Abstract
Stem-like T cells promote the efficacy of immunotherapy and are heterogeneous in stemness, with long-term (LT) stem-like T cells at the apex of this hierarchy. How the stemness hierarchy is regulated in chimeric antigen receptor (CAR) T cells and how it affects antitumor function are unclear. Here we show that BACH2 dose-dependently regulates LT stem-like differentiation and antitumor immunity of CAR T cells. LT stem-like CAR T cells that appear before infusion and re-emerge after tumor clearance have superior antitumor immunity and the greatest BACH2 expression. BACH2 promotes the antitumor response of CAR T cells and the LT stem-like transcriptional program. Temporal and quantitative induction of BACH2 expression in CAR T cells during manufacturing using chemical switches fine-tunes the degree of stemness and imprints greater control of solid tumors. Together, these data show that BACH2 dosage defines stemness hierarchy in CAR T cells and can be temporally and tunably controlled to optimize differentiation and antitumor efficacy.
Similar content being viewed by others
Main
Chimeric antigen receptor (CAR) T cells have been remarkably effective in treating several types of hematological malignancies1. However, only a subpopulation of individuals treated for cancer experience durable remission2,3. CAR T cell therapies have achieved limited successes in treating solid tumors4. Increasing evidence suggests that T cell-intrinsic dysfunction, also termed exhaustion, may contribute to antigen-positive relapse after CAR T cell therapy5. T cell exhaustion is induced by persistent stimulation by tumor antigen and tonic CAR signaling6,7,8,9,10,11. Exhaustion, characterized by the upregulation of immune checkpoints, including PD-1, reduces proliferation and effector function, dysregulates metabolism and impairs persistence, limiting the therapeutic potential of CAR T cells6,7,10,12. Thus, to improve the efficacy of CAR T cell therapies, it is critical to identify the molecular circuit that overcomes T cell exhaustion.
Exhausted T cells are a heterogeneous population consisting of TCF1+ stem-like T cells, also called progenitor exhausted T (Tpex) cells13,14,15,16,17,18,19,20,21,22,23,24,25,26. Stem-like T cells persist via self-renewal and replenish TCF1− terminally differentiated T cells13,14,15,16,17,18,19,20,21,22,23,24,25,26. Stem-like T cells exhibit superior proliferative capacity and mitochondrial fitness and are critical for the efficacy of immune checkpoint inhibitors (ICIs) and CAR T cells9,10,11,23,24,25,26,27,28,29. Differentiation of stem-like T cells is controlled by transcription factors, including TCF1, TOX, BACH2, FOXP1, MYB, FOXO1, KLF2 and NRF2, during cancer and infection13,14,17,18,19,20,21,26,30,31,32,33,34,35,36,37,38,39,40,41. Stem-like T cells are hierarchically organized with various degrees of stemness36,42,43,44. At the apex of this system is long-term (LT) stem-like T cells. Distinct from other stem-like lineages, LT stem-like T cells are protected from the epigenetic scar of exhaustion and preserve greater potential of LT renewal and proliferative burst in response to ICIs36,42,43. However, the molecular mechanism that establishes the hierarchy of stemness in T cells, especially in CAR T cells, is not completely understood.
Various methods have been developed to induce stem-like properties in CAR T cells, including overexpressing pro-stem transcription factors or naturally occurring oncogenes6,34,35,45. Yet, the constitutive overexpression approach may limit the adaptability and plasticity of T cells in response to different CAR signaling and environmental cues and risk transformation of CAR T cells. Thus, there is a need to improve CAR T cell efficacy by achieving tunable and temporal control of the stem-like program by CAR T cells.
Here, we show that BACH2 activity in preinfusion CAR T cells was highest among LT stem-like T cells, which were associated with superior antitumor responses in mice and humans. An LT stem-like CAR T cell subset developed after tumor clearance, upregulated the BACH2 signature and downregulated TOX expression. BACH2 drove expansion of CAR T cells in vivo and the LT stem-like T cell program. Through tunable control of BACH2 protein levels in GD2 CAR T cells with tonic signaling-driven exhaustion, we showed that the degree of stemness and exhaustion in CAR T cells is controlled by the level of BACH2. By chemically inducing BACH2 activity during manufacturing of GD2 CAR T cells, we showed that temporal BACH2 activation before infusion is sufficient to imprint greater immunity against solid tumors. In summary, our study demonstrates that BACH2 dosage establishes the hierarchy of CAR T cell stemness and can be temporally and tunably controlled to optimize the differentiation and antitumor immunity of CAR T cells.
Results
Preinfusion LT stem-like CAR T cells epigenetically activate BACH2
We sought to understand how preinfusion CAR T cells are programmed at transcriptional and epigenetic levels using joint single-cell assay for transposase accessible chromatin with sequencing and RNA sequencing (scATAC-seq + scRNA-seq). Mouse CD19-specific CAR T cells were generated from T cells of C57BL/6 mice as previously described16,46,47. CD8+ T cells sorted from preinfusion CD19 CAR T cells were used for scATAC-seq + scRNA-seq (Extended Data Fig. 1a). Single-cell transcriptomes and chromatin accessibility profiles of 4,860 cells were used for analysis. An average of 2,003 genes and 9,609 open chromatin regions (peaks) per cell were detected.
Unsupervised clustering partitioned preinfusion CD8+ CD19 CAR T cells into four clusters based on transcriptome (Fig. 1a). Cluster 0 showed a differentiated phenotype and expressed genes associated with cytotoxicity (Gzma and Nkg7), proinflammatory chemokine signaling (Ccr2, Ccr5 and Cxcr3) and transcriptional regulators of effector differentiation (Id2, Zeb2 and Tbx21; Fig. 1b,c and Extended Data Fig. 1b). Top marker genes of cluster 1 included those encoding transcriptional regulators such as Bach2, Foxp1, Lef1 and Foxo1 that drive differentiation of stem-like T cells15,16,34,35,48,49 (Fig. 1b,c and Extended Data Fig. 1b). The stem-like CD8+ subset in preinfusion CAR T cells also upregulated the expression of Sell and Ccr7 (Fig. 1c and Extended Data Fig. 1b). In addition, two proliferating clusters were found in preinfusion CD8+ CAR T cells (Fig. 1a–c and Extended Data Fig. 1c).
a–i, In vitro-expanded anti-mouse CD19 CAR T cells derived from C57BL/6 mice were analyzed with scATAC-seq + scRNA-seq. a, CD8+ CAR T cells displayed in a transcriptome-based UMAP plot (RNA UMAP). Effector-like (RNA 0 Teff-like), stem-like (RNA 1 Tstem-like) and proliferating (RNA 2 and RNA 3) clusters are defined by single-cell transcriptomes and color coded accordingly. b, Heat map of the top marker genes in each transcriptome-defined cluster. Columns represent individual cells. Rows represent genes. c, Single-cell expression of Bach2, Sell, Id2 and Mki67 shown in a UMAP plot. Color reflects the mRNA level of each gene. d, CD8+ CAR T cells projected on UMAP plots based on scATAC-seq (left, ATAC UMAP) and scRNA-seq profiles (right, RNA UMAP). Cells are color coded based on cluster IDs defined by the scATAC-seq profile, including transitory stem-like (ATAC 0 Ttrans), effector-like (ATAC 1 Teff-like), LT stem-like (ATAC 2 LT-Tstem-like) and proliferating (ATAC 3) clusters. e, Heat map of epigenetically activated genes in each scATAC-seq-defined cluster. The color code is defined by scATAC-seq reads in the promoter and gene body. Clusters are represented by columns and organized based on hierarchical clustering. f, Volcano plot showing genes with differential chromatin accessibility between transitory stem-like and LT stem-like subsets. Adjusted P values (Padj) were calculated using a two-sided Wilcoxon rank-sum test with Bonferroni correction to account for multiple comparisons. g, GSEA showing increased chromatin accessibility at published LT stem-like signature genes (GSE199839) in cluster ATAC 2 compared to cluster ATAC 0; NES, normalized enrichment score. The adjusted P value was calculated using the Benjamini–Hochberg procedure to correct for multiple testing. The red dashed line in the GSEA plot indicates the location of the maximum running enrichment score (ES). h, Heat map highlighting the differentially enriched transcription factor motifs in open chromatin regions across the clusters defined by the scATAC-seq profile shown in d. Color represents the degree of motif enrichment. i, Feature plots depicting the single-cell enrichment of BACH2, LEF1, TCF1 and TBX21 (T-bet) binding motifs in the open chromatin regions of CD8+ CAR T cells. Color intensity indicates the degree of motif enrichment.
Based on unsupervised clustering of scATAC-seq profiles, preinfusion CD8+ CD19 CAR T cells were segregated into four clusters and projected into either an ATAC-defined uniform manifold approximation and projection (UMAP) or RNA-defined UMAP (Fig. 1d). The stem-like CD8+ subset was further segregated into two subsets based on chromatin accessibility (Fig. 1d). One of these subsets displayed a transitory chromatin state between stem-like and effector-like subsets (Fig. 1e,f). The open chromatin profiles of the other stem-like subset showed epigenetic activation of the gene signature of LT stem-like T cells36,44, including Bach2, Sell and Lef1, and enhanced enrichment of the epigenetic signature associated with a previously defined long-lived stem-like T cell population generated through Asxl1 knockout (KO50; Fig. 1f,g and Extended Data Fig. 1d). Compared to the transitory stem-like cells, LT stem-like CAR T cells showed lower chromatin accessibility in genes associated with effector/exhaustion differentiation, such as Zeb2 and Havcr2 (Fig. 1e,f). Transcription factor motif analysis revealed higher activity of T-box transcription factors such as T-bet (TBX21) in the transitory subset than in the LT stem-like subset (Fig. 1h). In addition to enrichment of TCF1 and LEF1 motifs, LT stem-like CAR T cells showed higher activity of the transcriptional repressor BACH2, indicated by lower chromatin accessibility at BACH2 targets (Fig. 1h,i). Together, these data show that a subset of preinfusion CD8+ CAR T cells shows a chromatin state of LT stem-like T cells and epigenetic activation of the pro-stem transcription factor BACH2.
