Abstract
Sheath blight (ShB), caused by necrotrophic fungus Rhizoctonia solani, is one of the most serious rice diseases worldwide. To the best of our knowledge, no genes with high potential for rice ShB resistance breeding have been previously characterized. Here we identify a ShB resistance receptor-like kinase 1 (SBRR1) gene via a genome-wide association study. The SBRR1-R elite allele, containing a 256-bp insertion in its promoter, is preferentially present in indica varieties in geographical regions with highly favorable conditions for ShB development. Introduction of SBRR1-R into a commercial japonica rice variety significantly reduces yield loss under severe ShB disease pressure. Transcription factor bHLH57 specifically binds to the 256-bp sequence and accounts for highly induced expression and stronger resistance of SBRR1-R. Localization of SBRR1 on plasma membrane, aided by SBRR1-interaction-protein 1, and phosphorylation of SBRR1 are required for SBRR1 to rapidly upregulate downstream chitinase genes for resistance. These findings offer mechanistical insights into ShB resistance hidden in natural rice varieties.
Similar content being viewed by others
Main
The soil-borne necrotrophic fungal pathogen Rhizoctonia solani (R. solani) invades a broad range of plants, including staple food crops, such as rice, maize and potato, posing a great threat to global food security1,2. In rice, it mainly attacks leaf sheath, causing the sheath blight (ShB) disease. ShB is one of the most destructive diseases of rice worldwide and causes an average yield loss of ~10% to 30% each year—even up to 50% under favorable conditions3,4,5. With the widespread cultivation of semi-dwarf and nitrogen-insensitive varieties, rice ShB has increasingly become serious in most rice production regions5,6. Although no completely resistant rice germplasms have been described, a natural variation with clear ShB resistance among rice accessions has been reported7,8. A few varieties, like Tetep and YSBR1, have been repeatedly identified as having relatively high ShB resistance9,10. Also, indica rice was generally found more resistant than japonica rice11. However, resistance mechanisms underlying these varieties remain poorly understood.
Rice ShB resistance is controlled by multiple quantitative trait loci (QTLs) or genes, and more than 60 QTLs for ShB resistance have been detected so far5,12. However, although some of them present breeding potentials, to the best of our knowledge, no ShB resistance QTLs have been isolated by traditional map-based cloning strategy because of the difficulty in precisely phenotyping ShB resistance in a large segregating population5,9,12,13,14,15. Comparatively, the genome-wide association study (GWAS) provides advantages in mining genes, especially for complex quantitative traits in crops16,17, and has been used to identify several genes/alleles with breeding potential in rice17,18,19. Regarding ShB resistance, because the varieties in GWAS are all pure lines, their phenotypes can be evaluated using multiple plants to increase reliability. By using 259 diverse rice varieties, Wang et al.20 characterized two ShB resistance genes, OsRSR1 and OsRLCK5, which improve resistance by regulating reactive oxygen species homeostasis. A stable ShB resistance QTL was identified by GWAS and considered to regulate ShB resistance by suppressing reactive oxygen species accumulation21. Using GWAS, Li et al.22 cloned the ZmFBL41B73 gene that confers resistance to maize ShB and confirmed that its resistance mechanism against ShB via increasing lignin contents in cell walls is conserved in rice. Recently, Gao et al.23 found that disruption of the Ca2+-sensor gene ROD1 confers resistance to several rice diseases, including ShB, but leads to reduced grain yield; notably, they further identified a natural variation, ROD1(SNP1A), that enhances resistance without yield loss. However, ShB resistance genes isolated from natural varieties remain very limited.
Here, we identify an elite allele of the ShB resistance RLK1 (SBRR1) gene from natural varieties through GWAS. We then show great potential for the SBRR1-R allele in rice breeding and decipher its critical upstream regulator and downstream defense signals.
Results
GWAS identifies SBRR1 tightly associated with ShB resistance
To investigate resistance to R. solani in rice, we evaluated ShB resistance of 178 commercial rice cultivars from various regions of China, Japan and South Korea by artificial inoculation in the field (Supplementary Table 1). These cultivars showed disease scores ranging from 4.15 to 7.47, with most of them (78.09%) showing susceptibility and only 1.69% exhibiting moderate resistance (Extended Data Fig. 1a). We conducted a genotyping-by-sequencing analysis and identified a total of 109,444 high-quality single-nucleotide polymorphisms (SNPs) covering the whole rice genome. Structural analysis suggested that these varieties could be divided into two subpopulations, corresponding to indica and japonica subspecies (Extended Data Fig. 1b–e). In order to systematically survey ShB resistance genes, we then performed a GWAS for ShB resistance using the general linear model (GLM) approach controlling population structure and identified 48 SNPs that exceeded the suggestive significance threshold of association (P < 3.2 × 10−5; Fig. 1a and Supplementary Table 2). These significant SNPs correspond to seven regions located on chromosomes 1, 2, 4, 7, 8 and 11 (Supplementary Table 2).
a, Manhattan plot of the GWAS. The horizontal line depicts the Bonferroni-adjusted significance threshold (P = 3.2 × 10−5). The dotted box indicates the location of the two most significant SNPs. b, SBRR1 (ShB resistance RLK 1) -based association mapping and pairwise LD (linkage disequilibrium) analysis. Triangles denote Indels (insertions/deletions) and dots represent SNPs. The lead SNPs/Indels are highlighted in red. The SNPs/Indels showing strong LD with the lead SNPs/Indels are connected to the pairwise LD diagram with solid lines and highlighted with asterisks. c, ShB resistance levels of SBRR1-S and SBRR1-R haplotypes. n denotes the number of genotypes belonging to each haplotype group. The disease score distribution of each haplotype group is displayed as a bar plot. d, Comparison of SBRR1 expression between SBRR1-R and SBRR1-S varieties (n = 20). SBRR1 expression levels were determined for 20 SBRR1-S and 20 SBRR1-R varieties with and without R. solani infection. HPI, hours post inoculation. e, Transient expression assay for promoter activity in rice protoplasts. Left panel: schematic representation of different promoter versions for the reporter construct. Right panel: luciferase reporter assay (n = 6). pSBRR1-S and pSBRR1-R indicate SBRR1 promoter from ZD88 (Zhendao 88) and XWX7 (Xiangwanxian 7) varieties, respectively. Data in c–e are presented as means ± standard deviation (s.d.). Statistical significances in c and d were determined by a two-sided Student’s t-test. Different lowercase letters in e indicate a significant difference (P < 0.05) based on one-way analysis of variance (ANOVA) with Duncan’s multiple range test.
The two most significant (P = 1.3×10−8) SNPs, S94780 and S94782, and three other significant SNPs (S94831, S94832 and S94833) are localized on chromosome 11 in a linkage disequilibrium (LD) block (191 kb) (Fig. 1a, Supplementary Fig. 1a,b, Supplementary Table 1 and Supplementary Note 1). In this block, LOC_Os11g10290, encoding a G-type LecRLK (lectin receptor-like kinase) protein, was identified as the most likely candidate and named ShB Resistance RLK1 (SBRR1) hereafter (Supplementary Fig. 1c,d and Supplementary Note 1). Further, we resequenced the SBRR1 alleles (chromosome 11: 5586341–5592553 bp), including a 3,341-bp promoter region, a 96-bp 5′ untranslated region (UTR), a 2,463-bp coding region, a 143-bp 3′ UTR and a 150-bp downstream sequence from 20 relatively susceptible (disease score >6.5) and 20 relatively resistant (disease score <5.5) varieties then performed a gene-based association mapping (Supplementary Table 3). We identified one variant (SNP1653 or S94780 mentioned above) in the coding region and three variants (SNP-2697 or S94782 mentioned above, indel (insertion/deletion) −1,171 (a 131-bp insertion) and indel-946 (a 256-bp insertion)) in the promoter region that were significantly associated with ShB resistance (Fig. 1b). Based on the four significantly associated variants, two haplotypes of SBRR1 were further identified, and one of them (haplotype ‘A—131 bp—256 bp—T’) showed significantly lower ShB disease scores than the other haplotype (‘G— - — - — G’), which were hereafter named SBRR1-R and SBRR1-S, respectively (Fig. 1c and Supplementary Table 1).
As the three promoter variants may affect expression whereas SNP1653 located in the coding region does not cause an amino acid change, we measured SBRR1 RNA expression levels for the 20 SBRR1-S and 20 SBRR1-R varieties before and after R. solani inoculation. Expression of SBRR1-R was significantly higher than that of SBRR1-S before inoculation, and SBRR1-R was more strongly induced than SBRR1-S, reaching a level 2.8-fold as high as that of SBRR1-S 12 h post-inoculation (HPI) (Fig. 1d and Supplementary Table 4). We further confirmed the differences between SBRR1-S and SBRR1-R promoters by conducting a transient expression assay in rice protoplasts using a luciferase reporter gene (LUC) driven by SBRR1-S promoter (from Zhendao 88, ZD88), SBRR1-R promoter (from Xiangwanxian 7, XWX7) and three SBRR1-R variant promoters containing deletions in the two indel regions of the SBRR1-R promoter. The SBRR1-R promoter more effectively promoted LUC expression than the SBRR1-S promoter by nearly twofold (Fig. 1e). Furthermore, the higher LUC expression level from the SBRR1-R promoter was abolished by deletion of the 256-bp fragment (indel-946), but not indel-1171, indicating the importance of the 256-bp insertion in the higher expression of SBRR1-R. Together, these results suggest that the SBRR1-R allele may be an elite allele conferring ShB resistance.
