Abstract
The mammalian (m)SWI/SNF family of chromatin remodelers govern cell type-specific chromatin accessibility and gene expression and assemble as three distinct complexes: canonical BRG1-associated or BRM-associated factor (cBAF), poly(bromo)-associated BAF (PBAF) and noncanonical BAF (ncBAF). ARID1A and ARID1B are paralog subunits that specifically nucleate the assembly of cBAF complexes and are frequently co-mutated in highly aggressive dedifferentiated or undifferentiated endometrial carcinomas (DDEC/UECs). Here in cellular models and primary human tumors, we find that ARID1A and/or ARID1B (ARID1A/B) deficiency-mediated cBAF loss results in increased ncBAF and PBAF biochemical abundance and chromatin-level functions to maintain the DDEC oncogenic state. Furthermore, treatment with clinical-grade SMARCA4 and/or SMARCA2 ATPase inhibitors markedly attenuates DDEC cell proliferation and tumor growth in vivo and synergizes with carboplatin-based chemotherapy to extend survival. These findings reveal the oncogenic contributions of shifted mSWI/SNF family complex stoichiometry and resulting gene-regulatory dysregulation and suggest therapeutic utility of mSWI/SNF small molecule inhibitors in DDEC/UECs and other cBAF-disrupted cancer types.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All sequencing raw and processed data are available in the Gene Expression Omnibus database under the accession no. GSE268960 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE268960) or in the European Genome–phenome Archive under accession no. EGAS50000001004 (http://www.ega-archive.org/datasets/) All raw proteomics and MS datasets have been deposited to ProteomeXchange via PRIDE database under accession no. PXD058236. Source data are provided with this paper.
Code availability
No new code was generated for this work; all analyses have been performed according to the details described in Methods (‘RNA-seq data analysis’, ‘ATAC–seq data analysis’, ‘ChIP–seq data analysis’, ‘TMT–MS data analysis’ and ‘Statistics and reproducibility’) using available software packages.
References
Valencia, A. M. & Kadoch, C. Chromatin regulatory mechanisms and therapeutic opportunities in cancer. Nat. Cell Biol. 21, 152–161 (2019).
Dawson, M. A. & Kouzarides, T. Cancer epigenetics: from mechanism to therapy. Cell 150, 12–27 (2012).
Valencia, A. M. et al. Landscape of mSWI/SNF chromatin remodeling complex perturbations in neurodevelopmental disorders. Nat. Genet. 55, 1400–1412 (2023).
Kadoch, C. Diverse compositions and functions of chromatin remodeling machines in cancer. Sci. Transl. Med. 11, eaay1018 (2019).
Kadoch, C. & Crabtree, G. R. Mammalian SWI/SNF chromatin remodeling complexes and cancer: mechanistic insights gained from human genomics. Sci. Adv. 1, e1500447 (2015).
Santen, G. W. E. et al. Coffin-Siris syndrome and the BAF complex: genotype-phenotype study in 63 patients. Hum. Mutat. 34, 1519–1528 (2013).
Chen, C.-A. et al. Retrospective analysis of a clinical exome sequencing cohort reveals the mutational spectrum and identifies candidate disease-associated loci for BAFopathies. Genet. Med. 24, 364–373 (2022).
Versteege, I. et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature 394, 203–206 (1998).
Kadoch, C. & Crabtree, G. R. Reversible disruption of mSWI/SNF (BAF) complexes by the SS18-SSX oncogenic fusion in synovial sarcoma. Cell 153, 71–85 (2013).
Santen, G. W. E. et al. Mutations in SWI/SNF chromatin remodeling complex gene ARID1B cause Coffin–Siris syndrome. Nat. Genet. 44, 379–380 (2012).
Tsurusaki, Y. et al. Mutations affecting components of the SWI/SNF complex cause Coffin–Siris syndrome. Nat. Genet. 44, 376–378 (2012).
Ramos, P. et al. Small cell carcinoma of the ovary, hypercalcemic type, displays frequent inactivating germline and somatic mutations in SMARCA4. Nat. Genet. 46, 427–429 (2014).
