Abstract
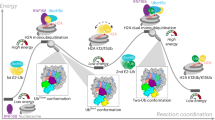
The activation of H2B K120 monoubiquitylation (H2BK120ub) by H2B S112 GlcNAcylation (H2BS112GlcNAc) has an important role in regulating transcriptional activation, yet its mechanism remains unclear. Here we chemically synthesized H2BS112GlcNAc-modified nucleosomes and quantitatively evaluated how H2BS112GlcNAc stimulates ubiquitylation by RNF20/RNF40–RAD6A E3–E2 enzymes. Cryo-electron microscopy determination of a chemically trapped RNF20/RNF40–RAD6A–Ub–H2BS112GlcNAc nucleosome complex revealed that the H2BS112GlcNAc moiety interacts with the E2 enzyme RAD6A but not the E3 ligase RNF20/RNF40. Mutagenesis and kinetics analyses demonstrated that H2BS112GlcNAc allosterically stimulates ubiquitin transfer from the RAD6A~Ub thioester to H2B K120 by enhancing the nucleophilicity of H2B K120. Structure‒activity relationship analysis further identified the essential roles of the C2 N-acetyl group and the β-configuration of C1 on the H2BS112GlcNAc moiety. These findings provide the structural evidence of histone posttranslational modification crosstalk involving O-GlcNAcylation and reveal how O-GlcNAcylation can allosterically stimulate enzyme activity through substrate modification.

This is a preview of subscription content, access via your institution
Access options





Similar content being viewed by others
Data availability
The cryo-EM maps and atomic model of RNF20/RNF40–RAD6A in complex with H2BS112GlcNAc nucleosome were deposited to the EM Data Bank and Protein Data Bank under accession codes EMD-62509 and PDB 9KQO, respectively. The three different conformational maps were deposited as additional maps to the EM Data Bank under accession code EMD-62509. Source data are provided with this paper.
References
Yang, X. Y. & Qian, K. V. Protein O-GlcNAcylation: emerging mechanisms and functions. Nat. Rev. Mol. Cell Biol. 18, 452–465 (2017).
Ma, J. F., Hou, C. Y. & Wu, C. Demystifying the O-GlcNAc code: a systems view. Chem. Rev. 122, 15822–15864 (2022).
Wu, C. et al. OGT and OGA: sweet guardians of the genome. J. Biol. Chem. 300, 107141 (2024).
Bond, M. R. & Hanover, J. A. O-GlcNAc cycling: a link between metabolism and chronic disease. Annu. Rev. Nutr. 33, 205–229 (2013).
Erickson, J. R. et al. Diabetic hyperglycaemia activates CaMKII and arrhythmias by O-linked glycosylation. Nature 502, 372–376 (2013).
Bolanle, I. O. & Palmer, T. M. Targeting protein O-GlcNAcylation, a link between type 2 diabetes mellitus and inflammatory disease. Cells 11, 705 (2022).
Bolanle, I. O., Riches-Suman, K., Williamson, R. & Palmer, T. M. Emerging roles of protein O-GlcNAcylation in cardiovascular diseases: insights and novel therapeutic targets. Pharmacol. Res. 165, 105467 (2021).
Wang, H. F. et al. Protein O-GlcNAcylation in cardiovascular diseases. Acta Pharmacol. Sin. 44, 8–18 (2023).
Yuzwa, S. A. & Vocadlo, D. J. O-GlcNAc and neurodegeneration: biochemical mechanisms and potential roles in Alzheimer’s disease and beyond. Chem. Soc. Rev. 43, 6839–6858 (2014).
Yuzwa, S. A. et al. Increasing O-GlcNAc slows neurodegeneration and stabilizes tau against aggregation. Nat. Chem. Biol. 8, 393–399 (2012).
Cheng, S. S., Mody, A. C. & Woo, C. M. Opportunities for therapeutic modulation of O-GlcNAc. Chem. Rev. 124, 12918–13019 (2024).