SBRR1 kinase phosphorylation is required for ShB resistance
To validate the involvement of SBRR1 in rice ShB resistance, we evaluated the phenotypes of a T-DNA insertion mutant sbrr1, SBRR1 knockout lines (sbrr1-ko1/2/3) and overexpression lines (SBRR1-OE1/2/3), and we confirmed that SBRR1 positively regulates ShB resistance (Fig. 2a,b, Extended Data Fig. 2, Supplementary Fig. 2–4 and Supplementary Note 2). As an annotated kinase, we also confirmed that SBRR1 kinase activity is required for its resistance (Fig. 2c,d and Supplementary Note 3). In a Phos-tag SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis) assay using SBRR1-OE plants, we found a clear bandshift for SBRR1 protein representing phosphorylated SBRR1 at 12 HPI with R. solani; this bandshift reached the highest level (approximately fourfold) at 18 HPI compared to 0 HPI in SBRR1-OE plants but was not detected in wild type (WT), indicating that phosphorylation of SBRR1 is induced by pathogen attack (Fig. 2e and Extended Data Fig. 3a–c).
a, ShB disease severity of SBRR1 knockout lines (sbrr1-ko1/2/3, T2 generation) 14 days after R. solani inoculation in a greenhouse. b, ShB resistance of SBRR1 overexpression lines (SBRR1-OE1/2/3, T3 generation). c, Immunoblotting assay using a pSer/Thr antibody to detect autophosphorylation of His-SBRR1-ICD (ICD, intracellular domain) and His-SBRR1-ICDK553E. Ponceau S stains His-SBRR1-ICD and His-SBRR1-ICDK553E proteins. P.S., Ponceau S staining. d, Complementation by WT (Com, T3 generation) and kinase-dead (KiD, T3 generation) constructs for the susceptible phenotype of the sbrr1 mutant (n = 18). e, Comparison of phosphorylation levels of SBRR1 protein in SBRR1-OE1 with or without R. solani infection. Total protein was fractionated in SDS-PAGE (sodium dodecyl sulfate polyacrylamide gel electrophoresis) containing Phos-tag or normal SDS-PAGE. ACTIN was used as a loading reference. f, LC-MS/MS (liquid chromatography-tandem mass spectrometry)-detected intensity ratios of phosphorylated (P-peptide) to un-phosphorylated peptide (UP-peptide) at S678/T682/T683/Y733 sites of SBRR1 protein in SBRR1-OE1 line with or without R. solani infection (n = 3). g, Immunoblot assay using a pSer/Thr antibody to detect autophosphorylation of His-SBRR1-ICDTT682/683AA. P.S., Ponceau S staining. h, ShB resistance of SBRR1TT682/683AA-OE lines (T2 generation) at 7 days after inoculation. Data in a, b, d, f and h are presented as means ± s.d. Statistical significances in a and b were determined by a two-sided Student’s t-test. Different lowercase letters in d and h indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range test. Phosphatase in c, e and g indicates calf intestinal alkaline phosphatase. Scale bar (a, b and d), 5 cm. The in vitro and in vivo phosphorylation assays were done independently three times.
We next used liquid chromatography-tandem mass spectrometry (LC-MS/MS) to detected amino acid residues phosphorylated upon R. solani infection. We found phosphorylation of SBRR1 residues S678, T682, T683, and Y733 in SBRR1-OE plants upon inoculation with R. solani; importantly, no phosphorylation of these residues was detected without R. solani infection. In particular, the phosphorylation levels of T682 and T683 were most notable, showing ratios (counts of phosphorylated peptide/unphosphorylated peptide detected in MS) of 1.8 and 2.8, respectively (Fig. 2f and Supplementary Fig. 5). We noticed that both T682 and T683 are located in the kinase activation loop of SBRR1, and in general, phosphorylation of one to three residues in the activation loop is required for kinase activity24,25. To evaluate the effects of T682 and T683 phosphorylation to SBRR1 kinase activity, we mutated both residues from threonine to alanine (TT682/683AA) and found that the His-SBRR1-ICDTT682/683AA (ICD, intracellular domain) protein completely abolished SBRR1 autophosphorylation (Fig. 2g and Extended Data Fig. 3d), suggesting that the phosphorylation of TT682/683 is essential for SBRR1 kinase activity. To extend these findings in vivo, we generated transgenic rice plants overexpressing SBRR1TT682/683AA under control of the 35S promoter (Extended Data Fig. 3e). We found that SBRR1TT68/2683AA-OE plants did not improve ShB resistance compared with WT, showing significantly lower resistance than SBRR1-OE plants (Fig. 2h). Taken together, these data confirm that the kinase activity of SBRR1 and phosphorylation of TT682/683 are required for SBRR1-mediated ShB resistance.
SBRR1-R is an elite allele mainly present in indica rice
To confirm the variation in the promoter region causes the difference in resistance between SBRR1-R and SBRR1-S, we transformed the sbrr1 mutant with elite promoter (EPro) construct pSBRR1XWX7:SBRR1DJ (SBRR1-R promoter from XWX7 driving SBRR1-S coding region from Dongjin (DJ)). The three EPro lines showed significantly higher inducible SBRR1 transcript levels, reaching levels approximately threefold as high as that of WT DJ after R. solani inoculation, indicating that the SBRR1-R promoter from XWX7 indeed more strongly responded to R. solani infection than the SBRR1-S promoter (Fig. 3a). When assessed for their resistance to R. solani, we found that the three EPro lines all showed lesion lengths (~15.22–15.70 cm) significantly shorter than WT (18.76 cm) and the sbrr1 mutant (23.84 cm) (Fig. 3b). The resistance phenotype in field test is consistent with that in greenhouse and no visible changes in major agronomic traits were found in the EPro transgenic lines compared with WT (Extended Data Fig. 4). Together, these data confirm that the natural variation in the promoter region accounts for stronger resistance of the SBRR1-R elite allele.
a, Expression levels of SBRR1 in EPro lines (pSBRR1XWX7:SBRR1DJ, EPro1/2/3, T3 generation) (n = 3). b, ShB resistance of EPro lines. Data in a and b are presented as means ± s. d. Statistical significance in a was determined by a two-sided Student’s t-test. Different lowercase letters in b indicate a significant difference (P < 0.05) based on one-way ANOVA with Duncan’s multiple range test. Scale bar, 5 cm. c, Geographic distributions of SBBR1-R and SBRR1-S among 2,478 cultivated rice varieties. Orange and green circles indicate SBRR1-R and SBRR1-S, respectively. d, The relative ratio of nucleotide diversity of cultivated rice to common wild rice in the whole rice chromosome 11. The dotted red line indicates the SBRR1 locus. The ratios of nucleotide diversity of japonica/O. rufipogon and indica/O. rufipogon are shown. The ratio in SBRR1 is higher than in flanking regions for japonica/O. rufipogon, whereas the ratio in SBRR1 is notably lower than in flanking regions for indica/O. rufipogon. e, The Tajima’s D analysis in the whole chromosome 11 of cultivated rice. Purple line and dots indicate the SBRR1 locus. The estimated Tajima’s D values for the SBRR1 locus are substantially positive in indica but near zero in japonica. f, Haplotype network of SBRR1 generated using the coding and upstream 1.5-kb sequences of SBRR1 from cultivated rice varieties and common wild rice. Black spots on the lines indicate mutational steps between two haplotypes.
After a comprehensive survey, we found differential presence of SBRR1-R in indica and japonica rice varieties (Supplementary Figs. 6 and 7a,b, Supplementary Tables 5 and 6 and Supplementary Note 4). Geographic distribution analysis showed that the SBRR1-R allele is mainly present in Southeast Asia, East Asia, South Asia and Africa, all of which have favorable temperature and humidity conditions for ShB disease development during rice growing periods (Fig. 3c).
To further probe the molecular evolution of SBRR1-R, we analyzed the relative ratio of nucleotide diversity in indica to wild rice and observed that it is notably lower in the SBRR1 locus than in flanking regions (Fig. 3d and Supplementary Table 7) and that the estimated Tajima’s D values for the SBRR1 locus are substantially positive in indica (Fig. 3e). These results suggest that SBRR1 might have experienced a balancing selection during indica rice domestication. By contrast, the locus of SBRR1 followed a neutral evolution during japonica rice domestication (Fig. 3d,e and Supplementary Table 7). Notably, we found that the SBRR1 allele was diverged in the common wild rice (Supplementary Table 9). Up to 71.43% of O. rufipogon I (Or-I, the progenitor of indica) varieties contained the SBRR1-R allele, whereas all sampled O. rufipogon III (Or-III, the progenitor of japonica) contained the SBRR1-S allele. Both phylogenetic data and haplotype network analysis results using 37 SBRR1 haplotypes revealed that the indica varieties containing SBRR1-R and the japonica varieties containing SBRR1-S had the closest relationship with Or-I and Or-III types of wild rice, respectively (Fig. 3f and Supplementary Fig. 7c), suggesting that SBRR1-R and SBRR1-S alleles are derived from Or-I and Or-III, respectively. Together, we conclude that the SBRR1-R allele originated from wild rice Or-I and then was inherited by most indica varieties located in the regions with more favorable conditions for R. solani infection.
SBRR1-R allele shows great potential for breeding
Because the SBRR1-R allele is almost absent in japonica rice cultivars, which are increasingly spreading toward south China due to the large demand of japonica rice, we tested the breeding potential of SBRR1-R in japonica rice against ShB. Based on the sequence variations in the promoter region between SBRR1-R and SBRR1-S, we developed a diagnostic marker for SBRR1-R (Extended Data Fig. 5a). Via marker assisted selection, we transferred SBRR1-R from YSBR1, a rice variety containing the SBRR1-R allele showing relatively high resistance to ShB10,26, into two widely planted japonica cultivars, Taigeng 394 (TG394) and Xudao 3 (XD3) separately (Extended Data Figs. 2a,b and 5a,b). At the BC5F5 generation, we obtained near-isogenic lines TG394-SBRR1R and XD3-SBRR1R carrying SBRR1-R in the TG394 and XD3 backgrounds, respectively (Extended Data Fig. 5b). TG394-SBRR1R showed significantly higher ShB resistance than WT TG394, with a 24.4% reduction in lesion length (19.3 versus 14.6 cm) in greenhouse assays (Fig. 4a). Similarly, XD3-SBRR1R also displayed significantly enhanced ShB resistance compared with XD3, with a 22.0% reduction in lesion length and no differences in main agronomic traits (Extended Data Fig. 5c,d).
