Abstract
CRISPR–Cas9, an RNA-guided immune system, functions specifically in bacteria while controlling autoimmunity. However, its application to genome editing often causes deleterious off-target cleavages. Here, by sequencing CRISPR RNAs (crRNAs), we discovered abasic modifications that naturally suppress off-target self-cleavages from activated Cas9 in Streptococcus pyogenes (SpCas9). Bacteriophage infection induces oxidative stress, preferentially oxidizing the 5′ end of crRNAs into abasic modifications. Mechanistically, abasic substitutions at the 5′ end reduce off-target effects by limiting base pairing while preserving SpCas9-interacting backbones to maintain on-target efficiency. Abasic extensions at the 5′ end reduce off-target effects by sterically constraining SpCas9 but retain on-target activity by avoiding extra base pairs. Moreover, these approaches can be combined (abasic substitution and extension), enhancing SpCas9 fidelity by increasing mismatch intolerance at the protospacer-adjacent motif-distal region and outperforming SpCas9 variants. Biologically inspired, we developed abasic chemical modifications for guide RNAs that improve CRISPR–Cas9 genome-editing specificity, demonstrating potential for in vivo application.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All raw sequencing data from synthetic abasic RNA (SRP532234), S. pyogenes crRNAs (SRP532228 and SRP532225), ARP-crRNA-seq (SRP532072), targeted sequencing of SF370 genome (SRP585061 and SRP585068), targeted sequencing of human genome (SRP466382 and SRP637086), CIRCLE-seq (SRP456533, SRP456527 and SRP456526), random mismatch libraries (SRP455464, SRP455558, SRP455560, SRP455565, SRP585086, SRP649377 and SRP649387) and RNA-seq (SRP466117, SRP532284 and SRP541971) were deposited to the SRA database. All FASTQ files are available online (http://clip.korea.ac.kr/abasic_crRNA/). Source data are provided with this paper.
Code availability
The Python scripts used in this work are available from GitHub (https://github.com/DotchGNU/crRNA-seq) and Zenodo (https://doi.org/10.5281/zenodo.17762942)80.
References
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Sander, J. D. & Joung, J. K. CRISPR–Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Doudna, J. A. & Charpentier, E. The new frontier of genome engineering with CRISPR–Cas9. Science 346, 1258096 (2014).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Brouns, S. J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964 (2008).
Marraffini, L. A. & Sontheimer, E. J. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322, 1843–1845 (2008).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9–crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl Acad. Sci. USA 109, E2579–E2586 (2012).
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Heler, R. et al. Cas9 specifies functional viral targets during CRISPR–Cas adaptation. Nature 519, 199–202 (2015).
Ishino, Y., Shinagawa, H., Makino, K., Amemura, M. & Nakata, A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J. Bacteriol. 169, 5429–5433 (1987).
Mojica, F. J. M., Diez-Villasenor, C., Garcia-Martinez, J. & Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology (Reading) 155, 733–740 (2009).
Mojica, F. J., Diez-Villasenor, C., Garcia-Martinez, J. & Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 60, 174–182 (2005).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Cho, S. W., Kim, S., Kim, J. M. & Kim, J. S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 230–232 (2013).
Tsai, S. Q. & Joung, J. K. Defining and improving the genome-wide specificities of CRISPR–Cas9 nucleases. Nat. Rev. Genet. 17, 300–312 (2016).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR–Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Lin, Y. et al. CRISPR/Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res. 42, 7473–7485 (2014).
Zhang, X. H., Tee, L. Y., Wang, X. G., Huang, Q. S. & Yang, S. H. Off-target effects in CRISPR/Cas9-mediated genome engineering. Mol. Ther. Nucleic Acids 4, e264 (2015).
Deveau, H., Garneau, J. E. & Moineau, S. CRISPR/Cas system and its role in phage–bacteria interactions. Annu. Rev. Microbiol. 64, 475–493 (2010).
Couvin, D. et al. CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res. 46, W246–W251 (2018).
Chen, X. & Wolin, S. L. Transfer RNA halves are found as nicked tRNAs in cells: evidence that nicked tRNAs regulate expression of an RNA repair operon. RNA 29, 620–629 (2023).
Bates, C. S., Montañez, G. E., Woods, C. R., Vincent, R. M. & Eichenbaum, Z. Identification and characterization of a Streptococcus pyogenes operon involved in binding of hemoproteins and acquisition of iron. Infect. Immun. 71, 1042–1055 (2003).
Nakamura, T. et al. Two-dimensional gel electrophoresis analysis of the abundance of virulent exoproteins of group A Streptococcus caused by environmental changes. Arch. Microbiol. 181, 74–81 (2004).
Yanagawa, H., Ogawa, Y. & Ueno, M. Redox ribonucleosides. Isolation and characterization of 5-hydroxyuridine, 8-hydroxyguanosine, and 8-hydroxyadenosine from torula yeast RNA. J. Biol. Chem. 267, 13320–13326 (1992).
Küpfer, P. A., Crey-Desbiolles, C. & Leumann, C. J. trans-Lesion synthesis and RNaseH activity by reverse transcriptases on a true abasic RNA template. Nucleic Acids Res. 35, 6846–6853 (2007).
Ide, H. et al. Synthesis and damage specificity of a novel probe for the detection of abasic sites in DNA. Biochemistry 32, 8276–8283 (1993).
