Abstract
Understanding conformational changes of the coronavirus spike protein is critical for developing broad-spectrum therapies. The pan-coronavirus epitope spike residues 815–825 (centred on the S2′ site) are buried in the prefusion spike but are transiently exposed upon ACE2 binding1,2. Here, using integrated functional and structural analyses, we demonstrate that 76E1, an antibody targeting spike residues 815–825, specifically recognizes an open early fusion intermediate conformation in which this epitope adopts a helical conformation, designated the S2′-helix. SARS-CoV-2 Omicron variants evade such antibodies via steric hindrance resulting from S2′-helix shifts and restricted S1–ACE2 distancing in the early fusion intermediate conformation, together with increased reliance on cathepsin-mediated entry that impairs 76E1 inhibition of S2′ cleavage. The H655Y mutation is central to this evasion. Antibody size directly affects its access to the S2′-helix. Crucially, antibody size minimization reversed the evasion mechanisms and significantly enhanced neutralizing activity against authentic Omicron variants and other human coronaviruses, including SARS-CoV-1 and HCoV-229E. These findings establish small-molecule targeting of the S2′-helix as a strategy for pan-coronavirus therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The cryo-EM density maps and atomic models were deposited in the Electron Microscopy Data Bank (EMDB) and the Protein Data Bank (PDB) with accession codes EMD-63002 and 9LDJ (WT SE-FIC–76E1), EMD-63000 and 9LD2 (the local S2–76E1 of the WT SE-FIC–76E1 structure), EMD-62992 (the local S2–76E1-top of the WT SE-FIC–76E1 structure), EMD-65168 and PDB 9VLT (XBB.1.5 SE-FIC–76E1), EMD-65166 and PDB 9VLS (the local S2–76E1 of the XBB.1.5 SE-FIC–76E1 structure), and EMD-65164 (the local S2–76E1-top of the XBB.1.5 SE-FIC–76E1 structure). The plasmid sequences used to generate the antibodies have been deposited at Figshare (https://doi.org/10.6084/m9.figshare.31383049 (ref. 50)). All other data supporting the findings of this study are available within the article and its supplementary files. Source data are provided with this paper.
References
Low, J. S. et al. ACE2-binding exposes the SARS-CoV-2 fusion peptide to broadly neutralizing coronavirus antibodies. Science 377, 735–741 (2022).
Sun, X. et al. Neutralization mechanism of a human antibody with pan-coronavirus reactivity including SARS-CoV-2. Nat. Microbiol. 7, 1063–1074 (2022).
Qu, P. K. et al. Enhanced neutralization resistance of SARS-CoV-2 Omicron subvariants BQ.1, BQ.1.1, BA.4.6, BF.7, and BA.2.75.2. Cell Host Microbe 31, 9–17.e3 (2023).
Wang, Q. et al. Alarming antibody evasion properties of rising SARS-CoV-2 BQ and XBB subvariants. Cell 186, 279–286.e278 (2023).
Chen, Y. J. et al. Broadly neutralizing antibodies to SARS-CoV-2 and other human coronaviruses. Nat. Rev. Immunol. 23, 189–199 (2023).
Ling, Z. Y., Yi, C. Y., Sun, X. Y., Yang, Z. & Sun, B. Broad strategies for neutralizing SARS-CoV-2 and other human coronaviruses with monoclonal antibodies. Sci. China Life Sci. 66, 658–678 (2023).
Dacon, C. et al. Broadly neutralizing antibodies target the coronavirus fusion peptide. Science 377, 728–735 (2022).
Bianchini, F. et al. Human neutralizing antibodies to cold linear epitopes and subdomain 1 of the SARS-CoV-2 spike glycoprotein. Sci. Immunol. 8, eade0958 (2023).
Marcink, T. C. et al. Intermediates in SARS-CoV-2 spike-mediated cell entry. Sci. Adv. 8, eabo3153 (2022).
