Abstract
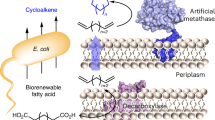
Artificial metalloenzymes (ArMs) integrated within whole cells have emerged as promising catalysts; however, their sensitivity to metal centers remains a systematic challenge, resulting in diminished activity and turnover. Here we address this issue by inducing in cellulo liquid–liquid phase separation through a self-labeling fusion protein, HaloTag–SNAPTag. This strategy creates membraneless, isolated liquid condensates within Escherichia coli as protective compartments for the assembly of ArMs using the same fusion protein. The approach allows for high ArM loading and stabilization by localizing the ArMs within the phase-separated regions. Consequently, the performance of ArM-based whole-cell catalysts is improved, with a demonstrated turnover per cell of up to 7.1 × 109 for the olefin metathesis reaction. Furthermore, we apply this to an engineered E. coli system in live mice, where host bacterial cells confine the metal catalytic species, and in a mouse colorectal cancer model, where ArM-containing whole-cell catalysts mediate concurrent reactions to activate prodrugs.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data supporting the findings of this study are available within the paper, the Extended Data Figs. 1–10 and the Supplementary Information. Original raw data are available as source data files that are provided with this paper or from the corresponding author upon request. Source data are provided with this paper.
References
Wittwer, M. et al. Engineering and emerging applications of artificial metalloenzymes with whole cells. Nat. Catal. 4, 814–827 (2021).
Chordia, S., Narasimhan, S., Lucini Paioni, A., Baldus, M. & Roelfes, G. In vivo assembly of artificial metalloenzymes and application in whole-cell biocatalysis. Angew. Chem. Int. Ed. 60, 5913–5920 (2021).
Bloomer, B. J., Clark, D. S. & Hartwig, J. F. Progress, challenges, and opportunities with artificial metalloenzymes in biosynthesis. Biochemistry 62, 221–228 (2023).
Coelho, P. S., Brustad, E. M., Kannan, A. & Arnold, F. H. Olefin cyclopropanation via carbene transfer catalyzed by engineered cytochrome P450 enzymes. Science 339, 307–310 (2013).
Vornholt, T. et al. Systematic engineering of artificial metalloenzymes for new-to-nature reactions. Sci. Adv. 7, abe4208 (2021).
Fischer, S., Ward, T. R. & Liang, A. D. Engineering a metathesis-catalyzing artificial metalloenzyme based on HaloTag. ACS Catal. 11, 6343–6347 (2021).
Key, H. M., Dydio, P., Clark, D. S. & Hartwig, J. F. Abiological catalysis by artificial haem proteins containing noble metals in place of iron. Nature 534, 534–537 (2016).
Zastrow, M. L., Peacock, A. F., Stuckey, J. A. & Pecoraro, V. L. Hydrolytic catalysis and structural stabilization in a designed metalloprotein. Nat. Chem. 4, 118–123 (2011).
Eda, S. et al. Biocompatibility and therapeutic potential of glycosylated albumin artificial metalloenzymes. Nat. Catal. 2, 780–792 (2019).
Kato, S., Onoda, A., Schwaneberg, U. & Hayashi, T. Evolutionary engineering of a Cp*Rh(III) complex-linked artificial metalloenzyme with a chimeric β-barrel protein scaffold. J. Am. Chem. Soc. 145, 8285–8290 (2023).
Renata, H., Wang, Z. J. & Arnold, F. H. Expanding the enzyme universe: accessing non-natural reactions by mechanism-guided directed evolution. Angew. Chem. Int. Ed. 54, 3351–3367 (2015).
Hyster, T. K. & Ward, T. R. Genetic optimization of metalloenzymes: enhancing enzymes for non-natural reactions. Angew. Chem. Int. Ed. 55, 7344–7357 (2016).
Huang, J. et al. Unnatural biosynthesis by an engineered microorganism with heterologously expressed natural enzymes and an artificial metalloenzyme. Nat. Chem. 13, 1186–1191 (2021).
Davis, H. J. & Ward, T. R. Artificial metalloenzymes: challenges and opportunities. ACS Cent. Sci. 5, 1120–1136 (2019).