Tanaka, M., Han, S., Kupfer, P. A., Leumann, C. J. & Sonntag, W. E. An assay for RNA oxidation induced abasic sites using the aldehyde reactive probe. Free Radic. Res. 45, 237–247 (2011).
Hahm, J. Y., Park, J., Jang, E. S. & Chi, S. W. 8-Oxoguanine: from oxidative damage to epigenetic and epitranscriptional modification. Exp. Mol. Med. 54, 1626–1642 (2022).
Dong, T. G. et al. Generation of reactive oxygen species by lethal attacks from competing microbes. Proc. Natl Acad. Sci. USA 112, 2181–2186 (2015).
Eom, S. et al. Widespread 8-oxoguanine modifications of miRNA seeds differentially regulate redox-dependent cancer development. Nat. Cell Biol. 25, 1369–1383 (2023).
Seok, H. et al. Position-specific oxidation of miR-1 encodes cardiac hypertrophy. Nature 584, 279–285 (2020).
Montes, M. & Huarte, M. o8G modifications rewire tumoral microRNAs. Nat. Cell Biol. 25, 1243–1244 (2023).
Hall, D. B., Holmlin, R. E. & Barton, J. K. Oxidative DNA damage through long-range electron transfer. Nature 382, 731–735 (1996).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014).
Fu, Y., Sander, J. D., Reyon, D., Cascio, V. M. & Joung, J. K. Improving CRISPR–Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Lee, H. S. et al. Abasic pivot substitution harnesses target specificity of RNA interference. Nat. Commun. 6, 10154 (2015).
Ramakrishna, S. et al. Surrogate reporter-based enrichment of cells containing RNA-guided Cas9 nuclease-induced mutations. Nat. Commun. 5, 3378 (2014).
Kleinstiver, B. P. et al. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016).
Jiang, F. et al. Structures of a CRISPR–Cas9 R-loop complex primed for DNA cleavage. Science 351, 867–871 (2016).
Cho, S. W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res 24, 132–141 (2014).
Huai, C. et al. Structural insights into DNA cleavage activation of CRISPR–Cas9 system. Nat. Commun. 8, 1375 (2017).
Okafor, I. C. et al. Single molecule analysis of effects of non-canonical guide RNAs and specificity-enhancing mutations on Cas9-induced DNA unwinding. Nucleic Acids Res. 47, 11880–11888 (2019).
Tsai, S. Q. et al. CIRCLE-seq: a highly sensitive in vitro screen for genome-wide CRISPR–Cas9 nuclease off-targets. Nat. Methods 14, 607–614 (2017).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR–Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Jung, C. et al. Massively parallel biophysical analysis of CRISPR–Cas complexes on next generation sequencing chips. Cell 170, 35–47 (2017).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Chen, J. S. et al. Enhanced proofreading governs CRISPR–Cas9 targeting accuracy. Nature 550, 407–410 (2017).
Lee, J. K. et al. Directed evolution of CRISPR–Cas9 to increase its specificity. Nat. Commun. 9, 3048 (2018).
Vakulskas, C. A. et al. A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells. Nat. Med. 24, 1216–1224 (2018).
Chung, S. H. et al. CRISPR-based VEGF suppression using paired guide RNAs for treatment of choroidal neovascularization. Mol. Ther. Nucleic Acids 28, 613–622 (2022).
Liang, C. et al. Tumor cell-targeted delivery of CRISPR/Cas9 by aptamer-functionalized lipopolymer for therapeutic genome editing of VEGFA in osteosarcoma. Biomaterials 147, 68–85 (2017).
Haapaniemi, E., Botla, S., Persson, J., Schmierer, B. & Taipale, J. CRISPR–Cas9 genome editing induces a p53-mediated DNA damage response. Nat. Med. 24, 927–930 (2018).
Wilhelm, S. M. et al. Preclinical overview of sorafenib, a multikinase inhibitor that targets both RAF and VEGF and PDGF receptor tyrosine kinase signaling. Mol Cancer Ther 7, 3129–3140 (2008).
Workman, R. E. et al. Anti-CRISPR proteins trigger a burst of CRISPR–Cas9 expression that enhances phage defense. Cell Rep. 43, 113849 (2024).
Stoltzfus, M. J., Workman, R. E., Keith, N. C. & Modell, J. W. A dynamic subpopulation of CRISPR–Cas overexpressers allows Streptococcus pyogenes to rapidly respond to phage. Nat. Microbiol. 9, 2410–2421 (2024).
Margolin, Y., Cloutier, J. F., Shafirovich, V., Geacintov, N. E. & Dedon, P. C. Paradoxical hotspots for guanine oxidation by a chemical mediator of inflammation. Nat. Chem. Biol. 2, 365–366 (2006).
Koizume, S., Inoue, H., Kamiya, H. & Ohtsuka, E. Neighboring base damage induced by permanganate oxidation of 8-oxoguanine in DNA. Nucleic Acids Res. 26, 3599–3607 (1998).
Saito, I. et al. Mapping of the hot spots for DNA damage by one-electron oxidation: efficacy of GG doublets and GGG triplets as a trap in long-range hole migration. J. Am. Chem. Soc. 120, 12686–12687 (1998).
Ryan, D. E. et al. Improving CRISPR–Cas specificity with chemical modifications in single-guide RNAs. Nucleic Acids Res. 46, 792–803 (2018).
