Abstract
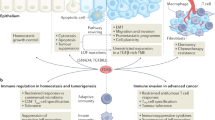
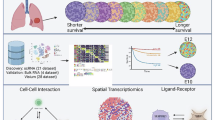
Transforming growth factor β (TGF-β) signaling in the tumor microenvironment predicts resistance to immune checkpoint blockade (ICB). While TGF-β inhibition enhances ICB efficacy in murine cancer models, clinical trials have yet to demonstrate benefit, underscoring the need to better understand its immunoregulatory roles across disease contexts. Using mouse models of advanced colorectal cancer and patient-derived data, we demonstrate that TGF-β impairs antitumor immunity at multiple levels in liver metastases. It acts directly on T cells to block recruitment of peripheral memory CD8+ T cells, thereby limiting the effectiveness of ICB. Concurrently, TGF-β instructs tumor-associated macrophages to suppress clonal expansion of newly arrived T cells by inducing SPP1 expression. This extracellular matrix protein promotes collagen deposition and accumulation of tumor-associated macrophages and fibroblasts, ultimately driving ICB resistance. Our findings reveal how TGF-β coordinates immunosuppressive mechanisms across innate and adaptive immune compartments to promote metastasis, offering new avenues to improve immunotherapy in colorectal cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Expression arrays and RNA-seq data are available at the Gene Expression Omnibus under the accession code GSE272376. Count matrices for single-cell RNA-seq experiments were deposited at the Gene Expression Omnibus under the accession code GSE273148. Source data are provided with this paper.
Code availability
No new code was developed in this study. The programs used to process and analyze the data are described in the relevant section of the Methods. Further information and requests for resources, biological material (MTOs) and reagents should be directed to the corresponding authors.
References
Cañellas-Socias, A., Sancho, E. & Batlle, E. Mechanisms of metastatic colorectal cancer. Nat. Rev. Gastroenterol. Hepatol. 21, 609–625 (2024).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015).
Tauriello, D. V. F. et al. TGFβ drives immune evasion in genetically reconstituted colon cancer metastasis. Nature 554, 538–543 (2018).
Tauriello, D. V. F., Sancho, E. & Batlle, E. Overcoming TGFβ-mediated immune evasion in cancer. Nat. Rev. Cancer 22, 25–44 (2022).
Soldevilla, B. et al. The correlation between immune subtypes and consensus molecular subtypes in colorectal cancer identifies novel tumour microenvironment profiles, with prognostic and therapeutic implications. Eur. J. Cancer 123, 118–129 (2019).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Nakanishi, Y. et al. Simultaneous loss of both atypical protein kinase C genes in the intestinal epithelium drives serrated intestinal cancer by impairing immunosurveillance. Immunity 49, 1132–1147 (2018).
Dodagatta-Marri, E. et al. α-PD-1 therapy elevates Treg/TH balance and increases tumor cell pSmad3 that are both targeted by α-TGFβ antibody to promote durable rejection and immunity in squamous cell carcinomas. J. Immunother. Cancer 7, 62 (2019).
Principe, D. R. et al. TGFβ blockade augments PD-1 inhibition to promote T-cell mediated regression of pancreatic cancer. Mol. Cancer Ther. 18, 613–620 (2018).
Holmgaard, R. B. et al. Targeting the TGFβ pathway with galunisertib, a TGFβRI small molecule inhibitor, promotes anti-tumor immunity leading to durable, complete responses, as monotherapy and in combination with checkpoint blockade. J. Immunother. Cancer 6, 47 (2018).
Terabe, M., et al. Blockade of only TGF-β 1 and 2 is sufficient to enhance the efficacy of vaccine and PD-1 checkpoint blockade immunotherapy. Oncoimmunology 6, e1308616 (2017).
Herzog, B. H. et al. Tumor-associated fibrosis impairs immune surveillance and response to immune checkpoint blockade in non-small cell lung cancer. Sci. Transl. Med. 15, eadh8005 (2023).
Usset, J. et al. Five latent factors underlie response to immunotherapy. Nat. Genet. 56, 2112–2120 (2024).
Batlle, E. & Massagué, J. Transforming growth factor-β signaling in immunity and cancer. Immunity 50, 924–940 (2019).
Kelly, A., Houston, S. A., Sherwood, E., Casulli, J. & Travis, M. A. Regulation of innate and adaptive immunity by TGFβ. Adv. Immunol. 134, 137–233 (2017).
Chakravarthy, A., Khan, L., Bensler, N. P., Bose, P. & Carvalho, D. D. D. TGF-β-associated extracellular matrix genes link cancer-associated fibroblasts to immune evasion and immunotherapy failure. Nat. Commun. 9, 4692 (2018).
Grout, J. A. et al. Spatial positioning and matrix programs of cancer-associated fibroblasts promote T-cell exclusion in human lung tumors. Cancer Discov. 12, 2606–2625 (2022).
Salmon, H. et al. Matrix architecture defines the preferential localization and migration of T cells into the stroma of human lung tumors. J. Clin. Invest. 122, 899–910 (2012).
Wang, Q., Strong, J. & Killeen, N. Homeostatic competition among T cells revealed by conditional inactivation of the mouse Cd4 gene. J. Exp. Med. 194, 1721–1730 (2001).
