Abstract
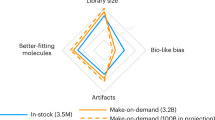
Virtual ligand libraries for ligand discovery have recently increased 10,000-fold. Whether this has improved hit rates and potencies has not been directly tested. Meanwhile, typically only dozens of docking hits are assayed, clouding hit-rate interpretation. Here we docked a 1.7 billion-molecule virtual library against β-lactamase, testing 1,521 new molecules and comparing the results to a 99 million-molecule screen where 44 molecules were tested. In a larger screen, hit rates improved twofold, more scaffolds were discovered and potency improved. Fifty-fold more inhibitors were found, supporting the idea that the large libraries harbor many more ligands than are being tested. In sampling smaller sets from the 1,521, hit rates only converged when several hundred molecules were tested. Hit rates and affinities improved steadily with docking score. It may be that as the scale of docking libraries and their testing grows, both ligands and our ability to rank them will improve.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The compounds docked in this study are freely available from the ZINC20 and ZINC22 databases, https://zinc20.docking.org and https://cartblanche22.docking.org. All compounds tested can be purchased from Enamine. Compound information including their ZINC ID, catalog ID, SMILES, DOCK score, ranking and affinity can be found in Supplementary Table 1. The synthetic procedures and purity information for the hits can be found in the Supplementary Note. Extensive docking-related files can be found at https://lsd.docking.org. DOCK3.8 is freely available for noncommercial research at https://dock.compbio.ucsf.edu/DOCK3.8/. A web-based version is available without restriction at https://blaster.docking.org/. X-ray structures and maps are available in the PDB under accession numbers 9C81 (Z4462773688), 9C6P (Z6615017509), 9C84 (Z6615020275) and 9DHL (Z6615017782), respectively. Source data are provided with this paper.
References
Lyu, J. et al. Ultra-large library docking for discovering new chemotypes. Nature 566, 224–229 (2019).
Gorgulla, C. et al. An open-source drug discovery platform enables ultra-large virtual screens. Nature 580, 663–668 (2020).
Stein, R. M. et al. Virtual discovery of melatonin receptor ligands to modulate circadian rhythms. Nature 579, 609–614 (2020).
Alon, A. et al. Structures of the sigma(2) receptor enable docking for bioactive ligand discovery. Nature 600, 759–764 (2021).
Sadybekov, A. A. et al. Synthon-based ligand discovery in virtual libraries of over 11 billion compounds. Nature 601, 452–459 (2022).
Fink, E. A. et al. Structure-based discovery of nonopioid analgesics acting through the α2A-adrenergic receptor. Science 377, eabn7065 (2022).
Singh, I. et al. Structure-based discovery of conformationally selective inhibitors of the serotonin transporter. Cell 186, 2160–2175.e17 (2023).
Gahbauer, S. et al. Docking for EP4R antagonists active against inflammatory pain. Nat. Commun. 14, 8067 (2023).
Sadybekov, A. V. & Katritch, V. Computational approaches streamlining drug discovery. Nature 616, 673–685 (2023).
Gorgulla, C. et al. A multi-pronged approach targeting SARS-CoV-2 proteins using ultra-large virtual screening. iScience 24, 102021 (2021).
Klarich, K., Goldman, B., Kramer, T., Riley, P. & Walters, W. P. Thompson sampling─an efficient method for searching ultralarge synthesis on demand databases. J. Chem. Inf. Model. 64, 1158–1171 (2024).
Walters, W. P. Virtual chemical libraries. J. Med. Chem. 62, 1116–1124 (2019).
Gorgulla, C., Jayaraj, A., Fackeldey, K. & Arthanari, H. Emerging frontiers in virtual drug discovery: from quantum mechanical methods to deep learning approaches. Curr. Opin. Chem. Biol. 69, 102156 (2022).
Lyu, J., Irwin, J. J. & Shoichet, B. K. Modeling the expansion of virtual screening libraries. Nat. Chem. Biol. 19, 712–718 (2023).
Weston, G. S., Blazquez, J., Baquero, F. & Shoichet, B. K. Structure-based enhancement of boronic acid-based inhibitors of AmpC beta-lactamase. J. Med. Chem. 41, 4577–4586 (1998).
Powers, R. A., Morandi, F. & Shoichet, B. K. Structure-based discovery of a novel, noncovalent inhibitor of AmpC beta-lactamase. Structure 10, 1013–1023 (2002).
Feng, B. Y., Shelat, A., Doman, T. N., Guy, R. K. & Shoichet, B. K. High-throughput assays for promiscuous inhibitors. Nat. Chem. Biol. 1, 146–148 (2005).
