Abstract
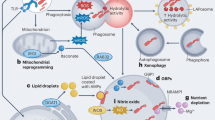
Pathogen-associated molecules can have both membrane-associated and intracellular receptors. Bacterial lipoproteins are recognized by Toll-like receptor 2, but it is unclear whether they can also be sensed by cytoplasmic receptors. Here we found that bacterial lipoproteins could be recognized in the cytoplasm of macrophages by cystathionine γ-lyase (CTH) and hydrolyzed into lipid chains containing sulfhydryl groups. The hydrolyzed lipid chains form molecules containing four acylated chains linked through disulfide bonds, which further cleave caspase-11 and activate the noncanonical inflammasome. Changing the redox environment in macrophages affects their recognition of bacterial lipoproteins. CTH-deficient primary and immortalized macrophages do not trigger activation of the noncanonical inflammasome in the presence of intracellular bacterial lipoproteins, while CTH-deficient mice exhibit attenuated immune responses to infection with Staphylococcus aureus and Listeria monocytogenes. Our findings elucidate the molecular mechanisms by which macrophages recognize intracellular bacterial lipoproteins, as well as the regulatory relationship between cellular redox levels and infection resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data are shown in the main figures and Extended Data files. The data supporting the findings of this study are available within the article and its supplementary materials. Source data are provided with this paper.
Code availability
This paper does not report original code.
References
Kawai, T., Ikegawa, M., Ori, D. & Akira, S. Decoding Toll-like receptors: recent insights and perspectives in innate immunity. Immunity 57, 649–673 (2024).
Wolf, A. J. & Underhill, D. M. Peptidoglycan recognition by the innate immune system. Nat. Rev. Immunol. 18, 243–254 (2018).
Philpott, D. J., Sorbara, M. T., Robertson, S. J., Croitoru, K. & Girardin, S. E. NOD proteins: regulators of inflammation in health and disease. Nat. Rev. Immunol. 14, 9–23 (2014).
Zhu, F. et al. The orphan receptor Nur77 binds cytoplasmic LPS to activate the non-canonical NLRP3 inflammasome. Immunity 56, 753–767 (2023).
Ryu, J.-K. et al. Reconstruction of LPS transfer cascade reveals structural determinants within LBP, CD14, and TLR4–MD2 for efficient LPS recognition and transfer. Immunity 46, 38–50 (2017).
Liu, Q. et al. Eukaryotic ADCY7 catalyzes the production of c-di-AMP to activate the NLRP3 inflammasome. Nat. Chem. Biol. 21, 1283–1291 (2025).
Mangan, M. S. J. et al. Targeting the NLRP3 inflammasome in inflammatory diseases. Nat. Rev. Drug Discov. 17, 588–606 (2018).
Swanson, K. V., Deng, M. & Ting, J. P. The NLRP3 inflammasome: molecular activation and regulation to therapeutics. Nat. Rev. Immunol. 19, 477–489 (2019).
He, W. et al. Gasdermin D is an executor of pyroptosis and required for interleukin-1β secretion. Cell Res. 25, 1285–1298 (2015).
Shi, J. et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature 526, 660–665 (2015).
Kayagaki, N. et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature 526, 666–671 (2015).
Hagar, J. A., Powell, D. A., Aachoui, Y., Ernst, R. K. & Miao, E. A. Cytoplasmic LPS activates caspase-11: implications in TLR4-independent endotoxic shock. Science 341, 1250–1253 (2013).
Kayagaki, N. et al. Noncanonical inflammasome activation by intracellular LPS independent of TLR4. Science 341, 1246–1249 (2013).
Lagrange, B. et al. Human caspase-4 detects tetra-acylated LPS and cytosolic Francisella and functions differently from murine caspase-11. Nat. Commun. 9, 242 (2018).
Shi, J. et al. Inflammatory caspases are innate immune receptors for intracellular LPS. Nature 514, 187–192 (2014).
Kieser, K. J. & Kagan, J. C. Multi-receptor detection of individual bacterial products by the innate immune system. Nat. Rev. Immunol. 17, 376–390 (2017).
McWhirter, S. M. & Jefferies, C. A. Nucleic acid sensors as therapeutic targets for human disease. Immunity 53, 78–97 (2020).
Buddelmeijer, N. The molecular mechanism of bacterial lipoprotein modification—how, when and why? FEMS Microbiol. Rev. 39, 246–261 (2015).
