Abstract
Understanding early embryonic development is fundamental for unraveling plant cell differentiation and organogenesis. Here we integrate multiomics data from 403 upland cotton ovules to identify 2,960 metabolic quantitative trait loci and 24,485 expression quantitative trait loci. A key locus, ME_A07, influencing 252 known metabolite levels and expression of 4,293 genes, with the MYB gene GhTT2_A07 identified as central regulator, potentially regulated by a 520 kb inversion. GhTT2_A07 orchestrated both primary and secondary metabolite biosynthesis, influencing agronomic traits. Another locus, ME_A06, driven by the MYB gene Proanthocyanidin Regulator (GhPAR), modulates proanthocyanin content and suggests an ecological adaptation. GhTT2_A07 and GhPAR exhibit both shared and distinct expression profiles, contributing variably to fiber quality and yield. These findings highlight the critical role of MYB genes in the early development of cotton ovules and fibers, offering comprehensive multiomics resources that advance cotton research and molecular breeding.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The raw RNA-seq data generated in this study are available in the NCBI BioProject database under accession number PRJNA776409. Genomic sequencing data can be accessed through the China National Center for Bioinformation (CNCB) under accession number PRJCA010331 or through the NCBI BioProject database under accession number PRJNA1277445, PRJNA412456, PRJNA497352, PRJNA605345 and PRJNA680449. Metabolomics data are also available from the CNCB under accession number PRJCA038211, and are also provided in the Supplementary Tables 2, 4 and 5. The assembled genome of the brown cotton cultivar ZMS81 can be downloaded from the CottonGVD database (https://db.cngb.org/cottonGVD/download/) or accessed through the NCBI BioProject database under accession number PRJNA1274689. Source data are provided with this paper.
Code availability
All software used in the study are publicly available and described in Methods and Reporting Summary.
References
Pichersky, E. & Lewinsohn, E. Convergent evolution in plant specialized metabolism. Annu. Rev. Plant Biol. 62, 549–566 (2011).
Sulpice, R. & McKeown, P. C. Moving toward a comprehensive map of central plant metabolism. Annu. Rev. Plant Biol. 66, 187–210 (2015).
Pichersky, E. & Gang, D. R. Genetics and biochemistry of secondary metabolites in plants: an evolutionary perspective. Trends Plant Sci. 5, 439–445 (2000).
Fiehn, O. Metabolomics - the link between genotypes and phenotypes. Plant Mol. Biol. 48, 155–171 (2002).
Crawford, B. C. W. et al. Genetic control of distal stem cell fate within root and embryonic meristems. Science 347, 655–659 (2015).
Maheshwari, P. Embryology of angiosperms as a field for research. Nature 156, 354–355 (1945).
Chen, W. et al. A novel integrated method for large-scale detection, identification, and quantification of widely targeted metabolites: application in the study of rice metabolomics. Mol. Plant 6, 1769–1780 (2013).
Fang, C. Y., Fernie, A. R. & Luo, J. Exploring the diversity of plant metabolism. Trends Plant Sci. 24, 83–98 (2019).
Polturak, G. et al. Transcriptome and metabolic profiling provides insights into betalain biosynthesis and evolution in Mirabilis jalapa. Mol. Plant 11, 189–204 (2018).
Nett, R. S., Lau, W. & Sattely, E. S. Discovery and engineering of colchicine alkaloid biosynthesis. Nature 584, 148 (2020).
Peng, Z. et al. Flavonoid biosynthetic and starch and sucrose metabolic pathways are involved in the pigmentation of naturally brown-colored cotton fibers. Ind. Crops Prod. 158, 113045 (2020).
Wang, L. et al. System level analysis of cacao seed ripening reveals a sequential interplay of primary and secondary metabolism leading to polyphenol accumulation and preparation of stress resistance. Plant J. 87, 318–332 (2016).
Ying, S. Y. et al. Trichome regulator SlMIXTA-like directly manipulates primary metabolism in tomato fruit. Plant Biotechnol. J. 18, 354–363 (2020).
Zhu, G. T. et al. Rewiring of the fruit metabolome in tomato breeding. Cell 172, 249–261 (2018).
