Abstract
Recent advancements in intravital microscopy have enabled the study of cell death in vivo under various experimental conditions, such as infection and cancer. However, the limited throughput of this technology, together with a lack of openly accessible datasets, affects the development of algorithms for the automatic detection and characterization of cell death, which in turn require the integration of extensive and curated datasets. To address these needs, we present a curated dataset of microscopy videos depicting the death of neutrophils, eosinophils, and dendritic cells, acquired in the spleen and in the lymph node of mice under inflammatory conditions. The dataset provides time-lapse imaging data, along with coordinates in space and time of cell death events displaying apoptotic–like morphodynamics, and 3D reconstruction of the cell morphology at each time point. Altogether, these data will be pivotal for developing computer vision and bioimage analysis methods to advance cell death research.
Similar content being viewed by others
Background and Summary
Cell death is pivotal in multiple physiological and pathological processes1,2. Every day, damaged or dysfunctional cells undergo apoptosis to accommodate new healthy cells, an essential mechanism in tissue development and homeostasis3,4,5. Moreover, cell death is also associated with the defense mechanism entitled to clear dysfunctional or infected cells, thus preventing the onset of pathological conditions like cancer and infectious diseases6. The fine regulation of this mechanism is necessary to limit inflammation7 and to prevent autoimmune disorders2. The death of immune cells is particularly important as it contributes to the initiation of the immune response8, and the selection of antigen-specific cells9, amongst others10.
From a morphological perspective, multiple types of cell death are characterized by the disruption of the cell membrane and the formation of multiple bodies. Amongst these, apoptosis presents hallmarks such as chromatin condensation, cell shrinkage, membrane blebbing, and the formation of apoptotic bodies11. The latter are membrane vesicles that encapsulate the cytoplasm preventing the leakage of alarming signals into the extracellular space12. This process is considered immunologically silent, as it does not further elicit an inflammatory response. Successively, upon the formation of apoptotic bodies, phagocytic cells are recruited locally to clear the resulting apoptotic vesicles13, promoting the end of this process. In this paper, we refer as “apoptotic-like” to all the cell death events that yield to the disruption of cell membrane and formation of apoptotic bodies.
Different imaging techniques can be used to study apoptotic-like cell death. In vitro imaging allows control over multiple experimental conditions, including cell density, migratory cues, or signaling pathways14,15,16. However, this modality cannot capture the complex behavior of the immune system within living tissues. Conversely, techniques such as 2-photon intravital microscopy (2P-IVM) enable the visualization of cells within their physiological environment by delivering in real time17,18,19, four-dimensional time-lapses for different channels (x, y, z, c, t)20,21. Therefore, 2P-IVM represents one of the most informative platforms for the physiological study of cell death in vivo22. However, acquiring high-quality 2P-IVM movies remains non-trivial due to technical and biological challenges such as the surgical procedure, the bleaching of the sample, and the stability of the imaging site, among others23,24,25. As a result, the potential to generate extensive apoptotic datasets in vivo remains limited, whereas the lack of accessibility prevents the reuse of existing 2P-IVM movies25,26. Consequently, the scarcity of 2P-IVM movies negatively impacts studies that require curated datasets, such as comparative analysis or algorithmic solutions. This shortage of data mainly affects the latter aspect, given that mainstream deep learning (DL) algorithms are known for being data-hungry27. Therefore, establishing publicly accessible 2P-IVM repositories, complete with metadata and annotations, becomes pivotal. Such an initiative would broaden the scope of fundamental research on cell death and enable the development of robust algorithms to quantify this biological process in vivo.
In this study, we address this need by providing a comprehensive dataset capturing cell death events with apoptotic-like morphology in vivo28. This dataset encompasses 2P-IVM movies featuring three immune cell type:, neutrophils, eosinophils, and dendritic cells imaged in various experimental settings associated with the inflammatory response. A schematic workflow is shown in Fig. 1A,B. In addition to the raw data, we provide manual annotations of cell death events, which were curated by multiple operators trained to detect the morphological hallmarks of the process (Fig. 2). Accordingly, each apoptotic-like event is first identified in 2P-IVM fields-of-view (Fig. 2A), and successively annotated for the duration of the process (Fig. 2B). The annotations include centroid-based trajectories of the apoptotic-like events, frame-by-frame semantic annotations describing the cell morphology at any given time step, and 3D volumetric reconstructions of the cellular surface. To determine if a cell presented apoptotic-like morphology, we strictly adhered to the observation of the morphological hallmarks of apoptosis, including shrinkage of the cell, membrane blebbing, and the formation of apoptotic bodies. As a result, the presented catalog is an invaluable resource for comparative studies and the development of analytical solutions for the classification, detection, and segmentation of apoptotic-like cell death in vivo.
