Abstract
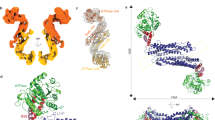
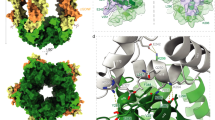
Dynamic assembly of the complex I signalosome mediated by three death domain (DD)-containing proteins—TNFR1, TRADD and RIPK1—is key for transmitting extracellular TNF stimuli to intracellular NF-κB signalling in controlling ‘live or die’ cell fate1. This signalling hub features the rapid recruitment of TRADD and RIPK1 after engagement of TNFR1 by TNF for the formation of complex I, followed by timed disassembly for transition into downstream signalling complexes2,3, but the mechanism driving the dynamic reversibility of complex I remains unclear. Here we captured the assembly core of complex I and determined its cryo-electron microscopy structure, showing a pentameric fibre comprising 31 DDs, with a single layer of a TRADD-DD pentamer sandwiched between multiple layers of TNFR1-DD and RIPK1-DD homopentamers. Structural analysis revealed a strong opposing electric dipole moment (EDM) generated by RIPK1-DD oligomerization relative to that of TNFR1-DD and TRADD-DD. Structure-guided mutagenesis in TNFR1–TRADD–RIPK1 pentameric fibres altering the EDM without affecting DD oligomerization demonstrated the role and mechanism of EDM in driving the dynamic reversibility mediating the rapid assembly and disassembly of complex I. Our study demonstrates a role for long-range interactions mediated by protein EDMs in driving the assembly and disassembly of super-signalling complex I for promoting NF-κB signalling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The atomic coordinates and cryo-EM density maps reported in this study have been deposited in the Protein Data Bank and Electron Microscopy Data Bank under accession codes 9V9C and EMD-64869 (TNFR1-DD filament), 9VGD and EMD-65047 (TRADD-DD filament), 9V9E and EMD-64870 (RIPK1-DD filament) and 9VIN and EMD-65094 (TNFR1-DD–TRADD-DD–RIPK1-DD ternary complex). Source data are provided with this paper.
References
Micheau, O. & Tschopp, J. Induction of TNF receptor I-mediated apoptosis via two sequential signaling complexes. Cell 114, 181–190 (2003).
Linkermann, A. & Green, D. R. Necroptosis. N. Engl. J. Med. 370, 455–465 (2014).
Ofengeim, D. & Yuan, J. Regulation of RIP1 kinase signalling at the crossroads of inflammation and cell death. Nat. Rev. Mol. Cell Biol. 14, 727–736 (2013).
Chen, G. & Goeddel, D. V. TNF-R1 signaling: a beautiful pathway. Science 296, 1634–1635 (2002).
Bradley, J. R. TNF-mediated inflammatory disease. J. Pathol. 214, 149–160 (2008).
Sun, Z. et al. Cooperation of TRADD- and RIPK1-dependent cell death pathways in maintaining intestinal homeostasis. Nat. Commun. 16, 1890 (2025).
Jiang, Y., Woronicz, J. D., Liu, W. & Goeddel, D. V. Prevention of constitutive TNF receptor 1 signaling by silencer of death domains. Science 283, 543–546 (1999).
Hsu, H., Huang, J., Shu, H. B., Baichwal, V. & Goeddel, D. V. TNF-dependent recruitment of the protein kinase RIP to the TNF receptor-1 signaling complex. Immunity 4, 387–396 (1996).
Hsu, H., Xiong, J. & Goeddel, D. V. The TNF receptor 1-associated protein TRADD signals cell death and NF-κB activation. Cell 81, 495–504 (1995).
Dondelinger, Y., Darding, M., Bertrand, M. J. & Walczak, H. Poly-ubiquitination in TNFR1-mediated necroptosis. Cell. Mol. Life Sci. 73, 2165–2176 (2016).
Li, X. et al. Ubiquitination of RIPK1 regulates its activation mediated by TNFR1 and TLRs signaling in distinct manners. Nat. Commun. 11, 6364 (2020).