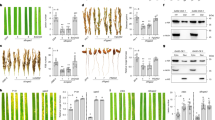
a, ShB lesions of TG394-SBRR1R (TG394, Taigeng 394) and TG394 measured 14 days after R. solani inoculation in greenhouse (n = 27). b, ShB phenotypes of TG394-SBRR1R and TG394 under severe disease conditions in the field. c–l, Comparison of disease score (c, n = 60), grain yield (d, n = 5), seed setting rate (e, n = 11), 1,000-grain weight (f, n = 11), panicle number (g, n = 11), grain number per panicle (h, n = 11), plant height (i, n = 10), growth period (j, n = 10), chalkiness rate (k, n = 10) and amylose content (l, n = 10) of TG394-SBRR1R and TG394 under slight and severe disease conditions in the field. Data in a and c–l are presented as means ± s.d. Statistical significance in a was determined by a two-sided Student’s t-test. Different lowercase letters in c–l indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range test. N.S. indicates no significant differences. Scale bar, 5 cm in a, 5 cm in b and 1 cm in k.
Furthermore, we conducted a field test for TG394 and TG394-SBRR1R under two ShB disease conditions: slight and severe ShB diseases (Fig. 4b–l). We found that under the severe ShB disease condition, TG394-SBRR1R showed an average ShB disease score of 6.13, significantly lower than TG394 (7.22), demonstrating a considerable resistance effect for SBRR1-R in the field trial (Fig. 4b,c). In the severe ShB condition, grain yield and grain quality of TG394 and TG394-SBRR1R lines were both significantly reduced, mainly attributed to decreased seed setting rate, 1,000-grain weight, increased chalkiness rate and amylose content (Fig. 4d–i). However, the TG394-SBRR1R line had significantly lower yield loss than TG394 upon severe ShB disease occurrence: TG394 yield was reduced from 1,404.0 g/1.32 m2 to 1,028.5 g/1.32 m2 (26.75% drop) and TG394-SBRR1R yield from 1,354.3 g/1.32 m2 to 1,126.6 g/1.32 m2 (16.82% drop). Comparatively, the SBRR1-R allele rescued 9.54% (98.1 g per 1.32 m2) of the 26.75% grain yield loss of TG394 due to reduced ShB disease severity (Fig. 4d). Further analysis showed that the rescued yield losses were mainly attributed to two yield components: seed setting rate (6.75%) and 1,000-grain weight (2.00%) (Fig. 4e,f). TG394-SBRR1R also displayed significantly lower loss in grain quality than TG394, including lower chalkiness rate (~3% reduction) and amylose content (~0.8% reduction), suggesting that SBRR1-R also contributed to retaining grain quality under severe ShB disease conditions (Fig. 4k,l). Under slight ShB disease conditions, TG394-SBRR1R showed no significant differences from TG394 in disease scores (Fig. 4c), major agronomical traits, grain yield associated traits and grain quality, indicating that SBRR1-R has no adverse impact on rice development (Fig. 4d–l and Extended Data Fig. 5d). Meanwhile, we also conducted a similar field trial using SBRR1-OE1 and WT plants and, in agreement, found that overexpressing SBRR1 rescued 13.2% of yield losses under severe disease conditions (Extended Data Fig. 6). Together, these results suggest that SBRR1-R carries great potential for rice breeding against ShB.
bHLH57 differentiates SBRR1-R and SBRR1-S expression
To investigate how the 256-bp insertion in the SBRR1-R promoter affects its higher ShB resistance (Fig. 1e), we performed a yeast one-hybrid (Y1H) screening assay using this insertion segment and identified transcription factor bHLH57 (basic helix-loop helix 57, LOC_Os07g35870) (Fig. 5a). By dividing the 256-bp region into two fragments for the Y1H assay, we further narrowed down the binding site of bHLH57 to a 121 bp region (pSBRR1-M2) that contains a ‘CACCGG’ motif putatively serving as a cis-motif for bHLH57 binding. Moreover, mutating ‘CACCGG’ to ‘AAACGG’ (pSBRR1-mutated-M2) abolished the interaction with bHLH57 (Fig. 5a). Notably, this motif is only present in the 256-bp insertion sequence of SBRR1-R but not in the SBRR1-S promoter or its coding genomic region. Furthermore, in electrophoretic mobility shift assays (EMSAs), we detected a slower migrating band for the 21-bp native probe carrying the CACCGG motif when incubated with a GST-bHLH57 (GST, glutathione S-transferase) fusion protein. Unlabeled WT competitors, but not mutated competitors, at 20-fold greatly reduced the binding to the probe, indicating specific binding to this sequence (Fig. 5b). To further confirm that bHLH57 binds to the SBRR1-R promoter in vivo, we carried out a chromatin-immunoprecipitation-quantitative PCR (ChIP-qPCR) assay using TG394-SBRR1R/bHLH57-OE plants (Supplementary Fig. 8). The results showed that bHLH57 specifically enriched the P2 fragment carrying the CACCGG motif of the SBRR1-R promoter (Fig. 5c). We next tested the effect of bHLH57 on SBRR1-R expression by conducting dual-luciferase assays in N. benthamiana. We found that addition of bHLH57 protein notably increased (by threefold) luciferase activities driven by the SBRR1-R promoter, but not by the SBRR1-S promoter (Fig. 5d), indicating that bHLH57 is able to activate SBRR1-R expression but not SBRR1-S expression. In addition, we found that bHLH57 protein was localized in the nucleus and mainly expressed in leaf sheaths and blades, which is well correlated with R. solani infection sites in rice (Extended Data Fig. 7). Together, these data indicate that bHLH57 activates SBRR1-R expression by directly binding to its 256-bp insertion sequence.
a, Y1H assay to identify potential bHLH57 (basic helix-loop helix 57) binding site. M1 and M2 represent two fragments of the 256-bp insertion. The CACCGG motif is found only in the M2 fragment. b, EMSA to confirm the direct binding of bHLH57 to CACCGG motif. GST-EV, glutathione S-transferase-empty vector. c, ChIP-qPCR assay for the enrichment of the P2 region of the SBRR1-R promoter by transcription factor bHLH57 (n = 3). Enriched values are normalized to the input. An unrelated OsActin1 intron region (ACTIN) was used as a negative control. IgG-immunoprecipitated DNA was used as a control. d, Dual-luciferase reporter assay to study the effects of bHLH57 on SBRR1-R gene expression (n = 6). 35S, cauliflower mosaic virus 35S promoter. GFP, green fluorescent protein. REN, Renilla luciferase. LUC, firefly luciferase. e, RNA expression levels of the SBRR1 gene in TG394, T394-SBRR1R and their bhlh57-ko plants (n = 3). HPI, hours post inoculation. f, ShB lesions of bHLH57 knockout lines (T1 generation) in TG394 and T394-SBRR1R backgrounds scored 14 days after R. solani inoculation in greenhouse (n = 24). g, ShB lesions of bHLH57 overexpression lines (T1 generation) in TG394 and T394-SBRR1R backgrounds scored 14 days after R. solani inoculation in greenhouse (n = 21). Data in c–g are presented as means ± s.d. Statistical significances in d and c were determined by a two-sided Student’s t-test. Different lowercase letters in e–g indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range test. Scale bar, 5 cm in f and g. The Y1H, dual-luciferase reporter and EMSAs were done independently three times.
To determine the contribution of bHLH57 to SBRR1-R-mediated ShB resistance, we used CRISPR/Cas9 to generate bHLH57 knockout lines in TG394 cultivar (carrying the SBRR1-S allele) and its near-isogenic line TG394-SBRR1R (with the SBRR1-R allele) background separately (Supplementary Fig. 9). We found that the RNA level of SBRR1 in TG394-SBRR1R plants was about seven times of that in TG394 at 12 HPI, whereas the level in TG394-SBRR1R/bhlh57-ko plants was comparable with those in TG394 and TG394/bhlh57-ko plants (Fig. 5e), indicating that the strong induction pattern of SBRR1 expression in SBRR1-R varieties depends on bHLH57. In agreement, we found that all TG394/bhlh57-ko and TG394-SBRR1R/bhlh57-ko lines displayed lower ShB resistance than their corresponding controls (Fig. 5f), indicating that bHLH57 positively regulates ShB resistance. Importantly, the fact that TG394-SBRR1R plants showed stronger ShB resistance than TG394 plants, whereas TG394-SBRR1R/bhlh57-ko plants displayed similar susceptible levels as TG394/bhlh57-ko plants, clearly showed that SBRR1-R-mediated ShB resistance requires bHLH57 (Fig. 5f). We also generated bHLH57 overexpression lines in TG394 and TG394-SBRR1R backgrounds (Supplementary Fig. 8) and found that TG394/bHLH57-OE lines displayed enhanced ShB resistance in comparison with TG394, which is consistent with a previous report showing that bHLH57 was involved in ShB resistance27. In addition, we found that TG394-SBRR1R/bHLH57-OE plants showed stronger resistance than both TG394-SBRR1R and TG394/bHLH57-OE plants (Fig. 5g), further confirming that bHLH57 and SBRR1-R depend on each other to further enhance ShB resistance. Collectively, these data indicate that bHLH57 is a positively master regulator in the expression of elite SBRR1-R in response to R. solani infection.