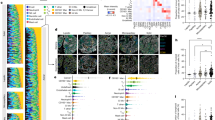
Dataset generation workflow. (A) Representative intravital acquisition set-up. Immune-relevant tissues from an anesthetized mouse were surgically exposed and subjected to 2P-IVM. Resulting in vivo acquisitions (B) are 4D sequential data composed of multiple focal planes per time point and multiple channels. Raw data are annotated by three individual operators tracking cells undergoing apoptotic-like death and generating a 3D mask. Dying cells are associated with a cell state between “blebbing” and “disrupting” that defines the phase of the ongoing process.
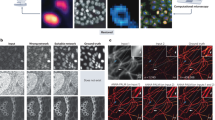
Dataset annotation of apoptotic events. (A) Micrograph depicting a MP-IVM acquisition in the lymph node of a mouse. Neutrophils expressing GFP (green) and a UV-inactivated influenza vaccine are labeled with DiD (red). (B) Dying cells from MP-IVM acquisitions are annotated frame by frame for the duration of the process according to their morphology (top). Frame annotation include semantic labels (pre-apoptotic, blebbing, disrupted), centroid tracking (middle), and 3D volumetric reconstruction of the cell (bottom).
Methods
Mice
Mice were maintained in a pathogen-free animal facility at the Institute for Research in Biomedicine (Bellinzona, Switzerland). Before imaging, mice were anesthetized with a cocktail of xylazine (10 mg/Kg) and ketamine (100 mg/Kg), as previously reported21. All experiments were conducted in compliance with the regulations set by the local authorities and were approved by the Swiss Federal Veterinary Office.
Intravital microscopy
All the videos were acquired through intravital two-photon microscopy (TrimScope, LaVision BioTec) in lymphoid organs (lymph nodes and spleen), under different experimental conditions detailed in Table 1. To acquire videos in the lymph nodes, the organ was exposed through minimally invasive surgery as previously described29. Then, time-lapse images were acquired using a custom upright microscope. To acquire videos in the spleen, mice were anesthetized and immobilized, while the spleen was accessed by making an incision through the skin and musculature as previously described30. After exposure of the organ, the spleen was moisturized by applying pre-warmed PBS. In both surgeries, 4D data encompassing multiple stacks, channels, and time points (Table 1, Fig. 1A) were generated. The imaging system employed two Ti:sapphire lasers (Chameleon Ultra I, Chameleon Ultra II, Coherent) and an optical oscillator that emitted light within the 1,010–1,340 nm range (Chameleon Compact OPO, Coherent). The output wavelength ranged between 690–1,080 nm, ontributing to probe excitation and tissue second-harmonic generation (SHG).
All the videos were acquired under inflammatory conditions. In all, except Neu7, Neu11, Neu17, and Den1, UV-inactivated PR8 virus was injected in the footpad 0 up to 24 hours prior to imaging. Neu7, Neu11, and Neu 17 videos were acquired in an experimental model of Vaccinia virus infection, whereas Den1 was acquired in a model of B16 melanoma metastasis in the lymph node. CFB and GFP-labeled neutrophils were adoptively transferred from a donor to a recipient animal which was CD11c-YFP in the case of Den1. Further details on the staining and experimental stimuli are available through Immunemap31.
Data processing
The microscopy data acquired from the videos were stored in HDF5 files, comprising uint8 or uint16 TIFFs. Prior to image analysis, the raw data underwent no preprocessing. For cell detection, tracking, and volumetric reconstruction of the microscopy videos, we employed Imaris imaging software (Oxford Instruments, v9.7.2). Subsequently, the data obtained from Imaris underwent further analysis using customized Matlab and Python scripts (refer to the code availability section for details).