Liu, Z. et al. Assembly and evolution of artificial metalloenzymes within E. coli Nissle 1917 for enantioselective and site-selective functionalization of C–H and C–C bonds. J. Am. Chem. Soc. 144, 883–890 (2022).
Wu, S., Zhou, Y., Gerngross, D., Jeschek, M. & Ward, T. R. Chemo-enzymatic cascades to produce cycloalkenes from bio-based resources. Nat. Commun. 10, 5060 (2019).
Okamoto, Y. et al. A cell-penetrating artificial metalloenzyme regulates a gene switch in a designer mammalian cell. Nat. Commun. 9, 1943 (2018).
Samanta, A., Sabatino, V., Ward, T. R. & Walther, A. Functional and morphological adaptation in DNA protocells via signal processing prompted by artificial metalloenzymes. Nat. Nanotechnol. 15, 914–921 (2020).
Yu, F. et al. Protein design: toward functional metalloenzymes. Chem. Rev. 114, 3495–3578 (2014).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2018).
Creus, M. et al. X-ray structure and designed evolution of an artificial transfer hydrogenase. Angew. Chem. Int. Ed. 47, 1400–1404 (2008).
Volker, T., Dempwolff, F., Graumann, P. L. & Meggers, E. Progress towards bioorthogonal catalysis with organometallic compounds. Angew. Chem. Int. Ed. 53, 10536–10540 (2014).
Sasmal, P. K. et al. Catalytic azide reduction in biological environments. ChemBioChem 13, 1116–1120 (2012).
Schunck, N. S. & Mecking, S. In vivo olefin metathesis in microalgae upgrades lipids to building blocks for polymers and chemicals. Angew. Chem. Int. Ed. 61, e202211285 (2022).
Jeschek, M. et al. Directed evolution of artificial metalloenzymes for in vivo metathesis. Nature 537, 661–665 (2016).
Zhao, J. et al. Genetic engineering of an artificial metalloenzyme for transfer hydrogenation of a self-immolative substrate in Escherichia coli’s periplasm. J. Am. Chem. Soc. 140, 13171–13175 (2018).
Jeschek, M., Panke, S. & Ward, T. R. Periplasmic screening for artificial metalloenzymes. Methods Enzymol. 580, 539–556 (2016).
Wilson, Y. M., Durrenberger, M., Nogueira, E. S. & Ward, T. R. Neutralizing the detrimental effect of glutathione on precious metal catalysts. J. Am. Chem. Soc. 136, 8928–8932 (2014).
Rottmann, P., Ward, T. & Panke, S. Compartmentalization—a prerequisite for maintaining and changing an identity. Chimia 70, 428–431 (2016).
Cao, S. et al. Dipeptide coacervates as artificial membraneless organelles for bioorthogonal catalysis. Nat. Commun. 15, 39 (2024).
Sun, Y. et al. Phase-separating peptides for direct cytosolic delivery and redox-activated release of macromolecular therapeutics. Nat. Chem. 14, 274–283 (2022).
Garabedian, M. V. et al. Designer membraneless organelles sequester native factors for control of cell behavior. Nat. Chem. Biol. 17, 998–1007 (2021).
Wei, S. P. et al. Formation and functionalization of membraneless compartments in Escherichia coli. Nat. Chem. Biol. 16, 1143–1148 (2020).
Alberti, S., Gladfelter, A. & Mittag, T. Considerations and challenges in studying liquid–liquid phase separation and biomolecular condensates. Cell 176, 419–434 (2019).
Boeynaems, S. et al. Protein phase separation: a new phase in cell biology. Trends Cell Biol. 28, 420–435 (2018).
Tong, X. et al. Liquid–liquid phase separation in tumor biology. Signal Transduct. Target. Ther. 7, 221 (2022).
Hilditch, A. T. et al. Assembling membraneless organelles from de novo designed proteins. Nat. Chem. 16, 89–97 (2024).
Hirose, T., Ninomiya, K., Nakagawa, S. & Yamazaki, T. A guide to membraneless organelles and their various roles in gene regulation. Nat. Rev. Mol. Cell Biol. 24, 288–304 (2023).
Wang, Y. et al. Phase‐separated multienzyme compartmentalization for terpene biosynthesis in a prokaryote. Angew. Chem. Int. Ed. 61, e202203909 (2022).
