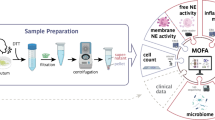
Abstract
Chronic obstructive pulmonary disease (COPD) is clinically and molecularly heterogeneous. To investigate COPD heterogeneity, we profiled lung tissue by single-nucleus RNA sequencing from 141 study participants (1,516,727 nuclei) and identified shifts in cell composition and emergent cell states that correlated with lung function, emphysema and composite symptom scores. Epithelial regenerative states peaked in early COPD and declined thereafter, whereas inflamed nonimmune cells and profibrotic/remodeling states, together with select immune populations, expanded with disease progression. Clustering study participants by the proportion of pathologic cells coupled with spatial transcriptomics identified distinct patterns of cellular co-occurrence within spatially localized niches. Proteomic analyses identified plasma biomarkers of cell states and their impact on the extracellular matrix. Mediation and cell communication analyses revealed cell-autonomous and intercellular communication networks associated with disease. These data define the cellular landscape of COPD heterogeneity, revealing molecular drivers and biomarkers that could inform therapeutic strategies.
Similar content being viewed by others
Main
Chronic obstructive pulmonary disease (COPD) is a leading cause of death in the United States1. Although defined by persistent airflow limitation, COPD presents with broad variation in the severity of obstruction, symptoms and exacerbation frequency2,3. Pathologic features, including chronic inflammation, tissue remodeling and emphysema, also vary widely and evolve over time, shaped by genetics and cumulative environmental exposures4. With few effective stratification tools, many studies have treated COPD as a single entity, limiting the discovery of mechanisms and targeted therapies. Despite the emergence of biomarkers such as polygenic risk scores and blood eosinophil counts, clinical characterization of COPD remains anchored in symptoms and spirometry. These coarse metrics fail to account for the underlying biological heterogeneity that shapes disease progression5,6,7.
Much of the field’s understanding derives from mouse and other reductionist models, which, while informative, do not capture the full cellular diversity or spatial organization of the human lung. COPD arises from a multicellular response to injury in which epithelial and endothelial damage triggers persistent inflammation and extracellular matrix (ECM) remodeling without effective repair4,8. These processes involve diverse structural and immune populations, yet the relative contributions of each, and their variation across individuals, remain unclear. Single-cell RNA sequencing of human lungs has begun to address this gap, identifying distinct cellular phenotypes, or ‘cell states’, in health and disease9,10,11, including regenerative states and those marked by metabolic dysregulation and impaired stress tolerance in COPD12,13,14,15. However, prior COPD studies often involved small, end-stage cohorts, sampling only part of the clinical spectrum and yielding fragmented insights loosely connected to established COPD pathobiology.
Here we address the challenge of deconvolving COPD heterogeneity by integrating single-nucleus RNA sequencing (snRNA-seq) with high-resolution spatial transcriptomics and paired lung proteomics and plasma proteomics. Nuclei isolation enabled sequencing of archived lung tissue from a large, well-phenotyped cohort, identifying canonical and disease-emergent cell states whose abundances correlate with airflow obstruction, symptom burden and/or emphysema severity. Unsupervised clustering of participants based on changes in cell and cell-state abundances revealed reproducible patterns of co-occurring cells that defined distinct cellular communities, such as inflammation-enriched and repair/remodeling-enriched groups. Spatial transcriptomics demonstrated that these aberrant populations also colocalize within shared tissue niches, and pathway analyses identified cell-autonomous and intercellular signaling programs shaping these spatially organized communities. Lung ECM proteomics linked specific aberrant cells to matrix remodeling, while plasma proteomics detected systemic biomarkers of these cell states. Together, these multi-omic data provide a spatially resolved portrait of COPD that links clinical heterogeneity to distinct pathogenic microenvironments and underlying molecular pathways.
Results
Patient cohorts and study design
We performed snRNA-seq on lung tissue from the Lung Transplant Research Consortium (LTRC), which includes 2,213 study participants (never-smokers, asymptomatic current/former smokers and individuals with COPD; pulmonary fibrosis excluded). From this cohort, we profiled 146 lung lobes obtained from 141 study participants, generating 1,516,727 high-quality cell transcriptomes. The median age was 63 years (interquartile range (IQR) = 10.5), smoking exposure 40 pack-years (IQR = 40) and years since quitting 5.7 (IQR = 9.8); 53.9% were female; median body mass index (BMI) was 26.07 kg m−2 (IQR = 7.12); 13 were never-smokers and 11 were current smokers. Clinical traits (Table 1) included the percentage predicted forced expiratory volume in 1 s (FEV1), the ratio FEV1/forced vital capacity (FEV1/FVC), diffusing capacity of the lung for carbon monoxide (DLCO) and Global Initiative for Chronic Obstructive Lung Disease (GOLD) stage. Emphysema burden was quantified in the sampled lobe by computed tomography (CT; % voxels Hounsfield units (HU) of ≤950) and by semiquantitative radiologist scoring. Symptoms were assessed using multiple questionnaires, including the St. George’s Respiratory Questionnaire (SGRQ). Composite symptom scores for dyspnea, cough, infection, wheezing and exacerbations were derived using principal component analysis (PCA) of responses to questionnaires across the full LTRC, anchoring traits to a larger and more representative population (Supplementary Fig. 1a–e).
Unsupervised clustering of snRNA-seq data resolved major cell types, annotated by automated label transfer and refined with marker genes from the human lung cell atlas and LungMAP (Extended Data Fig. 1 and Supplementary Table 1)9,16. Endothelial subsets included the following: arterial (GJA5, DKK2), venous (ACKR1), systemic/bronchial (COL15A1), lymphatic (PROX1, LYVE1) and two capillary subsets—gas-exchange aerocytes (HPGD, EDNRB) and general capillaries (FCN3, IL7R)17. Alveolar epithelial clusters comprised alveolar type 2 (AT2) cells (SFTPC, LAMP3) and alveolar type 1 (AT1) cells (SCEL, RTKN2). Airway epithelium segregated into secretory/club (SCGB1A1, SCGB3A2), goblet (MUC5B, SPDEF), ciliated (CFAP47, DNAH9) and basal cells (TRPC6, TP63). The mesenchyme featured alveolar fibroblasts (COL13A1, PDGFRA, NPNT) and adventitial fibroblasts (COL14A1, MFAP5, TWIST2)18. All major immune populations were represented, including alveolar macrophages (AM; SLC11A, INHBA), B cells (MS4A1) and T cells (THEMIS, ITK).
We hypothesized that shifts in canonical cell-type proportions and the emergence of discrete disease-associated cell states underlie COPD heterogeneity. We quantified cell and cell-state proportions within parent lineages (Supplementary Table 2) and correlated them with clinical metrics, accounting for compositional constraints and adjusting for age, sex, smoking status and BMI19. Analyses were performed on an individual basis, except for emphysema, which was assessed per lobe, with an additional adjustment for anatomical location. Only samples with ≥10 cells in the parent population (or ≥10 nonepithelial alveolar cells when the parent population included alveolar epithelial cells) were included to ensure robust proportion estimates. Quality-control (QC) thresholds are shown in Supplementary Figs. 2–4. Findings were validated or extended using four approaches on matched tissue—Xenium spatial transcriptomics (2-mm2 tissue microarrays, 38 matched formalin-fixed paraffin-embedded (FFPE) samples, 99 cores), immunofluorescence staining, label-free ECM proteomics (72 matched samples) and plasma proteomics (Olink Explore HT, 64 matched samples)—and a fifth approach in an independent Baylor cohort (n = 37; Supplementary Table 3) using GeoMx spatial transcriptomic deconvolution20. QC metrics for both spatial datasets are shown in Supplementary Figs. 5 and 6.
Inflamed nonimmune cell states in COPD
Beyond canonical cell identities, we identified disease-associated cell states within nonimmune lineages that resolved into the two following broad transcriptional archetypes: inflamed nonimmune cell states and repair/remodeling cell states. Below, we first define the transcriptional features of these cell states and validate their expansion in COPD. We then assess their relationships with clinical features before applying a similar analytic framework to immune cells. Finally, we integrate clinical, multi-omic and spatial data to construct a unified framework that connects these cell states to each other and disease.
Inflamed nonimmune cell states were identified across endothelial, epithelial and fibroblast lineages. Inflamed endothelial states (arteriali, gCapi and aerocyteᵢ) showed increased expression of inflammatory mediators (NFKB1, IRF1, IL6, IL32, CXCL10, CSF3) and enrichment for tumor necrosis factor (TNF), interferon-γ (IFNγ) and interleukin-1 (IL-1) signaling (Fig. 1a–c). Similar pathways were upregulated in inflamed epithelial and fibroblast states (Fig. 1c). Inflamed epithelial states included AT2i, AT1i and secretoryi cells, marked by elevated SGPP2, CSF3 and IL4R in AT2i, IL32 and IRF1 in AT1i and CXCL1/CXCL3/CXCL8 in secretoryi (Fig. 1d,e and Extended Data Fig. 2a). Inflamed fibroblast states (alveolar fibroblasti and adventitial fibroblasti) expressed NF-κB subunits, IRF1, PLAUR, ADAMTS4, VEGF and CD44, suggesting roles in angiogenesis and tissue remodeling in addition to inflammation (Fig. 1f–h)21,22. Immunofluorescence staining of matched FFPE samples showed colocalization of NF-κB and/or SOD2 with endothelial, alveolar epithelial and secretory markers in samples with higher proportions of inflamed cells (Fig. 1i–k, Extended Data Fig. 2b–e and Supplementary Fig. 7). Spatial deconvolution in the independent Baylor cohort similarly revealed increased inflamed nonimmune cells in tissue from study participants with more severe airflow obstruction and emphysema (Fig. 1l–n and Extended Data Fig. 2f–h).
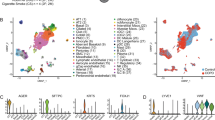
a, UMAP of endothelial cells (left) and cell states (right), including artery (n = 2,891), vein (n = 2,171), systemic circulation endothelial cells (systemic, n = 1,523), lymphatic (n = 7,925), cycling endothelial cells (Endodiv, n = 127), gCap (n = 9,131) and aerocytes (n = 8,001). Cell states including control (aerocytec, gCapc, arteryc) and inflamed (aerocytei, gCapi, arteryi) endothelial cells and angiogenic tip cells (gCaptip). b, Dot plot of endothelial marker genes. Size represents the proportion of expressing cells; color denotes scaled expression. c, Pathway enrichment analysis across inflamed cell states; x axis shows NES, dot size shows −log10(P) and color shows cell state; only pathways with FDR < 0.05 are shown (Benjamini–Hochberg, two-sided permutation tests). d, UMAP of epithelial populations, including goblet (n = 9,544), basal (n = 2,570), ABCs (n = 5,001), SCGB1A1+ secretory (n = 8,655), SCGB3A2+ secretory (n = 25,704) and AT2 subtypes—AT2div (n = 1,326), AT2SCGB3A2 (n = 23,375), AT2i (n = 148,699), AT2b (n = 330,033) and AT2s (n = 98,751). e, Dot plot of epithelial marker genes. Size represents proportion of expressing cells; color denotes scaled expression. f, UMAP of fibroblast subsets, including alv. (n = 36,968), adv. (n = 6,294), CTHRC1+ fibroblasts (n = 2,734), IR (n = 739), FRC (n = 524), PB (n = 953) and myofibroblasts (n = 1,057). Alv. and adv. fibroblasts further stratified into control (fibroblastc) and inflamed (fibroblasti) cell states. g, Dot plot of fibroblast marker genes. Size represents proportion of expressing cells; color denotes scaled expression. h, Heatmap of normalized gene expression in adv. and adv. fibroblasts, clustered by cell state and study participant. i–k, Immunofluorescence of NF-κB (red) and DAPI (blue), with (i) PECAM1 (green) in inflamed endothelial-enriched tissue (j), pro-SFTPC (pSFTPC; green) in AT2i-rich tissue and (k) AGER (green) in AT1i-rich tissue; scale bars = 100 μm. l–n, Proportion of inflamed cell states across GOLD stages identified after deconvolution of spatial transcriptomic data in the Baylor cohort—(l) gCapi, (m) AT2i and (n) alv. fibroblasti (no COPD, n = 11; GOLD stage I/II, n = 10; GOLD stage III/IV, n = 12). o, Feature plots of scaled LAMP3 and SCGB3A2 expression in epithelial cells. p, Immunofluorescence of KRT17 (red), CPA6 (green) and DAPI (blue); scale bar = 100 μm. q, Immunofluorescence of CTHRC1 (red), KRT17 (green) and DAPI (blue); scale bar = 50 μm. r, Proportion of CTHRC1+ fibroblasts (proportion of all fibroblasts) across GOLD stages in the Baylor cohort identified after deconvolution of spatial transcriptomic data (no COPD, n = 11; GOLD stage I/II, n = 10; GOLD stage III/IV, n = 12). All images representative of five participants per group (i–k,p,q). Group differences tested by Kruskal–Wallis with two-sided Wilcoxon post hoc tests and Benjamini–Hochberg FDR correction (l–n,r). Box plots depict the median (centerline), IQR (box) and 1.5× IQR whiskers. UMAP, Uniform Manifold Approximation and Projection; NES, normalized enrichment score; IFNγ, interferon-γ; gCap, general capillaries; alv, alveolar; adv, adventitial; IR, immune reticular; PB, peribronchial; expr, expression.