Wertz, I. E. & Dixit, V. M. Ubiquitin-mediated regulation of TNFR1 signaling. Cytokine Growth Factor Rev. 19, 313–324 (2008).
Iwai, K. LUBAC-mediated linear ubiquitination: a crucial regulator of immune signaling. Proc. Jpn Acad. Ser. B 97, 120–133 (2021).
Mifflin, L., Ofengeim, D. & Yuan, J. Receptor-interacting protein kinase 1 (RIPK1) as a therapeutic target. Nat. Rev. Drug Discov. 19, 553–571 (2020).
Sukits, S. F. et al. Solution structure of the tumor necrosis factor receptor-1 death domain. J. Mol. Biol. 310, 895–906 (2001).
Zhang, N., Yuan, W., Fan, J. S. & Lin, Z. Structure of the C-terminal domain of TRADD reveals a novel fold in the death domain superfamily. Sci. Rep. 7, 7073 (2017).
Ding, J. et al. Structural and functional insights into host death domains inactivation by the bacterial arginine GlcNAcyltransferase effector. Mol. Cell 74, 922–935 (2019).
Hsu, H., Shu, H. B., Pan, M. G. & Goeddel, D. V. TRADD-TRAF2 and TRADD-FADD interactions define two distinct TNF receptor 1 signal transduction pathways. Cell 84, 299–308 (1996).
Vangone, A. & Bonvin, A. M. Contacts-based prediction of binding affinity in protein-protein complexes. eLife 4, e07454 (2015).
Lim, C. S., Lee, J., Kim, J. W. & Lee, J. O. Highly ordered clustering of TNFα and BAFF ligand-receptor-intracellular adaptor complexes on a lipid membrane. Nat. Commun. 16, 5551 (2025).
Zhao, L., Fu, Q., Pan, L., Piai, A. & Chou, J. J. The diversity and similarity of transmembrane trimerization of TNF receptors. Front. Cell Dev. Biol. 8, 569684 (2020).
Sharp, K. A. & Honig, B. Electrostatic interactions in macromolecules: theory and applications. Annu. Rev. Biophys. Biophys. Chem. 19, 301–332 (1990).
Warshel, A. & Papazyan, A. Electrostatic effects in macromolecules: fundamental concepts and practical modeling. Curr. Opin. Struct. Biol. 8, 211–217 (1998).
Lockhart, D. J. & Kim, P. S. Internal stark effect measurement of the electric field at the amino terminus of an alpha helix. Science 257, 947–951 (1992).
Chan, F. K. et al. A domain in TNF receptors that mediates ligand-independent receptor assembly and signaling. Science 288, 2351–2354 (2000).
Karathanasis, C. et al. Single-molecule imaging reveals the oligomeric state of functional TNFα-induced plasma membrane TNFR1 clusters in cells. Sci. Signal. 13, eaax5647 (2020).
Chan, F. K. The pre-ligand binding assembly domain: a potential target of inhibition of tumour necrosis factor receptor function. Ann. Rheum. Dis. 59, i50–i53 (2000).
Naismith, J. H., Devine, T. Q., Brandhuber, B. J. & Sprang, S. R. Crystallographic evidence for dimerization of unliganded tumor necrosis factor receptor. J. Biol. Chem. 270, 13303–13307 (1995).
Kucka, K. & Wajant, H. Receptor oligomerization and its relevance for signaling by receptors of the tumor necrosis factor receptor superfamily. Front. Cell Dev. Biol. 8, 615141 (2020).
Ferrao, R. & Wu, H. Helical assembly in the death domain (DD) superfamily. Curr. Opin. Struct. Biol. 22, 241–247 (2012).
Wang, L. et al. The Fas-FADD death domain complex structure reveals the basis of DISC assembly and disease mutations. Nat. Struct. Mol. Biol. 17, 1324–1329 (2010).
Scott, F. L. et al. The Fas-FADD death domain complex structure unravels signalling by receptor clustering. Nature 457, 1019–1022 (2009).