Liu, B. et al. Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat. Cancer 3, 108–121 (2022).
Gueguen, P. et al. Contribution of resident and circulating precursors to tumor-infiltrating CD8+ T cell populations in lung cancer. Sci. Immunol. 6, eabd5778 (2021).
Wang, A. Z. et al. Glioblastoma-infiltrating CD8+ T cells are predominantly a clonally expanded GZMK+ effector population. Cancer Discov. 14, 1106–1131 (2024).
Yap, T. A. et al. First-in-human phase I study of a next-generation, oral, TGFβ receptor 1 inhibitor, LY3200882, in patients with advanced cancer. Clin. Cancer Res. 27, 6666–6676 (2021).
Jung, S. Y. et al. Pharmacokinetic characteristics of vactosertib, a new activin receptor-like kinase 5 inhibitor, in patients with advanced solid tumors in a first-in-human phase 1 study. Invest. New Drugs 38, 812–820 (2020).
Álvarez-Varela, A. et al. Mex3a marks drug-tolerant persister colorectal cancer cells that mediate relapse after chemotherapy. Nat. Cancer 3, 1052–1070 (2022).
Liu, Y. et al. Immune phenotypic linkage between colorectal cancer and liver metastasis. Cancer Cell 40, 424–437 (2022).
Lee, H.-O. et al. Lineage-dependent gene expression programs influence the immune landscape of colorectal cancer. Nat. Genet. 52, 594–603 (2020).
Qi, J. et al. Single-cell and spatial analysis reveal interaction of FAP+ fibroblasts and SPP1+ macrophages in colorectal cancer. Nat. Commun. 13, 1742 (2022).
Sathe, A. et al. Colorectal cancer metastases in the liver establish immunosuppressive spatial networking between tumor-associated SPP1+ macrophages and fibroblasts. Clin. Cancer Res. 29, 244–260 (2023).
Wu, Y. et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov. 12, 134–153 (2022).
Bill, R. et al. CXCL9:SPP1 macrophage polarity identifies a network of cellular programs that control human cancers. Science 381, 515–524 (2023).
Obradovic, A. et al. Single-cell protein activity analysis identifies recurrence-associated renal tumor macrophages. Cell 184, 2988–3005 (2021).
Ma, R.-Y., Black, A. & Qian, B.-Z. Macrophage diversity in cancer revisited in the era of single-cell omics. Trends Immunol. 43, 546–563 (2022).
Park, M. D. et al. TREM2 macrophages drive NK cell paucity and dysfunction in lung cancer. Nat. Immunol. 24, 792–801 (2023).
Kirschenbaum, D. et al. Time-resolved single-cell transcriptomics defines immune trajectories in glioblastoma. Cell 187, 149–165 (2024).
Yofe, I. et al. Spatial and temporal mapping of breast cancer lung metastases identify TREM2 macrophages as regulators of the metastatic boundary. Cancer Discov. 13, 2610–2631 (2023).
Zhang, L. et al. Single-cell analyses inform mechanisms of myeloid-targeted therapies in colon cancer. Cell 181, 442–459 (2020).
Lavin, Y. et al. Innate immune landscape in early lung adenocarcinoma by paired single-cell analyses. Cell 169, 750–765 (2017).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308 (2018).
Raghavan, S. et al. Microenvironment drives cell state, plasticity, and drug response in pancreatic cancer. Cell 184, 6119–6137 (2021).
Li, S. & Jakobs, T. C. Secreted phosphoprotein 1 slows neurodegeneration and rescues visual function in mouse models of aging and glaucoma. Cell Rep. 41, 111880 (2022).
Wang, Y. & Lui, W.-Y. Transforming growth factor-β1 attenuates junctional adhesion molecule-A and contributes to breast cancer cell invasion. Eur. J. Cancer 48, 3475–3487 (2012).
Shaim, H., et al. Targeting the αv integrin/TGF-β axis improves natural killer cell function against glioblastoma stem cells. J. Clin. Invest. 131, e142116 (2021).
Singh, N. N. & Ramji, D. P. Transforming growth factor-β-induced expression of the apolipoprotein E gene requires c-Jun N-terminal kinase, p38 kinase, and casein kinase 2. Arterioscler. Thromb. Vasc. Biol. 26, 1323–1329 (2006).
Croxford, A. L. et al. The cytokine GM-CSF drives the inflammatory signature of CCR2+ monocytes and licenses autoimmunity. Immunity 43, 502–514 (2015).
Wang, L. et al. Single-cell transcriptome analysis profiling lymphatic invasion-related TME in colorectal cancer. Sci. Rep. 14, 8911 (2024).
Moorman, A. et al. Progressive plasticity during colorectal cancer metastasis. Nature 637, 947–954 (2025).
Lund, S. A., Giachelli, C. M. & Scatena, M. The role of osteopontin in inflammatory processes. J. Cell Commun. Signal. 3, 311–322 (2009).
Icer, M. A. & Gezmen-Karadag, M. The multiple functions and mechanisms of osteopontin. Clin. Biochem. 59, 17–24 (2018).
Liaw, L. et al. Altered wound healing in mice lacking a functional osteopontin gene (Spp1). J. Clin. Invest. 101, 1468–1478 (1998).