Cromwell, C. R. et al. Incorporation of bridged nucleic acids into CRISPR RNAs improves Cas9 endonuclease specificity. Nat. Commun. 9, 1448 (2018).
Yin, H. et al. Partial DNA-guided Cas9 enables genome editing with reduced off-target activity. Nat. Chem. Biol. 14, 311–316 (2018).
Kim, D., Luk, K., Wolfe, S. A. & Kim, J. S. Evaluating and enhancing target specificity of gene-editing nucleases and deaminases. Annu. Rev. Biochem. 88, 191–220 (2019).
Seok, H., Lee, H., Jang, E. S. & Chi, S. W. Evaluation and control of miRNA-like off-target repression for RNA interference. Cell. Mol. Life Sci. 75, 797–814 (2018).
Park, J. et al. siAbasic: a comprehensive database for potent siRNA-6O sequences without off-target effects. Database (Oxford) 2018, bay109 (2018).
Beerens, D. et al. Survival strategies of Streptococcus pyogenes in response to phage infection. Viruses 13, 612 (2021).
Lyublinskaya, O. G. et al. Flow cytometric HyPer-based assay for hydrogen peroxide. Free Radic. Biol. Med. 128, 40–49 (2018).
Forman, H. J., Bernardo, A. & Davies, K. J. What is the concentration of hydrogen peroxide in blood and plasma? Arch. Biochem. Biophys. 603, 48–53 (2016).
Antoniali, G. et al. Mammalian APE1 controls miRNA processing and its interactome is linked to cancer RNA metabolism. Nat. Commun. 8, 797 (2017).
Kukurba, K. R. & Montgomery, S. B. RNA sequencing and analysis. Cold Spring Harb. Protoc. 2015, 951–969 (2015).
Zhang, L.-S. et al. m7G-quant-seq: quantitative detection of RNA internal N7-methylguanosine. ACS Chem. Biol. 17, 3306–3312 (2022).
Kang, S. H. et al. Prediction-based highly sensitive CRISPR off-target validation using target-specific DNA enrichment. Nat. Commun. 11, 3596 (2020).
Park, J., Lim, K., Kim, J. S. & Bae, S. Cas-analyzer: an online tool for assessing genome editing results using NGS data. Bioinformatics 33, 286–288 (2017).
Ruiz, S. et al. A genome-wide CRISPR screen identifies CDC25A as a determinant of sensitivity to ATR inhibitors. Mol. Cell 62, 307–313 (2016).
Liu, D. et al. Nuclear import of proinflammatory transcription factors is required for massive liver apoptosis induced by bacterial lipopolysaccharide. J. Biol. Chem. 279, 48434–48442 (2004).
Kim, G.-W. D. crRNA-seq. Zenodo https://doi.org/10.5281/zenodo.17762942 (2025).
Acknowledgements
We thank all members of the S.W.C. laboratory for their help and discussions. We thank E. Choi and E.-J. Lee for help in experiments with S. pyogenes and thank J. S. Yoo and J. W. Hur for their initial assistance in preliminary studies. We are especially grateful to the late S.-D. Yoo and the late Y.-S. Lee for motivating us to apply CRISPR–Cas9 to the treatment of liver cancer. We thank S. Bae for providing mammalian expression vectors of HypaCas9 and Sniper-Cas9, S. W. Nam for providing MIHA cells, J. S. Ko for providing EA.hy926 cells and B. S. Kim for providing 293T cells. This work was supported by the National Research Foundation of Korea grants funded by the Korea government (Ministry of Science and ICT and Ministry of Education; RS-2025-16652968, RS-2024-00442012 and RS-2023-00221182 (to S.W.C.) and RS-2025-24683408 (to D.G.)), from Korea University (to S.W.C.), from the Korea University–Korea Institute of Science and Technology (KIST) Graduate School of Converging Science and Technology Program and from the KIST institutional program (2V09840-23-P024, to S.W.C.).
Author information
Authors and Affiliations
Contributions
D.G. and G.D.K. performed the major experiments. D.G., G.D.K., M.P. and A.D.P. conducted the major bioinformatics analyses. A.D.P. analyzed the S. pyogenes genomes and CRISPR–Cas++ data. G.D.K. conducted the S. pyogenes experiments, crRNA sequencing and ARP-crRNA-seq analyses. G.D.K., J.K., H.O. and S.H.K. cultured S. pyogenes and propagated phage A1. D.G., M.P. and H.S.L. performed the in vitro SpCas9 assays. G.D.K. and H.W. performed the dot blot analyses. J.P. analyzed the sequencing results from oxidized artificial transcript fragments. D.G. and M.P. performed the molecular and cell biology experiments. S.W.C. performed the structural modeling and analyses. D.G., G.D.K., S.E. and N.S. performed the qPCR. D.G. and M.P. performed the surrogate reporter assays. J.K.H. modified and optimized the CIRCLE-seq protocol. D.G. performed the targeted genome sequencing and CIRCLE-seq; D.G., J.L. and S.H.A. analyzed the data. D.G. and M.P. conducted the experiments using random mismatch libraries; D.G., J.L., S.H.A. and M.P. analyzed the data. H.S.L. constructed the RNA-Seq libraries; A.D.P. and G.D.K. analyzed the data. S.E. performed the mouse experiments. S.W.C. supervised the study. D.G., G.D.K., E.-S.J. and S.W.C. conceptualized and designed the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Korea University has filed a Patent Cooperation Treaty application (PCT/KR2025/014286) covering part of the work presented in this article, listing D.G., H.S.L., E.-S.J. and S.W.C. as inventors. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Pranam Chatterjee and Nozomu Yachie for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Identification of abasic crRNAs reducing SpCas9-mediated off-target cleavage in S. pyogenes.
