Abstract
Phage therapy is an exciting strategy against antimicrobial-resistant bacterial infections, but critical knowledge gaps regarding its clinical application persist. Here we present a case study of a 22-year-old male patient with cystic fibrosis, presenting with a recurrent, invasive and ultimately lethal Bordetella bronchialis infection, who failed compassionate-use phage therapy. Using longitudinal clinical samples, we found that our patient harbored pre-existing antibodies against active prophages induced from the genome of the infecting pathogen. Notably, these antibodies may have contributed to clinical failure by cross-reacting with and effectively neutralizing therapeutic phage. We also uncovered bacterial heteroresistance, characterized by bacterial subpopulations from the initial infection with reduced phage susceptibility, as a possible further contributor to treatment failure. These findings highlight the intricate interplay between host immunology, bacterial genetic diversity and phage biology, bearing broad importance for clinical phage therapy. Future phage therapy patients, especially those with chronic infections, should be screened for antiphage immunity and bacterial heteroresistance before phage treatment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All bacterial and phage genomes, sequencing data and metadata derived from this study have been deposited on NCBI under the BioProject number PRJNA1180749 (Supplementary Table 10). Raw data from all figures have been compiled and presented into an available file. Source data are provided with this paper.
References
GBD 2021 Antimicrobial Resistance Collaborators Global burden of bacterial antimicrobial resistance 1990–2021: a systematic analysis with forecasts to 2050. Lancet 404, 1199–1226 (2024).
Dadgostar, P. Antimicrobial resistance: implications and costs. Infect. Drug Resist. 12, 3903–3910 (2019).
Gordillo Altamirano, F. L. & Barr, J. J. Phage therapy in the postantibiotic era. Clin. Microbiol. Rev. 32, e00066–18 (2019).
Gomez-Ochoa, S. A. et al. Efficacy of phage therapy in preclinical models of bacterial infection: a systematic review and meta-analysis. Lancet Microbe 3, e956–e968 (2022).
Uyttebroek, S. et al. Safety and efficacy of phage therapy in difficult-to-treat infections: a systematic review. Lancet Infect. Dis. 22, e208–e220 (2022).
Marchi, J., Zborowsky, S., Debarbieux, L. & Weitz, J. S. The dynamic interplay of bacteriophage, bacteria and the mammalian host during phage therapy. iScience 26, 106004 (2023).
Vandamme, P. A. et al. Bordetella bronchialis sp. nov., Bordetella flabilis sp. nov. and Bordetella sputigena sp. nov., isolated from human respiratory specimens, and reclassification of Achromobacter sediminum Zhang et al. 2014 as Verticia sediminum gen. nov., comb. nov. Int. J. Syst. Evol. Microbiol. 65, 3674–3682 (2015).
Spilker, T., Darrah, R. & LiPuma, J. J. Complete genome sequences of Bordetella flabilis, Bordetella bronchialis, and “Bordetella pseudohinzii”. Genome Announc. 4, e01132-16 (2016).
Bridel, S. et al. A comprehensive resource for Bordetella genomic epidemiology and biodiversity studies. Nat. Commun. 13, 3807 (2022).
Bonilla, N. et al. Phage on tap-a quick and efficient protocol for the preparation of bacteriophage laboratory stocks. PeerJ 4, e2261 (2016).
Subedi, D. et al. Rational design of a hospital-specific phage cocktail to treat Enterobacter cloacae complex infections. Nat. Microbiol. 10, 2702–2719 (2025).
Khatami, A. et al. Standardised treatment and monitoring protocol to assess safety and tolerability of bacteriophage therapy for adult and paediatric patients (STAMP study): protocol for an open-label, single-arm trial. BMJ Open 12, e065401 (2022).
Gembara, K. & Dabrowska, K. Phage-specific antibodies. Curr. Opin. Biotechnol. 68, 186–192 (2021).
Jerne, N. K. & Avegno, P. The development of the phage-inactivating properties of serum during the course of specific immunization of an animal: reversible and irreversible inactivation. J. Immunol. 76, 200–208 (1956).
Hodyra-Stefaniak, K. et al. Natural and induced antibodies against phages in humans: induction kinetics and immunogenicity for structural proteins of PB1-related phages. Phage 1, 91–99 (2020).