Klement, J. D. et al. An osteopontin/CD44 immune checkpoint controls CD8+ T cell activation and tumor immune evasion. J. Clin. Invest. 128, 5549–5560 (2018).
Hoeft, K. et al. Platelet-instructed SPP1+ macrophages drive myofibroblast activation in fibrosis in a CXCL4-dependent manner. Cell Rep. 42, 112131 (2023).
Li, G. et al. TGF-β-dependent lymphoid tissue residency of stem-like T cells limits response to tumor vaccine. Nat. Commun. 13, 6043 (2022).
Zinselmeyer, B. H. et al. PD-1 promotes immune exhaustion by inducing antiviral T cell motility paralysis. J. Exp. Med. 210, 757–774 (2013).
Mori, R., Shaw, T. J. & Martin, P. Molecular mechanisms linking wound inflammation and fibrosis: knockdown of osteopontin leads to rapid repair and reduced scarring. J. Exp. Med. 205, 43–51 (2008).
Levéen, P. et al. Induced disruption of the transforming growth factor β type II receptor gene in mice causes a lethal inflammatory disorder that is transplantable. Blood 100, 560–568 (2002).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Clausen, B. E., Burkhardt, C., Reith, W., Renkawitz, R. & Förster, I. Conditional gene targeting in macrophages and granulocytes using LysMCre mice. Transgenic Res. 8, 265–277 (1999).
Schindelin, J., et al. Fiji — an open platform for biological image analysis. Nat. Methods 9, 676–682 (2009).
Thevenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998).
Meijering, E. H. W., Niessen, W. J. & Viergever, M. A. Quantitative evaluation of convolution-based methods for medical image interpolation. Med. Image Anal. 5, 111–126 (2001).
Zuiderveld, K. Contrast limited adaptive histogram equalization. In Graphics Gems 474–485 (Elsevier, 1994).
Tinevez, J.-Y. et al. TrackMate: an open and extensible platform for single-particle tracking. Methods 115, 80–90 (2017).
Ershov, D. et al. TrackMate 7: integrating state-of-the-art segmentation algorithms into tracking pipelines. Nat. Methods 19, 829–832 (2022).
Zhang, G., He, B. & Weber, G. F. Growth factor signaling induces metastasis genes in transformed cells: molecular connection between Akt kinase and osteopontin in breast cancer. Mol. Cell. Biol. 23, 6507–6519 (2003).
Campeau, E., et al. A versatile viral system for expression and depletion of proteins in mammalian cells. PLoS ONE 4, e6529 (2009).
Krueger, F., James, F., Ewels, P., Afyounian, E. & Schuster-Boeckler, B. FelixKrueger/TrimGalore: v0.6.7 — DOI via Zenodo. Zenodo https://doi.org/10.5281/ZENODO.5127899 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. The R package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 47, e47 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Andreatta, M. et al. Semi-supervised integration of single-cell transcriptomics data. Nat. Commun. 15, 872 (2024).
Andreatta, M. & Carmona, S. J. UCell: robust and scalable single-cell gene signature scoring. Comput. Struct. Biotechnol. J. 19, 3796–3798 (2021).
Kang, J. B. et al. Efficient and precise single-cell reference atlas mapping with Symphony. Nat. Commun. 12, 5890 (2021).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Xie, Z., et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Chen, E. Y., et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Huang, R., et al. The NCATS BioPlanet — an integrated platform for exploring the universe of cellular signaling pathways for toxicology, systems biology, and chemical genomics. Front. Pharmacol. 10, 445 (2019).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Borcherding, N., Bormann, N. L. & Kraus, G. scRepertoire: an R-based toolkit for single-cell immune receptor analysis. F1000Res. 9, 47 (2020).
Cañellas-Socias, A. et al. Metastatic recurrence in colorectal cancer arises from residual EMP1+ cells. Nature 611, 603–613 (2022).
Therneau, T. M. coxme: mixed effects Cox models. R package v.2.2-16 (Cran, 2020).
Therneau, T. M., Grambsch, P. M. & Pankratz, V. S. Penalized survival models and frailty. J. Comput. Graph. Stat. 12, 156–175 (2003).
Pelka, K. et al. Spatially organized multicellular immune hubs in human colorectal cancer. Cell 184, 4734–4752 (2021).
Acknowledgements
We thank all members of the Batlle and Heyn laboratories for their support and discussions. We are grateful for the outstanding assistance from IRB Barcelona core facilities for biostatistics, histopathology, functional genomics and advanced digital microscopy as well as the flow cytometry and animal facilities of the Universitat de Barcelona, PCB and the CRG genomic unit. A.H. held a postdoctoral fellowship from European Union’s Horizon 2020 research and innovation program under Marie Skłodowska-Curie grant agreement no. 754510. A.P. held a postdoctoral fellowship from the Spanish Association against Cancer Scientific Foundation (FC AECC, INVES223171PRAD). M.S.-C. held a PhD fellowship from MINECO (FPI program) and E.M.-G. likewise from AGAUR, Generalitat de Catalunya (FI program). This work has been supported by the Fundación Olga Torres, Fundación La Caixa (LCF/PR/HR19/52160018), ERC Advanced Grant 884623, AECC (PRYGN223027BATL and PROYE18046BATL), the Spanish MICINN (PID2020-119917RB-I00), CRUK/AECC CRC STARS Alliance (GEACC248909TABE) and Worldwide Cancer Research (19-0005) (to E.B.), La Marató de TV3 (201903-30-31-32) (to E.B. and H.H.) and AGAUR 2021 SGR-001278. IRB Barcelona is the recipient of a Severo Ochoa Award of Excellence from MINECO. This project was also founded by the AECC Excellence Award to IRB Barcelona.