Reparative and remodeling cell states in COPD
We identified reparative and remodeling states across epithelial and endothelial lineages. Secretory/club cells comprised the following three subsets: proximal SCGB1A1+SCGB3A1+ cells; SCGB3A2+ cells residing distally with capacity to differentiate into alveolar epithelium; and proliferating secretory cells (Extended Data Fig. 3a,b)12,13,14. AT2 cells resolved into the following five populations (Fig. 1c–e): (1) homeostatic AT2b enriched for surfactant genes, HHIP and LAMP3; (2) inflamed AT2i; (3) AT2S with a transcriptional program suggesting AT2-to-AT1 transition potential (TEAD1, TEAD4, YAP1) and enrichment for adhesion, migration and wound-healing pathways23; (4) proliferative AT2div and (5) AT2SCGB3A2 coexpressing SCGB3A2 and canonical AT2 markers (Fig. 1o)12. Reparative endothelial populations included angiogenic gCaptip cells (gCaptip; KCNE3, ANGPT2) and proliferating endothelium (Endodiv), consistent with vascular repair (Fig. 1a,b)24,25,26.
We also identified CTHRC1+ fibroblasts, defined by CTHRC1 and high ECM gene expression (for example, COL1A1) and aberrant basaloid cells (ABCs) expressing KRT17, MMP7, GDF15 and CDKN1A but not KRT5, distinguishing them from basal cells11,27,28 (Fig. 1d–g). These cell states, previously associated with idiopathic pulmonary fibrosis (IPF) and other fibrotic lung diseases, were detected in samples from lobes without radiographic evidence of fibrosis and immunofluorescence costaining confirmed their colocalization in remodeling areas without fibroblastic foci (Fig. 1p,q and Extended Data Fig. 3c,d). CTHRC1+ fibroblasts were also increased in participants with greater disease severity in the Baylor cohort (Fig. 1r and Extended Data Fig. 3e). Whether the population we termed ABCs in COPD are transient intermediate cell states or persist as in IPF remains unknown29, but their co-occurrence with CTHRC1+ fibroblasts in COPD suggests convergent processes across diseases.
Loss of alveolar gas exchange and increased goblet cells in COPD
We then assessed how epithelial and endothelial composition varied with disease severity (Supplementary Table 4). Aerocytes were reduced in GOLD stage IV disease (false discovery rate (FDR) = 0.014) and in lungs with >75% emphysema (FDR = 0.031), and were inversely correlated with lobe-specific emphysema. In contrast, nonpulmonary endothelial populations were increased; systemic endothelial cells of the bronchial circulation were increased with >75% emphysema (FDR = 0.046) and correlated positively with lobe-specific emphysema, and lymphatic endothelial cells were increased in GOLD stage IV disease (FDR = 0.069; Fig. 2a–c). Among alveolar epithelial cells, AT1 cells were depleted in GOLD stage I/II and GOLD stage IV (FDR = 0.01, 0.02) and in >75% emphysema (FDR = 0.008), with abundance positively correlating with DLCO (Fig. 2c–e). Airway remodeling was also evident—goblet cells expanded in former and current smokers (FDR = 0.018, 0.017) and in GOLD stage II disease (FDR = 0.065; Fig. 2f), ciliated cells declined with falling FEV1 and SCGB3A2+ secretory cells decreased with increasing emphysema (Fig. 2c and Extended Data Fig. 3f,g). These findings outline a cellular trajectory in COPD consistent with goblet-cell hyperplasia and loss of ciliated cells, SCGB3A2+ secretory cells and cells involved in gas exchange (aerocytes, AT1)30,31,32.
a, Alluvial plots of median endothelial cell proportions stratified by semiquantitative lobe emphysema (n = 128) and GOLD stage (n = 123). b, Lobe emphysema (% attenuation of ≤950 HU) and proportion of aerocytes (proportion of endothelial cells; n = 101). c, Clinical traits and cell proportions; aerocytes and systemic endothelial cells (proportion of endothelial cells; n = 101 for lobe emphysema; n = 123 for other traits), AT1 cells (proportion of alveolar epithelial cells; n = 121), ciliated cells (proportion of airway epithelial cells; n = 137) and SCGB3A2+ secretory cells (proportion of secretory/club cells; n = 108). Colors denote cell states, dot size reflects enrichment and x axis shows partial Spearman correlation coefficient. d, Alluvial plots of median alveolar epithelial cell proportions stratified by semiquantitative lobe emphysema (n = 127) and GOLD stage (n = 122). e, AT1 proportion (among alveolar epithelial cells) and DLCO (% predicted; n = 114). f, Goblet cells (proportion of airway epithelial cells) stratified by smoking status (N, F and C) and GOLD stage (n = 137). g, Proportions of inflamed nonimmune cell states relative to their respective parent lineages, stratified by GOLD stage and semiquantitative lobe emphysema—aerocytei (n = 87,90), arteryi (n = 71,71), gCapi (n = 91,94), AT1i (n = 122,127), AT2i (n = 122,127), secretoryi (n = 137,142), alveolar fibroblasti (n = 117,122) and adventitial fibroblasti (n = 101,103). h, Clinical traits and inflamed nonimmune cell-state proportions relative to their parent populations. Colors denote cell states, dot size reflects enrichment and x axis shows partial Spearman correlation coefficient. i,j, Reparative epithelial populations (AT2div, AT2S, secretorydiv, AT2SCGB3A2; proportion of all cells) stratified by smoking status (n = 127; i) and GOLD stage (n = 127; j). k, Clinical traits and reparative epithelial cell states (AT2div, AT2S, secretorydiv, AT2SCGB3A2; proportion of all cells), reparative endothelial cells (Endodiv, gCaptip; proportion of all endothelial cells), PB fibroblasts (fibroblastPB; proportion of all fibroblasts) and myofibroblasts (proportion of all fibroblasts). Colors denote cell states, dot size reflects enrichment and x axis shows partial Spearman correlation coefficient. l,m, ABCs (l) and CTHRC1+ fibroblasts (m; proportion of all cells) stratified by GOLD stage (n = 127). n,o, FibroblastPB (n) and myofibroblasts (o) as a proportion of all fibroblasts stratified by GOLD stage (n = 127). Proportions (range 0–1) were multiplied by 100 for visualization and plotted on a log10(scale) (i,j,l,m). Differential composition by sccomp (Bayesian sum-constrained β binomial), adjusted for age, sex, BMI and smoking; emphysema analyses additionally adjusted for lobe (a–o). We report posterior discovery probabilities, FDR-adjusted (Benjamini–Hochberg). *FDR < 0.10, **FDR < 0.05, ***FDR < 0.01. Partial Spearman correlations between continuous clinical traits and cell-state proportions, adjusted for the same covariates, illustrate the direction and strength of associations (f,i,j,l,m–o). Box plots depict the median (centerline), IQR (box) and 1.5× IQR whiskers. N, never; F, former; C, current smokers; emph., emphysema; Sec, secretory.
Inflammatory, reparative and fibrotic cell-state dynamics
Inflammatory nonimmune populations were increased in COPD and expanded progressively with advancing GOLD stage and greater lobe-specific emphysema (Fig. 2g). Their proportion correlated positively with airflow limitation, DLCO, emphysema and symptom scores (Fig. 2h). Notably, their abundance varied widely among individuals with severe disease (Extended Data Fig. 3h), highlighting interindividual heterogeneity in inflamed nonimmune cells. No associations were observed with pack-years or quit-years, suggesting a cause for persistent inflammation in former smokers despite tobacco cessation.
Reparative epithelial states (AT2S, AT2SCGB3A2, AT2div, secretorydiv) were enriched in current and former smokers compared with never-smokers (Fig. 2i), consistent with their emergence after injury. These populations peaked in early COPD but declined with disease progression (Fig. 2j), and AT2div and AT2SCGB3A2 abundance inversely correlated with multiple measures of disease severity (Fig. 2k). Reparative endothelial states (Endodiv, gCaptip) were also reduced in advanced disease, and both inversely correlated with disease severity (Fig. 2k and Extended Data Fig. 3i,j). In contrast, ABCs and CTHRC1+ fibroblasts expanded with advancing GOLD stages and their proportions correlated with multiple disease metrics (Fig. 2k–m and Extended Data Fig. 3k,l). Together, these data point to a shift from early repair to persistent inflammatory and profibrotic responses with advancing disease.
The following two additional fibroblast subsets were identified: peribronchial fibroblasts (fibroblastPB; LGR5, ENTPD1) and myofibroblasts (WNT5A, ASPN, ACTA2, MYH11; Fig. 1f,g)28,33. They decreased with increasing GOLD stage and emphysema involvement, with loss correlating with disease severity (Fig. 2k,n,o). The loss of these regionally localized fibroblast populations may limit tissue-specific repair capacity or simply reflect tissue destruction in advanced disease.
Immune cells and lymphoid fibroblasts
We then profiled immune cells and their contributions to COPD progression. We identified AM, monocyte-derived macrophages, monocytes and two interstitial macrophage (IM) subsets—IM1, defined by canonical IM markers (STAB1, F13A1) and IMCHIT1 expressing profibrotic genes (CHIT1, CHI3L1, PLA2G7, GPNMB, MMP9, SPP1; Fig. 3a,b)34,35. Overall, IMs were increased in COPD, driven primarily by the profibrotic IMCHIT1 subset, which showed the strongest associations with severe airflow obstruction, high emphysema burden, smoking and other clinical metrics (Fig. 3c–e) and was validated by immunofluorescence staining in COPD lung tissue (Extended Data Fig. 4a and Supplementary Fig. 8). IM1 also contributed modestly, with enrichment in GOLD stage IV disease (Extended Data Fig. 4b). Several myeloid cell states were also associated with disease. We identified inflammatory AM states (AMCSF1, AMNR4A and AMinflam) demonstrating heightened cytokine and chemokine signaling, with each state variably linked to emphysema, GOLD stage and exacerbations (Fig. 3d–i and Extended Data Fig. 4c). Additional macrophage populations included AMp21 characterized by high expression of DNA damage response genes (TP53, CDKN1A); AMHSP enriched in heat shock protein genes; and AMIFIT expressed interferon-stimulated genes. AMp21 was increased in smokers and in study participants with <50% emphysema (Fig. 3j,k). Monocyte-derived macrophages were elevated in smokers and samples from lungs with <50% emphysema, while proliferative macrophages declined with advanced disease (Fig. 3l,m and Extended Data Fig. 4d,e). B cells and plasma cells were also increased in COPD across multiple contexts, whereas CD8+ T cells were positively correlated with pack-years. In contrast, mast cells and migratory dendritic cells (CCR7, FSCN1) were reduced in advanced disease (Fig. 3d and Extended Data Fig. 4f–j). We also identified two VCAM+ fibroblast populations associated with lymphoid structures—CXCL13+ fibroblast reticular cells (FRCs), specialized stromal important for germinal centers and B cell recruitment/antigen presentation36,37, and immune-regulating FRCs (IR fibroblasts) with elevated expression of IRF1, CCL2, CXCL10, CCL19 and IL33, implicating them in T-cell immunoregulation (Fig. 1f,g)38,39,40. FRCs and IR fibroblasts were increased with advanced GOLD stages and radiographic emphysema involvement (Fig. 3d,n,o).