Kazmierczak, Z. et al. Immune response to therapeutic staphylococcal bacteriophages in mammals: kinetics of induction, immunogenic structural proteins, natural and induced antibodies. Front. Immunol. 12, 639570 (2021).
Le, H. T. et al. Differences in phage recognition and immunogenicity contribute to divergent human immune responses to Escherichia coli and Klebsiella pneumoniae phages. Eur. J. Immunol. 55, e202451543 (2025).
Lusiak-Szelachowska, M., Miedzybrodzki, R., Fortuna, W., Borysowski, J. & Gorski, A. Anti-phage serum antibody responses and the outcome of phage therapy. Folia Microbiol. 66, 127–131 (2021).
Lusiak-Szelachowska, M. et al. Phage neutralization by sera of patients receiving phage therapy. Viral Immunol. 27, 295–304 (2014).
Dedrick, R. M. et al. Phage therapy of mycobacterium infections: compassionate use of phages in 20 patients with drug-resistant mycobacterial disease. Clin. Infect. Dis. 76, 103–112 (2023).
Lusiak-Szelachowska, M. et al. Antiphage activity of sera during phage therapy in relation to its outcome. Future Microbiol. 12, 109–117 (2017).
Van Belleghem, J. D., Dabrowska, K., Vaneechoutte, M., Barr, J. J. & Bollyky, P. L. Interactions between bacteriophage, bacteria, and the mammalian immune system. Viruses 11, 10 (2018).
Popescu, M., Van Belleghem, J. D., Khosravi, A. & Bollyky, P. L. Bacteriophages and the immune system. Annu. Rev. Virol. 8, 415–435 (2021).
Bichet, M. C., Patwa, R. & Barr, J. J. Protocols for studying bacteriophage interactions with in vitro epithelial cell layers. STAR Protoc. 2, 100697 (2021).
Haddock, N. L., Barkal, L. J. & Bollyky, P. L. Bacteriophage populations mirror those of bacterial pathogens at sites of infection. mSystems 8, e0049723 (2023).
Haddock, N. L. et al. Phage diversity in cell-free DNA identifies bacterial pathogens in human sepsis cases. Nat. Microbiol. 8, 1495–1507 (2023).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585 (2021).
Egido, J. E., Costa, A. R., Aparicio-Maldonado, C., Haas, P. J. & Brouns, S. J. J. Mechanisms and clinical importance of bacteriophage resistance. FEMS Microbiol. Rev. 46, fuab048 (2022).
Lebeaux, D. et al. A case of phage therapy against pandrug-resistant Achromobacter xylosoxidans in a 12-year-old lung-transplanted cystic fibrosis patient. Viruses 13, 60 (2021).
Blasco, L. et al. Case report: analysis of phage therapy failure in a patient with a Pseudomonas aeruginosa prosthetic vascular graft infection. Front. Med. 10, 1199657 (2023).
Jault, P. et al. Efficacy and tolerability of a cocktail of bacteriophages to treat burn wounds infected by Pseudomonas aeruginosa (PhagoBurn): a randomised, controlled, double-blind phase 1/2 trial. Lancet Infect. Dis. 19, 35–45 (2019).
Schooley, R. T. et al. Development and use of personalized bacteriophage-based therapeutic cocktails to treat a patient with a disseminated resistant Acinetobacter baumannii infection. Antimicrob. Agents Chemother. 61, e00954–17 (2017).
Gordillo Altamirano, F. L. et al. Phage-antibiotic combination is a superior treatment against Acinetobacter baumannii in a preclinical study. EBioMedicine 80, 104045 (2022).
Zhvania, P., Hoyle, N. S., Nadareishvili, L., Nizharadze, D. & Kutateladze, M. Phage therapy in a 16-year-old boy with Netherton syndrome. Front. Med. 4, 94 (2017).
Bertozzi Silva, J., Storms, Z. & Sauvageau, D. Host receptors for bacteriophage adsorption. FEMS Microbiol. Lett. 363, fnw002 (2016).
Happonen, L. J., Pajunen, M. I., Jun, J. W. & Skurnik, M. BtuB-dependent infection of the T5-like Yersinia phage varphiR2-01. Viruses 13, 2171 (2021).
Hong, J. et al. Identification of host receptor and receptor-binding module of a newly sequenced T5-like phage EPS7. FEMS Microbiol. Lett. 289, 202–209 (2008).