Author information
Authors and Affiliations
Contributions
E.B., A.H. and M.S.-C. conceived the study, coordinated experiments and wrote the manuscript. H.H. conceived and coordinated scRNA-seq experiments. A.P. coordinated experiments, analyzed scRNA-seq datasets and conceptualized data derived from genetically engineered mouse models. A.H. designed, performed and analyzed results involving myeloid cells and Spp1, as well as a subset of experiments on T cells; M.S.-C. designed, performed and analyzed a subset of experiments on T cells. J.B.-R., A.H. and M.S.-C. generated scRNA-seq datasets. S.P.-P. and X.H.-M. provided technical support with mouse work. M.S. performed immunohistochemistry (IHC) and immunofluorescence (IF). P.N. analyzed mouse scRNA-seq data and TCR datasets. L.M. analyzed bulk RNA-seq data. E.M.-G. analyzed human scRNA-seq datasets. J.N., G.C., D.M. and M.R.G. provided technical assistance with scRNA-seq experiments. S.T. developed the image analysis workflow to quantify T cell localization and motility from the migration assay; T.S., M.L.M. and E.M.H. performed image analysis and quantifications. A.R. and C.S.-Z. synthesized galunisertib. D.V.F.T. developed the MTO model and corrected the manuscript. E.S. provided logistic support and helped with figures and manuscript writing. A.C.-M. and C.S.-O.A. performed statistical analysis and analyzed patient datasets. E.B. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
H.H. is a cofounder and the chief scientific officer of Omniscope; a scientific advisory board member at NanoString, Bruker and Mirxes; and a consultant for Moderna; and has received honoraria from Genentech. E.B. and D.V.F.T. hold a patent on the synthesis and use of a TGFBRI small-molecule inhibitor. The laboratory of E.B. has received research funding from Merus, Incyte and Revolution Medicines. E.B. has received honoraria for consulting from Genentech. The other authors declare no competing financial interests.
Peer review
Peer review information
Nature Genetics thanks Karuna Ganesh, Jaeyop Lee, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 TGF-β limits the proliferation and accumulation of CD8+ TEFF TILs.
a, Expression levels of key marker genes defining the six CD8+ T cell clusters identified by 5′ scRNA-seq. b, Average expression of published gene signatures for T cell phenotypes (left) and general biological processes (right) across CD8+ clusters (see Supplementary Table 1). c, Cycling CD8+ TILs were defined by a cycling score > 0.2. Non-cycling cells were used to construct a reference map into which cycling cells were projected. d, Violin plots showing expression of Pdcd1, Cxcr3, Ly6a and Crtam across CD8+ clusters. Markers were selected based on differential expression, surface localization, and availability of validated antibodies. e, Flow cytometry gating strategy for CD8+ TIL subpopulations based on PD-1, CXCR3, SCA-1 and CRTAM. Naïve cells were PD-1-negative; T progenitor (TPR) were PD-1+ with high CXCR3 expression. PD-1hi cells were subdivided by SCA-1 and CRTAM expression into CRTAM+ T tissue resident (TTR), CRTAM−/SCA-1+ T effectors (TEFF), and CRTAM−/SCA-1− T exhausted (TEX) cells. f, CD8+ TIL composition in metastases following 3 days of treatment, analyzed by flow cytometry. Mice per group: vehicle, n = 26; galunisertib, n = 15; αPD-L1, n = 29; dual, n = 17. Mean ± S.D. g, TCR clonal size composition of scRNA-seq CD8+ clusters. Cells were categorized as single (=1), small (1 < x ≤ 5), medium (5 < x ≤ 20), large (20 < x ≤ 100), hyper expanded (100 < x ≤ 250) based on TCR sharing. h, Cell fate trajectories of expanded CD8+ TILs (>1 TCR copy) were inferred with Monocle 3, starting from naïve-like cells. i, Pseudotime trajectory values per CD8+ cluster. Box plots show median, IQR, and 1.5× IQR whiskers; outliers plotted individually. All statistical tests were two-sided and corrected for multiple comparisons where appropriate. Statistical analysis was performed using a generalized linear mixed-effects model (beta family) with Dunnett’s multiple comparisons (f).