Lin, S. C., Lo, Y. C. & Wu, H. Helical assembly in the MyD88-IRAK4-IRAK2 complex in TLR/IL-1R signalling. Nature 465, 885–890 (2010).
Park, H. H. et al. Death domain assembly mechanism revealed by crystal structure of the oligomeric PIDDosome core complex. Cell 128, 533–546 (2007).
Bertonati, C., Honig, B. & Alexov, E. Poisson-Boltzmann calculations of nonspecific salt effects on protein-protein binding free energies. Biophys. J. 92, 1891–1899 (2007).
Steeland, S. et al. Generation and characterization of small single domain antibodies inhibiting human tumor necrosis factor receptor 1. J. Biol. Chem. 290, 4022–4037 (2015).
Götzke, H. et al. The ALFA-tag is a highly versatile tool for nanobody-based bioscience applications. Nat. Commun. 10, 4403 (2019).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Felder, C. E., Prilusky, J., Silman, I. & Sussman, J. L. A server and database for dipole moments of proteins. Nucleic Acids Res. 35, W512–W521 (2007).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001).
Dolinsky, T. J., Nielsen, J. E., McCammon, J. A. & Baker, N. A. PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res. 32, W665–W667 (2004).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Heger, K. et al. OTULIN limits cell death and inflammation by deubiquitinating LUBAC. Nature 559, 120–124 (2018).
Lafont, E. et al. TBK1 and IKK ε prevent TNF-induced cell death by RIPK1 phosphorylation. Nat. Cell Biol. 20, 1389–1399 (2018).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Roberts, A., Trapnell, C., Donaghey, J., Rinn, J. L. & Pachter, L. Improving RNA-seq expression estimates by correcting for fragment bias. Genome Biol. 12, R22 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank M. Zhang for his help in electroscope maintenance and training; Y. Ge for her help in fluorescence microscope maintenance and training; and H. Jing and B. Shan for discussion about RNA-seq. Funding: this project was supported (to J.Y.) by the China National Natural Science Foundation (82188101), the Shanghai Municipal Science and Technology Major Project (grant no. 2019SHZDZX02), the Shanghai Science and Technology Development Funds (22JC1410400) and the Shanghai Key Laboratory of Aging Studies (19DZ2260400). J.J.C. was funded by the Science and Technology Commission of Shanghai Municipality (22JC1410500), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB1060000) and the National Natural Science Fund of China-Research Fund for International Scientists (82350710799). J.L. was supported by a grant from the National Natural Science Foundation of China (32071297).
Author information
Authors and Affiliations
Contributions
J. Yuan conceptualized and directed the project. J.L., J.J.C. and J. Yuan designed research. J.L., J.Z. and J.G. purified proteins, performed biochemical and cell biology studies. J.L. solved TRADD-DD filament and ternary DD complex structures with help from Y.H. in data collection. K.Z. solved the cryo-EM structures of RIPK1-DD and TNFR1-DD filaments with samples prepared by J.L.; J. Yang calculated Gibbs free energy and EDM of DDs. F.W., X.L., Zekai Li and B.L. helped in protein expression and purification. S.J. helped in cell biology studies. Zefei Li, J. Ye and Z.S. contributed cell lines. J.L., J.Z., J.G., K.Z., J. Yang, C.L., Y.Z., J.J.C. and J. Yuan analysed data. J.L., J.Z., J.G., J. Yang, J.J.C. and J. Yuan wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Cryo-EM structure determination of TNFR1-DD, TRADD-DD and RIPK1-DD helical filaments.
a, Fluorescence images of HeLa cells expressing mEGFP-tagged TNFR1-DD, TRADD-DD or RIPK1-DD. b, Flow chart for the cryo-EM structure determination of TNFR1-DD, TRADD-DD, RIPK1-DD filaments and the local resolution filtered map calculated with FSC 0.143 criterion. c, The FSC curve calculated between two half maps of the refined final density volume. d, Electrostatic potential surface representation of the cryo-EM structures of TNFR1-DD, TRADD-DD and RIPK1-DD filaments. Results are representative of three independent experiments for (a).
