Abstract
Notch signaling regulates cell fate decisions and has context-dependent tumorigenic or tumor suppressor functions. Although there are several classes of Notch inhibitors, the mechanical force requirement for Notch activation has hindered attempts to generate soluble agonists. To address this problem, we engineered synthetic Notch agonists (SNAGs) by tethering affinity-matured Notch ligands to proteins that internalize their targets. This bispecific format enables SNAGs to ‘pull’ on mechanosensitive Notch receptors, triggering their activation in the presence of desired biomarkers. We successfully developed SNAGs targeting six independent surface markers, including the tumor antigens PDL1, CD19 and HER2 and the immunostimulatory receptor CD40. HER2-SNAGs and CD19-SNAGs increased the expression of T cell activation markers and Notch target genes in cocultures with tumor cells, highlighting their potential for immunotherapeutic applications. These insights have broad implications for the pharmacological activation of mechanoreceptors and will expand our ability to modulate Notch signaling in biotechnology.

Similar content being viewed by others
Main
The Notch pathway is a cell-to-cell communication system that regulates embryonic development, tissue homeostasis and immune cell differentiation. The signaling of mechanosensitive Notch receptors is tightly regulated1 and aberrant Notch activity causes various human diseases2. For example, loss-of-function mutations in Notch components are linked to the development of aortic valve disease (Notch1), Alagille syndrome (Notch2 and Jagged1 (JAG1)), CADASIL (Notch3) and spondylocostal dysostosis (Delta-like ligand 3 (DLL3))3,4,5,6. In cancer, Notch functions as a tumor suppressor or oncogene depending on the cell type and both loss-of-function and hyperactivating mutations influence tumorigenesis and disease progression7. Notch is also pleiotropic with respect to its guidance of cell fate decisions, in that Notch activation stimulates either proliferation or differentiation in different stem cell populations8. These diverse functions suggest that Notch agonists and antagonists may each be viable therapeutics in certain biomedical contexts9.
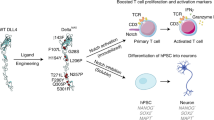
The role of Notch in T cell biology has led to the development of Notch-based strategies for enhancing cancer immunotherapy. Notch signaling is important for several natural stages of T cell maturation10 and ex vivo Notch activation is required for the differentiation of T cells from hematopoietic stem cells11. This latter function may potentially be used to generate allogeneic T cells for ‘off-the-shelf’ adoptive T cell or chimeric antigen receptor T cell therapies. More recently, Notch activation has been shown to enhance the antitumor function of fully mature, activated T cells12. Genetic overexpression of an activated form of Notch13 and culturing T cells in the presence of Notch1-specific antibodies14 or ligand-expressing cells15 were each associated with improved tumor clearance in various animal models of cancer. Detailed analysis of the T cells used in these studies revealed that these phenotypes were because of the Notch-stimulated induction of exhaustion-resistant or stem-like phenotypes.
At the molecular level, Notch receptors are massive (~290 kDa) transmembrane proteins that are activated by a distinctive, mechanical force-driven mechanism1,16,17,18. Notch signaling is initiated when a DLL or JAG ligand forms a trans-interaction with a Notch receptor on the surface of an adjacent cell1,19,20,21. Endocytosis of the ligand then generates a ‘pulling’ force that propagates to the negative regulatory region (NRR) of Notch16,22. This pulling destabilizes the NRR, which exposes internal cleavage sites for processing by the intramembrane proteases ADAM10 (S2 cleavage) and γ-secretase (S3 cleavage)23,24. Following these proteolytic events, the Notch intracellular domain (NICD) translocates to the nucleus to function as a transcriptional coactivator25.
Although Notch inhibitors are widely available, soluble agonists are challenging to engineer because they must somehow ‘pull’ on the Notch receptor despite lacking a method of force generation9,18. Multiple strategies have been developed to activate Notch receptors in vitro through mimicry of the physiological activation process. Notch signaling may be induced through coculture of Notch-expressing cells and ligand-expressing cells, by culturing Notch-expressing cells on plates coated with ligands or antibodies or through administration of ligand-coated microbeads14,26,27. Thus far, the only Notch agonist antibody, A13, functions by destabilizing the metastable NRR domain of Notch3 (refs. 28,29,30). Unfortunately, this NRR unfolding approach has been ineffective for receptor subtypes with stable NRRs (for example, Notch1, Notch2 and Notch4). Oligomerized ligands have also shown limited promise as agonists. For example, preclustering DLL1–Fc fusions with anti-Fc antibodies, JAG1-coated DNA origami structures and DLL4 proteins fused to designed trimeric scaffolds (C3-DLL4) stimulate low levels of Notch activation in vitro31,32,33.
Here we engineered bispecific proteins that stimulate activation of Notch signaling in desired cellular contexts. We successfully developed SNAGs that enhance the signaling of weakly activating JAG1 ligands and those that selectively activate Notch in the presence of the several immunologically relevant cell surface biomarkers. Furthermore, we show that SNAGs stimulate increased expression of Notch target genes and T cell activation markers in primary human T cells. The modularity and versatility of this SNAG platform provide a blueprint for the development of a diverse repertoire of Notch-based biologics.
Results
Soluble DLL4 ligand multimers do not activate Notch signaling
As an initial attempt to generate Notch agonists, we investigated whether soluble oligomers of an affinity-matured DLL4 ligand (DeltaMAX) activate Notch signaling34. DeltaMAX contains ten mutations that increase its affinity for human Notch receptors by 500-fold to 1,000-fold (KD = 24 to 54 nM), making it a more potent activator than DLL4 (KD > 10 µM) in coculture and plate-bound formats34. We hypothesized that this increased affinity, coupled with receptor crosslinking through multimerization, could introduce tension in the absence of an endocytic pulling force. To test this hypothesis, we incubated Notch1–Gal4 mCitrine reporter cells35 with soluble and immobilized DeltaMAX multimers (Fig. 1a–c). DeltaMAX dimers were generated through the C-terminal addition of a dimeric human IgG1 Fc domain (Fig. 1b) and tetramers were generated by premixing a 4:1 molar ratio of biotinylated DeltaMAX with streptavidin (SA) (Fig. 1c). We found that neither the monomers nor the multimers induced reporter activity. By contrast, the plated DeltaMAX ligands potently stimulated Notch1 activation (Fig. 1a–c). This indicates that the receptor crosslinking by DeltaMAX–Fc dimers and DeltaMAX–SA tetramers is insufficient for signaling activation.
a, Flow cytometry histogram overlay of Notch1 reporter cells stimulated by soluble or plated (nonspecifically adsorbed) DeltaMAX. The cartoon depicts the site-specifically biotinylated DeltaMAX (N–EGF5) construct. b, Histogram overlay of Notch1 reporter cells stimulated with soluble or plated DeltaMAX–Fc protein. c, Histogram overlay of Notch1 reporter cells stimulated with plated or soluble DeltaMAX–SA tetramers. d, Cartoon schematic depicting the ECDs of Notch1 and DLL4 interacting during canonical Notch activation. The NRR and ligand-binding domains (LBDs) of Notch1 (EGF domains 8–12) are shaded. e, Schematic of a generalized SNAG construct alongside a cartoon depicting SNAG-mediated Notch activation. f, Representation of the modular design of SNAGs. g–i, Top: schematic highlighting targets of SNAGs separated by canonical Notch ligands (g), tumor biomarkers (h) and B cell biomarker (i). Bottom: the corresponding endocytic mechanism.