Slawson, C. & Hart, G. W. O-GlcNAc signalling: implications for cancer cell biology. Nat. Rev. Cancer 11, 678–684 (2011).
Sakabe, K., Wang, Z. H. & Hart, G. W. -β-N-acetylglucosamine (O-GlcNAc) is part of the histone code. Proc. Natl Acad. Sci. USA 107, 19915–19920 (2010).
Dupas, T., Lauzier, B. & McGraw, S. O-GlcNAcylation: the sweet side of epigenetics. Epigenetics Chromatin 16, 49 (2023).
Hanover, J. A., Krause, M. W. & Love, D. C. Bittersweet memories: linking metabolism to epigenetics through O-GlcNAcylation. Nat. Rev. Mol. Cell Biol. 13, 312–321 (2012).
Chen, Q., Chen, Y., Bian, C., Fujiki, R. & Yu, X. TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 493, 561–564 (2013).
Fujiki, R. et al. GlcNAcylation of histone H2B facilitates its monoubiquitination. Nature 480, 557–560 (2011).
Xu, B. et al. Histone methyltransferase Dot1L recruits O-GlcNAc transferase to target chromatin sites to regulate histone O-GlcNAcylation. J. Biol. Chem. 298, 102115 (2022).
Wang, P. F. et al. OGT mediated histone H2B S112 GlcNAcylation regulates DNA damage response. J. Genet. Genomics 42, 467–475 (2015).
Wojcik, F. et al. Functional crosstalk between histone H2B ubiquitylation and H2A modifications and variants. Nat. Commun. 9, 1394 (2018).
Deng, Z. et al. Mechanistic insights into nucleosomal H2B monoubiquitylation mediated by yeast Bre1–Rad6 and its human homolog RNF20/RNF40-hRAD6A. Mol. Cell 83, 3080–3094 (2023).
Wisnovsky, S. & Bertozzi, C. R. Reading the glyco-code: new approaches to studying protein–carbohydrate interactions. Curr. Opin. Struct. Biol. 75, 102395 (2022).
Schmaltz, R. M., Hanson, S. R. & Wong, C. H. Enzymes in the synthesis of glycoconjugates. Chem. Rev. 111, 4259–4307 (2011).
Payne, R. J. & Wong, C. H. Advances in chemical ligation strategies for the synthesis of glycopeptides and glycoproteins. Chem. Commun. 46, 21–43 (2010).
Grogan, M. J., Pratt, M. R., Marcaurelle, L. A. & Bertozzi, C. R. Homogeneous glycopeptides and glycoproteins for biological investigation. Annu. Rev. Biochem. 71, 593–634 (2002).
Pratt, M. R. & Bertozzi, C. R. Synthetic glycopeptides and glycoproteins as tools for biology. Chem. Soc. Rev. 34, 58–68 (2005).
Lv, P. et al. O-GlcNAcylation modulates liquid-liquid phase separation of SynGAP/PSD-95. Nat. Chem. 14, 831–840 (2022).
Dong, S. et al. Recent advances in chemical protein synthesis: method developments and biological applications. Sci. China Chem. 67, 1060–1096 (2024).
Li, C. & Wang, L. X. Chemoenzymatic methods for the synthesis of glycoproteins. Chem. Rev. 118, 8359–8413 (2018).
Ye, F. R. et al. Synthetic homogeneous glycoforms of the SARS-CoV-2 spike receptor-binding domain reveals different binding profiles of monoclonal antibodies. Angew. Chem. Int. Ed. Engl. 60, 12904–12910 (2021).
Zhao, J. et al. Chemical synthesis creates single glycoforms of the ectodomain of herpes simplex virus-1 glycoprotein D. J. Am. Chem. Soc. 146, 2615–2623 (2023).
Balana, A. T. et al. O-GlcNAc forces an α-synuclein amyloid strain with notably diminished seeding and pathology. Nat. Chem. Biol. 20, 410–421 (2024).