Golomidova, A. K. et al. Branched lateral tail fiber organization in T5-like bacteriophages DT57C and DT571/2 is revealed by genetic and functional analysis. Viruses 8, 26 (2016).
Gordillo Altamirano, F. L. & Barr, J. J. Unlocking the next generation of phage therapy: the key is in the receptors. Curr. Opin. Biotechnol. 68, 115–123 (2021).
Gillis, A. & Mahillon, J. An improved method for rapid generation and screening of Bacillus thuringiensis phage-resistant mutants. J. Microbiol. Methods 106, 101–103 (2014).
Andersson, D. I., Nicoloff, H. & Hjort, K. Mechanisms and clinical relevance of bacterial heteroresistance. Nat. Rev. Microbiol. 17, 479–496 (2019).
Meng, J., Young, G. & Chen, J. The Rcs system in Enterobacteriaceae: envelope stress responses and virulence regulation. Front. Microbiol. 12, 627104 (2021).
Wall, E., Majdalani, N. & Gottesman, S. The complex Rcs regulatory cascade. Annu. Rev. Microbiol. 72, 111–139 (2018).
Chaudhry, W. et al. Mucoidy, a general mechanism for maintaining lytic phage in populations of bacteria. FEMS Microbiol. Ecol. 96, fiaa162 (2020).
Mutalik, V. K. et al. High-throughput mapping of the phage resistance landscape in E. coli. PLoS Biol. 18, e3000877 (2020).
Smith, L. M. et al. The Rcs stress response inversely controls surface and CRISPR–Cas adaptive immunity to discriminate plasmids and phages. Nat. Microbiol. 6, 162–172 (2021).
Burmeister, A. R., Tewatia, H. & Skinner, C. A tradeoff between bacteriophage resistance and bacterial motility is mediated by the Rcs phosphorelay in Escherichia coli. Microbiology 170, 001491 (2024).
Kortright, K. E., Chan, B. K. & Turner, P. E. High-throughput discovery of phage receptors using transposon insertion sequencing of bacteria. Proc. Natl Acad. Sci. USA 117, 18670–18679 (2020).
Acton, L. et al. Collateral sensitivity increases the efficacy of a rationally designed bacteriophage combination to control Salmonella enterica. J. Virol. 98, e0147623 (2024).
Bai, J., Raustad, N., Denoncourt, J., van Opijnen, T. & Geisinger, E. Genome-wide phage susceptibility analysis in Acinetobacter baumannii reveals capsule modulation strategies that determine phage infectivity. PLoS Pathog. 19, e1010928 (2023).
Nyenhuis, D. A., Nilaweera, T. D. & Cafiso, D. S. Native cell environment constrains loop structure in the Escherichia coli cobalamin transporter BtuB. Biophys. J. 119, 1550–1557 (2020).
Shearer, J., Jefferies, D. & Khalid, S. Outer membrane proteins OmpA, FhuA, OmpF, EstA, BtuB, and OmpX have unique lipopolysaccharide fingerprints. J. Chem. Theory Comput. 15, 2608–2619 (2019).
Halperin, S. A., Ferrieri, P., Gray, E. D., Kaplan, E. L. & Wannamaker, L. W. Antibody response to bacteriophage hyaluronidase in acute glomerulonephritis after group A streptococcal infection. J. Infect. Dis. 155, 253–261 (1987).
Fluckiger, A. et al. Cross-reactivity between tumor MHC class I-restricted antigens and an enterococcal bacteriophage. Science 369, 936–942 (2020).
Berkson, J. D. et al. Phage-specific immunity impairs efficacy of bacteriophage targeting vancomycin resistant Enterococcus in a murine model. Nat. Commun. 15, 2993 (2024).
Zborowsky, S. et al. Macrophage-induced reduction of bacteriophage density limits the efficacy of in vivo pulmonary phage therapy. Nat. Commun. 16, 5725 (2025).
Roach, D. R. et al. Synergy between the host immune system and bacteriophage is essential for successful phage therapy against an acute respiratory pathogen. Cell Host Microbe 22, 38–47 (2017).
Dewachter, L., Fauvart, M. & Michiels, J. Bacterial heterogeneity and antibiotic survival: understanding and combatting persistence and heteroresistance. Mol. Cell 76, 255–267 (2019).