Machado, A., Wu, Y., Yang, Y., Llewellyn, D. J. & Dennis, E. S. The MYB transcription factor GhMYB25 regulates early fibre and trichome development. Plant J. 59, 52–62 (2009).
Shan, C.-M. et al. Control of cotton fibre elongation by a homeodomain transcription factor GhHOX3. Nat. Commun. 5, 5519 (2014).
Walford, S.-A., Wu, Y., Llewellyn, D. J. & Dennis, E. S. Epidermal cell differentiation in cotton mediated by the homeodomain leucine zipper gene, GhHD-1. Plant J. 71, 464–478 (2012).
Wu, H. et al. Genetics and evolution of MIXTA genes regulating cotton lint fiber development. N. Phytol. 217, 883–895 (2018).
Du, X. et al. Resequencing of 243 diploid cotton accessions based on an updated A genome identifies the genetic basis of key agronomic traits. Nat. Genet. 50, 796–802 (2018).
He, S. et al. The genomic basis of geographic differentiation and fiber improvement in cultivated cotton. Nat. Genet. 53, 916–924 (2021).
Ma, Z. et al. Resequencing a core collection of upland cotton identifies genomic variation and loci influencing fiber quality and yield. Nat. Genet. 50, 803–813 (2018).
Haigler, C. H., Betancur, L., Stiff, M. R. & Tuttle, J. R. Cotton fiber: a powerful single-cell model for cell wall and cellulose research. Front. Plant Sci. 3, 104 (2012).
Tian, Y. & Zhang, T. MIXTAs and phytohormones orchestrate cotton fiber development. Curr. Opin. Plant Biol. 59, 101975 (2021).
Wendel, J. F., Brubaker, C., Alvarez, I., Cronn, R. & Stewart, J. M. in Genetics and Genomics of Cotton (ed. Paterson, A. H.) 3–22 (Springer, 2009).
Herrmann, K. M. The shikimate pathway as an entry to aromatic secondary metabolism. Plant Physiol. 107, 7–12 (1995).
Hohner, R. et al. Reduced arogenate dehydratase expression: ramifications for photosynthesis and metabolism. Plant Physiol. 177, 115–131 (2018).
Wang, X. C. et al. Arabidopsis MYB4 plays dual roles in flavonoid biosynthesis. Plant J. 101, 637–652 (2020).
Wang, Z. et al. TRANSPARENT TESTA2 regulates embryonic fatty acid biosynthesis by targeting FUSCA3 during the early developmental stage of Arabidopsis seeds. Plant J. 77, 757–769 (2014).
Nesi, N., Jond, C., Debeaujon, I., Caboche, M. & Lepiniec, L. The Arabidopsis TT2 gene encodes an R2R3 MYB domain protein that acts as a key determinant for proanthocyanidin accumulation in developing seed. Plant Cell 13, 2099–2114 (2001).
Sun, J., Sun, Y. & Zhu, Q. H. Breeding next-generation naturally colored cotton. Trends Plant Sci. 26, 539–542 (2021).
Kai, K. et al. Scopoletin is biosynthesized via ortho-hydroxylation of feruloyl CoA by a 2-oxoglutarate-dependent dioxygenase in Arabidopsis thaliana. Plant J. 55, 989–999 (2008).
Janowitz, T., Kneifel, H. & Piotrowski, M. Identification and characterization of plant agmatine iminohydrolase, the last missing link in polyamine biosynthesis of plants. FEBS Lett. 544, 258–261 (2003).
Brubaker, C. L. & Wendel, J. F. Reevaluating the origin of domesticated cotton (Gossypium hirsutum; Malvaceae) using nuclear restriction fragment length polymorphisms (RFLPs). Am. J. Bot. 81, 1309 (1994).
Fang, L. et al. Genomic analyses in cotton identify signatures of selection and loci associated with fiber quality and yield traits. Nat. Genet. 49, 1089–1098 (2017).
Yoo, M. J. & Wendel, J. F. Comparative evolutionary and developmental dynamics of the cotton (Gossypium hirsutum) fiber transcriptome. PLoS Genet. 10, e1004073 (2014).
Yan, Q. et al. Up-regulation of GhTT2-3A in cotton fibres during secondary wall thickening results in brown fibres with improved quality. Plant Biotechnol. J. 16, 1735–1747 (2018).