Riera, R. et al. Single-molecule imaging of glycan–lectin interactions on cells with Glyco-PAINT. Nat. Chem. Biol. 17, 1281–1288 (2021).
Fan, X. et al. Cell-type-specific labeling and profiling of glycans in living mice. Nat. Chem. Biol. 18, 625–633 (2022).
Mukherjee, M. M. et al. Selective bioorthogonal probe for N-glycan hybrid structures. Nat. Chem. Biol. 21, 681–692 (2025).
Tang, F. et al. Selective N-glycan editing on living cell surfaces to probe glycoconjugate function. Nat. Chem. Biol. 16, 766–775 (2020).
Ong, Q. et al. Spatiotemporal control of subcellular O-GlcNAc signaling using Opto-OGT. Nat. Chem. Biol. 21, 300–308 (2025).
Ai, H. S. et al. H2B Lys34 ubiquitination induces nucleosome distortion to stimulate Dot1L activity. Nat. Chem. Biol. 18, 972–980 (2022).
Tong, Z. et al. Synovial sarcoma X breakpoint 1 protein uses a cryptic groove to selectively recognize H2AK119Ub nucleosomes. Nat. Struct. Mol. Biol. 31, 300–310 (2024).
Ai, H. S., Pan, M. & Liu, L. Chemical synthesis of human proteoforms and application in biomedicine. ACS Cent. Sci. 10, 1442–1459 (2024).
Shi, Q. et al. Promotion of RNF168-mediated nucleosomal H2A ubiquitylation by structurally defined K63-polyubiquitylated linker histone H1. Angew. Chem. Int. Ed. Engl. 64, e202413651 (2025).
Ai, H. et al. Mechanism of nucleosomal H2A K13/15 monoubiquitination and adjacent dual monoubiquitination by RNF168. Nat. Chem. Biol. 21, 668–680 (2025).
Pedersen, S. L., Tofteng, A. P., Malik, L. & Jensen, K. J. Microwave heating in solid-phase peptide synthesis. Chem. Soc. Rev. 41, 1826–1844 (2012).
Qu, Q. et al. A highly efficient synthesis of polyubiquitin chains. Adv. Sci. 5, 1800234 (2018).
Vanier, G. S. Microwave-assisted solid-phase peptide synthesis based on the Fmoc protecting group strategy (CEM). Methods Mol. Biol. 1047, 235–249 (2013).
Fang, G. M. et al. Protein chemical synthesis by ligation of peptide hydrazides. Angew. Chem. Int. Ed. Engl. 50, 7645–7649 (2011).
Zheng, J. S., Tang, S., Qi, Y. K., Wang, Z. P. & Liu, L. Chemical synthesis of proteins using peptide hydrazides as thioester surrogates. Nat. Protoc. 8, 2483–2495 (2013).
Mitchell, S. A., Pratt, M. R., Hruby, V. J. & Polt, R. Solid-phase synthesis of O-linked glycopeptide analogues of enkephalin. J. Org. Chem. 66, 2327–2342 (2001).
Wan, Q. & Danishefsky, S. J. Free-radical-based, specific desulfurization of cysteine: a powerful advance in the synthesis of polypeptides and glycopolypeptides. Angew. Chem. Int. Ed. Engl. 46, 9248–9252 (2007).
Onishi, S. et al. Structure of the human Bre1 complex bound to the nucleosome. Nat. Commun. 15, 2580 (2024).
Zheng, Q. et al. A bifunctional molecule-assisted synthesis of mimics for use in probing the ubiquitination system. Nat. Protoc. 18, 530–554 (2023).
Pan, M. et al. Structural insights into Ubr1-mediated N-degron polyubiquitination. Nature 600, 334–338 (2021).