Extended Data Fig. 2 TGF-β blockade with alternative TGFBR1 inhibitors confirms therapeutic effects on T cells and TAMs.
a, Schematic of acute (3-day; endpoint day 18) and long-term (15-day; endpoint day 30) in vivo treatment of liver metastases (LiMs), established by injecting AKTP MTOs into the spleen of C57BL/6J mice. Mice were treated with monoclonal antibodies (αPD-L1/IgG) and/or vactosertib (or LY3200882 in h–n) starting on day 12 (long-term) or day 15 (acute). Antibodies were administered every 3 days. b, c, Number of liver metastases (b) and CD3+ T cell density (cells/mm2) (c) at day 30 in mice treated with vactosertib and/or αPD-L1. Mice per group: Vehicle, n = 5; αPD-L1, n = 4; vactosertib, n = 5; vactosertib + αPD-L1, n = 8. d, e, CD8+ TIL subpopulation (Naïve, TPR, TEFF, TEX) frequencies (d) and TEFF:TEX ratio (e) after 3 days of vactosertib treatment. f, g, Ki67+ percentages in total CD8+ T cells (f) and in subpopulations (g) by flow cytometry. Mice per group: Vehicle, n = 8; vactosertib, n = 6. h, Schematic as in (a), with animals being treated with LY3200882. i, j, Number of metastases (i) and CD3+ T cell density (cells/mm2) (j) at day 30. Mice per group: Vehicle, n = 7; αPD-L1, n = 7; LY3200882, n = 5; LY3200882 + αPD-L1, n = 7. k, l, CD8+ TIL subpopulation frequencies (k) and TEFF:TEX ratios (l) after 3 days of LY3200882. m, n, Ki67+ percentages in total CD8+ T cells (m) and in subpopulations (n) by flow cytometry. Mice per group: Vehicle, n = 6; LY3200882, n = 8. o, p, mRNA expression of selected genes in TAMs sorted from LiMs in mice treated with vactosertib (o) or LY3200882 (p) vs. vehicle. Mice: LY3200882, n = 5; vactosertib, n = 7; vehicle, n = 6. All data are Mean ± SEM. All statistical tests were two-sided with correction for multiple comparisons where applicable. Statistical analysis performed as following: negative binomial GLM (b, i), linear model with log transformation (c, f, j, m), box-cox transformation (d, g, k, o, p), and linear mixed-effects model with Dunnett’s test (e, l). Schematics created with BioRender.
Extended Data Fig. 3 TGF-β limits continuous CD8+ TIL tumor infiltration.
a, Flow cytometry analysis of CD8+ blood populations 3 days post-treatment. Mice per group: Vehicle = 6, galunisertib = 6, αPD-L1 = 8, dual = 5. Mean ± SD. b, CXCR3 expression in CD8+ blood populations. Same group sizes as (a). Mean ± SD. c, Schematic of T cell infiltration blockade using FTY720, administered 3 days pre-treatment. d, CD3+ T cells in 100 μl blood after FTY720 (empty dots) or saline (filled dots) pre-treatment, followed by galunisertib or vehicle. Mice per group: Vehicle = 3, vehicle + FTY720 = 2; galunisertib = 4, galunisertib + FTY720 = 5. Mean ± SEM. e, Schematic of anti-CXCR3 mAb treatment for 3 days in tumor-bearing mice. f, CD3+, CD4+, and CD8+ T cells in 200 μl blood after anti-CXCR3 or IgG. Mean ± SEM. g, h, Frequencies of naïve (CD62L+CD44−), central memory (TCM) (CD62L+CD44+) and effector memory (TEM) (CD62L−CD44+) subsets in CD8+ (g) and CD4+ (h) T cells. n = 3/condition. Mean ± SEM. i, Percentage of high motility T cells at 200 μm from metastasis edge. Animals were previously administered one dose of FTY720 or saline, 3 days pre-treatment. Mice per group/metastasis: Vehicle = 6/38; αPD-L1 = 5/34; Dual = 8/35; FTY720 + αPD-L1 = 3/20; FTY720 + Dual = 4/23. Box plots show median, IQR, and 1.5× IQR whiskers; outliers plotted individually. All statistical tests were two-sided and corrected for multiple comparisons where applicable. Statistical analysis performed as following: linear model with log transformation (f, g, h), generalized linear model with mouse as random effect (Poisson, log link) for (a), linear mixed model with box-cox transformation (b), and with Dunnett’s correction for (i). Schematics generated with BioRender.
Extended Data Fig. 4 Genetically targeting TGF-β signaling in T cells.
a, Schematic of the dLckCre; Rosa26mTmG;Tgfbr2fl/fl mouse model. dLckCre mice were crossed with Tgfbr2fl/fl mice to generate T cell–specific Tgfbr2 knockout mice, then bred with Rosa26mTmG mice. Resulting T cells were either Tgfbr2WT/TOM (expressing TBRII and mTomato) or Tgfbr2dLck-KO/GFP (lacking TBRII and expressing mGFP). LoxP sites are indicated by triangles. b, Recombination efficiency in peripheral blood T cells from 12-week-old male mice, measured as % GFP+ cells by flow cytometry. n = 6 mice/genotype. Mean ± SD. c, Tgfbr2 mRNA levels in sorted splenocytes from 12-week-old males, assessed by qPCR. n = 3 and 4 (left to right). Mean ± SD. d, Representative flow cytometry plot of surface TBRII expression in CD3+ splenocytes from WT C57BL/6J or dLckCre; Rosa26mTmG;Tgfbr2fl/fl mice, gated as Tomato+ (Tgfbr2WT/TOM) or GFP+ (Tgfbr2dLck-KO/GFP). e, Schematic of adoptive cell transfer (ACT) of primed Tgfbr2WT/TOM or Tgfbr2dLck-KO/GFP T cells into WT recipients, two weeks post-metastasis inoculation. f, Representative flow cytometry plot of CD8+ blood populations in donor splenocytes prior to ACT. g, Schematic of ACT using primed WT T cells stained with Red Cell Tracker (vehicle-treated) or CFSE (galunisertib-treated), transferred into WT recipients two weeks after liver metastasis inoculation. h, Tissue distribution of transferred CD8+ T cells detected by flow cytometry. Each dot represents the percentage of CFSE+ (Gal) or Red+ (Vehicle) cells among all transferred cells (CFSE+ + Red+) in a given tissue. n = 4 mice. All statistical tests were two-sided and corrected for multiple comparisons where appropriate. Statistical analysis in (h) was performed using a linear model with mouse as a random effect. Schematics created with BioRender.