Lohner, P. et al. Inside a shell-organometallic catalysis inside encapsulin nanoreactors. Angew. Chem. Int. Ed. 60, 23835–23841 (2021).
Liu, Y. et al. AgHalo: a facile fluorogenic sensor to detect drug-induced proteome stress. Angew. Chem. Int. Ed. 56, 8672–8676 (2017).
Liu, Y. et al. A HaloTag-based multicolor fluorogenic sensor visualizes and quantifies proteome stress in live cells using solvatochromic and molecular rotor-based fluorophores. Biochemistry 57, 4663–4674 (2018).
Fares, M. et al. A molecular rotor-based halo-tag ligand enables a fluorogenic proteome stress sensor to detect protein misfolding in mildly stressed proteome. Bioconjug. Chem. 29, 215–224 (2018).
Wilson, C., Lewis, K. A., Fitzkee, N. C., Hough, L. E. & Whitten, S. T. ParSe 2.0: a web tool to identify drivers of protein phase separation at the proteome level. Protein Sci. 32, e4756 (2023).
Farag, M., Borcherds, W. M., Bremer, A., Mittag, T. & Pappu, R. V. Phase separation of protein mixtures is driven by the interplay of homotypic and heterotypic interactions. Nat. Commun. 14, 5527 (2023).
Farag, M. et al. Condensates formed by prion-like low-complexity domains have small-world network structures and interfaces defined by expanded conformations. Nat. Commun. 13, 7722 (2022).
Lin, Y., Protter, D. S., Rosen, M. K. & Parker, R. Formation and maturation of phase-separated liquid droplets by RNA-binding proteins. Mol. Cell 60, 208–219 (2015).
Jin, X. et al. Membraneless organelles formed by liquid–liquid phase separation increase bacterial fitness. Sci. Adv. 7, eabh2929 (2021).
Yeong, V., Werth, E. G., Brown, L. M. & Obermeyer, A. C. Formation of biomolecular condensates in bacteria by tuning protein electrostatics. ACS Cent. Sci. 6, 2301–2310 (2020).
Peskett, T. R. et al. A liquid to solid phase transition underlying pathological huntingtin exon 1 aggregation. Mol. Cell 70, 588–601 (2018).
Watanabe, S. et al. Aggresome formation and liquid–liquid phase separation independently induce cytoplasmic aggregation of TAR DNA-binding protein 43. Cell Death Dis. 11, 909 (2020).
Kroschwald, S. et al. Promiscuous interactions and protein disaggregases determine the material state of stress-inducible RNP granules. eLife 4, e06807 (2015).
Chan, T. R., Hilgraf, R., Sharpless, K. B. & Fokin, V. V. Polytriazoles as copper(I)-stabilizing ligands in catalysis. Org. Lett. 6, 2853–2855 (2004).
Lu, Q. et al. A dense-shell macromolecular scaffold for catalyst- or substrate-guided catalysis in a cellular environment. ACS Mater. Lett. 2, 89–94 (2020).
Deng, Y. et al. A membrane-embedded macromolecular catalyst with substrate selectivity in live cells. J. Am. Chem. Soc. 145, 1262–1272 (2023).
Sivakumar, K. et al. A fluorogenic 1,3-dipolar cycloaddition. Org. Lett. 6, 4603–4606 (2004).
Bielawski, C. W. & Grubbs, R. H. Living ring-opening metathesis polymerization. Prog. Polym. Sci. 32, 1–29 (2007).
Geng, J. et al. Radical polymerization inside living cells. Nat. Chem. 11, 578–586 (2019).
Shen, Q. et al. Intracellular radical polymerization of paclitaxel-bearing acrylamide for self-inflicted apoptosis of cancer cells. ACS Mater. Lett. 3, 1307–1314 (2021).
Vidal, C., Tomas-Gamasa, M., Destito, P., Lopez, F. & Mascarenas, J. L. Concurrent and orthogonal gold(I) and ruthenium(II) catalysis inside living cells. Nat. Commun. 9, 1913 (2018).
Ofori Atta, L., Zhou, Z. & Roelfes, G. In vivo biocatalytic cascades featuring an artificial-enzyme-catalysed new-to-nature reaction. Angew. Chem. Int. Ed. 62, e202214191 (2023).