SBRR1 localization aided by SIP1 is crucial for its function
We identified a SBRR1-interacting protein (named SIP1, LOC_Os03g63480) via yeast two-hybrid (Y2H) and confirmed that SIP1 is required for SBRR1-mediated ShB resistance (Fig. 6, Supplementary Figs. 10 and 11 and Supplementary Note 5). To probe how SIP1 affects SBRR1-mediated resistance, we generated SBRR1 and SIP1 proteins truncated at different regions and assessed their interaction in Y2H system. We found that the plasminogen-apple-nematode (PAN) domain in the extracellular juxtamembrane region of SBRR1 and the aspartate-proline (DP)-repeat domain of SIP1 were required for their interaction (Extended Data Fig. 8). Because DP-repeat domain has been reported to affect protein transport from endoplasmic reticulum (ER) to plasma membrane (PM)28, we hypothesized that the subcellular localization of SBRR1 might be influenced by SIP1. To test this hypothesis, we analyzed the subcellular localization of SBRR1 and SIP1 in both WT rice protoplasts and N. benthamiana leaf epidermal cells. We found that the green fluorescence of the SBRR1-GFP (GFP, green fluorescent protein) protein was present on PM, merging well with the red signals of the PM marker, and that SIP1-GFP was partially co-localized with the ER marker and the PM marker (Fig. 7a, Supplementary Fig. 12a and Extended Data Fig. 9). We then observed the localization pattern of SBRR1-GFP in the sip1-ko1 mutant and found that most SBRR1-GFP signals (~81%) were localized to ER, with only ~19% localized on PM, indicating that the sip1-ko1 mutant severely affected the PM localization of SBRR1 (Fig. 7b,c and Supplementary Fig. 12b). We further generated an SBRR1 mutant (SBRR1ΔPAN) that lacks the PAN domain and found that almost all SBRR1ΔPAN-GFP signals (~95%) were co-localized with the ER marker and only ~5% co-localized with the PM marker in WT rice protoplasts (Fig. 7d,e and Supplementary Fig. 12c), suggesting that deletion of the PAN domain caused retention of SBRR1 in ER.
a–c, Y2H assay (a), in vitro GST (glutathione S-transferase) pull-down assay (b), and in vivo co-immunoprecipitation (Co-IP) assay using SBRR1-OE1 transgenic plants (c) for SBRR1 and SIP1 (SBRR1-interacting protein 1) interaction. ECD, extracellular domain. ICD, intracellular domain. BD, binding domain; AD, activity domain. The schematic diagram on the right side in a shows the different domains of SBRR1 and SIP1 proteins. Lower panel in b shows the protein abundance stained by CBB (Coomassie brilliant blue) in SDS-PAGE gel. FLAG, transgenic plants expressing a FLAG tag alone, and anti-PIP1;1 (plasma membrane intrinsic protein 1;1) were used as negative controls. The GST pull-down assay and Co-IP assay were performed three times independently with similar results. d, ShB lesions of SIP1 knockout and overexpression lines (T3 generation) scored 14 days after R. solani inoculation in greenhouse. e, ShB lesions of sbrr1-ko1/sip1-ko1, sbrr1-ko1/SIP1-OE1 and SBRR1-OE1/sip1-ko1 lines 14 days after R. solani inoculation in greenhouse. Data in d and e are presented as means ± s.d. Different lowercase letters in d and e indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range test. Scale bar, 5 cm in d and e. The Y2H, pull-down and Co-IP assays were done independently three times.
a, Subcellular localization of SBRR1 and SIP1 in WT protoplasts. b,c, Subcellular localization of SBRR1-GFP in WT and sip1-ko1 protoplasts, and its partition in PM (plasma membrane) and ER (endoplasmic reticulum). d,e, Subcellular localization of SBRR1ΔPAN-GFP (PAN, plasminogen-apple-nematode) in WT protoplasts and its partition in PM and ER. The images in a, b and d were taken under a laser scanning confocal microscope 12 h after transfection. Scale bar (a, b and d), 10 μm. The ratio of protoplasts with only PM-localized GFP signals to the total protoplasts with GFP signals was measured with 100 fluorescent cells in c and e. The protoplasts without clear PM-localized GFP signals show ER-localized GFP signals. Data in c and e are presented as means ± s.d (n = 4). Different lowercase letters in c indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range test. Statistical significance in e was determined by a two-sided Student’s t-test. f, SBRR1 protein distribution in two-phase partitioning. S, soluble fraction; M, microsomal fraction; U, upper phase of partitioning; L, lower phase of partitioning. The blot was probed with the antibody indicated to the right of each panel. PIP1;1 (plasma membrane intrinsic protein 1;1), plasma membrane marker; HSP82 (heat shock protein 82), cytoplasmic marker. The subcellular localization and two-phase partitioning assays were done independently three times.
In addition, we performed aqueous two-phase partitioning experiments using SBRR1-OE1 and SBRR1-OE1/sip1-ko1 lines and found that SBRR1 protein was all detected in the PM-enriched fraction (U, upper phase of partitioning) in SBRR1-OE1 plants; whereas in SBRR1-OE1/sip1-ko1 plants, less than one third of SBRR1 were detected in the PM-enriched fraction and more than two thirds of SBRR1 were found in the endomembrane-enriched fraction (L, lower phase of partitioning) (Fig. 7f; Supplementary note 6). This further confirmed that sip1-ko indeed influenced the PM localization of SBRR1. The loss of proper SBRR1 localization on PM due to lack of SIP1 may be the reason that SBRR1-OE1/sip1-ko1 showed a ShB resistance level close to WT level (Fig. 6e). Together, these data demonstrate that SIP1 facilitates SBRR1 accumulation on PM where SBRR1 functions.
SBRR1 confers ShB resistance mainly via chitinase genes
To further probe the resistance mechanism of SBRR1 to ShB, we performed an RNA-seq to compare transcript profiles of both sbrr1-ko1 and sip1-ko1 (most SBRR1 protein was retained in ER) plants with those of WT plants before and after inoculation with R. solani. We identified 1,137 and 760 differentially expressed genes (DEGs) responding specifically to R. solani for sbrr1-ko1 and sip1-ko1, respectively (Extended Data Fig. 10a; Supplementary Tables 9,10). Gene ontology (GO) analysis showed that these DEGs belong to many pathways. Interestingly, we noted that multiple chitinase gene-related pathways (chitin catabolic process, chitin binding and chitinase activity) and antioxidative pathways (oxidation-reduction process, oxidoreductase activity) were affected in both sbrr1-ko1 and sip1-ko1 plants (Extended Data Fig. 10a and Supplementary Tables 11 and 12). It has been widely validated that chitinases contribute importantly to ShB resistance by degrading chitin, the most abundant polymer of fungal cell walls9. Therefore, we compared the expression levels of all 19 chitinase (Chit) genes between sbrr1-ko1 and WT and between sip1-ko1 and WT at different timepoints post inoculation. Compared with WT plants, we found that the induced expression levels of Chit3 (Loc_Os04g41680), Chit4 (Loc_Os03g30470), Chit7 (LOC_Os06g51050), Chit12 (Loc_Os08g41100) and Chit17 (Loc_Os05g33130) in sbrr1-ko1 plants, and Chit3, Chit4, and Chit17 in sip1-ko1 plants were significantly lower, and among them Chit3 and Chit4 were changed most significantly. These data suggest that chitinase genes act downstream of SBRR1 in response to R. solani (Fig. 8a; Supplementary Fig. 13).
a, Chitinase (Chit) genes showing the most significant changes in RNA expression levels between WT and sbrr1-ko1 or sip1-ko1 line at different hours post R. solani inoculation (HPI) (n = 3). b, Chitinase activity in SBRR1 and SIP1 knockout and overexpression lines at different HPI (n = 3). c, ShB lesions of WT, SBRR1-OE1 and SBRR1-OE1/chit3chit4-ko lines (T2 generation) 14 days after R. solani inoculation in greenhouse (n = 24). d, ShB lesions of WT, Chit3-OE, sbrr1-ko1, sip1-ko1, sbrr1-ko1/Chit3-OE (T2 generation) and sip1-ko1/Chit3-OE (T2 generation) lines 14 days after R. solani inoculation in greenhouse. e, A model for SBRR1-mediated ShB resistance. In SBRR1-R varieties, transcription factor bHLH57 specifically binds to the ‘CACCGG’ motif in the 256-bp insertion of SBRR1-R promoter, elevating SBRR1 expression, reflected by mRNA. In cytoplasm, SIP1 protein, which likely acts as a molecular chaperone, interacts with the extracellular domain of SBRR1 to facilitate effective accumulation of SBRR1 protein on plasma membrane to function. As a kinase protein, SBRR1 phosphorylation is increased upon R. solani attack and required for ShB resistance. Lastly, quickly elevated expression of Chit genes accounts for the majority of SBRR1-mediated downstream defense against R. solani. In SBRR1-S varieties, due to lack of the 256-bp insertion in promoter, bHLH57 cannot effectively activate SBRR1 expression, and lower SBRR1 mRNA lead to lower accumulation of chitinases and susceptibility. Data in a–d are presented as means ± s.d. Different lowercase letters in b–d indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range test. N.S. indicates no significant differences. Scale bar, 5 cm in c and d.
Further, we measured chitinase activities in the knockout and overexpression lines of SBRR1 and SIP1 at different time points after inoculation. We found that chitinase activities were all significantly reduced (up to ~50% reduction) in sbrr1-ko and sip1-ko lines, but significantly increased in overexpression plants compared with WT plants, at 24 and 48 HPI (Fig. 8b). Since the SBRR1-R allele shows more rapidly induced SBRR1 expression levels than SBRR1-S, we also measured the chitinase activities and found that TG394-SBRR1R carried ~40% higher chitinase activity than TG394 at 48 HPI, but showed no differences in chitinase activity than TG394 without ShB inoculation (Extended Data Fig. 10b).
To confirm the contribution of chitinases to SBRR1-mediated resistance, we simultaneously knocked out Chit3 and Chit4, which showed the most significant changes in their transcript levels, in SBRR1-OE1 background. We found that the knockout plants (SBRR1-OE1/chit3-ko/chit4-ko) were significantly more susceptible to ShB than SBRR1-OE1 plants (~16.4 cm versuss 14.5 cm lesion length) but still moderately more resistant than WT plants (18.7 cm), indicating that the two chitinase genes are required for full function of SBRR1 overexpression (Fig. 8c and Supplementary Fig. 14). We also overexpressed Chit3 in sbrr1-ko1, sip1-ko1 and WT backgrounds and found that sbrr1-ko1/Chit3-OE and sip1-ko1/Chit3-OE plants clearly exhibited stronger resistance than sbrr1-ko1, sip1-ko1 and even WT plants but lower resistance than Chit3-OE plants in WT background (Fig. 8d and Supplementary Fig. 15), suggesting that Chit3-OE can restore ShB resistance in sbrr1-ko1 and sip1-ko1 plants. Together, these results demonstrate that chitinase genes play a critical role in SBRR1-mediated downstream defense response to R. solani infection (Fig. 8e).