Extended Data Fig. 5 Characterization of liver metastases TME and effects of TGF-β inhibition in the immune compartments.
a, Flow cytometry gating strategy for identifying major immune populations in liver metastases from AKTP MTO–injected C57BL/6J mice. After exclusion of doublets (FSC-H vs. FSC-A; not shown) and dead cells (Live/Dead Aqua), CD45+ leukocytes were analyzed. Neutrophils were identified as Ly6G+ (R1). Remaining cells were divided into CD11b+/CD11c+ myeloid (R1) and CD11b−/CD11c− lymphoid (Lympho) subsets. B cells were MHCII+ CD24+; T cells were MHCII− CD24−. Myeloid cells (R1) were further resolved by side scatter and MHCII expression. Monocytes: CD11bhi MHC II− Ly6Chi CD64int; eosinophils: MHCII− CD24+. MHC II+ cells (R3) were divided into macrophages (CD64+ CD24−) and dendritic cells (CD64− CD24+). Plots representative of 5 independent experiments. b, Schematic of acute in vivo treatment (3 days from day 15) with vehicle or galunisertib in mice bearing liver metastases. c, Absolute numbers of immune populations from metastases, normalized to tissue weight. Mice per group: vehicle, n = 9; galunisertib, n = 11. Mean ± SEM. d, Schematic of in vivo treatment with vehicle, αPD-L1, galunisertib or combination (Tx from day 15; one antibody dose on day 15; endpoint day 18). Tumors were macrodissected, digested, and EpCAM− cells (immune, CAFs and endothelial cells) were isolated for 3′ scRNA-seq. e, UMAP of 43,377 stromal cells from liver metastases, color-coded by stromal cell type. Mice per group: vehicle, n = 4; αPD-L1, n = 2; galunisertib, n = 2; dual, n = 2. f, Dot plot showing expression of canonical markers across 12 stromal clusters. Dot size reflects % of cells expressing the gene; color represents scaled average expression. All statistical tests were two-sided and corrected for multiple comparisons where appropriate. Statistical analysis in (c) used a linear model with box-cox transformation. Schematics generated with BioRender. N, neutrophils; Mo, monocytes; TAM, tumor-associated macrophages; DC, dendritic cells.
Extended Data Fig. 6 Effects of TGFBR1 inhibition and αPD-L1 on TAMs.
a, Canonical marker gene expression across all tumor-associated macrophage (TAM) subsets. Dot size indicates the proportion of cells expressing each gene; color reflects expression centered to the average across all TAMs (scale: −1 to 2). b, Canonical marker gene expression in C1q+ and Spp1+ TAM subsets, centered to their subset averages (scale: −2 to 2). Dot size represents the proportion of expressing cells per cluster. c, Proportions of TAM subsets across treatments (vehicle, αPD-L1, galunisertib, dual). Mice per group: vehicle, n = 4; αPD-L1, n = 2; galunisertib, n = 2; dual, n = 2. Horizontal lines indicate medians. d, UMAP of TAMs showing Trem2 expression across all treatments. Color scale reflects expression level. e, Volcano plot showing differentially expressed genes in TAMs between vehicle- and αPD-L1–treated groups (10x Genomics). Genes with P < 0.05 and log2(fold change) ≥ 0.85 are considered significant. NCM, non-classical monocytes; TAM, tumor-associated macrophages.
Extended Data Fig. 7 Functional characterization of LysMCre-mediated Tgfbr2 deletion across tumor stages and the impact of neutrophil depletion on ICB efficacy.