a, UMAP projection of macrophage and monocyte cells (left) and cell states (right) including AM (n = 163,726), IM (n = 9,037), MDM (n = 8,325) and monocytes (n = 1,686). b, Dot plot of select marker genes. Size reflects the proportion of expressing cells; color intensity denotes normalized expression. c, IMCHIT1 as a proportion of all macrophages stratified by smoking status (N, F and C; n = 140), GOLD stage (n = 140) and semiquantitative lobe-specific emphysema (n = 127). d, Clinical traits and immune cell states expressed as a proportion of their parent populations. Colors denote cell states, dot size reflects enrichment (calculated by sccomp) and the x axis shows the partial Spearman correlation coefficient. Macrophage expressed as a proportion of total macrophages, other immune cells as a proportion of total cells. e, Pathway enrichment analysis in inflammatory macrophage populations (AMCSF, AMNR4A and AMinflam). The x axis represents NES; dot size corresponds to −log10(P); color indicates cell state. Significance assessed using a two-sided permutation test; FDR was controlled using the Benjamini–Hochberg correction. f, AMCSF expressed as a proportion of all macrophages (stratified by GOLD stage; n = 140). g,h, AMCSF (g) and AMinflam (h) stratified by semiquantitative lobe-specific emphysema; n = 127) and expressed as a proportion of all cells. i, AMNR4A as a proportion of macrophages and lobe emphysema (% attenuation of ≤950 HU; n = 111). j–m, AMp21 (j,k), MDM (l) and Macdiv (m) expressed as a proportion of all cells (stratified by semiquantitative lobe-specific emphysema; n = 127) or as a proportion of all macrophages (stratified by GOLD or smoking status; n = 140). n,o, IR (fibroblastIR) (n) and FRC (fibroblastFRC) fibroblasts (o) expressed as a proportion of all cells, stratified by GOLD stages (n = 122) and lobe-specific radiographic emphysema (n = 127). Proportions (range 0–1) were multiplied by 100 for visualization and plotted on a log10(scale) (c,j,k,m,n,o). Differential composition by sccomp (Bayesian sum-constrained β binomial), adjusted for age, sex, BMI and smoking; emphysema analyses additionally adjusted for lobe (c,d,f–o). We report posterior discovery probabilities, FDR-adjusted (Benjamini–Hochberg). *FDR < 0.10, **FDR < 0.05, ***FDR < 0.01. Partial Spearman correlations between continuous clinical traits and cell-state proportions, adjusted for the same covariates, illustrate the direction and strength of associations (c,f–h,j–o). Box plots depict the median (centerline), IQR (box) and 1.5× IQR whiskers. MDM, monocyte-derived macrophages; Exacerb., exacerbation.
Cell composition across composite phenotypes
Single-trait analyses do not capture the relationships between cell populations and composite COPD phenotypes. We therefore clustered 110 categorical and 24 continuous variables from the LTRC cohort (spirometry, radiographic emphysema, smoking history, symptoms and functional capacity; Supplementary Table 5). We identified seven clinical clusters (Fig. 4a) representing composite COPD phenotypes and projected these phenotypes onto snRNA-seq data. The seven clusters mapped to readily identifiable clinical phenotypes spanning symptom burden and airflow impairment—cluster 7 (never-smokers and/or normal spirometry); clusters 1 and 6 (mild obstruction); clusters 2 and 5 (moderate obstruction); and clusters 3 and 4 (severe obstruction with substantial emphysema; Fig. 4b). Notably, clusters 2 and 4 had the greatest symptom burden. All clusters were represented in the snRNA-seq dataset, with proportions of 12.8% (cluster 1), 5.0% (cluster 2), 24.8% (cluster 3), 29.1% (cluster 4), 9.9% (cluster 5), 7.8% (cluster 6) and 11.3% (cluster 7). For downstream analysis, we merged clusters 1 + 6 and 2 + 5 to yield five categories that balanced phenotypic similarity with sample size (Supplementary Table 6) and, notably, preserved a distinction between high and low symptom burden among participants with severe disease. Using this composite-phenotype framework, we observed directional shifts in cell composition that were similar to those observed in single-trait analyses. For example, certain cell populations increased with advanced disease, including CTHRC1+ fibroblasts and IMCHIT1 macrophages, while other populations decreased with advanced disease, including AT1 cells and aerocytes (Fig. 4c,d). The key distinction revealed by the composite analysis was the relative specificity of the inflamed nonimmune cells to be selectively elevated in study participants with both severe obstruction and high symptom burden, whereas other cell populations tracked with physiologic impairment irrespective of symptoms.
a, Heatmap of normalized clinical trait values for 2,213 study participants grouped into seven clinical clusters. Clustering used 110 categorical and 24 continuous variables from the full LTRC cohort. Rows denote clinical trait clusters and columns denote selected traits; color represents the scaled values of each trait. b, Box plots of selected clinical variables. Boxes colored by LTRC clinical clusters identified in Fig. 4a. Box plots depict the median (centerline), IQR (box) and 1.5× IQR whiskers. c,d, Median-centered FC plots across LTRC clinical clusters. Clusters are labeled according to their predominant clinical features—NS/NS (cluster 7), mild obstruction (clusters 1 and 6), moderate obstruction (clusters 2 and 5), severe obstruction with LS (cluster 3) and severe obstruction with HS (cluster 4). Points represent the median FC in cell proportion increased (c) or decreased (d) relative to cluster 7 (NS/NS); error bars denote the IQR, and color denotes cell type. All proportions are relative to their level 3 parent lineages—endothelial (n = 122), alveolar epithelial (n = 122), airway epithelial (n = 137), fibroblast (n = 127) and macrophage (n = 140), except fibroblastIR and fibroblastFRC, which are expressed as a proportion of all cells (n = 122). e, Heatmap of the similarity matrix showing pairwise Spearman correlation coefficients between sample-level proportions of aberrant cell states relative to their parent populations. Cell states are grouped into four categories—inflamed nonimmune (red), remodeling (blue), tissue change (green) and immune (purple). Correlations are shown on a color scale from −1 (blue) to 1 (red). An asterisk denotes populations that decrease in disease; for these states, the correlation sign was inverted. f, Dot plot of median cell proportions after clustering samples into five cellular communities. Dot color reflects the cell-state groupings defined in Fig. 4e. Dot size and color intensity represent the median proportion z score. g, Distribution of clinical traits grouped and colored by the cellular communities identified in f. Communities are named for their enriched populations—community 1 = ‘mixed inflamed/remodeling’, community 2 = ‘healthy’, community 3 = ‘goblet/AMNR4A3’, community 4 = ‘immune’ and community 5 = ‘inflammatory’. Box plots depict the median (centerline), IQR (box) and 1.5× IQR whiskers. Statistical significance was first assessed using the Kruskal–Wallis test; traits with FDR < 0.05 were further evaluated using pairwise Dunn’s tests with Benjamini–Hochberg correction (c,d,g). Dot size (c,d) and symbols (g) indicate significant pairwise differences, with an asterisk (*) indicating comparison to community 2 and a hash (#) indicating comparison to community 3. Significance thresholds are as follows: *, # FDR < 0.05; **, ## FDR < 0.01; ***, ### FDR < 0.001. NS/NS, never-smoker/normal spirometry; LS, low symptoms; HS, high symptoms.
Co-occurrence of disease-associated cell states
We then postulated that disease-associated cells co-occur in discrete patterns independent of clinical classification. To investigate this, we generated a similarity matrix of proportional cell-type abundances across samples and applied spectral clustering to delineate communities of co-occurring cell states (Fig. 4e). Inflamed nonimmune populations exhibited strong intercorrelations and were positively associated with inflamed macrophage subsets. CTHRC1+ fibroblasts and ABCs correlated strongly with one another and with reparative and inflammatory subsets, consistent with mixed remodeling–inflammation programs. Adaptive immune populations were tightly correlated with each other and with IMCHIT1 macrophages. Clustering samples by aberrant cellular composition identified five discrete communities (Fig. 4f). Community 1 (‘mixed-inflammation/fibrosis’) enriched for inflammatory and remodeling populations; community 2 (‘healthy’) contained low abundances of disease-associated states; community 3 enriched for goblet cells and AMNR4A macrophages; community 4 (‘immune’) enriched for CD4+, CD8+, B cells and IMCHIT1; and community 5 (‘high-inflammatory’) highly enriched for inflamed nonimmune and inflammatory macrophage populations. Community 2 had the highest DLCO, FEV1 and FEV1/FVC; communities 1 and 5 had the lowest DLCO and FEV1 and the highest SGRQ-symptom scores, with community 5 showing the greatest wheezing and exacerbations (Fig. 4g). These data point to separable, biology-defined disease programs with overlapping but nonidentical clinical manifestations in COPD.
Spatially resolved cellular neighborhoods
We then sought to validate these co-occurrence patterns and determine whether these cells are organized within spatially resolved neighborhoods. Therefore, we applied the Xenium high-resolution spatial platform to profile 480 genes encompassing canonical lung cells, COPD-associated states and signaling mediators (Supplementary Table 7). Profiling included ≥2 cores per sample from 3 never-smokers, 3 former smokers with normal spirometry, 6 study participants with GOLD stage I/II, 8 with GOLD stage III and 18 with GOLD stage IV (Supplementary Table 8). Unsupervised clustering delineated canonical cells and COPD-associated states consistent with those defined by snRNA-seq, and their relative abundances closely mirrored paired snRNA-seq measurements (Fig. 5a and Extended Data Fig. 5a–f).
a, Comparison of aberrant cell proportions across snRNA-seq and Xenium modalities. Cell states were quantified as the fraction of each aberrant population relative to its parent lineage in both snRNA-seq and Xenium for matched participants (n = 38). Samples were ordered by the snRNA-seq-derived fraction and divided into quartiles (Q1–Q4, x axis). The y axis displays the corresponding Xenium-derived proportions for the same participants. Shown are inflamed AT2i per AT2 cells, inflamed AT1i per AT1 cells, inflamed fibroblasts (fibroi) per all fibroblasts, inflamed gCapi per gCap cells, inflamed macrophages (Mi) per macrophages, IMCHIT1 per macrophages, ABC per epithelial cells and CTHRC1+ fibroblasts per all fibroblasts. Box plots show the median, IQR and 1.5× IQR whiskers. Pairwise differences were assessed using two-sided Wilcoxon rank-sum tests with Benjamini–Hochberg correction. *FDR < 0.10, **FDR < 0.05, ***FDR < 0.01. b, Dot plot summarizing the distribution of cell types across spatially defined niches. Columns correspond to cell types and rows to niches, defined by their enriched populations—airway, large artery, low-immune parenchyma 1 and 2 (LI-parenchyma 1, LI-parenchyma 2), AM, inflamed, remodeling, CHIT1/CD4/CD8 and B cell niches. Dot size reflects the relative proportion of each cell type within each niche, and dot color indicates niche identity. c, Stacked barplots showing the proportion of cells assigned to each niche across disease severity groups, including NS/NS, GOLD stage I/II, GOLD stage III and GOLD stage IV. Colors correspond to niche identities as defined in b. d–f, Spatial localization of canonical and disease-associated cell types, cellular neighborhoods and gene expression across representative ROIs. Top, cellular niche maps based on cell-type classification and location. Each color denotes a distinct population. ROIs include (d) two distinct LI parenchymal and inflamed niches, (e) remodeling niches and (f) B cell and CHIT1/CD4/CD8 niches. Middle, cell-type overlay showing spatial proximity of cell populations. Cell types labeled by color. Bottom, spatial gene expression maps highlighting key inflammatory, remodeling and immune transcripts. Each dot represents a detected transcript labeled by color. Dot size and color density are scaled for visualization and reflect relative, not absolute transcript abundance. Scale bars = 200 μm (top), 100 μm or 50 μm (middle) and 100 μm or 50 μm (bottom).
To delineate spatial microenvironments, we constructed k-nearest neighbor (k-NN) graphs using Euclidean distances among cells, defining niches based on physical colocalization (not transcriptomic similarity). These included airway and large-artery niches and two parenchymal niches with low inflamed nonimmune cells (LI-parenchyma 1 and 2; Fig. 5b). Additional niches recapitulated cellular communities described above—an inflamed nonimmune niche; a remodeling niche enriched for CTHRC1+ fibroblasts and ABCs; an alveolar and inflamed macrophage niche; and two immune-rich niches containing IMCHIT1 macrophages, T cells and B cells. LI-parenchymal niches were predominant in never-smokers and individuals with preserved lung function, whereas increasing COPD severity was associated with expansion of inflammatory, remodeling and immune niches (Fig. 5c and Extended Data Fig. 6). Hematoxylin and eosin overlays of cells and transcripts or clustering transcripts independent of cell boundaries confirmed that these niches were enriched for inflamed nonimmune cells with elevated expression of inflammatory transcripts that were frequently adjacent to macrophage-rich niches expressing IL1B, IL1A, IL10RA and CXCL5 (Fig. 5d, Extended Data Fig. 7a–d and Supplementary Fig. 9). In remodeling niches, CTHRC1+ fibroblasts were often adjacent to ABCs, which displayed a distinct elongated morphology with characteristic gene expression (Fig. 5e)41, and commonly neighbored immune-rich B cell and CD4/CD8/IMCHIT1 (Fig. 5f). These data demonstrate that aberrant cells in COPD colocalize within spatially defined neighborhoods, providing a framework for understanding how coordinated cell–cell interactions drive tissue-level pathology.