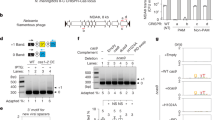
a, b, Genome-wide analysis of OTs in S. pyogenes (n = 107 strains); 481 crRNAs, located < 6 kb from Cas9 loci (mean = 4.5 per strain; a). Among 171 unique spacers, OTs were ordered by off-target count (b). Observed OTs (≤3MM = 13, ≤4MM = 48, ≤5MM = 235, ≤6MM = 1,607) exceeded expected frequencies (≤3MM = 1, ≤4MM = 7, ≤5MM = 73, ≤6MM = 568). c, d, in vitro SpCas9 assays for OTs (≤4MM) using synthetic crRNAs; blue, cleaved OTs; grey triangle, substrate; black triangle, product; cleavage ratio (%). OTs were named by assigning number of mismatch followed by unique alphabetical order (lower case). e–h, In vitro SpCas9 assays as in (c, d) except for OTs (≤6MM) from SF370 (crRNA1 (Spyo1h_001), e; crRNA4 (Spyo1h_004), f; crRNA6 (Spyo1h_006), g) and from other strains harboring the same crRNA spacer sequence as the Bruno strain (crRNAb, h). i, In vitro SpCas9 assays with small RNA fractions (SF370; cold RNA extraction25 to preserve crRNA-tracrRNA complex), reacted with supplemented tracrRNA to prevent defective complex dissociation (via masking potential cross-hybridization); on-targets for crRNA1 (T1), crRNA4 (T4), and crRNA6 (T6). crRNA6 showed marginal activity (0.9%), so only crRNA1 and crRNA4 were analyzed further. j, Cleavage efficiency (%) was quantified by capillary electrophoresis (Fragment analyzer) using concentrated samples for off-targets (for example, 9x). k, Endogenous crRNA estimated at ~0.4–1.2 nM in 2.5 µg small RNA. Assays with 0.1–1.0 nM synthetic crRNAs showed that 0.25 nM synthetic crRNA matched 2.5 µg small RNA on-target activity; cleavage efficiency (%) = cleaved product/total substrate (below gels). l, Stepwise RNA oxidation from all bases to abasic modification (Ø), with preferential guanine and cytosine oxidation (o8G and h5C) leading to G > T and C > T mutations in cDNA via the A-rule (Ø•A pairing during reverse transcription). m, ELISA quantification of abasic modifications in small RNA (Bruno); anaerobic (n = 3) versus aerobic cultures (n = 4),*P = 1.7 x 10−5. n, Dot blots of abasic small RNA (ARP) performed as in Fig. 1m, except SF370. o, Relative ratio of cDNA variant (%) in in vitro oxidized (Fenton reaction) synthetic crRNAs for SF370 (crRNA 1, 4, and 6; n = 2 for 0.1 mM and 0.5 mM H2O2) and crRNAb for Bruno (n = 7 for 0.1 mM (n = 3), 0.5 mM (n = 3), and 2 mM H2O2 (n = 1)), relative to non-oxidized synthetic crRNAs (Oxi/Non); n = 7; *P = 9.9 ×10−5 and 1.5 x 10−4, respectively (Oxi/Non > 2); relative to no change (Oxi/Non = 1). p, In vitro SpCas9 assays comparing between oxidized and non-oxidized synthetic crRNAs at varying H2O2 (2 mM, upper panels; 0.5 mM, lower panels) and crRNA concentrations (50 nM, upper panel; 10 nM, lower panel); normalized cleavage (cleavage ratio normalized to on-target); ratio of normalized cleavage (oxidized/non-oxidized). Of note, similar results were observed across H2O2 and crRNA concentrations, indicating that the reduced off-target cleavage is due to a specific mechanistic effect rather than impaired crRNA function. q, Abasic-induced cDNA variants including incomplete 3’ end of cDNA sequences, analysed using synthetic RNAs with abasic modification at position 7 (7Ø; left and middle panels) and position 5 (5Ø BamHI; right panel) and o8G at position 7 (7o8G), relative to unmodified (7 G). Of note, no deletions were detected at 7o8G, in contrast to 7Ø, 5Ø BamH1 and 5Ø RNA. Unless stated otherwise, P values are two-sided (t-test), n values denote biologically independent samples, and data represent mean ± s.d.
Extended Data Fig. 2 Abasic crRNA oxidation induced by bacteriophage infection reduces SpCas9 off-target cleavage and applies to guide RNAs.