Feng, B. Y. et al. A high-throughput screen for aggregation-based inhibition in a large compound library. J. Med. Chem. 50, 2385–2390 (2007).
Eidam, O. et al. Design, synthesis, crystal structures, and antimicrobial activity of sulfonamide boronic acids as beta-lactamase inhibitors. J. Med. Chem. 53, 7852–7863 (2010).
Babaoglu, K. et al. Comprehensive mechanistic analysis of hits from high-throughput and docking screens against beta-lactamase. J. Med. Chem. 51, 2502–2511 (2008).
Gorgulla, C. et al. VirtualFlow 2.0—the next generation drug discovery platform enabling adaptive screens of 69 billion molecules. Preprint at bioRxiv https://doi.org/10.1101/2023.04.25.537981 (2023).
Bender, B. J. et al. A practical guide to large-scale docking. Nat. Protoc. 16, 4799–4832 (2021).
Fassio, A. V. et al. Prioritizing virtual screening with interpretable interaction fingerprints. J. Chem. Inf. Model. 62, 4300–4318 (2022).
Wu, Y. et al. Identifying artifacts from large library docking. J. Med. Chem. 67, 16796–16806 (2024).
Cheng, Y. & Prusoff, W. H. Relationship between the inhibition constant (K1) and the concentration of inhibitor which causes 50 per cent inhibition (I50) of an enzymatic reaction. Biochem. Pharmacol. 22, 3099–3108 (1973).
McGovern, S. L., Helfand, B. T., Feng, B. & Shoichet, B. K. A specific mechanism of nonspecific inhibition. J. Med. Chem. 46, 4265–4272 (2003).
Feng, B. Y. & Shoichet, B. K. A detergent-based assay for the detection of promiscuous inhibitors. Nat. Protoc. 1, 550–553 (2006).
O’Donnell, H. R., Tummino, T. A., Bardine, C., Craik, C. S. & Shoichet, B. K. Colloidal aggregators in biochemical SARS-CoV-2 repurposing screens. J. Med. Chem. 64, 17530–17539 (2021).
Walters, W. P. & Namchuk, M. Designing screens: how to make your hits a hit. Nat. Rev. Drug Discov. 2, 259–266 (2003).
Tirado-Rives, J. & Jorgensen, W. L. Contribution of conformer focusing to the uncertainty in predicting free energies for protein-ligand binding. J. Med. Chem. 49, 5880–5884 (2006).
Irwin, J. J. & Shoichet, B. K. Docking screens for novel ligands conferring new biology. J. Med. Chem. 59, 4103–4120 (2016).
Ackloo, S. et al. CACHE (Critical Assessment of Computational Hit-finding Experiments): a public–private partnership benchmarking initiative to enable the development of computational methods for hit-finding. Nat. Rev. Chem. 6, 287–295 (2022).
Gentile, F. et al. Artificial intelligence-enabled virtual screening of ultra-large chemical libraries with deep docking. Nat. Protoc. 17, 672–697 (2022).
Yang, Y. et al. Efficient exploration of chemical space with docking and deep learning. J. Chem. Theory Comput. 17, 7106–7119 (2021).
Chen, Y., McReynolds, A. & Shoichet, B. K. Re-examining the role of Lys67 in class C beta-lactamase catalysis. Protein Sci. 18, 662–669 (2009).
Riley, B. T. et al. qFit 3: protein and ligand multiconformer modeling for X-ray crystallographic and single-particle cryo-EM density maps. Protein Sci. 30, 270–285 (2021).
Fischer, M., Coleman, R. G., Fraser, J. S. & Shoichet, B. K. Incorporation of protein flexibility and conformational energy penalties in docking screens to improve ligand discovery. Nat. Chem. 6, 575–583 (2014).
Word, J. M., Lovell, S. C., Richardson, J. S. & Richardson, D. C. Asparagine and glutamine: using hydrogen atom contacts in the choice of side-chain amide orientation. J. Mol. Biol. 285, 1735–1747 (1999).
Meng, E. C., Shoichet, B. K. & Kuntz, I. D. Automated docking with grid-based energy evaluation. J. Comput. Chem. 13, 505–524 (1992).
Gallagher, K. & Sharp, K. Electrostatic contributions to heat capacity changes of DNA-ligand binding. Biophys. J. 75, 769–776 (1998).
Sharp, K. A. Polyelectrolyte electrostatics: salt dependence, entropic, and enthalpic contributions to free energy in the nonlinear Poisson–Boltzmann model. Biopolymers 36, 227–243 (1995).
Mysinger, M. M. & Shoichet, B. K. Rapid context-dependent ligand desolvation in molecular docking. J. Chem. Inf. Model. 50, 1561–1573 (2010).
