Abstract
Amyotrophic lateral sclerosis (ALS) is categorized into ~10% familial and ~90% sporadic cases. While familial ALS is caused by mutations in many genes of diverse functions, the underlying pathogenic mechanisms of ALS, especially in sporadic ALS (sALS), are largely unknown. Notably, about half of the cases with sALS showed defects in mitochondrial respiratory complex IV (CIV). To determine the causal role of this defect in ALS, we used transcription activator-like effector-based mitochondrial genome editing to introduce mutations in CIV subunits in rat neurons. Our results demonstrate that neuronal CIV deficiency is sufficient to cause a number of ALS-like phenotypes, including cytosolic TAR DNA-binding protein 43 redistribution, selective motor neuron loss and paralysis. These results highlight CIV deficiency as a potential cause of sALS and shed light on the specific vulnerability of motor neurons, marking an important advance in understanding and therapeutic development of sALS.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All raw RNA-seq and snRNA-seq data can be accessed via the Gene Expression Omnibus (accession nos. GSE252875 and GSE275999). Processed gene expression count matrices have been deposited with Zenodo62. Proteomics data have been uploaded to the PRIDE database under accession no. PXD058902. Source data are provided with this paper.
Change history
21 March 2025
A Correction to this paper has been published: https://doi.org/10.1038/s41593-025-01941-2
References
Grad, L. I., Rouleau, G. A., Ravits, J. & Cashman, N. R. Clinical spectrum of amyotrophic lateral sclerosis (ALS). Cold Spring Harb. Perspect. Med. 7, a024117 (2017).
Talbot, K. Familial versus sporadic amyotrophic lateral sclerosis—a false dichotomy? Brain 134, 3429–3431 (2011).
Todd, T. W. & Petrucelli, L. Modelling amyotrophic lateral sclerosis in rodents. Nat. Rev. Neurosci. 23, 231–251 (2022).
Kim, G., Gautier, O., Tassoni-Tsuchida, E., Ma, X. R. & Gitler, A. D. ALS genetics: gains, losses, and implications for future therapies. Neuron 108, 822–842 (2020).
Zhou, W. & Xu, R. Current insights in the molecular genetic pathogenesis of amyotrophic lateral sclerosis. Front. Neurosci. 17, 1189470 (2023).
Morgan, S. & Orrell, R. W. Pathogenesis of amyotrophic lateral sclerosis. Br. Med. Bull. 119, 87–98 (2016).
Kiernan, M. C. et al. Amyotrophic lateral sclerosis. Lancet 377, 942–955 (2011).
Barber, S. C. & Shaw, P. J. Oxidative stress in ALS: key role in motor neuron injury and therapeutic target. Free Radic. Biol. Med. 48, 629–641 (2010).
Smith, E. F., Shaw, P. J. & De Vos, K. J. The role of mitochondria in amyotrophic lateral sclerosis. Neurosci. Lett. 710, 132933 (2019).
Dafinca, R., Barbagallo, P. & Talbot, K. The role of mitochondrial dysfunction and ER stress in TDP-43 and C9ORF72 ALS. Front. Cell. Neurosci. 15, 653688 (2021).
Yu, M. et al. Widespread mislocalization of FUS is associated with mitochondrial abnormalities in skeletal muscle in amyotrophic lateral sclerosis with FUS mutations. J.Neuropathol. Exp. Neurol. 81, 172–181 (2022).
Tsai, Y.-L. et al. ALS/FTD-associated protein FUS induces mitochondrial dysfunction by preferentially sequestering respiratory chain complex mRNAs. Genes Dev. 34, 785–805 (2020).
Ruan, Y. et al. CHCHD2 and CHCHD10 regulate mitochondrial dynamics and integrated stress response. Cell Death Dis. 13, 156 (2022).
Sasaki, S. & Iwata, M. Mitochondrial alterations in the spinal cord of patients with sporadic amyotrophic lateral sclerosis. J. Neuropathol. Exp. Neurol. 66, 10–16 (2007).
Crugnola, V. et al. Mitochondrial respiratory chain dysfunction in muscle from patients with amyotrophic lateral sclerosis. Arch. Neurol. 67, 849–854 (2010).