Pan, J. et al. The ATPase module of mammalian SWI/SNF family complexes mediates subcomplex identity and catalytic activity-independent genomic targeting. Nat. Genet. 51, 618–626 (2019).
Valencia, A. M. et al. Recurrent SMARCB1 mutations reveal a nucleosome acidic patch interaction site that potentiates mSWI/SNF complex chromatin remodeling. Cell 179, 1342–1356.e23 (2019).
Mashtalir, N. et al. Modular organization and assembly of SWI/SNF family chromatin remodeling complexes. Cell 175, 1272–1288.e20 (2018).
Mashtalir, N. et al. A structural model of the endogenous human BAF complex informs disease mechanisms. Cell 183, 802–817.e24 (2020).
Michel, B. C. et al. A non-canonical SWI/SNF complex is a synthetic lethal target in cancers driven by BAF complex perturbation. Nat. Cell Biol. 20, 1410–1420 (2018).
St Pierre, R. et al. SMARCE1 deficiency generates a targetable mSWI/SNF dependency in clear cell meningioma. Nat. Genet. 54, 861–873 (2022).
Wang, X. et al. BRD9 defines a SWI/SNF sub-complex and constitutes a specific vulnerability in malignant rhabdoid tumors. Nat. Commun. 10, 1881 (2019).
Mathur, R. et al. ARID1A loss impairs enhancer-mediated gene regulation and drives colon cancer in mice. Nat. Genet. 49, 296–302 (2017).
Wang, X. et al. SMARCB1-mediated SWI/SNF complex function is essential for enhancer regulation. Nat. Genet. 49, 289–295 (2017).
Tessier-Cloutier, B. et al. SWI/SNF-deficiency defines highly aggressive undifferentiated endometrial carcinoma. Hip Int. 7, 144–153 (2021).
Silva, E. G., Deavers, M. T., Bodurka, D. C. & Malpica, A. Association of low-grade endometrioid carcinoma of the uterus and ovary with undifferentiated carcinoma: a new type of dedifferentiated carcinoma?. Int. J. Gynecol. Pathol. 25, 52–58 (2006).
Santoro, A. et al. Clinico-pathological significance of TCGA classification and SWI/SNF proteins expression in undifferentiated/dedifferentiated endometrial carcinoma: a possible prognostic risk stratification. Gynecol. Oncol. 161, 629–635 (2021).
Coatham, M. et al. Concurrent ARID1A and ARID1B inactivation in endometrial and ovarian dedifferentiated carcinomas. Mod. Pathol. 29, 1586–1593 (2016).
Guan, B. et al. Mutation and loss of expression of ARID1A in uterine low-grade endometrioid carcinoma. Am. J. Surg. Pathol. 35, 625–632 (2011).
Hammer, P. M. et al. Molecular classification outperforms histologic classification in prognostication of high-grade endometrial earcinomas with undifferentiated and sarcomatous components. Am. J. Surg. Pathol. https://doi.org/10.1097/PAS.0000000000002250 (2024).
Travaglino, A. et al. TCGA molecular subgroups in endometrial undifferentiated/dedifferentiated carcinoma. Pathol. Oncol. Res. 26, 1411–1416 (2020).
Zhang, X. & Yu, M. Undifferentiated endometrial carcinoma: a selected immunohistochemical panel including PAX-8 and E-cadherin for aiding distinction from other endometrial carcinomas. Ann. Diagn. Pathol. 39, 36–41 (2019).
Cancer Genome Atlas Research Network et al. Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
Ghandi, M. et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569, 503–508 (2019).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Kelso, T. W. R. et al. Chromatin accessibility underlies synthetic lethality of SWI/SNF subunits in ARID1A-mutant cancers. eLife 6, e30506 (2017).
Papillon, J. P. N. et al. Discovery of orally active inhibitors of Brahma homolog (BRM)/SMARCA2 ATPase activity for the treatment of Brahma related gene 1 (BRG1)/SMARCA4-mutant cancers. J. Med. Chem. 61, 10155–10172 (2018).