Preinfusion LT stem-like CAR T cells have greater tumor control
Expression of Sell was highest in the LT stem-like subset of preinfusion CD8+ CAR T cells (Extended Data Fig. 2a). The CD62L+ LT stem-like subset and CD62L− subset were sorted from preinfusion CD8+ CD19 CAR T cells and transferred separately into mice bearing E2A–PBX1 pre-B cell acute lymphoblastic leukemia (ALL), as previously described16. Seven days after infusion, we analyzed progenies of CD62L+ LT stem-like CAR T cells and their CD62L− counterparts (Extended Data Fig. 2b). Notably, the expansion of LT stem-like CAR T cells after adoptive transfer was more than twofold higher than that of CD62L− cells in the spleen, bone marrow, liver and lung (Fig. 2a and Extended Data Fig. 2c). In addition, the percentage of CD62L+CX3CR1− stem-like cells was higher among progenies of LT stem-like CAR T cells than among progenies of CD62L− cells (Fig. 2b). Progenies of LT stem-like CAR T cells were more resistant to apoptosis, expressed lower levels of immune checkpoints, including PD-1 and TIM3, and showed decreased expression of TOX compared to progenies derived from CD62L− cells (Extended Data Fig. 2d,e). After infusion in mice bearing 9464D-GD2 neuroblastoma, LT stem-like GD2-specific (14g2a-E101K-28z)6 CAR T cells exhibited enhanced tumor control (Extended Data Fig. 2f). Together, these results suggest that LT stem-like CAR T cells in the infusion product resist apoptosis and exhaustion and show enhanced stemness, expansion and tumor control after treating tumor-bearing mice.
a,b, Mouse CD8+ CD19 CAR T cells coexpressing truncated human epidermal growth factor receptor (huEGFR) as a marker were sorted into CD62L+ and CD62L− fractions, which were then transferred into C57BL/6 mice bearing E2A–PBX1 B cell ALL (n = 3 mice per group). On day 7 after transfer, flow cytometry was performed (left), and statistics (right) of the number of CAR T cells (a) and percentage of CD62L+CX3CR1− CAR T cells (b) derived from CD62L+ or CD62L− preinfusion CAR T cells were determined. c–h, Analysis of published scRNA-seq data (GSE241783) of CD8+ preinfusion human CD19 CAR (axicabtagene ciloleucel) T cells from 40 individuals with relapsed/refractory B cell lymphoma. c, UMAP plot of CD8+ human preinfusion CD19 CAR T cells. Cells are color coded based on cluster IDs, including proliferating T cell clusters, effector-like T (Teff-like) cell cluster, type 2 T cell cluster, stem-like T (Tstem-like) cell cluster, IL-19+ T cell cluster, exhausted-like T (Tex-like) cell cluster, γδ T cell cluster, IL-17F+ T cell cluster and immune effector cell-associated neurotoxicity syndrome-associated T cell (IAC) cluster. d, Heat map of differentially expressed genes in each cluster defined in c. e, Single-cell expression of BACH2, TCF7, GATA3 and MKI67 in preinfusion human CAR T cells. f, Enrichment of the LT stem-like signature (GSE199839) in each cluster defined in c. g, Volcano plot of differentially expressed genes between CAR T cells from individuals showing a CR and those who did not (non-CR). h, GSEA showing enrichment of the LT stem-like signature (GSE199839) in CR versus non-CR CAR T cells. The red dashed line in the GSEA plot indicates the location of the maximum running enrichment score (ES). Data in a and b are representative of two independent experiments. Bar graphs represent the mean ± s.e.m. Circles in the bar graphs represent individual mice. Statistical significance was calculated with a two-sided Student’s t-test; FDR, false discovery rate.
Human preinfusion CAR T cells of complete responders upregulate BACH2
To understand the transcriptional signature correlating with the response to CAR T cell therapy in individuals with cancer, we used published single-cell RNA-sequencing (scRNA-seq) data of infusion products from 40 individuals with relapsed/refractory B cell lymphoma treated with CD19 CAR T cells (axicabtagene ciloleucel)51. Fifteen individuals achieved complete response (CR), whereas 20 individuals had progressive disease or a partial response (non-CR). A total of 309,392 cells passed quality control, with an average of 6,950 reads and 2,250 genes per cell. Unsupervised clustering identified 13 clusters, including both CD8+ and CD4+ subsets (Extended Data Fig. 2g–i).
Reclustering of CD8+ T cells defined ten clusters in the UMAP (Fig. 2c). Two proliferating clusters were identified (Fig. 2c,d and Extended Data Fig. 2j). An effector-like cluster upregulated TBX21 and GZMB expression (Fig. 2c,d and Extended Data Fig. 2j). A type 2 CD8+ subset52,53 expressed genes associated with type 2 helper T cells, such as GATA3, IL5 and IL13 (Fig. 2c–e and Extended Data Fig. 2j). We also identified a stem-like population that upregulated the expression of genes including BACH2, CD27, TCF7 and LEF1 (Fig. 2c–e and Extended Data Fig. 2j). Human stem-like preinfusion CAR T cells showed the highest level of enrichment of the LT stem-like signature compared to other subsets and downregulated BACH2 expression and the LT stem-like signature as they differentiated into the effector-like subset (Fig. 2f and Extended Data Fig. 2k,l). Of note, LT stem-like signature genes, including BACH2, SELL and TCF7, were upregulated in CD8+ T cells from individuals with a CR compared to those from individuals who did not show a CR (Fig. 2g). In addition, gene set enrichment analysis (GSEA) showed that the LT stem-like gene signature was enriched in CD8+ T cells from individuals who showed a CR compared with those without CR (Fig. 2h). Therefore, LT stem-like signature genes, including BACH2, are upregulated in human CD8+ T cells in the infusion product from individuals who showed a CR to CD19 CAR T cell therapy.
BACH2high LT stem-like CAR T cells develop after leukemia clearance
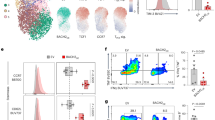
We previously showed that treatment with CD19 CAR T cells cleared leukemia in mice bearing E2A–PBX1 B cell ALL within 7 days of infusion16. To understand differentiation of CD19 CAR T cells after leukemia clearance, we performed scATAC-seq + scRNA-seq with CD19 CAR T cells from the spleens of mice inoculated with E2A–PBX1 20 days after CAR T cell infusion. scATAC-seq + scRNA-seq profiles were generated from 4,137 CD8+ CAR T cells with an average of 1,928 genes and 10,941 peaks detected in each cell. Unsupervised clustering defined seven subsets based on the scRNA-seq profile (Fig. 3a). Among major transcriptome-defined subsets was a stem-like subset (cluster 0) that expressed high levels of Il7r, Slamf6 and Tcf7 (Fig. 3a,b). Cluster 1 represented a natural killer (NK)-like subset16,54,55 and showed upregulation of genes including Zeb2, Klre1, Klrb1c (NK1.1) and Nkg7 (Fig. 3a,b and Extended Data Fig. 3a). Notably, cluster 2 cells showed upregulation of LT stem-like markers, including Bach2, Sell (CD62L), Foxp1, Lef1 and Satb1, and downregulation of the exhaustion marker Tox compared to stem-like CAR T cells (Fig. 3a–c and Extended Data Fig. 3b,c). Single-cell GSEA showed the highest enrichment of the LT stem-like signature in Bach2high cluster 2 cells, whereas enrichment of the LT stem-like signature and levels of Bach2 transcript were intermediate in the stem-like subset and low in the NK-like subset (Fig. 3c,d). Thus, the transcript level of Bach2 positively correlates with the degree of stemness in CAR T cells after tumor clearance.
a–g, scATAC-seq + scRNA-seq was performed with CAR T cells on day 20 after treatment in C57BL/6 mice that were inoculated with E2A–PBX1 and treated with CD19 CAR T cells. a, Transcriptome-based UMAP plot (RNA UMAP) showing CD8+ CD19 CAR T cells color coded based on cluster IDs, including stem-like (RNA 0 Tstem-like), NK-like (RNA 1 TNK-like), LT stem-like (RNA 2 LT stem-like), interferon-stimulated genes (RNA 3 ISG), proliferating (RNA 4), tissue-resident memory-like (RNA 5 TRM-like) and Nr4a1+ exhausted-like (RNA 6 Nr4a1+ Tex-like) clusters. b, Heat map of top marker genes in each transcriptome-based cluster defined in a. c, Violin plots showing single-cell expression of Bach2, Sell, Lef1, Foxp1, Satb1 and Tox in each transcriptome-based cluster defined in a. d, Violin plots showing enrichment of the LT stem-like signature in each transcriptome-based cluster defined in a. e, scATAC-seq-based UMAP plots showing CAR T cells color coded based on cluster IDs defined by chromatin state (left) or transcriptome (right). f, Volcano plot illustrating genes with differentially accessible (DA) chromatin between stem-like and LT stem-like CAR T cells. g, Volcano plot of differentially enriched (DE) transcription factor binding motifs between stem-like and LT stem-like CAR T cells. Adjusted P values in f and g were calculated using a two-sided Wilcoxon rank-sum test with a Bonferroni correction to account for multiple comparisons.
By connecting cis-regulatory elements to genes through correlating chromatin accessibility with transcription, we found that Bach2 was among the genes controlled by the highest number of enhancers (Extended Data Fig. 3d). Using FigR56, we connected transcription factors with their potential target genes. Notably, BACH2 was among the transcription factors with the highest number of potential targets (Extended Data Fig. 3e), suggesting that BACH2 is a key regulator of CAR T cells after leukemia clearance.