Manual tracking
The centroids of cells were manually annotated and tracked over time using the “Spots drawing” tool in Imaris (Bitplane). To ensure robustness, three operators independently and redundantly performed tracking without knowledge of reciprocal results. During the tracking process, the operators followed morphological criteria to identify cells showing apoptotic-like morphodynamics. Specifically, they focused on sequences exhibiting the characteristic hallmarks of apoptosis: 1) membrane blebbing, 2) subsequent formation of apoptotic bodies, and 3) eventual cell disruption (Fig. 2B). For each frame within the apoptotic-like sequences, the operators placed a centroid at the center of the cell using the Imaris “Spots” function, resulting in a connected track representing the apoptotic-like event (x, y, z, t, ID). Subsequently, the trajectories generated by each operator were consolidated into a ground truth using a majority consensus scheme that considered the duration and the location of the event. The starting and ending time points were determined by identifying overlaps among at least two of the three tracks of the annotators. For each time point, the two closest spatial coordinates were selected and averaged to create a final ground truth. Then, we performed volumetric 3D reconstruction using the Imaris “Surfaces” function. This reconstruction process generated precise 3D meshes of each cell undergoing apoptotic-like death, enhancing the accuracy and reliability of the dataset.
Semantic annotation
Three independent annotators labeled the state of the cells based on specific morphological criteria. To annotate the interval and dynamics of the cell death process, the annotators assigned to each cell, for each frame within the ground truth sequence, one of two semantic labels: “membrane blebbing” or “cell disruption (Table 2). To ensure robustness and consistency, a majority consensus scheme was employed to establish the final cell state. This scheme considered the overlap between annotations from at least two operators, promoting a unified and reliable consensus on the semantic label for each cell at every time point, hence minimizing potential biases and variations in the annotations.
Computation of quality metrics
To compute the signal-to-noise ratio (SNR) of the entries of the dataset, for each frame of the movies we computed a reference denoised image using a median filter. Successively, we used the SNR function of the Matlab Signal Processing toolbox to compare the reference image to the original one, computing an estimate of the SNR. This procedure was applied for each channel separately.
Computation of density metrics
The density metrics were computed using adaptive thresholding and morphological operations to separate foreground objects from the background in each channel. We then counted the number of cells visible at each time point in each channel. An estimate of the number of cells throughout the movies was generated by averaging these numbers over time. By knowing the spatial coordinates of the cells, we also estimated the average shortest distance between cells and a coefficient of clustering depicting the percentage of cells in close proximity.
Statistical analyses
Statistical analysis and plot visualization were performed using GraphPad Prism 8 (Graphpad, La Jolla, USA). All statistical tests were performed using the non-parametric Kruskal-Wallis test or Mann-Witney test. The significance of the p-value is described as * when p < 0.05, ** when p < 0.01, and *** when p < 0.001.
Data Record
Imaging data, metadata, and experimental details are available through the following channels: https://zenodo.org/records/14551288 or https://app.immunemap.org/cddb. The dataset on Zenodo28 comprises unprocessed raw microscopy videos, each characterized by a unique identifier (ID) and essential metadata. These videos encompass multiple channels and extend across five dimensions: spatial coordinates (x, y, z), time (t), and multiple channels (c). Raw videos are stored in HDF5 format and uploaded individually according to their cell type, for instance “Neu1.h5”. Within each HDF5 file, two types of annotations are included: 1) tracks coordinates of apoptotic-like events (x, y, z, t, ID), and 2) 3D volumetric reconstructions of cell deaths represented by meshes defined by vertices and edges. An additional folder labeled “Annotations” contains two sub-folders:
-
1.
Cell deaths
-
2.
Semantic annotations
The “Cell deaths” folder contains trajectories of apoptotic-like events in comma-separated value format (CSV). Each filename incorporates the identifier of the corresponding microscopy acquisition and the channel where apoptotic-like cells were observed, for instance “Neu1_ch1”. The “Semantic annotations” folder contains semantic labels (CSV) describing the morphological state of cells at various stages of the cell death process. These labels indicate whether dying cells exhibit “blebbing” or “disruptive” morphology and if they elicit an immune response. Start and ending frames are provided to denote the time interval of each state. Lastly, three additional folders contain cropped microscopy movies centered around dying cells:
-
1.
Dendritics_crops
-
2.
Eosinophils_crops
-
3.
Neutrophil_crops
Each cropped movie is named after the original uncropped acquisition and specifies the cell ID of the depicted cell death, such as “Neu1_cell1”.
The dataset on immunemap31 includes all the uncropped HD5F movies, providing complementary experimental details for each acquisition. In Supplementary Information we provide a minimal example in Python to download individual files using the Zenodo API. Moreover, data, metadata, experimental details and annotations (in JSON or CSV format) can be retrieved through the application programming interfaces offered by immunemap and described at https://www.immunemap.org/index.php/doc.