Wei, W. et al. Mitochondrial DNA point mutations and relative copy number in 1363 disease and control human brains. Acta Neuropathol. Commun. 5, 13 (2017).
Hirano, M. et al. Amyotrophic lateral sclerosis with ragged-red fibers. Arch. Neurol. 65, 403–406 (2008).
Wang, W. et al. A novel nonsense variant in MT-CO3 causes MELAS syndrome. Neuromuscul. Disord. 31, 558–565 (2021).
Guise, A. J. et al. TDP-43-stratified single-cell proteomics of postmortem human spinal motor neurons reveals protein dynamics in amyotrophic lateral sclerosis. Cell Rep. 43, 113636 (2024).
Tan, L. et al. A conditional knockout rat resource of mitochondrial protein-coding genes via a DdCBE-induced premature stop codon. Sci. Adv. 9, eadf2695 (2023).
Silva-Pinheiro, P. et al. A library of base editors for the precise ablation of all protein-coding genes in the mouse mitochondrial genome. Nat. Biomed. Eng. 7, 692–703 (2023).
DiMauro, S. & De Vivo, D. C. Genetic heterogeneity in Leigh syndrome. Ann. Neurol. 40, 5–7 (1996).
Rak, M. et al. Mitochondrial cytochrome c oxidase deficiency. Clin. Sci. 130, 393–407 (2016).
Schweingruber, C. & Hedlund, E. The cell autonomous and non-cell autonomous aspects of neuronal vulnerability and resilience in amyotrophic lateral sclerosis. Biology 11, 1191 (2022).
Zuko, A. et al. tRNA overexpression rescues peripheral neuropathy caused by mutations in tRNA synthetase. Science 373, 1161–1166 (2021).
Pan, J. Z. et al. ATF3 is a neuron-specific biomarker for spinal cord injury and ischaemic stroke. Clin. Transl. Med. 14, e1650 (2024).
Renthal, W. et al. Transcriptional reprogramming of distinct peripheral sensory neuron subtypes after axonal injury. Neuron 108, 128–144 (2020).
Neel, D. V. et al. Gasdermin-E mediates mitochondrial damage in axons and neurodegeneration. Neuron 111, 1222–1240 (2023).
Krach, F. et al. Transcriptome–pathology correlation identifies interplay between TDP-43 and the expression of its kinase CK1E in sporadic ALS. Acta Neuropathol. 136, 405–423 (2018).
Mitchell, J. C. et al. Overexpression of human wild-type FUS causes progressive motor neuron degeneration in an age- and dose-dependent fashion. Acta Neuropathol. 125, 273–288 (2013).
Graffmo, K. S. et al. Expression of wild-type human superoxide dismutase-1 in mice causes amyotrophic lateral sclerosis. Hum. Mol. Genet. 22, 51–60 (2013).
Jackson, M., Marks, L., May, G. H. W. & Wilson, J. B. The genetic basis of disease. Essays Biochem. 62, 643–723 (2018).
Guo, X. et al. High-frequency and functional mitochondrial DNA mutations at the single-cell level. Proc. Natl Acad. Sci. USA 120, e2201518120 (2023).
Parakatselaki, M. E. & Ladoukakis, E. D. mtDNA heteroplasmy: origin, detection, significance, and evolutionary consequences. Life 11, 633 (2021).
Nissanka, N. & Moraes, C. T. Mitochondrial DNA heteroplasmy in disease and targeted nuclease-based therapeutic approaches. EMBO Rep. 21, e49612 (2020).
Monani, U. R. et al. The human centromeric survival motor neuron gene (SMN2) rescues embryonic lethality in Smn−/− mice and results in a mouse with spinal muscular atrophy. Hum. Mol. Genet. 9, 333–339 (2000).
Murray, L. M., Beauvais, A., Gibeault, S., Courtney, N. L. & Kothary, R. Transcriptional profiling of differentially vulnerable motor neurons at pre-symptomatic stage in the Smn2b/− mouse model of spinal muscular atrophy. Acta Neuropathol. Commun. 3, 55 (2015).