Design of synthetic Notch agonists
To develop soluble Notch agonists, we engineered bispecific proteins that recapitulate the endocytosis-linked activation mechanism of DLL and JAG ligands (Fig. 1d). SNAGs were created by fusing DeltaMAX to the N terminus of biomarker-targeting antibody fragments through a flexible (GS)5 linker or by fusing DeltaMAX and antibody fragments to the N- and C- termini of a dimeric IgG1 Fc domain (Fig. 1e). These design concepts are intended to form a ‘molecular bridge’ between Notch-expressing cells and cells that express a given surface protein. Conceptually, SNAGs should then activate Notch if the enforced interactions induce endocytic or tensile force capable of unfolding the NRR. To this end, a modular SNAG platform (Fig. 1f) was developed to have exchangeable modules for biomarker-specific SNAGs against canonical Notch ligands (Fig. 1g), biomarkers upregulated in tumors (Fig. 1h) and an immunostimulatory biomarker (Fig. 1i).
SNAGs rescue the signaling of a signaling-deficient DLL4 mutant
To demonstrate proof of concept, we tested whether SNAGs could rescue the activity of a signaling-deficient DLL4 mutant. Loss-of-function DLL4 cells were generated by expressing a ‘headless’ DLL4 truncation where the Notch-binding C2 and DSL domains20,36 were replaced with a BC2 epitope tag (BC2-DLL4HL) (Fig. 2a and Supplementary Fig. 1a)37. BC2-SNAGs were then generated by fusing DeltaMAX to a BC2-specific nanobody (Fig. 2a,b and Supplementary Fig. 2a). We found that BC2-DLL4HL cells alone did not activate signaling in a Notch1–Gal4 mCitrine reporter assay, whereas the addition of 1–100 nM SNAGs stimulated a dose-dependent increase in reporter activity (Fig. 2c and Supplementary Fig. 3a–c). Monomeric BC2-SNAGs containing the (GS)5 linker (BC2-SNAG) stimulated a ~6-fold increase in Notch1 signaling, whereas dimeric BC2-SNAG–Fc fusion proteins (BC2-SNAGFc) were more effective and induced a ~10-fold increase (Fig. 2c). Importantly, administration of the monomeric or dimeric BC2-SNAGs alone did not substantially increase Notch1 reporter activity, indicating that a mixture of target-expressing and nonexpressing cells is required for SNAG-mediated activation (Fig. 2c).
a, Cartoon schematic depicting a SNAG binding to Notch1 and a loss-of-function DLL4 mutant. DLL4HL was generated by replacing the Notch-binding C2-DSL region with a BC2 peptide epitope recognized by the anti-BC2 nanobody. b, Cartoon schematic depicting the multivalent binding of BC2-SNAGFc to Notch1 and DLL4HL. c–e, Fluorescent reporter assays were used to evaluate SNAG-mediated activation of Notch1 in cocultures with HEK293T cells expressing DLL4HL (c), JAG1 (d) or JAG1-H268Q (e). Increasing concentrations (1 nM, 10 nM or 100 nM) of the indicated SNAGs were added to Notch1–Gal4 Citrine reporter cells alone or to a 1:1 mixture of Notch1 reporter cells and HEK293 cells expressing DLL4HL; fluorescence was measured by flow cytometry. A representative experiment from three biological replicates is shown. The MFI was normalized to the MFI of Notch1 reporter cells alone. Error bars represent the s.d. of n = 3 technical replicates, with the P value determined using a two-sided Student’s t-test shown above each comparison.
SNAGs bolster the activity of weakly signaling JAG1 ligands
DLL or JAG ligands preferentially signal through certain Notch receptor subtypes and JAG1 is a particularly weak activator of Notch1 (ref. 38). Therefore, we tested whether a JAG1-targeting SNAG (JAG1-SNAGFc) could potentiate JAG1–Notch1 signaling. In the JAG1-SNAGFc construct, DeltaMAX and a single-chain variable fragment (scFv) derived from the JAG1-targeting antibody B70 (ref. 39) were fused to the N and C termini of an IgG1 Fc domain as described above (Supplementary Fig. 2b). We then performed a signaling assay to measure the activation of Notch1 reporter cells by JAG1-overexpressing HEK293 cells (JAG1-293 cells) in the presence or absence of the SNAG. We found that addition of the JAG1-SNAGFc increased Notch1 reporter activity ~4-fold compared to JAG1-293 cells alone and JAG1-293 cells did not stimulate a significant increase in reporter activity (Fig. 2d and Supplementary Fig. 1b). We also evaluated the JAG1-SNAGFc with a JAG1-H268Q ‘nodder’ mutant that causes Alagille syndrome-like symptoms in mice by decreasing JAG1–Notch1 binding (Supplementary Fig. 1c). Addition of the JAG1-SNAGFc to cocultures of Notch1 and JAG1-H268Q cells increased JAG1 signaling up to sevenfold compared to Notch1 cells alone (Fig. 2e). These data indicate that SNAGs can function as ‘signaling enhancers’ by potentiating the activity of endogenous or mutated ligands.
SNAGs targeting tumor antigens activate Notch in mixed cell populations
We next tested whether SNAGs targeting the tumor antigens PDL1, CD19 or HER2 (Fig. 1h) can stimulate Notch activation. There is mounting evidence that Notch signaling enhances the function of activated T cells13,14,15 and SNAGs localized to the tumor microenvironment have the potential to stimulate localized activation of tumor-associated lymphocytes. For these SNAGs, the targeting arms were derived from antibody–drug conjugates (ADCs) that were preselected for their ability to induce target internalization. We hypothesized that SNAGs incorporating ADC antibodies could, thus, mimic the physiological endocytosis mechanism of DLL or JAG ligands.
We generated monomeric and dimeric PDL1-SNAGs by fusing DeltaMAX to an scFv derived from the ADC antibody atezolizumab40,41. In the monomeric PDL1-SNAG, DeltaMAX and the scFv were connected using a (GS)5 linker, whereas, in the dimeric PDL1-SNAG (PDL1-SNAGFc), DeltaMAX and the scFv were fused to the N and C termini of an IgG1 Fc domain (Fig. 1h and Supplementary Fig. 2a). Unexpectedly, addition of the monomeric PDL1-SNAG to a 1:1 mixture of Notch1 reporter cells and PDL1-expressing MDA-MB-231 cells did not activate Notch1 (Fig. 3a and Supplementary Fig. 1d). On the other hand, the dimeric PDL1-SNAGFc protein stimulated a ~7-fold increase in Notch1 signaling in the coculture, suggesting that multimerization or avidity enhancement may be required for SNAGs to effectively target biomarkers other than Notch ligands (Fig. 3a). Neither PDL1-SNAG nor PDL1-SNAGFc substantially increased Notch1 reporter activity in the absence of MDA-MB-231 cells. To test the importance of Notch-binding affinity in SNAG design, we generated a PDL1-SNAGFc incorporating wild-type DLL4 (Supplementary Fig. 2d), which binds to Notch1 with ~1,000-fold decreased affinity (KD = 24.7 µM) compared to DeltaMAX (KD = 24 nM). We found that the wild-type DLL4 SNAG was unable to activate Notch1 at all concentrations tested (Supplementary Fig. 4), indicating that the enhanced binding of the DeltaMAX variant is essential for SNAG function.