Technical Validation
Imaging acquisitions
Movies in the dataset were imaged with different microscopy settings. These included different voxel sizes, sampling rates, duration, and volume (Supplementary Table 3). As these parameters can influence the overall quality of the movies, we estimated the image quality of our dataset by computing the signal-to-noise (SNR) ratio for each acquisition (Fig. 3A). Higher SNR indicates better image quality, and reportedly, values above five are considered sufficiently good to distinguish foreground objects from the background. Hence, the measured average SNR of 10 can assert the overall quality of the dataset and guarantee that their entries provide reliable and interpretable results.
Data quality and validation. (A) Distribution of the signal-to-noise ratio of the different movies. (B) Prevalence of the sampling rates used for different microscopy acquisition. (C) Distribution of cell number per field-of-view. (D) Distribution of the shortest distance between each cell. (E) Distribution of cells involved in a local cluster formed by at least three cells in close proximity. (F) Ratios between the annotated timing of cell disruption and sequence duration. A value of 0.5 indicates that the cell disrupts in the middle of the annotation. (G,H) Scatterplots showing the total track distance of annotated apoptotic-like events (G) and their duration in frames (H). (I,J) Inter-operator consensus rate during the annotation of apoptotic-like events (I) and phases (J).
The sampling rate of the movies in our dataset varied between 15, 30, 45, and 60 seconds (Fig. 3B). This aspect is relevant when evaluating dynamic processes in time lapses. Specifically, estimations of general cell motility, such as speed, straightness and cell tracking, can be significantly influenced by the selected sampling rate23. Lower sampling rates may result in the loss of critical information, especially in highly dynamic processes like cell death, where rapid and short-lived changes in the morphology of the cell may occur. To address this concern and ensure an accurate depiction of fast and transient processes, our dataset encompasses movies with high sampling rates.
Aside from the acquisition settings, the interpretability of imaging data can be influenced by the biological content and complexity of the acquisitions. A prominent parameter in this regard is cell density, which significantly impacts the analysis. In time lapses with high cell density, potential side effects, such as cluttering, cell merging, and cell splitting, may arise. These factors challenge cell detection efforts and may hinder crucial information within the data. Consequently, excessive cell density may render specific algorithms incapable of extracting meaningful insights from the movies. Therefore, we estimated pivotal parameters related to cell density and aggregation for each channel in the dataset. These parameters encompass the overall cell number, the shortest distance between cells, and a clustering coefficient that indicates the percentage of cells in close proximity to at least three other cells (Supplementary Table 3, Fig. 3C–E). These measurements underscore the variability in the dataset, with movies ranging from 20 cells to movies containing up to 300 cells and exhibiting a clustering coefficient near 1. By analyzing such diverse time-lapses, researchers can effectively benchmark their detection algorithms on increasingly complex scenarios, ensuring the robustness and applicability of their methods across a wide range of biological conditions and cell densities.
Apoptotic-like events
In the absence of specific apoptotic reporters, the annotated cells might follow a pathway different than apoptosis. For this reason, we adopt the umbrella term “apoptotic-like” to indicate all cell death events sharing morphological similarity with apoptosis. While this does not ensure the annotation of a specific death pathway, the presented catalogue is an invaluable resource for the development of analytical solutions for the classification, detection, and segmentation of cell death presenting apoptotic features in vivo. Cell death events with apoptotic-like hallmarks are inherently diverse, exhibiting variability that can affect classification and detection algorithms. One major challenge arises from the distinct morphologies of cells, leading to a class heterogeneity32,33 that makes it difficult to identify consistent patterns within the same label in supervised classification. Additionally, the variable duration of sequences can pose a challenge to algorithms aimed at classifying the cell death process over time. Finally, the residual motility of cells, caused, for instance, by drifting, can further complicate the accurate identification of cell coordinates during the death process. To address these challenges and aid algorithm development, we estimated the variability between events in our dataset. We quantified the mean timing of cell disruption (Fig. 3F), which provides insights into the typical occurrence of this critical phase. Additionally, we assessed the average displacement of cells (Fig. 3G) and the overall duration of the disruption process (Fig. 3H), offering essential information to guide algorithmic solutions for cell death classification and detection. Hence, by considering these variabilities, researchers and practitioners can better design and fine-tune their algorithms for accurate classification and detection of apoptotic-like events.
