Abstract
Chronic stimulation in the tumor microenvironment can induce exhausted CD8+ T (Tex) cells that have limited tumoricidal activity. Here, we show an inverse correlation between signal strength and Tex cell differentiation by using patient-derived mutations in the T cell receptor (TCR)-signaling protein CARD11. Strong TCR signaling of the E134G mutant inhibits Tex cell differentiation and increases tumor growth. Conversely, reduced TCR signaling by the K215M mutant promotes Tex cell differentiation with better tumor control. These effects are a result of a restrained tumor-specific TCR clonal repertoire of Tex cells that reduces immunopathology but compromises tumoricidal activity. Mechanistically, CARD11 is a TCR signal-strength sensor, controlling the TCR repertoire of Tex cells by regulating the trafficking and homeostasis of the TCR complex. Expanding the TCR repertoire during Tex cell differentiation by fine-tuning the CARD11-mediated TCR signal strength reinvigorated antitumor function and indicates a strategy for improving cancer immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Sequencing data have been deposited in the NCBI database (ATAC-seq accession no. GSE292541, RNA-seq accession no. GSE292542, scRNA and scTCR-seq accession no. GSE292543). The human NSCLC dataset was accessed in the Gene Expression Omnibus database under accession code GSE179994. Other databases utilized in this study included the Molecular Signatures Database (https://www.gsea-msigdb.org/gsea/datasets.jsp), the GO website (http://www.geneontology.org/) and KEGG (https://www.genome.jp/kegg/). Source data are provided with this paper.
References
Hashimoto, M. et al. CD8 T cell exhaustion in chronic infection and cancer: opportunities for interventions. Annu. Rev. Med. 69, 301–318 (2018).
Giles, J. R., Globig, A.-M., Kaech, S. M. & Wherry, E. J. CD8+ T cells in the cancer-immunity cycle. Immunity 56, 2231–2253 (2023).
McLane, L. M., Abdel-Hakeem, M. S. & Wherry, E. J. CD8 T cell exhaustion during chronic viral infection and cancer. Annu. Rev. Immunol. 37, 457–495 (2019).
Li, H. et al. Dysfunctional CD8 T cells form a proliferative, dynamically regulated compartment within human melanoma. Cell 176, 775–789.e18 (2019).
Philip, M. & Schietinger, A. CD8+ T cell differentiation and dysfunction in cancer. Nat. Rev. Immunol. 22, 209–223 (2022).
ElTanbouly, M. A. & Noelle, R. J. Rethinking peripheral T cell tolerance: checkpoints across a T cell’s journey. Nat. Rev. Immunol. 21, 257–267 (2021).
Liu, Y. & Zheng, P. Preserving the CTLA-4 checkpoint for safer and more effective cancer immunotherapy. Trends Pharmacol. Sci. 41, 4–12 (2020).
Wu, J. E. et al. In vitro modeling of CD8+ T cell exhaustion enables CRISPR screening to reveal a role for BHLHE40. Sci. Immunol. 8, eade3369 (2023).
Thome, M., Charton, J. E., Pelzer, C. & Hailfinger, S. Antigen receptor signaling to NF-κB via CARMA1, BCL10, and MALT1. Cold Spring Harb. Perspect. Biol. 2, a003004 (2010).
Jun, J. E. & Goodnow, C. C. Scaffolding of antigen receptors for immunogenic versus tolerogenic signaling. Nat. Immunol. 4, 1057–1064 (2003).
Snow, A. L. et al. Congenital B cell lymphocytosis explained by novel germline CARD11 mutations. J. Exp. Med. 209, 2247–2261 (2012).
Lenz, G. et al. Oncogenic CARD11 mutations in human diffuse large B cell lymphoma. Science 319, 1676–1679 (2008).
Wei, Z. et al. Pathogenic CARD11 mutations affect B cell development and differentiation through a noncanonical pathway. Sci. Immunol. 4, eaaw5618 (2019).
Leong, Y. A. et al. CXCR5(+) follicular cytotoxic T cells control viral infection in B cell follicles. Nat. Immunol. 17, 1187–1196 (2016).