Mead, R. J., Shan, N., Reiser, H. J., Marshall, F. & Shaw, P. J. Amyotrophic lateral sclerosis: a neurodegenerative disorder poised for successful therapeutic translation. Nat. Rev. Drug Discov. 22, 185–212 (2023).
Obrador, E. et al. Oxidative stress, neuroinflammation and mitochondria in the pathophysiology of amyotrophic lateral sclerosis. Antioxidants 9, 901 (2020).
Le Gall, L. et al. Molecular and cellular mechanisms affected in ALS. J. Pers. Med. 10, 101 (2020).
McCauley, M. E. & Baloh, R. H. Inflammation in ALS/FTD pathogenesis. Acta Neuropathol. 137, 715–730 (2019).
McGrath, M. S., Zhang, R., Bracci, P. M., Azhir, A. & Forrest, B. D. Regulation of the innate immune system as a therapeutic approach to supporting respiratory function in ALS. Cells 12, 1031 (2023).
Held, A. et al. iPSC motor neurons, but not other derived cell types, capture gene expression changes in postmortem sporadic ALS motor neurons. Cell Rep. 42, 113046 (2023).
Thonhoff, J. R., Jordan, P. M., Karam, J. R., Bassett, B. L. & Wu, P. Identification of early disease progression in an ALS rat model. Neurosci. Lett. 415, 264–268 (2007).
Kraeuter, A. K., Guest, P. C. & Sarnyai, Z. The open field test for measuring locomotor activity and anxiety-like behavior. Methods Mol. Biol. 1916, 99–103 (2019).
Pang, S. et al. Integrin β1/FAK/SRC signal pathway is involved in autism spectrum disorder in Tspan7 knockout rats. Life Sci. Alliance 6, e202201616 (2022).
Sims, S. et al. Absent LH signaling rescues the anxiety phenotype in aging female mice. Mol. Psychiatry 28, 3324–3331 (2023).
Dong, W. et al. Ablation of C9orf72 together with excitotoxicity induces ALS in rats. FEBS J. 288, 1712–1723 (2021).
Gu, L., Liu, Y.-J., Wang, Y.-B. & Yi, L.-T. Role for monoaminergic systems in the antidepressant-like effect of ethanol extracts from Hemerocallis citrina. J. Ethnopharmacol. 139, 780–787 (2012).
Yankelevitch-Yahav, R., Franko, M., Huly, A. & Doron, R. The forced swim test as a model of depressive-like behavior. J. Vis. Exp. 97, 52587 (2015).
Zhang, Z., Theurkauf, W. E., Weng, Z. & Zamore, P. D. Strand-specific libraries for high throughput RNA sequencing (RNA-Seq) prepared without poly(A) selection. Silence 3, 9 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Zhong, J. et al. PCDHA9 as a candidate gene for amyotrophic lateral sclerosis. Nat. Commun. 15, 2189 (2024).
Liu, B. et al. Monosialoganglioside protects against bupivacaine-induced neurotoxicity caused by endoplasmic reticulum stress in rats. Drug Des. Devel. Ther. 13, 707–718 (2019).
El-Sayyad, H. I., Amin, A. H., El-Ghawet, H. A., El-Shahari, E. A. & El-Gebaly, A. Role of pomegranate juice and atorvastatin in ameliorating spinal neurotoxicity of Wistar rats maternally fed on hypercholesterolemic diet. J. Mol. Biomark. Diagn. 8, 1–9 (2017).
Bang, M. L. et al. Nebulin-deficient mice exhibit shorter thin filament lengths and reduced contractile function in skeletal muscle. J. Cell Biol. 173, 905–916 (2006).
Arnold, W. D. et al. Electrophysiological motor unit number estimation (MUNE) measuring compound muscle action potential (CMAP) in mouse hindlimb muscles. J. Vis. Exp. 103, 1–8 (2015).
Ni, J. et al. Mitochondrial genome variations are associated with amyotrophic lateral sclerosis in patients from mainland China. J. Neurol. 269, 805–814 (2022).
Cheng, M. & Li, K. Mitochondrial CIV deficiency recapitulates amyotrophic lateral sclerosis. Zenodo https://doi.org/10.5281/zenodo.14436071 (2024).