Operator annotations
To assess the inter-operator variability during the annotation process, we introduced a consensus ratio, which quantifies the proportion of annotations that achieved agreement between at least two of the three operators involved. The mean consensus ratio for annotating apoptotic-like events was 0.6, indicating a moderate level of agreement among the operators and highlighting the intrinsic challenge of visually detecting apoptotic-like events based on morphological cues (Fig. 3I). However, when annotating the specific phases (Fig. 3J), the mean consensus ratio significantly increased to 0.9, reflecting the agreement between the operators in identifying the distinct phases.
Usage Notes
The proposed dataset represents a valuable resource for researchers seeking to characterize cell death. Cell biologists can utilize the presented dataset as a reference for the death of leukocytes in vivo, enabling the generation of pilot and comparative studies. In addition, they can gain valuable insights into different properties of the process, such as morphology, duration, and the recruiting of neighboring cells. A protocol for opening, downloading, and analyzing the dataset is provided in Supplementary Information and the code repository.
The heterogeneity of the videos included in this dataset (cell type, staining, organ, experimental conditions) should be considered in comparative studies. We recommend the users to always verify the physical parameters of the videos being compared, including volume and temporal sampling rate, as these can introduce a non-linear bias on metrics to quantify cellular morphodynamics such as cell speed. Moreover, the annotation of events relied exclusively on morphological hallmarks, without the usage of reporters for specific cell death modalities. While this is a major limitation of the study, as we cannot pinpoint the specific cell death modality, the dataset provides valuable information to develop computer vision-based methods for cell death quantification.
With access to the raw data and annotations, computer scientists can address the specific tasks of cell death classification, detection, and segmentation. To create a dataset for supervised-learning tasks, users can access the raw microscopy videos and their corresponding annotations provided in Excel format. By utilizing the annotations, it is possible to locate and track the centroids of individual apoptotic-like events over time (x, y, z, t). Using this information, users can define a suitable region around each cell to generate single-cell crops and the respective labels (0 for healthy, 1 for apoptotic). It is important to remark that identifying cell death events in this dataset relied on the analysis of video sequences. Therefore, we recommend that users analyze the entire temporal sequence.
The creation of personalized training datasets offers the possibility to train custom deep learning architectures for the classification of images depicting apoptotic-likw cells based on their morphology (Fig. 4A left). To accomplish this task, a wide array of deep learning architectures is available, such as convolutional networks (CNNs)34, capsule networks35, and vision transformers36,37, among others. Accordingly, recent studies demonstrated the successful classification of a variety of cell death types in static images38,39. Additionally, the presented dataset is best suited for the temporal classification of apoptotic-like sequences (Fig. 4A right), as the latter representation is expected to encapsulate more information than single-frame images. To this end, deep learning architectures tailored for activity recognition40, such as 3DCNNs41, recurrent networks42, or optical flow-based methods43, can be employed. Recurrent architectures have demonstrated effectiveness in classifying sequences acquired in vitro1, showcasing their potential for this task. To achieve whole sequence classification, users can extract consecutive frames as cropped time-lapses, thus preserving the temporal dependencies of the events. With this information, users can train models capable of effectively analyzing the temporal dependencies and patterns exhibited by cells during cell death.
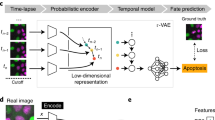
Dataset usage for deep learning applications. (A) Schematic representation of the classification task applied to single image crops depicting dying cells (left) and to apoptotic-like sequences (right). Single image crops can be classified with the aid of convolutional networks, mapping an image to a corresponding binary value labelling healthy (0) or apoptotic-like (1) cells. Similarly, sequences images can be classified with activity recognition architectures. (B) Schematic depiction of a detection routine applied to full microscopy images containing healthy and dying cells. Input images are processed with a detector that predicts the spatial coordinates of apoptotic-like cells (red bounding boxes). (C) Schematic representation of the image segmentation task applied to full microscopy images depicting dying cells. Input microscopy images are processed with an auto encoder-decoder architecture which learns to map the input into a binary mask that highlights target cells only.