He, R. et al. Follicular CXCR5-expressing CD8(+) T cells curtail chronic viral infection. Nature 537, 412–428 (2016).
Wu, T. et al. The TCF1–Bcl6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness. Sci. Immunol. 1, eaai8593 (2016).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8(+) T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Thommen, D. S. et al. A transcriptionally and functionally distinct PD-1+CD8+ T cell pool with predictive potential in non-small-cell lung cancer treated with PD-1 blockade. Nat. Med. 24, 994–1004 (2018).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Beltra, J.-C. et al. Developmental relationships of four exhausted CD8+ T cell subsets reveals underlying transcriptional and epigenetic landscape control mechanisms. Immunity 52, 825–841.e8 (2020).
Duhen, T. et al. Co-expression of CD39 and CD103 identifies tumor-reactive CD8 T cells in human solid tumors. Nat. Commun. 9, 2724 (2018).
Oliveira, G. et al. Phenotype, specificity and avidity of antitumour CD8+ T cells in melanoma. Nature 596, 119–125 (2021).
Liu, B. et al. Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat. Cancer 3, 108–121 (2022).
Zhang, L. et al. Lineage tracking reveals dynamic relationships of T cells in colorectal cancer. Nature 564, 268–272 (2018).
Gaide, O. et al. CARMA1 is a critical lipid raft-associated regulator of TCR-induced NF-κB activation. Nat. Immunol. 3, 836–843 (2002).
Lamason, R. L., Kupfer, A. & Pomerantz, J. L. The dynamic distribution of CARD11 at the immunological synapse is regulated by the inhibitory kinesin GAKIN. Mol. Cell 40, 798–809 (2010).
Luton, F., Legendre, V., Gorvel, J. P., Schmitt-Verhulst, A. M. & Boyer, C. Tyrosine and serine protein kinase activities associated with ligand-induced internalized TCR/CD3 complexes. J. Immunol. 158, 3140–3147 (1997).
Evnouchidou, I. et al. IRAP-dependent endosomal T cell receptor signalling is essential for T cell responses. Nat. Commun. 11, 2779 (2020).
Phelan, J. D. et al. A multiprotein supercomplex controlling oncogenic signalling in lymphoma. Nature 560, 387–391 (2018).
Evnouchidou, I., Caillens, V., Koumantou, D. & Saveanu, L. The role of endocytic trafficking in antigen T cell receptor activation. Biomed. J. 45, 310–320 (2022).
Ouchida, R. et al. A lysosomal protein negatively regulates surface T cell antigen receptor expression by promoting CD3ζ-chain degradation. Immunity 29, 33–43 (2008).
Chiffelle, J. et al. T-cell repertoire analysis and metrics of diversity and clonality. Curr. Opin. Biotechnol. 65, 284–295 (2020).
Altman, J. D. et al. Phenotypic analysis of antigen-specific T lymphocytes. Science 274, 94–96 (1996).
Simoni, Y. et al. Bystander CD8+ T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557, 575–579 (2018).
Kataoka, T. et al. Concanamycin A, a powerful tool for characterization and estimation of contribution of perforin- and Fas-based lytic pathways in cell-mediated cytotoxicity. J. Immunol. 156, 3678–3686 (1996).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Westcott, P. M. K. et al. Mismatch repair deficiency is not sufficient to elicit tumor immunogenicity. Nat. Genet. https://doi.org/10.1038/s41588-023-01499-4 (2023).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med. 25, 1251–1259 (2019).
Han, J. et al. TCR repertoire diversity of peripheral PD-1+CD8+ T cells predicts clinical outcomes after immunotherapy in patients with non-small cell lung cancer. Cancer Immunol. Res. 8, 146–154 (2020).
Piehl, N. et al. Cerebrospinal fluid immune dysregulation during healthy brain aging and cognitive impairment. Cell 185, 5028–5039.e13 (2022).
Shakiba, M. et al. TCR signal strength defines distinct mechanisms of T cell dysfunction and cancer evasion. J. Exp. Med. 219, e20201966 (2022).