The LT stem-like subset showed a chromatin state distinct from that of stem-like T cells (Fig. 3e). Compared to stem-like T cells, LT stem-like cells displayed greater chromatin accessibility at genes including Bach2, Satb1 and Lef1 and showed elevated enrichment of the epigenetic signature of long-lived stem-like T cells generated through Asxl1 KO50 (Fig. 3f and Extended Data Fig. 3f). Transcription factor motif analysis revealed that, compared to stem-like CAR T cells, LT stem-like CAR T cells showed higher enrichment of motifs of pro-stem transcription factors, including TCF1 and LEF1, and reduced motif enrichment of AP1, NFAT and RUNX (Fig. 3g). In addition, LT stem-like CAR T cells showed enhanced activity of the transcriptional repressor BACH2, manifested by reduced chromatin accessibility at BACH2 binding sites (Fig. 3g). Therefore, our data suggest that LT stem-like CAR T cells emerge after tumor clearance and demonstrate the highest level of BACH2 activation and repression of TOX.
A subset of CAR T cells maintains LT stem-like fate following rechallenge
CD19 CAR T cells generated from a Tcf7-green fluorescent protein (Tcf7-GFP) reporter mouse were adoptively transferred into E2A–PBX1 leukemia-bearing mice. On day 7 after infusion, Tcf7-GFP+ CD8+ CAR T cells and Tcf7-GFP− CD8+ CAR T cells were isolated and transferred separately to a second cohort of leukemia-bearing mice. Seven days after treatment of the second cohort, approximately tenfold more progeny of Tcf7-GFP+ CD8+ CAR T cells accumulated in the spleen and bone marrow of treated mice than progeny of Tcf7-GFP− CD8+ CAR T cells (Fig. 4a and Extended Data Fig. 4a). Whereas most of the progeny from Tcf7-GFP+ CD8+ CAR T cells terminally differentiated into Tcf7-GFP−TIM3+, a small fraction of cells derived from Tcf7-GFP+ donors retained a Tcf7-GFP+TIM3− phenotype (Fig. 4b and Extended Data Fig. 4b). By contrast, the terminally differentiated Tcf7-GFP− donors only gave rise to Tcf7-GFP−TIM3+ progeny (Fig. 4b and Extended Data Fig. 4b). A small population of Tcf7-GFP+ progeny retained expression of the LT stem-like marker CD62L, whereas few Tcf7-GFP− progeny expressed CD62L (Fig. 4c and Extended Data Fig. 4c). Surface levels of PD-1 and TIM3 were higher in Tcf7-GFP− progeny than in Tcf7-GFP+ progeny (Fig. 4d,e and Extended Data Fig. 4d,e). To assess the recall response of LT stem-like CAR T cells after leukemia clearance, we sorted CD62L+Tcf7-GFP+ CD8+ LT stem-like CAR T cells and their CD62L−Tcf7-GFP− counterparts on day 24 after CAR T cell treatment of mice that were previously inoculated with E2A–PBX1 and transferred them separately to a second cohort of leukemia-bearing mice. On day 7 after infusion, CD8+ CAR T cells derived from LT stem-like cells exhibited greater expansion, enhanced ability to retain the CD62L+CX3CR1− stem-like phenotype and lower levels of PD-1 and TIM3 (Extended Data Fig. 4f–i). Thus, our results demonstrate that a subset of post-treatment CAR T cells preserves proliferative potential and LT stem-like differentiation during the secondary response against tumor antigen.
a–e, E2A–PBX1-bearing mice were treated with CD19 CAR T cells coexpressing truncated huEGFR generated from a Tcf7-GFP reporter mouse. Tcf7-GFP+ and Tcf7-GFP− CD8+ CAR T cells were sorted on day 7 after infusion and transferred separately to a second batch of E2A–PBX1-bearing mice (n = 5 mice per group). a, Left: representative flow cytometry plot of splenic CD8+ CAR T cells (huEGFR+CD45.1+) in mice that received Tcf7-GFP+ or Tcf7-GFP− CD8+ CAR T cells. Right: numbers of progeny from Tcf7-GFP+ or Tcf7-GFP− CD8+ CAR T cells in the spleen. b, Left: representative flow cytometry plot of splenic Tcf7-GFPhighTIM3low CD8+ CAR T cells in mice that received Tcf7-GFP+ or Tcf7-GFP− CD8+ CAR T cells. Right: frequencies of splenic Tcf7-GFPhighTIM3low CD8+ CAR T cells derived from Tcf7-GFP+ or Tcf7-GFP− donors. c, Left: representative flow cytometry plot of splenic CD62LhighTIM3low CD8+ CAR T cells in mice that received Tcf7-GFP+ or Tcf7-GFP− CD8+ CAR T cells. Right: frequencies of splenic CD62LhighTIM3low CD8+ CAR T cells derived from Tcf7-GFP+ or Tcf7-GFP− donors. d,e, Expression of PD-1 (d) and TIM3 (e) in splenic CD8+ CAR T cells derived from Tcf7-GFP+ and Tcf7-GFP− donors. Data are representative of two independent experiments. Bar graphs represent the mean ± s.e.m. Circles in the bar graphs represent individual mice. Statistical significance was calculated with a two-sided Student’s t-test; MFI, mean fluorescence intensity.
BACH2 promotes antitumor immunity and the LT stem-like program of CAR T cells
To determine the role of BACH2 in the antitumor response of CAR T cells in vivo, we generated Bach2loxP/loxP CreERT2 CD19 CAR T cells with induced deletion of Bach2 (Bach2 induced KO (iKO)) and compared them with Bach2loxP/loxP (control) CD19 CAR T cells after adoptive transfer to E2A–PBX1-bearing mice. BACH2 deficiency led to a greater than twofold reduction in the number of CD8+ CAR T cells and an approximately fivefold reduction in the number of CD4+ CAR T cells in the spleen on day 7 after infusion (Fig. 5a,b). BACH2 deficiency significantly reduced the frequency of TCF1+ cells and the expression of the LT stem-like marker CD62L in CD8+ CAR T cells (Fig. 5c,d). Immune checkpoints, including TIM3 and PD-1 as well as TOX, were upregulated in Bach2-iKO CD8+ CAR T cells (Fig. 5e–g). Similarly, BACH2 deficiency decreased the numbers of CD8+ CAR T cells and CD4+ CAR T cells in both the spleens and tumors of B16-CD19-bearing mice (Extended Data Fig. 5a–d). Bach2-iKO CD8+ CAR T cells showed decreased expression of markers associated with stemness (Extended Data Fig. 5e,f). Next, we performed scRNA-seq with 2,496 control and 4,243 Bach2-iKO CD8+ CAR T cells (Fig. 5h and Extended Data Fig. 5g). BACH2 deficiency caused a clear shift in the scRNA-seq profiles of CD8+ CAR T cells (Fig. 5h). Bach2-iKO CD8+ CAR T cells contained fewer stem-like cells (cluster 1; Fig. 5i,j). Comparison between control and Bach2-iKO stem-like CD8+ CAR T cells showed that BACH2 deficiency downregulated the expression of stem/memory markers, including Ccr7 and Id3, and upregulated AP1 transcription factors, which are linked to exhaustion of CAR T cells6, including Junb (Extended Data Fig. 5h). Notably, BACH2-deficient stem-like CD8+ CAR T cells downregulated the LT stem-like gene signature compared to control stem-like CAR T cells (Fig. 5k). To identify genomic targets of BACH2, we performed BACH2 chromatin immunoprecipitation with sequencing (ChIP–seq) with CD8+ CD19 CAR T cells. BACH2 was primarily bound to enhancers in intronic or intergenic regions (Extended Data Fig. 5i). BACH2 binding sites showed enrichment of motifs associated with BACH2 and AP1 transcription factors, such as JunB (Extended Data Fig. 5j). JunB ChIP–seq in CAR T cells revealed that BACH2 overexpression reduced genomic binding by JunB at BACH2 binding sites with differential JunB binding (Extended Data Fig. 5k). Using guide RNA (gRNA) targeting Junb, we showed that JunB-deficient CD8+ CAR T cells exhibited a higher frequency of the CD62L+CX3CR1− stem-like subset (Extended Data Fig. 5l). To evaluate the recall response of BACH2-deficient CAR T cells, we collected control and Bach2-iKO CAR T cells from mice inoculated with E2A–PBX1 on day 20 after CAR T cell treatment and transferred them separately to a second cohort of E2A–PBX1-bearing mice. On day 7 after infusion, BACH2 deficiency in CD8+ CAR T cells impaired the recall response, reduced the frequency of the stem-like subset and increased expression of PD-1 and TIM3 (Extended Data Fig. 5m–p). Together, these data show that BACH2 expression in CAR T cells is necessary for the LT stem-like transcriptional program and antitumor response.
a–g, CD8+ T cells were isolated from Bach2loxP/loxP (control) and Bach2loxP/loxP CreERT2 (Bach2-iKO) mice after tamoxifen treatment and retrovirally transduced with CD19 CAR coexpressing truncated huEGFR. E2A–PBX1-bearing mice treated with control or Bach2-iKO CD19 CAR T cells were analyzed on day 7 after infusion (control: n = 7 mice, Bach2 iKO: n = 8 mice). a, Representative flow cytometry plots (left, gated on CD8+ cells) and the number (right) of splenic CD8+ CAR T cells (CD8+huEGFR+) in each group. b, Representative flow cytometry plots (left, gated on CD4+ cells) and the number (right) of splenic CD4+ CAR T cells (CD4+huEGFR+) in each group. c, Left: TCF1 expression in splenic CD8+ CAR T cells in each group. Right: frequency of TCF1+ cells among splenic CD8+ CAR T cells in each group. d–g, Expression of CD62L (d), TIM3 (e), PD-1 (f) and TOX (g) in splenic CD8+ CAR T cells in each group. Data are representative of two independent experiments. Bar graphs represent mean ± s.e.m. Circles represent individual mice. Statistical significance was calculated with a two-sided Student’s t-test. h–k, B16-CD19-bearing mice were treated with control or Bach2-iKO CD19 CAR T cells. scRNA-seq was performed with control or Bach2-iKO splenic CD8+ CAR T cells on day 8 after infusion. h, UMAP plot of combined scRNA-seq data from control or Bach2-iKO CD8+ CAR T cells. Cells are color coded based on the genotype. i, Left: UMAP plot of control or Bach2-iKO CD8+ CAR T cells color coded based on cluster IDs. Right: frequencies of cells from different clusters in control or Bach2-iKO CD8+ CAR T cells. j, Single-cell enrichment of the stem-like gene signature (GSE202543) shown in a UMAP plot (left) and violin plot (right). k, Enrichment of the LT stem-like gene signature (GSE199839) in control stem-like (cluster 1) versus Bach2-iKO stem-like (cluster 1) CAR T cells. The adjusted P value was calculated using the Benjamini–Hochberg procedure to correct for multiple testing. The red dashed line in the GSEA plot indicates the location of the maximum running enrichment score (ES).