Song, Y. T. et al. In situ architecture and membrane fusion of SARS-CoV-2 Delta variant. Proc. Natl Acad. Sci. USA 120, e2213332120 (2023).
Grunst, M. W. et al. Structure and inhibition of SARS-CoV-2 spike refolding in membranes. Science 385, 757–765 (2024).
Xing, L. et al. Early fusion intermediate of ACE2-using coronavirus spike acting as an antiviral target. Cell 188, 1297–1314.e1224 (2025).
Jackson, C. B., Farzan, M., Chen, B. & Choe, H. Mechanisms of SARS-CoV-2 entry into cells. Nat. Rev. Mol. Cell Biol. 23, 3–20 (2022).
Meng, B. et al. Altered TMPRSS2 usage by SARS-CoV-2 Omicron impacts infectivity and fusogenicity. Nature 603, 706–714 (2022).
Willett, B. J. et al. SARS-CoV-2 Omicron is an immune escape variant with an altered cell entry pathway. Nat. Microbiol. 7, 1161–1179 (2022).
Hu, B. et al. Spike mutations contributing to the altered entry preference of SARS-CoV-2 omicron BA.1 and BA.2. Emerg. Microbes Infect. 11, 2275–2287 (2022).
Kakizaki, M. et al. The respective roles of TMPRSS2 and cathepsins for SARS-CoV-2 infection in human respiratory organoids. J. Virol. 99, e0185324 (2025).
Furnon, W. et al. Phenotypic evolution of SARS-CoV-2 spike during the COVID-19 pandemic. Nat. Microbiol. 10, 77–93 (2025).
Shi, W. et al. Cryo-EM structure of SARS-CoV-2 postfusion spike in membrane. Nature 619, 403–409 (2023).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280.e278 (2020).
Koch, J. et al. TMPRSS2 expression dictates the entry route used by SARS-CoV-2 to infect host cells. EMBO J. 40, e107821 (2021).
Hu, Y. B., Dammer, E. B., Ren, R. J. & Wang, G. The endosomal–lysosomal system: from acidification and cargo sorting to neurodegeneration. Transl. Neurodegener. 4, 18 (2015).
Fedotov, S., Alexandrov, D., Starodumov, I. & Korabel, N. Stochastic model of virus–endosome fusion and endosomal escape of pH-responsive nanoparticles. Mathematics 10, 375 (2022).
Li, C. et al. Broad neutralization of SARS-CoV-2 variants by an inhalable bispecific single-domain antibody. Cell 185, 1389–1401.e18 (2022).
Addetia, A. et al. Neutralization, effector function and immune imprinting of Omicron variants. Nature 621, 592–601 (2023).
Kimura, I. et al. The SARS-CoV-2 spike S375F mutation characterizes the Omicron BA.1 variant. iScience 25, 105720 (2022).
Pastorio, C. et al. Determinants of Spike infectivity, processing, and neutralization in SARS-CoV-2 Omicron subvariants BA.1 and BA.2. Cell Host Microbe 30, 1255–1268.e5 (2022).
Qu, P. et al. Determinants and mechanisms of the low fusogenicity and high dependence on endosomal entry of omicron subvariants. mBio 14, e0317622 (2023).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, 281–292.e286 (2020).
Modjarrad, K. et al. Preclinical characterization of the Omicron XBB.1.5-adapted BNT162b2 COVID-19 vaccine. npj Vaccines 9, 229 (2024).
Peacock, T. P. et al. The altered entry pathway and antigenic distance of the SARS-CoV-2 Omicron variant map to separate domains of spike protein. Preprint at bioRxiv https://doi.org/10.1101/2021.12.31.474653 (2022).
Yi, C. Y. et al. Key residues of the receptor binding motif in the spike protein of SARS-CoV-2 that interact with ACE2 and neutralizing antibodies. Cell Mol. Immunol. 17, 621–630 (2020).
Yi, C. Y. et al. Comprehensive mapping of binding hot spots of SARS-CoV-2 RBD-specific neutralizing antibodies for tracking immune escape variants. Genome Med. 13, 164 (2021).