Zhao, F., Hicks, C. W. & Wolberger, C. Mechanism of histone H2B monoubiquitination by Bre1. Nat. Struct. Mol. Biol. 30, 1623–1627 (2023).
de Oliveira, J. F. et al. Mechanistic insights revealed by a UBE2A mutation linked to intellectual disability. Nat. Chem. Biol. 15, 62–70 (2019).
Horn-Ghetko, D. et al. Ubiquitin ligation to F-box protein targets by SCF–RBR E3–E3 super-assembly. Nature 590, 671–676 (2021).
Baek, K. et al. NEDD8 nucleates a multivalent Cullin–RING–UBE2D ubiquitin ligation assembly. Nature 578, 461–466 (2020).
Yunus, A. A. & Lima, C. D. Lysine activation and functional analysis of E2-mediated conjugation in the SUMO pathway. Nat. Struct. Mol. Biol. 13, 491–499 (2006).
Weake, V. M. & Workman, J. L. Histone ubiquitination: triggering gene activity. Mol. Cell 29, 653–663 (2008).
Deng, Z. H., Ai, H. S., Lu, C. P. & Li, J. B. The Bre1/Rad6 machinery: writing the central histone ubiquitin mark on H2B and beyond. Chromosome Res. 28, 247–258 (2020).
Fierz, B. et al. Histone H2B ubiquitylation disrupts local and higher-order chromatin compaction. Nat. Chem. Biol. 7, 113–119 (2011).
Sun, Z. W. & Allis, C. D. Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature 418, 104–108 (2002).
Briggs, S. D. et al. Gene silencing: trans-histone regulatory pathway in chromatin. Nature 418, 498 (2002).
Ng, H. H., Xu, R. M., Zhang, Y. & Struhl, K. Ubiquitination of histone H2B by Rad6 is required for efficient Dot1-mediated methylation of histone H3 lysine 79. J. Biol. Chem. 277, 34655–34657 (2002).
Pavri, R. et al. Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II. Cell 125, 703–717 (2006).
McGinty, R. K., Kim, J., Chatterjee, C., Roeder, R. G. & Muir, T. W. Chemically ubiquitylated histone H2B stimulates hDot1L-mediated intranucleosomal methylation. Nature 453, 812–816 (2008).
Kim, J. et al. RAD6-Mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137, 459–471 (2009).
Zhao, J. et al. O-GlcNAcylation stimulates the deubiquitination activity of USP16 and regulates cell cycle progression. J. Biol. Chem. 300, 107150 (2024).
Han, C. et al. O-GlcNAcylation of SIRT1 enhances its deacetylase activity and promotes cytoprotection under stress. Nat. Commun. 8, 1491 (2017).
Liu, J. et al. Tandem intermolecular [4 + 2] cycloadditions are catalysed by glycosylated enzymes for natural product biosynthesis. Nat. Chem. 15, 1083–1090 (2023).
Tarrant, M. K. et al. Regulation of CK2 by phosphorylation and O-GlcNAcylation revealed by semisynthesis. Nat. Chem. Biol. 8, 262–269 (2012).
Dias, W. B., Cheung, W. D., Wang, Z. & Hart, G. W. Regulation of calcium/calmodulin-dependent kinase IV by O-GlcNAc modification. J. Biol. Chem. 284, 21327–21337 (2009).
Salguero, A. L. et al. Multifaceted regulation of Akt by diverse C-terminal post-translational modifications. ACS Chem. Biol. 17, 68–76 (2022).
Schwein, P. A. et al. Writing and erasing O-GlcNAc on casein kinase 2 alpha alters the phosphoproteome. ACS Chem. Biol. 17, 1111–1121 (2022).
Raj, R., Lercher, L., Mohammed, S. & Davis, B. G. Synthetic nucleosomes reveal that GlcNAcylation modulates direct interaction with the FACT complex. Angew. Chem. Int. Ed. Engl. 55, 8918–8922 (2016).