Remillard, D. et al. Degradation of the BAF complex factor BRD9 by heterobifunctional ligands. Angew. Chem. Int. Ed. Engl. 56, 5738–5743 (2017).
de Miguel, F. J. et al. Mammalian SWI/SNF chromatin remodeling complexes promote tyrosine kinase inhibitor resistance in EGFR-mutant lung cancer. Cancer Cell 41, 1516–1534.e9 (2023).
Fiskus, W. et al. BRG1/BRM inhibitor targets AML stem cells and exerts superior preclinical efficacy combined with BET or menin inhibitor. Blood 143, 2059–2072 (2024).
Patel, S. P. et al. 1128P A phase I dose escalation and expansion study of FHD-286, a novel BRG1/BRM (SMARCA4/SMARCA2) inhibitor, for the treatment of metastatic uveal melanoma. Ann. Oncol. 34, S677–S678 (2023).
Eskander, R. N. et al. Pembrolizumab plus chemotherapy in advanced endometrial cancer. N. Engl. J. Med. 388, 2159–2170 (2023).
Davis, J. M. et al. Long-term disease-free survival with chemotherapy and pembrolizumab in a patient with unmeasurable, advanced stage dedifferentiated endometrial carcinoma. Gynecol. Oncol. Rep. 53, 101380 (2024).
Wilson, M. R. et al. ARID1A and PI3-kinase pathway mutations in the endometrium drive epithelial transdifferentiation and collective invasion. Nat. Commun. 10, 3554 (2019).
Suryo Rahmanto, Y. et al. Inactivation of Arid1a in the endometrium is associated with endometrioid tumorigenesis through transcriptional reprogramming. Nat. Commun. 11, 2717 (2020).
Guan, B., Wang, T.-L. & Shih, I.-M. ARID1A, a factor that promotes formation of SWI/SNF-mediated chromatin remodeling, is a tumor suppressor in gynecologic cancers. Cancer Res. 71, 6718–6727 (2011).
Reske, J. J. et al. SWI/SNF inactivation in the endometrial epithelium leads to loss of epithelial integrity. Hum. Mol. Genet. 29, 3412–3430 (2020).
Nakayama, R. T. et al. SMARCB1 is required for widespread BAF complex-mediated activation of enhancers and bivalent promoters. Nat. Genet. 49, 1613–1623 (2017).
Wang, Z. et al. Dual ARID1A/ARID1B loss leads to rapid carcinogenesis and disruptive redistribution of BAF complexes. Nat. Cancer 1, 909–922 (2020).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385.e18 (2018).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Zhang, Z. et al. Clinicopathological and prognostic significance of SWI/SNF complex subunits in undifferentiated gastric carcinoma. World J. Surg. Oncol. 20, 383 (2022).
Tessier-Cloutier, B. et al. Loss of switch/sucrose non-fermenting complex protein expression in undifferentiated gastrointestinal and pancreatic carcinomas. Histopathology 77, 46–54 (2020).
Chambers, L. et al. Bridging the gap from bench to bedside: a call for in vivo preclinical models to advance endometrial cancer and cervical cancer immuno-oncology research. Clin. Cancer Res. 30, 2905–2909 (2024).
Dawe, C. J., Banfield, W. G., Morgan, W. D., Slatick, M. S. & Curth, H. O. Growth in continuous culture, and in hamsters, of cells from a neoplasm associated with acanthosis nigricans. J. Natl Cancer Inst. 33, 441–456 (1964).
Crowe, A. R. & Yue, W. Semi-quantitative determination of protein expression using immunohistochemistry staining and analysis: an integrated protocol. Bio. Protoc. 9, e3465 (2019).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Springer Science & Business Media, 2009).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2016).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Acknowledgements
We thank all the members of C.K.’s laboratory and Y.W.’s and D.G.H.’s laboratories for assistance and discussion. Specifically, we thank S. Jain for assistance with ARID1A PiggyBac construct cloning and E. Li for isogenic cell line generation, as well as C. Young Shin and L. Hoang for assistance with DDEC microdissection. We also thank Z. Herbert, M. Sullivan and R. Garuti of the DFCI Molecular Biology Core Facility for help with high-throughput sequencing. J.S.L. is supported by the National Institutes of Health (NIH) Reproductive Scientist Development Program (grant no. K12-HD000849-37) and the GOG Foundation. This work was supported in part by an NIH DP2 New Innovator Award (to C.K.) and the Howard Hughes Medical Institute, Canadian Institute of Health Research (grant nos. PJT-462168 and PJT-197923 to Y.W.), Terry Fox New Frontiers Program Project Grants (nos. 1116 to Y.W. and 1082 to D.G.H). Y.W. is a recipient of the North Family Health Research Award. D.G.H. is supported by a Canada Research Chair in Molecular and Genomic Pathology. We also appreciate the generous support from the British Columbia Cancer Foundation and the VGH and UBC Hospital Foundation.