a, Schematic of the LysMCre; Rosa26mTmG; Tgfbr2fl/fl mouse model. LysMCre mice were crossed with Tgfbr2fl/fl mice to generate a myeloid-specific Tgfbr2 knockout colony, then bred with Rosa26mTmG mice. Cre-mediated recombination excises mTomato, enabling mGFP expression. Triangles indicate loxP sites. b, Absolute numbers of immune populations (by flow cytometry as depicted in Extended Data Fig. 5a) in peripheral blood of naïve in LysMCre; Tgfbr2fl/fl vs. LysMCre; Tgfbr2+/+ mice, normalized per µl. c, d, Frequencies of naïve, central memory (TCM), and effector memory (TEM) CD8+ T cells (c) and proportion of CXCR3+ CD8+ T cells (d). n = 6 per group. Mean ± SEM. e, Schematic of liver metastasis generation via intrasplenic injection of AKTP MTOs in LysMCre; Tgfbr2fl/fl vs. control mice. f, GFP+ recombined immune populations in macrodissected liver metastases assessed by flow cytometry. Mice per group: LysMCre; Tgfbr2+/+, n = 7; LysMCre; Tgfbr2fl/fl, n = 5. Mean ± SEM. g, Tgfbr2 mRNA expression in TAMs sorted from liver metastases at day 18, normalized to Ppia and Tbp. Mice per group: LysMCre; Tgfbr2+/+, n = 4; LysMCre; Tgfbr2fl/fl, n = 3. Mean ± SEM. h, i, IHC (CD3) and IF (CD8, Ki67) images and quantification from LiMs 17 days post-injection. Mean ± SEM. Scale bars: CD3, 500 µm; CD8/Ki67, 50 µm. Arrowheads indicate Ki67+CD8+ cells. j, Relative expression of selected genes in TAMs from LiMs at day 18. Mice per group: LysMCre; Tgfbr2+/+, n = 8; LysMCre; Tgfbr2fl/fl, n = 7. Mean ± SEM. k, Schematic of treatment of established LiMs (Tx from day 12) with αPD-L1 (every 3 days), αLy6G (daily), or α-rat κ (every other day). l, m, Number of liver metastases (l) and CD3+ T cell densities (m) at day 24. Mean ± SEM. n, Neutrophil ablation validated via IF for LCN2+ cells. Mice per group: n = 5, 5, 5, 7, 5. Mean ± SEM. Statistical tests were two-sided and corrected for multiple comparisons. Statistical analysis performed as following, linear model (d); linear model with random effects after box-cox (b, f, g, j) or log transformation (c, h, m, n); and negative binomial GLM (l). Schematics created with BioRender. N, neutrophils; Mo, monocytes; TAM, tumor-associated macrophages; DC, dendritic cells.
Extended Data Fig. 8 Dual targeting of Tgfbr2 in CCR2+ Myeloid Cells and ICB induces tumor regression via T cell expansion and downregulation of immunosuppressive TAM signatures.
a, Schematic of the Ccr2CreERT2; Rosa26mTmG; Tgfbr2fl/fl mouse model. Ccr2CreERT2 mice were crossed with Tgfbr2fl/fl mice for inducible myeloid-specific Tgfbr2 deletion, then bred with Rosa26mTmG mice. Tamoxifen-inducible Cre excises mTomato, enabling mGFP expression. Triangles indicate loxP sites. b, Schematic of liver metastasis generation via intrasplenic AKTP MTO implantation in Ccr2CreERT2; Tgfbr2fl/fl vs. Ccr2CreERT2; Tgfbr2+/+ mice. Tamoxifen was administered days 12–17. c, Flow cytometry of GFP recombination in macrodissected liver metastases. Ccr2CreERT2; Tgfbr2+/+, n = 9; Ccr2CreERT2; Tgfbr2fl/fl, n = 7. Mean ± SEM. d, Tgfbr2 mRNA levels in TAMs sorted from liver metastases at day 18, normalized to Ppia. Ccr2CreERT2; Tgfbr2+/+, n = 7; Ccr2CreERT2; Tgfbr2fl/fl, n = 8. Mean ± SEM. e, Schematic of treatment with tamoxifen (days 12–17) and antibodies (IgG/αPD-L1, every 3 days, days 12–24) in mice with established liver metastases. f, g, Number of liver nodules (f) and CD3+ T cell densities (cells/mm2) (g) at endpoint (day 24). Mice per group: n = 7, 5, 5, and 5 mice from left to right. Mean ± SEM. h, Percentage of Ki67+ proliferating cells within CD8+ T cells. Mice per group: Ccr2CreERT2; Tgfbr2+/+, n = 5; Ccr2CreERT2; Tgfbr2fl/fl, n = 5. Mean ± SEM. i, j, Representative images of CD3 IHC (i) and IF for CD8 and Ki67 (j) used for quantification. Scale bars: CD3, 500 µm; CD8/Ki67, 50 µm. Arrowheads denote Ki67+CD8+ cells. k, Relative expression of selected genes in TAMs sorted from liver metastases at day 18, normalized to Ppia. Ccr2CreERT2; Tgfbr2+/+, n = 8; Ccr2CreERT2; Tgfbr2fl/fl, n = 7. Mean ± SEM. All tests were two-sided and corrected for multiple comparisons. Statistical analyses used linear models after box-cox transformation (c, d, k), linear model with random effects after log transformation (g), and Poisson GLM with log link (f). Schematics created with BioRender. B, B cells; T, T cells; N, neutrophils; Mo, monocytes; TAM, tumor-associated macrophages; DC, dendritic cells.
Extended Data Fig. 9 Differential enrichment of TAM-HR gene signature expression in primary CRC tumors and liver metastases.
a, UMAP of whole tumors (stroma + epithelial cells) from the Moorman dataset, annotated by major cell types. b, Proportions of cell populations in primary colorectal cancer (CRC) and matched chemotherapy-naive liver metastases (n = 9 pairs) in the Moorman dataset. c, UMAP of whole tumors split by primary CRC and liver metastasis in the Moorman dataset, colored by TAM-HR signature expression. d, UMAP of myeloid subpopulations in the Moorman dataset. e, Proportions of myeloid subpopulations comparing primary CRC and liver metastasis. f, Fold change of TAM-HR genes in macrophages from liver metastases versus colon tumors. g, UMAP of tumor microenvironment (TME) from macrodissected liver metastases in C57BL/6J mice after 3 days of treatment (Tx started day 14 post-MTO inoculation). Groups: vehicle (n = 4), αPD-L1 (n = 2), galunisertib (gal, n = 2), dual (n = 2). h, UMAP of TME showing Spp1 expression comparing vehicle and galunisertib-treated mice. i, UMAP of monocytes and TAMs annotated by cluster. j, UMAP of monocytes and TAMs showing Spp1 expression comparing control and galunisertib-treated mice (n as in g). k, UMAP of cancer-associated fibroblasts (CAFs) annotated by cluster (vehicle, αPD-L1, gal, dual; n = 2 mice each). l, UMAP of CAFs showing Spp1 expression comparing vehicle and galunisertib treatment.