a, Increasing concentrations (1 nM, 10 nM or 100 nM) of DeltaMAX–Fc, PDL1-SNAG or PDL1-SNAGFc were added to Notch1–Gal4 mCitrine reporter cells alone or in a 1:1 coculture with PDL1-expressing MDA-MB-231 cells and fluorescence was measured by flow cytometry. b, MDA-MB-231 cells were incubated with PDL1-SNAGFc, soluble DeltaMAX–Fc or plated DeltaMAX–Fc and Notch1 activation was assessed by Western Blot using an antibody to the cleaved NICD. Data represent a single experiment (n = 1). c, Increasing concentrations (1 nM, 10 nM or 100 nM) of DeltaMAX–Fc or CD19-SNAGFc were added to Notch1 reporter cells or 1:1 mixtures of reporter cells and CD19-expressing 3T3 cells. d, Increasing concentrations (1 nM, 10 nM or 100 nM) of DeltaMAX–Fc or HER2-SNAGFc were added to Notch1 reporter cells or 1:1 mixtures of reporter cells and HER2-expressing SK-BR-3 cells. e, Increasing concentrations (1 nM, 10 nM or 100 nM) of trimeric CD40-SNAG or DeltaMAX–Fc were added to reporter cells alone or 1:2 mixtures of Notch1 reporter cells and CD40-expressing OCI-Ly3 cells. f), Notch1 reporter cells were stimulated with plated DeltaMAX–Fc, DLL4–Fc, EDTA, DLL4-expressing HEK293T cells and PDL1-SNAGFc in the presence of MDA-MB-231 cells and fluorescence was measured using flow cytometry. The dashed line denotes the activation level induced by cellular activation with HEK293T-DLL4 cells. Ligands were adsorbed at 100 nM concentrations, EDTA was added to a concentration of 1 mM and soluble PDL1-SNAGFc was added to a concentration of 100 nM. For a,c–f, a representative experiment from three biological replicates is shown. The MFI was normalized to the MFI of Notch1 reporter cells alone. Error bars represent the s.d. of n = 3 technical replicates, with the P value determined using a two-sided Student’s t-test shown above each comparison (P = 6.467 × 10−7 in comparison of DeltaMAX–Fc to CD40-SNAG at 10 nM).
SNAGs do not activate signaling on cells expressing both Notch1 and PDL1
Given the ubiquitous expression of Notch1 in mammalian cells, it is conceivable that SNAGs could activate signaling when Notch1 and the target protein are both present on the cell surface. To test this possibility, we cultured MDA-MB-231 cells in the presence of soluble DeltaMAX–Fc, PDL1-SNAGFc or immobilized DeltaMAX–Fc and monitored the levels activated Notch1 by western blot (Fig. 3b). We found that the plated DeltaMAX–Fc protein stimulated high levels of Notch1 activation, whereas PDL1-SNAGFc did not induce signaling over the background levels observed for soluble DeltaMAX–Fc alone (Fig. 3b). The inability of SNAGs to activate Notch1 in MDA-MB-231 cells suggests that the present design does not enable sufficient intercellular crosslinking in cultures of cells expressing both Notch1 and the biomarker.
Development of SNAGs targeting CD19, HER2 and CD40
To generate a SNAG targeting the B lymphocyte antigen CD19, we fused an scFv derived from the CD19-targeting ADC loncastuximab42 to the C terminus of DeltaMAX–Fc (CD19-SNAGFc) (Supplementary Fig. 2a). CD19-SNAG was then added to Notch1 reporter cells or to cocultures of Notch1 reporter cells and CD19-overexpressing 3T3 fibroblast cells (Supplementary Fig. 1e). We found that the CD19-SNAGFc protein stimulated up to a sixfold increase in reporter activity in the coculture compared to untreated Notch1 cells (Fig. 3c). To generate a SNAG targeting the breast cancer antigen HER2 (HER2-SNAGFc), we replaced the CD19-targeting arm with an scFv derived from the HER2-targeting ADC trastuzumab43 (Supplementary Fig. 2c). Addition of HER2-SNAGFc to a mixed culture of Notch1 reporter cells and HER2-expressing SK-BR-3 breast cancer cells induced a sixfold increase in reporter activity (Fig. 3d and Supplementary Fig. 1f) at the highest concentration tested (100 nM), which is similar to the level of activation we observed for the PDL1-SNAGFc and the CD19-SNAGFc constructs (Fig. 3a,c). In the absence of biomarker-expressing cells, neither CD19-SNAGFc nor HER2-SNAGFc stimulated a significant increase in signaling compared to DeltaMAX–Fc alone (Fig. 3c,d). Our collective development of PDL1-SNAGs, CD19-SNAGs and HER2-SNAGs demonstrates that SNAGs can facilitate signaling by engaging cell surface proteins beyond endogenous ligands.
We also generated a SNAG targeting CD40 (CD40-SNAG), an immunostimulatory receptor that undergoes endocytosis upon binding to CD40 ligand (CD40L)44,45. We hypothesized that a CD40-SNAG could be used to activate Notch in CD40− T cells in the presence of CD40+ B cells, as these cell types are known to colocalize in germinal centers, the tumor microenvironment and peripheral blood46. To generate a CD40-SNAG, we fused DeltaMAX and the extracellular domain (ECD) of CD40L to the N and C termini of a trimeric leucine zipper47 (Fig. 1i and Supplementary Fig. 2e), respectively. This trimeric scaffold was selected instead of an Fc domain for CD40-SNAG because CD40L is naturally a homotrimer. We found that the addition of CD40-SNAG robustly activated Notch in mixed cultures of Notch1 reporter cells and CD40-expressing OCI-Ly3 cells (Supplementary Fig. 1g) but only weakly in Notch1 reporter cells alone (Fig. 3e). The OCI-Ly3 cells used in this assay are nonadherent, indicating that SNAGs also facilitate Notch activation between adherent and suspension cells.
Comparison of SNAGs to conventional methods of Notch activation
To assess the relative effectiveness of SNAGs, we compared Notch activation levels of the soluble PDL1-SNAGFc and various conventional methods of Notch agonism. In this assay, Notch reporter cells were stimulated with plated DeltaMAX–Fc, plated DLL4–Fc, EDTA, DLL4-expressing HEK293T cells and PDL1-SNAGFc in the presence of PDL1-expressing cells (Fig. 3f). In the plate-bound format, 100 nM concentrations of each ligand were nonspecifically adsorbed to the surfaces and 100 nM concentrations of SNAG were added to the coculture. We found that, in the presence of PDL1-expressing cells, PDL1-SNAGFc induced higher levels of reporter activity than all methods of activation except for the plated DeltaMAX–Fc protein, which has been shown to exhibit superagonist activity34. Notably, PDL1-SNAGFc stimulated higher levels of signaling than DLL4-overexpressing cells, which typically serve as a benchmark for Notch activation. The DLL4–Fc protein did not detectably activate Notch1 in this format, which may be because of the lack of C-terminally anchored coupling required for optimal signaling with wild-type ligands34,48.