Extended Data Fig. 2 Cryo-EM structure determination of TNFR1-DD, TRADD-DD and RIPK1-DD ternary super complex.
a, Flow chart for the first attempt of cryo-EM structure determination of co-expressed TNFR1-DD/TRADD-DD/RIPK1-DD ternary complex, which failed to yield high resolution density map. b, Flow charts for the purification of co-expressed TNFR1-DD/TRADD-DD/RIPK1-DD complexes without and with K596E RIPK1-DD treatment. c-d, Size-exclusion chromatographs and SDS-PAGE analysis of complexes purified as in b. e, Size-exclusion chromatographs of GB1-tagged WT RIPK1-DD and K596E RIPK1-DD protein, whose molecular weight was determined by size-exclusion chromatography coupled with multi-angle static light scattering. f, Distribution of the length of filaments of TNFR1-DD/TRADD-DD/RIPK1-DD complexes purified as procedure in b. Data are presented as mean ± SEM for 100 filaments in each group. g, Flow chart for the cryo-EM structure determination of co-expressed TNFR1-DD/TRADD-DD/RIPK1-DD ternary complex treated with K596E RIPK1-DD. h, The FSC curve calculated between two half maps of the refined final density volume. Results are representative of three independent experiments for (c-f). The P-value in f is derived from a two-tailed Welch’s t test for comparison between two groups and presented as * (P < 0.05), ** (P < 0.01), *** (P < 0.001), **** (P < 0.0001).
Extended Data Fig. 3 Structural basis for the unique assembly of TNFR1-DD/TRADD-DD/RIPK1-DD ternary complex.
a-e, Cartoon representation of the association between interfaces a and d of TNFR1-DD, TRADD-DD and RIPK1-DD, with interacting residues shown as sticks and interface a shown as electrostatic potential surface. f-j, Cartoon representation of the association between interfaces b and e among TNFR1-DD, TRADD-DD and RIPK1-DD, with interacting residues shown as sticks and interface b shown as electrostatic potential surface. k-o, Cartoon representation of the association between interfaces c and f among TNFR1-DD, TRADD-DD and RIPK1-DD, with interacting residues shown as sticks and interface c shown as electrostatic potential surface. p, Energetically favourable and unfavorable assembly pathways derived based on the calculated Gibbs free energy changes (ΔG) of pairwise interactions among TNFR1-DD, TRADD-DD and RIPK1-DD pentamer layers.
Extended Data Fig. 4 Both inter-layer and intra-layer DD association are critical for Complex I formation and signalling, part A.
a, Fluorescence imaging of TNFR1-KO HeLa cells with individual expression of R376E and E398R TNFR1-DD mutants fused with mEGFP and mCherry tag, respectively. b, Fluorescence imaging of TNFR1-KO HeLa cells with co-expression of the R376E and E398R mutants in (a). c, Heatmap analysis of RNAseq of the expression level of genes induced by TNFα stimulation (100 ng/mL) for 60 min in TNFR1-KO HeLa cells reconstituted with either individual expression of WT, R376E, and E398R TNFR1 or co-expression of R376E and E398R TNFR1 mutants. d, TIRF imaging of individual or co-expression of full-length R376E and E398R TNFR1 mutants fused with fluorescence tag in TNFR1-KO HeLa cells after 5 min of TNFα stimulation (20 ng/mL). mNeonGreen and Halo (TMR) channels are shown in green and red, respectively. e, Quantitation of the puncta density on cell membranes formed by fluorescence-tagged TNFR1 variants in TNFR1-KO HeLa cells in (d). Data are presented as mean ± SEM for 100 areas in each group. f, Fluorescence imaging of RIPK1-KO HeLa cells reconstituted with individual expression of E620R and R621E RIPK1-DD mutants fused with mEGFP and mCherry tag, respectively. g, Fluorescence imaging of RIPK1-KO HeLa cells reconstituted by co-expression of the E620R and R621E RIPK1-DD mutants in (f). h, Heatmap analysis of RNAseq of the expression levels of genes induced by TNFα (100 ng/mL) for 60 min in RIPK1-KO HeLa cells reconstituted with WT or indicated RIPK1 variants. Results are representative of three independent experiments (a, b, d-g) or of three biological replicates (c & h). The P-value in (e) is derived from a two-tailed Welch’s t test for comparison between two groups.