Chemical control of BACH2 tunes CAR T cell differentiation and function
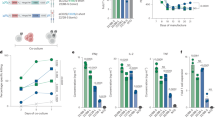
To fine-tune BACH2 expression, we fused BACH2 with a destabilizing domain (DD–BACH2). DD-fused protein is targeted for proteasomal degradation unless stabilized by the small molecule Shield-1 (ref. 57; Fig. 6a). Varying the dose of Shield-1 tuned the amount of BACH2 protein in T cells (Extended Data Fig. 6a). To determine how the level of BACH2 affects the stemness and exhaustion of CAR T cells, we used an in vitro T cell exhaustion model in which persistent tonic signaling of GD2-specific CAR 14g2a-E101K-28z caused by antigen-independent aggregation drives T cell exhaustion6. GD2 CAR T cells were co-transduced with a control construct or a construct overexpressing wild-type BACH2 or DD–BACH2. DD–BACH2 CD8+ GD2 CAR T cells were treated with 0, 100 or 1,000 nM Shield-1. Constitutive overexpression of BACH2 (BACH2 OE) increased the frequency of the CD62L+ stem-like subset and downregulated the expression of immune checkpoints, including PD-1 and TIM3 (Fig. 6b–d). Notably, DD–BACH2 CD8+ GD2 CAR T cells treated with 100 or 1,000 nM Shield-1 showed significantly higher frequencies of the CD62L+ stem-like subset and lower levels of PD-1 than control CD8+ GD2 CAR T cells (Fig. 6b,c). DD–BACH2 downregulated TIM3 in CAR T cells in a Shield-1 dose-dependent manner (Fig. 6d). Remarkably, a low level of exogenous BACH2 in untreated DD–BACH2 CD8+ GD2 CAR T cells was sufficient to increase the frequency of stem-like T cells and decrease PD-1 and TIM3 expression, albeit to a lesser extent than Shield-1-treated cells (Fig. 6b–d and Extended Data Fig. 6a). Thus, the degree of stemness and exhaustion in GD2 CAR T cells is quantitatively controlled by the expression level of BACH2.
a, Schematic illustration of chemically controlling the protein level of DD–BACH2 by Shield-1. b–d, CD8+ GD2 CAR T cells were co-transduced with empty murine stem cell virus (MSCV)-IRES-GFP (pMIG) plasmid (control) or a pMIG plasmid overexpressing DD–BACH2 or wild-type BACH2 (BACH2 OE). DD–BACH2 GD2 CAR T cells were cultured with 0, 100 or 1,000 nM Shield-1 (n = 3 independent cultures per group). Expression of CD62L (b), PD-1 (c) and TIM3 (d) in CD8+ CAR T cells was evaluated. e, Schematic illustration of inducing nuclear translocation of BACH2–ERT2 with 4-OHT. f–h, CD8+ GD2 CAR T cells co-transduced with empty pMIG (control) or pMIG with a BACH2–ERT2 insert were cultured with 1 μM 4-OHT (n = 3 independent cultures per group). Levels of CD62L (f), PD-1 (g) and TIM3 (h) in CD8+ GD2 CAR T cells were determined. i, Tumor growth in mice bearing 9464D-GD2 neuroblastoma after treatment of control GD2 CAR T cells or BACH2–ERT2 GD2 CAR T cells described in f (n = 5 mice per group). Data are representative of two independent experiments. Bar graphs represent the mean ± s.e.m. Statistical significance in b–d was determined by one-way ANOVA with a Bonferroni test to correct for multiple comparisons. Statistical significance in f–h was calculated with a two-sided Student’s t-test. Statistical significance in i was determined by two-way ANOVA with a Tukey test to correct for multiple comparisons. Schematics in a and e were created using BioRender.
To determine the effect of transient induction of DD–BACH2 on stemness, DD–BACH2 CD8+ GD2 CAR T cells were first cultured with Shield-1 before removing Shield-1 from culture. The withdrawal of Shield-1 reduced BACH2 expression to the baseline level within 24 h (Extended Data Fig. 6b). However, even after the loss of DD–BACH2 expression due to Shield-1 withdrawal, GD2 CAR T cells partially retained expression of the stem-like marker CD62L and maintained lower expression levels of immune checkpoints such as PD-1 and TIM3 (Extended Data Fig. 6c–e).
Next, we sought to determine whether chemically inducing BACH2 expression enhances stemness in human CAR T cells. Human GD2 CAR T cells were co-transduced with a control construct or a DD–BACH2 construct. DD–BACH2 human GD2 CAR T cells were cultured with or without 1 µM Shield-1. Induction of DD–BACH2 expression by Shield-1 in human CD8+ GD2 CAR T cells increased the expression of stem/memory markers CCR7 and CD127 (interleukin-7 receptor-α (IL-7Rα)) and downregulated immune checkpoints TIM3 and LAG3 (Extended Data Fig. 6f–i). Thus, inducing BACH2 expression through a chemical switch enhances stemness and suppresses exhaustion in human CAR T cells.
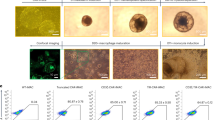
We next sought to determine whether inducing BACH2 activity during CAR T cell manufacturing by controlling BACH2 nuclear translocation improves the stemness and antitumor immunity of CAR T cells. BACH2 was fused to ERT2 that has enhanced reactivity to tamoxifen and minimal reactivity to endogenous estrogen to control the nuclear translocation of the BACH2–ERT2 fusion protein but not the endogenous BACH2 protein. Wild-type CD8+ GD2 CAR T cells co-transduced with a control construct or a BACH2–ERT2 construct were cultured with 4-hydroxytamoxifen (4-OHT) to induce nuclear translocation of BACH2–ERT2 (Fig. 6e). Compared to control CD8+ GD2 CAR T cells, BACH2–ERT2 GD2 CAR T cells showed an increase in the frequency of the CD62L+ stem-like subset and a reduction in the levels of PD-1 and TIM3 (Fig. 6f–h). Next, we transferred control or BACH2–ERT2 CD8+ GD2 CAR T cells cultured with 4-OHT into mice bearing 9464D-GD2 neuroblastoma38,58 (Fig. 6i). Mice treated with CD8+ GD2 CAR T cells programmed by BACH2 during manufacture exhibited greater tumor control than those that received control CAR T cells (Fig. 6i). Together, these data suggest that chemically harnessing the level and timing of BACH2 expression in CAR T cells quantitatively controls the stem-like and exhaustion differentiation programs and improves control of solid tumors.
Discussion
Single-cell omics revealed that stem-like T cells are a heterogenous population containing subsets with different degrees of stemness, exhaustion and functionality. In cancer and chronic infection, an LT stem-like T cell subset has the highest capacity to persist, repopulate terminally exhausted T cells and proliferate in response to ICIs36,42,43,44. LT stem-like T cells also resist the exhaustion program and maintain a lower level of TOX than Tpex cells42, underscoring the important clinical relevance of LT stem-like T cells. However, how the hierarchy of T cell stemness is programmed is not fully defined. The transcription factor MYB is essential for the differentiation of CD62L+ LT stem-like T cells, whereas MYB deficiency led to a hyper-response by CD8+ T cells and T cell-driven immunopathology36. Our results suggest that the position of T cells in the spectrum from stemness to exhaustion is determined by the level of BACH2. However, unlike MYB, loss of BACH2 did not enhance the response of terminally differentiated T cells but instead impaired expansion by CAR T cells. Therefore, BACH2 and MYB may regulate LT stem-like T cells through different downstream mechanisms. In healthy humans, two populations of stem cell-like memory T cells resemble LT stem-like T cells and Tpex cells, respectively49. Interestingly, BACH2 is highly expressed by activated human LT stem-like T cells, whereas activated Tpex cells express higher levels of TOX and immune checkpoints PD-1 and TIGIT49. Of note, our data showed that BACH2 deficiency increased TOX expression in CAR T cells. Although LT stem-like T cells formed a transcriptionally and epigenetically distinct subset in CAR T cells after clearing tumors in vivo, the LT stem-like subset in preinfusion CAR T cells was more distinct at the epigenetic level than at the transcriptional level. Thus, it is possible that before antigen experience, a subset of preinfusion CAR T cells is epigenetically predisposed to develop into LT stem-like T cells. Future mechanistic studies are needed to determine how the distinct chromatin states in preinfusion CAR T cells impact their antitumor response during adoptive cell therapy.