Wiktor, M. et al. Structural insights into the mechanism of the membrane integral N-acyltransferase step in bacterial lipoprotein synthesis. Nat. Commun. 8, 15952 (2017).
Paul, B. D., Sbodio, J. I. & Snyder, S. H. Cysteine metabolism in neuronal redox homeostasis. Trends Pharmacol. Sci. 39, 513–524 (2018).
Kolluru, G. K., Shackelford, R. E., Shen, X., Dominic, P. & Kevil, C. G. Sulfide regulation of cardiovascular function in health and disease. Nat. Rev. Cardiol. 20, 109–125 (2023).
McBean, G. J., Aslan, M., Griffiths, H. R. & Torrão, R. C. Thiol redox homeostasis in neurodegenerative disease. Redox Biol. 5, 186–194 (2015).
Zhang, H.-F., Klein Geltink, R. I., Parker, S. J. & Sorensen, P. H. Transsulfuration, minor player or crucial for cysteine homeostasis in cancer. Trends Cell Biol. 32, 800–814 (2022).
Sartorio, M. G., Pardue, E. J., Feldman, M. F. & Haurat, M. F. Bacterial outer membrane vesicles: from discovery to applications. Annu. Rev. Microbiol. 75, 609–630 (2021).
Nguyen, M.-T., Matsuo, M., Niemann, S., Herrmann, M. & Götz, F. Lipoproteins in gram-positive bacteria: abundance, function, fitness. Front. Microbiol. 11, 582582 (2020).
Smithers, L., Olatunji, S. & Caffrey, M. Bacterial lipoprotein posttranslational modifications. New insights and opportunities for antibiotic and vaccine development. Front. Microbiol. 12, 788445 (2021).
Craven, R. R. et al. Staphylococcus aureus α-hemolysin activates the NLRP3-inflammasome in human and mouse monocytic cells. PLoS ONE 4, e7446 (2009).
Vande Walle, L. & Lamkanfi, M. Inflammasomes: caspase-1-activating platforms with critical roles in host defense. Front. Microbiol. 2, 3 (2011).
Beam, J. E. et al. Inflammasome-mediated glucose limitation induces antibiotic tolerance in Staphylococcus aureus. iScience 26, 107942 (2023).
Meunier, E. et al. Caspase-11 activation requires lysis of pathogen-containing vacuoles by IFN-induced GTPases. Nature 509, 366–370 (2014).
Goers, L. et al. Shigella IpaH9.8 limits GBP1-dependent LPS release from intracytosolic bacteria to suppress caspase-4 activation. Proc. Natl Acad. Sci. USA 120, e2218469120 (2023).
Bowran, K. & Palmer, T. Extreme genetic diversity in the type VII secretion system of Listeria monocytogenes suggests a role in bacterial antagonism. Microbiology (Reading) 167, mic.0.001034 (2021).
Ghosh, S. et al. Host AAA-ATPase VCP/p97 lyses ubiquitinated intracellular bacteria as an innate antimicrobial defence. Nat. Microbiol. 10, 1099–1114 (2025).
Korea, C. G. et al. Staphylococcal Esx proteins modulate apoptosis and release of intracellular Staphylococcus aureus during infection in epithelial cells. Infect. Immun. 82, 4144–4153 (2014).
Cao, X. Self-regulation and cross-regulation of pattern-recognition receptor signalling in health and disease. Nat. Rev. Immunol. 16, 35–50 (2016).
Man, S. M. & Jenkins, B. J. Context-dependent functions of pattern recognition receptors in cancer. Nat. Rev. Cancer 22, 397–413 (2022).
Liu, Q. et al. The TET3 inflammasome senses unique long HSV-1 proteins for virus particle budding from the nucleus. Cell. Mol. Immunol. 21, 1322–1334 (2024).
Li, W. et al. Adipose triglyceride lipase suppresses noncanonical inflammasome by hydrolyzing LPS. Nat. Chem. Biol. 20, 1434–1442 (2024).
Hara, H. et al. The NLRP6 inflammasome recognizes lipoteichoic acid and regulates gram-positive pathogen infection. Cell 175, 1651–1664 (2018).
Man, S. M. et al. IRGB10 liberates bacterial ligands for sensing by the AIM2 and caspase-11–NLRP3 inflammasomes. Cell 167, 382–396 (2016).
Fernández-Rodríguez, C. et al. Structural basis of the inhibition of cystathionine γ-lyase from Toxoplasma gondii by propargylglycine and cysteine. Protein Sci. 32, e4619 (2023).