Rappazzo, C. G. et al. Broad and potent activity against SARS-like viruses by an engineered human monoclonal antibody. Science 371, 823–829 (2021).
Scheres, S. H. W. RELION: Implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Tan, Y. Z. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Pettersen, E. F. et al. UCSF chimera - a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
He, J. H., Li, T. & Huang, S. Y. Improvement of cryo-EM maps by simultaneous local and non-local deep learning. Nat. Commun. 14, 3217 (2023).
Mannar, D. et al. Altered receptor binding, antibody evasion and retention of T cell recognition by the SARS-CoV-2 XBB.1.5 spike protein. Nat. Commun. 15, 1854 (2024).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of. Acta Crystallogr. D 66, 486–501 (2010).
Afonine, P. V. et al. Real-space refinement in for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Chen, V. B. et al. All-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Bao, Z. Pan-CoV epitope sterically occluded in an open early spike fusion intermediate. Figshare https://doi.org/10.6084/m9.figshare.31383049 (2026).
Acknowledgements
We thank the Center of Cryo-Electron Microscopy, Core Facility of Shanghai Medical College, Fudan University, for the support on cryo-EM data collection. This work was supported by the Shanghai Municipal Science and Technology Major Project ZD2021CY001 to X.S. and Y.X.; the National Natural Science Foundation of China 92469108 to L.S., 92369113 to Y.W. and 32270991 to C.Y.; the National Key R&D Program of China 2025YFA1016500 to X.S., 2021YFC2302500 to L.S., 2024YFA1803100 to Y.X.; the Shanghai Eastern Talent Plan QNWS2024070 to X.S.; and the R&D Program of Guangzhou Laboratory SRPG22-003 to L.S.
Author information
Authors and Affiliations
Contributions
X.S., L.S., Y.W. and Y.X. initiated and supervised the study. X.S. and Z.B. designed and performed most of the functional experiments with assistance from X.W., J.B., H.M. and Y.L. L.S. and Z. Liu performed structural studies with assistance from X.J. and Z.C. Y.W., L.Z. and Z.Z. performed the authentic virus neutralization studies. Y.X., C.Y., Z. Ling and Z.H. provided plasmid and antibody-related materials and valuable suggestions. X.S., L.S., Y.W., Z.B., Z. Liu and Y.X. analysed the data. X.S., L.S. Z. Liu and Z.B. wrote the paper. Y.W. and Y.X. revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The patent application for antibody 76E1 (International Publication No. WO2022148374A1) was filed by the Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences, and includes X.S., C.Y. and Z. Ling as inventors. The patent has been granted in China (CN202110008031.1) and Japan (JP2023541020), and is pending in the European Union (EP22736536) and the United States (US18270917). This patent covers antibody 76E1 and its related sequences. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Shan-Lu Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 SDS-PAGE analysis of S2′-helix-targeting antibodies in various formats and SARS-CoV-2 entry-associated proteases.
a, SDS-PAGE and Coomassie Blue staining analysis of S2′-helix-targeting antibodies (76E1, COV44-62 and C77G12) in three formats, the full-length antibody, Fab and scFv, under reducing conditions. b, SDS-PAGE and Western Blot analysis of cell membrane-bound TMPRSS2 with a C-terminal his tag expressed in 293 T cells under non-reducing conditions. The full-length TMPRSS2 protein comprises the N-terminal intracellular domain, transmembrane domain (TMD), and extracellular region. Within the extracellular domain, TMPRSS2 features the LDLR-A domain, class A SRCR domain, and the C-terminal serine peptidase. The TMPRSS2 precursor undergoes autocleavage at the R255-L256 site, situated between the SRCR domain and the serine peptidase. Despite this cleavage, the two domains remain covalently linked by a disulfide bond, yielding an active protease with an approximate molecular weight of 55 kDa. c, The pro-CTSL was activated in a pH 5.0 buffer containing 400 mM sodium acetate and 4 mM EDTA for 20 min under non-reducing conditions, resulting in the conversion of most pro-CTSL into its activated form, with an approximate molecular weight of 23 kDa. Data are representative of 2 experiments.