Wen, T. et al. Linkage and association mapping reveals the genetic basis of brown fibre (Gossypium hirsutum). Plant Biotechnol. J. 16, 1654–1666 (2018).
Tan, H. et al. A Crucial Role of GA-Regulated Flavonol Biosynthesis in Root Growth of Arabidopsis. Mol. Plant 12, 521–537 (2019).
Zhang, Z. et al. GmPIN1-mediated auxin asymmetry regulates leaf petiole angle and plant architecture in soybean. J. Integr. Plant Biol. 64, 1325–1338 (2022).
Zhu, L., Jiang, B., Zhu, J. & Xiao, G. Auxin promotes fiber elongation by enhancing gibberellic acid biosynthesis in cotton. Plant Biotechnol. J. 20, 423–425 (2022).
Han, L.-B. et al. The dual functions of WLIM1a in cell elongation and secondary wall formation in developing cotton fibers. Plant Cell 25, 4421–4438 (2013).
Bar-Peled, M. & O’Neill, M. A. Plant nucleotide sugar formation, interconversion, and salvage by sugar recycling. Annu. Rev. Plant Biol. 62, 127–155 (2011).
Richard, P. & Hilditch, S. d-Galacturonic acid catabolism in microorganisms and its biotechnological relevance. Appl. Microbiol. Biotechnol. 82, 597–604 (2009).
Vaughn, K. C. & Turley, R. B. The primary walls of cotton fibers contain an ensheathing pectin layer. Protoplasma 209, 226–237 (1999).
Pang, C.-Y. et al. Comparative proteomics indicates that biosynthesis of pectic precursors is important for cotton fiber and Arabidopsis root hair elongation. Mol. Cell. Proteom. 9, 2019–2033 (2010).
Yao, Y. et al. Comparative proteomic and biochemical analyses reveal different molecular events occurring in the process of fiber initiation between wild-type allotetraploid cotton and its fuzzless-lintless mutant. PLoS ONE 10, e0117049 (2015).
Di Ferdinando, M., Brunetti, C., Fini, A. & Tattini, M. Flavonoids as antioxidants in plants under abiotic stresses. Int. J. Mol. Sci. 14, 3540–3555 (2013).
Daryanavard, H., Postiglione, A. E., Muhlemann, J. K. & Muday, G. K. Flavonols modulate plant development, signaling, and stress responses. Curr. Opin. Plant Biol. 72, 102350 (2023).
Brown, D. E. et al. Flavonoids act as negative regulators of auxin transport in vivo in Arabidopsis. Plant Physiol. 126, 524–535 (2001).
Gou, J.-Y., Wang, L.-J., Chen, S.-P., Hu, W.-L. & Chen, X.-Y. Gene expression and metabolite profiles of cotton fiber during cell elongation and secondary cell wall synthesis. Cell Res. 17, 422–434 (2007).
Naoumkina, M., Hinchliffe, D. J., Turley, R. B., Bland, J. M. & Fang, D. D. Integrated metabolomics and genomics analysis provides new insights into the fiber elongation process in Ligon lintless-2 mutant cotton (Gossypium hirsutum L.). BMC Genomics 14, 155 (2013).
Yang, Z. E. et al. Extensive intraspecific gene order and gene structural variations in upland cotton cultivars. Nat. Commun. 10, 2989 (2019).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–U110 (2010).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinf. 12, 323 (2011).
Silva, I. T., Rosales, R. A., Holanda, A. J., Nussenzweig, M. C. & Jankovic, M. Identification of chromosomal translocation hotspots via scan statistics. Bioinformatics 30, 2551–2558 (2014).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 9, 559 (2008).
Chen, C. J. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Smoot, M. E., Ono, K., Ruscheinski, J., Wang, P.-L. & Ideker, T. Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27, 431–432 (2011).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 26, 1641–1650 (2009).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Ge, X. et al. Efficient genotype-independent cotton genetic transformation and genome editing. J. Integr. Plant Biol. 65, 907–917 (2023).
Gu, Z., Huang, C., Li, F. & Zhou, X. A versatile system for functional analysis of genes and microRNAs in cotton. Plant Biotechnol. J. 12, 638–649 (2014).
Lin, R. et al. Transposase-derived transcription factors regulate light signaling in Arabidopsis. Science 318, 1302–1305 (2007).