Klein, S. et al. Comparative genomic reveals clonal heterogeneity in persistent Staphylococcus aureus infection. Front. Cell. Infect. Microbiol. 12, 817841 (2022).
Bartell, J. A. et al. Evolutionary highways to persistent bacterial infection. Nat. Commun. 10, 629 (2019).
Stachurska, X., Roszak, M., Jablonska, J., Mizielinska, M. & Nawrotek, P. Double-layer agar (DLA) modifications for the first step of the phage-antibiotic synergy (PAS) identification. Antibiotics 10, 1306 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595 (2017).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Aziz, R. K. et al. The RAST server: rapid annotations using subsystems technology. BMC Genomics 9, 75 (2008).
Terzian, P. et al. PHROG: families of prokaryotic virus proteins clustered using remote homology. NAR Genom. Bioinform. 3, lqab067 (2021).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962 (2016).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Feldgarden, M. et al. AMRFinderPlus and the Reference Gene Catalog facilitate examination of the genomic links among antimicrobial resistance, stress response, and virulence. Sci. Rep. 11, 12728 (2021).
Bin Jang, H. et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat. Biotechnol. 37, 632–639 (2019).
Nayfach, S. et al. Metagenomic compendium of 189,680 DNA viruses from the human gut microbiome. Nat. Microbiol. 6, 960–970 (2021).
Cook, R. et al. INfrastructure for a PHAge REference Database: identification of large-scale biases in the current collection of cultured phage genomes. Phage 2, 214–223 (2021).
Turner, D. et al. Abolishment of morphology-based taxa and change to binomial species names: 2022 taxonomy update of the ICTV bacterial viruses subcommittee. Arch. Virol. 168, 74 (2023).
Jespersen, M. C., Peters, B., Nielsen, M. & Marcatili, P. BepiPred-2.0: improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res. 45, W24–W29 (2017).
Wick, R. R. et al. Trycycler: consensus long-read assemblies for bacterial genomes. Genome Biol. 22, 266 (2021).
Wick, R. R. & Holt, K. E. Polypolish: Short-read polishing of long-read bacterial genome assemblies. PLoS Comput. Biol. 18, e1009802 (2022).
Schwengers, O. et al. Bakta: rapid and standardized annotation of bacterial genomes via alignment-free sequence identification. Microb. Genom. 7, 000685 (2021).
Hawkey, J. et al. ISMapper: identifying transposase insertion sites in bacterial genomes from short read sequence data. BMC Genomics 16, 667 (2015).
Tonkin-Hill, G. et al. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21, 180 (2020).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Yu, G. Using ggtree to visualize data on tree-like structures. Curr. Protoc. Bioinform. 69, e96 (2020).
Acknowledgements
This work was supported by the Alfred Hospital and Monash University funding of VICPhage and the Monash Phage Foundry, and by Monash eResearch capabilities, including M3. J.J.B. and A.Y.P. thank the Australian National Health and Medical Research Council (NHMRC) Investigator L1 grant (no. 2023/GNT2026130 awarded to J.J.B.) and Practitioner Fellowship (grant no. APP1117940 awarded to A.Y.P.) and the Frontier Health Medical Research support (grant no. RFRHPI000017 awarded to J.J.B. and A.Y.P. as part of the Phage Australia Network). The Monash Antibody Discovery Platform is supported by funding from Bioplatforms Australia (enabled by NCRIS). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. We thank R. Burrows (University of Melbourne and Melbourne Water) for his help establishing the collaboration that granted us access to the raw sewage samples used to isolate øSimón. We thank C. Rootes, C. Pace and D. Siauw for useful feedback on the structure and readability of the manuscript, and R. Patwa for her help in logistics and laboratory management. We thank S. Usúcar Gordillo, after whom phage Simón was named.
Author information
Authors and Affiliations
Contributions
Conceptualization by F.G.A., J.J.B. and A.Y.P. Laboratory experimental work by F.G.A., D.S., M.Bu., D.M.P., M.P., D.Ko., M.J.R., K.P., J.W. and H.R. Genomics and bioinformatics by D.S., M.Be., M.Bu., S.D. and J.H. Clinical monitoring and follow-up by F.G.A., S.F.K., B.J.G., Y.H., D.Ke., T.K. and A.Y.P. Data curation, analysis and visualization by F.G.A. and A.Y.P. Provision of resources by C.R., J.J.B. and A.Y.P. Original preparation of the manuscript by F.G.A. and A.Y.P. All authors reviewed, edited and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Robert Schooley and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Lia Parkin, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Immune and inflammatory markers during phage therapy.
