b) of ZH11 and snrk1β1a in SZ, PJ, and DG fields. The main agronomic traits, including plant height (c), effective tiller number (d), 1,000 grains weight (e) and grain weight per hill (f) were investigated. g, snrk1β1a mutants produce less diseased grains and have higher single-hill yield in PJ and DG disease nurseries. h, snrk1β1a shows early heading. The rice lines were grown in the SZ and DG fields, and the heading time was investigated. i, The hydroponic experiments showed that the snrk1β1a line is more sensitive to 50 mM NaCl compared to the WT ZH11 and SnRK1β1A-OE line. Data represent mean ± s.d.; n = biologically independent samples in the graphs. One-way ANOVA followed by post-hoc Tukey tests were used in (c–f,h); the same letters in (c–f) indicate no significantly different groups, and the letters in (h) indicate significantly different groups. Two-tailed Student’s t tests were employed in (g,i), *P < 0.05, **P < 0.01. For exact P values, see Source Data. All experiments were repeated three times with similar results.
Extended Data Fig. 7 Gas2 prevents 26S proteasome–mediated degradation of SnRK1β1A and drives its nuclear localization.
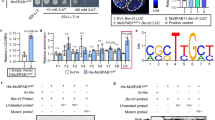
a, SnRK1β1A–3Flag was more accumulated in rice protoplasts co-expressing Gas2–3HA and GFP, but not in those co-expressing GFP. Western blot (WB) was conducted using anti-Flag, anti-HA, and anti-GFP antibodies. Cycloheximide (CHX) was added to the protein extracts and incubated for 4 h to inhibit protein synthesis. b, 2HA–Gas2 reduced the degradation of purified GST-SnRK1β1A by total protein extracts from rice leaves. Proteins were detected within a 0–90 min time course by WB using anti-GST and anti-HA antibodies, with anti-Actin as a loading control. c,d, SnRK1β1A-3HA with Gas2–3Flag or GUS–3Flag (c), SnRK1β1AK248R–3HA, SnRK1β1AK267R–3HA and SnRK1β1AK275R–3HA (d) were transiently expressed in N. benthamiana leaves. The proteins were immunoprecipitated with anti-HA beads to detect their ubiquitination levels by WB using an anti-Ubiquitin antibody. e, Δgas2 knockout the mutants in RB22 isolate were generated and checked by PCR. Genomic DNA isolated from the WT strain RB22 and the Δgas2 mutants was amplified by PCR using specific primers. f, Overexpression of SnRK1β1A in rice complements the defects of ΔUvgas2. The ZH11, snrk1β1a and SnRK1β1A-OE lines were inoculated with the ΔUvgas2 mutant and its WT strain JS60-2. False smut balls (FSB) were counted. One-way ANOVA followed by post-hoc Duncan tests were used, and the letters indicate significantly different groups. g, 3HA–SnRK1β1A signals were detected in the cytoplasm of rice leaf cells without infection but in both the nuclei and the cytoplasm after infection by M. oryzae. The WT M. oryzae strain induced more nuclear localization of 3HA–SnRK1β1A than its Δgas2 mutants. 3HA–SnRK1β1A was detected by the anti-HA antibody. anti-Actin and anti-H3 served as the cytoplasmic and nuclear markers, respectively. h, Electrostatic surface potential of Gas2 predicted by PyMol. Positively charged and negatively charged surfaces of Gas2 are displayed in blue and red, respectively. The positively charged side chains of amino acids are indicated as putative nuclear localization signals (NLSs). i, Mutation strategy of predicted NLSs in Gas2. Gas2mNLS1 and Gas2mNLS2 were obtained by mutating all lysine/arginine to alanine in NLS1 and NLS2, respectively. j, Subcellular localizations of Gas2–GFP and its NLS mutants in N. benthamiana. Green fluorescence was observed at 36 h post-infiltration of the vectors using confocal microscopy. Scale bars, 25 μm. k,l, Y2H assays (k) and LCI assays (l) showed Gas2mNLS1 fails to interact with SnRK1β1A.The relative luciferase activity was measured, and the protein expression was confirmed by immunoblot analysis with anti-HA and anti-cluc antibodies, respectively. Data represent mean ± s.d.; n = biologically independent samples in the graphs. Experiments were repeated at least twice (a–d) or three times (f,g,j–l) with similar results.