ECM proteomics links aberrant cell states to remodeling
We then hypothesized that these aberrant cell states influence ECM composition in distinct ways. We performed proteomic profiling of matched decellularized lung tissue, clustering proteins into modules based on their co-occurrence across samples and correlated these ECM modules with aberrant cell states and clinical phenotypes (Fig. 6a). Specific ECM modules were strongly linked to aberrant populations—high collagen-producing CTHRC1+ fibroblasts and inflamed nonimmune cells correlated positively with module ME8 (fibrillar collagens I, III, V) and negatively with modules ME22 (keratins) and ME9 (laminins, type IV collagens). Inflamed nonimmune cells positively correlated with ME5 enriched for inflammatory signaling and neutrophil degranulation, and inversely correlated with ME12 (SERPINA1, inhibitors of complement/coagulation), suggesting a bidirectional relationship in which reduced antiprotease activity may allow excessive injury and inflammation, while inflamed microenvironments may promote SERPINA1 depletion. IMs, particularly IMCHIT1, positively correlated with modules enriched for COPD-related proteases (MMP12, cathepsins), implicating IMs as a source of these enzymes. Full protein lists, correlations and module memberships are provided in Supplementary Table 9a,b. PCA of ECM proteomes independently segregated samples into the same high-severity communities identified by snRNA-seq (community 1, ‘mixed-inflammation/fibrosis’; community 5, ‘high-inflammatory’), underscoring the critical relationship between these microenvironments and ECM remodeling proteins (Fig. 6b).
a, Heatmap showing Spearman correlations between modules of ECM proteins, detected by label-free MS, and clinical traits or aberrant cell populations. Modules were defined using WGCNA. Correlations are represented using a color scale from −1 (blue) to 1 (red). Only values with Benjamini–Hochberg-adjusted (FDR) Spearman correlation of <0.05 are displayed. Representative proteins of highlighted modules are shown. b, PCA plot of ECM protein profiles, with each sample colored by snRNA-seq-defined cellular communities (Fig. 4f,g). Axes represent the first two PCs. c, Heatmap showing Spearman correlations between modules of Olink-detected plasma proteins and clinical traits or aberrant cell populations. Modules defined using WGCNA. Correlations are represented using a color scale from −1 (blue) to 1 (red). Only values with Benjamini–Hochberg-adjusted (FDR) Spearman correlation of <0.05 are displayed. d, Dot plot of enriched pathways in module 2 (ME2). The x axis shows the NES; dot size corresponds to −log10(P). Over-representation was assessed with a one-sided Fisher’s exact test. All displayed pathways were significantly enriched (FDR < 0.05) after Benjamini–Hochberg correction. e, Distribution of ME2 eigengene values across snRNA-seq-defined cellular communities (Fig. 4f,g). Box plots depict the median (centerline), IQR (box) and 1.5× IQR whiskers. Boxes colored by snRNA-seq-defined cellular communities, as shown in b. Statistical significance was assessed using the Kruskal–Wallis test with Dunn’s post hoc correction and Benjamini–Hochberg (FDR) correction for multiple comparison testing; *FDR < 0.05.
Plasma proteomics identifies aberrant cell-state biomarkers
To assess whether aberrant states are reflected in circulation, we profiled matched plasma from 64 study participants using the Olink Explore HT panel (5,420 protein biomarkers) and applied a module-based approach. Plasma module ME2 correlated with inflamed arterial and gCap endothelial cells (Fig. 6c), was enriched for inflammation-related pathways (including C-type lectin receptor, TNF and IL-17 signaling; Fig. 6d), and included NF-κB regulators (CHUK, SIRT6, LRRFIP1), lymphoid and myeloid transcription factors (NFATC1, NFATC3, CEBPB), cytokine signaling mediators (TNF, STAT2, STAT5B), innate immune effectors (CLEC6A, TRAF3) and apoptosis-related proteins (CASP8, CASP10, PDCD5; Supplementary Table 10a,b). The ME2 eigenscore was higher in samples from community 5 (‘high-inflammatory’) compared to other communities, suggesting that ME2 may represent a noninvasive biomarker of high-inflammatory tissue states (Fig. 6e).
Cell-autonomous and cell signaling pathways in COPD
We hypothesized that disease progression is sustained by dysregulated cell-autonomous programs and aberrant cell–cell signaling. To test this, we derived cell-specific gene expression modules and assessed their associations with clinical traits, adjusting for sex, BMI, age and smoking exposure (Extended Data Fig. 8a). Smoking exposure was summarized using principal components (PCs) from pack-years, quit-years and smoking status (Supplementary Fig. 10). We then applied mediation analysis to determine whether the effects of smoking and age on clinical outcomes were transmitted through these expression programs or cell-type abundance (Fig. 7a–c). Modules mediating or associated with disease were enriched for inflammatory signaling, growth and repair (WNT, TGF/BMP, growth factors), and aging pathways (telomere maintenance, DNA damage response, mTOR signaling, autophagy, stress responses), along with cell migration, ECM interactions, cell death, cell cycle regulation and protein metabolism (Fig. 7d and Extended Data Fig. 8b). Mediation analysis of cell-state abundance revealed IMCHIT1 mediate the negative effects of smoking on dyspnea, FEV1 and FEV1/FVC, whereas AT1 cells protect against dyspnea (Fig. 7c).
a, Mediation analysis model. b, Sankey plot depicting cell-specific transcriptional modules mediating the effect of smoking or age on clinical traits. c, Sankey plot depicting cell proportion changes mediating the effect of smoking on clinical traits. d, Dot plot illustrating pathway enrichment of gene modules identified through marginal-effect analyses (circles) or mediation analyses (triangles) across key clinical traits in COPD. Clinical traits are represented along the panels on the x axis, where the direction of the effect estimate is multiplied by −log10(Penrichment). Clinical traits grouped into spirometry (FEV1, FEV1/FVC), emphysema (emph), exacerbations (wheezing, infections) and symptoms (cough, dyspnea, SGRQ symptoms). Pathways are displayed on the y axis. Pathways grouped into biologic themes represented by y-axis color labels (dark blue, inflammatory; orange, remodeling and repair; pink, aging hallmarks; light blue, other; yellow, cell migration and cell–ECM interactions). Significance assessed using a two-sided permutation test. The size of each dot indicates the degree of pathway enrichment, and colors correspond to specific cell types. All displayed pathways were significantly enriched within corresponding cell types after Benjamini–Hochberg correction (FDR < 0.05). e,f, Aberrant ligand–receptor signaling. e, Receiver-focused analysis. Rows represent ligand–receptor pairs (or ligands alone when the cognate receptor was not profiled with Xenium). Columns indicate either ligand-expressing (left) or receptor-expressing (right) cell types. Color intensity in the left reflects normalized ligand expression across sender populations measured by snRNA-seq. Right, the normalized difference in hotspot prevalence between inflamed and noninflamed neighborhoods, calculated as (hotspot % in inflamed − hotspot % in noninflamed)/(hotspot % in inflamed + hotspot % in noninflamed). For IMCHIT1, contrasts were computed relative to IM1. f, Sender-focused analysis. Rows represent ligand–receptor pairs (or ligands alone if the receptor was not profiled with Xenium). All ligands shown were significantly upregulated in inflamed versus noninflamed senders, IMCHIT1 versus IM1 or FibroCTHRC1 versus other fibroblasts (Wilcoxon rank-sum test, FDR < 0.05). Left, the log2(FC) in ligand expression relative to the matched control population. Right, the normalized difference in hotspot prevalence using the same formula as in e. Only ligand–receptor interactions with significant differences in both hotspot prevalence (two-proportion z test; FDR < 0.05) and hotspot intensity (Moran’s I z score, Wilcoxon rank-sum test; FDR < 0.05) are shown.
To delineate signaling that sustains aberrant states, we performed ligand–receptor inference on snRNA-seq, treating aberrant cells as both recipients and sources of inflammatory/remodeling cues. We then validated these interactions using spatial transcriptomic data by quantifying spatial autocorrelation with local Moran’s I, which showed greater ligand–receptor colocalization with aberrant cells or with increasing disease severity. Inflamed nonimmune cells received increased IL4R, IL33, IL6, IL-1, IFN and TNF signaling relative to noninflamed counterparts, while IMCHIT1 macrophages received greater SEMA3A, CSF1, IL4R and TNF signaling than IM1 (Fig. 7e and Supplementary Table 11a). As signal-producing cells, inflammatory endothelial cells were prominent sources of CXCL10, CXCL11 and CSF3; inflammatory fibroblasts (alveolar and reticular) expressed high levels of CCL2, VEGF, IL15 and IL33; CTHRC1+ fibroblasts expressed FGF7 and TGFB1; and inflamed macrophages expressed CXCL3, CXCL5, IL1B, IL1A and CCL2 (Fig. 7f and Supplementary Table 11b). Collectively, these findings delineate cell-type-specific, spatially organized signaling networks that likely sustain inflammatory or remodeling microenvironments and promote disease progression.
Discussion
This study delineates an evolving cellular landscape in COPD that is closely linked to clinical features, disease progression and ECM remodeling. We identify shifts in canonical cell types and the emergence of pathologic cell states that assemble into spatially organized microenvironments. This includes an ‘inflammatory microenvironment’ enriched in nonimmune cell states with pro-inflammatory transcriptional programs and a ‘remodeling microenvironment’ composed of reparative and profibrotic cells. Quantifying the abundance of these populations across disease severity revealed that early reparative responses give way to persistent inflammatory and fibrotic states, with inflamed nonimmune cells strongly associated with increased symptoms and higher exacerbation scores. These findings provide a mechanistic framework linking disease-associated cell states to clinical phenotypes, while the incorporation of matched plasma biomarkers suggests opportunities for patient stratification and targeted therapy.
The inflammatory landscape of COPD emerges as a spatially organized, multicellular network in which structural cells adopt pro-inflammatory transcriptional programs and engage in reciprocal signaling with macrophages and other immune cells. Both individual inflamed states and the broader inflammatory microenvironment were increased in severe disease, particularly among participants with high symptom burden and exacerbation scores. Mechanistically, pathway analyses indicate that these inflammatory niches are sustained by a combination of cell-autonomous dysfunction, including activation of aging-related pathways, and paracrine signaling between structural and immune cells. Defining this subtype of COPD pathobiology as a spatially organized inflammatory network provides a basis for improving therapeutic targeting by revealing the cell states and microenvironments most relevant to disease biology. For example, IL4R, which is clinically targetable with dupilumab, was elevated across multiple cell types and correlated with a broad inflammatory cytokine program rather than a type-2-restricted milieu. Although prior studies identify a history of exacerbations as the strongest predictor of future events42, our cross-sectional design did not allow us to determine whether the inflammatory microenvironment’s association with exacerbation scores is causal.
A second key finding is a population shift consistent with aberrant repair. While COPD is widely recognized as a disease of impaired regeneration, direct evidence in human lung tissue has been limited12,43,44,45. We observed an expansion of regenerative epithelial populations in early disease that plateaued or declined with progression, while ABCs and CTHRC1+ fibroblasts increased, suggesting a transition from regeneration to maladaptive remodeling. These observations position COPD as a disorder of impaired regeneration, highlight conserved mechanisms that may be therapeutically targetable and support a biology-driven disease classification. The overlap of populations in COPD and IPF implies shared injury-repair programs but does not explain the divergence between IPF and COPD. One clear difference is their relative abundance in disease, but activation state, spatial organization or microenvironmental cues may also have a role. It also remains unclear if ABCs in COPD are transitional intermediates that resolve or nonproductive cells that accumulate and drive disease. Interestingly, ABCs were predominantly parenchymal and their relative scarcity near airways may reflect alveolar specificity, but this may have occurred due to undersampling of airway regions or an airway intermediate not captured by our Xenium panel10,29.