Dann, G. P. et al. ISWI chromatin remodellers sense nucleosome modifications to determine substrate preference. Nature 548, 607–611 (2017).
Seenaiah, M., Jbara, M., Mali, S. M. & Brik, A. Convergent versus sequential protein synthesis: the case of ubiquitinated and glycosylated H2B. Angew. Chem. Int. Ed. Engl. 54, 12374–12378 (2015).
Xu, T. H. et al. Structure of nucleosome-bound DNA methyltransferases DNMT3A and DNMT3B. Nature 586, 151–155 (2020).
Zheng, Q. et al. An E1-catalyzed chemoenzymatic strategy to isopeptide-N-ethylated deubiquitylase-resistant ubiquitin probes. Angew. Chem. Int. Ed. Engl. 59, 13496–13501 (2020).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. Biol. Crystallogr. 60, 2126–2132 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Biol. Crystallogr. 75, 861–877 (2019).
Acknowledgements
This study was supported by the National Key R&D Program of China (2025YFA1310000/2023YFA0915300 to M.P.), the National Natural Science Foundation of China (22207065 to Y.L., 22137005, 92253302, 22227810, T2488301 to L.L., 22277073 to M.P., 32501108 to H.A.), the Shanghai Natural Science Foundation (25ZR1402193 to H.A.), the Shanghai Frontiers Science Center of Drug Target Identification and Delivery (ZXWH2170101 to H.A.), the Shanghai Rising-Star Program (22QA1404900 to M.P.), the Shanghai Pilot Program for Basic Research - Shanghai Jiao Tong University (21TQ1400224 to M.P.), the Taishan Scholars Program (tsqn202306268 to Y.L.) and the New Cornerstone Investigator Program (to L.L.). Z.D. acknowledges support from the Shuimu Tsinghua Scholars Program and National Facility for Translational Medicine (Shanghai). We acknowledge the Tsinghua University Branch of China National Center for Protein Sciences for cryo-EM data screening and collection.
Author information
Authors and Affiliations
Contributions
Conceptualization, H.A., Z.D., M.P. and L.L. Design of experiments, Z.D., H.A. and L.L. Synthesis of glycosylated histone H2B variants, H.A. and S.T. Synthesis of ubiquitylation intermediate mimic, Z.D., H.A., L.Z. and S.T. Synthesis of glycobased amino acid building blocks, Y.L. and X.D. Cloning and protein purification, Y.D., Z.D., M.S., L.Z., S.T., Q.S. and Z.T. Cryo-EM sample preparation, Z.D. Cryo-EM data processing and model building, H.A. and Z.D. Biochemical assays, Z.D. and Y.D. Visualization and figure preparation, Z.D. Writing—original draft, H.A. and Z.D. Writing—review and editing, all authors. Funding and supervision, H.A., M.P. and L.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Yunpeng Feng, Mingxuan Wu and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Structure and synthetic intermediates of H2BS112GlcNAc.
a, Chemical structure of O-GlcNAc modified on H2B S112 residue. The distance from the H2B S112 αC atom to the C4-OH hydrogen of O-GlcNAc is 8.5 Å based on its atomic model. b, In our reported structure of the RNF20/RNF40–RAD6A–nucleosome complex (PDB: 8IEJ), a semi-transparent green spherical with a radius of 8.5 Å is centered on the H2B S112 αC atom. Within this structural model, O-GlcNAc modified on H2B S112 may potentially interact with RNF20 or RAD6A. c, Representative RP-HPLC trace (214 nm) of purified 1. d, Representative ESI-MS spectra and deconvoluted MS spectra of purified 1. e, Representative RP-HPLC trace (214 nm) of crude 2, crude 3, and purified 3. f, Representative ESI-MS spectra and deconvoluted MS spectra of crude 2. g, Representative ESI-MS spectra and deconvoluted MS spectra of purified 3. h, RP-HPLC monitoring of peptide hydrazide ligation from 1 and 3 to generate 4. Peaks of 3 and 4 are indicated. i, Representative RP-HPLC trace (214 nm) of purified 4. j, Representative ESI-MS spectra and deconvoluted MS spectra of purified 4.