Extended Data Fig. 8 SnRK1β1A interacts with SnRK1α1 and inhibits its nuclear localization.
a–d, Interaction of SnRK1β1A with SnRK1α1 detected by Y2H assays (a), LCI assays (b), GST-pulldown assays (c) and BiFC assays (d). The relative luciferase activity was measured and expression of proteins was confirmed by immunoblot analysis with anti-HA and anti-cluc antibodies in (b). Scale bars in (d), 25 μm. e, SnRK1β1A interacts with SnRK1α1 in the cytoplasm under normal conditions. SnRK1α1–3Flag/SnRK1β1A–GFP or SnRK1α1–3Flag/GFP (control) were co-expressed in rice protoplasts. Total proteins were fractionated into cytoplasmic (Cyto) and nuclear (Nuc) compartments and immunoprecipitated with anti-Flag beads, respectively, followed by immunoblotting analysis using anti-HA, anti-GFP, and anti-Flag antibodies. anti-Actin and anti-H3 serving as the cytoplasmic and nuclear markers, respectively. f, Schematic representation of the SnRK1α1 and SnRK1β1A structure and gene editing sites in the snrk1β1asnrk1α1 double mutant. Bold letters indicate PAM, “-” indicates nucleotide deletion, and red letter indicates nucleotide insertion. g, SnRK1β1A inhibited nuclear localization of SnRK1α1 in N. benthamiana cells. SnRK1α1–GFP co-expressed with SnRK1β1A–RFP or free RFP in N. benthamiana. Green fluorescence was observed after 36 h of infiltration using confocal microscopy. Scale bar, 25 μm. h, WB showing reduced distribution of SnRK1α1–3Flag in the nucleus when SnRK1β1A–3Flag was co-expressed in N. benthamiana leaves. An anti-Flag antibody was used to detect SnRK1α1 in the cytoplasmic and nuclear fractionations, with anti-Actin and anti-H3 as the cytoplasmic and nuclear markers, respectively. The nuclear to cytoplasmic ratio (g,h) was determined using ImageJ software. Two-tailed Student’s t tests were employed, *P < 0.05. For exact P values, see Source Data. i, WT M. oryzae strain RB22 was more efficient than its Δgas2 mutant to reduce nuclear distributions of SnRK1α1 in the ZH11 and snrk1β1a but not in SnRK1β1A-OE line. WB was performed using the protein fractions extracted from rice leaves at 24 h post-inoculation (hpi) with M. oryzae strains and Mock treatment (H2O containing 0.025% Tween20), respectively. anti-SnRK1α1 was used for the detection, with anti-Actin and anti-H3 serving as the cytoplasmic and nuclear markers, respectively. Nuclear-to-cytoplasmic ratios were estimated by ImageJ software. j, SnRK1β1A inhibited the kinase activity of SnRK1α1. Both proteins were expressed and purified from Escherichia coli and the AMARA peptide was used as a substrate. Data represent mean ± s.d.; n = biologically independent samples in the graphs. One-way ANOVA followed by post-hoc Tukey tests were conducted; letters denote significantly different groups. Experiments were repeated at least twice in (c–e) or three times in (a,b,g–j) with similar results.