Endocytosis is required for SNAG-mediated Notch activation
Ligand endocytosis is important for Notch activation49 and this process is regulated by ubiquitination of DLL or JAG ICDs by the E3 ligase Mind bomb 1 (refs. 50,51,52). To determine whether endocytosis occurs with a SNAG targeting a surface protein other than a natural Notch ligand, we performed an immunofluorescence endocytosis assay using CD19-SNAGFc in CD19-expressing cells. CD19-SNAGFc coupled with a fluorescent secondary antibody (Alexa Fluor 647-labeled anti-Fc) bound strongly to the surface of the CD19-expressing cells when the mixture was incubated on ice (Fig. 4a and Supplementary Fig. 5a–c). Following CD19-SNAGFc binding, incubation at 37 °C permitted the resumption of cellular activity, including endocytosis. Visualizing the cells after a 15-min incubation at 37 °C showed that the majority of bound CD19-SNAGFc was internalized (Fig. 4b and Supplementary Fig. 5e–d). Performing the assay in the presence of the dynamin-dependent endocytosis inhibitor Dynasore blocked SNAG uptake, further supporting our observation that SNAGs are internalized (Fig. 4c).
a, Representative immunofluorescence images of fluorescently labeled CD19-SNAGs (magenta) used to stain the surface of CD19-expressing 3T3 cells that were kept on ice. CD19-SNAGs visualized by Alexa Fluor anti-Fc 647. To visualize the contours of the cells, the actin cytoskeleton was stained using phalloidin 488 (green). Nuclei were counterstained by Hoechst 33342 (blue). Scale, 20 mm. b,c, Representative immunofluorescence images of fluorescently labeled CD19-SNAGs used to stain the surface of CD19-expressing 3T3 cells in the absence (b) or presence (c) of Dynasore, followed by washing away unbound SNAGs and subjecting the cells to a 15-min incubation in a 37 °C incubator to resume cellular processes, including endocytosis. After 15 min, the cells were fixed and stained in parallel with the no endocytosis samples. Scale, 20 mm. A representative staining is shown for each experimental condition from one of two biological replicates. d–g, Flow cytometry histogram overlays depicting Notch1 reporter activity induced by soluble CD19-SNAGFc, BC2-SNAG or BC2-SNAGFc in the presence or absence of Dynasore. d, Notch1 reporter cells were cocultured with CD19-overexpressing 3T3 cells. In e,f, Notch1 reporter cells were cocultured with HEK293 cells expressing DLL4HL. g, Flow cytometry histogram overlay depicting Notch1 reporter activity induced by immobilized BC2-SNAG in the presence or absence of Dynasore. A representative histogram is shown for each experimental condition from one biological replicate for CD19-SNAGFc (soluble), two biological replicates for BC2-SNAG (soluble) or three biological replicates for both BC2-SNAGFc (soluble) and BC2-SNAG (plated).
To test whether endocytosis is necessary for SNAG signaling, we coadministered SNAGs with Dynasore. We found that Dynasore completely ablated the activity of CD19-SNAGFc in cocultures of Notch1-expressing and CD19-expressing cells, indicating that endocytosis is required for SNAG-mediated activation using CD19 as a biomarker (Fig. 4d). We further found that BC2-SNAGs targeting BC2-DLL4HL were unable to activate Notch1 in cocultures of Notch1 and BC2-DLL4HL cells in the presence of Dynasore, confirming that endocytosis is also required for SNAG-mediated rescue of DLL4 signaling (Fig. 4e,f.) Interestingly, we found that immobilized SNAGs were also unable to activate Notch1 in the presence of Dynasore, suggesting that endocytosis in the Notch receptor cell is essential for Notch activation by plated ligands (Fig. 4g). These studies demonstrate that Notch activation by plated ligands, SNAGs targeting a DLL4 loss-of-function mutant and SNAGs targeting tumor antigens each depend on endocytosis.
SNAGs stimulate the expression of T cell activation markers
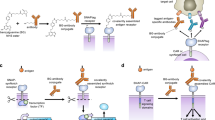
To investigate whether bispecific SNAGs influence the behavior of activated T cells, we established in vitro cocultures of primary human CD8⁺ T cells with tumor cells expressing the SNAG targets PDL1 or HER2 (Fig. 5a). The cocultures were then treated with PDL1-SNAGFc or HER2-SNAGFc and then activated with anti-CD28-coated or anti-CD3-coated beads. We found that the addition of PDL1-SNAGFc or HER2-SNAGFc but not DeltaMAX–Fc increased the proportion of granzyme B (GrB)-expressing CD8⁺ T cells, which is indicative of a cytotoxic phenotype. At the transcriptional level, SNAG-treated T cells showed elevated expression of GZMB and IFNG, key markers of effector function, and HES4, a canonical Notch target gene (Fig. 5b,c). In the PDL1-SNAGFc-treated cocultures, GZMB, IFNG and HES4 were upregulated 18-fold, 37-fold and 3-fold, respectively, relative to untreated controls. Similarly, treatment with HER2-SNAGFc resulted in 4-fold, 52-fold and 16-fold increases in GZMB, IFNG and HES4 expression, respectively. This increased expression could be additionally found at the protein level, as intracellular staining followed by flow cytometry revealed increased GrB and interferon-γ (IFNγ) expression in SNAG-treated T cells (Supplementary Fig. 6a,b).
PBMCs were used to isolate naïve CD8+ T cells that were treated with 100 nM DeltaMAX–Fc, PDL1-SNAGFc or HER2-SNAGFc or not treated (control) in the presence of PDL1+ MDA-MB-231 cells or HER2+ SK-BR-3 cells, followed by human CD3 and CD28 stimulation for 48 h before isolation. a, Schematic depicting a SNAG engaging the target biomarkers expressed on a tumor cell and Notch receptors of a CD8+ T cell. b,c, The first bar chart shows the percentage of CD8+ and GrB+ T cells and subsequent three bar charts show the relative mRNA levels for GZMB, IFNG and HES4 normalized to the control; each bar represents the fold change. Error bars represent the s.d. of n = 3 technical replicates, with the P value determined using a two-sided Student’s t-test shown above each comparison.
Discussion
The development of soluble agonists has been an enduring challenge in the Notch field9,53. The SNAG platform described here offers a solution to this problem and provides a framework for the development of a diverse array of Notch activating biologics. Such agents have a wide range of potential translational applications, particularly in cancer where Notch functions as a tumor suppressor7, T cell manufacturing11,27, T cell immunotherapy13,14,15, wound healing54 and other areas of regenerative medicine. These first-generation SNAGs were engineered using an Fc fusion format used in clinically viable protein drugs, which may also help to accelerate in vivo translation.
In their present form, SNAGs facilitate potent activation of Notch signaling in mixed populations of cells. It is possible that these designs may be further affinity-tuned to improve SNAG function. For example, the DeltaMAX arm (KD = 24 nM) and the JAG1-targeting, PDL1-targeting and HER2-targeting arms (KD = 0.9 nM, 1.8 nM and 5 nM, respectively) bind with similar affinities34,39,55,56. Lowering the affinity of the Notch-binding arm may, therefore, improve specificity and tissue distribution, as observed for bispecific inhibitory antibodies57 and T cell engagers58. On the other hand, we also demonstrated that wild-type DLL4 was ineffective when incorporated into SNAGs, suggesting that there is a lower limit for the allowable affinity range. Additionally, higher-order oligomers beyond the dimeric and trimeric SNAG scaffolds tested here may lead to increased signaling potency. Future studies will focus on optimizing affinity and multimerization to maximize signaling while maintaining favorable biochemical properties.