Hwang, E. T. & Lee, S. Multienzymatic cascade reactions via enzyme complex by immobilization. ACS Catal. 9, 4402–4425 (2019).
Mokhtari, R. B. et al. Combination therapy in combating cancer. Oncotarget 8, 38022–38043 (2017).
Stritzker, J. et al. Tumor-specific colonization, tissue distribution, and gene induction by probiotic Escherichia coli Nissle 1917 in live mice. Int. J. Med. Microbiol. 297, 151–162 (2007).
Harimoto, T. et al. A programmable encapsulation system improves delivery of therapeutic bacteria in mice. Nat. Biotechnol. 40, 1259–1269 (2022).
Nasibullin, I. et al. Synthetic prodrug design enables biocatalytic activation in mice to elicit tumor growth suppression. Nat. Commun. 13, 39 (2022).
Acknowledgements
We would like to acknowledge the Joint National Natural Science Foundation of China – Israel Science Foundation (NSFC-ISF) Research Grant (22361142830 to Y.B. and 3119/23 to N.G.L.), the Natural Science Foundation of Hunan Province (grants 2022RC1107 and 2024JJ2010 to Y.B.) and the Hunan Provincial Innovation Foundation for Postgraduates (grant CX20220389 to T.W.) for providing funding support. We would like to thank Prof. Q. Min of Nanjing University and the Analysis and Testing Center of Hunan University for their assistance in characterizations.
Author information
Authors and Affiliations
Contributions
T.W. and Y.B. proposed the idea, designed the experiments, analyzed the results, collated the experimental data, prepared the figure panels and tables, and drafted the paper. T.W. and Y.F. cloned the plasmids and expressed the proteins (HaloTag, HS, HSTruc and HC). T.W., X.C. and Y.F. performed the animal studies. T.W., Y.F., G.H. and Z.C. prepared the cofactors and prodrugs. T.W. and Y.D. conducted the TON and catalytic kinetics measurements. T.W. performed the FRAP studies with the help of Y.W. and performed all other experiments. A.Y. offered ideas for validating the intracellular LLPS mechanism. T.W., X.F., N.G.L. and Y.B. revised the paper. T.W., X.F., N.G.L. and Y.B. contributed to funding collection. X.F. and Y.B. provided laboratory spaces and necessities for the experiments. Y.B. supervised the project. All authors read, discussed and analyzed the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Sylvestre Bonnet, Takakazu Nakabayashi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Illustrations on the self-labeling property and structural analysis of HS protein.
a, Mechanistic illustration of the self-labeling reactions of HaloTag–SNAPTag fusion protein. HaloTag is a modified haloalkane dehalogenase that can have its Asp106 residue irreversibly displace the chloride on a chloroalkyl substrate. SNAPTag is a truncated and modified human O6-alkylguanine-DNA alkyltransferase that can react with an O6-benzylguanine (BG) substrate with its Cys145 residue and displace the O6-benzylguaninyl moiety. The two coupling reactions are both bioorthogonal and efficient. b, Phase-separating region prediction made by ParSe v2 tool. The (EAAAK)4 linker sequence was predicted to be the major intrinsically disordered region (IDR) that promotes LLPS of the protein.
Extended Data Fig. 2 Additional data showing the phase-separation and in cellulo compartmentalization of HS–E. coli.
a, Cartoon illustration and confocal microscope images of compartmentalized HS–E. coli cells, triggered by Tris–Cl3. Bacterial genomic DNA was excluded from the protein compartments. The curve figure on the right denotes the fluorescent intensity of pixels along the line indicated by the yellow arrow in the fluorescently labeled image. For the kinetics of LLPS compartment formation in HS–E. coli, see Supplementary Fig. 1. Scale bar = 5 μm. b, Photographs of phase-separated HS protein in vitro in dextran-70 solution, which provides a crowding environment. Various controls without phase separation are also provided. Phase-separated and homogeneous samples are labeled in blue and black, respectively. c, Confocal microscope images of compartmentalized HS–E. coli cells, triggered by TTA–Cl3 (35 µM). Rho–BG was used to label the HS compartments. n ≥ 3 independent experiments; representative images are shown for each experiment described in the figure. Scale bar = 5 µm.