Extended Data Fig. 9 Gas2 is unable to interact with SnRK1α1 but forms a complex with SnRK1β1A and SnRK1α1.
a,b, Y2H (a) and LCI (b) assays showing Gas2 unable to interact with SnRK1α1. c, BiFC assays showing that Gas2 enhanced the interaction of SnRK1β1A with SnRK1α1 in the nucleus. Scale bars, 25 μm. d, The formation of a Gas2-SnRK1β1A-SnRK1α1 tripartite complex with enhanced SnRK1β1A–GFP co-precipitation in GAS2-OE lines. SnRK1α1–3Flag and SnRK1β1A–GFP were co-expressed in rice protoplasts derived from NPB and GAS2-OE lines. Total protein extracts were immunoprecipitated with anti-Flag beads, followed by immunoblotting analysis using anti-HA, anti-GFP, and anti-Flag antibodies. Band intensities were quantified by ImageJ software. Experiments were repeated at least twice (d) or three times (a–c) with similar results.
Extended Data Fig. 10 Transcriptome analysis of ZH11, snrk1β1a, and SnRK1β1A-OE with or without M. oryzae infection.
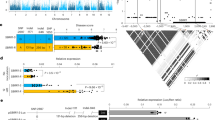
a, Venn diagrams showing differentially expressed genes [DEGs; change folds ≥ 2.0, adjusted P-value (padj) < 0.01)] in snrk1β1a and SnRK1β1A-OE compared to ZH11, with and without M. oryzae infection. Differential gene expression was assessed using the Wald test in DESeq2 (two-sided), with multiple-comparison adjustment using the Benjamini–Hochberg method. Overlapped are SnRK1β1A-suppressed genes (up-regulated in snrk1β1a but down-regulated in SnRK1β1A-OE), and SnRK1β1A-activated genes (down-regulated in snrk1β1a but up-regulated in SnRK1β1A-OE), respectively. b, Hierarchical clustered heatmap of SnRK1β1A-suppressed and -activated genes identified in (a); colours represent fold changes. c, KEGG enrichment analysis of the top-5 overlapping terms shared between all DEGs in (b) and the SnRK1β1A-suppressed genes. Statistical significance was determined using Fisher’s exact test (two-sided) with padj < 0.05 after Benjamini–Hochberg correction. d, Schematic illustration of enrichment of diterpenoid phytoalexins (DPs) biosynthesis genes among the SnRK1β1A-suppressed genes (in red). GGDP, geranylgeranyl diphosphate; CDP, copalyl diphosphate; SDPD, sandaracopimaradiene. DPs biosynthesis genes sourced from modified from Lin et al.53, Zhan et al.54, and Liu et al.55. e, Heat map showing that DPs biosynthetic genes (left) and pathogenicity-related genes (right) are down-regulated by SnRK1β1A. Gene expression levels were presented using the -2 − 2 scale method by normalizing DESeq2’s vst read counts. f, RT-qPCR validated the expression of selected genes shown in (e). g, Higher transcription levels of flowering-related genes HAF1 and HD1 in snrk1β1a than in WT ZH11. Data represent mean ± s.d.; n = biologically independent samples in the graphs. Two-tailed Student’s t tests were employed, *P < 0.05, **P < 0.01. For exact P values, see Source Data. Experiments were repeated at least three times (f,g) with similar results. Schematic in d adapted with permission from ref. 55, Wiley, under a Creative Commons licence CC BY-NC-ND 4.0.
Supplementary information
Supplementary Fig. 1 (download PDF )
Uncropped blots and gel images.
Supplementary Tables 1–6 (download ZIP )
Supplementary Table 1: Gas2 homologs in rice fungal pathogens. Supplementary Table 2: Gas1 homologs in rice fungal pathogens. Supplementary Table 3: Transcription profiles of ZH11, snrk1β1a and SnRK1β1A-OE with or without infection by M. oryzae. Supplementary Table 4: DEGs in the ZH11, snrk1β1a and SnRK1β1A-OE without infection by M. oryzae. Supplementary Table 5: DEGs in the ZH11, snrk1β1a and SnRK1β1A-OE with infection by M. oryzae. Supplementary Table 6: Primers used in this study.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yuan, G., Lu, X., Wang, X. et al. Inactivating SnRK1β1A promotes broad-spectrum disease resistance in rice. Nature (2026). https://doi.org/10.1038/s41586-026-10273-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10273-5