Stem-like or memory-like phenotype in premanufacture T cells or preinfusion CAR T cells correlates with favorable outcomes in CAR T cell therapy9,10,11. CAR T cells produced from enriched naive/memory T cells show a greater antitumor efficacy28,59. Instead of adding a cell enrichment step to the CAR T cell manufacturing process, recent studies developed strategies to improve antitumor efficacy by CAR T cells through overexpressing signaling molecules that promote T cell stemness and/or repress exhaustion6,34,35,45. Transcription factors dictate T cell differentiation and function and are ideal targets for programming CAR T cells to acquire stemness and resist exhaustion. However, instead of functioning in a binary manner, transcription factor activation is often under tight temporal and quantitative control. The precise level and timing of pro-stem transcription factors may differ for different CAR designs and tumor types. In this study, we developed two strategies to chemically control the protein level or nuclear translocation of BACH2 to allow quantitative and temporal control of CAR T cell differentiation. We further showed that temporary activation of BACH2 by a small-molecule drug during manufacturing of exhaustion-prone GD2 CAR T cells limited tonic signaling-driven exhaustion and enhanced control of a solid tumor. Future studies are warranted to evaluate similar strategies to control other transcriptional regulators of T cell differentiation in CAR T cells against different malignancies. In addition to improving tumor control, transcriptional regulators can be targeted in therapeutic T cells to achieve desirable cytokine profiles and/or prevent toxicity.
In summary, we have shown that the hierarchical structure of stem-like T cells is quantitatively controlled by the level of BACH2. Through controlling BACH2 activity in a temporal and tunable manner by small-molecule drugs, we have engineered CAR T cells of which differentiation can be fine-tuned to improve tumor control. Thus, our study shed light on a new avenue to enhance the efficacy of T cell-based immunotherapy through dynamically controlling transcriptional regulators of T cell differentiation.
Methods
Mice
Male and female C57BL/6J (B6, strain 000664), B6.SJL-Ptprca Pepcb/BoyJ (B6 CD45.1, strain 002014) and B6(Cg)-Tcf7tm1Hhx/J (Tcf7-GFP reporter, strain 030909) mice were purchased from The Jackson Laboratory. Bach2loxP/loxP and Bach2loxP/loxP CreERT2 mice were described in our previous study15. All mice used in the experiments of this study were between 6 and 12 weeks old, age and sex matched and maintained on a B6 genetic background. Mice were fed an irradiated 2916 Teklad Global Rodent Diet and were kept in specific pathogen-free facilities with a 12-h light/12-h dark cycle. The room temperature range was 20–25 °C. Humidity was maintained at 30–70%. All animal procedures were approved by the Institutional Animal Care and Use Committee at University of Texas Southwestern Medical Center under protocol numbers 103162 and 103111.
Tumor inoculation
Syngeneic tumor cell lines E2A–PBX1 B cell ALL, B16-CD19 melanoma and 9464D-GD2 neuroblastoma are in a B6 background and have been described in previously published studies46,58,60. E2A–PBX1 cells were intravenously inoculated at 1 × 106 per mouse, whereas B16-CD19 and 9464D-GD2 cells were subcutaneously injected at 1 × 106 or 2 × 106 per mouse, respectively. Mice were euthanized if the tumor size exceeded 2 cm in diameter, if the tumor burden exceeded 10% of body weight or if animals exhibited >20% loss in body weight. Mice were also euthanized if tumors caused notable clinical signs, including pain, impaired mobility or evidence of metastasis. At no point during the study were these humane endpoints exceeded.
Transfection and retroviral transduction
For transducing mouse T cells, retroviral plasmid, pCL-Eco plasmid, Opti-MEM (Thermo Fisher Scientific) and TransIT-293 Transfection Reagent (Mirus Bio) were mixed, incubated and added to HEK293T cell cultures (ATCC, CRL-3216). Two days after transfection, culture supernatants with retroviruses were collected and centrifuged to remove cell debris. Mouse T cells activated by plate-bound anti-CD3 and anti-CD28 were transduced by a mixture of retroviruses and 8 µg ml−1 polybrene via spinoculation at 32 °C for 90 min. Anti-mouse CD19 CAR, described in our previous study16, was derived from the monoclonal antibody 1D3 and contains sequences from mouse CD28 and CD3ζ. An MSCV retroviral plasmid expressed anti-mouse CD19 CAR and a truncated huEGFR separated by a P2A peptide. The MSGV plasmid with an anti-GD2 CAR insert was a generous gift from C. Mackall (Stanford University) and was used in our previous study38. pMIG with a human BACH2 OE cassette was described by us previously15. A previously described BACH2–ERT2 cassette61 was inserted into pMIG to generate a retroviral construct with tamoxifen-responsive BACH2. To generate DD–BACH2 pMIG plasmid, a destabilization domain derived from a mutant FKBP12 (ref. 57) was fused to BACH2. A gRNA targeting Junb (ACGCAGCGGCGGGATACGGT) was cloned into SL21-VEX plasmid, a gift from J. Shi (University of Pennsylvania). For transducing human T cells, retroviral supernatants were generated using the 293GP packaging cell line (Takara, 631512). Briefly, 293GP cells were co-transfected with retroviral and RD114 envelope plasmids using TransIT-293 Transfection Reagent (Mirus Bio) in Opti-MEM (Thermo Fisher Scientific). Viral supernatants were collected 48 h after transfection and centrifuged to remove cellular debris. Human T cells purified from the peripheral blood mononuclear cells of healthy donors with an EasySep Human T Cell Isolation kit (StemCell Technologies) were activated with Dynabeads Human T-Expander CD3/CD28 (Gibco) at a 3:1 bead:cell ratio and transduced on days 2 and 3 after activation. Non-tissue culture-treated 24-well plates were coated overnight at 4 °C with RetroNectin (25 μg ml−1; Takara) in PBS, washed twice with PBS and blocked with 2% bovine serum albumin for 15 min. Retroviral supernatants were added, and plates were centrifuged at 2,000g for 2 h at 32 °C before the addition of T cells. On day 4 after activation, beads were removed by magnetic separation using an EasySep Magnet (StemCell Technologies).
Radiation and cell transfer
One day before CAR T cell transfer, mice were sublethally irradiated (500 cGy) as previously described16,38,47. CD19 CAR T cells or GD2 CAR T cells were adoptively transferred into mice through tail vein injection.
Tamoxifen treatment
As we previously described15, Bach2loxP/loxP CreERT2 mice were intraperitoneally injected with tamoxifen dissolved in sunflower seed oil to induce deletion of the floxed genomic segment. Control (Bach2loxP/loxP) mice were also treated with tamoxifen.
Flow cytometry and cell sorting
Flow cytometry and fluorescence-activated cell sorting (FACS) in this study were performed with the following antibodies and dyes. Anti-mouse CD45.1–Alexa Fluor 700 (A20, 1:200), anti-mouse CD4–FITC (RM4-5, 1:200), anti-mouse CD8a–BV605 (53-6.7, 1:200), anti-human EGFR–PE/Cy7 (AY13, 1:200), anti-mouse PD-1–PE (RMP1-30, 1:200), anti-mouse KLRG1–BV421 (2F1, 1:200), anti-mouse TIM3–APC (RMT3-23, 1:100), anti-mouse TIM3–BV605 (RMT3-23, 1:100), anti-mouse TIM3–PE/Cy7 (RMT3-23, 1:100), anti-mouse CX3CR1–FITC (SA011F11, 1:200), anti-mouse CX3CR1–PE/Cy7 (SA011F11, 1:800), anti-human CD4–PerCP/Cy5.5 (OKT4, 1:400), anti-human CD366 (TIM3)–PE (F38-2E2, 1:200), anti-human CD197 (CCR7)–PE/Cy7 (G043H7, 1:200), anti-human CD8–Alexa Fluor 700 (SK1, 1:400), anti-human CD39–APC/Cy7 (A1, 1:200), anti-human CD127 (IL-7Rα)–APC/Cy7 (A019D5, 1:200), anti-human CD62L–BV421 (DREG-56, 1:200) and anti-human CD45RA–BV785 (HI100, 1:200) were purchased from BioLegend. Anti-mouse CD62L–PerCP/Cy5.5 (MEL-14, 1:200), anti-TOX–PE (TXRX10, 1:100), goat anti-rabbit IgG (H + L) APC (1:500), anti-human CD223 (LAG3)–PE (3DS223H, 1:200), anti-human CD279 (PD-1)–PE/Cy7 (eBioJ105 (J105), 1:200), LIVE/DEAD Fixable Aqua Dead Cell Staining kit (1:250) and LIVE/DEAD Fixable Near-IR Dead Cell Stain kit (1:400) were purchased from Thermo Fisher. Anti-mouse granzyme B–BV421 (GB11, 1:200) and anti-active caspase-3–FITC (C92-605, 1:10) were from BD Biosciences. Anti-TCF1 (C63D9, 1:200) was purchased from Cell Signaling Technology. Flow cytometry was performed with a Cytek Aurora using SpectroFlo (v3.0.3). FACS was performed with a BD FACSAria II using BD FACSDIVA (v9.0.1). Analysis was performed with FlowJo 10.10.0.
Western blotting
Proteins were extracted from cells using RIPA Lysis and Extraction Buffer (Thermo Scientific) supplemented with Protease Inhibitor Cocktail (Thermo Scientific) and used for SDS–PAGE. The proteins were subsequently transferred to a PVDF membrane (Fisher Scientific) using a wet transfer method. The blots were blocked with 5% bovine serum albumin in Phosphate-Buffered Saline with Tween 20 and probed overnight at 4 °C with primary antibodies to BACH2 (1:2,000; Abcam, 7A4) and β-actin (1:1,000; Cell Signaling Technology, 8H10D10). Following washing, the blots were incubated with secondary horseradish peroxidase-conjugated antibodies (Cell Signaling Technology) at a 1:5,000 dilution for 60 min. Protein bands were visualized using enhanced chemiluminescent substrate (Thermo Scientific), according to the manufacturer’s instructions, on a ChemiDoc MP Imaging System (Bio-Rad).