Users can tackle a task of increased complexity with respect to the classification problem by detecting dying cells within uncropped fields-of-view containing multiple cells (Fig. 4B). Accordingly, detecting apoptotic-like events amid other cells poses more significant challenges due to potential cell overlaps and background clutters44. Moreover, the temporal aspect of live cell imaging introduces new challenges, including morphological changes and sudden displacements. As a result, the algorithmic detection of cell death events in live cell imaging remains unexplored. To address this aspect, researchers can use the presented dataset to explore novel approaches for detecting apoptotic-like events in time-lapse microscopy.
Finally, the presented dataset is also suitable for cell segmentation (Fig. 4C) through the utilization of encoder-decoder architectures45. These DL networks have proven effective in several bio-imaging segmentation tasks, thus holding promising applications for segmenting cell death based on their morphology. In this regard, our dataset enables 3D and 2D cell segmentation by providing the 3D binary masks of the events over time. Consequently, researchers can apply 2D or 3D segmentation using each frame of the movies as an independent training sample.
Code availability
Code repository is available at: https://github.com/AlainPulfer/Leuko-Death.git.
The “Matlab code” folder contains functions to read acquisition data and annotations. The re-distributed “ImarisReader” (https://github.com/PeterBeemiller/ImarisReader) can read time-lapse acquisition stored within HDF5 files. The “getMask” function can retrieve the 2D and 3D masks of apoptotic-like events stored in the HDF5 files. The “signal2noise” function provides a measurement for the average and peak signal-to-noise in a specified channel of the acquisition (xyzct). The function “cellCount” estimates the average number of cells over time in a channel of interest. It additionally computes a mean distance between cells that is used to derive a clustering coefficient, which in turns measures the percentage of cells involved in local clusters.
The “LeukoDeath” folder provides different python functions to read, inspect, and display acquisition data as well as annotations. A detailed Jupiter Notebook illustrates how to use the source code to load movies into 2D and 3D Python arrays, manipulate these to access individual and sequential frames, and match the movie content with the respective annotations to generate datasets for classification tasks.
References
Mobiny, A., Lu, H., Nguyen, H. V., Roysam, B. & Varadarajan, N. Automated Classification of Apoptosis in Phase Contrast Microscopy Using Capsule Network. IEEE Transactions on Medical Imaging 39(1), 1–10, https://doi.org/10.1109/TMI.2019.2918181 (2020).
Elmore, S. Apoptosis: A Review of Programmed Cell Death. Toxicologic Pathology https://doi.org/10.1080/01926230701320337 (2007a).
Opferman, J. T. Apoptosis in the development of the immune system. In Cell Death and Differentiation (Vol. 15, Issue 2), https://doi.org/10.1038/sj.cdd.4402182 (2008).
Rathmell, J. C. & Thompson, C. B. Pathways of apoptosis in lymphocyte development, homeostasis, and disease. In Cell (Vol. 109, Issue 2 SUPPL. 1), https://doi.org/10.1016/S0092-8674(02)00704-3 (2002).
Elmore, S. Apoptosis: A Review of Programmed Cell Death. In Toxicologic Pathology (Vol. 35, Issue 4, pp. 495–516), https://doi.org/10.1080/01926230701320337 (2007b).
Hotchkiss, R. S., Strasser, A., McDunn, J. E. & Swanson, P. E. Mechanisms of disease: Cell death. New England Journal of Medicine 361(16), 1570–1583, https://doi.org/10.1056/NEJMra0901217 (2009).
Yang, Y., Jiang, G., Zhang, P. & Fan, J. Programmed cell death and its role in inflammation. In Military Medical Research (Vol. 2, Issue 1), https://doi.org/10.1186/s40779-015-0039-0 (2015).
Chatziandreou, N. et al. Macrophage Death following Influenza Vaccination Initiates the Inflammatory Response that Promotes Dendritic Cell Function in the Draining Lymph Node. Cell Reports, 18, https://doi.org/10.1016/j.celrep.2017.02.026 (2017).
Gonzalez, S. F. et al. Trafficking of B cell antigen in lymph nodes. Annual Review of Immunology 29, 215–233, https://doi.org/10.1146/annurev-immunol-031210-101255 (2011).
Pérez-Figueroa, E., Álvarez-Carrasco, P., Ortega, E. & Maldonado-Bernal, C. Neutrophils: Many Ways to Die. In Frontiers in Immunology (Vol. 12), https://doi.org/10.3389/fimmu.2021.631821 (2021).
Tang, D., Kang, R., Berghe, T. V, Vandenabeele, P. & Kroemer, G. The molecular machinery of regulated cell death. In Cell Research (Vol. 29, Issue 5), https://doi.org/10.1038/s41422-019-0164-5 (2019).