Several limitations warrant consideration. First, snRNA-seq detects similar numbers of unique genes as single-cell RNA sequencing but is less sensitive for cytoplasmic transcripts and low-abundance cell types, including T cells, potentially underestimating their contributions46. While we captured certain T-cell subsets, further associations were deferred pending independent validation. Second, integration across samples, inherent to snRNA-seq workflows, may obscure rare or participant-specific features. Third, technical variability (nuclear isolation, ambient RNA, tissue quality, processing time) can introduce bias despite our efforts to mitigate these through a standardized single-operator protocol, ambient RNA removal and enrollment from a large National Institutes of Health (NIH)-sponsored study with uniform procedures. Fourth, sample representativeness is limited by parenchymal sampling (under-representing airway compartments), the absence of precise anatomic coordinates within lobes and the inclusion of early-stage samples from lobectomies for malignancy. While prior reports describe an increase in AT2SCGB3A2 cells in COPD, we observed a modest decline in advanced disease12,13. This may have arisen from methodological differences, such as the narrow transcriptional window captured by snRNA-seq compared to protein detection by immunostaining, differences in cohort composition, smoking status or disease severity. Finally, snRNA-seq of samples from the same lobe yielded consistent cellular compositions, whereas we identified differences between lobes suggestive of regional heterogeneity even within participants. While the study was underpowered to confirm this, correlations between inflammatory plasma biomarkers and inflamed endothelial cells suggest participant-level inflammatory pathways transcend regional variation, meriting validation in larger cohorts.
Despite these limitations, this study advances a biology-anchored framework for COPD. Clinically, it highlights the potential for mechanism-guided stratification, in which aberrant cell states and their surrounding tissue microenvironments can be used to define therapeutic targets. The findings also support more precise clinical trial design by enabling participant enrichment based on tissue or plasma biomarkers, allowing treatments to be aligned with the dominant biological processes present in each patient. Furthermore, the ability to track cell-state shifts as pharmacodynamic readouts, together with longitudinal monitoring through serial biomarker trajectories, provides a roadmap for moving COPD care toward a precision medicine approach.
Methods
Compliance with ethical regulations
This study complied with all relevant ethical regulations. Lung tissue and plasma were obtained from the National Heart, Lung, and Blood Institute-sponsored LTRC47. The LTRC protocol was approved by local institutional review boards; all participants provided written informed consent; and the study was approved by the Yale Institutional Review Board (protocols 200003839 and 2000039474). Deidentified clinical data and tissue samples were obtained from the Biologic Specimen and Data Repository Coordinating Center (BioLINCC).
Study cohort and clinical data
LTRC clinical center staff conducted standardized face-to-face interviews using LTRC forms (https://biolincc.nhlbi.nih.gov/studies/ltrc/) covering demographics, medical/family history, smoking, therapies, symptoms, environmental/occupational exposures and validated questionnaires. Spirometry and DLCO testing were performed in accordance with the 2005 American Thoracic Society/European Respiratory Society recommendations48; predicted values for FEV1 were based on ref. 49; DLCO reference equations followed the standards without race/ethnic adjustment50. DLCO was hemoglobin (Hgb)-adjusted as—corrected DLCO = measured DLCO × (10.22 + Hgb)/(1.7 × Hgb). Of 3,066 participants without primary pulmonary fibrosis, duplicate entries were removed, prioritizing completeness and proximity to surgery. We excluded 386 with missing symptom questionnaires, 273 with missing spirometry and an additional 194 with missing smoking exposure variables, yielding a final cohort of 2,213. Missing Hgb was imputed with sex-specific means. We imputed ‘99’ for quit-years, ‘0’ pack-years for never-smokers and ‘0’ quit-years for active smokers. Follow-up questions related to symptom frequency or severity were imputed ‘0’ if the primary symptom was absent. For semiquantitative CT variables with missing lobe-specific/region-specific entries, we applied a hierarchical fill—(1) within-lung/lobe medians across regions; (2) if unavailable, contralateral-lung same lobe or (3) same-lung median of other lobes. For missing lobe-specific quantitative emphysema (HU = ≤950) where semiquantitative scores indicated no emphysema, we imputed the median among samples without emphysema.
Quantitative representation of symptoms
Questionnaire items were grouped into clinically relevant domains—cough, wheezing, exacerbations, infection and dyspnea. To obtain a single quantitative measure for each domain, we performed PCA on all LTRC participants (n = 2,213). Severity scales were recoded so higher values always reflected greater severity; binary items were coded 0/1 (no/yes). The first PC of each domain served as the domain score. Domain scores for infection and cough were multiplied by −1, so higher values uniformly indicated worse symptoms.
Clinical-trait-based clustering of the LTRC dataset
We retained 110 categorical and 24 continuous variables present in all participants (Supplementary Table 5). Continuous features included FEV1/FVC, % predicted FEV1, % predicted FVC, % predicted DLCO, smoking metrics (cigarettes/day, pack-years, years since cessation) and age. Summarized semiquantitative CT features included emphysema, airway inflammation, mosaic attenuation, nodules and ground glass. Participant-reported symptoms were assessed with the SGRQ (symptoms, activity, impacts, total) and SF-12, as well as additional questionnaire items (Supplementary Fig. 1) contributing to dyspnea, cough, infection, exacerbation and wheezing. Variable clustering was performed with the ClustOfVar package (v1.1)51. For categorical variable clusters, we retained the top five PCs (each explaining >85% of the variance in all but one cluster), yielding 55 PCs; for continuous clusters, we retained the top 12 PCs. Distance matrices (x) were computed using pairwise Euclidean distances of PC loadings among participants for categorical and clinical variables separately, which was further transformed into a similarity matrix w = exp(−x). The categorical and continuous similarity matrices were fused using similarity network fusion (k = 20 neighbors; T = 20 iterations)52. The fused similarity w′ was converted to a dissimilarity d = −log(2w′) for patient clustering using the partitioning around medoids method53.
snRNA-seq
Human snap-frozen lung tissue samples were requested from BioLINCC. Nuclei isolation was performed using the Chromium Nuclei Isolation Kit (10x Genomics) per the manufacturer’s instructions54. Aliquots were stained with DAPI, nuclei were counted using the Thermo Fisher Scientific Countess II FL, and approximately 20,000 nuclei per sample were loaded on Chip M and processed on the Chromium X controller. Libraries were prepared with the Chromium Next GEM Single Cell 3′ HT kit (v3.1) and QC assessed with the Agilent Bioanalyzer High Sensitivity DNA chip. Of 164 experiments, 9 did not meet QC for sequencing. Libraries were sequenced on the NovaSeq 6000 System (paired end = 100 bp), targeting approximately 50,000 reads per cell. Samples from 146 lobes and 141 participants were sequenced (five samples from the same participant, different lobes).
Computational processing
Reads were processed with Cell Ranger (v7.1; GRCh38) to generate feature-barcode matrices55. Ambient RNA was removed with CellBender (v0.2; false-positive rate = 0.01; 150 epochs)56. Six additional samples were excluded for low quality. Using Seurat (v5.0.1), data were log-normalized; 2,000 variable genes (excluding antisense and mitochondrial genes) were selected; scaling regressed out percent mitochondrial reads and feature counts57. Mitochondrial thresholds were 2% for all cells except 3% for T cells. PCA was performed, batch effects were corrected with Harmony (v1.2.0)58 and a shared-nearest-neighbor graph was built for Louvain clustering, while embeddings were visualized with Uniform Manifold Approximation and Projection. Doublets were identified with DoubletFinder (v2.0), and we applied an iterative clustering approach to remove additional clusters identified as doublets or cellular debris.
Cell-type annotation
Initial annotations were assigned via reference-based label transfer using Azimuth (v0.5.0)59 then refined by manual review of differential expression and canonical markers curated from the human lung cell atlas and LungMAP9,16,34. Clusters were inspected for technical artifacts (gene/feature counts, mitochondrial fraction, antisense, participant-specific). Canonical lineages (alveolar/airway epithelium, endothelium, fibroblasts and immune populations) were verified by marker expression obtained using the Seurat FindAllMarkers function. We identified emergent cell states by increasing clustering resolution, then annotated them based on distinct transcriptomic signatures and prior literature; inflammatory nonimmune states were labeled by enrichment of inflammatory pathways and AT2SCGB3A2 (refs. 15,34), AT2S (refs. 12,13,14), ABC10,11, fibroblast populations13,16,28,33,38, gCapS25,26,60 and IMCHIT1 (ref. 35) by marker genes identified in prior studies.
Cell-type proportion and compositional analyses
We quantified COPD-associated shifts in cellular composition using proportional abundances of hierarchically annotated states. Cells were organized into the following four levels: level 4 (all cells), level 3 (parent lineages—alveolar epithelium, airway epithelium, endothelium, fibroblasts, macrophages, monocytes, lymphocytes, mast cells, dendritic cells, mesothelium), level 2 (canonical cell types within each lineage) and level 1 (intermediate/emergent states). To address the unit-sum constraint of single-cell proportions, we used scComp (v1.9)19. Denominators were selected to match the biological question. For inflamed nonimmune states, proportions were computed within the corresponding level-2 canonical cell type to capture lineage-intrinsic inflammation. For canonical cell-type comparisons, proportions were calculated within the parent level-3 lineage. For reparative/remodeling epithelial states and immune populations, we reported tissue-level representation as a proportion of all cells. For macrophages, we analyzed both all-cell and level-3 lineage denominators consistently by trait—continuous traits, smoking and GOLD-used level-3 denominators; emphysema category used all-cell denominators. Samples were excluded if there were <10 cells in the relevant parent population (or <10 nonepithelial alveolar cells for analyses involving alveolar epithelium). Partial Spearman correlations with continuous traits were adjusted for age, sex, BMI, smoking status (yes/no) and lobe (for emphysema) using ppcor (v1.1).
Pathway enrichment analyses for gene expression profiles and protein data
Differential expression used Seurat’s FindMarkers (minimal log2 fold change (FC) = 0.20; minimal expression = 0.075). Gene-set enrichment used fGSEA (v3.19) with Molecular Signatures Database (MSigDB, v2023.2), focusing on hallmark, curated and Gene Ontology pathways gene sets. Ranked lists were based on average log2(FC); normalized enrichment scores were computed with minimum and maximum gene-set sizes of 15 and 500. Redundancy was reduced using collapsePathways as well as manual curation. Pathways with FDR < 0.05 were considered significant (figures show unadjusted P values). For protein-level enrichment, we used g:Profiler (v0.2.3)61.
Clustering patients by cell-type proportions
Participants were clustered by proportions of aberrant cells. To ensure stable estimates, we included samples with ≥5 cells for each of gCap, aerocyte, alveolar fibroblast, AT2, AT1, secretory/club and macrophage. Spectral clustering (SNFtools, v2.3.1) was performed using between-participant Euclidean distances based on cell-type proportions and the affinity matrix was constructed using 15 NNs and a local variance of 0.5 (ref. 52).
Mediation analysis
Exposures were smoking and aging. Smoking exposure was summarized by PCA across four variables (ever-smoker, active smoker, pack-years, quit-years) using the first PC. For each cell type, aggregated gene information was computed as follows: (1) retain samples with ≥10 cells per cell type and exclude cell types with fewer than 30 samples after filtering as well as genes expressed in less than 7.5% of cells; (2) aggregate counts to pseudobulk per sample using Seurat’s AggregateExpression function and (3) identify coexpression modules using WGCNA (see below) with the first principal component (module eigengene) serving as the aggregated gene information. Outcomes included FEV1, FEV1/FVC, DLCO, symptom domain scores (dyspnea, cough, infection, exacerbation, wheezing), lobe-specific quantitative emphysema (HU = ≤950), lobe-specific semiquantitative emphysema and SGRQ symptoms. For each module/exposure/outcome, we fit generalized linear regression, binary probit regression and ordered probit regression for continuous, binary and ordered outcomes, respectively. BMI and sex were included as covariates together with smoking or aging (unless serving as the exposure).
Blood proteomics
Plasma samples from 64 individuals were profiled using the Olink Explore HT. Relative protein abundance was reported as normalized protein expression (log2 scaled). Internal controls (incubation, extension (primary normalization) and amplification) and external controls (plate control, sample control, negative control) were used per Olink guidelines; all samples passed QC. Final normalized protein expression values were generated using Olink Explore software (v6.7.2).
Lung decellularization
Decellularization was optimized for lung tissue62,63. Tissue was rinsed in PBS + Ca2+/Mg2+ for 10 min and then for 30 min. Samples were treated with 0.0035% Triton X-100 for 60 min, washed in Benzonase buffer for 20 min, incubated with Benzonase (20U ml−1; Sigma-Aldrich, E1014) for 60 min, exposed to 1 M NaCl for 20 min and rinsed in PBS for 8 min. Samples underwent graded washes with sodium deoxycholate (Sigma-Aldrich, D6750) at 0.01%, 0.05% and 0.1% for 30 min each, followed by PBS for 20 min. Samples were washed in Benzonase buffer for 20 min, treated with Benzonase overnight to ensure nucleic acid removal. The decellularized tissue was then washed in PBS for 20 min, followed by a 20-min wash in 0.5% Triton X-100, and five PBS washes, each lasting 20 min. Benzonase buffer consists of 50 mM Tris–HCl (pH 8.0), 1 mM MgCl2 and 0.1 mg ml−1 BSA in deionized water.