Extended Data Fig. 5 Both inter-layer and intra-layer DD association are critical for Complex I formation and signalling, part B.
a-g, Western blotting analysis of TNFα (100 ng/mL) induced Complex I and NF-κB activation in TNFR1-KO, TRADD-KO or RIPK1-KO HeLa cells reconstituted with indicated mutants of TNFR1, TRADD and RIPK1. The relative levels of the indicated proteins normalized to β-actin are shown below blots (a-g). Results are representative of three independent experiments (a-g).
Extended Data Fig. 6 Modelling of high-order TNFR1/TNFα clustering suggests that the ECD-TMD linker length is barely long enough for a tetramer-of-trimer cluster or 12 TNFR1s.
a, Tilt of ~51° between two trimers of TNFα/TNFR1 complex in the cryo-EM structure (PDB entry 8ZUI) shown with 3-fold axis of each TNFα/TNFR1 trimer (yellow stick). b-c, Modelling of 5 interacting trimers of TNFα/TNFR1 complex on a flat membrane shows that the distance between ECD and TMD in the most peripheral TNFR1 exceeds the length of ECD-TMD linker. Note: The intracellular linker (~120 residues) between the TNFR1 TMD and DD is long enough to support more than 12 TNFR1 DDs that can be added to the TNFR1-TRADD-RIPK1 ternary complex. However, the extracellular linker between the ECD and TMD is only ~10 nm long (aa 184-211, 28 residues). Modelling based on the cryo-EM structure of the tetramer-of-trimer TNFR1/TNFα indicates that expanding the multimer-of-trimer interactions beyond tetramer-of-trimer would result in TNFR1-ECD of the most peripheral trimer rotating away from the TMD by at least 13 nm, which cannot be accommodated by the ECD-TMD linker unless the membrane is severely deformed.
Extended Data Fig. 7 Electric dipole moment modulates the dynamics of Complex I.
a, Three sites (T579, T580 and T583, shown as spheres) on the periphery of RIPK1-DD helical oligomer are selected for mutation to change the EDM of RIPK1-DD without affecting DD-DD interactions. b, Negative-stain EM micrographs of fibres formed in vitro by WT, 2D, 3D or 3 R mutant RIPK1-DD purified from E. coli. c, Representative 2D images of aligned and averaged DD fibres and their Fourier-transformed power spectrum. d, Fluorescence imaging of fibres formed by mEGFP-tagged WT, 2D, 3D or 3 R mutant RIPK1-DD in RIPK1-KO HeLa cells. e-g, Quantification of A20 mRNA level by qRT-PCR in RIPK1-KO HeLa cells reconstituted with WT, 3 R or 3D RIPK1 variants induced by 1 ng/mL (e), 10 ng/mL (f) or 100 ng/mL (g) TNFα for 60 min. Data are presented as mean ± SEM for three biological replicates in each group. h, Heatmap analysis of RNAseq of the expression levels of genes induced by TNFα (10 ng/mL) for 60 min in RIPK1-KO HeLa cells reconstituted with WT, 3 R or 3D RIPK1 variants. i, Snap-shots of TIRF live imaging of RIPK1-KO HeLa cells expressing WT, 3 R and 2D RIPK1 tagged with mNeonGreen upon TNFα stimulation (100 ng/mL). j-l, Distribution of life-time of RIPK1 puncta induced by TNFα in RIPK1-KO HeLa cells expressing WT, 3 R or 2D RIPK1. Data are plotted for 100 puncta in each group. Results are representative of three independent experiments (b, d-g & i) or of three biological replicates (h). The P-value in (e-g) is derived from a two-tailed Welch’s t test for comparison between two groups and presented as * (P < 0.05), ** (P < 0.01), *** (P < 0.001), **** (P < 0.0001).