Extended Data Fig. 3 Experimental results showing the cause and composition of the formed LLPS compartments in E. coli.
a, Results from PAGE and confocal microscopy indicated HaloTag could be dimerized like HS with the aid of TTA–Cl3, but HaloTag-expressing E. coli (H-E. coli) did not show LLPS phenomenon 6 h or 18 h after cross-linker treatment. HS–E. coli showed pronounced LLPS under the same condition. Scale bar = 5 μm. b, Truncated HS protein with (EAAAK)4 linker (HaloTag–SNAPTagTruc) showed LLPS and compartment formation upon TTA–Cl3 treatment in E. coli. A HaloTag–cutinase fusion with the same (EAAAK)4 linker also showed similar LLPS phenomenon upon TTA–Cl3 treatment. Scale bar = 5 μm. c, Phase-separating region prediction made by ParSe v2 tool for HaloTag–SNAPTagTruc and HaloTag–Cutinase fusions. The (EAAAK)4 linker sequence was again predicted to be the major intrinsically disordered region (IDR) that promotes LLPS of the proteins. These results demonstrated the importance of this (EAAAK)4 linker sequence in promoting protein LLPS, which was required by the construction of catalytic compartments in cells. d,e, Gel analysis results of cross-linked HS units formed in vitro (d) and in cellulo (e), using TTA–Cl3 or Tris–Cl3 as the cross-linker and LLPS promoter. Conditions marked with + indicate the corresponding reagent is used, and − indicates unused. The in vitro assembly was performed using purified HS protein, while for the in cellulo assembly, the HS-expressing E. coli was treated with the cross-linker, lysed and analyzed on PAGE. n = 3 independent experiments; representative images are shown for each experiment described in a,b,d,e.
Extended Data Fig. 4 Additional explorations on the nature of HS protein’s LLPS.
a, Fluorescence recovery from the in vitro FRAP experiment indicating the liquid-like nature of the phase-separated HS protein. Additional in cellulo FRAP results and related discussions are provided in Supplementary Fig. 3 in the Supplementary Information. Scale bar = 50 µm. b, Confocal microscopy images of live compartmentalized HS–E. coli stained by Nile red (0.01% wt/vol) or thioflavin T (50 μM) for 1 h at 25 °C. The staining pattern suggested that the LLPS compartments should have provided a relatively hydrophobic microenvironment, which selectively uptakes the more hydrophobic dye, Nile red. n = 3 independent experiments; representative results are shown for each experiment described in a,b. Scale bar = 5 µm. c, LLPS compartment (labeled with Rho–BG) disruption by incubating with 1,6-hexanediol (1,6-HD, 20% wt/vol). 1,6-HD is known to partially dissolve phase-separated proteins by disrupting hydrophobic interactions within the droplets. This result suggested that hydrophobic interaction might be the primary driving force for the LLPS of HS protein. n = 4 biological replicates; representative images are shown. Scale bar = 5 μm. d, Statistics on 1,6-HD’s disruptive effect on LLPS compartments at different stages. n = 4 biological replicates. Data are presented as mean ± s.d.
Extended Data Fig. 5 Characterization of ArMs assembled in E. coli cells.
a, Analysis of HS protein’s assembly and self-labeling conjugation in vitro and in cellulo. HS: in vitro reactions using purified HS protein; HS–E. coli: reactions conducted in live HS–E. coli and the cell lysate was used for analysis. For the fluorescence scanning result of SDS–PAGE, HS was labeled with Rho–BG or TMR-Cl. Conditions for each individual lane were indicated in the legend above. A detailed protocol is provided in the Supplementary Methods—Gel-based visualization of the in vitro HS assembly (Extended Data Figs. 2d and 5a). Please note the cross-linked HS cannot be labeled by TMR-Cl or HG–Ru–Cl because the chloride-reactive site is taken by the alkyl chloride cross-linker. n = 3 independent experiments; a representative image is shown. b, Ruthenium content analysis of the protein bands cut from the SDS–PAGE gel by ICP-MS. The results indicated high specificity of the conjugation of ruthenium cofactor with HS. n = 3 biological replicates; a representative image is shown. c, TEM–EDS images of HS–E. coli containing Cu–HS–ArMs, showing the enrichment of copper in the LLPS compartments. n = 3 independent experiments; representative images are shown. d, Copper content analysis of the protein bands cut from the SDS–PAGE gel by ICP-MS. The results indicated the necessity of TTA ligands for copper to be incorporated into the protein scaffold and the specificity of copper binding to the TTA–HS scaffolds. n = 3 biological replicates; a representative image is shown. All data are presented as mean ± s.d. where relevant.