Label-free mass spectrometry (MS)
Decellularized tissue was denatured in urea containing ammonium bicarbonate, reduced with dithiothreitol, and alkylated using iodoacetamide to prevent disulfide bond formation. Proteins were digested with PNGase F at 37 °C overnight on a shaker to cleave N-linked glycans. Proteins were then digested with Lys-C for 4 h at 37 °C, followed by overnight digestion with trypsin. Digestion was quenched with 20% trifluoroacetic acid. Samples were desalted using BioPureSPN PROTO 300 C18 Macro spin columns (The Nest Group), eluted and dried using a SpeedVac. The dried peptides were resuspended in MS loading buffer before liquid chromatography–tandem MS (LC–MS/MS). LC–MS/MS was performed on a Thermo Fisher Scientific Q Exactive HFX equipped with a Waters ACQUITY M-Class UPLC system using a binary solvent system. Trapping was performed at 5 µl min−1 using a Waters Symmetry C18 trap column (100 Å, 5 µm, 180 µm × 20 mm). Peptides were separated at 37 °C using an ACQUITY UPLC Peptide BEH C18 column (130 Å, 1.7 µm, 75 µm × 250 mm) and eluted at 300 nl min−1. MS was acquired in profile mode over the m/z range of 350–1,500. Data-dependent LC–MS/MS were acquired in centroid mode on the top 20 precursors per MS scan. To monitor instrument performance, heavy-labeled synthetic peptide standards (Retention Time Calibration Mix, Thermo Fisher Scientific) were injected into every experimental sample. Sample injection sequences were randomized to minimize the risk of drift in false-positive or false-negative signals. Approximately 250 ng per sample was loaded onto the LC column, with four blank runs performed among sample injections to control for carryover. Data analysis was conducted using Proteome Discoverer software (v2.5; Thermo Fisher Scientific). For protein identification, data were searched using SEQUEST HT (Thermo Fisher Scientific) (UniProt human). Search parameters included tryptic peptides of ≤2 missed cleavages, 10-ppm precursor mass tolerance, 0.02 Da fragment mass tolerance and variable modifications of oxidation on methionine and carbamidomethylation on cysteine. A decoy database was searched to determine the FDR. Label-free quantification used a feature mapper node, aligned chromatographic features and a precursor ion quantifier node to normalize protein abundances based on total peptide amount. Only proteins with an FDR of ≤5% and at least two unique peptides were included in statistical analyses.
Weighted gene coexpression network analysis (WGCNA) for proteins
WGCNA (v1.73) was run separately for (1) cell-type-specific pseudobulk gene expression, (2) decellularized ECM proteomics and (3) blood proteomics, using the same analysis stages. For each dataset, we computed biweight midcorrelations to build a signed network, selected the soft-thresholding power to approximate scale-free topology (genes, data-driven target R2 ≳ 0.9; ECM, β = 12; blood, β = 8), converted correlations to an adjacency matrix and then to a topological overlap matrix—a measure of shared network connectivity among feature pairs that emphasizes nodes with common neighbors. Modules were detected by dynamic tree cutting with dataset-matched settings (genes—minimum module size = 10, split depth = 1; ECM—minimum module size = 20, split depth = 3; blood—minimum module size = 20, split depth = 4) and a merge threshold of 0.25. Module eigengenes (first PC per module) were correlated with clinical traits using Spearman correlation and per-feature module membership (kME) were calculated.
Immunofluorescence staining
FFPE sections were dewaxed (xylene), rehydrated (graded ethanol) and subjected to antigen retrieval (Tris–EDTA, pH 9; NB900-62085) at 98 °C for 20 min. Slides were permeabilized (0.3% Triton/PBS-T) for 10 min, blocked for 10 min (CAS-Block; Life Technologies, 008120) and incubated with primary antibodies overnight at 4 °C, followed by fluorophore-conjugated secondary antibodies (1 h, room temperature), and subsequently mounted in Vectashield with DAPI (H-1800). Images were acquired with a Leica Stellaris 8 Falcon confocal microscope (×20 objective). Primary/secondary antibody details are provided in Supplementary Table 12.
Xenium spatial transcriptomics
We performed Xenium in situ analysis (10x Genomics) on tissue microarrays constructed from 38 matched participants (2 mm2, two to three cores per participant), sampling parenchyma and, where possible, airway and/or large vessel regions. All samples underwent histopathologic review. FFPE sections (5 µm) were processed on Xenium slides following the manufacturer’s protocols (CG000578, Rev C) with a custom gene panel. FFPE tissue sections were deparaffinized by incubating slides at 60 °C, followed by sequential immersion in xylene, ethanol (100%, 96%, 70%) and nuclease-free water. Slides were then dried, assembled into Xenium cassettes and treated with a Decrosslinking Buffer. The assembled cassettes were incubated at 80 °C in a thermal cycler to release RNA for downstream analyses. After hybridization, the slides were imaged and processed using the standard pipeline of the Xenium Analyzer (v3.2.0), with transcript assignment to individual cells performed using the 10x Genomics multimodal segmentation algorithm. QC thresholds included Phred Q > 20 for transcripts; cells required nFeature of >5, nCounts of >9 and nuclear area of 6–80 µm². Downstream analysis was performed in Seurat (v5.0.1) including normalization, PCA, neighbor graph construction, clustering and Uniform Manifold Approximation and Projection. Cell types were annotated by marker expression. Spatial niche analysis was conducted using Seurat’s BuildNicheAssay function (k neighbors = 30, k niches = 10). The dominant cell types within each niche were explored and visualized, along with the distribution of these niches across the sample. Images were generated using Xenium Explorer (v3.2.0).
Deconvolution of the GeoMx spatial transcriptomics secondary cohort (Baylor cohort) for validation
We reanalyzed data obtained from an independent cohort that underwent spatial transcriptomic profiling of lung tissue samples (GSE237120). A full description has been previously described64. Participants were recruited at the University of Arizona (2019–2021), and samples were obtained from lung volume reduction surgery, transplant for severe emphysema or resection of a solitary peripheral nodule (resection samples of >10 cm from the nodule), no active respiratory infection at surgery. GeoMx Digital Spatial Profiler Whole Transcriptome Atlas 18,753 RNA probes was used with structural markers (SYTOX13, Pan-CK-Alexa-532, CD45-Alexa-594). Regions of interest (ROIs) were selected by freehand annotation. Libraries were prepared per the manufacturer’s instructions and sequenced (Illumina NextSeq 500). FastQ files were converted to DCC for analysis with GeoMxTools (v3.8.0; default settings). ROIs were excluded if they failed any of the following criteria: raw reads of <1,000; trimming rate of <80%; stitching rate of <80%; alignment rate of <80%; sequencing saturation of <50% (computed as 1 − deduplicated reads/aligned reads × 100); negative-control geometric mean (nontargeting probes) of <1; no-template control count of >1,000; nuclei count of <100; segmented area of <5,000 µm2 and <1% genes detected. The deletion rates, from highest to lowest, were as follows: vessel (61%), airway (55%), follicle (47%) and parenchyma (32%). A total of 467 ROIs from these four tissue locations passed QC and were included in downstream analyses, resulting in the exclusion of samples (33 samples retained). Cell-type deconvolution was conducted using spatialDWLS65 implemented in the giotto R package (v1.1.2) with default parameters66 with our snRNA-seq data as reference. Reference sample criteria were total cells of ≥10,000, ≥10 cells in each parent lineage and ≥5 cells for each cell state. Four reference samples met criteria and were downsampled to 100 cells per cell type, with final reported proportions averaged across references. Differences in cell-type proportions across GOLD and emphysema categories were assessed.
Ligand–receptor interaction inference
Ligand–receptor signaling was inferred using NicheNet (v2.1)67. Ligand activity was quantified as the area under the precision–recall curve (AUPR), measuring how well each ligand’s downstream regulatory network predicted the expression of differentially expressed genes in receiver cells. We implemented two complementary analytical strategies:
-
1.
Receiver-oriented analysis (sender-agnostic mode)—structural cell types, as well as AM and IM, were treated individually as receiver populations. Within each, differentially expressed genes between the inflamed versus noninflamed cells (for structural cells and macrophages), IMCHIT1 versus IM1 or CTHRC1+ fibroblast versus non-CTHRC1+ fibroblasts were identified using Seurat v5’s FindMarkers function (min.pct = 0.05, |log2(FC) | ≥ 1, FDR < 0.01) and defined as the target gene set for NicheNet analysis. Ligand activity was computed in sender-agnostic mode, and the top 30 ligands per receiver based on AUPR were retained for further evaluation.
-
2.
Sender-oriented analysis (sender-specific mode)—the same structural populations were then designated as sender cells, with all other cell types considered potential receivers. Upregulated ligands between inflamed and noninflamed sender cells were identified using the same approach (min.pct = 0.05, log2(FC) ≥ 0.25, FDR < 0.01). To evaluate the association between inflamed sender cell frequency and gene expression in each receiver cell populations, we fit negative binomial generalized linear mixed models with inflamed sender cell proportion as a fixed effect and participants as a random effect, adjusting for age, sex, BMI and smoking status. The models were implemented using the iDESC function (iDESC, v0.1.0)68, without including a term for dropout rate. Genes with regression coefficient β > 1 and FDR < 0.01 were used as the target gene set for sender-specific ligand activity scoring. The top 30 ligands per receiver population were retained based on AUPR. For both strategies, analyses were restricted to samples containing ≥5 cells per structural cell type. Because lymphoid fibroblasts (IR fibroblasts and FRCs) were not identified specifically in our spatial dataset, they were incorporated into the fibroblasti population.
Spatial validation of ligand–receptor interactions using Xenium dataset
To validate predicted ligand–receptor interactions in situ, we performed bivariate local Moran’s I analysis on matched spatial transcriptomic data to quantify spatial co-enrichment. Cell coordinates were used to construct a spatial weight matrix based on k-NN with k = 30, using the spdep package (v1.2-8) with row-standardized weights. The list of ligand–receptor pairs identified from our initial analysis was further refined by cross-referencing with the curated ligand–receptor database in CellChat (v2.0)69, and excluding ECM-related interactions. Spatial colocalization was assessed under two conditions:
-
1.
Ligand–receptor analysis—if the receptor for a given ligand was expressed in ≥10% receiver cells, ligand and receptor expression vectors were z-score normalized (with ligand and receptor expression masked to their specific cell type) and their spatial association quantified with the bivariate local Moran’s I statistic.
-
2.
Ligand–cell-type analysis—if the receptor for a given ligand was not part of the 480 Xenium gene panel, a binary indicator of receiver cell identity was used in place of receptor expression if that receiver cell was known to express the receptor in >10% participants.
Bivariate local Moran’s I assumes that closely related variables tend to show similar spatial patterns in neighboring regions. For each cell i, we define the statistic as:
where \({I}_{i}^{\left({xy}\right)}\) is the bivariate local Moran’s I index for cell i, which quantifies the spatial association between the expression of ligand x in cell i and the expression of receptor y or cell-type indicator y in its neighboring cells j; \({z}_{x,i}\) is the z-score normalized ligand x expression in cell i; \({z}_{y,j}\) is (1) either z-score normalized receptor y expression in neighboring cell j, or (2) a binary indicator indicating whether cell j belongs to a specific receiver cell type, which was subsequently z-score normalized across all cells; \({w}_{{ij}}\) is the spatial weight between cells i and j, derived from the k-NN graph. Cell-type pairs with the same sender and receiver were excluded. Sender cells with bivariate local Moran’s I z scores of >2 were defined as spatial hotspots. Differences among groups were assessed by comparing hotspot prevalence with a two-proportion z test and hotspot intensity (Moran’s I z scores) with a Wilcoxon rank-sum test. Interactions meeting significance in both tests (FDR < 0.05) were considered spatially enriched.