Wei, X. et al. GhRCD1 promotes cotton tolerance to cadmium by regulating the GhbHLH12-GhMYB44-GhHMA1 transcriptional cascade. Plant Biotechnol. J. 22, 1777–1796 (2024).
Acknowledgements
We thank Z. Lin of Huazhong Agricultural University for providing the cotton seeds of GhTT2_A07-OE lines and receptor plants, and R. Yang (the Sanya Institute of Nanjing Agricultural University) and X. Li (Institute of Cotton Research, Chinese Academy of Agricultural Sciences) for assisting in sample collection. We thank the National Cotton Germplasm Midterm Bank (Anyang) for providing the seeds. We thank the National Supercomputing Center in Zhengzhou for providing the computing resources. We thank the Wuhan Metware Biotechnology Co., Ltd. (Wuhan, China) for metabolomic results. This work was supported by the National Natural Science Foundation of China (grant nos. 32122062 to S.H., 32401819 to X.Z.), the National Key Research and Development Program of China (grant nos. 2022YFD1200300 to X.D., 2021YFF1000101 to S.H. and 2022YFF1001400 to Xueke Gao), Xinjiang Science and Technology Major Program (2023A02001 and 2024A02002), the Youth Innovation Program of Chinese Academy of Agricultural Sciences (grant no. Y2023QC23 to Xueke Gao), the Natural Science Foundation of Henan Province (252300421076), the Agricultural Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences and Xinjiang Tianchi Talents Program (S.H. and L.M.).
Author information
Authors and Affiliations
Contributions
X.D., L.M., J.C. and S.H. conceived of and conceptualized the study. X.Z. performed the bioinformatics analysis and data analysis. X.T. and Xueke Gao designed and performed the experiments. G.S., P.D., X. Wang, S.L., J.P., H.Z. and Xu Gao performed the bioinformatics analysis. Z. Peng, X. Geng, H.L., Z.W., Z. Pan, Y.J., B.C., L.W., D.H., A.X., C.D. and B.P. participated in the phenotype data investigation. X. Wei performed the EMSA experiment. X.Z., X.D., L.M., J.C. and S.H. wrote and revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Xueying Guan, Tianzhen Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Distributions of metabolite variation and heritability.
a,b, Coefficient of variation (CV) (a) and broad-sense heritability (H2) (b) for all detected metabolites. Histograms show the distribution of metabolites (vertical axes) against CV and H2 values (horizontal axes), respectively.
Extended Data Fig. 2 Genetic diversity of 403 accessions based on genotype data.
a, Population structure analysis when K = 2, 3 and 4. The x-axis represents the 403 accessions, with G1 (red), G2 (yellow), and G3 (blue) distinguished by different colors. b, Principal component analysis (PCA) conducted on all 403 accessions. c, The phylogenetic tree of the 403 accessions. The colored fiber cottons (brown) were marked.
Extended Data Fig. 3 Analysis of eQTL and their regulated genes.
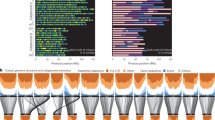
a, The distribution of eQTL and eGenes, which eQTL regulated. b, Comparison of association significance among local eQTL (n = 8,367), distal eQTL in the same chromosome (distal_intralChr, n = 4,921), and distal eQTL in the different chromosome (distal_interChr, n = 89,352). Box limits and center lines indicate quartiles and medians, respectively. Whiskers denote 1.5× interquartile range and points show outliers. Significances are tested by the two-tailed Student’s t-test. c-d, GO enrichment of Dt-eGenes, which eQTL regulated in subgenome A (c), and subgenome A excluding ME_A07 (d).
Extended Data Fig. 4 The downstream genes regulated by GhTT2_A07 and GhPAR.
a, The Yeast One-Hybrid (Y1H) assay indicates that GhTT2_A07 binds to the promoter regions of GhG6PDH, GhSK1, GhADT6, GhDFR, GhANR, GhLAR, and GhGT. b, The electrophoretic mobility shift assay (EMSA) has confirmed GhTT2_A07’s binding to the promoters of GhSK1, GhANR, and GhLAR in vitro. c, The Yeast One-Hybrid (Y1H) assay demonstrates GhPAR’s binding to the promoter regions of GhDFR, GhANR, GhLAR, and GhGT. d, The electrophoretic mobility shift assay (EMSA) confirms GhPAR’s binding to the promoters of GhDFR, GhANR, GhLAR, and GhGT in vitro. Each experiment was conducted with three independent biological replicates (a-d).