Extended Data Fig. 2 Sample purification and cryo-EM workflows for identifying distributions of various species from different datasets.
a, Size exclusion chromatography, SDS-PAGE, and negative stain EM micrographs of wild-type (WT, black) and XBB.1.5 (orange) S. Data are representative of 3 experiments. b-c, Size exclusion chromatography and SDS-PAGE of soluble ACE2 (b) and 76E1-Fab (c). Data are representative of 2 experiments. d-f, Data processing workflows for identifying distributions of various species from the randomly generated sub-datasets from dataset 1 (WT-ACE2-76E1, 30 min) (d), dataset 2 (XBB.1.5-ACE2-76E1, 10 min) (f), and dataset 3 (XBB.1.5-ACE2-76E1, 30 min) (e).
Extended Data Fig. 3 Cryo-EM workflow and map quality analysis of WT S-ACE2-76E1 Fab complex.
a, Cryo-EM data processing workflow (dataset 1). The final cryo-EM maps are colored according to the local resolution estimated by cryoSPARC with the FSC = 0.143 criterion. b, The result of 3D classification indicates that 76E1-Fab is not associated with prefusion S. c, Gold standard FSC curves of the refined 3D reconstructions in WT SE-FIC-76E1. d, The WT SE-FIC-76E1 structure fits into the 6.21 Å map. Three protomers of S and ACE2s are colored as in Fig. 2a, with 76E1-Fabs in orange. e, The S2′-helix in IL770 and variable regions of 76E1-Fab fit into the local S2-76E1 map at a resolution of 3.73 Å, with IL770, HC, and LC in lime, tomato, and orange, respectively. f-i, Representative density in gray surface representation from the local S2-76E1 map.
Extended Data Fig. 4 Cryo-EM workflow and map quality analysis of XBB.1.5 S-ACE2-76E1 Fab complex.
a, Cryo-EM data processing workflow. The final cryo-EM maps are colored according to the local resolution estimated by cryoSPARC with the FSC = 0.143 criterion. b, Gold standard FSC curves of the refined 3D reconstructions in XBB.1.5 SE-FIC-76E1. c, The XBB.1.5 SE-FIC-76E1 structure fits into the 5.65 Å map. d, The S2′-helix in IL770 and variable regions of 76E1-Fab fit into the local S2-76E1 map at a resolution of 4.14 Å. e-h, Representative density in gray surface representation from the local S2-76E1 map. Colors of various components in (c-h) correspond to those in Extended Data Fig. 3d–i, respectively.
Extended Data Fig. 5 Neutralization mechanism of 76E1 revealed by comparing WT SE-FIC-76E1 structure with apo D614G SE-FIC.
a, Schematic of SARS-CoV-2 S2 fragment. Segments excluded from ectodomain expression construct are indicated with dashed boxes. Segments include: S1/S2; S2′ cleavage site; β718-729; 3H, three-helix segment; IL770, intermediate loop 770; FPPR, fusion peptide proximal region; FP, fusion peptide; HR1, heptad repeat 1; CH, central helix; CD, connector domain; HR2, heptad repeat 2; TM, transmembrane anchor; CT, cytoplasmic tail; and inverted triangles for glycans. b, Surface representation of local XBB.1.5 S2-76E1 structure (left) and magnified view of dashed boxes (right), shown as in Fig. 2b,c. c-e, Interfaces between 76E1-Fab and S2′-helix in WT SE-FIC-76E1. Hydrogen bonds (dotted lines) between S2′-helix and HC (c) or LC (d), and hydrophobic interaction (e) are shown. Key residues are indicated as sticks. f, Structural comparison of WT S2′-helix-76E1 structures resolved by cryo-EM (color) and crystallography (PDB 7X9E, gray). g, Overall structure of apo D614G SE-FIC (PDB 8Z7P) shown as in Fig. 2a. h, Structural comparison of S2 in apo D614G SE-FIC and WT SE-FIC-76E1. Magnified view of dashed boxes indicates a slight movement of flexible areas in IL770. i, Collisions occur between 76E1 in WT SE-FIC-76E1 and ACE2 in apo D614G SE-FIC. j, Structural comparison of trimeric S1-ACE2 in apo D614G SE-FIC (surface) and WT SE-FIC-76E1 (ribbon), shown as in Fig. 2h. k, Same as (j), but only S1-ACE2A is shown with a 90° rotation; meanwhile, S1-ACE2A in apo D614G SE-FIC rotates ~8° counterclockwise around the axis. Arrows in arcs show ACE2’s rotations relative to trimeric SD2 centroid, after S1-ACE2 rotating counterclockwise (~8°) and shifting downward (~ 6 Å). Arrows indicate S1-ACE2s’ movements. In (h-k), apo D614G SE-FIC is in gray; WT SE-FIC-76E1 is colored as in Fig. 2a. Structures are aligned by invariant CH-CD.