Statistics and reproducibility
Sample sizes were determined by sample availability. No statistical method was used to predetermine sample size. Studies were nonrandomized and unblinded, but samples were randomized to sequencing or processed in balanced analytic batches, grouped by GOLD when possible. Data were excluded only by prespecified QC (failed libraries/runs, assay-specific metrics and minimum cells per parent lineage). Immunofluorescence images are representative of tissue samples from five participants per group. Differential cell composition was modeled with sccomp, adjusting for age, sex, BMI and smoking status, with emphysema-related analyses additionally adjusted for anatomic lobe; partial Spearman correlations (adjusted for the same covariates) summarized direction and strength of associations with continuous traits (Figs. 2a–o and 3c,d,f–o). Exact P values of compositional testing are provided in Supplementary Table 4. Groupwise comparisons of variables across clinical clusters or cellular communities were assessed using Kruskal-Wallis tests with post hoc Dunn’s tests (Fig. 4c,d,g; Fig. 6g), while pairwise comparisons used two-sided Wilcoxon rank-sum tests (Figs. 1l–n,r and 7e,f). Pathway enrichment analyses used gene set enrichment–based methods (Figs. 1c, 3e, 6d and 7d). Correlations with WGCNA modules and across aberrant cell populations were assessed using Spearman correlations (Figs. 4e and 6a,c). All inferential statistical tests were two-sided, with multiple testing controlled using Benjamini–Hochberg correction; exact n is reported in the legends.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Raw and processed data are deposited in the NCBI SRA under BioProject PRJNA1301244 and in GEO under accessions GSE310058 and GSE313006. Data from the Baylor cohort are available with accession GSE237120.
Code availability
Custom scripts for this project are available on GitHub (https://github.com/SaulerLab/COPD_snRNA-seq) and Zenodo (https://doi.org/10.5281/zenodo.17635478)70.
References
Ahmad, F.B. & Anderson, R.N. The leading causes of death in the US for 2020. JAMA 325, 1829–1830 (2021).
Han, M. K. et al. Chronic obstructive pulmonary disease phenotypes: the future of COPD. Am. J. Respir. Crit. Care Med. 182, 598–604 (2010).
Hansel, N. N. et al. In-home air pollution is linked to respiratory morbidity in former smokers with chronic obstructive pulmonary disease. Am. J. Respir. Crit. Care Med. 187, 1085–1090 (2013).
Hogg, J. C. & Timens, W. The pathology of chronic obstructive pulmonary disease. Annu. Rev. Pathol. 4, 435–459 (2009).
Bhatt, S. P. et al. Dupilumab for COPD with type 2 inflammation indicated by eosinophil counts. N. Engl. J. Med. 389, 205–214 (2023).
Moll, M. et al. Chronic obstructive pulmonary disease and related phenotypes: polygenic risk scores in population-based and case-control cohorts. Lancet Respir. Med. 8, 696–708 (2020).
Cho, M. H., Hobbs, B. D. & Silverman, E. K. Genetics of chronic obstructive pulmonary disease: understanding the pathobiology and heterogeneity of a complex disorder. Lancet Respir. Med. 10, 485–496 (2022).
Tuder, R. M. & Petrache, I. Pathogenesis of chronic obstructive pulmonary disease. J. Clin. Invest. 122, 2749–2755 (2012).
Sikkema, L. et al. An integrated cell atlas of the lung in health and disease. Nat. Med. 29, 1563–1577 (2023).
Adams, T. S. et al. Single-cell RNA-seq reveals ectopic and aberrant lung-resident cell populations in idiopathic pulmonary fibrosis. Sci. Adv. 6, eaba1983 (2020).
Habermann, A. C. et al. Single-cell RNA sequencing reveals profibrotic roles of distinct epithelial and mesenchymal lineages in pulmonary fibrosis. Sci. Adv. 6, eaba1972 (2020).
Basil, M. C. et al. Human distal airways contain a multipotent secretory cell that can regenerate alveoli. Nature 604, 120–126 (2022).
Kadur Lakshminarasimha Murthy, P. et al. Human distal lung maps and lineage hierarchies reveal a bipotent progenitor. Nature 604, 111–119 (2022).
Rustam, S. et al. A unique cellular organization of human distal airways and its disarray in chronic obstructive pulmonary disease. Am. J. Respir. Crit. Care Med. 207, 1171–1182 (2023).
Sauler, M. et al. Characterization of the COPD alveolar niche using single-cell RNA sequencing. Nat. Commun. 13, 494 (2022).
Sun, X. et al. A census of the lung: CellCards from LungMAP. Dev. Cell 57, 112–145 (2022).
Gillich, A. et al. Capillary cell-type specialization in the alveolus. Nature 586, 785–789 (2020).
Ghonim, M. A., Boyd, D. F., Flerlage, T. & Thomas, P. G. Pulmonary inflammation and fibroblast immunoregulation: from bench to bedside. J. Clin. Invest. 133, e170499 (2023).
Mangiola, S. et al. sccomp: robust differential composition and variability analysis for single-cell data. Proc. Natl Acad. Sci. USA 120, e2203828120 (2023).
Rojas-Quintero, J. et al. Spatial transcriptomics resolve an emphysema-specific lymphoid follicle B cell signature in COPD. Am. J. Respir. Crit. Care Med. 209, 48–58 (2023).
Li, Y. et al. Severe lung fibrosis requires an invasive fibroblast phenotype regulated by hyaluronan and CD44. J. Exp. Med. 208, 1459–1471 (2011).
Boyd, D. F. et al. Exuberant fibroblast activity compromises lung function via ADAMTS4. Nature 587, 466–471 (2020).
DiGiovanni, G. T. et al. Epithelial Yap/Taz are required for functional alveolar regeneration following acute lung injury. JCI Insight 8, e173374 (2023).
Deckelbaum, R. A. et al. The potassium channel Kcne3 is a VEGFA-inducible gene selectively expressed by vascular endothelial tip cells. Angiogenesis 23, 179–192 (2020).
Del Toro, R. et al. Identification and functional analysis of endothelial tip cell-enriched genes. Blood 116, 4025–4033 (2010).
Mason, E. C. et al. Activation of mTOR signaling in adult lung microvascular progenitor cells accelerates lung aging. J. Clin. Invest. 133, e171430 (2023).
Franzen, L. et al. Mapping spatially resolved transcriptomes in human and mouse pulmonary fibrosis. Nat. Genet. 56, 1725–1736 (2024).
Tsukui, T. et al. Collagen-producing lung cell atlas identifies multiple subsets with distinct localization and relevance to fibrosis. Nat. Commun. 11, 1920 (2020).
Wang, F. et al. Regulation of epithelial transitional states in murine and human pulmonary fibrosis. J. Clin. Invest. 133, e165612 (2023).
Willemse, B. W. M., Postma, D. S., Timens, W. & ten Hacken, N. H. T. The impact of smoking cessation on respiratory symptoms, lung function, airway hyperresponsiveness and inflammation. Eur. Respir. J. 23, 464–476 (2004).
Lumsden, A. B., McLean, A. & Lamb, D. Goblet and Clara cells of human distal airways: evidence for smoking induced changes in their numbers. Thorax 39, 844–849 (1984).
Hogg, J. C. et al. Micro-computed tomography measurements of peripheral lung pathology in chronic obstructive pulmonary disease. Proc. Am. Thorac. Soc. 6, 546–549 (2009).
Madissoon, E. et al. A spatially resolved atlas of the human lung characterizes a gland-associated immune niche. Nat. Genet. 55, 66–77 (2023).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Reyfman, P. A. et al. Single-cell transcriptomic analysis of human lung provides insights into the pathobiology of pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 199, 1517–1536 (2019).
Rodda, L. B. et al. Single-cell RNA sequencing of lymph node stromal cells reveals niche-associated heterogeneity. Immunity 48, 1014–1028 (2018).
Conlon, T. M. et al. Inhibition of LTbetaR signalling activates WNT-induced regeneration in lung. Nature 588, 151–156 (2020).
Malhotra, D. et al. Transcriptional profiling of stroma from inflamed and resting lymph nodes defines immunological hallmarks. Nat. Immunol. 13, 499–510 (2012).
Brown, F. D. et al. Fibroblastic reticular cells enhance T cell metabolism and survival via epigenetic remodeling. Nat. Immunol. 20, 1668–1680 (2019).
Abe, Y. et al. A single-cell atlas of non-haematopoietic cells in human lymph nodes and lymphoma reveals a landscape of stromal remodelling. Nat. Cell Biol. 24, 565–578 (2022).
Chioccioli, M. et al. Stem cell migration drives lung repair in living mice. Dev. Cell 59, 830–840 (2024).
Hurst, J. R. et al. Susceptibility to exacerbation in chronic obstructive pulmonary disease. N. Engl. J. Med. 363, 1128–1138 (2010).
Kasahara, Y. et al. Inhibition of VEGF receptors causes lung cell apoptosis and emphysema. J. Clin. Invest. 106, 1311–1319 (2000).
Hu, Y. et al. Airway-derived emphysema-specific alveolar type II cells exhibit impaired regenerative potential in COPD. Eur. Respir. J. 64, 2302071 (2024).
Puchelle, E., Zahm, J. M., Tournier, J. M. & Coraux, C. Airway epithelial repair, regeneration, and remodeling after injury in chronic obstructive pulmonary disease. Proc. Am. Thorac. Soc. 3, 726–733 (2006).
Koenitzer, J. R., Wu, H., Atkinson, J. J., Brody, S. L. & Humphreys, B. D. Single-nucleus RNA-sequencing profiling of mouse lung. Reduced dissociation bias and improved rare cell-type detection compared with single-cell RNA sequencing. Am. J. Respir. Cell Mol. Biol. 63, 739–747 (2020).
Yang, I. V. et al. Relationship of DNA methylation and gene expression in idiopathic pulmonary fibrosis. Am. J. Respir. Crit. Care Med. 190, 1263–1272 (2014).
Pellegrino, R. et al. Interpretative strategies for lung function tests. Eur. Respir. J. 26, 948–968 (2005).
Hankinson, J. L., Odencrantz, J. R. & Fedan, K. B. Spirometric reference values from a sample of the general U.S. population. Am. J. Respir. Crit. Care Med. 159, 179–187 (1999).
Crapo, R. O. & Morris, A. H. Standardized single breath normal values for carbon monoxide diffusing capacity. Am. Rev. Respir. Dis. 123, 185–189 (1981).
Chavent, M., Kuentz-Simonet, V., Liquet, B. & Saracco, J. ClustOfVar: an R package for the clustering of variables. J. Stat. Softw. 50, 1–16 (2012).
Wang, B. et al. Similarity network fusion for aggregating data types on a genomic scale. Nat. Methods 11, 333–337 (2014).
Kaufman, L. & Rousseeuw, P. J. Finding Groups in Data: An Introduction to Cluster Analysis (Wiley, 1990).
Wang, M. et al. Single-nucleus multi-omic profiling of human placental syncytiotrophoblasts identifies cellular trajectories during pregnancy. Nat. Genet. 56, 294–305 (2024).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat. Methods 20, 1323–1335 (2023).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Gaskill, C. F. et al. Disruption of lineage specification in adult pulmonary mesenchymal progenitor cells promotes microvascular dysfunction. J. Clin. Invest. 127, 2262–2276 (2017).
Kolberg, L., Raudvere, U., Kuzmin, I., Vilo, J. & Peterson, H. gprofiler2—an R package for gene list functional enrichment analysis and namespace conversion toolset g:Profiler. F1000Res 9, ELIXIR-709 (2020).
Calle, E. A. et al. Targeted proteomics effectively quantifies differences between native lung and detergent-decellularized lung extracellular matrices. Acta Biomater. 46, 91–100 (2016).
Petersen, T. H., Calle, E. A., Colehour, M. B. & Niklason, L. E. Matrix composition and mechanics of decellularized lung scaffolds. Cells Tissues Organs 195, 222–231 (2012).
Rojas-Quintero, J. et al. Spatial transcriptomics resolve an emphysema-specific lymphoid follicle B cell signature in chronic obstructive pulmonary disease. Am. J. Respir. Crit. Care Med. 209, 48–58 (2024).
Dong, R. & Yuan, G.-C. SpatialDWLS: accurate deconvolution of spatial transcriptomic data. Genome Biol. 22, 145 (2021).
Dries, R. et al. Giotto: a toolbox for integrative analysis and visualization of spatial expression data. Genome Biol. 22, 78 (2021).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Liu, Y. et al. iDESC: identifying differential expression in single-cell RNA sequencing data with multiple subjects. BMC Bioinformatics 24, 318 (2023).
Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell-cell communication from single-cell transcriptomics. Nat. Protoc. 20, 180–219 (2025).
Zhang, Y., Nouws, J. & Wei, H.-H. SaulerLab/COPD_snRNA-seq: aberrant cellular communities underlying disease heterogeneity in chronic obstructive pulmonary disease (v1.0.0). Zenodo https://doi.org/10.5281/zenodo.17635478 (2025).