Author information
Authors and Affiliations
Contributions
J.S.L.: conceptualization, methodology, validation, formal analysis, investigation, data curation, writing, visualization and project administration. G.X., A.P., D.S.G., S.C., W.W.F., D.S.N. and B.G.: methodology, validation, formal analysis and data curation. A.W.Y., K.S.C. and A.S.: formal analysis, data curation, visualization and methodology. J.A.P.: methodology, validation, formal analysis, software and data curation. J.Q. and S.P.G.: resources. J.L.H.: data curation and resources. D.J.K.: methodology, investigation, analysis, data curation, resources and manuscript editing. Y.W.: methodology, validation, formal analysis, investigation, data curation, supervision, conceptualization, cell line-screening efforts, manuscript editing, project administration and funding acquisition. D.G.H.: resources, supervision and funding acquisition. C.K.: conceptualization, methodology, investigation, resources, data curation, writing, visualization, supervision, project administration and funding acquisition.
Corresponding author
Ethics declarations
Competing interests
C.K. is the Scientific Founder, Scientific Advisor to the Board of Directors, Scientific Advisory Board member, shareholder and consultant for Foghorn Therapeutics, Inc., serves on the Scientific Advisory Board of Nereid Therapeutics and is a consultant for Google Ventures. C.K. is also a member of the Molecular Cell and Cell Chemical Biology editorial boards. J.S.L is a shareholder for AI Proteins Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 mSWI/SNF gene mutations in dedifferentiated endometrial carcinomas.
A. Summary of whole-exome sequencing profiling of uterine dedifferentiated carcinomas (GENIE cohort, n = 22 cases), indicating mutation frequencies and type of mSWI/SNF, POLE, TP53, and other gene mutations. B. Summary of whole-exome sequencing profiling of endometrial/ovarian dedifferentiated carcinomas (Dana Farber dataset; n = 35 cases), indicating mutation frequencies and type of mSWI/SNF, POLE, TP53, and other genes. C. Summary of mutations in TCGA (n = 515 cases; serous/mixed/endometrioid carcinoma). Features are indicated in legend; Tumor type, tumor grade, mutation type and histology are indicated. D. Pie chart showing breakdown of mSWI/SNF gene mutations in DDEC cases from DFCI, Coatham et al., and TCGA. E. Mutual exclusivity analyses performed for mSWI/SNF, POLE, and TP53 on TCGA dataset of serous/mixed/endometrioid dedifferentiated carcinomas. F. Mutational characteristics for endometrial carcinoma cell lines used in this study. mSWI/SNF and top-mutated genes are shown (CCLE). *indicates damaging mutations. G. Immunoblot performed on nuclear extract (input) and IgG and anti-SMARCA4 immunoprecipitation from VOA1066 cells. mSWI/SNF components and TBP are shown. H. TMT-MS shown for SMARCA4 IP (and IgG control IP) in VOA1066 cells. I, J. Summary of n = 75 DDEC cases analyzed by IHC, indicating positive/negative protein staining status of ARID1A, ARID1B, SMARCA4, SMARCA2, and SMARCB1. K. SMARCA4 IHC performed on n = 3 ARID1A/ARID1B dual-mutated DDEC cases. Undifferentiated and well-differentiated compartments are indicated.
Extended Data Fig. 2 cBAF reassembly controls genome-wide redistribution of mSWI/SNF distribution and chromatin accessibility in DDEC cells.