Extended Data Fig. 6 Additional details of the apo D614G SE-FIC, WT SE-FIC-76E1, and XBB.1.5 SE-FIC-76E1 structures.
a, Surface representation of apo D614G SE-FIC, WT SE-FIC-76E1 and XBB.1.5 SE-FIC-76E1 structures. ACE2, 76E1, and S1 are colored in white, orange, and dark gray, respectively. S2 is in dim gray, of which IL770, FPPR, FP, and HR1 are colored as in Extended Data Fig. 5a, with S2′-helix in red. The distance is defined by the gap between the plane containing the centroid of each S2′-helix and another plane containing the centroid of each SD2. Using trimeric SD2 centroid as the vertex, the angle is defined by the deflection of one ACE2 molecule centroid in relation to the 3-fold axis. b, Left: Top view of the superposition of trimeric S1-ACE2 structures from apo D614G SE-FIC (gray) and WT SE-FIC-76E1 (colored as in Fig. 2a) after D614G-S1-ACE2 rotating (~8°) around the axis and shifting downward (~6 Å). Right: Side view of the superposition of S1-ACE2A. c, Left: Top view of the superposition of trimeric S1-ACE2 structures from WT SE-FIC-76E1 (gray) and XBB.1.5 SE-FIC-76E1 (colored as in Fig. 2a) after WT-S1-ACE2 rotating (~ 4°) around the axis and shifting upward (~3 Å). Right: Side view of the superposition of S1-ACE2A. In (b-c), arrows in arcs show rotations of ACE2A and each domain in S1A, relative to the trimeric SD2 centroid, while SD2 domain remains unchanged. d, Overall structure of the S2 trimer in the postfusion conformation (PDB 8FDW) shown as a ribbon diagram, colored as in Extended Data Fig. 5a. The S2′-helix epitope is indicated by a dashed box colored in lime. e, Clashes between the 76E1-Fab from XBB.1.5 SE-FIC-76E1 and the ACE2 from WT SE-FIC-76E1 can be avoided by reducing the antibody size into scFv.
Extended Data Fig. 7 Comparison of S2′-cleavage efficiency by CTSL and CTSB.
Western blot analysis of dose-dependent S2′ cleavage on cell-surface WT or BF.7 S by CTSL or CTSB, with or without ACE2-Fc priming. Cleavage EC50 values were calculated from band intensity ratios S2′/(S2′ + S2). Results demonstrate markedly stronger cleavage by CTSL compared to CTSB across the concentration range tested. Data are representative of two independent experiments. Uncropped gel images are shown in Supplementary Fig. 1. The samples derive from the same experiment and that gels were processed in parallel.