Acknowledgements
We thank J. Lin, P. Guo, Y. Yu and J. Qiu for critical comments. We acknowledge the financial support provided by the Ministry of Science and Technology of China (grant nos. 2019YFA0508701 and 2022YFA1303300 to X.R.Z.), the National Natural Science Foundation of China (no. 82450106 to X.R.Z), the Haihe Laboratory of Cell Ecosystem Innovation Fund (no. HH23KYZX0003 to X.R.Z. and no. HH24KYZX0007 to Y.W.M.), the National Key R&D Program of China (no. 2022YEF0203200 to D.L.), the CAMS Innovation Fund for Medical Sciences (grant no. 2021-I2M-1-024 to Y.W.M. and grant no. 2022-I2M-1-020 to D.L.) and the Non-profit Central Research Institute Fund of the Chinese Academy of Medical Sciences (no. 2023-PT180-01 to Y.W.M.).
Author information
Authors and Affiliations
Contributions
X.R.Z. and Y.W.M. planned the studies and analyzed all the data. M.C., D.L. and Y.W. performed all the experiments except for the bioinformatics analysis. X.W.T. and Y.W. performed all the RNA-seq and proteomics. K.X.L. and J.Z. performed the bioinformatics analysis. X.L.Q. carried out the gene editing of rats. C.Z.Y., K.Q.J. and W.W. performed the patient studies. J.L.W. provided the blood sequencing data for the patients with ALS. J.Y.F. performed the electromyography analysis. X.Z., W.N.K., J.X.M. and H.J.L. generated the mutant rats and performed the rat genotyping. J.X.M., K.R.L. and Y.H.W. conducted the animal behavior tests. P.P.W. and Q.S.L. performed the BN-PAGE analysis. C.Y.S. conducted the human brain analyses. X.R.Z. and X.-D.F. wrote the paper. All authors discussed the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Xiongwei Zhu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
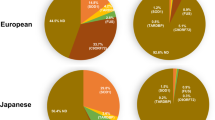
Extended Data Fig. 1 Defective CIV in ALS patients.
a, Three representative sALS patients with frameshift mutations in their COXI and COXIII subunits. Patient 2 carried a 6289 C insertion (as illustrated) besides a 6692 A deletion as in patient 1 (not shown). Patient 3 carried a 9487 T deletion. b, Section of formalin-fixed cerebellar tissues stained for COXI from six non-demented controls, four C9-ALS/FTD, and two sALS patients. Scale bar, 50 μm. c, Section of formalin-fixed cerebellar tissues stained for COXII from seven non-demented controls, five C9-ALS/FTD, and two sALS patients. Scale bar, 50 μm.
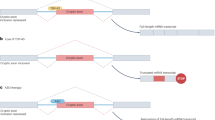
Extended Data Fig. 2 Neuronal-specific targeting of mtDNA.
a, Illustration of the strategy for generating neuronal-specific COXII and COXIII mutant rats. Crossing COXIII/IILSL rats with NeuN-Cre rats results in DdCBE expression in mature neurons to target mtCOXs. The conversion of the targeted nucleotide to ‘TAA’ creates a premature termination codon for COXII or COXIII. LSL: loxp-stop-loxp, MTS: mitochondrial targeted sequence. b, Design overview of DdCBEs for targeting rat COXI G5396, COXII C7021 and COXIII G8645. The green text highlights the mitochondrial sequences targeted by the base editors for specific editing. c, Onset and progression of disease phenotype with COXIILSL/COXIINeuN-Cre, and COXIIILSL/COXIIINeuN-Cre rats. Data are presented as mean ± SEM. The statistical test used was Log-rank test. d, The survival curve of COXILSL/COXINeuN-Cre, COXIILSL/COXIINeuN-Cre and COXIIILSL/COXIIINeuN-Cre rats. The statistical test used was Log-rank test. e, Design overview of DdCBEs for targeting rat ND1 G2996, ATP8 G7783. The green text highlights the mitochondrial sequences targeted by the base editors for specific editing. f, Survival curves of ND1LSL/ND1NeuN-Cre and ATP8LSL/ATP8NeuN-Cre rats. Data are presented as mean ± SEM. The statistical test used was Log-rank test.