Extended Data Fig. 6 In vitro performance evaluation of HS–ArMs in hostile environments.
a, Performance comparison of small molecular ruthenium complex (HG–Ru–Cl) and Ru–HS–ArMs, when mediating the fluorogenic RCM reaction of HVPA, in PBS or wild-type E. coli cell lysate. b, Performance comparison of small molecular copper complex (Cu–BTTAA) and Cu–HS–ArMs in mediating the fluorogenic CuAAC between AzHCou and 4-EA, in PBS or wild-type E. coli cell lysate. For a and b, the reaction schemes are provided on the bottom left, and the yields of the products were calculated based on the detected fluorescence, using the solution of the pure product at the concentration of presumed 100% yield. The product solutions were prepared according to the media and reagents used for each experiment to eliminate the impact of the changes in the product’s fluorescence intensity in different environments. For both experiments, standalone small molecular catalysts showed significantly hindered activity in wild-type E. coli lysate, while the assembled HS–ArMs showed much higher resistance despite slower kinetics. c, Evaluation of the GSH resistance of HG–Ru–Cl (small-molecule catalyst) and Ru–HS–ArM (aggregating macromolecular catalyst). HG–Ru–Cl showed completely no resistance toward GSH, while the resistance to GSH increased significantly for Ru–HS–ArM in an in vitro phase-separating condition. All data are presented as mean ± s.d. (n = 3 independent experiments). ***t ≤ 0.001, **t ≤ 0.01 and *t ≤ 0.05 in two-tailed Student’s t-test.
Extended Data Fig. 7 Supporting experimental results for intracellular ROMP mediated by Ru–HS–ArM in compartmentalized HS–E. coli.
a, Chemical scheme of the ROMP of NB-Ant monomer. b, Fluorescence emission spectra of NB-Ant (left) and poly(NB-Ant) (right) in different DMF/water mixtures. The results clearly indicated the aggregation-induced emission (AIE) property of the polymerized anthracene units. Concentration: 0.1 mg/mL, Ex = 370 nm. c, 1H NMR spectra overlay of the monomer (NB-Ant) and the product polymer (poly(NB-Ant)) involved in the intracellular ROMP in HS–E. coli. The characteristic backbone alkene peak (5.4–5.8 ppm) and the broadening of the peaks clearly indicated the successful formation of polynorbornene. d, Comparison of the molar mass distribution of the poly(NB-Ant) obtained from living ROMP (performed in dichloromethane) and in cellulo ROMP mediated by Ru–HS–ArM in artificial compartments.
Extended Data Fig. 8 Additional cartoon illustrations and data showing the site isolation and spatial control of the catalyst through the construction of LLPS-based catalytic compartments.
a, Confocal microscopic images showing the in-compartment catalysis by Cu–HS–ArMs in the LLPS regions and in-cytoplasm catalysis by Cu–BTTAA when LLPS compartments were created by the TTA-free cross-linker, Tris–Cl3. Cu–BTTAA could not enter the compartments due to its high hydrophilicity. Scale bar = 5 µm. b, Loss of capability in tandem reaction catalysis as the ruthenium and copper catalytic centers were spatially separated in the compartmentalized E. coli cells. The separation of Cu and Ru catalysts significantly hindered the two-step reaction (Fig. 4d), and the fluorescent final product could hardly be observed. H.–G. catalyst: Hoveyda–Grubbs catalyst. n = 3 independent experiments; representative results are shown. Scale bar = 5 µm.