Singhaviranon, S., Dempsey, J. P., Hagymasi, A. T., Mandoiu, I. I. & Srivastava, P. K. Low-avidity T cells drive endogenous tumor immunity in mice and humans. Nat. Immunol. 26, 240–251 (2025).
Saeidi, A. et al. T-cell exhaustion in chronic infections: reversing the state of exhaustion and reinvigorating optimal protective immune responses. Front. Immunol. 9, 2569 (2018).
Jhunjhunwala, S., Hammer, C. & Delamarre, L. Antigen presentation in cancer: insights into tumour immunogenicity and immune evasion. Nat. Rev. Cancer 21, 298–312 (2021).
Yang, K., Halima, A. & Chan, T. A. Antigen presentation in cancer — mechanisms and clinical implications for immunotherapy. Nat. Rev. Clin. Oncol. 20, 604–623 (2023).
Garcia, J. et al. Naturally occurring T cell mutations enhance engineered T cell therapies. Nature 626, 626–634 (2024).
Acknowledgements
We acknowledge L. Ye and Y. Liu for their ideas and suggestions about this project. We thank all the staff in the Institutional Center for Shared Technologies and Facilities of SINH, CAS, for technical assistance. This study is supported by research funding from the National Natural Science Foundation of China (nos. 82050001, 82330051 and 92478203 to W.L.; nos. 82071866, 82271762 and 82241227 to Y. Zhang; and nos. 82072557 and 82372855 to H.L.) and the National Key Research and Development Project of MOST, China (2023YFC2411403 to Y. Zhang and 2021YFC2500900 to H.L.).
Author information
Authors and Affiliations
Contributions
Y.H., Q.Z., Y.Q., L.H., B.W., H.Z., L.Y., Z.H. and Y.L. performed the experiments. Y.H. Q.Z., Y.Q., S.M. and Y. Zang performed the data and statistical analysis. H.L., Y. Zhang, P.Z., M.J.L. and W.L. designed the study; and M.J.L. and W.L. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Nick Bernard, in collaboration with the rest of the Nature Immunology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CARD11 mutations show distinct impact on CD8+ TILs.
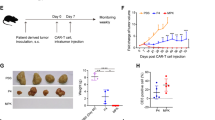
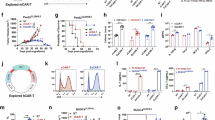
(a) Images of the B16-F10 tumor and (b) the MC-38 tumor in WT and Card11 mutant mice, 16 days after incubation. (c) The proportion of B16-F10 CD45+ TILs in WT (n = 28), E134G (n = 23) and K215M mice (n = 20) and (d) MC-38 CD45+ TILs in WT (n = 12), E134G (n = 11) and K215M mice (n = 13) were analyzed by flow cytometry (n > =11). (e) UMAP visualization of scRNA-seq data from B16-F10 CD45+ TILs. The clusters are identified by the expression of the maker genes. (f) Dot plots showing the expression of representative signature genes from selected gene sets across clusters of WT CD45+ TILs, based on scRNA-seq analysis. The color scale indicates the expression score. Circle size indicates the percentage of cells that express that gene. (g) Pie charts showing the composition of immune cells in WT, E134G, and K215M CD45+ TILs. (h) UMAP visualization of scRNA-seq data from B16-F10 CD8+ TILs. The clusters are identified by unsupervised clustering. (i) The relative proportion of CD8+ TIL subsets in WT, E134G, and K215M mutant mice was indicated by different color bands based on scRNA-seq. Four subsets contain distinct clusters identified in Fig. 1c. (j) Visualization of Pdcd1, Havcr2, Tcf7, and Tox expression on CD8+ TILs based on scRNA-seq. (k) PD-1 and TIM3 expression on CD8+ TILs in WT and Card11 KO mice were analyzed by flow cytometry. (l) PD-1 and TCF1 expression on CD8+ TILs in WT and Card11 KO mice were analyzed by flow cytometry. (m) The frequency of Tpre, Tpex, and Tex populations in WT and Card11 KO mice were analyzed by flow cytometry (n = 5). (n) The frequency of PD-1low, PD-1med, and PD-1high CD45.2+ CD8+ TILs in BM chimeras of WT (n = 6) /E134G (n = 9)/ K215M mice (n = 10). (o) The frequency quantification of tumor-infiltrating PD-1low, PD-1med, and PD-1high CD45.2+ OT-I cells from WT (n = 7) /E134G (n = 4)/ K215M mice (n = 7). in co-transfer models. (p) Heatmap of peak intensity for all differentially accessible regions in WT Tpre, WT Tex and K215M Tex samples (q) Diagram of the gene targeting strategy of conditional knock-in of Card11-K215M mutation mice. (r) The B16-F10 tumor image of E8I-Cre, K215Mf/+ and K215Mf/+ E8I-Cre mice 16 days after incubation. Data represent mean ± SEM of n > 3 biological replicates. P values were determined using unpaired two-tailed Student’s t-tests.