Ishii, I. et al. Cystathionine γ-lyase-deficient mice require dietary cysteine to protect against acute lethal myopathy and oxidative injury. J. Biol. Chem. 285, 26358–26368 (2010).
Liu, Q. et al. Identification and application of a panel of constitutive promoters for gene overexpression in Staphylococcus aureus. Front. Microbiol. 13, 818307 (2022).
Acknowledgements
We thank Y. Chen and F. Shao for their contributions in providing the cell lines used in this study. This work was supported by the National Natural Science Foundation of China (92369104, 82271790, 22174003, 92569301, 82595923 and U25A20654), the Beijing Natural Science Foundation (JG23028), the National Key R&D Program of China (2022YFC2302900, 2022YFA1304500 and 2021YFA1300202), Strategic Priority Research Programs of the Chinese Academy of Sciences (XDB1470000), the CAS Project for Young Scientists in Basic Research (YSBR-010) and the Fok Ying Tung Education Foundation (171015) to P.X.
Author information
Authors and Affiliations
Contributions
Q.L., C.W. and M.L. performed experiments and analyzed data; C.K., Z.Z., X.C., X.G., Z.L., C.Z., D.J., X.S. and S.W. performed experiments and analyzed data; P.X. initiated the study, designed and performed experiments, analyzed data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Anirban Banerjee, Rongbin Zhou and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Ioana Staicu, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Prokaryotic lipoproteins activate the non-canonical inflammasome.
a, Coomassie blue staining of lipoproteins purified from E. coli and S. aureus. b, Limulus Amebocyte Lysate (LAL) assay for endotoxin contamination in E. coli lipoproteins crude or LPS removal (–LPS). c, LAL assay comparing endotoxin levels in E. coli and S. aureus lipoprotein. SA-Lpp, S. aureus lipoprotein. d, Immunoblots of BMDMs from Nlrp3+/+ and Nlrp3−/− mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with cell-penetrating peptides (100 ng/μl) alone or together with SA-Lpp (100 ng/μl) for 16 h. Sup., supernatant; SA-Lpp, S. aureus lipoprotein. e, Quantification of pyroptosis in BMDMs from Nlrp3+/+ and Nlrp3−/− mice treated as in d. ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. CPP, cell-penetrating peptides; SA-Lpp, S. aureus lipoprotein. f, Immunoblots of BMDMs from wild-type mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with cell-penetrating peptides (100 ng/μl) alone or together with SA-Lpp for 16 h, SA-Lpp cloned, expressed in indicated host and purified. g-i, Quantification of pyroptosis in BMDMs from wild-type mice treated as in f. ELISA assays of IL-1β in supernatants (g), LDH release assays of cell death in supernatants (h) and ATP assays of cell viability in cell pellets (i). In b, c, e and g-i, data are pooled from three independent experiments (n = 3) and shown as means ± SD. Unpaired two-tailed Student’s t-test was used. Exact P values are indicated in the figures. Data are representative of three independent experiments with similar results in a-i.
Extended Data Fig. 2 Bacterial lipoproteins activate the non-canonical inflammasome in a CTH-dependent manner.
a, Quantification of pyroptosis in iBMDMs primed with 1 μg/ml Pam3CSK4 overnight before transfected with cell-penetrating peptides (100 ng/μl) alone or together with SA-Lpp (100 ng/μl) for 16 h. iBMDMs infected with lentiviruses carrying sgRNAs against indicated genes. ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. b, Immunoblots of Cth−/− iBMDMs rescued with CTHWT and CTHK211R, then primed with 1 μg/ml Pam3CSK4 overnight before transfected with cell-penetrating peptides (100 ng/μl) alone or together with SA-Lpp (100 ng/μl) for 16 h. Sup., supernatant; SA-Lpp, S. aureus lipoprotein. c, Quantification of cell death and cytokine secretion in Cth−/− iBMDMs rescued with CTHWT and CTHK211R, then treated as in b. ELISA of IL-1β and IL-18 in supernatants, LDH release of cell death in supernatants, ATP assays of cell viability in cell pellets, ELISA of TNF-α in supernatants, ELISA of IL-6 in supernatants. CPP, cell-penetrating peptides; SA-Lpp, S. aureus lipoprotein. d, Quantification of cell death and cytokine secretion in BMDMs from Cth+/+ and Cth−/− mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with cell-penetrating peptides (100 ng/μl) alone or together with SA-Lpp (100 ng/μl) for 16 h. ATP assays of cell viability in cell pellets, ELISA assays of TNF-α and IL-6 in supernatants. CPP, cell-penetrating peptides; SA-Lpp, S. aureus lipoprotein. e, Quantification of cell death and cytokine secretion in BMDMs from Cth+/+ and Cth−/− mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with DOTAP alone or together with 10 μg/ml Pam3CSK4 for 16 h. ATP assays of cell viability in cell pellets, ELISA assays of TNF-α and IL-6 in supernatants. In c-e, data are pooled from three independent experiments (n = 3) and shown as means ± SD. Unpaired two-tailed Student’s t-test was used. Exact P values are indicated in the figures. Data are representative of three independent experiments with similar results in a-e.