A. Density sedimentation analyses performed on AN3CA ARID1A/B dual-deficient cells in +Dox (ARID1A rescue) conditions using 10-30% glycerol gradients. Immunoblot for selected mSWI/SNF subunits are shown; -Dox pair shown in Fig. 1e. B. PCA plot showing -/+ Dox (ARID1A rescue) conditions in AN3CA and VOA1066 cells. C. Volcano plots showing proteins (peptide signal) significantly increased and decreased upon rescue with ARID1A in AN3CA and VOA cell line systems. mSWI/SNF complex components are highlighted. D. Clustered heatmap demonstrating logFC of TMT mass-spectrometry signal in AN3CA and VOA1066 cells by subunit. E. Immunofluorescence imaging for pan-cytokeratin (green) with DAPI counterstain in both DDEC cell lines in -/+ dox conditions; scale bar= 20um, Dox treatment of 72 h, representative of n = 3 experimental replicates. F. Colony formation assays performed in AN3CA and VOA1066 cell lines following 14 days of Dox treatment (ARID1A rescue). G. Quantification of colony formation assays shown in (F), p-values determined with two-tailed Students t-test, error bars represent mean ± S.E.M. H. Bar graphs depicting number of peaks across antibodies profiled in ChIP-seq experiments. Venn diagrams depicting BRD9-GLTSCR1 (ncBAF) merged peaks in -/+ ARID1A rescue conditions in AN3CA and VOA1066 cell lines. I. Distance-to-TSS stacked bar graphs indicating peak localization for gained, lost, and retained SMARCA4 sites. J. HOMER motif analyses performed over gained and lost SMARCA4 sites in VOA1066 cells. K. PCA plot performed on ATAC–seq (broad peaks) datasets in -/+ Dox (ARID1A rescue) conditions in AN3CA and VOA1066 cells. L. Bar graphs depicting number of ATAC-seq peaks in AN3CA and VOA1066 cells in both conditions. M. Heatmaps reflecting ChIP-seq for GLTSCR1 and BRD9 as well as ATAC-seq in VOA1066 over ATAC-seq sites. Gained, lost and retained peak sets are indicated. N. Top, Overlap between de novo gained ARID1A peaks and BRD9/GLTSCR1 peaks prior to ARID1A restoration. Bottom, pie charts reflecting percent of ncBAF target sites replaced by cBAF following ARID1A restoration. O. Metaplot showing altered PBRM1 occupancy over TSS sites (top 20% expressed genes) in -/+ dox conditions. P. PBRM1 peaks in VOA1066 cells in -/+ dox conditions.
Extended Data Fig. 3 Gene regulatory impact of cBAF restoration in DDEC cell lines and primary tumors.
A. Immunoblot for indicated mSWI/SNF components following ARID1A restoration (72 h). B. PCA plots performed on RNA-seq in AN3CA and VOA1066 cell lines following ARID1A restoration (n = 3 experimental replicates). C. Heatmaps indicating clustering of -Dox and +Dox (+HA-ARID1) conditions. D. GSEA analyses performed on AN3CA and VOA cells (ARID1A rescue/no rescue); NES, Normalized Enrichment Score. E. Clustered heatmap indicating genes up and downregulated following ARID1A rescue. F. Venn diagrams indicating cell line-specific and shared up- and down-regulated genes upon ARID1A rescue. G. Volcano plots showing up- (red) and down- (blue) regulated proteins (by TMT-MS) following 72 h of ARID1A restoration in AN3CA and VOA1066 DDEC cell lines. H. Representative ChIP-seq and ATAC-seq tracks at the CDH1, CLDN4, TONSL, and BRCA2 gene loci with corresponding bar graphs depicting gene expression (RNA-seq).
Extended Data Fig. 4 Concordant and distinct impacts of ARID1A, ARID1B, or dual paralog restoration in DDEC cell lines.