Extended Data Fig. 2 CARD11 mutants affect the differential trajectory of CD8+ TILs.
(a) Four subsets of CD8+ TILs sorted from B16-F10 tumor (cell surface SLAMF6 substitutes intracellular TCF1) in WT were i.v. injected into Rag2−/− recipient mice bearing B16-F10 tumors, respectively. 10 days later, the TCF1 and PD-1 expression on CD8+ TILs from donor were analyzed by flow cytometry. (b) Dot plot showing the expression of typical marker genes from selected gene sets (colored rectangle gate) across four subsets of WT CD8+ TILs, based on scRNA-seq transcriptome analysis. The color scale indicates the expression score. Circle size indicates the percentage of cells that express the gene. (c-f) The frequency of PRF1 (n = 7) (c), GzmB (n = 7) (d), Annexin V (n = 10) (e), and Ki-67 (n = 11) (f) in different subsets. (g) UMAP visualization of scRNA-seq data from human NSCLC patient-derived CD8+ T cells (GSE179994). The Tmem subset represents the memory CD8+ TILs, which is absent in murine tumor models. The clusters identified by the expression of maker genes as aforementioned (left). Monocle-3 trajectory analysis of CD8+ TILs’ differentiation. Cells are colored according to their corresponding pseudotime (right). Data represent mean ± SEM of n > 3 biological replicates. P values were determined using unpaired two-tailed Student’s t-tests.
Extended Data Fig. 3 TCR signaling is differentially influenced in Card11 mutant mice.
(a-b) Flow cytometry analysis of CD69 and CD25 expression on soluble (a) and plate-coated (b) anti-CD3/28 antibodies-activated WT and Card11 mutant CD8+ T cells after 24 h (n = 3). (c) CD44 and CD62L expression on WT and Card11 mutant splenic CD8+ T cells were assessed by flow cytometry. (d) CD44 and PD-1 expression on WT and Card11 mutant splenic CD8+ T cells were also evaluated (representative of 3 independent experiments for (c) and (d)). (e) Apoptotic cell proportions in WT and Card11 mutant CD8+ T cells activated by soluble antibodies for 24 h were measured by flow cytometry. Quantification was on the right (n = 3). (f) Apoptotic cell proportions in WT and Card11 mutant CD8+ T cells activated by plate-bound antibodies for 24 h were measured by flow cytometry. Quantification was shown on the right (n = 5). (g) Ki-67+ cell proportions in WT and Card11 mutant CD8+ T cells activated by soluble antibodies for 24 h were measured by flow cytometry. Quantification was shown on the right (n = 3). (h) Cell proliferations of WT and Card11 mutant CD8+ T cells activated by plate-bound antibodies were measured by CFSE staining at indicated time points. (i) Diagram of the gene targeting strategy of conditional knock-in of Card11-E134G mutation mice. (j-k) Flow cytometry analysis of CD69 and CD25 expression on soluble (n = 4) (j) and plate-coated (k) anti-CD3/28 antibodies-activated E134Gflox/+ and E8ICre E134Gflox/flox CD8+ T cells after 24 h (n = 3). (l-m) Flow cytometry analysis of CD69 and CD25 expression on soluble (n = 4) (l) and plate-coated (n = 3) (m) anti-CD3/28 antibodies-activated K215Mflox/+ and E8ICre K215Mflox/flox CD8+ T cells after 24 h (representative of 3 independent experiments). (n) Monocle-2 trajectory analysis of CD8+ TILs differentiation revealing divergent trajectories in WT CD8+ TILs. (o) The repeat experiment of Fig. 3m. Subset Ttr-1 (PD-1med SLAMF6− KLRC1+) and Ttr-2 (PD-1med SLAMF6− KLRC1−) from WT CD8+ TILs were purified and reinjected into tumor-bearing Rag2−/− recipient mice, respectively. 10 days later, PD-1 and TCF1 expression on donor cells in the tumor and naïve CD8+ T cells were analyzed. The dot plots are shown. (p) The repeat experiment of Fig. 3n. Subset Ttrans from E134G and K215M mutant CD8+ TILs were injected into tumor-bearing Rag2−/− recipient mice, respectively. PD-1 and TCF1 expressions on donor cells in the tumor were analyzed after 10 days. (q) Clonotype tracing analysis of CD8 TCR in the indicated cluster. (r) The scheme of the TCR signal strength-dependent differentiation model of Tex in WT mice. Data represent mean ± SEM of n > 3 biological replicates. P values were determined using unpaired two-tailed Student’s t-tests.