One surprising observation was that PDL1-SNAGs did not activate signaling on cells expressing both PDL1 and Notch1. We speculate that these SNAGs engage the two targets in cis on the surface of a single cell, as opposed to bridging PDL1 and Notch1 proteins between cells; moreover, cis interactions do not introduce sufficient tension to unfold the NRR. This may be attributed to the restricted diffusion of SNAGs in the two-dimensional environment of the membrane, which can promote preferential cis interactions by increasing the local concentration. Previous studies showed that cis inhibition of Notch signaling occurs when ligands and receptors are expressed on the same cell35,59 and it appears that SNAGs are similarly unable to activate Notch in this context. Regardless, the ability of SNAGs to mediate unidirectional signaling enables highly selective targeting, which could minimize the risks of potential toxicity from global Notch agonism.
There has been longstanding interest in using Notch to manufacture off-the-shelf CD4+ or CD8+ T cells for immunotherapy60. By contrast, the use of Notch agonists to bolster the effector function of mature or activated T cells emerged more recently and the advantages of this strategy have not been thoroughly vetted61. Here, we showed that the addition of PDL1-SNAGs or HER2-SNAGs can stimulate Notch signaling in CD8+ T cells cocultured with tumor cells expressing the target biomarkers. T cells stimulated in this fashion also had increased expression of the T cell activation markers GrB and IFNγ. These findings suggest that injected SNAGs may have the potential enhance the function of adoptively transferred T cells in vivo similarly to what has been achieved ex vivo with immobilized agonists; however, these predictions will require validation in animal models.
An important consideration in SNAG design, especially in the context of immunotherapy, is the ubiquitous expression of Notch receptors in nearly all cell types. In addition to effector T cells, regulatory T cells, myeloid-derived suppressor cells, macrophages and other immune cells are all typically present in the tumor microenvironment. Activation of Notch may not be desirable in all of these populations and has the potential to induce a range of immunostimulatory and immunosuppressive phenotypes. The ubiquitous expression of Notch proteins may also hinder the tissue distribution of SNAGs or result in Notch inhibition in the absence of a relevant biomarker. Further modifications may, thus, be necessary to maximize on-target Notch activation with SNAGs or alternative agonists scaffolds.
While SNAGs are effective in mixed cell populations, the development of ‘unconditional’ agonists (that is, those that do not rely on a secondary target) remains a challenging problem. The NRR of Notch3 appears to be uniquely susceptible to antibody-mediated destabilization29,30, whereas the engineering of agonists targeting other Notch receptors with more stable NRRs may require alternative solutions. In vitro activation of Notch1 with ligands fused to DNA origami structures, Fc-clustered DLL1 proteins and trimerization scaffolds suggests that induced receptor clustering may be an effective strategy31,32,33. The recent development of the trimeric C3-DLL4 agonists is the most promising among these approaches, as it is capable of stimulating Notch in T cell bioreactors and ameloblasts derived from induced pluripotent stem cells33,62. However, in vitro use of this method requires multiple rounds of doxycycline induction of Notch expression over a prolonged 96-h incubation period to activate Notch. Despite these limitations, the development of SNAGs and the related technologies above represent a key first step toward the widespread implementation of Notch agonists for basic and translational research.
Methods
Protein expression and purification
All SNAG sequences were cloned into a pAcGP67A vector for insect cell production containing an N-terminal gp67 signal peptide and C-terminal 8×His tag. Monomeric SNAGs were generated by fusing a truncated version of the DeltaMAX protein spanning from the N terminus to EGF5 (N–EGF5) fused to a biomarker-targeting scFv or nanobody using a flexible (GS)5 linker. Dimeric SNAGFc constructs were generated by fusing DeltaMAX (N–EGF5) and the biomarker-targeting module to the N and C termini of a human IgG1 Fc domain, respectively. All SNAGFc constructs contained short GSG linkers between the Fc sequence and DeltaMAX or the targeting module. Published sequences of atezolizumab, trastuzumab and loncastuximab63 were converted into scFv format before being incorporated into SNAGs and the sequence of the BC2-specific nanobody37 was obtained from the Protein Data Bank (PDB 5VIN). Each scFv was generated by fusing the C terminus of the variable heavy domain to the N terminus of the variable light domain with a (GGGGS)3 linker. Biotinylated DeltaMAX (N–EGF5) protein was generated through enzymatic modification of a C-terminal biotin acceptor peptide (BirA tag) as previously described34. The DLL4HL mutant was generated by replacing the C2 and DSL domains of human DLL4 with the BC2 peptide sequence, which was connected to the N terminus of EGF1 by a short GSG linker. The DLL4HL construct was cloned into a pLenti-IRES-Puro vector for mammalian expression.
All SNAG constructs in this study were expressed for by infecting Trichoplusia ni insect cell cultures (Expression Systems) at a density of 2 × 106 cells per ml with recombinant Baculovirus. Culture supernatants were isolated after 48 h and proteins were purified by nickel and size-exclusion chromatography. Biotinylated proteins were site-specifically modified using BirA ligase and excess biotin was removed by purifying the proteins on a size-exclusion column. Protein purity was assessed by SDS–PAGE using TGX 12% precast gels (Bio-Rad). All proteins were flash-frozen in liquid nitrogen and stored at −80 °C following purification.
Cell culture and generation of cell lines
Mammalian cells were cultured at 37 °C with a humidified atmosphere of 5% CO2, washed with Dulbecco’s PBS (DPBS; Corning) and detached with trypsin–EDTA 0.25% (Gibco) for subculturing or cell-based assays. Notch reporter CHO-K1 N1-Gal4 cells were a gift from M. Elowitz (California Institute of Technology)35. Briefly, transfections of HEK293T cells were carried out with packaging vectors VSV-G and d8.9 in the presence of polyethyleneimine at a ratio of 4:1 (DNA to polyethyleneimine). HER2+ SK-BR-3 cells, human CD19-overexpressing 3T3 cells, PDL1+ MDA-MB-231 and CD40+ OCI-Ly3 cells were gifts from B. Czerniecki, F. Locke, E. Lau and J. Cleveland, respectively (Moffit Cancer Center). HEK293T cells, SK-BR-3 cells, 3T3 mouse fibroblasts and MDA-MB-231 cells were cultured in high-glucose DMEM (Cytiva) supplemented with 10% FBS (peak serum) and 2% penicillin–streptomycin (Gibco). Puromycin (5 µg ml−1) was added to HEK293T cell cultures to maintain homogeneous populations of receptor-expressing cells. CHO-K1 N1-Gal4 cells were cultured in α-MEM (Cytiva) supplemented with 10% FBS (peak serum), 2% penicillin–streptomycin (Gibco), 400 µg ml−1 zeocin (Alfa Aesar) and 600 µg ml−1 geneticin (Gibco). Expression of receptors on the cell surface was confirmed using flow cytometry (BD Accuri C6 plus), by staining the cell lines with anti-hDLL4 PE (Biolegend; 1:100), anti-hJAG1 APC (Biolegend; 1:100), anti-hPDL1 FITC (Biolgend; 1:100) and anti-hHER2 (Cell Signaling Technologies; 1:100) followed by anti-IgG Alexa Fluor 488 (Biolegend; 1:200), anti-hCD19 FITC (Biolegend; 1:100) and anti-hCD40 PE (Biolegend; 1:100) in DMEM supplemented with 10% FBS for 1 h at 4 °C.