A. Immunoblot for indicated mSWI/SNF components following ARID1A, ARID1B or dual ARID1A/B restoration (72 h). B. Bar graphs indicating quantitative densitometry of blots in (I). C. Venn diagrams indicating overlap of downregulated genes (left) and sites with decreased accessibility (right) following ARID1A, ARID1B or ARID1A/B dual rescue. D. GSEA (C2 gene set) performed on AN3CA cells with ARID1A, ARID1B or dual paralog rescue relative to no rescue (-Dox) in AN3CA cells. E. Z-score heatmap indicating genes upregulated, downregulated and corresponding genes in undifferentiated (UD) and well-differentiated (WD) primary tumor compartments. F. Hallmark GSEA in AN3CA, VOA1066 cell lines following ARID1A rescue (relative to - Dox) and DDEC UD tumor compartment, normal endometrium relative to DDEC WD tumor compartment (patient); NES, Normalized Enrichment Score.
Extended Data Fig. 5 Cell proliferation assays and CRISPR screening in cBAF-disrupted endometrial cell lines.
A. Colony formation assays performed on AN3CA cells bearing shRNAs targeting either BRD9 or ARID2. B. Immunoblot for ARID2, BRD9 and GAPDH in VOA1066 cells treated with indicated shRNA conditions. C. Bar graph depicting AUC values in indicated conditions in VOA1066 colony formation assays. D. Immunoblot for shRNA-mediated suppression of subunits indicated in AN3CA cells. E. Bar graph depicting AUC values in indicated conditions in AN3CA colony formation assays. F. Normalized read counts from genome-scale CRISPR screen (TKOv3 library). G. Box and whisker plots indicating impact across gene sets indicated on X-axis. H. Colony formation assays performed on HEC1B cells and HEC1B cells with ARID1B KO (ARID1B-BKO) treated with the indicated sgRNAs. I. Quantification of colony formation assays from (H) (relative confluency) at Day 10 post infection. J. Colony formation assays performed on HEC1B cells and HEC1B cells with ARID1B KO (ARID1B-BKO) treated with the indicated sgRNAs. K. Quantification of colony formation assays from (J) (relative confluency) at Day 10 post infection. L. Venn diagram depicting overlap between HEC1B ARID1B KO-specific dependencies and genes downregulated by ARID1 paralog restoration in AN3CA cell lines. p-values determined with two-tailed Students t-test for C, E, I and K. Error bars represent mean ± S.E.M.
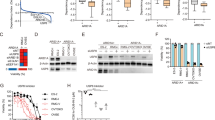
Extended Data Fig. 6 SMARCA4/2 ATPase inhibition and BRD9 degradation attenuate oncogenic gene regulation and proliferation in DDEC cells.
A. Cell proliferation measurements for VOA1066 cells upon treatment with either Cmp14 or dBRD9-A, compared to DMSO control. n = 3 experimental replicates; Significance values are indicated. B. Principal component analysis (PCA) performed on RNA-seq in VOA1066 and AN3CA cells in conditions indicated. C. Volcano plots indicating up- (red) and down- (blue) regulated genes upon either Cmp14 or dBRD9-A in VOA1066 cells; 72 h treatment. D. GSEA Hallmark performed on RNA-seq profiles in VOA1066 cells treated with Cmp14 and dBRD9-A, relative to DMSO. E. GSEA C2 gene set analysis performed in AN3CA cells treated with the indicated conditions. F. Venn diagram indicating overlap of upregulated genes (logFC>1; p < 0.01) in Cmp14 and dBRD9-A conditions in VOA1066 cells. G. Venn diagram indicated overlap of downregulated (top) and upregulated (bottom) genes (logFC>1; p < 0.01) in AN3CA cells. H. Representative ChIP-seq and ATAC-seq tracks at the GDF15 and EMP1 loci. I. PCA performed on ATAC-seq data in AN3CA and VOA1066 cells. J. Venn diagrams showing overlap of ATAC-seq peaks in AN3CA and VOA1066 cells in conditions indicated. K. HOMER motif analysis over sites with retained acccessibility (no change) in AN3CA and VOA1066 cells with dBRD9-A conditions. L. Venn diagrams indicating overlap between accessible peaks (ATAC-seq) in DMSO and Cmp14 conditions in each cell line. M. Homer motif analyses over sites with lost accessibility upon Cmp14 treatment. N. GSEA C2 gene set analysis of genes with altered accessibility withiin 2kB in AN3CA cells treated with either Cmp14 or dBRD9-A. O. Graph indicating relative confluency of HEC1B and HEC1B ARID1B KO cells across concentrations of FHD-286. P. Colony formation assays performed on HEC1B and HEC1B ARID1B KO cells treated with FHD-286 across concentrations indicated.