Coleman, R. G., Carchia, M., Sterling, T., Irwin, J. J. & Shoichet, B. K. Ligand pose and orientational sampling in molecular docking. PLoS ONE 8, e75992 (2013).
Stein, R. M. et al. Property-unmatched decoys in docking benchmarks. J. Chem. Inf. Model. 61, 699–714 (2021).
Tingle, B. I. et al. ZINC-22—a free multi-billion-scale database of tangible compounds for ligand discovery. J. Chem. Inf. Model. 63, 1166–1176 (2023).
Eidam, O. et al. Fragment-guided design of subnanomolar beta-lactamase inhibitors active in vivo. Proc. Natl Acad. Sci. USA 109, 17448–17453 (2012).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Liebschner, D. et al. Polder maps: improving OMIT maps by excluding bulk solvent. Acta Crystallogr. D 73, 148–157 (2017).
Acknowledgements
This work is supported by US National Institutes of Health (NIH) grant nos. R35GM122481 (to B.K.S.), GM71896 (to J.J.I.) and GM145238 (to J.S.F.) and a Damon Runyon Postdoctoral Research Fellowship (F.L.). We thank ChemAxon for JChem, OpenEye Scientific software for Omega and Schrödinger LLC for the Maestro suite. We thank G. Meigs and J. Holton for their assistance at Beamline 8.3.1 at the Advanced Light Source, operated by UCSF with NIH grant nos. R01 GM124149 for technology development and P30 GM124169 for beamline operations, and the Integrated Diffraction Analysis Technologies program of the US Department of Energy Office of Biological and Environmental Research. We thank K. Srinivasan for his assistance on data collection. The Advanced Light Source (Berkeley, CA, USA) is a national user facility operated by Lawrence Berkeley National Laboratory on behalf of the US Department of Energy under contract number DE-AC02-05CH11231, Office of Basic Energy Sciences.
Author information
Authors and Affiliations
Contributions
F.L. conducted the docking screens and the ligand optimization assisted by S.F.V. and advised by B.K.S. F.L. and I.S.G. conducted the in vitro enzymatic assays, with early assistance from S.F.V. F.L. determined the structures by X-ray crystallography, with assistance from V.B. and X.X., advised by J.S.F. F.L. and O.M. did the analysis with advice from M.S.S. Aggregation studies were conducted by K.F.-V. and I.S.G. J.J.I. developed and prepared the make-on-demand library assisted with large library docking strategies. D.S.R. and Y.S.M. supervised compound synthesis of Enamine compounds purchased from the ZINC22 database and the 46 billion catalog library.
Corresponding authors
Ethics declarations
Competing interests
B.K.S. is a founder of Epiodyne, Inc.; BlueDolphin, LLC; and Deep Apple Therapeutics, Inc., and serves on the SAB of Schrodinger LLC and of Vilya Therapeutics, and on the SRB of Genentech. J.J.I. cofounded Deep Apple Therapeutics, Inc., and BlueDolphin, LLC. J.S.F. is a consultant for, has equity in and receives research support from Relay Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Artem Cherkasov, Tyuji Hoshino and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Molecules with artifactually favorable scores disrupt the distribution of docking scores and concentrate among the top-ranking docked molecules.
a, DOCK scores of molecules against AmpC. b, DOCK scores of molecules against σ2 receptor4.
Extended Data Fig. 2 Concentration-response curves for 17 of the new docking-derived AmpC inhibitors.
Nitrocefin was kept at a constant concentration of 100 μM (for positive control ZINC549719643, new inhibitors Z6615018018, Z6615017509, Z6615022372, Z6615017782, Z6615020275, Z6615019214 and Z6615014610) or 50 μM (for positive control ZINC339304163, new inhibitors Z2275216423, Z6615017736, Z2940316600, Z2940315182, Z6615019960, Z2940322517, Z6615017422, Z6615015266, Z6615016774 and Z6615155291). The estimated Ki is calculated based on the Kd of nitrocefin (180 μM) calculated from a Lineweaver-Burk analysis. The previously reported Ki for ZINC549719643 is 77 nM1 and for ZINC339304163 is 1.25 μM1. Data represent mean ± s.d.s from three biological replicates.
Extended Data Fig. 3 Lineweaver-Burk plots of seven of the new AmpC inhibitors (a-g).
ZINC339304163 is a positive control inhibitor identified in a previous docking campaign1.
Extended Data Fig. 4 Electron density omit maps of the AmpC inhibitors.
a-d, Polder omit maps of the inhibitors (3σ).