Extended Data Fig. 8 The acidic environment within endosomes does not significantly contribute to Omicron’s evasion from 76E1-FL.
a, ELISA-based binding curves of 76E1-FL antibody to SECD proteins of WT SARS-CoV-2 and Omicron BF.7 under different low pH conditions. pH 7.4 was included as the control. n = 3. Data are shown as mean ± s.e.m. b, Binding affinity of 76E1-FL to Omicron BF.7 SECD protein as measured by BLI under different low pH conditions. pH 7.4 was included as the control. The curves were fit in red color. c, Exposure of 293T-S cells that bound with the 76E1-ACE2 complex to low pH conditions did not significantly elicit shedding of the 76E1 antibody. 76E1 and ACE2 pre-treated 293T-S cells were subjected to various low pH buffers. Following exposure, cells retaining 76E1 antibody binding after washing were stained with streptavidin-conjugated BV421 and subsequently analyzed via FCM. d, Exposure of ACE2-bound 293T-S cells to low pH conditions does not significantly induce ACE2 shedding. The 293T-S cells were treated with ACE2-biotin protein and subsequently detected using streptavidin-conjugated BV421, followed by analysis via FCM. Relative MFI values of each low pH compared to pH 7.4 was shown. n = 3. Data are shown as mean ± s.e.m. Data representative of two experiments. Schematics in c,d created in BioRender; Bao, Z. https://BioRender.com/vbdqpeh (2026).
Extended Data Fig. 9 Size reduction of 76E1 to scFv reverses antibody evasion caused by H655Y mutation-mediated epitope shielding and viral entry pathway change.
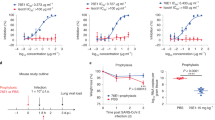
a, Neutralizing curves of 76E1-FL, Fab and scFv at equal molar concentrations against pseudoviruses of SARS-CoV-2 WT, WT-H655Y and reverted Y655H in BF.7 in 293T-ACE2 cells. b-c, Neutralization curves of 76E1-FL (b) and scFv (c) against SARS-CoV-2 WT, WT-H655Y, BF.7 and BF.7-Y655H pseudoviruses in 293T-ACE2 and Caco-2 cells, respectively. a-c, n = 3. Data: mean ± s.e.m; representative of two experiments.
Extended Data Fig. 10 Proposed model for Omicron escape and intermediate spike recognition to S2′-helix-targeting antibodies.
a, SARS-CoV-2 WT: Entry occurs primarily via TMPRSS2-mediated plasma membrane fusion. ACE2 receptor binding induces an early fusion intermediate conformation (E-FIC) of the spike trimer, exposing the S2′-helix epitope. Subsequent downward translation and counterclockwise rotation of S1-ACE2 along the 3-fold axis, followed by outward ACE2 expansion, generates an expanded cavity permitting 76E1-Fab binding. We called this S state as “open E-FIC”. This binding inhibits TMPRSS2-mediated S2′ cleavage and subsequent refolding to the postfusion state. b, Omicron variants: While the TMPRSS2-mediated surface pathway remains important for Omicron entry, its reduced cleavage efficiency at the S2′ site causes an enhanced preference toward CTSL-dependent endocytic entry pathways. Although ACE2 binding exposes the S2′-helix, conformational constraints prevent 76E1-Fab binding at the WT-like E-FIC state of Omicron. Substantial outward expansion of ACE2, which requires surmounting higher conformational energy barriers, is essential for 76E1 engagement. This reduced epitope accessibility, combined with viral entry pathway change of Omicron enables dual evasion mechanisms to 76E1. c, Therapeutic strategy: Antibody size reduction (e.g., to scFv) overcomes both steric constraints and entry pathway-dependent evasion, enabling enhanced neutralization across CoVs. Schematics created in BioRender; Zhao, B. https://BioRender.com/bdoubfb (2026).
Supplementary information
Supplementary Information (download PDF )
This file includes Supplementary Figs. 1–7 and Supplementary Tables 1–6
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bao, Z., Liu, Z., Zhang, Z. et al. Steric hindrance of antibody binding in an Omicron spike fusion intermediate. Nature (2026). https://doi.org/10.1038/s41586-026-10462-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10462-2