a, Positional distribution of 3′ cDNA ends of crRNA sequencing (read count, %) in SF370 (crRNA1, 4 and 6) was compared (read count, %) between bacteriophage A1 infection (+infection; n = 3) and mock (n = 3; lower graphs), quantitated as Δrelative enrichment (upper graphs), difference of relative enrichment (log2 ratio normalized to synthetic gRNA) between infection and mock; *P = 0.022, 3.7 x 10−4, 0.007, 0.008, 0.019, 4.1 x 10−4, 0.011 and 0.012, respectively (crRNA1); *P = 0.031, 0.039, 0.001, and 0.006, respectively (crRNA4); *P = 0.008 and 0.005, respectively (crRNA6). b, Similar positional distribution analyses as in (a), except examining abasic-induced cDNA variants (deletion, >T, >C) in ARP-crRNA Seq; bacteriophage A1 infection (+infection; n = 2) versus mock (endogenous; n = 2). c, d, Positional analysis of abasic-induced deletions (%) in cDNA of oxidized random RNA fragments, produced by shearing artificial transcripts followed by Fenton reaction. Sequencing data were retrieved and processed as previously described34 (c). Dot blot (d, upper panel) detected abasic modification (ARP) in the oxidized random RNA fragments (SYBR Gold). Deletions in reads were quantified only when other reads (aligned to either 5′ or 3′ ends) displayed Ø>T mutation signatures (G > T, A > T, or C > T) at the same relative positions as the deletion (n = 469,912 (mock) and 491,458 reads (Fenton) in PE150). Deletion frequencies were averaged using a 7-nt sliding window from both the 5′ and 3′ ends across all fragments (shaded area, s.d.; d, lower graphs); n = 7; *P < 0.01 versus mock (P values provided in Source Data). e, Targeted deep sequencing quantifying substitution ratio (relative to mock) at increasing distances from the OT5b cleavage site (n = 3, different distances; ± 20, 30 and 40 bp) in bacteriophage A1–infected SF370 (MOI = 0.1; 1 h post-infection) in the presence of antioxidant NAC pretreatment (5 mM for 24 h). f, g, in vitro SpCas9 assays using synthetic gRNAs targeting human loci (T2, T3, and EMX1) after in vitro oxidation (Fenton reaction, 2 mM H2O2; oxi), assessing on-target (f, left panel) and off-target cleavage (f, right panel); normalized cleavage (%) relative to on-target; n = 3; *P = 3.4 x 10−4, 9.2 x 10−6,1.4 x 10−5, 1.9 ×10−4, 4.0 x 10−5, 0.004, 0.004, 1.1 x 10−4, 2.3 x 10−5, 0.002 and 0.006, respectively; relative to non-oxidized synthetic gRNA (non); average ratio of normalized cleavage (Oxi/Non). Off-target sequences and representative capillary electrophoresis images (g). h, Quantification of RNA stability for abasic T2 gRNA (0.001 nM) containing rSpacer (rØØX20) versus dSpacer (ØØX20) in HEK293T cells, measured by qPCR. Relative expression indicates the abundance of modified T2 gRNA normalized to unmodified T3 gRNA; n = 3, *P = 0.015. Unless stated otherwise, P values are two-sided (t-test), n values denote biologically independent samples, and data represent mean ± s.d.
Extended Data Fig. 3 Abasic substitution in gRNAs reduces off-target activity as much as in tru-gRNA.
a–d, in vitro SpCas9 assays for off-target sites (OTs) of T2 (a, b), T3 (c) and EMX1 (d) using gRNA (X20), tru-gRNA (X17) and Ø3X17 at various concentrations (n = 3 different doses for T2 and T3; n = 1 for EMX1 due to low off-target activity); OT sequences show mismatches in red, PAM in lowercase, and sites aligned to abasic substitution (positions 18-20) underlined (a–d, upper panels). Details of OT sequences for T2 (OT2-LAMA3 and OT2-24, a; OT2-FMN1, b), T3 (OT3-COMDA, OT3-20, OT3-MAX, and OT3-4, c, left to right) and EMX1 (OT4-HCN1, OT4-10 and OT4-MFAP1, d) in Supplementary Table 19. Cleaved fragments (*) visualized by capillary electrophoresis (a, c and d) or agarose gel (b); cleavage rates (%) shown below. e, f, Surrogate reporter assays for T2 and T3 off-target sites using gRNA (X20), tru-gRNA (X17), and Ø3X17 at increasing concentrations (0, 0.02, 0.1, 0.5, 2.5, 15, and 50 nM). Flow cytometry results (log10 GFP vs. log10 RFP; n = 10,000 cells) at 50 nM shown (upper panels); ratio of GFP+ in gate (%; n = 6); relative editing (%, GFP+ / RFP+). EC50 and Emax values are listed below each panel (n = 6); *P = 1.5 x 10−4, 1.3 x 10−6, 1.3 x 10−8 and 2.1 x 10−6, respectively (tru-gRNA for OT2-LAMA3; e); *P = 1.7 x 10−5, 3.8 x 10−8, 1.4 x 10−9 and 7.3 x 10−6, respectively (Ø3X17 for OT2-LAMA3; e); *P = 0.003, 4.3 x 10−4 and 1.9 x 10−13, respectively (tru-gRNA for OT2-24; e); *P = 0.004, 4.9 x 10−4 and 7.3 x 10−9, respectively (Ø3X17 for OT2-24; e); *P = 1.4 x 10−4, 1.2 × 10−7, 1.9 x 10−7 and 1.4 × 10−10, respectively (tru-gRNA for OT3-COMDA; f); *P = 4.7 x 10−4, 1.3 x 10−7, 1.2 x 10−8 and 4.9 x 10−12, respectively (Ø3X17 for 3-COMDA; f); *P = 7.0 x 10−4, 1.6 x 10−6, 1.9 x 10−8 and 3.3 x 10−12, respectively (tru-gRNA for OT3-20; f); *P = 7.0 x 10−5, 2.8 x 10−8, 1.8 x 10−8 and 4.2 x 10−10, respectively (ØX17 for 3-20; f); *P = 2.1 x 10−4, 4.7 x 10−7 and 3.9 x 10−5, respectively (tru-gRNA for OT3-MAX; f); *P = 2.6 x 10−5, 3.4 x 10−9 and 2.2 x 10−10, respectively (Ø3X17 for 3-MAX; f); *P = 7.4 x 10−9, 1.2 x 10−7 and 7.8 x 10−9, respectively (tru-gRNA for OT3-4; f); *P = 3.1 x 10−5, 3.8 x 10−7 and 2.5 × 10−8, respectively (Ø3X17 for 3-4; f); relative to unmodified gRNA (X20). Unless stated otherwise, P values are two-sided (t-test), n values denote biologically independent samples, and data represent mean ± s.d.