Discussion
Currently, available genes with breeding values for ShB resistance are very limited. Here, we identified an elite natural allele, SBRR1-R, which shows a significant effect on ShB resistance. We found that SBRR1-R is most likely inherited from O. rufipogon I and is dominantly distributed in indica rice varieties from the regions with more favorable temperature and humidity conditions for R. solani infection, implying the importance of this allele in combating the pathogen (Fig. 3 and Supplementary Fig. 7). By using a functional marker specific to the 256-bp insertion, we successfully introduced SBRR1-R into two commercial japonica rice varieties and found it significantly increased ShB resistance in both backgrounds. Importantly, under severe ShB conditions, this elite allele was found to rescue up to 9.54% of the total 26.75% yield loss. Certainly, this conclusion still needs more large-scale field tests in more rice varieties with different genetic backgrounds, but it suggests a great potential for SBRR1-R in developing ShB resistant varieties (Fig. 4 and Extended Data Fig. 5).
In summary, we propose a working model for SBRR1-mediated ShB resistance (Fig. 8e). In SBRR1-R varieties, transcription factor bHLH57 acts as a positively master regulator, directly binding to the 256-bp insertion sequence of SBRR1-R promoter to effectively elevate SBRR1-R expression upon R. solani infection. In cytoplasm, SIP1 likely functions as a molecular chaperone to facilitate SBRR1 protein for effective accumulation on PM, where SBRR1 protein goes through R. solani-induced phosphorylation events to activate SBRR1 function. With regard to the downstream mechanism leading to ShB resistance, the rapid activation of Chit3 and Chit4 genes accounts for the majority of SBRR1-mediated resistance. But activation of other genes, such as those involved in antioxidative pathways, may also contribute to the full SBRR1-mediated ShB resistance. However, how SBRR1 is phosphorylated and then transduces the signal into the nucleus to activate the expression of chitinase genes and other genes remains to be further elucidated. We hypothesize that SBRR1 relays the intracellular signaling directly or indirectly to downstream transcription factor(s) that in turn regulate the expression of chitinase and other defense genes to counteract R. solani infection. In SBRR1-S varieties, SBRR1 is expressed at lower levels due to the lack of bHLH57-promoted expression, leading to lower levels of chitinases and other relevant proteins and to ShB susceptibility. Together, our findings not only provide an elite allele for rice ShB resistance breeding but also discover a molecular module mediated by SBRR1-R against necrotrophic pathogen R.solani in natural rice varieties.
Methods
Plant materials
The 178 rice cultivars used for GWAS were selected from different regions of China (148), South Korea (15) and Japan (15) (Supplementary Table 1). The sbrr1 mutant (3D-50196L) that carries a T-DNA insertion located 19 bp downstream of the ATG start codon of SBRR1 was isolated from a collection of T-DNA insertion lines (Extended Data Fig. 2c,d; http://orygenesdb.cirad.fr/)29. WT SBRR1 was from DJ, a japonica variety. The YSBR1 variety carrying high ShB resistance, derived from the progeny of a japonica/indica hybrid developed by pedigree breeding10,26, was the donor of the SBRR1-R allele. Two temperate japonica cultivars Taigeng 394 (TG394) and Xudao 3 (XD3) carrying the SBRR1-S allele were used as recipients of SBRR1-R.
Evaluation of ShB resistance
R. solani isolate RH-9 with strong pathogenicity was used to inoculate rice10. The isolate was first grown on potato dextrose agar medium at 28 °C for 3 days. The fungal blocks (0.7 cm in diameter) were then transferred to potato dextrose broth medium containing wood veneers with a thickness of 0.8 mm and a length of 1.0 cm and grown at 28 °C until the hyphae twined around the wood veneers. The wood veneers colonized with hyphae were used as inoculum.
For inoculation in the field, the inoculum was inserted into the third sheath from the top of a plant at the late tillering stage; three main tillers in each plant were inoculated. Disease index was recorded 30 days after heading with more than 30 sheaths each genotype using a 0–9 disease scoring system13. For inoculation in greenhouse, plants were grown in plots in a natural environment until the early booting stage, then transferred to a greenhouse with a relative humidity of 75%-85% under long-day conditions (14 h light at 30 °C/10 h darkness at 24 °C) for 2 days before inoculation. Three main tillers in each plant were inoculated using the same method as field inoculation. Lesion lengths were measured 7 or 14 days after inoculation.
GWAS for rice ShB resistance
GWAS was performed with 178 rice varieties using 109,444 high-quality SNPs with a minor allele frequency of ≥2% and a missing rate of <25% (Supplementary Table 1). Two kinds of univariate GWAS models, CMLM and BLINK, were used to evaluate ShB resistance-SNP associations using the Genomic Association and Prediction Integrated Tool (GAPIT)30. The genome-wide significant threshold of GWAS (P = 3.2 × 10−5) was determined by the minimum Bayes factor calculated using the following formula: minimum Bayes factor = −e∗P∗ln(P)31. The Manhattan and QQ plots for GWAS were generated using the R package CMplot32. LD blocks were defined with the Solid Spine method, and the LD heatmap was constructed using the R package “LDBlockShow”33.
Luciferase reporter assays
The 3,123 bp promoter regions of SBRR1 were amplified from ZD88 (a SBRR1-S variety) and XWX7 (a SBRR1-R variety) individually. These two fragments and different deleted versions derived from were cloned into the pGreenII 0800-LUC vector; Renilla luciferase was used as an internal control34. These two vectors were used to transfect rice protoplasts via PEG-mediated transformation35. Luciferase activity was measured using the Dual-Luciferase reporter assay system (Promega, E1910). Ratios of LUC to Ren activity were calculated to determine relative promoter activity. Six biological replicates were included. The primers used are listed in Supplementary Table 13.
Nucleotide diversity and evolutionary analyses
From the rice XWX7 reference genome, a DNA sequence containing the 256 bp Indel-946 marker and 2,000 bp upstream and 2,000 bp downstream regions (a total of 4,256 bp) was extracted as the reference sequence, named Sequence1. The second-generation sequencing data of all 430 varieties with Illumina sequencing depth >20× from the 3 K rice population were aligned to Sequence1 using the Bowtie2 software. The results were analyzed using Samtools, and a Perl script was used to count the number of reads successfully covering the 256 bp endpoints in the sequencing data of the 430 varieties. Based on this comprehensive analysis, whether these varieties contain the indel-946 sequence was determined. Simultaneously, the Bowtie2 sequence alignment, Samtools, and Perl script analysis were used to determine the SNP type of the SNPG/A site for the 430 varieties. By comparing the indel-946 marker type and the SNPG/A marker type of the 430 varieties, it was determined whether these two markers are closely linked.
The genomic sequences of 26 wild rice and 2,478 cultivated rice accessions were obtained from OryzaGenome (http://viewer.shigen.info/oryzagenome/)36 and the Rice Functional Genomics and Breeding Database (http://www.rmbreeding.cn/Snp3k)37, respectively. The distributions of SBRR1-R and SBRR1-S in the two rice subspecies were compared using the chi-square test. The geographical information of cultivated varieties was obtained from the Rice Functional Genomics and Breeding Database and marked on map using Cartopy package v0.20.0 (ref. 38) in Python v3.6.0 to observe geographic distribution of SBRR1-R and SBRR1-S. The map layer of precipitation from 1970 to 2000 ad was obtained from WorldClim39. The average FST values in each 400-kb window were estimated at chromosome 11 between indica and japonica subspecies using VCFtools v0.1.16 (ref. 40). The nucleotide diversity (π) and Neutral test (Tajima’s D) of each population were calculated in 400-kb window using VCFtools v0.1.16 (ref. 40). Phylogenetic trees and haplotype networks were constructed for all sites with minor allele frequency ≥0.01 of the coding sequences of SBRR1 and their upstream 1.5 kb. The phylogenetic tree for SBRR1 was inferred using the UPGMA method in MEGA v11 (ref. 41). The haplotype network was calculated using pegas package v1.2 (ref. 42) in R v4.1.2 software. The haplotypes that contained more than 3 rice accessions were displayed using plotting module matplotlib v3.6.0 (ref. 43) in Python v3.6.0.
Field trial for rice grain yield and quality analysis
Rice lines TG394, DJ, TG394-SBRR1R and SBRR1-OE1 were included in field trial for yield evaluation. TG394-SBRR1R, carrying the SBRR1-R elite allele, is a near-isogenic line of TG394. SBRR1-OE1 is a transgenic overexpression line in DJ background. Two ShB disease conditions/treatments (slight disease and severe disease) were tested. The slight disease condition was conducted by spraying thifluzamide (a fungicide against R. solani) at tillering, jointing and booting stages to suppress ShB disease occurrence. The severe disease condition was performed via artificial inoculation with R. solani on six tillers per plant to ensure severe disease development. The inoculation procedure was the same as described above. For TG394 and TG394-SBRR1R rice lines, each treatment consisted of six replications/plots, and each plot was 2.37 m long and 1.38 m wide. For DJ and SBRR1-OE1, each treatment consisted of five replications/plots with the same plot size as above. Each plot comprised 10 rows with 12 plants per row (11.55 cm × 23.76 cm rowing space). To reduce marginal effects, only the six inside rows with middle eight plants per row (48 plants per plot covering around 1.32 m2) were disease-rated and collected for grain yield and quality measurement. Grain yield per plot, seed setting rate, 1,000-grain weight, panicle number, grain number per panicle, plant height, growth period, chalkiness rate and amylose content were measured for all plot samples.
In vitro phosphorylation assay
To compare the autophosphorylation activities of SBRR1-ICD (ICD, intracellular domain), SBRR1-ICDK553E, SBRR1-ICDS678A and SBRR1-ICDTT682/683AA, their coding sequences were individually ligated into the pET28a vector to generate corresponding plasmids. Fusion proteins were expressed in Escherichia coli strain BL21 and purified using Ni-Sepharose beads (GE Healthcare, GE17-5268-01). Phosphorylation assays were performed by adding beads containing purified fusion proteins in 60 μl kinase buffer (10 mM ATP, 20 mM MgCl2, 20 mM MnCl2, 40 mM HEPES (pH 7.5), 2 mM DTT, phosphatase inhibitor cocktail and proteinase inhibitor cocktail) and incubating for 90 min at 30 °C. The samples were subsequently separated on SDS-PAGE gels and transferred to nitrocellulose membranes, followed by immunoblot analysis with a pSer/Thr antibody (ECM Biosciences, PP2551; 1:1,000 in bovine serum albumin). Horseradish peroxidase (HRP)-conjugated anti-rabbit IgG (EASYBIO, BE0101; 1:3,000 in bovine serum albumin) was used as the secondary antibody. Primer sequences used are listed in Supplementary Table 13.