Notch stimulation with DeltaMAX multimers
On day 1, biotinylated DeltaMAX, DeltaMAX tetramers formed with SA or DeltaMAX–Fc was reconstituted in DPBS and adsorbed to tissue culture 96-well plates (Costar) for 1 h at 37 °C. The wells were then washed three times with 200 μl of DPBS to remove unbound proteins. Next, CHO-K1 N1-Gal4 cells were detached with trypsin–EDTA 0.25% (Gibco) and manually counted. Appropriate dilutions were prepared in α-MEM to ensure 30,000 CHO-K1 N1-Gal4 cells per well in a volume of 50 µl. Cells were transferred to the ligand-coated plates and cultured for 24 h at 37 °C in 5% CO2. On day 2, CHO-K1 N1-Gal4 cells were washed with 200 µl of DPBS, detached with 30 µl of trypsin–EDTA 0.25% and quenched with 170 µl of α-MEM. Finally, cells were resuspended and the H2B–mCitrine signal was measured by flow cytometry (BD Accuri C6 plus). CHO-K1 N1-Gal4 cells alone were used as the control. The measurements represent the mean fluorescence intensity (MFI) as the fold change of Notch activation ± s.d. of three technical replicates. Notch activation was normalized to wells containing CHO-K1 N1-Gal4 cells alone.
Notch activation with SNAGs in coculture of cells expressing the target biomarker
On day 1, cells expressing the target receptor of the SNAG (signal-sending cells) were detached with trypsin–EDTA and counted manually; dilutions were prepared such that 50 µl of DMEM containing 15,000 signal-sending cells were added to the wells of a tissue culture 96-well plate. The next day, CHO-K1 N1-Gal4 reporter cells (signal-receiving cells) were detached with trypsin–EDTA and 50 µl of α-MEM containing 30,000 cells was added to the tissue culture 96-well plate containing the signal-sending cells after combining with the indicated DeltaMAX or SNAG protein. For the CD40-SNAG experiments, the difference was that an equal number of signal-sending and signal-receiving cells were added same day. Wells without signal-sending cells were used to determine background activation of Notch by DeltaMAX and SNAGs. When testing inhibition of endocytosis, 80 µM of dynamin inhibitor I (Dynasore, Sigma) was added to the mixture of Notch reporter cells with protein and added to the tissue culture 96-well plate containing the signal-sending cells. Notch activation was measured as previously described.
Comparison of SNAGs to conventional methods of Notch activation
On day 1, cells expressing the canonical Notch ligand, DLL4 and PDL1+ MDA-MB-231 cells expressing the target receptor of the SNAG were detached with trypsin–EDTA and counted manually; dilutions were prepared such that 50 µl of DMEM containing 15,000 signal-sending cells was added to wells of a tissue culture 96-well plate. The next day, DeltaMAX-Fc and DLL4–Fc in DPBS were adsorbed to tissue culture 96-well plates (Costar) for 1 h at 37 °C. The wells were then washed three times with 200 μl of DPBS to remove unbound proteins. Next, CHO-K1 N1-Gal4 cells were detached with trypsin–EDTA 0.25% (Gibco) and manually counted. Appropriate dilutions were prepared in α-MEM to ensure 30,000 CHO-K1 N1-Gal4 cells per well in a volume of 50 µl. Cells were transferred to wells with adsorbed ligands and signal-sending cells only or after combining with DeltaMAX–Fc and EDTA (1.0 mM final concentration) or in coculture with PDL1-SNAGFc with or without DAPT (3 µM final concentration) and cultured for 24 h at 37 °C in 5% CO2. On day 2, CHO-K1 N1-Gal4 cells were washed with 200 µl of DPBS, detached with 30 µl of trypsin–EDTA 0.25% and quenched with 170 µl of α-MEM. Finally, cells were resuspended and the H2B–mCitrine signal was measured by flow cytometry (BD Accuri C6 plus). CHO-K1 N1-Gal4 cells alone were used as the control. The measurements represent the MFI as the fold change of Notch activation ± s.d. of three technical replicates. Notch activation was normalized to wells containing CHO-K1 N1-Gal4 cells alone.
Western blot detection of Notch1 activation by the PDL1-SNAGFc in MDA-MB-231 cells
DeltaMAX (100 nM protein in 600 µl of DPBS) was nonspecifically adsorbed to a single well of a 12-well plate for 1 h at 37 °C as a positive control for Notch1 activation. The positive control well and three additional wells were then seeded with 200 × 103 MDA-MB-231 cells. The plate was centrifuged at 400g for 4 min to ensure that cells were retained at the bottom of each well; then, the media of all wells were discarded. In the first uncoated well, 600 µl of DMEM was added as a negative control. The second well was filled with 600 µl of medium containing 100 nM of DeltaMAX–Fc to monitor Notch1 activation by soluble ligand. The third was filled with 600 µl of medium containing 100 nM PDL1-SNAGFc. The following day, the media were aspirated from all four wells and the samples were resuspended in 60 µl of Laemmli sample buffer with 5% β-mercaptoethanol to lyse cells, followed by boiling at 100 °C for 4 min. Lastly, the samples were analyzed by western blotting using equal protein amounts of cell lysates separated by SDS–PAGE (12% Mini-PROTEAN TGX precast protein gels, Bio-Rad) and transferred to PVDF membranes using an iBlot2 gel transfer device (Thermo Fisher Scientific). The membranes were blocked in 3% BSA + 0.1% TBS–Tween. Primary antibodies were anti-Notch1 (D1E11 rabbit monoclonal antibody, Cell Signaling Technology; 1:1,000), anti-cleaved Notch1 (Val1744 rabbit monoclonal antibody, Cell Signaling Technology; 1:1,000) and anti-β-actin (rabbit polyclonal antibody, Cell Signaling Technology; 1:1,000). Secondary antibody anti-rabbit IgG conjugated to horseradish peroxidase (goat polyclonal antibody, Vector Laboratories; 1:8,000) was used for the detection of proteins using SuperSignal West Pico PLUS chemiluminescent substrate (Thermo Fisher Scientific). Images were acquired using a Chemidoc Imaging System and analyzed with Image-Lab version 6 software (Bio-Rad).
Immunofluorescence cell staining
For endocytosis assays, cells were grown on glass-like polymer bottoms in 24-well black-frame plates (Cellvis). For visualization of CD19-SNAGFc protein binding, 500 nM protein was preincubated with anti-Fc 647 (Alexa Fluor) at 1:200 dilution for 1 h on rotation at 4 °C. The CD19-SNAGFc–647 solution was added to cells on ice that were further kept at 4 °C for 1 h. For endocytosis, the incubation was followed by washing away nonbound CD19-SNAGFc–647 with PBS and 37 °C DMEM was added to the cells followed by a 15-min incubation in a 37 °C incubator. After incubation of CD19-SNAGFc–647 with or without endocytosis, the cells were fixed in 3% paraformaldehyde and permeabilized with 0.15% Triton X-100 in PBS for 10 min at room temperature. Nonspecific binding was blocked by incubation in 3% BSA in PBS with 0.05% Triton X-100 and 0.1 M glycine for 60 min at room temperature. Cells were further stained for filamentous actin with Alexa 488 conjugated to phalloidin (Invitrogen) for 45 min to visualize contours of the individual cells. Hoechst 33342 (Invitrogen) was used to counterstain nuclei. Images were acquired using a Keyence BZ-X710 microscope using a Nikon Plan Apo ×20 objective. The far-red channel (magenta) was processed with the dehaze function in the BZ-X710LE analyzer software. A minimum of 100 cells were imaged for each condition.