Extended Data Fig. 4 Abasic extension in gRNA outperforms GGX20 in target specificity.
a, b, in vitro SpCas9 assays for T2, T3 and EMX1 on-targets using gRNA (X20), GGX20 and ØØX20 at varying concentration: T2 (n = 2 (10, 25, and 50 nM), a; n = 2 (25, 50 and 100 nM), b), T3 and EMX1 (n = 1 (10 nM), 3 (25 nM), and 3 (50 nM), a; n = 2 (25, 50 and 100 nM), b). Cleaved fragments (asterisks) in capillary electrophoresis (upper panel); quantitation of cleavage (%; lower graphs). c–g, in vitro SpCas9 assays for T2 off-targets: OT2-LAMA3 (c, left panel), OT2-24 (c, right panel), and OT2-FMN1 (d), for T3 off-targets: OT3-COMDA (e, left panel), OT3-20 (e, right panel), OT3-MAX (f, left panel), and OT3-4 (f, right panel), and for EMX1 off-targets: OT4-HCN1 (g, upper panel), OT4-10 (g, middle panel), and OT4-MFAP1 (g; lower panel). OT sequences show mismatches in red, PAM in lowercase, and sites aligned to abasic extension (positions 21-22) underlined (c–g, upper panels); cleaved fragments (asterisks) in agarose gel electrophoresis or capillary electrophoresis (c–g, lower panels). Unless stated otherwise, P values are two-sided (t-test), n values denote biologically independent samples, and data represent mean ± s.d.
Extended Data Fig. 5 Abasic substitution and extension (ØXØ) in gRNA improves SpCas9 target specificity.
a, b, in vitro SpCas9 assays for T2, T3, and EMX1 on-targets using ØXØ and unmodified gRNAs at varying concentration (n = 2 different series of doses; 10, 25 and 50 nM, a; 25, 50 and 100 nM, b); cleaved fragments (asterisks) in capillary electrophoresis analyses (a, b, upper panels); quantitation of cleavage (%; a, b, lower graphs). c–g, in vitro SpCas9 assays for T2 off-targets: OT2-LAMA3 (c, left panel), OT2-24 (c, right panel), and OT2-FMN1 (d), for T3 off-targets: OT3-COMDA (e, left panel), OT3-20 (e, right panel), OT3-MAX (f, left panel), and OT3-4 (f, right panel), and for EMX1 off-targets: OT4-HCN1 (g, upper panel), OT4-10 (g, middle panel), and OT4-MFAP1 (g, lower panel). OT sequences show mismatches in red, PAM in lowercase, and sites aligned to abasic extension (positions 21-22) underlined (c–g, upper panels); cleaved fragments (asterisks) in agarose gel electrophoresis or capillary electrophoresis (c–g, lower panels). Off-target sequences are shown with mismatches highlighted in red, PAM sequence in lower case (c–g, upper panels); cleaved fragments (*) in agarose gel electrophoresis or capillary electrophoresis (c–g, lower panels). Unless stated otherwise, P values are two-sided (t-test), n values denote biologically independent samples, and data represent mean ± s.d.
Extended Data Fig. 6 Genome-wide analyses of CRISPR-Cas9 off-targets reduced by ØXØ in T2 gRNA.