Extended Data Fig. 5 Comparative analysis of hit rates from large-scale and small-scale AmpC screens with statistical validation.
a, The hit rates (number of actives/total tested) of the 1.7 Billion screen (blue bar; 8.26%) versus the 99 Million screen (orange bar; 2.27%) with a hit defined as less than 100 µM. b, The hit rates (number of actives/total tested) of the 1.7 Billion screen (blue bar; 2.47%) versus the 99 Million screen (orange bar; 2.27%) with a hit defined as less than 30 µM. c, The hit rates of all manually picked molecules of the 1.7 Billion screen (blue bar; 21.14%) versus the 99 Million screen (orange bar; 11.4%). d, The hit rates of the top 44 manually picked molecules of the 1.7 Billion screen (blue bar; 47.7%) versus the 99 Million screen (orange bar; 11.4%). e, Hit rates from the manually picked, experimentally tested molecules of the 99 Million and 1.7 Billon screens (44 and 626 molecules, respectively), referred to as the “Small” and “Big” screens. For each set, 44 or 626 molecules were resampled for 10,000 bootstrap iterations, and the mean of the resampled hit rates is shown in parenthesis. P-values for the null hypothesis that the difference between two resampled distributions is zero are provided. For panels a-d, a two-sided Z-test was used to compare the hit rates of the two screens, under the assumption that the data followed a normal distribution. For panel e, P-values were obtained from a one-tailed non-parametric bootstrap test (10,000 iterations) comparing the means of the resampled distributions, with no assumption of normality.
Extended Data Fig. 6 The impact of testing fewer molecules on hit rate confidence.
a, For 327 molecules tested against the σ2 receptor, each sample size is randomly drawn 30 times and the resulting hit rates were plotted. The error bars represent s.d.s of the hit rates. b, The impact of randomly purchasing 44 and 139 molecules out of 327 molecules for testing on hit rates with different affinity cutoffs. Each sample size is drawn 30 times and the resulting hit rates were plotted. The error bars represent s.d.s of the hit rates. c, For 371 molecules tested against the D4 receptor, each sample size is randomly drawn 30 times and the resulting hit rates were plotted. The error bars represent s.d.s of the hit rates. d, The impact of randomly purchasing 44 and 139 molecules out of 371 molecules for testing on hit rates with different affinity cutoffs. Each sample size is drawn 30 times and the resulting hit rates were plotted. Data represent mean ± s.d.s of the hit rates.
Extended Data Fig. 7 Examples of the new warheads and chemotypes from the AmpC screen, in their docked poses in the enzyme active site.
a, docked pose of Z6615021877 (Ki = 121 μM). b, docked pose of Z2607647274 (Ki = 47 μM). c, docked pose of Z6615146667 (Ki = 173 μM). d, docked pose of Z6615020742 (Ki = 184 μM). e, docked pose of Z2610488449 (Ki = 12 μM). f, docked pose of Z4173922012 (Ki = 230 μM). g, docked pose of Z6615146331 (Ki = 214 μM). h, docked pose of Z6722203632 (Ki = 465 μM). i, docked pose of Z5389129999 (Ki = 298 μM). The Ki values for Z6615021877 and Z2610488449 were calculated using Lineweaver-Burk plots, while the rest were determined based on the three-point inhibition assays.
Extended Data Fig. 8 Docking poses of the some of the top scoring molecules.
Docking poses of ZINCop00000kUi3Y, ZINCov000006qjGM, ZINCpM00000d7IVN, ZINCpw000006Kp2I, ZINCqs000002TbmO and ZINCpa00000sPJnu are shown.
Extended Data Fig. 9 Hit rate of experimentally tested compounds plotted against DOCK scores with different affinity cutoffs.
a, Hit rates of all compounds tested (1,447 well-behaved molecules among 1,521 purchased) plotted against DOCK scores with four different affinity cutoffs: < 400, <137, <40 and <13 μM. b, Hit rates of manually picked compounds (687 compounds) plotted against DOCK scores with four different affinity cutoffs: <400, <137, <40 and <13 μM.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 2–4, Data 5 and Table 6.
Supplementary Table 1 (download XLSX )
Supplementary Table 1: Molecules tested against AmpC β-lactamase.
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, F., Mailhot, O., Glenn, I.S. et al. The impact of library size and scale of testing on virtual screening. Nat Chem Biol 21, 1039–1045 (2025). https://doi.org/10.1038/s41589-024-01797-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-024-01797-w
This article is cited by
-
Deciphering DEL pocket patterns through contrastive learning
Nature Communications (2026)
-
Unfreezing structural biology for drug discovery
Nature Chemical Biology (2026)
-
SLICE (SMARTS and Logic In ChEmistry): fast generation of molecules using advanced chemical synthesis logic and modern coding style
Journal of Cheminformatics (2025)
-
Navigating structure-based drug discovery with emerging innovations in physics- and knowledge-based approaches
npj Drug Discovery (2025)