Extended Data Fig. 5 Characterization of GhTT2_A07 genetic engineered lines.
a, Fiber morphology of GhTT2_A07 overexpression (GhTT2_A07-OE) lines compared to the wild type. b, Comparative analysis of fiber strength, fiber micronaire, and the expression levels of downstream genes regulated by GhTT2_A07 between GhTT2_A07-OE lines and Jin668. c, Fiber morphology of GhTT2_A07-silenced lines compared to the control plants (acc. ZMS81). d, Comparative analysis of the expression levels of downstream genes regulated by GhTT2_A07 between GhTT2_A07-silenced lines and the control plants. Data are presented as mean values +/- SD (b, d). Statistical significances were determined using the two-tailed Student’s t-test (b, d). e, Comparison of GA3 content between GhTT2_A07-silenced lines and the control plants. All comparative analyses were conducted using three independent biological replicates (b, d-e).
Extended Data Fig. 6 Generation and characterization of GhPAR knockout (GhPAR-KO) lines using CRISPR/Cas9.
a, CRISPR/Cas9-mediated generation of GhPAR-KO lines using two distinct single-guide RNAs (sgRNA1 and sgRNA2), with the protospacer-adjacent motif (PAM) highlighted in blue. b. Comparison of fiber strength and fiber micronaire between the wild types, GhPAR-KO-1 lines and GhPAR-KO-4 lines. c, Sequence alterations in GhPAR-KO lines, with deletions and insertions represented by dashes and red texts, respectively. d. qRT-PCR analysis comparing the expression levels of GhDFR, GhANR, GhLAR, and GhGT in ovules (5 DPA) of the wild type, GhPAR-KO-1 lines and GhPAR-KO-4 lines. e, Quantification of various auxins and gibberellin (GA) in ovules (5 DPA) of the wild type, GhPAR-KO-1 lines and GhPAR-KO-4 lines. Data are presented as mean values +/- SD (b, d). Statistical significances are determined using a two-tailed Student’s t-test (b, d). All comparative analyses were conducted with three independent biological replicates (b, d-e).
Extended Data Fig. 7 GWAS for coumarins and identification of the candidate gene GhF6’H1.
a, Manhattan plots showing the SV-GWAS and SNP-GWAS results for scopoline (mws1077_P), scopoletin (pme2993), and scoparone (pme3569). The red rectangle denotes the genomic location of the trait locus on chromosome D08. Comparison of the relative content of three metabolites between two haplotypes. b, Local Manhattan plot (top) and LD heat map (bottom) between 65,705,310 and 65,723,000 bp. The statistical analysis was performed with Bonferroni correction. c, Haplotypes of coumarin locus in the 403 panel. d, Comparison of GhF6’H1 expression between the two haplotypes. n represents the sample size in each haplotype. Box limits and center lines indicate quartiles and medians, respectively, whiskers denote 1.5× interquartile range and points show outliers (a, d). Significances are tested by the two-tailed Student’s t-test (a, d). e, Association between D08 haplotypes and genotypes of the 9.3 kb deletion (DEL) and SNP in the GWAS population. The mapping reads were analyzed using both manual observation (IGV) and software (DELLY).
Extended Data Fig. 8 GWAS for phenolamides and identification of the candidate gene GhAIH.
a, Manhattan plots showing the GWAS results for N-acetylputrescine (pme2693) and agmatine (pmb0501). The red rectangle denotes the genomic location of the trait locus on chromosome D07. b, Local Manhattan plot (top) and LD heat map (bottom) between 6,835,563 and 6,895,484 bp. c, Haplotypes of phenolamide locus in the 403 panel. d-e, Comparison expression levels of GhAIH (d), and the relative content of two metabolites (e), between the two haplotypes. n represents the sample size in each haplotype. Significances are tested by the two-tailed Student’s t-test. Data are presented as median with interquartile range (d). Box limits and center lines indicate quartiles and medians, respectively, whiskers denote 1.5× interquartile range and points show outliers (e). f, Association between D07 haplotypes and genotypes of the 45 bp deletion and SNP in the GWAS population determined by resequencing mapping reads. The significance threshold is P = 0.05/n (where n is the independent SNP number), and the statistical analysis was performed with Bonferroni correction (a-b).