Treatment of CD8+ T cells with HER2-SNAGFc and PDL1-SNAGFc
Human CD8⁺ T cells were purified from peripheral blood mononuclear cells (PBMCs) using the MagniSort human CD8+ T cell enrichment kit (Invitrogen, 8804-6812-74) according to the manufacturer’s protocol. Naive CD8⁺ T cells were then cocultured at a 1:1 ratio with PDL1⁺ MDA-MB-231 cells or HER2⁺ SK-BR-3 cells, with or without DeltaMAX–Fc or SNAGFc. To enhance cell–cell contact, cultures were subjected to a short centrifugation at 300g for 1 min. T cells were activated by adding human CD3 and CD28 Dynabeads (Gibco, 11161D). After 48 h of stimulation, cells were treated with BD GolgiStop protein transport inhibitor (BD Biosciences, 554724) for 4 h. Fixation and permeabilization were performed using the BD Cytofix/Cytoperm fixation and permeabilization kit (BD Biosciences, 554714) according to the manufacturer’s protocol. Cells were then stained with human anti-IFNγ (APC), anti-GrB (FITC) and anti-CD8 (PE) antibodies and DAPI followed by flow cytometry analysis. Data were quantified by FlowJo 10 software. To quantify relative mRNA levels, the total RNA was extracted using TRIzol reagent (Invitrogen, 15596026). Complementary DNA (cDNA) synthesis was performed using 500 ng of purified RNA with the Verso cDNA synthesis kit (Thermo Fisher Scientific, AB1453B). qPCR was conducted using primers specific for IFNγ, GrB, HES4 and B2M. Gene expression levels were normalized to B2M as an internal control and fold changes were calculated relative to the control condition.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated in this study are available within the article; additional raw data are available from the corresponding author upon request. Source data are provided with this paper.
References
Sprinzak, D. & Blacklow, S. C. Biophysics of Notch signaling. Annu. Rev. Biophys. 50, 157–189 (2021).
Zhou, B. et al. Notch signaling pathway: architecture, disease, and therapeutics. Signal. Transduct. Target. Ther. 7, 1–33 (2022).
Garg, V. et al. Mutations in NOTCH1 cause aortic valve disease. Nature 437, 270–274 (2005).
Li, L. et al. Alagille syndrome is caused by mutations in human Jagged1, which encodes a ligand for Notch1. Nat. Genet. 16, 243–251 (1997).
Joutel, A. et al. Notch3 mutations in CADASIL, a hereditary adult-onset condition causing stroke and dementia. Nature 383, 707–710 (1996).
Bulman, M. P. et al. Mutations in the human Delta homologue, DLL3, cause axial skeletal defects in spondylocostal dysostosis. Nat. Genet. 24, 438–441 (2000).
Radtke, F. & Raj, K. The role of Notch in tumorigenesis: oncogene or tumour suppressor? Nat. Rev. Cancer 3, 756–767 (2003).
Hori, K., Sen, A. & Artavanis-Tsakonas, S. Notch signaling at a glance. J. Cell Sci. 126, 2135–2140 (2013).
Andersson, E. R. & Lendahl, U. Therapeutic modulation of Notch signalling—are we there yet? Nat. Rev. Drug Discov. 13, 357–378 (2014).
Brandstadter, J. D. & Maillard, I. Notch signalling in T cell homeostasis and differentiation. Open Biol. 9, 190187 (2019).
Schmitt, T. M. & Zúñiga-Pflücker, J. C. Induction of T cell development from hematopoietic progenitor cells by Delta-like-1 in vitro. Immunity 17, 749–756 (2002).
Kelliher, M. A. & Roderick, J. E. NOTCH signaling in T-cell-mediated anti-tumor immunity and T-cell-based immunotherapies. Front. Immunol. 9, 1718 (2018).
Sierra, R. A. et al. Rescue of Notch-1 signaling in antigen-specific CD8+ T cells overcomes tumor-induced T-cell suppression and enhances immunotherapy in cancer. Cancer Immunol. Res 2, 800–811 (2014).
Wilkens, A. B. et al. Notch1 signaling during CD4+ T-cell activation alters transcription factor networks and enhances antigen responsiveness. Blood 140, 2261–2275 (2022).
Kondo, T. et al. Notch-mediated conversion of activated T cells into stem cell memory-like T cells for adoptive immunotherapy. Nat. Commun. 8, 15338 (2017).
Meloty-Kapella, L., Shergill, B., Kuon, J., Botvinick, E. & Weinmaster, G. Notch ligand endocytosis generates mechanical pulling force dependent on dynamin, epsins, and actin. Dev. Cell 22, 1299–1312 (2012).
Wang, X. & Ha, T. Defining single molecular forces required to activate Integrin and Notch signaling. Science 340, 991–994 (2013).
Gordon, W. R. et al. Mechanical allostery: evidence for a force requirement in the proteolytic activation of Notch. Dev. Cell 33, 729–736 (2015).
Rebay, I. et al. Specific EGF repeats of Notch mediate interactions with Delta and Serrate: implications for Notch as a multifunctional receptor. Cell 67, 687–699 (1991).
Luca, V. C. et al.Structural basis for Notch1 engagement of Delta-like 4. Science 347, 847–853 (2015).
Luca, V. C. et al. Notch–Jagged complex structure implicates a catch bond in tuning ligand sensitivity. Science 355, 1320–1324 (2017).
Gordon, W. R. et al. Structural basis for autoinhibition of Notch. Nat. Struct. Mol. Biol. 14, 295–300 (2007).
De Strooper, B. et al. A presenilin-1-dependent γ-secretase-like protease mediates release of Notch intracellular domain. Nature 398, 518–522 (1999).
Brou, C. et al. A novel proteolytic cleavage involved in Notch signaling: the role of the disintegrin-metalloprotease TACE. Mol. Cell 5, 207–216 (2000).
Schroeter, E. H., Kisslinger, J. A. & Kopan, R. Notch-1 signalling requires ligand-induced proteolytic release of intracellular domain. Nature 393, 382–386 (1998).
Varnum-Finney, B. et al. Immobilization of Notch ligand, Delta-1, is required for induction of notch signaling. J. Cell Sci. 113, 4313–4318 (2000).
Trotman-Grant, A. C. et al. DL4-μbeads induce T cell lineage differentiation from stem cells in a stromal cell-free system. Nat. Commun. 12, 5023 (2021).
Li, K. et al. Modulation of Notch signaling by antibodies specific for the extracellular negative regulatory region of Notch3. J. Biol. Chem. 283, 8046–8054 (2008).
Tiyanont, K., Wales, T. E., Siebel, C. W., Engen, J. R. & Blacklow, S. C. Insights into Notch3 activation and inhibition mediated by antibodies directed against its negative regulatory region. J. Mol. Biol. 425, 3192–3204 (2013).
Xu, X. et al. Insights into autoregulation of Notch3 from structural and functional studies of its negative regulatory region. Structure 23, 1227–1235 (2015).
Ilagan, M. X. G., Lim, S., Fulbright, M., Piwnica-Worms, D. & Kopan, R. Real-time imaging of Notch activation with a luciferase complementation-based reporter. Sci. Signal. 4, rs7(2011).
Smyrlaki, I. et al. Soluble and multivalent Jag1 DNA origami nanopatterns activate Notch without pulling force. Nat. Commun. 15, 465 (2024).
Mout, R. et al. Design of soluble Notch agonists that drive T cell development and boost immunity. Cell 188, 1–15 (2025).
Gonzalez-Perez, D. et al. Affinity-matured DLL4 ligands as broad-spectrum modulators of Notch signaling. Nat. Chem. Biol. 19, 9–17 (2023).
Sprinzak, D. et al. Cis-interactions between Notch and Delta generate mutually exclusive signalling states. Nature 465, 86–90 (2010).