Extended Data Fig. 3 CTH binds bacterial lipoproteins.
a, Immunoblots of BMDMs from wild-type mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with DOTAP alone or together with 200 ng/ml LPS or 10 μg/ml Pam3CSK4, or transfected with cell penetrating peptides alone or together with SA-Lpp (100 ng/μl) for 16 h. Sup., supernatant; SA-Lpp, S. aureus lipoprotein. b, Immunoblots of BMDMs from Cth+/+ and Cth−/− mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with 200 ng/ml LPS for 16 h. Otherwise, cells primed with 1 μg/ml LPS for 3 h then stimulated with 2 mM ATP for 30 min or 2 μg/ml poly(dA:dT) for 4 h. Sup., supernatant. c, RT-PCR analysis of indicated gene in BMDMs from Cth+/+ and Cth−/− mice primed with 1 μg/ml Pam3CSK4 overnight. d, Immunoblots of BMDMs from Tlr2flox/flox (Tlr2+/+) and Tlr2flox/flox; Lyz2-Cre (Tlr2−/−) mice primed with 10 μg/ml poly(I:C) overnight before transfected with DOTAP together with 10 μg/ml Pam3CSK4 for 16 h. Sup., supernatant. e, Quantification of pyroptosis in BMDMs from Tlr2+/+ and Tlr2−/− mice treated as in d. ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. In c and e, data are pooled from three independent experiments (n = 3) and shown as means ± SD. Unpaired two-tailed Student’s t-test was used. Exact P values are indicated in the figures. Data are representative of three independent experiments with similar results in a-e.
Extended Data Fig. 4 CTH binds to Pam3CSK4 and bacterial lipoprotein.
a, Coomassie blue staining of recombinant CTH proteins purified from HEK293F cells. b, Pulldown assay of BMDM lysates from wild-type mice with biotin-Pam3CSK4 or biotin-GFP (control) using streptavidin beads and immunoblotted for CTH. c, Pulldown assay of BMDM lysates from wild-type mice with his-tagged Lpp purified from S. aureus or HEK293F cells. NLRP3 fragment (aa 600–920) as a control. Data are representative of three independent experiments with similar results in a-c.
Extended Data Fig. 5 Lipid chains are removed from bacterial lipoproteins by CTH.
a, Immunoblots of his-tagged SA-Lpp, blocked with maleimide, separated by SDS-PAGE, and transferred to a membrane. Membrane incubated with or without recombinant CTH, then labeled with biotin-maleimide. Sulfhydryl groups detected by anti-biotin immunoblot. SH signal, sulfhydryl groups; SA-Lpp, S. aureus lipoprotein. b, Immunoblots of his-tagged SA-Lpp, blocked with maleimide, then reacted with CTH (solution). Reaction products separated by SDS-PAGE and SH signal immunoblotted with anti-biotin. SA-Lpp, S. aureus lipoprotein. c, Biochemical identification of Pam3CSK4 degradation products, followed Pam3CSK4 incubated with or without recombinant CTH, analyzed by HPLC. Eluted fractions analyzed by mass spectrometry. d, Immunoblots of BMDMs from wild-type, Casp11−/− and Casp1−/− mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with DOTAP alone or together with 10 μg/ml Pam3CSK4 for 16 h. Cells cultured in fresh medium containing gentamycin (100 μg/ml) with or without 0.1 mM β-mercaptoethanol (β-ME) for 16 h. e, Quantification of pyroptosis in BMDMs from wild-type, Casp11−/− and Casp1−/− mice treated as in d, ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. f, Immunoblots of BMDMs from wild-type, Casp11−/− and Casp1−/− mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with DOTAP alone or together with 10 μg/ml Pam3CSK4 for 16 h. Cells cultured in fresh medium containing gentamycin (100 μg/ml) with or without 0.1 mM H2O2 for 16 h. g, Quantification of pyroptosis in BMDMs from wild-type, Casp11−/− and Casp1−/− mice treated as in f, ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. In e and g, data are pooled from three independent experiments (n = 3) and shown as means ± SD. Unpaired two-tailed Student’s t-test was used. Exact P values are indicated in the figures. Data are representative of three independent experiments with similar results in a-g.