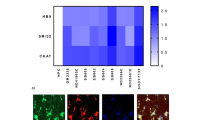
Extended Data Fig. 3 Characterization of neuronal-specific targeting of COXII/III.
a-b, DdCBE-mediated G-to-A conversion frequency at COXII C7021 and COXIII G8645 analyzed by deep sequencing. n = 3 rats/group. Data were presented as means ± SEM. c, DdCBE-mediated G-to-A conversion frequency at COXIII G8645 analyzed by deep sequencing. n = 3 rats. Data were presented as means ± SEM. d, BN-PAGE followed by Western blotting analysis to analyze the abundance of complex II, V and IV in the brain of COXIIILSL/COXIIINeuN-Cre rats, n = 3 rats/group. e, SDS-PAGE followed by Western blotting to analyze the protein level of CIV subunits in the brain, n = 3 rats/group. f, BN-PAGE followed by Western blotting to analyze the abundance of complex II, V and IV in thoracic spinal cord of COXIIILSL/COXIIINeuN-Cre rats, n = 3 rats/group. g-h, SDS-PAGE followed by Western blotting to analyze the protein level of CIV subunits in the thoracic spinal cord and lumbar spinal cord of COXIIILSL/COXIIINeuN-Cre rats, n = 3 rats/group. i, SDS-PAGE followed by Western blotting to analyze the protein level of CIV subunits in the lumbar spinal cord of COXIILSL/COXIINeuN-Cre rats, n = 3 rats/group. j-k, Body weight changes in male and female COXIILSL/COXIINeuN-Cre and COXIIILSL/COXIIINeuN-Cre rats. Data are presented as mean ± SEM. The statistical test used was Log-rank test.
Extended Data Fig. 4 Motor-related behavior of neuronal CIV deficient rats.
a-b, Behavioral tests conducted on COXIIILSL and COXIIINeuN-Cre rats to evaluate the locomotor ability at the age of 3 weeks and 1 month. Data are presented as mean ± SEM. The statistical test performed was a two-sided unpaired Student’s t-test.
Extended Data Fig. 5 Deficient COXII or COXIII impairs spinal cord structures in adult rats.
a-b, Electron microscopic images of the ventral lumbar roots in comparison with dorsal roots from COXIIINeuN-Cre/ COXIIILSL and COXIINeuN-Cre/COXIILSL rats at the end-stage disease. Scale bars: (a) Ventral root 10 µm and 2 µm, Dorsal root 5 µm and 1 µm (b) Ventral root 10 µm and 2 µm, Dorsal root 5 µm and 2 µm. All experiments were performed independently three times with similar results.
Extended Data Fig. 6 Denervation and atrophy of skeletal muscle in neuronal specific COXII deficient rats.
a-b, Representative images of gastrocnemius (GA) muscle and tibialis anterior (TA) muscle of 5-week COXIILSL and COXIINeun-Cre rats and quantified the muscle weights are presented on the right. Scale bar, 1 cm. Data are presented as mean ± SEM. The statistical test performed was a two-sided unpaired Student’s t-test. c, Representative cross sections of WGA-stains transverse sections of tibialis anterior (TA) muscle in COXIILSL and COXIINeun-Cre rats. scale bar, 200 μm. All experiments were performed independently three times with similar results. d, Quantification of averages cross-section area (CSA) of tibialis. Data are presented as mean ± SEM. The statistical test performed was multiple unpaired Student’s t-tests. e, The representative image shows the neuromuscular junction (NMJ) in hindlimb skeletal muscles of COXIILSL and COXIINeuN-Cre. Motor endplates were stained with a-bungarotoxin to visualize AchR and SV2 was used to label neuromuscular synapses. Scale bar, 20 μm. All experiments were performed independently three times with similar results. f, Quantification of denervated neuromuscular junction (NMJ) in COXIINeuN-Cre rats. Data are presented as mean ± SEM. The statistical test performed was multiple unpaired Student’s t-tests. g-i, Representative images and quantification of CMAPs evoked by supramaximal stimulation of sciatic nerves and the quantification of MUNEs for COXIILSL and COXIINeuN-Cre. Data are presented as mean ± SEM. The statistical test performed was a two-sided unpaired Student’s t-test.