Extended Data Fig. 10 Spp1 effects on CD8+ T cell proliferation in vivo and in vitro.
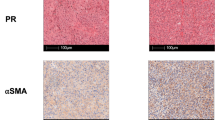
a, Schematic of in vivo generation of established liver metastases (LiMs) by injecting AKTP MTOs in spleens of Spp1−/− and wild-type (WT) mice. b, Representative immunofluorescence (IF) images of CD8 and Ki67 with respective quantification in (c). Scale bars: 50 μm. Mice per group: WT, n = 5; Spp1−/−, n = 5 mice. Mean ± SEM. Arrowheads indicate Ki67+CD8+ T cells. d, In vitro experimental design: CD3+ T cells were isolated from spleen and mesenteric lymph nodes of C57BL/6J mice by magnetic sorting, stained with CFSE, and cultured with αCD3, αCD28, IL-2, and indicated recombinant osteopontin (OPN) concentrations for 3 days. e, f, CFSE intensity histogram (e) and quantification of CD8+ T cell division (f) after 72 h with or without recombinant OPN (1, 2.5, 5 µg/ml). Representative of >3 independent experiments. g, Absolute numbers of major immune populations from liver metastases normalized to tissue weight at day 18 (schematics in a). Mice per group: WT, n = 6; Spp1−/−, n = 7 mice. Mean ± SEM. h, Contour plots and gating strategy for flow cytometry of liver metastases identifying major populations. Live leukocytes (CD45+) were distinguished from epithelial cells (EpCAM+), and EpCAM−CD45− cells subdivided into endothelial cells (CD31+) and cancer-associated fibroblasts (CAFs, PDGFRβ+). Quantification of PDGFRβ+ CAFs normalized to tissue weight in WT and Spp1−/− mice (WT, n = 6; Spp1−/−, n = 7). Mean ± SEM. i, Schematic of LiMs generated by AKTP MTOs in Spp1−/− and WT mice sacrificed at days 7, 14, 18, and 26. j, Representative IF images of α-SMA and E-cadherin staining over metastatic progression (quantification and animal number in Fig. 8k). This experiment used at least three animals per timepoint and was independently repeated three times. Scale bars: 50 μm. All statistical tests were two-sided with multiple comparison correction. Statistical analyses included mixed-effect linear models (c), linear models with random effects after Box-Cox transformation (g), and linear models (h). Schematics created with BioRender. B, B cells; T, T cells; N, neutrophils; Mo, monocytes; TAM, tumor-associated macrophages; DC, dendritic cells; OPN, osteopontin.
Supplementary information
Supplementary Tables 1–13 (download XLSX )
Supplementary Table 1. Signatures used in scRNA-seq analysis for T cells. Supplementary Table 2. Descriptives of clinical information for the metacohort. Supplementary Table 3. Genes predicting disease relapse (All-HR signature). Supplementary Table 4. TAM-HR signature. Supplementary Table 5. Effect of galunisertib on the TAM-HR gene expression profile. Supplementary Table 6. Effect of myeloid TGFBR2 ablation on the TAM-HR gene expression profile. Supplementary Table 7. Comparison of TAM-HR gene expression in TAMs from primary and metastatic human CRC tumors. Supplementary Table 8. Primers used for mouse genotyping. Supplementary Table 9. Antibodies for flow cytometry staining, immunohistochemistry and immunofluorescence staining. Supplementary Table 10. Primers used for RT–qPCR. Supplementary Table 11. Upregulated GSEA terms in Spp1+ TAMs (BioPlanet 2019). Supplementary Table 12. Downregulated GSEA terms in Spp1+ TAMs (BioPlanet 2019). Supplementary Table 13. Downregulated GSEA terms in C1q+ TAMs (BioPlanet 2019).
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 1.
Source Data Fig. 2 (download XLSX )
Statistical source data for Fig. 2.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 3.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 4.
Source Data Fig. 5 (download XLSX )
Statistical source data for Fig. 5.
Source Data Fig. 6
Statistical source data for Fig. 6.
Source Data Fig. 7 (download XLSX )
Statistical source data for Fig. 7.
Source Data Fig. 8 (download XLSX )
Statistical source data for Fig. 8.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 9 (download JPG )
Statistical source data for Extended Data Fig. 9.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data for Extended Data Fig. 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Henriques, A., Salvany-Celades, M., Nieto, P. et al. TGF-β builds a dual immune barrier in colorectal cancer by impairing T cell recruitment and instructing immunosuppressive SPP1+ macrophages. Nat Genet 57, 3050–3065 (2025). https://doi.org/10.1038/s41588-025-02380-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02380-2