Extended Data Fig. 9 Experimental data supporting the validity of the prodrug-based combination therapy, mediated by compartmentalized HS–E. coli whole-cell catalysts.
a, An illustration showing the setup of the experiment, in which Cu–HS–E. coli and Pd–HS–E. coli-mediated CuAAC and Proc-removal reactions in live mice. b, The images of live HS–E. coli containing fluorescent products in the mouse feces, collected during the experiment described in a. These results clearly demonstrated the activity of such whole-cell catalysts in mouse intestines. See the Whole-cell catalysis by compartmentalized, ArM-containing HS–E. coli in mouse intestine subsection in the Supplementary Information for detailed protocol. n = 5 biological replicates for each reaction; representative results are shown. Scale bar = 5 µm. c, Heatmap showing the cytotoxicity of doxorubicin (Dox, IC50 = 4 µM), farnesyl umbelliferone (FUmb, IC50 = 64 µM) and their combinations toward HCT116 cells. The experiment found Dox and FUmb showing the highest synergy at a 1:32 (wt/wt) ratio. At this ratio, the IC50 of Dox/FUmb combinations was measured to be 0.7 µM (Dox) + 23 µM (FUmb) through curve-fitting. d, Schematic illustration on the in vitro validation of Pd and Ru whole-cell catalyst’s capability of activating Pro-Dox and Pro-FUmb. Prodrugs were added to solution containing HS–E. coli catalysts, and after 24 h, the supernatant was collected for HPLC characterization. The prodrug activation reactions are shown on the right. e, HPLC analysis results indicated that the whole-cell catalysts successfully mediated the desired abiotic transformation in the experiment depicted in d, and the products found in the supernatant indicated that the activated drugs could diffuse out of the bacterial cells. Concentrations used during incubations: 25 µM for Pd, 100 µM for Ru, 10 µM for Pro-Dox, 100 µM for Pro-FUmb, whole-cell catalyst OD = 0.6.
Extended Data Fig. 10 Additional data from the synergistic cancer treatment study using HS–E. coli-based whole-cell abiotic catalysts.
a, Statistics on the number of tumors in intestine for all the mice upon death or killing in the mouse colorectal cancer model study. n = 6 biological replicates. b, Statistics on the number of tumors in intestine for mice that survived past day 8 in the mouse colorectal cancer model study. n = 1–5 biological replicates, based on the number of mice survived in each group. c, Mean spleen volumes of the mice survived past day 8 in the mouse colorectal cancer model study. n = 1–5 biological replicates, based on the number of mice survived in each group. d, Representative photographs of mouse organs in the 5 treatment groups. Large tumors are marked by white arrows (markings are not exhaustive, detailed statistics are provided in a). These results clearly demonstrated the effectiveness and reduced toxicity brought by the combination therapy, assisted by whole-cell catalysts that activate drugs at the disease site. **, *** indicate t ≤ 0.01, 0.001 in two-tailed Student’s t-test, respectively. All data are presented as mean ± s.d. where relevant.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–28, Tables 1–5 and Methods.
Supplementary Data 1 (download ZIP )
Supporting data for Supplementary Figs. 1b, 2b, 3b, 4c,d, 7a,b, 8c,e, 9b,d, 10b,d and 13.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 1d.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 3b.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 4e.
Source Data Fig. 5 (download XLSX )
Statistical source data for Figs. 5b–e.
Source Data Fig. 6 (download XLSX )
Statistical source data for Figs. 6b–d.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed gel image for Extended Data Figs. 3a,d,e.
Source Data Extended Data Fig. 4 (download XLSX )
Raw data for Extended Data Fig. 4a and statistical source data for Extended Data Fig. 4d.
Source Data Extended Data Fig. 5 (download XLSX )
Unprocessed gel image for Extended Data Figs. 5a,b,d and statistical source data for Extended Data Figs. 5b,d.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data for Extended Data Figs. 6a–c.
Source Data Extended Data Fig. 7 (download XLSX )
Raw data for Extended Data Fig. 7b.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data for Extended Data Fig. 10a–c.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, T., Chen, X., Fei, Y. et al. Artificial metalloenzyme assembly in cellular compartments for enhanced catalysis. Nat Chem Biol 21, 779–789 (2025). https://doi.org/10.1038/s41589-024-01819-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-024-01819-7
This article is cited by
-
De novo design and evolution of an artificial metathase for cytoplasmic olefin metathesis
Nature Catalysis (2025)