Extended Data Fig. 7 RNA-Seq and quantitative-mass spectrometry analysis of COXIIINeuN-Cre rats.
a, GO term enrichment analysis of significantly up-regulated genes in lumber spinal cord of COXIIINeuN-Cre rat. Padj<0.05 and |log2FC | >1. b & d, GSEA plots showing the enrichment profiles of cell death genes and immune response genes.Padj<0.05. c, Heatmap showing upregulated DEGs related to the cellular response to oxidative stress and downregulated DEGs related to neuroactive ligand receptor interactions in COXIIINeuN-Cre rats. padj<0.05 and |log2FC | >1. e-f, Volcano plots showing the proteins with significantly altered abundance in lumbar spinal cord of COXIIINeuN-Cre rats relative to scrambled controls. P < 0.05 and |log2FC | >1.
Extended Data Fig. 8 ATP and development progression analysis of COXIIINeuN-Cre rats.
a-b,The representative image shows the TDP-43 at 2, 4 and 6 weeks in spinal cords of COXIIILSL and COXIIINeuN-Cre rats. Scale bar, 20 μm. c, ATP production in lumber spinal cord of COXIIILSL and COXIIINeuN-Cre rats. Data are presented as mean ± SEM. The statistical test performed was a two-sided unpaired Student’s t-test. d, Representative confocal fluorescence microscopic analysis showing the expression of GFAP and IBA1 in the lumbar spinal cord of COXIIILSL and COXIIINeuN-Cre rats. All experiments were performed independently three times with similar results.Scale bar, 500 μm. e, Electron microscopic images showing demyelination in the lumbar spinal cord. All experiments were performed independently three times with similar results. Scale bar, 2 μm. f, The temporal map of development progression in ALS rat model.
Extended Data Fig. 9 Changes in ALS-related proteins of COXIINeuN-Cre and COXIIINeuN-Cre rats.
a-b, SDS-PAGE followed by Western blotting to analyze the protein level of ALS-related protein in COXIINeuN-Cre and COXIIINeuN-Cre rats, n = 3 rats/group.
Supplementary information
Supplementary Information (download PDF )
Supplementary Tables 1 and 2. Table 1: Human sample information. Table 2: Primers used for genotyping, Sanger sequencing, deep sequencing and qPCR.
Supplementary Video 1 (download MP4 )
Video for COXIINeuN-Cre rat and COXIILSL control rat at 3 weeks of age in a horizontal view angle.
Supplementary Video 2 (download MP4 )
Video for COXIINeuN-Cre rat and COXIILSL control rat at 1 month of age in a horizontal view angle.
Supplementary Video 3 (download MP4 )
Video for COXIINeuN-Cre rat and COXIILSL control rat at 1.5 month of age in a horizontal view angle.
Supplementary Video 4 (download MP4 )
Video for COXIIINeuN-Cre rat and COXIIILSL control rat at 3 weeks of age in a horizontal view angle.
Supplementary Video 5 (download MP4 )
Video for COXIIINeuN-Cre rat and COXIIILSL control rat at 1 month of age in a horizontal view angle.
Supplementary Video 6 (download MP4 )
Video for COXIIINeuN-Cre rat and COXIIILSL control rat at 2 months of age in a horizontal view angle.
Supplementary Video 7 (download MP4 )
Video for COXINeuN-Cre rat and COXILSL control rat at 2 weeks of age in a vertical view angle.
Source data
Source Data Figs. 1–5 and Extended Data Figs. 2–4, 6, 8 and 9 (download ZIP )
Source data files for the main and extended data figures.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cheng, M., Lu, D., Li, K. et al. Mitochondrial respiratory complex IV deficiency recapitulates amyotrophic lateral sclerosis. Nat Neurosci 28, 748–756 (2025). https://doi.org/10.1038/s41593-025-01896-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41593-025-01896-4
This article is cited by
-
Mutant CHCHD10 disrupts cytochrome c oxidation and activates mitochondrial retrograde signaling
EMBO Molecular Medicine (2025)