Mycelia-containing cellulose acetate membrane inoculation
To collect large amounts of inoculated rice tissues with similar disease progression, a new inoculation method, named mycelia-containing cellulose acetate membrane inoculation, was developed. Briefly, a fungal block (0.7 cm in diameter) was transferred to a cellulose acetate membrane placed on potato dextrose broth medium and grown at 28 °C until R. solani hyphae spread throughout the entire membrane. The cellulose acetate membrane colonized with hyphae was used to tightly cover 14-day-old SBRR1-OE1 seedlings lying flat on a tray. The roots of the seedlings were wrapped with damp gauze to keep them moist. A blank cellulose acetate membrane without R. solani served as a control. Finally, the entire setup was placed in an incubator with 80%-95% relative humidity under long-day conditions (14 h light at 30 °C/10 h darkness at 28 °C). After inoculation, most plant tissues that were close to the cellulose acetate membrane showed similar disease progression and symptoms, which greatly facilitated the collection of large amounts of inoculated rice tissues at similar disease degree.
In vivo phosphorylation assay
To analyze the phosphorylation status of SBRR1 in vivo, protein samples were extracted from 14-day-old SBRR1-OE1 seedlings at different time points after R. solani inoculation and separated in SDS-PAGE gels with or without 50 μM Phos-tag acrylamide (NARD Institute, AAL-107) and 100 μM MnCl2. The samples were then transferred to NC membranes, followed by immunoblot analysis with an anti-FLAG monoclonal antibody (Sigma-Aldrich, F3165; 1:5,000 in PBS). HRP-conjugated anti-mouse IgG (EASYBIO, BE0102; 1:3,000 in PBS) were used as the secondary antibodies. The ImageJ 1.53c (https://imagej.net/ij/ij/download.html) was used to evaluate the gray value of western blotting.
LC-MS/MS analysis for phosphorylation sites
For identification of SBRR1 phosphorylation sites by LC-MS/MS, anti-FLAG agarose (Sigma-Aldrich, A4596) was used to immunoprecipitate SBRR1-FLAG protein from SBRR1-OE1 plants at 18 HPI with R. solani or mock (inoculation with water). Denatured proteins were separated in 10% SDS-PAGE and followed by trypsin digestion overnight before LC-MS/MS analysis (Beijing Biotech-Pack Scientific). LC-MS/MS analysis was performed using a Q-Exactive mass spectrometer (Thermo Fisher Scientific) with a nanoflow HPLC instrument (Thermo Fisher Scientific, EASY-nLC1200 system). Phosphorylated peptides were identified and quantified based on the extracted ion chromatogram peak area using MaxQuant software (1.6.2.10). The intensity ratios at phosphorylation sites represent the normalized intensity of the phosphorylated peptide divided by the normalized intensity of the same but un-phosphorylated peptide. Three biological repeats were performed.
One-hybrid assays in yeast
The full-length cDNA sequence of transcription factor gene bHLH57 was amplified and fused in frame with the GAL4 activation domain in pGADT7-Rec2 (Clontech), forming pGADT7-bHLH57. Then, the fusion construct was used with the reporter plasmid (containing SBRR1-R promoter fused to HIS2) to co-transform Y187 yeast cells (Clontech). Sequences of the primers are listed in Supplementary Table 13. DNA-protein interaction was determined by the growth of the transformants on the nutrient-deficient medium with 50 mM 3-amino-1,2,4-triazole (3-AT), following the manufacturer’s manual (Clontech).
EMSA
For double-stranded DNA probes, equal amounts of forward and reverse strands, whose 5′-ends were labeled with biotin, were mixed, boiled then slowly cooled to 25 °C. Different amounts of unlabeled probes (20-, 100- and 200-fold over labeled probes) were incubated with biotin-labeled probes at 28 °C in binding buffer for 20 min using an EMSA kit (Thermo Fisher Scientific, 20148 Light Shift Chemiluminescent EMSA). The samples were separated by nondenaturing electrophoresis using 6% native acrylamide gels in 0.5 × Tris-borate-EDTA (TBE) buffer under 110 V for 1.5 h at 4 °C. Then DNA probes were transferred to a nylon membrane (Beyotime, FFN10) in 0.5 × TBE by running at 380 mA for 45 min at 4 °C. The biotin-labeled double-stranded DNA on nylon membrane was detected by chemiluminescent assays according to the manufacturer’s instructions (Thermo Fisher Scientific, 89880 Chemiluminescent Nucleic acid detection module kit).
ChIP-qPCR
Three-week-old seedlings of TG394-SBRR1R/bHLH57-OE1 transgenic rice were fully ground into a powder with liquid nitrogen and suspended in buffer to perform ChIP assay. After protein-DNA cross-linking with formaldehyde, nuclei were isolated and chromatin DNA was broken into 200- to 1,000-bp fragments by sonication. The DNA-protein complexes were immunoprecipitated with an anti-FLAG monoclonal antibody (Sigma-Aldrich, F3165; 1:200 in PBS) and enriched using immunomagnetic beads. The collected complexes were de-crosslinked to release the DNA fragments which were used for subsequent qPCR analysis following purification. FLAG monoclonal antibody conjugated to magnetic beads was used to immunoprecipitate protein-DNA complex. Precipitated DNA fragments were analyzed by qPCR to evaluate the enrichment. OsACTIN was used as a negative control. Primer sequences for this experiment are provided in Supplementary Table 13.
Protein interaction analysis
The Y2H assay was conducted to screen for interacting proteins of SBRR1 and analyze their interaction regions. The different truncated deletions of SBRR1 and SIP1 were cloned into pGBKT7 and pGADT7 vectors, respectively. Yeast transformation and screening process were performed according to the manufacturer’s instructions (Clontech).
For the GST pull-down assay, the SBRR1-ECD (ECD, extracellular domain) and full-length SIP1 coding sequence fragments were inserted into pET28a and pGEX4T-2 vectors, respectively. His- and GST-tagged recombinant proteins were expressed in BL21 cells and purified using Ni-Sepharose beads (GE Healthcare, GE17-5268-01) and glutathione beads (GE Healthcare, GE17-0756-04), respectively. GST-SIP1- or GST-coupled beads were used to capture His-SBRR1-ECD. Proteins were separated on SDS-PAGE gels and immunoblotted with an antibody against GST (ABclonal, AE027; 1:3,000 in PBS) or His (ABclonal, AE028; 1:3,000 in PBS).
For Co-IP assay, total proteins were extracted from leaves of 14-day-old SBRR1-OE1 (Ubi-SBRR1-3FLAG) and Ubi-3FLAG (negative control) transgenic rice plants using protein extraction buffer (50 mM HEPES (pH 7.5), 150 mM KCl, 1 mM DTT, 0.4% Triton-X 100 and proteinase inhibitor cocktail). Anti-FLAG agarose (Sigma-Aldrich, A4596) was used to immunoprecipitate the SBRR1 complex, and the complex was detected with FLAG monoclonal antibody (Sigma-Aldrich, F3165; 1:5,000 in PBS) or SIP1 (Beijing Protein Innovation; 1:500 in PBS). HRP-conjugated anti-mouse IgG (EASYBIO, BE0102; 1:3,000 in PBS) were used as the secondary antibodies. Primers used above are listed in Supplementary Table 13.
Subcellular localization
Full-length SBRR1 cDNA, mutant SBRR1ΔPAN and SIP1 cDNA were individually cloned into the PAN580-GFP vector to fuse with GFP at the N-terminus of GFP. The SBRR1-GFP, SIP1-GFP and SBRR1ΔPAN-GFP constructs were separately used to co-transform rice protoplasts with PM marker (PIP2;1-mCherry)44 or ER marker (mCherry ER-Rkcd3-959)45. Meanwhile, GFP was also fused to the C-terminus of SBRR1 and SIP1 in the pCAMBIA1305.1 vector, then introduced into N. benthamiana leaves. Fluorescence images were observed using a laser scanning confocal microscope (ZEISS LSM 700). All primer sequences used are listed in Supplementary Table 13.
RNA-seq and functional classification of DEGs
RNA-seq was used to compare transcriptomic profiles between WT and sbrr1-ko1 or sip1-ko1 before and after inoculation with R. solani. The screening criteria for DEGs were fold change = 2 and P < 0.05. Gene ontology analysis was performed using DAVID Resources 6.7 (http://david.abcc.ncifcrf.gov/)46.
Determination of chitinase activity
Chitinase activities of leaf sheath samples collected at different times after R. solani inoculation were measured by a colorimetric method47. Briefly, to 0.25 g rice sample was added 2 ml of 0.05 M acetic acid buffer (pH 5.0), and the sample was centrifuged at 16,099g at 4 °C for 15 min. Each reaction mixture of 1.2 ml contained 0.4 ml crude enzyme extract, 0.4 ml of 0.05 M acetic acid buffer (pH 5.0) and 0.4 ml of 1% chitin solution. After incubation at 37 °C for 1 h, the reaction mixture was immediately centrifuged at 2,795g for 30 min at 4 °C. Then 0.2 ml saturated borax solution were added to 0.4 ml supernatant, and the mixture was boiled for 7 min. After cooling, 2 ml glacial acetic acid and 1 ml of 1% P-dimethylaminobenzaldehyde solution were added. After incubation at 37 °C for 15 min, the absorbance value at 585 nm was measured. One enzyme activity unit is defined as the amount of 1 mg N-acetylglucosamine produced from chitin decomposition per gram of tissue per hour.
Inclusion and ethics
Data of this study exclusively come from rice, without involving any animal experiments. Planting of transgenic materials and artificial inoculation are subject to strict regulation. All experimental data are included in the Data availability section.