Sample preparation for scRNA-seq
We generated scRNA-seq libraries using a Chromium Next GEM Single Cell 5′ kit (10x Genomics). Briefly, T cells labeled with TotalSeq Hashtag Antibody (BioLegend) from three biological replicates were sorted, which were then combined for each group. Following a wash step, cells were loaded onto a Chromium Single Cell Chip G to produce barcoded DNA. After DNA amplification, we prepared gene expression libraries using large cDNA fragments and cell surface protein libraries (HTO) from small DNA fragments (~200 bp). Gene expression and HTO libraries were multiplexed and sequenced on an Illumina NovaSeq 6000 using the same cycle settings described above.
Sample preparation for scATAC-seq + scRNA-seq
We generated scRNA-seq and scATAC-seq libraries using a Chromium Next GEM Single Cell Multiome ATAC + Gene Expression Reagent kit (10x Genomics), following the manufacturer’s user guide. Specifically, we sorted cells for nuclei isolation and followed the Low Cell Input Nuclei Isolation protocol from 10x Genomics to minimize nuclei loss. We tagmentated the isolated nuclei and captured them using the 10x Genomics Chromium Single Cell Controller. After GEM cleanup and preamplification PCR, we prepared gene expression and ATAC libraries separately. Sequencing of the gene expression libraries was performed on an Illumina NovaSeq 6000 using the following sequencing configuration: read 1 (26 cycles), i7 index (10 cycles), i5 index (10 cycles) and read 2 (90 cycles), targeting 30,000 reads per cell. The ATAC libraries were sequenced with read 1 (50 cycles), i7 index (8 cycles), i5 index (24 cycles) and read 2 (49 cycles), targeting 30,000 reads per cell.
Sample preparation for ChIP–seq
Ten million mouse CD19 CAR T cells transduced with pMIG or pMIG-BACH2 (human) were used to generate ChIP–seq libraries. A truChIP Chromatin Shearing kit (Covaris, 520154) was used for chromatin cross-linking, nuclei isolation and chromatin shearing. Ten micrograms of anti-BACH2 (Cell Signaling Technologies, 80775, clone D3T3G, 1:100) or anti-JunB (Cell Signaling Technologies, 3753, clone C37F9, 1:100) was used to pull down sheared chromatin from CD8+ T cells. ChIP–seq libraries were constructed using a published ChIPmentation protocol62. We sequenced the ChIP–seq libraries on an Illumina NextSeq 550, generating at least 20 million reads per sample.
Data analysis for scRNA-seq
For human CAR T cell samples, the unique molecular identifier (UMI) count matrix for each sample was downloaded from the GSE241783 dataset. We used R package Seurat (v4.4.0) for the subsequent analysis. We merged cells from 40 individuals and filtered the cells with the following criteria: 200–7,000 detected genes, 1,000–15,000 detected RNA molecules and less than 15% mitochondrial genes. To focus on CD8+ CAR T cells, we selected CAR T cells expressing either CD8A or CD8B and not expressing CD4. Monocle 3 (ref. 63) was used to reconstruct developmental trajectories and infer transcriptional dynamics. The preprocessed Seurat object was imported into Monocle 3 while preserving the original UMAP embedding. Developmental lineages were inferred by applying the ‘learn_graph’ function, which fits a principal graph onto the UMAP manifold to capture the underlying cellular topology. Cells were then ordered along pseudotime using ‘order_cells’, with biologically defined root populations designated as starting points.
For mouse CAR T cell scRNA-seq fastq files, we used Cell Ranger (v6.0.0) to align FASTQ files to the mm10 reference genome and to quantify barcodes and UMIs for both the gene expression and hashtag libraries with the cellranger count pipeline. We combined the cells from different samples using the merge function and retained those with 1,000–3,500 detected genes, 2,000–15,000 detected RNA molecules and less than 5% mitochondrial gene content.
After quality control, we log normalized the data using a scale factor of 10,000. During the ScaleData function, we regressed out the effects of cell cycle, the number of detected genes and mitochondrial gene content. T cell antigen receptor and immunoglobulin genes were excluded from the top 2,000 variable genes identified with FindVariableFeatures, and the remaining genes were used for principal component analysis via RunPCA. We used the top 20 principal components to calculate neighbors, cluster the cell types and generate the UMAPs. We used the FindAllMarkers function or FindMarkers (min.pct = 0.1, logfc.threshold = 0.1) to determine markers genes in each cluster and the HTODemux function to determine Hashtag antibody labeling of each biological replicate. We used pheatmap (v1.0.12) or the DoHeatmap function to generate heat maps. We calculated gene set enrichment at the single-cell level using the AddModuleScore function. Additionally, we performed GSEA for two clusters of cells using the clusterProfiler package (v4.12.6).
Data analysis for scATAC-seq + scRNA-seq
scATAC-seq + scRNA-seq data were analyzed as previously described16. Cell versus gene matrices of UMI counts for gene expression assays and cell versus fragment matrixes for ATAC assays were generated from fastq files aligned to mm10 using cellranger-arc (v2.0.0). We performed subsequent analyses using Signac (v1) and Seurat (v4). Cells were retained if they had 1,000–4,000 detected genes, 15,000–40,000 detected ATAC fragments and less than 8% UMIs mapped to the mitochondrial genome. Additionally, we excluded cells with fewer than 50% of ATAC reads mapped to peaks, a nucleosome signal greater than 2 and a transcription start site enrichment score below 2. Using the approach outlined in ‘Data analysis for scRNA-seq’, we analyzed the scRNA-seq data to calculate UMAPs and identify clusters based on gene expression. For the scATAC-seq data, peaks were called using MACS2, and those located in the mm10 genomic blacklist or nonstandard chromosomes were removed. The FeatureMatrix function was used to quantify counts within the peaks, generating a peak assay for downstream analyses. After normalizing the data, top variable features were identified using FindTopFeatures. Latent semantic indexing was performed using RunSVD, and components 2 through 30 were used to calculate UMAPs and identify clusters. We used the geneactivity function to generate RNA activity assays and the chromVAR function to generate the motif assay. Differentially accessible peaks, RNA activity and motif activity were identified using FindAllMarkers or FindMarkers. We used the AddModuleScore function to calculate enrichment of the open chromatin signature of Asxl1-KO stem-like T cells. The number of correlated genes was determined using the dorcJPlot function from FigR (v0.1.0).
Data analysis for ChIP–seq
We analyzed the ChIP–seq data as previously described14. We mapped the ChIP–seq reads to the mouse genome (mm10) with Bowtie 1.1.1. We called peaks with MACS (v1.4.2; default P value threshold of 1 × 10−4). We used HOMER version 4.9 for peak annotation and transcription factor motif enrichment. We visualized transcription factor binding profiles using Integrative Genomics Viewer.
Statistical analysis
For statistical analysis, Prism (GraphPad, v10.4.0) and R (v4.1.3) were used. A two-tailed Student’s t-test was used for calculating the statistical significance between two experimental conditions. Data distribution was assumed to be normal, but this was not formally tested. A one-way or two-way ANOVA was used for determining statistical significance among more than two experimental groups. A P value of <0.05 was considered statistically significant. No statistical methods were used to predetermine sample sizes, but our sample sizes are similar to those reported in previous publications16,38,39. Mice used for all experiments were randomly assigned to each group. Data collection and analysis were not performed blind to the conditions of the experiments. No data points were excluded from the statistical analyses.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNA-seq and scATAC-seq + scRNA-seq data have been deposited in the Gene Expression Omnibus under the accession code GSE283635. Raw and processed data for scRNA-seq of axicabtagene ciloleucel infusion products from individuals with relapsed or refractory large B cell lymphoma are deposited under the accession code GSE241783, and ATAC-seq data from control and ASXL1-deficient stem-like P14 CD8+ T cells have been published elsewhere and are under accession code GSE274863 (refs. 50,51). Source data are provided with this paper.
References
June, C. H. & Sadelain, M. Chimeric antigen receptor therapy. N. Engl. J. Med. 379, 64–73 (2018).
Brown, C. E. & Mackall, C. L. CAR T cell therapy: inroads to response and resistance. Nat. Rev. Immunol. 19, 73–74 (2019).
Shah, N. N. & Fry, T. J. Mechanisms of resistance to CAR T cell therapy. Nat. Rev. Clin. Oncol. 16, 372–385 (2019).
Martinez, M. & Moon, E. K. CAR T cells for solid tumors: new strategies for finding, infiltrating, and surviving in the tumor microenvironment. Front. Immunol. 10, 128 (2019).
Gumber, D. & Wang, L. D. Improving CAR-T immunotherapy: overcoming the challenges of T cell exhaustion. eBioMedicine 77, 103941 (2022).
Lynn, R. C. et al. c-Jun overexpression in CAR T cells induces exhaustion resistance. Nature 576, 293–300 (2019).
Long, A. H. et al. 4-1BB costimulation ameliorates T cell exhaustion induced by tonic signaling of chimeric antigen receptors. Nat. Med. 21, 581–590 (2015).
Finney, O. C. et al. CD19 CAR T cell product and disease attributes predict leukemia remission durability. J. Clin. Invest. 129, 2123–2132 (2019).
Chen, G. M. et al. Integrative bulk and single-cell profiling of premanufacture T-cell populations reveals factors mediating long-term persistence of CAR T-cell therapy. Cancer Discov. 11, 2186–2199 (2021).
Fraietta, J. A. et al. Determinants of response and resistance to CD19 chimeric antigen receptor (CAR) T cell therapy of chronic lymphocytic leukemia. Nat. Med. 24, 563–571 (2018).
Deng, Q. et al. Characteristics of anti-CD19 CAR T cell infusion products associated with efficacy and toxicity in patients with large B cell lymphomas. Nat. Med. 26, 1878–1887 (2020).
Zebley, C. C. et al. CD19-CAR T cells undergo exhaustion DNA methylation programming in patients with acute lymphoblastic leukemia. Cell Rep. 37, 110079 (2021).
Wu, T. et al. The TCF1–BCL6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness. Sci. Immunol. 1, eaai8593 (2016).
Yao, C. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8+ T cell persistence in chronic infection. Nat. Immunol. 20, 890–901 (2019).