Extended Data Fig. 2 Design and synthesis of the nucleosomal H2B ubiquitylation mimic SP1 complex.
a, Schematic representation of H2B ubiquitylation by human RNF20/RNF40–RAD6A complex on H2BS112GlcNAc nucleosome. The heterodimeric RNF20/RNF40 RING domains bind RAD6A~Ub thioester non-covalently (indicated by a dashed gray arrow) to form the tetrahedron transition state. b, Schematic representation of the engineered mimic (SP1) of H2B ubiquitylation intermediate. The RNF20 RING domain is covalently fused with RAD6A by SICC strategy. The covalent linker and the non-covalent interactions between RNF20 RING and RAD6A are indicated as a solid green line and a dashed gray arrow, respectively. c, Amino acid sequence of histone H2B K120C mutant. The ligation sites A58 and A97, the glycosylation site S112, and the conjugating site K120C are indicated by red, light sea green, and orange solid circles, respectively. d, Convergent synthetic route of probe 17.
Extended Data Fig. 3 Mass spectrometry of probe 17 synthetic intermediates.
a–l, Representative ESI-MS spectra and deconvoluted MS spectra of synthetic intermediates 6 (a), 8 (b), 7 (c), 9 (d), 10 (e), 11 (f), 12 (g), UbMesNa (h), 13 (i), 14 (j), 15 (k), and 16 (l) shown in Extended Data Fig. 2.
Extended Data Fig. 4 Cryo-EM data processing for RNF20/RNF40–RAD6A–NCPGlcNAc complex.
a, Representative micrograph from the dataset. b, CTF estimation of the micrograph shown in a. c, Representative 2D averages. d, Data processing flowchart. e, The final reconstruction colored by local resolution. f, Euler angle distributions of the final reconstruction. g, Fourier shell correlation curve calculated from final refinement. The resolution at FSC = 0.143 cutoff was highlighted. h, Fourier shell correlation curve for model-map correlation shown at FSC = 0.5.
Extended Data Fig. 5 Sample cryo-EM densities and conformational flexibility analysis.
a, Nucleosmal 147-bp DNA density with atomic model fitted. b, Histone octamer density with cartoon model fitted. c, Sample densities of histone H2A, H2B, H3, and H4, and nucleosomal SHL0 DNA with atomic model fitted. d, Cryo-EM density and model of the H2B αC helix (residues 103–125) modified with O-GlcNAc. e–h, RNF20 RING domain (g) and RNF40 RING domain (h) densities p with cartoon model fitted. i, Conformational flexibility analysis of the complex. Cryo-EM maps and models of three distinct states are shown. j, Structural alignment of the three conformational states, highlighting the rotation of the RNF20/RNF40 RING dimer and the movement of ubiquitin.
Extended Data Fig. 6 Biochemical investigations of H2BS112GlcNAc-promoted H2BK120ub generation.