Statistics and reproducibility
All quantitative data are presented as mean ± s.d. from at least three independent experiments. Sample sizes (n) and P values are shown in each figure and figure legend. IBM SPSS Statistics (v.23) was used for statistical analysis. Statistical differences between two groups were analyzed by two-sided Student’s t-test. Statistical significances between more than two groups were analyzed by ANOVA with Duncan’s multiple range test. Results with a P value < 0.05 were considered statistically significant. Luciferase reporter assays, Y1H assays, phosphorylation assays, EMSAs, Y2H assays, Co-IP, GST pull-down, subcellular localization, confocal microscopy and two-phase partitioning were done independently at least three times with similar results. No statistical method was used to predetermine the sample size, but our sample sizes are similar to those reported in previous publications23,48. No data were excluded from the analyses. Data distribution was assumed to be normal, but this was not formally tested.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are available in the article and its Supplementary Information files. The data of 109,444 high-quality SNPs covering the whole rice genome of 178 commercial rice cultivars used in this study and GWAS statistics are publicly available from figshare repository (https://doi.org/10.6084/m9.figshare.25265239.v2). Genomic DNA sequences for SBRR1 from the cultivars XWX7, DJ, YSBR1 and ZD88 can be found in the National Center for Biotechnology Information (NCBI) GenBank under accessions PV423243, PV423242, PV423244 and PV423245, respectively. The phosphorylation MS data has been submitted to figshare repository (https://doi.org/10.6084/m9.figshare.27939462.v1). The RNA-seq data have been deposited in the NCBI Sequence Read Archive under accession code PRJNA953858. Source data are provided with this paper.
Code availability
All software used in this study is publicly available, as described in Methods and Reporting Summary.
References
Zhen, A. et al. The evolution and pathogenic mechanisms of the rice sheath blight pathogen. Nat. Commun. 4, 1424 (2013).
Okubara, P. A., Dickman, M. B. & Blechl, A. E. Molecular and genetic aspects of controlling the soilborne necrotrophic pathogens Rhizoctonia and Pythium. Plant Sci. 228, 61–70 (2014).
Marchetti, M. A. & Bollich, C. N. Quantification of the relationship between sheath blight severity and yield loss in rice. Plant Dis. 75, 773–775 (1991).
Lee, F. N. & Rush, M. C. Rice sheath blight: a major rice disease. Plant Dis. 67, 829–832 (1983).
Senapati, M. et al. Rhizoctonia solani Kühn pathophysiology: status and prospects of sheath blight disease management in rice. Front. Plant Sci. 3, 88116 (2022).
Singh, P., Mazumdar, P., Harikrishna, J. A. & Babu, S. Sheath blight of rice: a review and identification of priorities for future research. Planta 250, 1387–1407 (2019).
Chen, Z. et al. Identification of new resistance loci against sheath blight disease in rice through genome-wide association study. Rice Sci. 26, 21–31 (2019).
Zhang, F. et al. Genome-wide association analysis of the genetic basis for sheath blight resistance in rice. Rice 12, 93 (2019).
Molla, K. A. et al. Understanding sheath blight resistance in rice: the road behind and the road ahead. Plant Biotechnol. J. 18, 895–915 (2020).
Cao, W. et al. Suppressing chlorophyll degradation by silencing OsNYC3 improves rice resistance to Rhizoctonia solani, the causal agent of sheath blight. Plant Biotechnol. J. 20, 335–349 (2022).
Jia, L. et al. Searching for germplasm resistant to sheath blight from the USDA rice core collection. Crop Sci. 51, 1507–1517 (2011).
Wang, Y. et al. Fine mapping and candidate gene analysis of qSB12YSB, a gene conferring major quantitative resistance to rice sheath blight. Theor. Appl. Genet. 136, 246 (2023).
Zuo, S. et al. Fine mapping of qSB-11LE, the QTL that confers partial resistance to rice sheath blight. Theor. Appl. Genet. 126, 1257–1272 (2013).
Zuo, S. et al. Fine-mapping of qSB-9TQ, a gene conferring major quantitative resistance to rice sheath blight. Mol. Breed. 34, 2191–2203 (2014).
Eizenga, G. C., Jia, M. H., Pinson, S. R., Gasore, E. R. & Prasad, B. Exploring sheath blight quantitative trait loci in a Lemont/O. meridionalis advanced backcross population. Mol. Breed. 35, 1–19 (2015).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967 (2010).
Chen, R. et al. Rice functional genomics: decades’ efforts and roads ahead. Sci. China Life Sci. 65, 33–92 (2022).
Li, W. et al. A natural allele of a transcription factor in rice confers broad-spectrum blast resistance. Cell 170, 114–126 (2017).
Liu, Y. et al. Genomic basis of geographical adaptation to soil nitrogen in rice. Nature 590, 600–605 (2021).
Wang, A. et al. Identification of rice (Oryza sativa L.) genes involved in sheath blight resistance via a genome-wide association study. Plant Biotechnol. J. 19, 1553–1566 (2021).
Oreiro, E. G. et al. Genome-wide associations and transcriptional profiling reveal ROS regulation as one underlying mechanism of sheath blight resistance in rice. Mol. Plant Microbe. 33, 212–222 (2020).
Li, N. et al. Natural variation in ZmFBL41 confers banded leaf and sheath blight resistance in maize. Nat. Genet. 51, 1540–1548 (2019).
Gao, M. et al. Ca2+ sensor-mediated ROS scavenging suppresses rice immunity and is exploited by a fungal effector. Cell 184, 5391–5404 (2021).
Taylor, S. S. et al. Structural framework for the protein kinase family. Annu. Rev. Cell Biol. 8, 429–462 (1992).
Lemmon, M. A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Feng, Z. et al. iTRAQ-based quantitative proteomics analysis of defense responses triggered by the pathogen Rhizoctonia solani infection in rice. J. Integr. Agr. 21, 139–152 (2022).
Liu, J. et al. OsbHLH057 targets the AATCA cis‑element to regulate disease resistance and drought tolerance in rice. Plant Cell Rep. 41, 1285–1299 (2022).
Chen, L. et al. The Hop/Sti1-Hsp90 Chaperone complex facilitates the maturation and transport of a PAMP receptor in rice innate immunity. Cell Host Microbe 7, 185–196 (2010).
Jeon, J. S. et al. T-DNA insertional mutagenesis for functional genomics in rice. Plant J. 22, 561–570 (2000).
Lipka, A. E. et al. GAPIT: genome association and prediction integrated tool. Bioinformatics 28, 2397–2399 (2012).
Goodman, S. N. Of p-values and Bayes: a modest proposal. Epidemiology 12, 295–297 (2001).
Yin, L. et al. rMVP: a memory-efficient, visualization-enhanced, and parallel-accelerated tool for genome-wide association study. Genom. Proteom. Bioinf. 19, 619–628 (2021).
Dong, S. et al. LDBlockShow: a fast and convenient tool for visualizing linkage disequilibrium and haplotype blocks based on variant call format files. Brief. Bioinform. 22, bbaa227 (2021).
Hellens, R. P. et al. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 1, 13 (2005).
Chern, M. et al. A rice transient assay system identifies a novel domain in NRR required for interaction with NH1/OsNPR1 and inhibition of NH1mediated transcriptional activation. Plant Methods 8, 6 (2012).
Ohyanagi, H., Obayashi, T. & Yano, K. Plant and Cell Physiology’s 2016 Online Database Issue. Plant Cell Physiol. 57, 1–3 (2016).
Sun, C. et al. RPAN: rice pan-genome browser for ~3000 rice genomes. Nucl. Acids Res. 45, 597–605 (2017).
Elson, P. et al. SciTools/cartopy: v0.22.0. Zenodo (2023).
Fick, S. E. & Hijmans, R. J. WorldClim 2: new 1-km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 37, 4302–4315 (2017).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Koichiro, T., Glen, S. & Sudhir, K. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Paradis, E. pegas: an R package for population genetics with an integrated-modular approach. Bioinformatics 26, 419–420 (2010).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Lee, H. K. et al. Drought stress-induced Rma1H1, a RING membrane-anchor E3 ubiquitin ligase homolog, regulates aquaporin levels via ubiquitination in transgenic Arabidopsis plants. Plant Cell 21, 622–641 (2009).
Nelson, B. K., Cai, X. & Nebenfuhr, A. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J. 51, 1126–1136 (2007).
Wei, H., Sherman, B. & Lempicki, R. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucl. Acids Res. 37, 1–13 (2009).
Boller, T., Gehri, A., Mauch, F. & Vogeli, U. Chitinase in bean leaves: induction by ethylene, purification, properties, and possible function. Planta 157, 22–31 (1983).
Zhu, M. et al. The ZmCPK39–ZmDi19–ZmPR10 immune module regulates quantitative resistance to multiple foliar diseases in maize. Nat. Genet. 56, 2815–2826 (2024).
Acknowledgments
We thank M. Chern for extensive editing. This study was supported by the project of National Key R&D Program of China (2024YFD1200600 to S.M.Z.), Zhongshan Biological Breeding Laboratory (ZSBBL-KY2023-01 to S.M.Z.), Seed Industry Revitalization Project of Jiangsu Province (JBGS[2021]001 to S.M.Z.), the National Natural Science Foundation of China (31701057 to Z.M.F., U24A20405 to H.X.K. and 32401889 to P.G.), Pinduoduo-China Agricultural University Research Fund (PC2023B02001 to J.L.), the Natural Science Foundation of Jiangsu Province (BK20240497 to P.G.), Open Project of Jiangsu Key Laboratory for Eco-Agricultural Biotechnology around Hongze Lake (HZHLAB2202 to Z.M.F.) and a Project Funded by Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD to S.M.Z.).
Author information
Authors and Affiliations
Contributions
S.M.Z., Z.F.Y. and Z.M.F. conceived the project and designed the experiments. S.M.Z. and Z.F.Y. supervised this project. Z.M.F., P.G., G.D.W., H.X.K., J.H.Z., W.Y.X., R.J.C., R.J., X.L.W., Z.G.W., H.M.Z., J.Q.Z., Y.F.Z., K.M.H., Q.Y.S., Y.T. Z., Y.H.X., X.L., X.J.C., Z.X.C. and C.Y. performed the experiments. Z.M.F., P.G., G.D.W., H.X.K., J.H.Z. and R.J.C. participated in the result analysis. S.-W.Z., J.L. and G.-L.W. participated in project discussions and manuscript revision. Z.M.F., P.G., G.D.W., H.X.K. and S.M.Z. wrote the original draft of the manuscript. All authors reviewed and edited the final draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Gitta Coaker, Zhiyong Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Distribution of disease scores and genetic structure of the 178 tested rice cultivars.
a, Distribution of disease scores in different rice varieties inoculated with R. solani in the field. HR, highly resistant; R, resistant; MR, moderately resistant; MS, moderately susceptible; S, susceptible; HS, highly susceptible. b, Optimal K values calculated by ADMIXTURE. The x-axis shows different values of K. The y-axis indicates cross-validation (CV) error. c, Visualization of population structure when K is 2, 3, or 4. d, Principal components analysis (PCA) scree plot. The x-axis shows the principal components. The numbers on the dots represent the percentage of the explained variance per principal component. The y-axis shows the eigen value. e, PCA scatterplot. PC1 and PC2 refer to the first and second principal components (PC), respectively. Triangles and circles represent indica and japonica varieties, respectively.