Extended Data Fig. 9 The effect assessment of two MYB homologous genes.
a, Changes in the contents of metabolites due to changes in GhPAR and GhTT2_A07 expression. The rounded rectangle shadow represents metabolites located on ME_A07, and the rectangular shadow represents metabolites on ME_A06. Asterisk indicates DAMs between Group I and Group II. The red-highlighted genes represent critical genes in the metabolic pathway. The solid-line arrows indicate genes verified to be regulated by GhTT2_A07 or GhPAR. b, The relative content of PA monomers and dimers among different haplotypes. The red and blue box indicates a significant increase and decrease in the accumulation of corresponding metabolites in A06HapII or A07 HapII genotypes. The sample size for each haplotype combination, from left to right, is represented as n = 343, 29, 18, 6. Box limits and center lines indicate quartiles and medians, respectively. Whiskers denote 1.5× interquartile range and points show outliers. Significances are tested by the two-tailed Student’s t-test.
Extended Data Fig. 10 Haplotype combination effects of ME_A06 and ME_A07 loci in the F2 population.
a, Fiber morphology of the parent lines Z128 (shorter fiber, carrying A06HapII and A07HapII) and Liao 96-23-30 (longer fiber, carrying both A06HapI and A07HapI), and F2 population (with a range of fiber colors and haplotype combinations). b-c, Heatmap of haplotypes on ME_A06 locus (b) and ME_A07 locus (c) in the F2 population. d, Comparison in agronomic traits among F2 individuals with different haplotype combinations. The sample size for each haplotype combination, from left to right, is represented as n = 23, 30, 19, 25. Box limits and center lines indicate quartiles and medians, respectively. Whiskers denote 1.5× interquartile range and points show outliers. All statistical significances were determined using the two-tailed Student’s t-test.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–19 and source data for Supplementary Figs. 14 and 17.
Supplementary Tables 1–21 (download XLSX )
Supplementary Table 1. The 32 representative Upland cotton accessions used for the MS2T library construction in this study. Supplementary Table 2. The 4,050 metabolites detected in the MS2T library constructed in this study. Supplementary Table 3. The 403 Upland cotton accessions used in this study. Supplementary Table 4. The raw dataset of metabolites. Supplementary Table 5. The 2,951 metabolites detected in 403 Upland cotton accessions in the study. Supplementary Table 6. The DAMs between Group I (WC) and Group II (brown cotton). Supplementary Table 7. The mGWAS signals (−log P > 7.31) for all the metabolites. Supplementary Table 8. The list of mQTL hotspots. Supplementary Table 9. Pearson’s correlation of 498 known metabolites and fiber traits (P < 10−9). Supplementary Table 10. GO enrichment of genes in the tan module. Supplementary Table 11. The information of genes in the tan module. Supplementary Table 12. The list of eQTL hotspots. Supplementary Table 13. The colocalization results of mGWAS and eGWAS. Supplementary Table 14. Pearson’s correlation of colocated metabolites and eGenes (P < 10−8). Supplementary Table 15. GO enrichment of 4,293 eGenes, which were correlated significantly with 252 metabolites and colocated on the ME_A07. Supplementary Table 16. Loci colocalized for fiber traits, genes and metabolites. Supplementary Table 17. Comparison of Fiber traits located on ME_A07 between two haplotypes. Supplementary Table 18. Comparison of key metabolites between two haplotypes. Supplementary Table 19. Comparison of metabolites located on ME_A07 between two haplotypes. Supplementary Table 20. The primer sequences used in this study. Supplementary Table 21. Whole-genome resequencing information of F2 population used in this study.
Source data
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, X., Tian, X., Gao, X. et al. Integrated metabolomic and transcriptomic analyses identify MYB genes regulating key metabolites and agronomic traits in upland cotton Gossypium hirsutum. Nat Genet 57, 2819–2830 (2025). https://doi.org/10.1038/s41588-025-02363-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02363-3