a, b, Preparation of T2 and EMX1 gRNA-delivered HEK293 cells for targeted genome sequencing to detect off-targets. After transfection with SpCas9-gRNA complex and the corresponding on-target surrogate reporter, cells were sorted by FACS: autofluorescence signals (Q3) were excluded (a, left graph); transfected cells (RFP+; Q1 and Q2 in a) were gated; CRISPR-active cells (RFP+/GFP+; P3) were sorted for T2 (gRNA and ØXØ; a, middle graphs) and EMX1 (gRNA and ØXØ; b, left graphs). On-target editing (%) of VEGFA T2 locus was validated by Surveyor nuclease assay (n = 1; a, b, right gel images); cleaved fragments (asterisk); cleavage (%). Sorted cells were used for targeted sequencing. c, Targeted sequencing for on-targets (n = 1 sequencing library) from transient delivery of unmodified gRNA, ØXØ, tru-gRNA (tru-) and ØØX20 into HEK293 cells. Indel frequencies (%; ± 30 bp from the cleavage site) are in the upper table; combined relative indel (%) normalized to unmodified gRNA (100%) is shown below. ØXØ showed similar on-target editing to ØØX20 and higher than tru-gRNA. d, e, OT2-24 and OT4-10 loci from gRNA-delivered HEK293 cells could not be analyzed due to multiple unreported substitution mutations (mutations in lowercases; d); these prevented primer design and efficient PCR amplification. Repeated attempts to design primers based on the reference human genome failed to efficiently amplify these loci by PCR, and the resulting amplicons showed an extremely low indel rate (e). f, Statistical analysis of CIRCLE-seq data (ØXØ and T2 gRNA at 50 nM; Supplementary Table 13) processed by CIRCLE-seq analytical software48. Confident CIRCLE-seq sites were stringently selected by read-count thresholds (right graph) determined statistically by comparing the distribution with cleavage regions having the highest mismatch scores (P < 0.1 relative to mismatch score = 6 as null hypothesis (n = 767), two-sided; left graph). g, h, CIRCLE-seq shows reduction of off-target sites with ØXØ versus unmodified gRNA (T2 at 10 nM or 50 nM) with different selection stringencies: all (no P value cutoff), P < 0.1, and P < 0.05, relative to mismatch score = 6. Venn diagrams display off-target overlaps for 50 nM (g, upper panels) and 10 nM (g, lower panels) datasets. Bar plots show ØXØ/gRNA ratios (%) for 50 nM (h, upper graph) and 10 nM (h, lower graph). i, CIRCLE-seq results from ØXØ and unmodified gRNA (T2 at 50 nM) for validated off-targets in targeted sequencing (Fig. 4g, h). Normalized cleavage (%) was calculated as the ratio of read counts (reads per million mapped reads; RPM) relative to the on-target site and further normalized to match indel ratio (%) from targeted sequencing (Fig. 4f). Unless stated otherwise, n values denote biologically independent samples.
Extended Data Fig. 7 CIRCLE-seq analyses for T3 and EMX1 gRNAs after ØXØ modification.
a, b, Comparative analyses of CIRCLE-seq off-target sites between unmodified and ØXØ gRNAs as in Extended Data Fig. 6g, h except for T3 (50 nM); Venn diagrams (a) and bar plots (b). c, CIRCLE-seq results from ØXØ and unmodified gRNA (T3 at 50 nM) for validated off-targets in targeted sequencing (Fig. 4g, i) as analyzed in Extended Data Fig. 6i. Results from CIRCLE-seq experiments with unmodified or ØXØ gRNA targeting VEGFA (T3 at 50 nM). Confident CIRCLE-seq sites (n = 140 sites; P < 0.01 relative to mismatch score = 6) are shown. Sites are displayed as absent (grey) or with reduced normalized cleavage rates (“norm”, %), based on normalized read counts (“reads”; reads per million mapped reads, RPM) relative to the on-target site, and scaled to match indel ratios (%) from targeted sequencing (Fig. 4f); the T3 on-target site is shown in red.
Extended Data Fig. 8 Comparative analyses of off-target sequences between gRNA and ØXØ using random mismatch libraries.
a, b, Sequencing strategy using a 1-mismatch (1MM) target library of T2 after reacting with T2 gRNA complexed with SpCas9 in vitro. The 1MM T2 library was generated by pooling 20 synthetic oligonucleotides, each carrying a different single random nucleotide position (from positions 1 to 20; a, upper panel). Substrate abundance (log2 ratio) was calculated by comparing normalized sequencing reads from SpCas9 treatment with gRNA (gRNA or ØXØ) to those without gRNA (cont; a, lower panels). Residual 1MM target libraries after in vitro SpCas9 reaction were subjected to sequencing (n = 1 sequencing library; b). c–g, Sequencing analysis of 1MM library (T2) after in vitro SpCas9 reaction with ØXØ versus unmodified gRNA. Composition of 1MM target libraries (pie charts; c), including on-target and spike-in control (non-cleavable sequence), after treatment with SpCas9 complexed with either unmodified gRNA or ØXØ (T2 at 50 nM; c, middle and right), relative to cont (without gRNA; c, left). Distribution (d) and scatter plots (e) of substrate abundance comparing gRNA (log2(gRNA/cont)) and ØXØ (log2(ØXØ/cont)); P value from Kolmogorov–Smirnov test (two-sided; d). 1MM sequences are ranked by depletion (mediated by SpCas9-induced cleavage), represented with ratios (relative abundance; %) normalized to control (gRNA/cont or ØXØ/cont; e); T2 on-target in red, off-targets (<6.5% relative abundance) in orange (e). ΔSubstrate abundance (log2(ØXØ/gRNA), difference of substrate abundance between ØXØ and unmodified gRNA) is ranked and shown as a heat map by mismatch position (f). Normalized substrate abundance, defined as the ratio of substrate abundance normalized as a percentage relative to the on-target (0%), was compared by position between T2 gRNA and ØXØ for 1MM sites (g). Of note, ØXØ showed reduced cleavage of 1MM sequences at PAM-distal regions, particularly near 5′ positions bearing the abasic modification, compared with unmodified gRNA. h, Construction of random mismatch libraries containing up to 3MM by pooling 20 different ‘doped’ synthetic oligonucleotides for T2 (left panel), T3 (middle panel), and EMX1 (right panel).