Saraste, A.& Pulkki, K. Morphologic and biochemical hallmarks of apoptosis. In Cardiovascular Research (Vol. 45, Issue 3), https://doi.org/10.1016/S0008-6363(99)00384-3 (2000).
Doran, A. C., Yurdagul, A. & Tabas, I. Efferocytosis in health and disease. In Nature Reviews Immunology (Vol. 20, Issue 4), https://doi.org/10.1038/s41577-019-0240-6 (2020).
Isherwood, B. et al. Live cell in vitro and in vivo imaging applications: Accelerating drug discovery. In Pharmaceutics (Vol. 3, Issue 2). https://doi.org/10.3390/pharmaceutics3020141 (2011).
Kramer, N. et al. In vitro cell migration and invasion assays. In Mutation Research - Reviews in Mutation Research (Vol. 752, Issue 1), https://doi.org/10.1016/j.mrrev.2012.08.001 (2013).
Pijuan, J. et al. In vitro cell migration, invasion, and adhesion assays: From cell imaging to data analysis. Frontiers in Cell and Developmental Biology, 7(JUN). https://doi.org/10.3389/fcell.2019.00107 (2019).
Grootveld, A. K. et al. Apoptotic cell fragments locally activate tingible body macrophages in the germinal center. Cell, 186(6). https://doi.org/10.1016/j.cell.2023.02.004 (2023).
Mayer, C. T. et al. The microanatomic segregation of selection by apoptosis in the germinal center. Science, 358(6360), https://doi.org/10.1126/science.aao2602 (2017).
Mesa, K. R. et al. Niche-induced cell death and epithelial phagocytosis regulate hair follicle stem cell pool. Nature, 522(7554), https://doi.org/10.1038/nature14306 (2015).
Secklehner, J., Lo Celso, C. & Carlin, L. M. Intravital microscopy in historic and contemporary immunology. In Immunology and Cell Biology (Vol. 95, Issue 6), https://doi.org/10.1038/icb.2017.25 (2017).
Sumen, C., Mempel, T. R., Mazo, I. B. & Von Andrian, U. H. Intravital microscopy: Visualizing immunity in context. In Immunity (Vol. 21, Issue 3), https://doi.org/10.1016/j.immuni.2004.08.006 (2004).
Kranich, J. et al. In vivo identification of apoptotic and extracellular vesicle-bound live cells using image-based deep learning. Journal of Extracellular Vesicles, 9(1), https://doi.org/10.1080/20013078.2020.1792683 (2020).
Beltman, J. B. et al. Analysing immune cell migration. Nature Reviews. Immunology 9(11), 789–798, https://doi.org/10.1038/nri2638 (2009).
Pizzagalli, D. U. et al. Data Descriptor: Leukocyte Tracking Database, a collection of immune cell tracks from intravital 2-photon microscopy videos. Scientific Data 5, 1–13, https://doi.org/10.1038/sdata.2018.129 (2018).
Sinaci, A. A. et al. From Raw Data to FAIR Data: The FAIRification Workflow for Health Research. Methods of Information in Medicine, 59(6), https://doi.org/10.1055/s-0040-1713684 (2020).
Jacobsen, A. et al. A generic workflow for the data fairification process. Data Intelligence, 2(1–2), https://doi.org/10.1162/dint_a_00028 (2020).
Adadi, A. A survey on data‐efficient algorithms in big data era. Journal of Big Data, 8(1), https://doi.org/10.1186/s40537-021-00419-9 (2021).
Pulfer, A. et al. An in vivo microscopy dataset capturing leukocyte cell death. Zenodo https://doi.org/10.5281/zenodo.14551288 (2025).
Masedunskas, A. et al. Intravital microscopy: A practical guide on imaging intracellular structures in live animals. BioArchitecture, 2(5), https://doi.org/10.4161/bioa.21758 (2012).
Di Pilato, M. et al. Neutrophil subtypes shape HIV-specific CD8 T-cell responses after vaccinia virus infection. Npj Vaccines, 6(1), https://doi.org/10.1038/s41541-021-00314-7 (2021).
Pizzagalli, D. U. et al. Systematic analysis of immune cell motility leveraging Immunemap, an open intravital microscopy atlas. https://doi.org/10.1101/2024.12.02.626343 (2024).