a, SYBR-Gold-stained EMSA gels analyzing the binding affinity of human RNF20/RNF40 to NCPunmod. (top) or NCPGlcNAc (bottom). A red line with circular endpoints indicates the RNF20/RNF40 concentration (1,280 nM) that shifts approximately half of the free nucleosomes (NCPunmod. or NCPGlcNAc) to form complexes. b, Cryo-EM density and models of H2BS112GlcNAc–RAD6A Q93 interaction. c, SDS-PAGE characterization of wild-type (WT) RAD6A and its Q93A/I, K66A/I mutants. d, Representative fluorescent and CBB-stained gel images of in vitro H2B ubiquitylation assay analyzing activities of WT RAD6A and its mutants on NCPunmod. or NCPGlcNAc substrates. e, Quantified data of RAD6A mutant assay shown in d. Data show each replicate and error bars indicate mean ± SD from n = 3 independent replicates. f, Bar graph depicting fold changes in ubiquitylation activity, calculated as the ratio of NCPGlcNAc to NCPunmod. levels (NCPGlcNAc/NCPunmod.), derived from panel e. A two-sided, unpaired Student’s t-test was employed to calculate p-values; ns means not significant (p > 0.05). g, Representative fluorescent and CBB-stained gel images of single-turnover assay analyzing H2B ubiquitylation rates on NCPGlcNAc or NCPunmod. at a defined range of RAD6A concentrations. h,i Representative fluorescent and CBB-stained gel images of single-turnover assay analyzing H2B ubiquitylation rates on NCPGlcNAc or NCPunmod. at a pH range of 6.0–9.0 (h) or 7.0–7.5 (i). j, Representative fluorescent and CBB-stained gel images of single-turnover assay analyzing ubiquitin transfer from RAD6A~Ub thioester to nucleosomal H2B K120. k, Quantified data of single-turnover assay shown in j. Data shows the results from n = 2 independent replicates. The histone H4 bands in the CBB-stained gel image were used as the loading control in d and g–k. Assays were performed as n = 3 independent replicates in g–i.
Extended Data Fig. 7 Chemically synthesized monoglycosylated H2B variants were incorporated into nucleosomes.
a, Amino acid sequence of histone H2B. The ligation sites A58 and A97, and the glycosylation site S112 are indicated by red and light sea green solid circles, respectively. b, Convergent synthetic route of monoglycosylated H2B variants (G-H2B). The chemical structures of O-D-pyranoses studied here are shown at the bottom. c, Size-exclusion chromatograms of H2A/G-H2B histone dimers. d, Representative CBB-stained gel image of purified H2A/G-H2B histone dimers. e, Representative CBB-stained SDS-PAGE (top) and SYBR-Gold-stained native PAGE (bottom) gel images of purified H2B-monoglycosylated NCPs for biochemical assays. Two independent PAGE analyses in d and e were performed with similar results.
Extended Data Fig. 8 Mass spectrometries of monoglycosylated H2B synthetic intermediates.
a–e, Representative ESI-MS spectra and deconvoluted MS spectra of synthetic intermediates of H2BS112-β-GlcN (a), H2BS112-β-Glc (b), H2BS112-β-GalNAc (c), H2BS112-α-GlcNAc (d), and H2BS112-β-Xyl (e).
Extended Data Fig. 9 Structure-activity relationship analysis of the O-linked GlcNAcylation on H2B using synthetic H2B-monoglycosylated NCP substrates.
a, Representative fluorescent and CBB-stained gel images of in vitro H2B ubiquitylation assay analyzing activities of RNF20/RNF40–RAD6A on different H2B-monoglycosylated NCP substrates. b, Representative fluorescent and CBB-stained gel images of single-turnover assay analyzing H2B ubiquitylation rates on different H2B-monoglycosylated NCP substrates at a defined range of RAD6A concentrations. c, Representative fluorescent and CBB-stained gel images of single-turnover assay analyzing H2B ubiquitylation rates on different H2B-monoglycosylated NCP substrates at a pH range of 6.0–8.5. Assays were performed as n = 3 independent replicates in a–c.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2, Figs. 1–12, Note and Uncropped blots for Supplementary Fig. 1.
Source data
Source Data Fig. 1 (download PDF )
Unprocessed western blots and/or gels.
Source Data Fig. 2 (download PDF )
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots and/or gels.
Source Data Extended Data Fig. 9 (download PDF )
Unprocessed western blots and/or gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Deng, Z., Tao, S., Du, Y. et al. Allosteric activation of RNF20/RNF40–RAD6A-mediated H2BK120 monoubiquitylation by H2BS112 GlcNAcylation. Nat Chem Biol (2026). https://doi.org/10.1038/s41589-025-02109-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41589-025-02109-6