Extended Data Fig. 6 Sulfhydryl-containing lipid chains are linked by disulfide bonds.
a, Immunoblots of caspase-11 cleavage after incubation with lipid A (50 ng/μl), SDG (10 ng/μl) or DDG (10 ng/μl). b, Immunoblots of BMDMs from Cth+/+ and Cth−/− mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with DOTAP alone or together with 10 μg/ml SDG or DDG for 16 h. c, Surface plasmon resonance analysis of caspase-11 binding to lipid A, SDG or DDG, GST as control. d, Immunoblots of BMDMs from wild-type, Casp11−/− and Casp1−/− mice primed with 1 μg/ml Pam3CSK4 overnight before infected with live S. aureus at an MOI of 10 for 6 h. Cells cultured in fresh medium containing gentamycin (100 μg/ml) with or without 0.1 mM H2O2 for 16 h. e, Quantification of pyroptosis in BMDMs from wild-type, Casp11−/− and Casp1−/− mice treated as in d. ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. f, Quantification of pyroptosis in BMDMs from wild-type, Casp11−/− and Casp1−/− mice primed with 1 μg/ml Pam3CSK4 overnight before infected with live S. aureus at an MOI of 10 for 6 h. Cells cultured in fresh medium containing gentamycin (100 μg/ml) with or without PAG (0.6 mM) for 16 h. LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. In e and f, data are pooled from three independent experiments (n = 3) and shown as means ± SD. Unpaired two-tailed Student’s t-test was used. Exact P values are indicated in the figures. Data are representative of three independent experiments with similar results in a-f.
Extended Data Fig. 7 CTH is essential for sensing cytoplasmic bacterial lipoproteins.
a, Immunoblots of peritoneal macrophage from Cth+/+ and Cth−/− mice primed with 1 μg/ml Pam3CSK4 overnight before infected with live S. aureus at an MOI of 10 for 6 h. Cells cultured in fresh medium containing 100 μg/ml gentamycin. b, Quantification of pyroptosis in peritoneal macrophage from Cth+/+ and Cth−/− mice treated as in a. ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. c, Immunoblots of BMDMs from wild-type and Gbpchr3 mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with DOTAP alone or together with 10 μg/ml Pam3CSK4 for 16 h. d, Quantification of pyroptosis in BMDMs from wild-type and Gbpchr3 mice treated as in c. ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. e, Immunoblots of BMDMs from wild-type and Gbpchr3 mice primed with 1 μg/ml Pam3CSK4 overnight before infected with live S. aureus at an MOI of 10 for 6 h. Cells cultured in fresh medium containing 100 μg/ml gentamycin. f, Quantification of pyroptosis in BMDMs from wild-type and Gbpchr3 mice treated as in e. ELISA assays of IL-1β in supernatants, LDH release assays of cell death in supernatants and ATP assays of cell viability in cell pellets. g-h, Immunoblots and quantification of cleaved caspase-11 in BMDMs from Vcp+/+ and Vcpflox/flox; Lyz2-Cre (Vcp−/−) mice primed with 1 μg/ml Pam3CSK4 overnight before infected with wild-type or ∆EsxA Listeria (g) or S. aureus (h) at an MOI of 10 for 6 h. Cells cultured in fresh medium containing 100 μg/ml gentamycin. Cells lysed and immunoblotted 16 h post infection. Relative band intensity between cleaved caspase-11 and β-actin calculated. i, Immunoblots of BMDMs from Nlrp6+/+ and Nlrp6−/− mice primed with 1 μg/ml Pam3CSK4 overnight before transfected with cell-penetrating peptides (100 ng/μl) alone or together with SA-Lpp (100 ng/μl) for 16 h. j, Immunoblots of human macrophages from NLRP7+/+ and NLRP7−/− human macrophages treated as in i. In b, d, f-h, data are pooled from three independent experiments (n = 3) and shown as means ± SD. Unpaired two-tailed Student’s t-test was used. Exact P values are indicated in the figures. Data are representative of three independent experiments with similar results in a-j.