Extended Data Fig. 4 Intracellular translocation of CARD11 after TCR stimulation.
(a) Diagram of the gene targeting strategy of Card11-Flag knock-in mice. (b) Naïve CD8+ T cells from Card11-Flag knock-in mice were untreated or treated with anti-CD3/28 agonist antibodies for 30 min. The localization of CARD11–FLAG (Green) on the cell plasma membrane (DiD far-red Dye) was detected by confocal. (c) The quantification of colocalization efficiency in (b) (n = 13) is shown. (d) CD8+ T cells were stimulated with increasing concentration of agonist antibodies. The images showing localization of CARD11–FLAG (green) and Rab7 (red). DAPI staining (blue) outlined by white dots. (e) The diagram showing the sediment saved as the pellet and the upper phase saved as the endo-lysosome in a centrifuge tube (top). Immunoblots assessed proteins in the late endo-lysosome fractions before and after one hour of anti-CD3/CD28 stimulation (bottom). (f) The diagram showing full-length CARD11 with each specific domain indicated. (g) Immunofluorescence images showing CARD11 (green) localization and Rab7 (red) in activated CD8+ T cells from WT and CARD11 mutant mice. (h) Immunofluorescence images showing localization of CARD11 (green) and Rab11 (red) in activated CD8+ T cells from WT and CARD11 mutant mice. Data represent mean ± SEM of n > 3 biological replicates. P values were determined using unpaired two-tailed Student’s t-tests.
Extended Data Fig. 5 CARD11 regulates the homeostasis of the TCR complex.
(a) Immunofluorescence images showing localization of TCRβ (red) and Rab5 (n = 19), Rab7 (n = 20), and Rab11 (n = 17), (green) in WT naïve CD8+ T cells (left). Quantification of the colocalization efficiency was shown (right). (b) Co-IP experiments using CARD11 antibody in WT CD8+ T cells before (-) and after (+) anti-CD3/CD28 antibody stimulation. (c) Immunofluorescence images of naïve CD8+ T cells showing colocalization of TCRβ (red) and LAMP1 (green) in WT and Card11 KO mice. (d) Quantification of the colocalization efficiency between TCRβ and Rab11 in WT CD8+ T cells before (-) and after (+) anti-CD3/CD28 stimulation for one hour (n = 30). (e) Quantification of the colocalization efficiency between TCRβ and LAMP1 in WT CD8+ T cells before (-) and after (+) anti-CD3/CD28 stimulation for one hour (n = 10). (f) WT naïve CD8+ T cells were stimulated with a gradient concentration of anti-CD3/CD28 antibodies. Flow cytometry evaluated the total TCRβ level (n = 3). (g) The total TCRβ level evaluated in (f) was quantified (n = 3). (h) WT CD8+ T cells were untreated or treated with anti-CD3/CD28 antibodies. TCR-stimulated cells were further treated with or without BafA1. Then, the total TCRβ level was evaluated by flow cytometry (n = 3). (i) WT and Card11 mutant CD8+ T cells were stimulated with anti-CD3/CD28 antibody for the indicated time. The cell surface TCRβ level was measured by flow cytometry, and the MFI of 0 min in each group was set as standard (n = 3). (j) WT, Card11 E134G mutant and KO CD8+ T cells were treated with increasing concentrations of lysosome inhibitor BafA1 for 5 h and then