Yao, C. et al. BACH2 enforces the transcriptional and epigenetic programs of stem-like CD8+ T cells. Nat. Immunol. 22, 370–380 (2021).
Zhu, Z. et al. FOXP1 and KLF2 reciprocally regulate checkpoints of stem-like to effector transition in CAR T cells. Nat. Immunol. 25, 117–128 (2024).
Huang, Z. et al. IL-27 promotes the expansion of self-renewing CD8+ T cells in persistent viral infection. J. Exp. Med. 216, 1791–1808 (2019).
Ji, Y. et al. miR-155 harnesses Phf19 to potentiate cancer immunotherapy through epigenetic reprogramming of CD8+ T cell fate. Nat. Commun. 10, 2157 (2019).
He, R. et al. Follicular CXCR5-expressing CD8+ T cells curtail chronic viral infection. Nature 537, 412–428 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Leong, Y. A. et al. CXCR5+ follicular cytotoxic T cells control viral infection in B cell follicles. Nat. Immunol. 17, 1187–1196 (2016).
Ferrando-Martinez, S. et al. Accumulation of follicular CD8+ T cells in pathogenic SIV infection. J. Clin. Invest. 128, 2089–2103 (2018).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Siddiqui, I. et al. Intratumoral TCF1+PD-1+CD8+ T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211 (2019).
Kurtulus, S. et al. Checkpoint blockade immunotherapy induces dynamic changes in PD-1−CD8+ tumor-infiltrating T cells. Immunity 50, 181–194 (2019).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8+ T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013 (2018).
Sabatino, M. et al. Generation of clinical-grade CD19-specific CAR-modified CD8+ memory stem cells for the treatment of human B-cell malignancies. Blood 128, 519–528 (2016).
van Bruggen, J. A. C. et al. Chronic lymphocytic leukemia cells impair mitochondrial fitness in CD8+ T cells and impede CAR T-cell efficacy. Blood 134, 44–58 (2019).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8+ T cell exhaustion. Nature 571, 211–218 (2019).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Seo, H. et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8+ T cell exhaustion. Proc. Natl Acad. Sci. USA 116, 12410–12415 (2019).
Chan, J. D. et al. FOXO1 enhances CAR T cell stemness, metabolic fitness and efficacy. Nature 629, 201–210 (2024).
Doan, A. E. et al. FOXO1 is a master regulator of memory programming in CAR T cells. Nature 629, 211–218 (2024).
Tsui, C. et al. MYB orchestrates T cell exhaustion and response to checkpoint inhibition. Nature 609, 354–360 (2022).
Gautam, S. et al. The transcription factor c-Myb regulates CD8+ T cell stemness and antitumor immunity. Nat. Immunol. 20, 337–349 (2019).
Zhu, Z. et al. The redox sensor KEAP1 facilitates adaptation of T cells to chronic antigen stimulation by preventing hyperactivation. Sci. Immunol. 9, eadk2954 (2024).
Zhu, Z. et al. Aging compromises terminal differentiation program of cytotoxic effector lineage and promotes exhaustion in CD8+ T cells responding to coronavirus infection. Aging Cell 24, e70109 (2025).
Fagerberg, E. et al. KLF2 maintains lineage fidelity and suppresses CD8 T cell exhaustion during acute LCMV infection. Science 387, eadn2337 (2025).
Tsui, C. et al. Lymph nodes fuel KLF2-dependent effector CD8+ T cell differentiation during chronic infection and checkpoint blockade. Nat. Immunol. 26, 1752–1765 (2025).
Huang, Q. et al. The primordial differentiation of tumor-specific memory CD8+ T cells as bona fide responders to PD-1/PD-L1 blockade in draining lymph nodes. Cell 185, 4049–4066 (2022).
Beltra, J. C. et al. Developmental relationships of four exhausted CD8+ T cell subsets reveals underlying transcriptional and epigenetic landscape control mechanisms. Immunity 52, 825–841 (2020).
Heuser, C. & Gattinoni, L. c-Myb redefines the hierarchy of stem-like T cells. Nat. Immunol. 23, 1405–1407 (2022).
Garcia, J. et al. Naturally occurring T cell mutations enhance engineered T cell therapies. Nature 626, 626–634 (2024).
Jacoby, E. et al. CD19 CAR immune pressure induces B-precursor acute lymphoblastic leukaemia lineage switch exposing inherent leukaemic plasticity. Nat. Commun. 7, 12320 (2016).
Yang, Y. et al. TCR engagement negatively affects CD8 but not CD4 CAR T cell expansion and leukemic clearance. Sci. Transl. Med. 9, eaag1209 (2017).
Utzschneider, D. T. et al. Early precursor T cells establish and propagate T cell exhaustion in chronic infection. Nat. Immunol. 21, 1256–1266 (2020).
Galletti, G. et al. Two subsets of stem-like CD8+ memory T cell progenitors with distinct fate commitments in humans. Nat. Immunol. 21, 1552–1562 (2020).
Kang, T. G. et al. Epigenetic regulators of clonal hematopoiesis control CD8 T cell stemness during immunotherapy. Science 386, eadl4492 (2024).
Li, X. et al. A single-cell atlas of CD19 chimeric antigen receptor T cells. Cancer Cell 41, 1835–1837 (2023).
Bai, Z. et al. Single-cell CAR T atlas reveals type 2 function in 8-year leukaemia remission. Nature 634, 702–711 (2024).
Feng, B. et al. The type 2 cytokine Fc-IL-4 revitalizes exhausted CD8+ T cells against cancer. Nature 634, 712–720 (2024).
Good, C. R. et al. An NK-like CAR T cell transition in CAR T cell dysfunction. Cell 184, 6081–6100 (2021).
Giles, J. R. et al. Shared and distinct biological circuits in effector, memory and exhausted CD8+ T cells revealed by temporal single-cell transcriptomics and epigenetics. Nat. Immunol. 23, 1600–1613 (2022).
Kartha, V. K. et al. Functional inference of gene regulation using single-cell multi-omics. Cell Genom. 2, 100166 (2022).
Banaszynski, L. A., Chen, L. C., Maynard-Smith, L. A., Ooi, A. G. & Wandless, T. J. A rapid, reversible, and tunable method to regulate protein function in living cells using synthetic small molecules. Cell 126, 995–1004 (2006).
Aiken, T. J. et al. Mechanism of effective combination radio-immunotherapy against 9464D-GD2, an immunologically cold murine neuroblastoma. J. Immunother. Cancer 10, e004834 (2022).
Larson, S. M. et al. CD19/CD20 bispecific chimeric antigen receptor (CAR) in naive/memory T cells for the treatment of relapsed or refractory non-Hodgkin lymphoma. Cancer Discov. 13, 580–597 (2023).
Hu, B. et al. Augmentation of antitumor immunity by human and mouse CAR T cells secreting IL-18. Cell Rep. 20, 3025–3033 (2017).
Swaminathan, S. et al. BACH2 mediates negative selection and p53-dependent tumor suppression at the pre-B cell receptor checkpoint. Nat. Med. 19, 1014–1022 (2013).
Schmidl, C., Rendeiro, A. F., Sheffield, N. C. & Bock, C. ChIPmentation: fast, robust, low-input ChIP–seq for histones and transcription factors. Nat. Methods 12, 963–965 (2015).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Acknowledgements
We thank A. Guzman and A. Mobley at the University of Texas Southwestern Medical Center Flow and Mass Cytometry Facility for outstanding support. We thank T. Fry (University of Colorado) for providing the MSCV-mCD19-CD28z-tEGFR plasmid and E2A–PBX1 cell line, C. June (University of Pennsylvania) for sharing the B16-CD19 cell line, C. Mackall (Stanford University) for sharing the MSGV-HA-28z (GD2 CAR) plasmid and P. Sondel (University of Wisconsin) for sharing the 9464D-GD2 cell line. We thank the following funding support: AI158294, AG083398 and AG056524 from National Institutes of Health, Clinic & Laboratory Integration Program and Lloyd J. Old STAR Program from the Cancer Research Institute, V Scholar Award from the V Foundation, Grant for Junior Faculty from the American Federation for Aging Research (AFAR) and a New Investigator Award from Hevolution/AFAR to T.W. and AI154450 from the National Institutes of Health, RR210035 and RP250282 from Cancer Prevention and Research Institute of Texas and HT94252310801 from the Department of Defense to C.Y. The funders had no influence on the design of this study, data analysis or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Study design: T.W. and C.Y. Methodology: Z.Z., T.H., Y.L., S.W., C.Y. and T.W. Investigation: Z.Z., T.H., Y.L., S.W., J.H., S.S.S., K.Y., C.Y. and T.W. Data curation: Z.Z., T.H., Y.L., S.W., C.Y. and T.W. Writing and editing: T.W. and C.Y. Visualization: T.H. and C.Y. Funding acquisition: T.W. and C.Y. Supervision: T.W. and C.Y.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Nick Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 scATAC-seq+scRNA-seq analysis of pre-infusion CAR T cells.
a, Gating strategy for sorting in vitro cultured anti-murine CD19 CAR T cells. b, c, scATAC-seq+scRNA-seq analysis was performed as in Fig. 1. b, Violin plots showing the mRNA levels of selected genes in each transcriptome-defined cluster in Fig. 1a. c, A feature plot (left) and violin plot (right) showing the enrichment of cell-cycle gene signature in pre-infusion CD19 CAR T cells. d, Single-cell enrichment of the open chromatin signature associated with Asxl1 KO long-lived stem-like T cells (GSE274863) in LT stem-like CAR T cells (ATAC_2) versus transitory stem-like CAR T cells (ATAC_0) defined in Fig. 1d. The P value was calculated using a two-sided Student’s t-test.