Levels of C-reactive protein (a), white blood cells (b), and neutrophils (c) in the patient over the course of the clinical history. Grey shaded zones in each panel represent the 30-day period of phage administration, whereas the coloured shaded zones represent the normal ranges for each marker. Each data point is a single clinical measurement. Blue arrows demark the day of patient death.
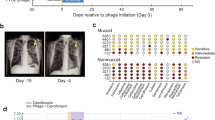
Extended Data Fig. 2 Immuno-TEM (Transmission Electron Microscopy).
Additional micrographs of phage øSimón incubated with patient’s serum from before phage therapy (Day 0) or after 21 days of phage therapy, and a gold-labelled anti-human IgG detection antibody (black dots) (left and middle columns, respectively). Assorted controls for the experiment on the right column. Scale bars depicted at the bottom corner of every micrograph. Images are representative of at least 100 observations from two experiments each.
Extended Data Fig. 3 øSimón neutralisation is mediated by anti-øSimón antibodies.
a: Correlation between the number of successive passages of serum on øSimón-coated wells, and the ELISA signal for anti-øSimón antibodies (n = 2 biological replicates). Simple linear regression with 95% CI shaded zones depicted in magenta. Using the equation from the regression (top right), it was predicted that the signal for anti-øSimón antibodies would reach the level of the vehicle control after 10 passages. b: Neutralisation assay of a 106 pfu/ml dose of øSimón after 60 min of incubation in day 0 patient serum 1:100 in PBS (black circles), or the same serum sample after 12 passages for anti-øSimón antibody depletion (magenta squares). The values for the postdepletion column were corrected by subtracting the amount of residual øSimón from the passages used for antibody depletion. Bars are medians ± 95% CI; n = 4 biological replicates; Mann-Whitney test; two-tailed. c: Bactericidal effect of patient serum, control serum or a PBS control over a B. bronchialis population. Bacteria were counted before (black circles) and after (magenta triangles) 30 minutes of treatment (bars are medians; n = 3 biological replicates; two-tailed two-way ANOVA: ns: not significant). d: The bacterial populations from panel (c) were used in an adsorption assay with øSimón. There were no differences in øSimón’s ability to adsorb to any of these bacterial cells after 80 minutes of coincubation (median ± 95% CI; n = 3 biological replicates).
Extended Data Fig. 4 In vitro mechanism of resistance against phage øSimón.
a: Representation of the btuB gene sequence in wt B. bronchialis (top) and a mutant resistant to øSimón (bottom). The red arrow and box indicate the insertion of an alanine at position 599, leading to a premature stop codon in the phage-resistant bacterium. b: Bacterial growth curves of wt B. bronchialis and the BtuB-deficient variant presented in panel (a), with and without phage (mean ± SD; n = 3 biological replicates). cand d: Bacterial growth curves of wt and BtuB-deficient B. bronchialis, respectively, in serial 1:2 dilutions of heart infusion (HI) broth in PBS (mean ± SD; n = 3 biological replicates). e: For the dilutions from panels (c) and (d) that supported B. bronchialis growth, comparison of the area under the curve between wt (grey) and BtuB-deficient (pink) isolates (bars are medians; n = 3 biological replicates; 2-way ANOVA; two-tailed). ns: not significant. f: Adsorption assay of øSimón to wt or BtuB-deficient B. bronchialis over 90 minutes (median ± 95% CI; n = 3 biological replicates; Kruskal-Wallis test).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–3 and Tables 1 and 2 (Supplementary Tables 3–10 uploaded as separate Excel file ‘Supplementary Tables 1’).
Source data
Source Data Figs. 1–6, Source Data Extended Data Figs. 1, 3 and 4. (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gordillo Altamirano, F., Subedi, D., Beiers, M. et al. Cross-reactive anti-prophage antibodies and bacterial heteroresistance implicated in phage therapeutic failure. Nat Med (2026). https://doi.org/10.1038/s41591-026-04301-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41591-026-04301-0