stimulated with anti-CD3/28 stimulation for 12 h. CD69 and CD25 expression level was measured by flow cytometry (n = 3). (k-l) qPCR analysis of mRNA level of CD25 (n = 3) (k), and CD69 (n = 3) (l) expression in the WT and Card11 E134G mutant CD8+ T cells treated with BafA1 for 18 h. (m) Quantification of Laptm5 expression by qPCR in CD8+ T cells after transduced with EV and Laptm5 knockdown shRNA (n = 8). (n) Quantification of the normalized TCRβ level by flow cytometry on CD8+ T cells transduced with EV and Laptm5 knockdown shRNA (n = 4). Data represent mean ± SEM of n > 3 biological replicates. P values were determined using unpaired two-tailed Student’s t-tests.
Extended Data Fig. 6 Tex exhibits potent tumor-specific cytotoxic activity.
(a) Body weights of the Rag2-/- mice transferred with PBS (control), PD-1low CD8+ Tpre population cells (n = 3). (b) Sorted PD-1high OT-I T cells from B16-OVA tumors grown in WT mice were incubated with original B16-OVA cells with or without Concanamycin A in vitro. The killing efficacy was evaluated (n = 6). (c) Dot plot showing the expression of typical marker genes from selected gene sets across five subsets of CD8+ TILs in human NSCLC, based on scRNA-seq transcriptome analysis (d) Sorted PD-1low, PD-1med, and PD-1high TIM3+ CD8+ TILs from three NSCLC patients were incubated with sorted tumor cells in vitro. The killing efficacy was evaluated by the flow cytometry of the remaining live tumor cells. Three independent experiments based on 3 biopsies. Tumor cells alone served as NC. Data represent mean ± SEM of n > 3 biological replicates. P values were determined using unpaired two-tailed Student’s t-tests.
Extended Data Fig. 7 The TCR diversity of Tex influences the antitumor efficacy.
(a) The differentially expressed genes of CD8+ TIL cluster between WT and K215M mutant mice was indicated by different color bands based on scRNA-seq. Top 10 gene were shown. (b) Flow cytometry analysis of PD-1 and OVA Tetramer expression on CD8+ TILs in WT and Card11 mutant mice with B16-OVA tumors (left). The quantification of OVA Tetramer+ cells in WT and K215M mutant mice (n = 4) (right). (c) Donut plots show the clone frequency of Tex of CD8+ TILs from B16-OVA tumors. Each color in the inner ring represents a clone except the non-expanded clones (too many clones to split). Each color in the outer ring represents clones with the same amplification level. The number indicates the percentage of the most hyperexpanded clone in total Tex and the CDR3 sequences of this top clone were shown below. (d) The scheme of the TCR signal strength-dependent clone selection in tumors. Data represent mean ± SEM of n > 3 biological replicates. P values were determined using unpaired two-tailed Student’s t-tests.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2 and Fig. 1.
Supplementary Table 3 (download XLSX )
Reagents information in the study.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Unprocessed immunoblots and statistical source data.
Source Data Fig. 4 (download XLSX )
Unprocessed immunoblots and statistical source data.
Source Data Fig. 5 (download XLSX )
Unprocessed immunoblots and statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Unprocessed immunoblots and statistical source data
Source Data Extended Data Fig. 5 (download XLSX )
Unprocessed immunoblots and statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, Y., Zhao, Q., Qin, Y. et al. CARD11 signaling regulates CD8+ T cell tumoricidal function. Nat Immunol 26, 1113–1126 (2025). https://doi.org/10.1038/s41590-025-02192-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02192-w