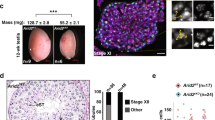
Extended Data Fig. 7 mSWI/SNF pharmacologic inhibition attenuates DDEC tumor growth in vivo.
A. H&E and IHC for ARID1A, ARID1B, SMARCA4 and SMARCA2 in the XVOA14590 PDX model system; scale bars 200um. B. Individual mouse tumor volume measurements (mm3) in XVOA14590 PDX model treated with either Cmp14 or dBRD9-A. C. Representative tumor images for XVOA14590 PDX model treated with vehicle control, Cmp14, or dBRD9-A; tumors isolated at Day 16, end of study. D. Immunoblot performed on whole-cell extracts for BRD9, CRBN and beta-actin in XVOA14590 PDX treated with dBRD9-A confirming degradation relative to vehicle. E. PCA analysis performed on RNA-seq profiles derived from n = 5 tumors in each condition indicated. F. Volcano and dot plots representing differentially expressed genes in Cmp14 vs DMSO cell lines (left) and dBRD9-A treated cells (right). G. GSEA Hallmark analyses performed on n = 6 primary tumors from XVOA14590 at Day 16 post treatment in vivo. H. Tumor weight measurements (g) for VOA1066 CDX treated with either Cmp14 (top) or FHD-286 (bottom). I. (Left) Cell proliferation experiments of VOA, RL95-2 and SNGM cells (by % confluency) using single-agent and FHD-286 and carboplatin combination. (Right) D-R-(LOWE) synergy plots using Combenefit studies performed in 2D culture. J. Percent change in body weight for XVOA14590 PDX model over days of treatment with single-agent and FHD-286 and carboplatin combinations. K. Individual mouse tumor volume measurements (mm3) in VOA1066 CDX model treated with vehicle, single-agent FHD-286, carboplatin, or combination FHD-286+carboplatin.
Supplementary information
Supplementary Information (download PDF )
Supplementary Note and Tables 1–8.
Supplementary Tables (download XLSX )
Supplementary Table 1 Data and calculations for all TCGA, DFCI, MSKCC endometrial carcinoma cases used in IHC. Supplementary Table 2 Features of DDEC cell lines used in the study. Supplementary Table 3 TMT–MS analysis for SMARCA4-bound complexes in VOA1066 cells. Supplementary Table 4 TMT–MS analysis for AN3CA and VOA1066 cell lines ± ARID1A WT rescue (+Dox). Supplementary Table 5 Clinicopathological information of patient samples used in LCM experiments and RNA-seq. Supplementary Table 6 Genome-scale CRISPR screening in HEC1B WT and ARID1A/B KO conditions. Supplementary Table 7 Recombinant DNA constructs utilized in this study. Supplementary Table 8 Antibodies used in this study for various methods.
Source data
Source Data Figs. 1, 2 and 5–7 and Extended Data Figs. 2, 4 and 5–7 (download XLSX )
Statistical source data supporting Figs. 1d,h, 2f, 5b, 6b, 7b,c,f,g,i,j and Extended Data Figs. 2g, 4b, 5b,c,e,i,k, 6a,j,o and 7h,i,k.
Source Data Figs. 1, 2, 4 and 5 and Extended Data Figs. 1–5 and 7 (download PDF )
Unprocessed western blots and/or gels supporting Figs. 1c,e, 2b,c, 4b and 5a,c and Extended Data Figs. 1g, 2a, 3a, 4a, 5b,d and 7d.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
St. Laurent, J.D., Xu, G.D., Ying, A.W. et al. Shifted assembly and function of mSWI/SNF family subcomplexes underlie targetable dependencies in dedifferentiated endometrial carcinomas. Nat Genet 57, 2743–2755 (2025). https://doi.org/10.1038/s41588-025-02333-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02333-9