Extended Data Fig. 2 Expression of SBRR1 in four SBRR-S or SBRR1-R varieties and ShB resistance of T-DNA insertion mutant sbrr1.
a, Differences in the promoter region of SBRR1 among Dongjin, TG394, XD3 (Xudao 3) and YSBR1. ATG represents the start codon. b, Comparison of SBRR1 expression among the four varieties before and after R. solani inoculation (n = 3). c, T-DNA insertion site in sbrr1. P1, P2, and P3 indicate primers used for verification of the insertion. RB (right border) and LB (left border) indicate the right and left borders of T-DNA, respectively. d, PCR verification of the insertion site. e, Comparison of ShB severity between sbrr1 and WT in the field. Data in b and e are presented as means ± s.d. Statistical significances in b and e were determined by a two-sided Student’s t test. Scale bar, 10 cm.
Extended Data Fig. 3 Phosphorylation levels of SBRR1 at different time points post inoculation with R. solani and SBRR1TT682/683AA protein levels in SBRR1TT682/683AA-OE lines.
a, Comparison of phosphorylation levels of SBRR1 protein in SBRR1-OE1 plants at different time points post R. solani inoculation. Total protein was separated in SDS-PAGE containing Phos-tag or normal SDS-PAGE. b, Immunoblotting assay using a pSer/Thr antibody to detect phosphorylation levels of SBRR1 protein in SBRR1-OE1 line with or without R. solani infection. c, Comparison of phosphorylation levels of SBRR1 protein in SBRR1-OE1 and WT plants with or without R. solani infection. Total protein was separated in SDS-PAGE containing Phos-tag acrylamide or normal SDS-PAGE. ACTIN was used as a loading reference. d, Immunoblotting assay using a pSer/Thr antibody to detect autophosphorylation of His-SBRR1-ICDS678A. e, SBRR1TT682/683AA protein levels in SBRR1TT682/683AA-OE lines. Phosphatase in b and d indicates calf intestinal alkaline phosphatase. ACTIN in a, b, c and e was used as a loading reference. The in vitro and in vivo phosphorylation assays were done independently three times.
Extended Data Fig. 4 ShB disease scores (a) and main agronomic traits (b) of SBRR1 elite promoter (EPro) lines in field tests.
Data are presented as means ± s.d (Disease score, n = 72; plant height, panicle number and grain number per panicle, n = 10; 1000-grain weight, n = 3). Different lowercase letters in a indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range tests. N.S. indicates no significant differences. Scale bar, 10 cm.
Extended Data Fig. 5 MAS scheme for transferring SBRR1-R, and ShB lesions and main agronomic traits of XD3 and XD3-SBRR1R.
a, Indel marker Indel-SBRR1-R specific to the SBRR1-R allele. b, MAS (marker assisted selection) scheme for introducing SBRR1-R from YSBR1 into temperate japonica cultivars TG394 and XD3. TG394-SBRR1R and XD3-SBRR1R represent near isogenic lines harboring SBRR1-R in the background of TG394 and XD3, respectively. c, Comparison of ShB lesion length of XD3 and XD3-SBRR1R measured 14 days after R. solani inoculation in greenhouse (n = 27). Scale bar, 5 cm. d, Comparison of plant height (n = 10), whole growth period (n = 10), panicle number (n = 10), grain number per panicle (n = 10), seed setting rate (n = 10) and 1000-grain weight (n = 3) of XD3 and XD3-SBRR1R. Data in c and d are presented as means ± s.d. Statistical significances in c and d were determined by a two-sided Student’s t test.
Extended Data Fig. 6 Overexpression of SBRR1 significantly reduces grain yield losses caused by ShB under severe disease condition.
a, ShB phenotypes of SBRR1-OE1 and WT under severe disease conditions in the field. Scale bar, 10 cm. b-i, Comparison of disease score (b, n = 60), grain yield (c, n = 5), seed setting rate (d, n = 11), 1000-grain weight (e, n = 11), panicle number (f, n = 11), grain number per panicle (g, n = 11), plant height (h, n = 10), growth period (i, n = 10) of SBRR1-OE1 and WT under slight and severe disease conditions in the field. Data in b-i are presented as means ± s.d. Different lowercase letters in b-e indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range tests. N.S. indicates no significant differences.
Extended Data Fig. 7 Subcellular localization and expression pattern of bHLH57.
a, Subcellular localization of bHLH57 in WT rice protoplasts. The images were taken under a laser scanning confocal microscope 12 hours after transfection. Scale bar, 10 μm. b, RNA levels of bHLH57 in different tissues detected by reverse transcription-quantitative PCR (RT-qPCR). Data are presented as means ± s.d (n = 3). The subcellular localization assays were done independently three times.
Extended Data Fig. 8 The PAN domain of SBRR1 and the DP-repeat domain of SIP1 are required for their interaction in the Y2H assay.
a, Western blot detection for the expression of Myc (c-Myc epitope tag)/HA (hemagglutinin epitope tag) fusion proteins in yeast. These proteins (labeled on each of the figures) were used in Fig. 6a. By using Myc/HA antibodies, each of the Myc/HA fusion proteins was detected essentially as a single major band representing the full-length GFP fusion protein in rice protoplasts, consistent with the predicted protein size. b, Y2H assays for the interaction of truncated SBRR1 and SIP1 proteins. c, Western blot analysis for confirming the expression of Myc/HA fusion proteins in yeast. These proteins (labeled on each of the figures) were used in Extended Data Fig. 8b. By using Myc/HA antibodies, each of the Myc/HA fusion proteins was detected essentially as a single major band representing the full-length GFP fusion protein in rice protoplasts, consistent with the predicted protein size. d, In vitro GST pull-down assay for the interaction between the PAN domain of SBRR1 and DP-repeat domain (DP, aspartate-proline) of SIP1. Lower panel shows the protein abundance stained by CBB (coomassie brilliant blue) in SDS-PAGE gel. The Y2H and pull-down assays were done independently three times.
Extended Data Fig. 9 Subcellular localization of SBRR1-GFP and SIP1-GFP proteins in N. benthamiana leaf epidermal cells.
Plasmolysis was achieved by incubating cells in 5% NaCl. The red arrows indicate positions of shrinking plasma membranes. Scale bar, 50 μm. The subcellular localization assays were done independently three times.
Extended Data Fig. 10 Go analysis of RNA-seq transcript profiles between sbrr1-ko1 or sip1-ko1 and WT, and Chitinase activities in TG394 and TG394-SBRR1R.
a, Comparisons of RNA-seq transcript profiles between sbrr1-ko1 and WT and between sip1-ko1 and WT without or with R. solani inoculation (12 HPI). b, Chitinase activities in TG394 and TG394-SBRR1R without (0 h) or with R. solani inoculation (48 HPI). Data are presented as means ± s.d (n = 3). Different lowercase letters indicate significant differences (P < 0.05) based on one-way ANOVA with Duncan’s multiple range tests.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–6, Supplementary Figs. 1-–5 and Unprocessed western blots for Supplementary Figs. 2, 8, 11 and 12.
Supplementary Tables 1–13 (download XLSX )
Supplementary Table 1.The 178 varieties used in assay for ShB resistance. Supplementary Table 2. SNPs significantly associated with ShB resistance. Supplementary Table 3. SBRR1 sequence variation in 20 relatively susceptible (S) and 20 relatively resistant (R) varieties. Supplementary Table 4. RNA expression levels of SBRR1 in SBRR1-S and SBRR1-R lines post inoculation with R. solani. Supplementary Table 5. Verification of the tight linkage between Indel-946 and SNPG/A. Supplementary Table 6. Information for accessions and SBRR1 alleles in Oryza sativa from the 3 K rice genome resource. Supplementary Table 7. The statistical test for the relative ratio of nucleotide diversity of cultivated rice to common wild rice between the SBRR1 locus and flanking genomic regions. Supplementary Table 8. Information for accessions and SBRR1 alleles in Oryza rufipogon from OryzaGenome. Supplementary Table 9. Genes differentially expressed between sbrr1-ko1 and WT that are specifically responsive to R. solani infection. Supplementary Table 10. Genes differentially expressed between sip1-ko1 and WT that are specifically responsive to R. solani infection. Supplementary Table 11. Gene Ontology (GO) enrichment analysis of the identified differentially expressed genes between sbrr1-ko1 and WT. Supplementary Table 12. Gene Ontology (GO) enrichment analysis of the identified differentially expressed genes between sip1-ko1 and WT. Supplementary Table 13. Primers used in this study.
Supplementary Data 1 (download XLSX )
Statistical source data for Supplementary Figs. 1–4 and 8, 10, 11, 13, 15.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed western blots and gels.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download PDF )
Unprocessed western blots.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download PDF )
Unprocessed western blots and gels.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download PDF )
Unprocessed western blots.
Source Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed gels.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots and gels.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed gels.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download PDF )
Unprocessed western blots and gels.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, Z., Gao, P., Wang, G. et al. Natural variation in SBRR1 shows high potential for sheath blight resistance breeding in rice. Nat Genet 57, 2004–2016 (2025). https://doi.org/10.1038/s41588-025-02281-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02281-4
This article is cited by
-
Characterization of the genetic architecture of adult plant resistance to sharp eyespot in Chinese wheat germplasm
Theoretical and Applied Genetics (2026)
-
Rice breeding against sheath blight is now feasible: a breakthrough discovery of SBRR1-mediated sheath blight resistance from natural rice germplasm
Stress Biology (2025)