Acknowledgements
M.S. is supported by NIH (grants R01HL155948 and R21HL173512) and DOD (grants W81XWH2210629 and HT94252310034). P.H. is supported by a Fellowship and grants from the National Health and Medical Research Council of Australia (1175134, 2010287, 2008937 and 2015613), Australian Research Council (200101058 and 230101156) and UTS. S.S.S. and M.S.B.R. are supported by the National Institute of General Medical Sciences (T32 GM086287). N.E.B. is supported by the National Library of Medicine (R01HL145372). X.Y. is supported by the National Library of Medicine (R01 LM014087). N.K. is supported by the National Heart, Lung, and Blood Institute (NHLBI; F30HL162459, R01HL127349, R01HL141852, U01HL145567, UH3 TR002445 and R21HL161723), and a grant from the Three Lakes Foundation. C.D.C. is supported by the National Institute of General Medical Sciences (HT94252310034). F.P. is supported by NHLBI (R01HL149744) and Baylor College of Medicine funds outside the submitted work. The authors thank the Keck MS & Proteomics Resource at the Yale School of Medicine for providing the necessary mass spectrometers and the accompanying biotechnology tools, supported in part by the Yale School of Medicine and the Office of the Director, NIH (S10OD02365101A1, S10OD019967 and S10OD018034). Funds for the MS analyses were supported in part by the Yale/NIDA Neuroproteomics Center (P30 DA018343). The authors appreciate the help of L. Charette and C. Akbar. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
M.S. conceived the study with considerable input from J.M. and X.Y. Y.Z., H.W., J.N., S.-H.K., J.P.N., S.M.P., W.J., W.W., J.L., A.Y.Z., R.M.B., S.L., A.T.O., N.E.B., T.T.L., J.M., X.Y. and M.S. analyzed data. J.N. performed sequencing and immunofluorescence staining. W.W., F.C. and T.T.L. performed mass spectrometry. Y.Z., H.W., J.N. and M.S. wrote the manuscript. R.J.H. and X.Z. evaluated histologic samples. F.P. provided data. S.S.S., J.P.N., P.H., C.D.C., C.B., J.G., S.M.C., E.L.H., T.T.L., N.E.B., M.S.B.R., S.M., R.J.H., N.K., F.P., J.M. and X.Y. provided technical support, conceptual advice and/or edited the manuscript. All authors provided intellectual input toward the manuscript and approved the submission.
Corresponding author
Ethics declarations
Competing interests
M.S. has received consulting fees and honorarium from Sanofi-Regeneron and has received grant funding from Genentech. He has a financial interest in Crosswalk Health. N.K. was a consultant to Pliant, AstraZeneca, CSL Behring, Galapagos, GSK, Merck, Thyron and Boehringer Ingelheim over the last 3 years, reports equity in Pliant and received research grants to his laboratory from Veracyte, Boehringer Ingelheim, BMS and AstraZeneca. He has patents on new therapies and biomarkers in pulmonary fibrosis. F.P. has received grants from Victory Houston and Boehringer Ingelheim, and consulting fees from Sanofi-Regeneron, Verona Pharma and Genentech for advisory board participation. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Robert Hall, Ani Manichaikul and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single-nuclear RNA sequencing of lung tissue samples from LTRC study participants with and without COPD.
a, UMAP showing 1,516,727 nuclei across 31 lung cell types. b, Heatmap displaying normalized expression levels of key markers across various cell types. Rows represent specific genes, while columns are hierarchically clustered by cell type and sample. Gene expression values are unit normalized (0–1) across rows. ABC = aberrant basaloid cell; AT1 = alveolar type 1 pneumocyte; AT2 = alveolar type 2 pneumocyte; gCap = general capillary cell; DC = dendritic cell; NK = natural killer cell; ccMyeloid = myeloid cells in cell cycle.
Extended Data Fig. 2 Inflamed structural cell states in COPD.
a, UMAP of epithelial cells, highlighting inflamed AT1 (AT1ᵢ). b, Immunofluorescence for SOD2 (red), PRX (green), and DAPI (blue) in representative fields (representative of 5 subjects/group) from samples with high versus low gCapᵢ/aerocyteᵢ abundance. c, Immunofluorescence for NF-κB (red), AGER (green), and DAPI from samples with high versus low AT1ᵢ (representative of 5 subjects/group). d, Immunofluorescence for NF-κB (red), pro-SFTPC (green), and DAPI (representative of 5 subjects/group) from samples with high versus low AT2ᵢ. e, Immunofluorescence for SOD2 (red), SCGB3A2 (green), and DAPI (representative of 5 subjects/group) from samples with high versus low secretoryᵢ. Scale bars, 100 μm (b–e). f–h, Proportion of inflamed cell states (gCapᵢ, AT2ᵢ, and alveolar fibroblastᵢ) within their parent populations, stratified by lobe-specific radiographic emphysema (% voxels ≤950 HU) in the Baylor cohort (low <5% emphysema, n = 20; high >5% emphysema, n = 13), as identified after deconvolution of spatial transcriptomic data. Box-and-whisker plots show the median (centerline), interquartile range (box), and whiskers extending to 1.5× IQR.
Extended Data Fig. 3 Aberrant cell states in COPD.
a, UMAP of airway epithelial cells. b, Dot plot of selected marker genes. c, Immunofluorescence for CPA6 (green), KRT17 (red), and DAPI (blue). Scale bars, 100 μm (top) and 50 μm (bottom). d, Immunofluorescence for CTHRC1 (red), KRT17 (green), and DAPI (blue). Scale bars, 100 μm. e, Proportion of CTHRC1⁺ fibroblasts among all fibroblasts, stratified by lobe emphysema in the Baylor cohort (low <5% emphysema, n = 20; high >5% emphysema, n = 13). f–h, Per-subject associations between cell-type abundance and clinical features: (f) ciliated epithelial cells (proportion of airway epithelial cells) vs. FEV₁ % predicted (n = 137). (g) SCGB3A2⁺ secretory (proportion of secretory cells) vs. lobe emphysema (n = 108). (h) DLCO % predicted vs. arteryᵢ (n = 73), aerocyteᵢ (n = 83), gCap (n = 87), AT1ᵢ (n = 122), AT2ᵢ (n = 114), adventitial fibroblastᵢ (n = 93), and alveolar fibroblastᵢ (n = 110), as a proportion of their parent population. i–l, Box plots of cell-type abundance: (i) Endodiv (proportion of endothelial cells) by GOLD stage (n = 123); (j) gCaptip (proportion of gCaps) by semiquantitative lobe emphysema (n = 128) and GOLD stage (n = 123); (k) aberrant basaloid cells (ABCs; proportion of all cells) by smoking status (N: never, F: former, C: current; n = 122); (l) CTHRC1⁺ fibroblasts (proportion of all cells) by smoking status (n = 122) and semiquantitative lobe emphysema (n = 127). c,d, Immunofluorescence staining representative of 5 subjects. i–k, Proportions multiplied by 100 and plotted on a log10 scale. f–l, Differential composition by sccomp (Bayesian sum-constrained β binomial with posterior probabilities converted to Benjamini–Hochberg adjusted probabilities), adjusted for age, sex, BMI, and smoking status; emphysema traits additionally adjusted for lobe (*FDR < 0.10, **FDR < 0.05, ***FDR < 0.01). e,i–l, Box plots depict the median (centerline), interquartile range (box), and 1.5× IQR whiskers.
Extended Data Fig. 4 Associations between immune cell populations and clinical traits in COPD.
a, Immunofluorescence staining for CD3 (red), CD19 (green), and CHIT1 (yellow) in lung tissue from study participants without COPD and GOLD IV COPD; nuclei are stained with DAPI (blue). Scale bars, 500 μm. b, Box plots of IM1 as a proportion of all macrophages stratified by GOLD stage (n = 140). c, Correlation between AMCSF (proportion of all macrophages) and exacerbation scores (n = 140). d, MDM as a proportion of all cells stratified by semiquantitative lobe emphysema (n = 127). e, Box plots of Macdiv as a proportion of all cells stratified by semiquantitative lobe emphysema (n = 127). f, Mast cells as a proportion of all cells stratified by lobe-specific semiquantitative emphysema (n = 127). g, B/plasma cells as a proportion of all cells stratified by smoking status (N: never, F: former, C: current), GOLD stage (n = 122), and semiquantitative lobe emphysema (n = 127). h, Migratory dendritic cells (mDCs) as a proportion of all cells, stratified by GOLD stage (n = 122) and semiquantitative lobe emphysema (n = 127). i, UMAP of lymphocyte cell populations, including CD4⁺ T cells, CD8⁺ T cells, regulatory T cells (Treg), mucosal-associated invariant T cells (MAIT), B cells, plasma cells, and natural killer (NK) cells. j, Feature plots of representative marker genes for immune cell populations shown in i. b,e,f,g,h, Proportions multiplied by 100 and plotted on a log10 scale. b–h, Differential composition by sccomp (Bayesian sum-constrained β binomial with posterior probabilities converted to Benjamini–Hochberg adjusted probabilities), adjusted for age, sex, BMI, and smoking status; emphysema traits additionally adjusted for lobe. (*FDR < 0.10, **FDR < 0.05, ***FDR < 0.01). b,d–h, Box plots depict the median (centerline), interquartile range (box), and 1.5× IQR whiskers.
Extended Data Fig. 5 Xenium spatial transcriptomics-defined cell populations.
a, UMAP of 706,966 high-confidence cells profiled by high-resolution Xenium in situ sequencing. Only cells passing quality control and assigned confident cell-type annotations are shown; an additional 101,907 QC passed cells lacking high-confidence labels are not displayed. b–e, Feature plots of selected marker genes: (b) epithelial and related markers; (c) endothelial and fibroblast markers; (d) myeloid and lymphoid markers; (e) B cell markers and inflammatory genes (IL6, CSF3, CXCL10, IL4R, CCL2, ADAMTS4). f, Dot plot of marker genes for cell populations identified in the Xenium spatial transcriptomics analysis. Dot size indicates the proportion of expressing cells; color denotes scaled expression.
Extended Data Fig. 6 Spatial niche maps of each tissue microarray (TMA) with clinical annotations.
Each TMA core is colored according to its assigned spatial niche. Right-side glyphs annotate each core’s clinical status: semiquantitative emphysema involvement (0 = none, 1 = <50%, 2 = 50–75%, 3 = >75%; circles) and GOLD stage (0, I/II, III, IV; triangles).
Extended Data Fig. 7 Differentially expressed and spatially resolved gene expression across inflammatory and macrophage niches.
a, Heatmap of scaled differentially expressed genes (rows) across spatial niches (columns), computed in a cell-agnostic manner (no cell labels or boundary stains). Niches include airway, large vessel (LV), low-immune parenchyma 1 and 2 (LI-parenchyma 1, LI-parenchyma 2), alveolar macrophages (AM), inflamed, remodeling, CHIT1/CD4/CD8 T-cell, and B-cell. Rows are cell types. b, Spatial expression maps (arbitrary units) overlaid on H&E for two pairs of adjacent TMA cores—one enriched for inflamed niches and the other enriched for low-immune (LI) parenchyma niches. Dot size and density are scaled for visualization and reflect relative, not absolute, transcript abundance. Scale bars, 1000 μm. c, Cell-type-resolved maps highlighting macrophage populations (MARCO, MCEMP1) and differentially expressed genes within macrophage niches (CXCL5, IL1B, IL1A, IL10RA). Scale bars, 500 μm. d, Colocalization of differentially expressed genes identified within macrophage niches and their spatial proximity to inflamed niches. Dot size and density are scaled for visualization and reflect relative, not absolute, transcript abundance. Scale bars, 1000 μm.
Extended Data Fig. 8 Cellular pathways and signaling networks associated with clinical outcomes and aberrant cell states.
a, Sankey diagram illustrating significant correlations between gene expression modules and clinical traits. b, Dot plot illustrating pathway enrichment of gene modules identified through marginal-effect analyses (circles) or mediation analyses (triangles) across key clinical traits in COPD. Clinical traits are represented along the panels on the x-axis, where the direction of the effect is multiplied by the −log10(enrichment P-value), and are grouped into spirometry (FEV1, FEV1/FVC), emphysema (emph), exacerbations (exacerbations, wheezing, infections), and symptoms (cough, dyspnea, SGRQ-symptoms). The pathways, categorized into overarching biological themes (inflammatory, remodeling and repair, aging, cell-migration, cell-ECM), are displayed on the y-axis. The size of each dot indicates the degree of pathway enrichment. Colors correspond to specific cell types. All displayed pathways are significantly enriched in their corresponding cell types after multiple-testing correction (Benjamini–Hochberg FDR < 0.05).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–10.
Supplementary Tables (download XLSX )
Supplementary Tables 1–12.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Y., Wei, H., Nouws, J. et al. Aberrant cellular communities underlying disease heterogeneity in chronic obstructive pulmonary disease. Nat Genet 58, 376–391 (2026). https://doi.org/10.1038/s41588-025-02480-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41588-025-02480-z