Pang, G., Shen, C., Cao, L. & Van Den Hengel, A. Deep Learning for Anomaly Detection: A Review. In ACM Computing Surveys (Vol. 54, Issue 2), https://doi.org/10.1145/3439950 (2021).
Van Valen, D. A. et al. Deep Learning Automates the Quantitative Analysis of Individual Cells in Live-Cell Imaging Experiments. PLoS Computational Biology, 12(11), https://doi.org/10.1371/journal.pcbi.1005177 (2016).
Bhatt, D. et al. Cnn variants for computer vision: History, architecture, application, challenges and future scope. In Electronics (Switzerland) (Vol. 10, Issue 20), https://doi.org/10.3390/electronics10202470 (2021).
Kwabena Patrick, M. et al. Capsule Networks – A survey. In Journal of King Saud University - Computer and Information Sciences (Vol. 34, Issue 1), https://doi.org/10.1016/j.jksuci.2019.09.014 (2022).
Han, K. et al. A Survey on Vision Transformer. IEEE Transactions on Pattern Analysis and Machine Intelligence, 45(1), https://doi.org/10.1109/TPAMI.2022.3152247 (2023).
He, K. et al. Transformers in medical image analysis. In Intelligent Medicine (Vol. 3, Issue 1), https://doi.org/10.1016/j.imed.2022.07.002 (2023).
Jin, J. et al. Machine Learning Classifies Ferroptosis and Apoptosis Cell Death Modalities with TfR1 Immunostaining. ACS Chemical Biology, 17(3). https://doi.org/10.1021/acschembio.1c00953 (2022).
Verduijn, J., Van der Meeren, L., Krysko, D. V. & Skirtach, A. G. Deep learning with digital holographic microscopy discriminates apoptosis and necroptosis. Cell Death Discovery, 7(1), https://doi.org/10.1038/s41420-021-00616-8 (2021).
Yeole, C., Singh, H., Waykole, H. & Deshpande, A. Deep Neural Network Approaches for Video Based Human Activity Recognition. International Journal of Innovative Science and Research Technology, 6(6) (2021).
Vrskova, R., Hudec, R., Kamencay, P. & Sykora, P. Human Activity Classification Using the 3DCNN Architecture. Applied Sciences (Switzerland), 12(2), https://doi.org/10.3390/app12020931 (2022).
Mohd Noor, M. H., Tan, S. Y. & Ab Wahab, M. N. Deep Temporal Conv-LSTM for Activity Recognition. Neural Processing Letters, 54(5), https://doi.org/10.1007/s11063-022-10799-5 (2022).
Weinzaepfel, P., Revaud, J., Harchaoui, Z. & Schmid, C. DeepFlow: Large displacement optical flow with deep matching. Proceedings of the IEEE International Conference on Computer Vision. https://doi.org/10.1109/ICCV.2013.175 (2013).
Vrigkas, M., Nikou, C. & Kakadiaris, I. A. A review of human activity recognition methods. In Frontiers Robotics AI (Vol. 2, Issue NOV), https://doi.org/10.3389/frobt.2015.00028 (2015).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. Lecture Notes in Computer Science (Including Subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). https://doi.org/10.1007/978-3-319-24574-4_28 (2015).
Acknowledgements
Swiss National Science Foundation grants 176124 (A.P., D.U.P., M.P., T.V., S.G.), 310030_163336 (P.A., M.T.), SystemsX.ch 2013124 (D.U.P., R.K., S.G.), Biolink 189699 (S.G., D.U.P., P.C.B., E.P.), swissuniversities RE2VITAL grant (S.F.G., R.K., D.U.P., A.P., E.P.), USI FIR (D.U.P.).
Author information
Authors and Affiliations
Contributions
Conceptualization: A.P., D.U.P., S.G.; Methodology: A.P., D.U.P.; Experiments: M.P., N.G., T.V., M.D.P., P.A., D.U.P., S.G.; Data annotation: A.P., D.U.P., P.C.B., T.V., P.A.; Data analysis and visualization: A.P.; Figures: A.P., D.U.P.; Writing original draft: A.P., P.C.B., D.U.P., S.G.; Revision: A.P., D.U.P., E.P., S.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pulfer, A., Pizzagalli, D.U., Segura, M.P. et al. An in vivo microscopy dataset for the characterization of leukocyte death. Sci Data 12, 593 (2025). https://doi.org/10.1038/s41597-025-04632-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-04632-6