Extended Data Fig. 9 Analysis of ØXØ-induced positional intolerance and prevention of mismatched off-targets that cause DNA damage and cellular toxicity.
a–f, Positional intolerance of gRNA to mismatched off-targets assessed using random mismatch libraries (≤3MM); unmodified gRNA (upper graph) versus ØXØ (lower graph). Normalized substrate abundance, defined as the ratio of substrate abundance normalized as a percentage relative to the on-target (0%), was compared by position for 1MM off-targets: T2 (a), T3 (b), and EMX1 (c) and for 2MM (d–f, left graphs) or 3MM off-targets (d–f, right graphs): T2 (d), T3 (e) and EMX1 (f). Mismatch positions are shown as a cumulative sum of positional 1MM and also stratified by mismatch type: transitions (faint, dotted) and transversions (solid). g–i, Effects of ØXØ modification in VEGFA-targeting T2 gRNA on cellular toxicity, examined after transfection of SpCas9-gRNA complex (control, unmodified gRNA, and ØXØ). After initiating tube formation, the morphology of EA.hy926 (human endothelial cell line) was examined (g); whole-dish images (g, upper images) and zoomed views (g, lower images; biological replicates of Fig. 6b); scale bars, 400 μm. h, images of the transfected EA.hy926 cells; scale bars, 100 μm. i, Apoptotic cell death in MIHA cells after transfecting tru-gRNA and GGX20, analyzed by Annexin V and PI staining (cytometry, n = 5000 cells); quantitation of cell death (%; right graph) as categorized (same color display) in cytometry results (left panels); n = 3; *P = 9.2 x 10−4, 0.006, and 0.003, respectively; relative to control. Unless stated otherwise, P values are two-sided (t-test), n values denote biologically independent samples, and data represent mean ± s.d.
Extended Data Fig. 10 Effects of ØXØ modification on VEGFA-targeting T2 gRNA in vivo.
a, Examination of in vivo liver toxicity in mice by CRISPR-Cas9. SpCas9 expression plasmid (0.95 mg/kg) was delivered to mouse liver as polyethyleneimine (PEI) complex with different gRNA (2.5 mg/kg; control, T2 gRNA, or ØXØ; n = 5, 4, 3 independent mice) via tail-vein injection (I.V.; n = 2 injections; a, upper panel). Considering difference in the delivery, relative caspase 3/7 activity (%; a, lower graph) was quantified from replicate measurements (different liver tissue sections from the same mouse) normalized to SpCas9 mRNA levels (qPCR results in Fig. 6i); *P = 0.02 versus control. b, Images of mouse liver sections stained with H&E (upper images) and TUNEL (lower images); replicate of zoomed-in regions as indicated in Fig. 6k; scale bars, 100 μm. c, in vivo liver toxicity experiments in mice, as performed in (a) except using half the dose of SpCas9 plasmid (0.475 mg/kg) and gRNA (1.25 mg/kg) including tru-gRNA and GGX20 with increased numbers of I.V. injection (n = 3 injections; upper panel); relative caspase 3/7 activity (%) normalized to liver SpCas9 mRNA (lower graph); n = 3 (cont, tru-gRNA) or n = 4 (gRNA, ØXØ, and GGX20) independent mice; *P = 0.049 versus gRNA; n.s, not significant (P > 0.05). d, Images of T2 gRNA-, tru-gRNA-, and GGX20-delivered mouse liver sections stained with H&E (upper images) and TUNEL (lower images); scale bars, 100 μm. e, Delivery of T2 gRNA (control or ØXØ) with SpCas9 vector (n = 8 injections, I.V. from 9 M to 10 M) into DEN-treated mouse liver (intraperitoneal injection (i.p.) at postnatal day 14 (p14); upper panel). Considering difference in the delivery, Vegfa mRNA expression (ratio; lower graph) was quantified using qPCR relative to control, normalized to SpCas9 mRNA levels (qPCR results in Fig. 6n); n = 4 independent mice; *P = 0.025 versus control. f, Images of control- and T2 ØXØ-delivered mouse liver sections stained with H&E (left images), Ki67 (middle images) and Trichrome (right images); replicate results of Fig. 6p; scale bars, 100 μm. g, Depending on redox state (with or without NAC pre-treatment), growth curves of S. pyogenes SF370 were measured (OD600 nm) during Phage A1 infection; x-axis, time (hr) post infection; n = 3; *P = 0.018, 0.013, 0.002, 0.003 and 9.2 x 10−4 (infection); *P = 0.01, 5.4 x 10−4, 0.001, 4.4 x 10−4 and 3.0 x 10−4 (NAC); *P = 0.02, 5.3 x 10−5, 1.7 × 10−4, 4.3 x 10−4, 1.9 ×10−4, 6.8 x 10−5 (NAC + infection), relative to mock. h, Positional sequence analysis of crRNAs in S. pyogenes (n = 199 unique spacers) from the CRISPR-Cas + + database; positional base frequencies shown with WebLogo3. Unless stated otherwise, P values are two-sided (t-test), n values denote biologically independent samples, and data represent mean ± s.d.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–8.
Supplementary Tables (download XLSX )
Supplementary Tables 1–22.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 1 (download PDF )
Uncropped capillary electrophoresis images and unprocessed dot blots.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download PDF )
Unprocessed dot blots and uncropped capillary electrophoresis images.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download PDF )
Unprocessed gels.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Source Data Extended Data Figs. 1–10 (download PDF )
Uncropped capillary electrophoresis images and unprocessed gels and dot blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gu, D., Kim, GW.D., Park, M. et al. Abasic CRISPR RNAs inherently harness fidelity of SpCas9 for genome editing. Nat Chem Biol (2026). https://doi.org/10.1038/s41589-026-02139-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41589-026-02139-8