Cordle, J. et al. A conserved face of the Jagged/Serrate DSL domain is involved in Notch trans-activation and cis-inhibition. Nat. Struct. Mol. Biol. 15, 849–857 (2008).
Braun, M. B. et al. Peptides in headlock—a novel high-affinity and versatile peptide-binding nanobody for proteomics and microscopy. Sci. Rep. 6, 19211 (2016).
Andersson, E. R. et al. Mouse model of Alagille syndrome and mechanisms of Jagged1 missense mutations. Gastroenterology 154, 1080–1095 (2018).
Lafkas, D. et al. Therapeutic antibodies reveal Notch control of transdifferentiation in the adult lung. Nature 528, 127–131 (2015).
Powles, T. et al. MPDL3280A (anti-PD-L1) treatment leads to clinical activity in metastatic bladder cancer. Nature 515, 558–562 (2014).
Xiao, D. et al. Development of bifunctional anti-PD-L1 antibody MMAE conjugate with cytotoxicity and immunostimulation. Bioorg. Chem. 116, 105366 (2021).
Zammarchi, F. et al. ADCT-402, a PBD dimer-containing antibody drug conjugate targeting CD19-expressing malignancies. Blood 131, 1094–1105 (2018).
Lewis Phillips, G. D. et al. Targeting HER2-positive breast cancer with trastuzumab–DM1, an antibody–cytotoxic drug conjugate. Cancer Res. 68, 9280–9290 (2008).
Yellin, M. J. et al. CD40 molecules induce down-modulation and endocytosis of T cell surface T cell–B cell activating molecule/CD40-L. Potential role in regulating helper effector function. J. Immunol. 152, 598–608 (1994).
Hernandez, M. G. H., Shen, L. & Rock, K. L. CD40–CD40 ligand interaction between dendritic cells and CD8+ T cells is needed to stimulate maximal T cell responses in the absence of CD4+ T cell help. J. Immunol. 178, 2844–2852 (2007).
Mitchison, N. A. T-cell–B-cell cooperation. Nat. Rev. Immunol. 4, 308–312 (2004).
Naito, M. et al. CD40L-Tri, a novel formulation of recombinant human CD40L that effectively activates B cells. Cancer Immunol. Immunother. 62, 347–357 (2013).
Andrawes, M. B. et al. Intrinsic selectivity of Notch 1 for Delta-like 4 over Delta-like 1. J. Biol. Chem. 288, 25477–25489 (2013).
Parks, A. L., Klueg, K. M., Stout, J. R. & Muskavitch, M. A. Ligand endocytosis drives receptor dissociation and activation in the Notch pathway. Development 127, 1373–1385 (2000).
Koo, B.-K. et al. An obligatory role of Mind bomb-1 in notch signaling of mammalian development. PLoS ONE 2, e1221 (2007).
McMillan, B. J. et al. A tail of two sites: a bipartite mechanism for recognition of Notch ligands by Mind bomb E3 ligases. Mol. Cell 57, 912–924 (2015).
Cao, R. et al. Structural requirements for activity of Mind bomb1 in Notch signaling. Structure 32, 1667–1676 (2024).
Medina, E., Perez, D. H., Antfolk, D. & Luca, V. C.New tricks for an old pathway: emerging Notch-based biotechnologies and therapeutics. Trends Pharmacol. Sci. 44, 934–948 (2023).
Bonnici, L., Suleiman, S., Schembri-Wismayer, P. & Cassar, A. Targeting signalling pathways in chronic wound healing. Int. J. Mol. Sci. 25, 50 (2024).
Tan, S. et al. Distinct PD-L1 binding characteristics of therapeutic monoclonal antibody durvalumab. Protein Cell 9, 135–139 (2018).
Troise, F., Cafaro, V., Giancola, C., D’Alessio, G. & De Lorenzo, C. Differential binding of human immunoagents and Herceptin to the ErbB2 receptor. FEBS J. 275, 4967–4979 (2008).
Mazor, Y. et al. Enhanced tumor-targeting selectivity by modulating bispecific antibody binding affinity and format valence. Sci. Rep. 7, 40098 (2017).
Haber, L. et al. Generation of T-cell-redirecting bispecific antibodies with differentiated profiles of cytokine release and biodistribution by CD3 affinity tuning. Sci. Rep. 11, 14397 (2021).
del Álamo, D., Rouault, H. & Schweisguth, F. Mechanism and significance of cis-inhibition in Notch signalling. Curr. Biol. 21, R40–R47 (2011).
Boyd, N. et al. ‘Off-the-shelf’ immunotherapy: manufacture of CD8+ T cells derived from hematopoietic stem cells. Cells 10, 2631 (2021).
Mensurado, S., Blanco-Domínguez, R. & Silva-Santos, B. The emerging roles of γδ T cells in cancer immunotherapy. Nat. Rev. Clin. Oncol. 20, 178–191 (2023).
Patni, A. P. et al. Designed soluble Notch agonist drives human ameloblast maturation for tooth regeneration. Preprint at bioRxiv https://doi.org/10.1101/2025.04.03.646929 (2025).
Abanades, B. et al. The Patent and Literature Antibody Database (PLAbDab): an evolving reference set of functionally diverse, literature-annotated antibody sequences and structures. Nucleic Acids Res. 52, D545–D551 (2024).
Acknowledgements
This project was supported by the National Institutes of Health (NIH) R35GM133482 (V.C.L.), R01CA233512 (P.C.R), R01CA262121 (P.C.R.), R01CA273034 (P.C.R), P01CA250984 project number 4 (P.C.R) and the Sigrid Juselius Foundation (D.A.). V.C.L. is a Rita Allen Scholar. Shared resources were provided by the Moffitt Cancer Center Support Grant (NIH P30CA076292).
Author information
Authors and Affiliations
Contributions
V.C.L. and D.H.P. wrote the paper. V.C.L., D.H.P, D.A., P.C.R. and S.C. designed the experiments. D.H.P. cloned the SNAG constructs, purified the proteins and performed the signaling assays. E.M. and D.G.P. generated the DeltaMAX constructs. D.A. performed the western blot analysis and immunofluorescence endocytosis assays. S.C. performed the T cell coculture assays. X.E.B. and D.A.D. assisted with the T cell assays. A.A.R. generated the CD40-SNAG construct. V.C.L. supervised the project and edited the paper.
Corresponding author
Ethics declarations
Competing interests
V.C.L. is a consultant on unrelated projects for Cellestia Biotech, Remunix and Curie.Bio. V.C.L. and D.H.P. have filed provisional patents (63/548,615 and 63/663,744) based on the described technology. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–6.
Supplementary Data 1 (download XLSX )
Supporting data for Supplementary Fig. 4.
Supplementary Data 2 (download XLSX )
Supporting data for Supplementary Fig. 6.
Source data
Source Data Fig. 1 (download XLSX )
Numerical source data.
Source Data Fig. 2 (download XLSX )
Numerical source data.
Source Data Fig. 3 (download XLSX )
Numerical source data and unprocessed western blots.
Source Data Fig. 4 (download XLSX )
Numerical source data.
Source Data Fig. 5 (download XLSX )
Numerical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Perez, D.H., Antfolk, D., Chang, S. et al. Engineering synthetic agonists for targeted activation of Notch signaling. Nat Chem Biol 22, 392–401 (2026). https://doi.org/10.1038/s41589-025-02030-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41589-025-02030-y