Extended Data Fig. 8 CTH is beneficial for the host to resist bacterial infections.
a, Kaplan–Meier survival curve of wild-type, Lyz2CreCthflox/flox (CTH cKO), Casp11−/− and Casp1−/− mice challenged intraperitoneally with poly(I:C) (2 mg/kg) for 6 h, then injected intraperitoneally with 2 ×106 CFU Listeria monocytogenes (n = 6 mice per group). b, Quantification of bacterial loads in spleen, liver and kidney of mice treated as in a. Bacterial loads were measured at 72 h post-infection (n = 5 mice per group). Casp1−/− mice were excluded due to complete mortality by day 3. ND, not detected. c, ELISA assay of serum levels of IL-1β in mice treated as in a. Serum levels of IL-1β determined at 24 h post infection (n = 5 mice per group). d, Kaplan–Meier survival curve of wild-type mice challenged intraperitoneally with poly(I:C) (2 mg/kg) for 6 h, then injected intraperitoneally with 5 × 108 CFU of S. aureus with or without PAG (50 mg/kg/day) (n = 8 mice per group). e, Quantification of bacterial loads in spleen, liver and kidney of wild-type mice treated as in d. Bacterial loads were measured at 72 h post-infection (n = 5 mice per group). f, ELISA assay of serum levels of IL-1β in wild-type mice treated as in d. Serum levels of IL-1β determined at 24 h post infection (n = 5 mice per group). In a and d, Gehan-Breslow-Wilcoxon test was used for survival analysis, n = 6 per group for a and n = 8 per group for d. In b, c, e and f, data are pooled from three independent experiments (n = 5) and shown as means ± SD. Unpaired two-tailed Student’s t-test was used. Exact P values are indicated in the figures. Data are representative of three independent experiments in a-f.
Supplementary information
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 1c,e.
Source Data Fig. 1 (download PDF )
Unprocessed western blots for Fig. 1b,d.
Source Data Fig. 2 (download XLSX )
Statistical source data for Fig. 2d,f,h.
Source Data Fig. 2 (download PDF )
Unprocessed western blots for Fig. 2b,c,e,g.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 3f–h.
Source Data Fig. 3 (download PDF )
Unprocessed western blots for Fig. 3a–c,e.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 4e–g.
Source Data Fig. 4 (download PDF )
Unprocessed western blots for Fig. 4a–d.
Source Data Fig. 5 (download XLSX )
Statistical source data for Fig. 5b,e,g.
Source Data Fig. 5 (download PDF )
Unprocessed western blots for Fig. 5a,d,f.
Source Data Fig. 6 (download XLSX )
Statistical source data for Fig. 6a–h,j.
Source Data Fig. 6 (download PDF )
Unprocessed western blots for Fig. 6i.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data for Extended Data Fig. 1b,c,e,g–i.
Source Data Extended Data Fig. 1 (download PDF )
Unprocessed western blots for Extended Data Fig. 1a,d,f.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data for Extended Data Fig. 2a,c–e.
Source Data Extended Data Fig. 2 (download PDF )
Unprocessed western blots for Extended Data Fig. 2b.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data for Extended Data Fig. 3c,e.
Source Data Extended Data Fig. 3 (download PDF )
Unprocessed western blots for Extended Data Fig. 3a,b,d.
Source Data Extended Data Fig. 4 (download PDF )
Unprocessed western blots for Extended Data Fig. 4a–c.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data for Extended Data Fig. 5e,g.
Source Data Extended Data Fig. 5 (download PDF )
Unprocessed western blots for Extended Data Fig. 5a,b,d,f.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data for Extended Data Fig. 6c,e,f.
Source Data Extended Data Fig. 6 (download PDF )
Unprocessed western blots for Extended Data Fig. 6a,b,d.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data for Extended Data Fig. 7b,d,f–h.
Source Data Extended Data Fig. 7 (download PDF )
Unprocessed western blots for Extended Data Fig. 7a,c,e,g–j.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data for Extended Data Fig. 8a–f.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Q., Wang, C., Li, M. et al. Cytosolic CTH senses bacterial lipoproteins and drives noncanonical inflammasome activation. Nat Immunol (2026). https://doi.org/10.1038/s41590-026-02511-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41590-026-02511-9