Extended Data Fig. 8 Intracellular Complex I assembly restricts extracellular TNFα engagement.
a-c, Fluorescence-activated cell sorting (FACS) of AlexaFlour-647 labelled TNFα stimulated wildtype, TRADD-KO and RIPK1-KO HeLa cells. The Q3 fraction represents the TNFα-bound cells. d-e, FACS sorting of AlexaFlour-647 labelled TNFα stimulated RIPK1-KO HeLa cells reconstituted with wildtype or E626R-RIPK1. f-g, Western-blot showing the expression level of TNFR1, TRADD and RIPK1 in these cells. h-i, Co-immunoprecipitation of TNFR1 with FLAG-TNFα in the presence of nanobody Nb70 that competes with TNFα for binding to TNFR1. Results are representative of three independent experiments (a-i).
Extended Data Fig. 9 Structural modelling and electric dipole moment calculation of DR3-TRADD-RIPK1 and MyD88-IRAK4-IRAK2 DD complexes.
a, Modelling of DR3-TRADD-RIPK1 ternary DD complex based on DR3-DD alignment with TNFR1-DD in our cryo-EM structure of the TNFR1-TRADD-RIPK1 DD complex. b-c, Electrostatic potential surface and calculated EDMs of the DR3-TRADD-RIPK1 ternary DD complex. d, The electrostatic potential of the “bottom” face of the DR3-DD pentamer is complementary to the “top” face of the TRADD-DD pentamer. e-f, Cystal structure of the MyD88-IRAK4-IRAK2 complex (PDB ID: 3MOP) and its electrostatic potential surface. g, Calculated EDMs of the MyD88-IRAK4-IRAK2 complex. h, Electrostatic potential of DD layers in MyD88-IRAK4-IRAK2 shows charge complementation.
Extended Data Fig. 10 Structural analysis shows the basis for the specificity of DD adaptors for different receptors.
a-b, Cryo-EM structure of Fas-FADD complex and its electrostatic potential surface. c, Calculated EDMs of the Fas-FADD complex. d-e, Structural modelling of TRAIL-R2-FADD complex and its electrostatic potential surface. f, Calculated EDMs of the TRAIL-R2-FADD complex. g-h, The “bottom” face of the TNFR1-DD pentamer has a negatively charged periphery, which complements the “top” face of the TRADD-DD pentamer. i-j, In the Fas-DD/FADD-DD complex, the “bottom” face of the Fas-DD pentamer shows charge complementation with the “top” face of the FADD-DD pentamer, which is incompatible with the “top” face of the TRADD-DD pentamer. k-l, Structural modelling of the DD pentamer DD from TRAIL-R2 or TRAIL-R1. The “bottom” faces of both receptors are compatible with the “top” face of the FADD-DD pentamer, but incompatible with TRADD-DD.
Supplementary information
Supplementary Video 1 (download MP4 )
Animation of the proposed model of electric dipole moment driving the assembly and timed dissociation of TNF-induced complex I. TNF binding to TNFR1 initiates receptor clustering to form tetramer-of-trimeric complexes and the formation of intracellular DD pentamers. The TNFR1-DD pentamer layers seed the recruitment of TRADD-DDs and RIPK1-DDs, mediated mostly by complementary charge–charge interactions. The cumulative electric dipole moment of RIPK1-DD layers exerts a repulsion between RIPK1-DD and TNFR1-DD–TRADD-DD, which causes the timed release of RIPK1-DD oligomers for activating downstream signalling pathways.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, J., Zhao, J., Gao, J. et al. Electric dipole moment drives the dynamics of the TNFR1 complex I signalosome. Nature (2026). https://doi.org/10.1038/s41586-026-10304-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10304-1