Extended Data Fig. 2 LT stem-like pre-infusion CAR T cells upregulate BACH2.
a, Single-cell expression of Sell in murine pre-infusion CD8+ CD19 CAR T subsets from the experiment in Fig. 1. b, Schematic illustration of the experiment in Fig. 2a, b. c–e, Experiment was set up as in b. On day 7 after infusion, numbers of CD8+ CAR T cells in the spleen, bone marrow, liver and lung (c), frequencies of active caspase-3+ CD8+ CAR T cells (d), and levels of PD1, TIM3 and TOX in CD8+ CAR T cells (e) were determined in E2A–PBX1-bearing mice treated with CD62L+ (n = 9 mice) or CD62L− (n = 6 mice) CD19 CAR T cells. Bar graphs represent the Mean ± SEM. Circles in the bar graphs represent individual mice. f, Tumor growth in mice bearing 9464D-GD2 neuroblastoma after treatment of CD62L+ (n = 5 mice) or CD62L− (n = 5 mice) CD8+ GD2 CAR T cells. Data are presented as Mean ± SEM. g–l, Re-analyzed published scRNA-seq data (GSE241783) of pre-infusion human CD19 CAR T cells from 40 relapsed/refractory B-cell lymphoma patients. g, h, UMAP plots of total (CD4+ and CD8+) pre-infusion human CAR T cells color-coded based on cluster IDs (g) or patient IDs (h). i, Single-cell expression of CD4 (upper) and CD8A (lower). j, Single-cell expression of selected genes in each cluster of CD8+ pre-infusion human CD19 CAR T cells. Clusters are defined in Fig. 2c. k, Single-cell trajectory plot showing the lineage relationships among T cell subsets. l, Expression of BACH2 and LT stem-like gene signature along the pseudotime from the stem-like cluster (Tstem-like) to the effector-like (Teff-like) cluster. Statistical significance in c–e was calculated with a two-sided Student’s t-test. Statistical significance in f was determined by two-way ANOVA. Data in c–f are representative of two independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
Extended Data Fig. 3 scATAC-seq+scRNA-seq of CAR T cells after leukemia clearance.
a–e, scATAC-seq+scRNA-seq experiment is described in Fig. 3. a, Single-cell expression of selected genes in each transcriptome-defined cluster. b, A volcano plot of differentially expressed (DE) genes between stem-like and LT stem-like CAR T cells. Adjusted P values (adj P) were calculated using a two-sided Wilcoxon rank-sum test with Bonferroni correction to account for multiple comparisons. c, The protein level of TOX in stem-like (CD62Lint PD1+) and LT stem-like (CD62Lhi PD1−) CD8+ CD19 CAR T cells from E2A-PBX1 inoculated mice (n = 6 mice) on day 20 after CAR T-cell infusion. Circles represent individual mice. Statistical significance was calculated with a two-sided paired Student’s t-test. Data are representative of two independent experiments. *P < 0.05. d, Genes in CD8+ CD19 CAR T cells are ranked by the number of correlated scATAC-seq peaks. Genes with more than five correlated peaks are highlighted in red. e, Transcription factors (TFs) in CD8+ CD19 CAR T cells are ranked by the number of target genes. f, Single-cell enrichment of the open chromatin signature associated with Asxl1 KO long-lived stem-like T cells (GSE274863) in LT stem-like CAR T cells (ATAC_3) versus stem-like CAR T cells (ATAC_0) as defined in Fig. 3e. The P value was calculated using a two-sided Student’s t-test.
Extended Data Fig. 4 Antitumor responses of stem-like CAR T cells during rechallenge.
a–e, Experimental setup is described in Fig. 4. a, The numbers of progenies from Tcf7-GFP+ or Tcf7-GFP− CD8+ CAR T cells in the bone marrow (n = 5). b, The frequencies of bone marrow Tcf7-GFPhighTIM3low CD8+ CAR T cells derived from Tcf7-GFP+ or Tcf7-GFP− donors (n = 5). c, The frequencies of bone marrow CD62LhighTIM3low CD8+ CAR T cells derived from Tcf7-GFP+ or Tcf7-GFP− donors (n = 5). d, e, Expression of PD1 (d) and TIM3 (e) in bone marrow CD8+ CAR T cells derived from Tcf7-GFP+ or Tcf7-GFP− donors (n = 5). f–i, CD19 CAR T cells co-expressing a truncated human EGFR (huEGFR) were generated from a Tcf7-GFP reporter mouse and infused into E2A-PBX1-bearing mice. On day 24 post-infusion, CD62L+Tcf7-GFP+ and CD62L−Tcf7-GFP− CD8+ CAR T cells were sorted and transferred separately to a second batch of E2A-PBX1-bearing mice. Recall responses by CAR T cells were analyzed on day 7 post-infusion. f, Representative flow plots (left) and numbers (right) of CD8+ CAR T cells (huEGFR+CD45.1+) derived from CD62L+Tcf7-GFP+ or CD62L−Tcf7-GFP− CD8+ CAR T cells (n = 5). g, Representative flow plots (left) and frequencies (right) of CD62L+CX3CR1− stem-like CD8+ CAR T cells derived from CD62L+Tcf7-GFP+ or CD62L−Tcf7-GFP− donors (n = 5). h, i, Expression of PD1 (h) and TIM3 (i) in progenies derived from CD62L+Tcf7-GFP+ or CD62L−Tcf7-GFP− donors (n = 5). Data are representative of two independent experiments. n, number of mice per group. Bar graphs represent the Mean ± SEM. Circles in the bar graphs represent individual mice. Statistical significance was calculated with a two-sided Student’s t-test. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
Extended Data Fig. 5 BACH2 deficiency impairs antitumor responses by CAR T cells.
a–f, B16-CD19-bearing mice treated with control or Bach2 iKO CD19 CAR T cells were examined on day 8 post-infusion (n = 4 mice per group). a, Representative flow plots and numbers of splenic CD8+ CAR T cells. b, Numbers of splenic CD4+ CAR T cells. c, Representative flow plots and numbers of tumor-infiltrating CD8+ CAR T cells. d, Numbers of tumor-infiltrating CD4+ CAR T cells. e, f, TCF1 (e) and CD62L (f) expression in control and Bach2 iKO CD8+ CAR T cells. g, UMAP plots of control or Bach2 iKO CD8+ CAR T cells from different mice (n = 3 mice per group). h, Volcano plots of differentially expressed genes between control stem-like (cluster 1) and Bach2 iKO stem-like (cluster 1) CAR T cells. Adjusted P values (adj P) were calculated using a two-sided Wilcoxon rank-sum test with Bonferroni correction for multiple comparisons. i, Frequencies of genomic features containing BACH2 binding sites revealed by BACH2 ChIP-seq. j, Transcription factor binding motifs enriched at BACH2 binding sites determined with HOMER (findMotifsGenome.pl) using BACH2 ChIP-seq data. k, Histogram of JunB binding centered around BACH2 binding sites that exhibited differential JunB binding in control versus BACH2 OE CD8+ CAR T cells revealed by JunB ChIP-seq. P-value = 1.193e-05. l, Representative flow plots and frequencies of the stem-like subset in control (n = 7 mice) and Junb gRNA (n = 8 mice) construct-transduced CD8+ CD19 CAR T cells in E2A-PBX1-bearing mice on day 7 post-infusion. m–p, Control and Bach2 iKO CD8+ CD19 CAR T cells from E2A-PBX1-bearing mice on day 20 post-infusion were transferred separately into a second cohort of E2A-PBX1-bearing mice (n = 5 mice per group) and analyzed on day 7 post-transfer. Numbers (m), frequencies of the stem-like subset (n), and expression of PD1 (o) and TIM3 (p) of splenic control and Bach2 iKO CD8+ CD19 CAR T cells are shown. Data in a–f, l–p are representative of two independent experiments. Bar graphs represent the Mean ± SEM. Circles represent individual mice. Statistical significance in a–f, l–p was calculated with a two-sided Student’s t-test. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
Extended Data Fig. 6 Control the protein level of DD-BACH2 through Shield-1.
a, Mouse CD8+ T cells were transduced with a pMIG plasmid (control), a pMIG-BACH2 plasmid (BACH2 OE), or a pMIG-DD-BACH2 plasmid (DD-BACH2). DD-BACH2 CD8+ T cells were cultured with various concentrations of Shield-1. Western blots were used to assess the protein level of BACH2. b, The protein level of BACH2 in mouse CD8+ T cells transduced with a pMIG-DD-BACH2 construct that were cultured with or without 1uM Shield-1 or cultured with 1uM Shield-1 before removing Shield-1 from culture for indicated number of days. c–e, Frequencies of CD62L+ cells (c) and expression of PD1 (d) and TIM3 (e) in mouse CD8+ GD2 CAR T cells co-transduced with a pMIG-DD-BACH2 construct that were continuously cultured with or without 1uM Shield-1 or cultured with 1uM Shield-1 followed by withdrawal of Shield-1 for two days (n = 4 independent cultures per group). Statistical significance was calculated with one-way ANOVA with Šídák’s multiple comparisons test. f–i, Human GD2 CAR T cells were co-transduced with a pMIG plasmid (control) or a pMIG-DD-BACH2 plasmid. DD-BACH2 GD2 CAR T cells were cultured with or without 1uM Shield-1 (n = 3 independent cultures per group). The protein levels of CCR7 (f), CD127 (g), TIM3 (h) and LAG3 (i) in control human CD8+ GD2 CAR T cells or DD-BACH2 human CD8+ GD2 CAR T cells cultured with or without Shield-1 are shown. Statistical significance was calculated with one-way ANOVA with Tukey’s multiple comparisons test. Data in a–i are representative of two independent experiments. Bar graphs represent the Mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.0001.
Supplementary information
Source data
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download JPG )
Unprocessed gel images.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hu, T., Zhu, Z., Luo, Y. et al. BACH2 dosage establishes the hierarchy of stemness and fine-tunes antitumor immunity in CAR T cells. Nat Immunol 27, 425–435 (2026). https://doi.org/10.1038/s41590-025-02388-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02388-0
This article is cited by
-
BACH2 the future
Nature Immunology (2026)